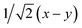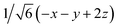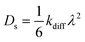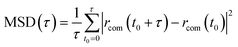Open Access Article
Open Access ArticleA consistent dynamics view on nanoporous catalysts in action across length and time scales
Veronique Van Speybroeck *,
Massimo Bocus
*,
Massimo Bocus ,
Wei Chen
,
Wei Chen and
Pieter Cnudde
and
Pieter Cnudde
Center for Molecular Modeling, Ghent University, Technologiepark 46, 9052 Zwijnaarde, Belgium. E-mail: veronique.vanspeybroeck@ugent.be
First published on 17th March 2026
Abstract
This paper demonstrates the need for a consistent dynamics view of all processes from the nano- to mesoscale with quantum accuracy to design industrial catalysts. This stems from the observation that when a feed of molecules passes through a crystalline nanoporous framework, the following events occur: molecules enter the crystal, diffuse to active sites where they adsorb, desorb, or react, and finally products exit. These events span vastly different time and length scales—from picoseconds to hours and nanometers to micrometers. Length and time phenomena are entangled—the dynamics is altered by modifications at the length scale—leading to spatiotemporal phenomena. Current modeling practices treat reactions and diffusion inconsistently, often combining quantum methods for reactions with classical force fields for diffusion, and frequently neglect true dynamics under operating conditions. Industrial catalysts are highly sensitive to process conditions; some active sites form only at elevated temperatures, and reactive intermediates can change nature under these conditions. First-principles molecular dynamics captures these effects but becomes prohibitively expensive when relying on quantum methods like Density Functional Theory for force evaluations. In this paper, we demonstrate—through the study of key intermediates in CO2-to-hydrocarbon conversion within small-pore zeolites—that diffusion must also be treated at the quantum level, as molecules can interact with active sites and even react while diffusing. Diffusion may become strongly hindered and compete with reactive events. In later case studies, we present proof-of-concept results for deriving diffusion coefficients in small-pore zeolites using machine learning potentials (MLPs) trained on quantum data. For benzene methylation in large-pore zeolites, we develop reactive MLPs to construct multidimensional free-energy surfaces, revealing competing pathways. Overall, the various cases underscore the need for a consistent dynamic description with quantum accuracy across scales. MLPs offer a promising route to obtain kinetic and transport properties from the nano- to mesoscale at reduced cost, however their integration in overall multiscale models and reaction–diffusion frameworks will be necessary to reach industrially relevant scales.
1. Introduction
The performance of heterogeneous catalysts is a prime example where an intricate interplay of various processes—adsorption, reaction, diffusion, materials restructuring—spanning a wide range of length and time scales dictates the catalytic performance. The catalytic function is highly dependent on the operating conditions such as temperature, moisture, and the presence of water.1,2 A prime example of this complexity is found in C1 catalysis,3 where molecules like CO2 and CH3OH are converted into high-value olefins and other chemical building blocks. Methanol-to-olefin reactions typically employ Brønsted acidic zeolites,4 often modified post-synthetically, while CO2 conversion relies on multifunctional catalysts combining metallic and zeolitic components.5,6 The desired catalytic function only emerges at operating conditions, in this sense catalysts are highly dynamical, where active sites are formed within the reactive environment, and the catalyst restructures at certain conditions.2,7 It is now well established that it is essential to capture these dynamics when modelling industrial heterogeneous catalysts.1,8 Within this discussion paper, focus is set on nanoporous heterogeneous catalysts namely zeolites having a Brønsted catalytic function. Evidence will be given that we need a consistent dynamics view of all processes – adsorption, reaction, diffusion, surface barriers – from the nano- to the mesoscale with quantum accuracy to design industrial catalysts.To illustrate the complexity and the events taking place during a catalytic process, consider the various steps taking place when a feed of molecules is sent over a crystalline catalytic particle. In an experiment, a feed of molecules flows over a nanoporous material, initiating a cascade of steps as schematically shown in Fig. 1a.9 The molecules first enter the crystal, sometimes after interacting with external active sites,10 and then diffuse through the lattice in search of active sites where adsorption and reaction can occur. Intracrystalline diffusion may be hindered by structural constraints such as narrow windows or by the presence of bulky co-adsorbed species.11 Once a reaction takes place, the products desorb from the active site and migrate toward the exit before leaving the material. These events span an enormous range of time scales—from picoseconds for molecular vibrations to seconds or even hours for activated processes. In principle every event is also characterized by its own characteristic time scales, indicated with τ in the figure.
 | ||
| Fig. 1 (a) Schematic representation of various steps taking place during a typical catalytic trajectory, where molecules are sent over a heterogeneous catalyst. Molecules enter the crystal and diffuse through the pores to arrive at the active site to adsorb/react. Diffusion in/out the crystal can be hindered due to external active sites, other molecules or narrow windows in the crystal. Every event is also characterized by its own characteristic time scales, indicated with τ. (b) Brief illustration of all individual events and the various kinetic and transport properties that may be derived from first principles simulations. (a) is adapted from ref. 9, copyright 2026, the Annual Review of Physical Chemistry. | ||
In today’s modelling practice, each of the events is often treated in an isolated way and with tailored methodologies, i.e. reactions obviously need to be treated with quantum mechanical (QM) methods, however this can be done using static or dynamic methodologies.1,12 Diffusion is most commonly treated with classical force fields as these processes take place on longer length and time scales.13 There is a growing evidence that reaction, diffusion, and adsorption are dynamically entangled under realistic, time-dependent operating conditions.1,2 Modelling the full catalytic progression via first-principles QM based approaches that fully account for the dynamics is to date not feasible. Also from the experimental point of view, it is still not possible to make a “molecular movie” with the most advanced spectroscopic or imaging techniques and to follow on the fly the breaking and formation of chemical bonds with atomic scale resolution.14–18 To evolve towards predictive and rational design, it is essential to devise methods that capture the dynamics of catalysts in action across length and timescales. New methods are needed that enable the connection between nanoscale chemical rearrangements dictated by electronic restructuring of bonds to mesoscale transport phenomena. With the rise of machine learning methods,19–22 new avenues are opened to integrate effects from the nano- to the mesoscale. In this Faraday Discussions contribution, we argue the need for obtaining a consistent dynamics view from the nano- to the mesoscale with quantum accuracy to design industrial catalysts.23 Machine learning methods are central to this vision but are on their own not sufficient to reach this ambition. More in particular, one can now derive Machine Learning Potentials (MLPs) to obtain the energies and forces with so-called quantum accuracy, i.e. the same accuracy as the underlying QM data on which they are trained, however at a much lower computational cost compared to the brute force QM calculations.23,24 This allows simulation of timescales and length scales of at least three orders of magnitude larger than when pristine density functional theory methods are used. Yet today, MLPs are not fully part of a current catalysis workflow with a high-dimensional free energy surface where many competitive reactions taking place simultaneously. Major methodological developments are necessary to couple MLPs with kinetic models to describe complex dynamical phenomena in the broad range of length and time scales relevant for heterogeneous catalysts.
Within this Faraday Discussions article, we describe first results on how MLPs can be instrumental to obtain kinetics for all events from the local to the crystal particle level with quantum accuracy. Fig. 1b shows the individual events taking place during a successful catalytic trajectory and the kinetic and transport properties needed to set up integrated reaction diffusion models based on underlying kinetics and characteristic time scales obtained with QM accuracy. Within this figure one observes reactions, which are local events taking place on the (sub)nanometer level and are typically described by QM methods. For reactions one needs to be able to calculate the reaction rates. Still today, reaction kinetics of activated processes are most often obtained using a static approach, based on a few points on the Potential Energy Surface (PES) and application of Transition State Theory (TST).25,26 Such an approach accounting only for local information around the (meta)stable states gives a too approximate view of complex transformations at operating conditions and does not capture broad transition state regions, competitive pathways, and configurational mobility at higher temperatures.27–31 Enhanced sampling MD (ESMD) simulations are now extensively used to account for operating conditions, however in combination with QM evaluation of energies these are very expensive.1 Additionally they rely mostly on collective variables (CVs) able to distinguish between various minima and their definition is far from trivial.32–36 Most importantly, current ESMD simulations determine the kinetics primarily within the limits of TST neglecting barrier recrossings.1,25,37 With the rise of MLP based approaches, it is expected that dynamically correct reaction rates will be possible to be determined in a more routine way, also for complex catalytic transformations. Within this Faraday Discussions article we show first proof of concepts results where we derived a MLP for zeolite catalyzed reactions, which enables sampling of competitive pathways in higher dimensionalities at a feasible computational cost.
Intracrystalline diffusion is an essential element of the catalytic trajectory but takes place on longer length scales, which is one of the reasons why it is usually described by classical force fields. One major drawback of such description is that specific host–guest interactions are not accounted for and also that potential reactive events taking place during a transport process are not captured.38–40 Diffusion within nanoporous catalysts can be hindered or non-hindered, i.e. diffusion itself can become an activated event and becomes competitive with reactive events. To characterize diffusion one typically needs to calculate diffusion coefficients, either collective or self-diffusion coefficients, that quantify how fast molecules spread out in a medium due to random thermal motion or a concentration gradient.41 Diffusion coefficients can be calculated by various approaches. Self-diffusion coefficients are typically calculated from the mean square displacement (MSD) of adsorbate molecules from very long MD runs.13,42 This approach is feasible when the molecules are not severely hindered during transport. For hindered diffusion, the diffusion itself becomes activated and it is necessary to rely on methods for activated processes.38,39 The interested reader is referred to dedicated reviews for more information on the topic.1,13,41 Comparison between experimental and theoretical diffusion coefficients needs to be done with extreme care as was pointed out nicely by Kärger et al.43,44 Apart from direct comparison of diffusion coefficients with experiment, one can also obtain information on residence times of probe molecules in zeolite catalysts using pulse-response temporal analysis of products (TAP) methodology.45,46 However various effects contribute to the measured TAP results, such as surface barriers, intracrystalline diffusion, adsorption at the active site and finally kinetics.47–49 From an experimental point of view, it is very difficult to disentangle all these contributions.38,50 With the development of consistent dynamical models for all events from the nano- to the mesoscale – as discussed in this paper – it might become possible in the future to build bottom-up models starting from the atomic scale and determine the various components contributing to the TAP data. Herein, we show first proof of concept results on how MLPs can be instrumental to simulate on longer length and time scales diffusion and how self-diffusion coefficients can be estimated from long molecular dynamics approaches with QM accuracy.
Fig. 1b also highlights events related to adsorption and kinetics at the external surface of zeolite catalysts. In principle it is necessary to also obtain kinetics of surface permeation at QM level of theory on realistic external surfaces. Experimental studies clearly indicate the presence of surface barriers, often attributed to pore blocking or amorphous outer layers, yet their precise molecular origin remains unknown.51 Obtaining spatially resolved information on the structure and nature of the external surface under operating conditions is currently impossible, leaving the location of active sites and their interaction with key intermediates unresolved.51 Also within this area machine learning potentials could be instrumental to simulate surface barrier phenomena, however further progress will also depend on construction of realistic surface models, however this discussion is beyond the scope of this paper.
The three case studies presented in this paper, give evidence for the claim that we need a consistent dynamics view from the nano- to the mesoscale with quantum accuracy to design industrial catalysts and show how MLPs can open new possibilities to model complex catalytic events with QM accuracy.
The first case study examines the diffusion of key intermediates—CO, formaldehyde, and ketene—through the small-pore zeolite H-SSZ-13, which is a crucial step in the CO2-to-hydrocarbons process. In this material, diffusion is strongly hindered, and in some cases, species like ketene do not simply diffuse but can also react while passing through the eight-ring windows.40 This demonstrates that diffusion must be described using quantum mechanical methods capable of capturing explicit interactions with active sites and possible transformations. However, such simulations require long timescales and extended length scales, making standard QM approaches impractical for complex catalytic systems.
The second case study provides a proof of concept for overcoming these limitations by training a machine learning potential (MLP) to model the diffusion of C2 olefins and paraffins in H-SSZ-13. When properly trained, MLPs enable the calculation of converged free-energy profiles between cages and allow the derivation of diffusion coefficients.
The third case study demonstrates the potential of machine learning potentials (MLPs) for exploring rare events in zeolite catalysis under realistic operating conditions. We focus on the methanol-to-olefins process and revisit the methylation of benzene by methanol in a large-pore H-SSZ-24 zeolite.30,31 This reaction can proceed via two competing mechanisms—a direct, concerted methyl transfer and a stepwise route through a surface-bound methoxide—both of which may coexist depending on operating conditions.52 In earlier work based on first-principles molecular dynamics, we showed that these pathways are accessible and that their relative preference is strongly influenced by the local environment, including methanol clustering and framework flexibility, resulting in a complex free-energy landscape.31 This seminal study highlighted the importance of entropic contributions and the complexity of the free-energy surface at elevated temperatures, revealing mechanistic competition that static models cannot capture. While these insights were groundbreaking, the simulations were computationally demanding and not broadly applicable to zeolite catalysis. MLPs hold the promise to make these insights much more accessible, provided the MLPs are trained in a correct way to capture the essential chemistry. Within this paper, we revisit the benzene methylation in H-SSZ-24 by deriving an MLP using active learning. We show how it becomes possible to simulate extensively longer within reasonable computational resources and can explore the free energy surface in a higher dimensional space.
These case studies illustrate how machine learning potentials (MLPs) enable dynamic information with quantum-level accuracy for individual steps of a catalytic trajectory. However, achieving a fully consistent dynamic view of an entire catalytic process—including all relevant time scales—requires more than MLPs alone. With MLP based methods, it is not possible to reach the micrometer length scales or the seconds-long time scales needed to mimic real catalytic experiments. To bridge this gap, consistent kinetic and transport data must be integrated into reaction–diffusion or overall kinetic models at the level of the entire catalyst particle. In the final conclusion and perspective section, we reflect on strategies to connect detailed insights from individual events within a catalytic cycle to a holistic description of the catalyst particle and its behavior under operating conditions. Ultimately, the long-term vision is to model the time evolution of real catalytic systems with complex feedstocks under realistic conditions, and we outline future avenues to move towards this goal.
2. Methods
Within this section various methodologies are described necessary to model each of the individual case studies. Various methodologies will be used varying from QM based simulations, enhanced sampling methodologies, derivation of Machine Learning Potentials (MLPs) both for reactive and non-reactive events.2.1 Methodology to describe diffusion of three unsaturated oxygenates with first principle molecular dynamics simulations
Diffusion of CO, formaldehyde, ketene and small hydrocarbons in H-SSZ-13 can be seen as a hopping process where the guest species mainly resides in the spacious cages and only occasionally jumps to a neighboring cage, through the connecting 8-ring window, as shown in Fig. 2a. As the probability to observe diffusion hops in regular MD simulations is quite low, enhanced sampling techniques are required to properly sample the entire diffusion path. To sample the rare events, umbrella sampling (US) simulations are carried out at elevated temperature representative for the process at hand. This procedure allows construction of free energy profiles for the diffusion event between two neighboring cages, A and B. A collective variable is defined as the orthogonal projection of the center-of-mass of the diffusing guest species onto the normal of the 8-ring window plane, thus unambiguously representing the diffusion coordinate, ξ. When ξ < 0, the guest hydrocarbon resides in cage A, around ξ = 0, it is traveling through the 8-ring window and for ξ > 0, it resides in cage B. First-principles molecular dynamics simulations are carried out in the NVT ensemble at a temperature of 673 K, which is controlled by a chain of five Nosé–Hoover thermostats.53 The integration time step was set to 0.5 fs. The revised Perdew–Burke–Ernzerhof functional54,55 with additional Grimme D3 dispersion corrections,56 that is, the revPBE-D3 functional, was used together with a triplet-zeta (ζ) valence polarized (TZVP) basis set57 and Goedecker–Teter–Hutter (GTH) pseudopotentials.58 During the self-consistent field procedure, the plane wave energy cutoff and the SCF convergence criterion were set to 360 Ry and 10−8 a.u. The collective variable range [−6.0 Å, 6.0 Å] is divided into a set of 25 equidistant umbrella sampling windows with equal spacing of 0.5 Å, for each of which an independent MD simulation of 30 ps is carried out. The MD sampling is restricted to each window individually by introducing a harmonic bias potential:
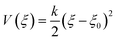 | (1) |
 | ||
| Fig. 2 Collective variables of three typical unsaturated oxygenates’ diffusion and reaction in SSZ-13. (a) CV1 (ξ) to describe the ketene diffusion, (b) CV2 to describe the protonation of CO with BAS, (c) CV3 to describe the protonation of formaldehyde with BAS, (d) CV4 to describe the protonation of ketene with BAS. (a) and (d) are reproduced under the terms of the CC BY 4.0 license from ref. 40. Copyright 2026 Elsevier. | ||
In the presence of BAS, CO, ketene, and formaldehyde may spontaneously react to new species along the diffusion pathway, i.e., by protonation into either a cationic intermediate or surface-bound species (Fig. 2b–d). Clearly, this interconversion between adsorbed intermediates presents an orthogonal degree of freedom and cannot be described solely by the diffusion coordinate (CV1). To accurately distinguish between neutral, cationic intermediates, and surface-bound species along the diffusion coordinate, a second collective variable is required to track these species. By considering both a diffusion coordinate as the first CV and a reaction coordinate as the second CV, the entangled effects of diffusion and reaction can be clearly revealed. Notably, enhanced sampling was not performed along the second CV, and the resultant 2D free energy surface can simply be regarded as a two-dimensional extension of the classical 1D free energy profile, unfolded along a second CV. Therefore, the obtained 1D free energy profile along the diffusion coordinate (CV1) is de-projected into a 2D free energy surface along a second coordinate describing the reaction using the deprojection module implemented in the ThermoLIB software.59
In detail, CO may be protonated to a formal cation (HCO+) and further form the surface formate (Zeo–HCO) described by CV2 in Fig. 2b; HCHO is able to be protonated to a hydroxymethyl cation (CH2OH+) and further form a surface hydroxymethyl (Zeo–CH2OH)60 described by CV3 in Fig. 2c; ketene can be easily protonated to an acylium ion and further form surface acetate described by CV4 in Fig. 2d. To ensure each of these states are properly visited, thorough sampling is required, resulting in the addition of the following umbrella windows: (i) 25 windows (between −6 Å ≤ ξ ≤ 6 Å) to sample the diffusion of neutral CO, formaldehyde, and ketene; (ii) 13 windows in proximity to the acid site (between −3 Å ≤ ξ ≤ 3 Å) in which the cationic forms are visited, and (iii) 7 windows in close proximity to the acid site (between −1.5 Å ≤ ξ ≤ 1.5 Å) in which surface-bound species are sampled.
2.2 Methodology to describe diffusion with machine learning potentials in confined spaces
In order to properly describe the specific interactions between the diffusing guest olefins and paraffins with the zeolite wall and the acid sites, a quantum level description is required. However, performing MD simulations at a DFT level of theory is computationally expensive and simulation times are typically restricted to the picosecond order. Nevertheless, as diffusion takes place on a longer length and timescale, much longer simulation times are necessary to ensure a sufficient sampling of the entire phase space of the diffusing molecules which typically exhibit a high degree of rotational and translation freedom. Therefore, in order to obtain reliable diffusion barriers, a machine learning potential (MLP) is trained for the diffusion of C2 olefins and paraffins in H-SSZ-13. The potential is trained with the MACE architecture61 based on an active learning scheme using the in-house developed Psiflow package.62 The active learning loop consists consecutively of (i) a sampling step (with the previously trained MLP) to generate new configurations for the dataset using i-PI,63,64 (ii) a DFT force/energy evaluation on the new configurations at the revPBE-D3 level of theory54–56 with a TZVP-MOLOPT basis set65 using CP2K66 and (iii) a training step of the expanded dataset with the MACE architecture (2 interaction layers, 128 features, cut-off radius of 6 Å). To ensure the potential is capable of handling a broad variety of cases, a versatile set of configurations are incorporated in the dataset, encompassing ethene and ethane diffusion in H-SSZ-13 with varying acid site densities and locations. The potential is trained up to an RMSE well below 1.0 meV per atom and 40 meV Å−1 for the energies and forces respectively and validated by benchmarking against the results of DFT simulations. Ultimately, the MLP is used to perform umbrella sampling (US) simulations for a total simulation time of 1 ns per window, extending well beyond the limits of DFT simulations, and free energy profiles for the diffusion event between two neighboring cages are constructed. The procedure of the simulations and the definition of the diffusion CV are similar to those described in Section 2.1. Here, the diffusion coordinate is divided into 53 equally spaced windows in the range [−6.5 Å, 6.5 Å] for each of which a biased MD simulation is carried out with the i-PI package,63 interfaced with PLUMED,67 in the NVT ensemble at 600 K, controlled by a Langevin thermostat with time constant 100 fs.2.3 Methodology to describe benzene methylation reaction with machine learning potentials
(1) A concerted methylation, in which a methyl group is transferred from one of the methanol molecules directly to benzene (I → III in Fig. 3a).
(2) A step-wise methylation, where a methyl group of one of the methanol molecules is first transferred to the framework (I → II in Fig. 3a), forming a surface-methoxide-species (SMS). The SMS acts subsequently as a methylating agent for benzene to form a toluenium ion (II → III in Fig. 3a).
(3) The two methanol molecules can condense forming dimethyl ether (I or II to IV in Fig. 3a). The latter can then act as a methylating agent either directly with benzene or with the SMS.
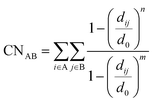
To be able to describe (at least qualitatively) all these processes, we selected as collective variables (CVs) 2 linear combinations of 3 coordination numbers (CNs). The general formula for the CNs is given below:
| State | max[CN(Cme,i, Car)] | min[CN(Cme,i, Ome,i)] | max[CN(Cme,i, Ozeo)] | |
|---|---|---|---|---|
| I | Reactant: 2 methanol molecules + benzene | 0 | 1 | 0 |
| II | Surface methoxide species, water, methanol, benzene | 0 | 0 | 1 |
| III | Toluenium ion, methanol | 1 | 0 | 0 |
| IV | Benzene, dimethyl ether, water | 0 | 0 | 0 |
With this definition of collective variables, the three intermediates I, II, and III can be seen as three vertices of an equilateral triangle in the space defined by these three CNs, located at 1 on the x, y, or z axis, as schematically shown in Fig. 3b. Intermediate IV, involving dimethyl ether, will be at the origin as all CNs will be about 0. Performing enhanced sampling simulations in three dimensions is normally associated with major convergence issues and, therefore, we opted in the first instance to reduce the dimensionality of the problem by defining two new collective variables CV1 and CV2 defined by the equations shown in Fig. 3b. The choice can be understood as a projection of the 3-dimensional space on the plane passing through (1, 0, 0), (0, 1, 0), and (0, 0, 1), in which the origin is projected on the origin of the new system (blue axes in Fig. 3b). The 2-dimensional CV space defined in this way was used both for the active learning procedure and for subsequent production runs. However once we derived a converged MLP, we also performed enhanced simulations in terms of the three collective variables, to obtain a full picture of all possible intermediates and transition state regions. We also note that a set of half-quadratic walls was placed starting from 1.15 Å (with a force constant of 1000 kJ mol−1 Å−2) on all the C–H bonds of benzene and the CH3 groups of methanol, to prevent unwanted proton transfer reactions.
We initially attempted to perform well-tempered metadynamics based on the MACE-mp0 foundational model to explore the whole reaction phase space.68 However, we observed that proton-transfer reactions were excessively favored (e.g., leading to methane formation), resulting in chemically irrelevant pathways. For instance, methyl cations extracted a proton from two other methanol molecules to form methane and formaldehyde, or from benzene, producing compounds that are not chemically feasible. Therefore, we decided to train a MACE model from scratch via active learning within Psiflow. The generation of new structures required four separated loops (steps 1–4 in Fig. 4) to achieve a satisfactory phase space coverage.
First, we began by performing 25 ps of well-tempered metadynamics with MACE-mp0 (ref. 68) at 800 K with 240 independent walkers (hills height: 5 kJ mol−1; hills sigma: 0.05 along both CVs; bias factor: 15; deposition pace: 250 steps) to generate an initial dataset of 1194 structures. Unless differently specified, the timestep for this and all subsequent simulations was set to 0.5 fs and temperature control was achieved by Langevin dynamics, with a time constant of 100 fs. The bias was applied by PLUMED67 and i-PI63,64 was used to drive the dynamics. A structure was saved every 5 ps. The initial dataset generated in this way was evaluated with our reference level of theory. All first-principles evaluations were performed with CP2K 2024.1 (ref. 66) at a PBE-D3(BJ)/MOLOPT-TZV2P level of theory (note that the double polarization functions were not used for Si and Al being electron-deficient).55,56,65 The cutoff for the secondary plane-wave basis set used by CP2K was set to 900 Ry to achieve well-converged forces. GTH pseudopotentials58 were used to smooth the electron density in proximity of the atomic nuclei.
The dataset and model obtained from this first set of simulations with MACE-mp0 were used as input for the actual active learning, which was organized in four steps (Fig. 4):
• Step 1: the well-tempered metadynamics simulations for the 240 independent walkers were continued for 5 active learning loops, propagating each time the walkers for 10 ps and collecting the final structures from the trajectories for the subsequent evaluation, dataset expansion and new model training. A uniform temperature ramp between 300 and 1000 K was applied to the simulations to expand the explored phase space boundaries.
• Step 2: since many walkers from Step 1 entered uninteresting regions of phase space (e.g., methane formation), we restarted all walkers from the reasonable, manually built initial structure of the reactants and added 3 extra active learning loops at 800 K, propagating each walker for 25 ps for each loop. The well-tempered metadynamics bias was also reinitialized.
• Step 3: the transition state region is subjected to limited sampling in metadynamics. Therefore, we constructed some additional structures representing the transition states and minima of interest. We defined a grid of 2-dimensional quadratic umbrella biases covering the CV-regions of interest (that is, the triangle defined by the three minima in Fig. 3). Each umbrella was initialized with the structure closest to its center in CV space. In the first online learning loop, we propagated the walkers for 2500 steps at 10 K, with a timestep of 0.05 fs. This mimics a geometry optimization, and allows the walkers to relax if they start far from an umbrella center. Then, the temperature was raised to 300 K, the timestep set back to its default value of 0.5 fs, and each walker propagated for 5 ps per loop. Four additional loops were performed in this way, simultaneously raising the temperature to 1000 K.
• Step 4: most of the simulations tend to sample TSs involving dimethyl ether. To augment the sampling of TSs involving methanol, we hand-picked a set of umbrella centers sampling the “direct” paths between the minima (i.e., not passing by the center of the triangle in Fig. 3), performing 3 additional on-line learning loops at 800 K with a propagation time of 10 ps per loop.
The final model from this procedure, trained on the 6169 collected structures (randomly split 90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 into training and validation sets), achieved a root mean squared error of 1.2 meV per atom and 1.1 meV per atom on the energies of the training and validation sets, respectively, and a root mean squared error of 39.8 and 42.0 meV Å−1 on the forces of the training and validation sets, respectively. This final model was then used for all subsequent production runs. Because of the qualitative nature of the presented results, no additional checks on the model accuracy were performed, i.e. generating independent test sets to further validate the MLP. Evidently, we visually inspected the generated trajectories to ensure (i) that they were stable and (ii) that no obviously new regions of phase space were explored.
10 into training and validation sets), achieved a root mean squared error of 1.2 meV per atom and 1.1 meV per atom on the energies of the training and validation sets, respectively, and a root mean squared error of 39.8 and 42.0 meV Å−1 on the forces of the training and validation sets, respectively. This final model was then used for all subsequent production runs. Because of the qualitative nature of the presented results, no additional checks on the model accuracy were performed, i.e. generating independent test sets to further validate the MLP. Evidently, we visually inspected the generated trajectories to ensure (i) that they were stable and (ii) that no obviously new regions of phase space were explored.
3. Results and discussion
3.1 Evidence for entanglement between reaction and diffusion and the need for a QM description to account for specific interactions with active sites
Herein we study the diffusion of three different types of unsaturated oxygenates (i.e., CO, formaldehyde (HCHO), and ketene (CH2CO)) in the pores of zeolite H-SSZ-13 with DFT based molecular dynamics simulations. These highly reactive oxygenates are important intermediates in the CO2-to-hydrocarbons conversion which can typically originate on the surface of the metal oxide part of the bifunctional catalyst, prior to migrating into the zeolite crystals.69 We explicitly show how specific interactions with active sites and hydrogen bonding interactions may alter the transport properties. Such effects were earlier already observed for unsaturated hydrocarbons with the Brønsted acid site (BAS) in zeolites, but are here shown to be more generally important.38,70 Furthermore, in the following, we will show how diffusion and reaction barriers in zeolites can become strongly intertwined, although these processes were traditionally viewed as independent, with markedly different barrier heights.40To understand the different degrees of diffusion–reaction entanglements, we consider the diffusion of these intermediates between two adjacent cages of H-SSZ-13 with one BAS on the connecting 8R window, and the resulting free energy profiles are displayed in Fig. 5. First, CO, a stable molecule with a weak affinity for protons (PA: 594.0 kJ mol−1), is difficult to protonate to a cation, and therefore crosses the 8R window in the form of a neutral species (Fig. 5a) by overcoming a barrier of 36.4 kJ mol−1 (Fig. 5d). Secondly, for formaldehyde (HCHO), with a higher PA value of 712.9 kJ mol−1, a weak entanglement between diffusion and reaction is observed, as indicated by the extension of the sampling along the reaction coordinate on the 2D free energy surface (Fig. 5b). However, the protonation of HCHO to CH2OH+ did not produce any local minima. Therefore, the diffusion of HCHO in H-SSZ-13 will occur predominantly its neutral form with a small barrier of 16.1 kJ mol−1 (Fig. 5e), although its cationic form (CH2OH+) may occasionally occur. Finally, ketene (CH2CO), with the highest PA value of 825.3 kJ mol−1, has a well-known high activity to react with the BAS, and the low protonation barrier of ketene (27.3 kJ mol−1 in Fig. 5f) into an acylium ion or surface-bound acetate leads to a strong diffusion–reaction entanglement as displayed in Fig. 5c. Based on the analysis of different diffusion–reaction pathways in Fig. 5f,40 ketene preferably diffuses from cage A to cage B in its neutral form (barrier of 20.4 kJ mol−1). However, diffusion as an acylium ion (CH3CO+) was also found to be kinetically feasible (with a barrier of 25.9 kJ mol−1) and strongly competitive with the pathway of the neutral ketene. Note that for those species that can form a hydrogen bond with the Brønsted acidic proton, like HCHO and ketene, the free energies of the adsorbed species in cages A and B are not the same. The Brønsted acid site (BAS) at the 8-ring window preferentially faces cage B, which leads to stronger hydrogen-bonding interactions with HCHO or ketene in cage B than in cage A. As a result, the free energy in cage B is slightly lower than that in cage A. Some snapshots along the minimum free energy coordinate are shown in Fig. 5d–f. Finally the variation in pathways from CO to HCHO and ketene suggests that this reaction–diffusion entanglement is primarily determined by the difference between the diffusion and reaction barriers. Overall, the increase in competition between diffusion and reaction from CO to ketene reflects the complexity of the diffusion behavior of highly active intermediates and reaffirms the necessity of applying a highly accurate QM accuracy methodology to comprehensively investigate both reaction and diffusion under operating conditions.
 | ||
| Fig. 5 The increasing degree of diffusion–reaction entanglement for unsaturated oxygenate mobility and reaction in H-SSZ-13. The 2D free energy profiles along the diffusion and reaction coordinates for (a) CO, (b) HCHO, and (c) CH2CO in H-SSZ-13 zeolite with one BAS on the 8R window at 673 K. The diffusion coordinate is a collective variable that describes the diffusion of guest molecules from cage A to cage B in H-SSZ-13, while the reaction coordinate is a collective variable that describes the reaction of guest molecules with the BAS of H-SSZ-13. The lines on 2D free energy surfaces are the minimum free energy paths of CO, HCHO, and ketene diffusion, and the free energy of these paths is shown in (d) CO, (e) HCHO, and (f) ketene. The reactions are described on the left side of each panel in the figure and Fig. 2b–d. Fig. 5c and f are reproduced under the terms of the CC BY 4.0 license from ref. 40. Copyright 2026 Elsevier. | ||
3.2 Intracrystalline diffusion in confined spaces accounting for specific interactions at QM accuracy using machine learning potentials
Within the Methanol-to-Hydrocarbons (MTH) process, ethene and propene are key products that must diffuse through the zeolite crystals. With time on stream, the catalyst evolves, and bulkier hydrocarbons accumulate within the pores, creating additional transport barriers for smaller species.12,71 Previous work showed that Brønsted acid sites facilitate the diffusion of ethene and propene through favourable interactions with their double bonds. Diffusion is further influenced by operating conditions such as high loadings. Moreover, reactive intermediates like ketene and formaldehyde, present during the catalytic cycle, can undergo reactions while diffusing, as was shown in the previous section. It is thus essential to use methodologies that allowing capturing these specific interactions; however as diffusion typically takes place on longer length and time scales, one also needs methodologies to calculate the energies more efficiently compared to the DFT level. Here we showcase the first results for the diffusion of ethene and ethane through H-SSZ-13 using a machine learning potential (MLP). The details of the derivation of the MLP are given in Section 2.2. From these simulations we deduce qualitative insights into the effect of acid sites on the diffusion behaviour and moreover we show how diffusion constants can be derived from the simulations. These diffusion constants can then serve in an overall kinetics model for the catalyst particle, see the introduction and final section.The influence of the presence of Brønsted acid sites (BAS) on the diffusivities of ethene and ethane in H-SSZ-13 is investigated by performing MLP-US simulations at 600 K to quantify diffusion energy profiles for hopping between two neighboring cages. These simulations can be performed on a small simulation cell and with the MLP simulation lengths up to 1 ns per window can be achieved. Three different zeolite models are considered, containing either 0, 1 or 2 acid sites on the 8-ring window through which the guest molecule jumps. The resulting free energy profiles are depicted in Fig. 6a. Note that by performing MLP simulations, smooth and properly converged free energy profiles can be obtained with significantly reduced error bars compared to analogous simulations at DFT level of theory. The barrier for hopping through the narrow 8-ring window is mainly determined by entropy for ethane and ethene, which are both much smaller than the ring surface area, as was also shown in our earlier work.39 In the absence of acid sites, ethene and ethane exhibit a similar diffusion barrier. However, the presence of BAS in the framework has a considerable impact on the diffusivities. For a single acid site on the 8-ring, the barrier for ethene diffusion is reduced by more than 50% which can be attributed to the stabilizing π–H interactions between the olefinic double bond and the acid proton. These favorable interactions remain intact while the olefin travels through the 8-ring, thus reducing the overall diffusion barrier. This effect is further magnified when two acid sites are located oppositely on the 8-ring window as this orientation allows the formation of a double π–H interaction (see Fig. 6b). In contrast, for ethane, no clear beneficial effect of acid sites is observed. Weak physisorption interactions of the paraffin with the acid proton result only in a minor reduction of the diffusion barrier. In conclusion, the presence of acid sites in the zeolite framework may have a considerable influence on the relative diffusion rates of light olefins and paraffins.
Diffusion constants Ds can be derived from the free energy profiles by applying transition state theory to obtain rate constants for diffusion, kdiff. These rate constants can be used to solve the so-called kinetic master equation or applied in a random walk model, with hopping distance λ, as developed by Smit and coworkers.72,73
| MSD(t) = 2dDst + b |
We used both strategies to derive the diffusion constant at 600 K for ethene in SSZ-13 without acid sites. According to transition state theory and the random walk model, a diffusion constant of 1.92 × 10−11 m2 s−1 is obtained while the MSD method predicts a value of 6.88 × 10−11 m2 s−1, within the same order of magnitude. To track the MSD of the ethene guest molecule, extended simulation lengths on a large simulation cell are required. Such simulations become feasible thanks to the MLP which was derived on DFT calculations on a small unit cell model (Fig. 7a). Regular MD simulations of 10 ns are carried out on a large 2 × 2 × 2 simulation cell. To improve the statistics, the final MSD curves are obtained as averages of 20 independent MD simulations in a lag time range of 5 ns. The results are plotted in Fig. 7b, wherein 2 cases are shown, one with no BAS in the simulation cell and one with 50 BAS randomly distributed in the simulation cell. The computed diffusion constants confirm the earlier observed trend that in the presence of acid sites in the framework, ethene diffusion is subject to a significant speed-up compared to a framework without acid sites. The applied procedure with the MLPs has great potential to extend to higher loadings, a variety of acid site densities. If the MLP is trained to cover complex chemistries, it becomes feasible to derive self-diffusion coefficients but also collective diffusion coefficients for industrial catalysts.
3.3 New directions for studying complex reaction environments and competing reactions via machine learning potentials based MD simulations
The rapid development of new machine learning potentials (MLP) architectures, along with advanced algorithms that efficiently generate diverse training sets on the fly, has opened new avenues to dynamically sample rare events. To illustrate this capability, we examine a complex case study that, ten years ago, was at the limit of what could be achieved with ab initio molecular dynamics.30,31 The reaction under investigation is the methylation of benzene in H-SSZ-24 (AFI topology) in the presence of two methanol molecules, as explained in the methodology section. This reaction offers a set of four chemically distinct intermediates (I: adsorbed benzene with two methanol molecules, II: adsorbed benzene at a surface methoxide species, SMS, and methanol, III: a methylated benzene, toluenium ion, and water, IV: adsorbed benzene and dimethyl ether) and both stepwise and concerted pathways connecting these intermediates (Fig. 3). Hereafter we study the various competitive pathways using the set of collective variables defined in Section 2.3.1. In the original study, the reaction network—excluding dimethyl ether formation—was explored using three collective variables (CVs) in a metadynamics simulation, producing a reasonably converged free-energy landscape after 235 ps of simulation time, albeit only by imposing substantial bias walls to restrict the accessible phase space. Considering a pace of roughly 10 seconds per step on a few dozen CPU cores, the total wall time for such a simulation would amount to nearly two continuous months.Two collective variable choices are possible (Fig. 3b): the first is a set of three coordination numbers (CNs), similar to what was done in the original study. For the second, one can imagine to project this 3-dimensional space on a 2-dimensional plane by means of linear combinations of the original CNs. While this significantly reduces the complexity of the problem at hand, it will also “merge” different states to similar regions of the 2-dimensional CV space. For an exhaustive explanation of the CVs definition, we refer back to the methods section.
Using the trained model, we began by performing eight independent well-tempered metadynamics simulations67,74 in the 3-dimensional CV space. Each simulation was propagated for 1 ns. The resulting free-energy surfaces are reported in Fig. 8b in terms of the three CVs defined in Section 2.3.1. It is interesting to note that all four minima can be explored by the walkers, however convergence in 3-dimensions is extremely challenging. This shows very clearly when looking at the free energy surfaces of the single simulations, where none managed to visit all four minima and all only visited 2 or 3 of them. The major issue in this system lies in its conformational freedom, where long timescales are needed for the reacting partners to “find each other”. To illustrate qualitatively the expected 3D free energy surface we took the mean of the 8 independent well-tempered metadynamics simulations. The resulting 3D-surface, shown in Fig. 8b, reveals—as anticipated from the collective variable definition—three minima positioned at the vertices of an equilateral triangle in the space defined by the three CNs. Each minimum lies around 1 along the x-, y-, or z-axis, as was already schematically depicted in Fig. 3b.
 | ||
Fig. 8 (a) Reaction scheme of the benzene methylation reaction by two methanol molecules in H-SSZ-24. The reactants (I) can undergo concerted methylation towards the products (III). Alternatively, a surface methoxide species can be formed first (II) or the two methanol molecules can condense to dimethyl ether (IV). To illustrate the wide variety of transition paths connecting the various minima, various snapshots of the direct methylation, methoxide and dimethyl ether formation are shown. (b) Mean 3-dimensional free energy surface obtained from a set of 8 independent well-tempered metadynamics simulations. The separate profiles are reported on the right, showcasing how none of the simulations has visited the four minima even after 1 ns of simulation time. (c) 2-Dimensional free energy surface obtained from umbrella sampling simulations in terms of CV1 and CV2, defined as CV1 = 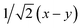 and CV2 = and CV2 = 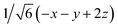 , where x, y, z were defined in Section 2.3.1 and schematically in Fig. 3b. , where x, y, z were defined in Section 2.3.1 and schematically in Fig. 3b. | ||
To illustrate the wide variety of transition paths connecting the various minima we show some snapshots of the direct methylation, methoxide and dimethyl ether formation in Fig. 8a. It is clear that various possible transition paths exist due to the large configurational freedom and the participation of one or either two methanol molecules in the transition paths. Note that it is not possible anymore to define one transition state as is often done in static simulation methods, but that a region of transition paths exists. As mentioned in the original publication of De Wispelaere et al., this is a very challenging reaction given the large conformational freedom in the large pore zeolite.31 The high degree of conformational flexibility and the various competing paths, makes a faster pace in the metadynamics bias deposition pace also non-ideal, as one risks forcing bond breaking before the reactive partners have the correct spatial disposition creating unphysical transition states.
To improve the convergence of the final free-energy surface, we constructed a fine grid of 2-dimensional umbrellas based on the following collective variables: CV1 = 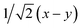 and CV2 =
and CV2 = 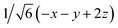 which were defined in Section 2.3.1 and performed umbrella-sampling simulations.75,76 We used an initial force constant of 1500 kJ mol−1, propagating each umbrella for 250 ps, and subsequently added a second set of umbrellas with a tighter force constant (2500 kJ mol−1) propagated for 125 ps to fill residual gaps in the two-dimensional CV space. The thus obtained 2D free-energy surface is shown in Fig. 8c. The four expected minima are clearly distinguishable, although we emphasize that these results are purely qualitative: the strong “compression” of phase space into only two dimensions prevents a clean separation of the different reactive channels (for example, the I → II pathway cannot be distinguished from the I → IV → II route). Obtaining reliable, quantitative results would require a more suitable CV selection or restricting the analysis to specific reaction steps. Nevertheless, this example demonstrates how accessible it has become to investigate complex mechanistic landscapes with MLPs. The production runs amount to almost 175 ns of total simulation time, but required only a few days of actual user time. These numbers were unthinkable at the time of the original study and highlight how MLPs enable more extensive exploration and improved convergence in simulations of zeolite catalysis.
which were defined in Section 2.3.1 and performed umbrella-sampling simulations.75,76 We used an initial force constant of 1500 kJ mol−1, propagating each umbrella for 250 ps, and subsequently added a second set of umbrellas with a tighter force constant (2500 kJ mol−1) propagated for 125 ps to fill residual gaps in the two-dimensional CV space. The thus obtained 2D free-energy surface is shown in Fig. 8c. The four expected minima are clearly distinguishable, although we emphasize that these results are purely qualitative: the strong “compression” of phase space into only two dimensions prevents a clean separation of the different reactive channels (for example, the I → II pathway cannot be distinguished from the I → IV → II route). Obtaining reliable, quantitative results would require a more suitable CV selection or restricting the analysis to specific reaction steps. Nevertheless, this example demonstrates how accessible it has become to investigate complex mechanistic landscapes with MLPs. The production runs amount to almost 175 ns of total simulation time, but required only a few days of actual user time. These numbers were unthinkable at the time of the original study and highlight how MLPs enable more extensive exploration and improved convergence in simulations of zeolite catalysis.
Finally it is interesting to comment on how far it would be possible to also deduce kinetic constants from the simulations which can afterwards be fed into an overall kinetics model. Rate constants are mostly derived from enhanced sampling simulations using transition state theory where barrier recrossings are neglected. The underlying reason for this assumption can be traced back to the high computational expense, especially when the energies and forces are calculated with expensive Density Functional Theory calculations or other first principles methods. With the use of MLPs, it becomes possible to obtain dynamically corrected rates explicitly accounting for barrier recrossings using the reactive flux formalism.25,77–79 Thousands of MD trajectories atop of the trajectories need to be determined to evaluate whether they end in the reactant or product basin.37 With the MLP based methodologies it will also become possible in the future to evaluate reaction rates using path based methods that also necessitate the generation of a lot of trajectories. One example is the usage of Replica Exchange Transition Interface Sampling (RETIS) where an ensemble of reactive paths is generated with a Monte Carlo procedure.80,81 These methods sample the transition path without prior knowledge of the transition state and without using CVs.82–85 So far the method has not been applied to complex reactions under investigation here, due to the computational expense, but new studies are underway also from some of the present authors to showcase the first successful application of these methods to calculate dynamically accurate reaction rates for complex zeolite catalysed reactions.
4. Conclusions and perspectives
The three case studies presented in this paper underscore the need for a consistent dynamical view across length and time scales to rationally design industrial catalysts. In the first case, we illustrated the diffusion of key intermediates in the CO2-to-hydrocarbons process within small-pore zeolites. Evidence was given that diffusion itself must be treated with quantum-mechanical (QM) methods, as reactive intermediates can interact explicitly with Brønsted acid sites and may even react while diffusing. Furthermore diffusion may become hindered and may in some circumstances compete with reactions. In current modelling practice reactions are typically treated with QM based methods whereas diffusion is treated with classical force fields, as the latter are taking place on longer length and time scales. This inconsistent and decoupled description of reaction and diffusion for complex industrial catalysts is clearly inadequate and fails to capture a catalyst in action. The necessity of a coupled description of reaction and diffusion was also emphasized recently by Liu et al.2,86In the second and third case studies we illustrated how machine learning potentials (MLPs) enable a path forward to obtain dynamic information with quantum-level accuracy for individual steps of a catalytic trajectory, namely for reactive and diffusion events.
For reactions in industrial catalysts, which often occur under extreme temperatures and pressures, it is essential to account for operating conditions.1,8,29,87 The most natural way to do so is to use first principles molecular dynamics simulations, where the catalytic reaction and their intermediates are followed on the fly at the actual reaction conditions.1 With the rise of MLPs, such simulations become much more accessible as shown in this paper. However as reactions are rare events, one needs to rely on enhanced sampling molecular dynamics simulations.74,88–91 As has become clear from case study 3 in this paper, proper choice of CVs can become rather cumbersome.27 An alternative would be to rely on path sampling methods, but their straightforward application to industrially relevant catalysts was to date out of reach due to the large computational cost to generate many paths at the DFT level of theory.81–85,92,93 However with the rise of MLPs, we foresee a more frequent usage of such methods with the rise of reliable MLPs that are able to capture the chemistry of the complex catalyst.
Clearly describing all events of a catalytic trajectory consistently from the nanoscale to the catalyst particle level, poses enormous challenges. However we argue that it is necessary to obtain such a consistent dynamics view across length and time scales. This is especially true as catalysts evolve continuously across length and time scales.23 During a catalytic cycle, new active sites can be formed dynamically, a process that is highly dependent on the operating conditions. Furthermore the dynamics at the crystal-particle level are strongly influenced by spatial gradients, crystal size, and morphology.17 A typical example of such a highly dynamical complex system is an MTO catalyst. A recent review paper by Liu et al. also clearly emphasized how diffusion, reaction network evolution, coke buildup/regeneration and catalyst evolution interact continuously, with each process influencing the others dynamically.2 Within this paper, we focused on the spatial heterogeneities observed within the catalyst particle level, ranging from Å to µm and argue that future modelling studies should focus on obtaining a consistent dynamics view of all events within the crystal particle level with QM accuracy. Catalytic solids used in industrial applications are further embedded within binders or other supports and within the industrial reactor, however these heterogeneities fall beyond the scope of this paper.94 Even at the level of the catalyst particle, where various events are intertwined, namely molecular diffusion, adsorption–desorption, chemical reactions and also surface barriers, substantial gradients in the concentration of reactants and products. Every spatial modification in the catalyst leads to temporal gradients and change of dynamics. Given this complexity, future modeling studies should fully embrace the spatial heterogeneities present in the crystal from the nanoscale local level to the crystal particle level.
However, achieving a fully consistent dynamic view of an entire catalytic process—including all relevant time scales—requires more than MLPs alone. With MLPs one can extend accessible length and time scales with roughly three orders of magnitude with current computational power, compared to accessible length and time scales of hundreds of picoseconds and a few nm accessible with DFT. Therefore, information on individual reactions and diffusion events must be integrated into multiscale models that bridge length and time scales in order to reach the scales of the catalyst particle level. Various models exist to tackle the multi-length/time scale problem, such as microkinetic modeling, kinetic Monte Carlo (kMC) models and heat and mass transport models to account for the inhomogeneities in temperatures and reactant concentrations in the reactor, but none of them account for the kinetics of all events taking place at the crystal particle level with quantum accuracy.95–99 Furthermore within the set of existing multiscale models, major advances are still necessary to make them applicable for complex industrial catalysts and to fully account for the dynamic evolution of the catalyst, spatiotemporal gradients and the interplay between diffusion and reaction. There exist also other phenomenological models to describe the combined effects of reaction and diffusion, however in this case one does not (or to a limited extent) use atomistic scale input.100–102
Overall, the results presented in this paper highlight the need for a consistent dynamics description with quantum accuracy across length and time scales to enable rational catalyst design. Machine learning potentials (MLPs) offer a promising pathway toward this goal by providing kinetic and transport properties for events from the nano- to mesoscale with quantum-level accuracy at significantly reduced computational cost. In the future these data can possibly be integrated into multiscale models such as kMC or reaction–diffusion frameworks, which are essential to reach the length and time scales relevant for industrial catalysts.
Author contributions
Veronique Van Speybroeck: conceptualization, funding acquisition, methodology, supervision, resources, project administration, visualization, writing – original draft, writing – review & editing. Massimo Bocus: investigation, formal analysis, software, methodology, visualization, data curation, writing – review & editing. Wei Chen: investigation, formal analysis, software, methodology, visualization, data curation, writing – review & editing. Pieter Cnudde: investigation, formal analysis, software, methodology, visualization, data curation, writing – review & editing.Conflicts of interest
There are no conflicts to declare.Data availability
All computational data generated in this study have been deposited in the Zenodo repository and are accessible at https://doi.org/10.5281/zenodo.18172885 under open access terms.Acknowledgements
V. V. S. is grateful for funding obtained from the Research Fund of Ghent University (BOF) for financial support. M. B. acknowledges the Fund for Scientific Research Flanders for funding for a junior postdoctoral fellowship (Grant Number 1269725N). P. C. acknowledges the Fund for Scientific Research Flanders for funding for a senior postdoctoral fellowship (Grant Number 1252525N). W. C. acknowledges the European Union’s Horizon 2022 Research and Innovation Program for a Marie Skłodowska-Curie Grant (Grant Number 101104138). V. V. S. acknowledges funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme 101201153 (ERC-AdG-2024, TIME-project). The resources and services used in this work were provided by the VSC (Flemish Supercomputer Center), funded by the Research Foundation - Flanders (FWO) and the Flemish Government. We also acknowledge the EuroHPC Joint Undertaking for awarding this project access to the EuroHPC supercomputer LUMI, hosted by CSC (Finland) and the LUMI consortium through a EuroHPC Regular Access Call (project EHPC-REG-2024R02-213).Notes and references
- V. Van Speybroeck, M. Bocus, P. Cnudde and L. Vanduyfhuys, ACS Catal., 2023, 13, 11455–11493 CrossRef PubMed.
- S. F. Lin, H. Li, P. Tian, Y. X. Wei, M. Ye and Z. M. Liu, J. Am. Chem. Soc., 2025, 147, 11585–11607 CrossRef CAS PubMed.
- J. Bao, G. Yang, Y. Yoneyama and N. Tsubaki, ACS Catal., 2019, 9, 3026–3053 CrossRef CAS.
- E. Alvarez, N. Guillou, C. Martineau, B. Bueken, B. Van de Voorde, C. Le Guillouzer, P. Fabry, F. Nouar, F. Taulelle, D. de Vos, J. S. Chang, K. H. Cho, N. Ramsahye, T. Devic, M. Daturi, G. Maurin and C. Serre, Angew. Chem., Int. Ed., 2015, 54, 3664–3668 CrossRef CAS PubMed.
- A. Dokania, A. Ramirez, A. Bavykina and J. Gascon, ACS Energy Lett., 2019, 4, 167–176 CrossRef CAS.
- T. Cordero-Lanzac, I. Capel Berdiell, A. Airi, S.-H. Chung, J. L. Mancuso, E. A. Redekop, C. Fabris, L. Figueroa-Quintero, J. C. Navarro de Miguel, J. Narciso, E. V. Ramos-Fernandez, S. Svelle, V. Van Speybroeck, J. Ruiz-Martínez, S. Bordiga and U. Olsbye, JACS Au, 2024, 4, 744–759 CrossRef CAS PubMed.
- B. Baumgartner, A. Wach, X. Ye, E. Ploetz and B. M. Weckhuysen, Adv. Mater., 2025, 37, 2415135 CrossRef CAS PubMed.
- L. Bonati, D. Polino, C. Pizzolitto, P. Biasi, R. Eckert, S. Reitmeier, R. Schlogl and M. Parrinello, Proc. Natl. Acad. Sci. U. S. A., 2023, 120, e2313023120 CrossRef CAS PubMed.
- V. Van Speybroeck, W. Temmerman, M. Bocus and L. Vanduyfhuys, Annu. Rev. Phys. Chem., 2026, 77, 371–395 CrossRef CAS PubMed.
- O. Knio, H. J. Fang, S. E. Boulfelfel, S. Nair and D. S. Sholl, J. Phys. Chem. C, 2020, 124, 15241–15252 CrossRef CAS.
- D. Dubbeldam, S. Calero, T. L. M. Maesen and B. Smit, Angew. Chem., Int. Ed., 2003, 42, 3624–3626 CrossRef CAS PubMed.
- V. Van Speybroeck, K. De Wispelaere, J. Van der Mynsbrugge, M. Vandichel, K. Hemelsoet and M. Waroquier, Chem. Soc. Rev., 2014, 43, 7326–7357 RSC.
- B. Smit and T. L. M. Maesen, Chem. Rev., 2008, 108, 4125–4184 CrossRef CAS PubMed.
- C. Vogt and B. M. Weckhuysen, Nat. Rev. Chem., 2022, 6, 89–111 CrossRef PubMed.
- B. M. Weckhuysen, Catal. Chem. Biol., Proc. Int. Solvay Conf. Chem., 24th, 2018, 205–220, DOI:10.1142/9789813237179_0031.
- B. M. Weckhuysen, Angew. Chem., Int. Ed., 2009, 48, 4910–4943 CrossRef CAS PubMed.
- I. L. C. Buurmans and B. M. Weckhuysen, Nat. Chem., 2012, 4, 873–886 CrossRef CAS PubMed.
- F. Meirer and B. M. Weckhuysen, Nat. Rev. Mater., 2018, 3, 324–340 CrossRef.
- P. Rowe, V. L. Deringer, P. Gasparotto, G. Csanyi and A. Michaelides, J. Chem. Phys., 2020, 153, 19 CrossRef PubMed.
- I. Batatia, D. P. Kovács, G. N. C. Simm, C. Ortner and G. Csányi, arXiv, 2023, preprint, arXiv:2206.07697, DOI:10.48550/arXiv.2206.07697.
- S. Batzner, A. Musaelian, L. Sun, M. Geiger, J. P. Mailoa, M. Kornbluth, N. Molinari, T. E. Smidt and B. Kozinsky, Nat. Commun., 2022, 13, 2453 CrossRef CAS PubMed.
- A. Musaelian, S. Batzner, A. Johansson, L. Sun, C. J. Owen, M. Kornbluth and B. Kozinsky, Nat. Commun., 2023, 14, 579 CrossRef CAS PubMed.
- V. Van Speybroeck, Philos. Trans. R. Soc., A, 2023, 381, 20220239 CrossRef PubMed.
- M. Bocus, S. Vandenhaute and V. Van Speybroeck, Angew. Chem., Int. Ed., 2025, 64, e202413637 CrossRef CAS PubMed.
- D. Frenkel and B. Smit, Understanding Molecular Simulation, Academic Press, Elsevier, 2nd edn, 2002 Search PubMed.
- B. Peters, in Reaction Rate Theory and Rare Events Simulations, ed. B. Peters, Elsevier, Amsterdam, 2017, pp. 227–271, DOI:10.1016/B978-0-44-456349-1.00010-6.
- G. M. Piccini, M. S. Lee, S. F. Yuk, D. F. Zhang, G. Collinge, L. Kollias, M. T. Nguyen, V. A. Glezakou and R. Rousseau, Catal. Sci. Technol., 2022, 12, 12–37 RSC.
- G. Collinge, S. F. Yuk, M.-T. Nguyen, M.-S. Lee, V.-A. Glezakou and R. Rousseau, ACS Catal., 2020, 10, 9236–9260 CrossRef CAS.
- P. Cnudde, K. De Wispelaere, L. Vanduyfhuys, R. Demuynck, J. Van der Mynsbrugge, M. Waroquier and V. Van Speybroeck, ACS Catal., 2018, 8, 9579–9595 CrossRef CAS PubMed.
- K. De Wispelaere, S. Bailleul and V. Van Speybroeck, Catal. Sci. Technol., 2016, 6, 2686–2705 RSC.
- K. De Wispelaere, B. Ensing, A. Ghysels, E. J. Meijer and V. Van Speybroeck, Chem.–Eur. J., 2015, 21, 9385–9396 CrossRef CAS PubMed.
- H. Sidky, W. Chen and A. L. Ferguson, Mol. Phys., 2020, 118, e1737742 CrossRef.
- W. Chen, H. Sidky and A. L. Ferguson, J. Chem. Phys., 2019, 150, 214114 CrossRef PubMed.
- W. Chen, A. R. Tan and A. L. Ferguson, J. Chem. Phys., 2018, 149, 072312 CrossRef PubMed.
- Y. Wang, J. M. Lamim Ribeiro and P. Tiwary, Curr. Opin. Struct. Biol., 2020, 61, 139–145 CrossRef CAS PubMed.
- F. Noe and C. Clementi, Curr. Opin. Struct. Biol., 2017, 43, 141–147 CrossRef CAS PubMed.
- M. Bocus, R. Goeminne, A. Lamaire, M. Cools-Ceuppens, T. Verstraelen and V. Van Speybroeck, Nat. Commun., 2023, 14, 1008 CrossRef CAS PubMed.
- P. Cnudde, E. A. Redekop, W. L. Dai, N. G. Porcaro, M. Waroquier, S. Bordiga, M. Hunger, L. D. Li, U. Olsbye and V. Van Speybroeck, Angew. Chem., Int. Ed., 2021, 60, 10016–10022 CrossRef CAS PubMed.
- P. Cnudde, R. Demuynck, S. Vandenbrande, M. Waroquier, G. Sastre and V. Van Speybroeck, J. Am. Chem. Soc., 2020, 142, 6007–6017 CrossRef CAS PubMed.
- W. Chen, P. Cnudde and V. Van Speybroeck, J. Catal., 2026, 453, 116546 CrossRef CAS.
- J. Kärger, D. M. Ruthven and D. N. Theodorou, Diffusion in Nanoporous Materials, Wiley-VCH Verlag GmbH & Co. KGaA, 2012 Search PubMed.
- A. Ghysels, S. L. C. Moors, K. Hemelsoet, K. De Wispelaere, M. Waroquier, G. Sastre and V. Van Speybroeck, J. Phys. Chem. C, 2015, 119, 23721–23734 CrossRef CAS.
- T. Titze, A. Lauerer, L. Heinke, C. Chmelik, N. E. R. Zimmermann, F. J. Keil, D. M. Ruthven and J. Kärger, Angew. Chem., Int. Ed., 2015, 54, 14580–14583 CrossRef CAS PubMed.
- S. Hwang and J. Kärger, Adsorption, 2020, 26, 1001–1013 CrossRef CAS.
- M. Mortén, Ł. Mentel, A. Lazzarini, I. A. Pankin, C. Lamberti, S. Bordiga, V. Crocellà, S. Svelle, K. P. Lillerud and U. Olsbye, ChemPhysChem, 2018, 19, 484–495 CrossRef PubMed.
- E. A. Redekop, A. Lazzarini, S. Bordiga and U. Olsbye, J. Catal., 2020, 385, 300–312 CrossRef CAS.
- E. Yoda, J. N. Kondo and K. Domen, J. Phys. Chem. B, 2005, 109, 1464–1472 CrossRef CAS PubMed.
- S. S. Gao, Z. Q. Liu, S. T. Xu, A. M. Zheng, P. F. Wu, B. Li, X. S. Yuan, Y. X. Wei and Z. M. Liu, J. Catal., 2019, 377, 51–62 CrossRef CAS.
- J. Han, Z. Liu, H. Li, J. Zhong, W. Zhang, J. Huang, A. Zheng, Y. Wei and Z. Liu, ACS Catal., 2020, 10, 8727–8735 CrossRef CAS.
- V. Shostak, E. Redekop and U. Olsbye, Catal. Today, 2022, 417, 113785 CrossRef.
- B. C. Bukowski, F. J. Keil, P. I. Ravikovitch, G. Sastre, R. Q. Snurr and M. O. Coppens, Adsorption, 2021, 27, 683–760 CrossRef CAS.
- J. S. Martinez-Espin, K. De Wispelaere, M. W. Erichsen, S. Svelle, T. V. W. Janssens, V. Van Speybroeck, P. Beato and U. Olsbye, J. Catal., 2017, 349, 136–148 CrossRef CAS.
- C. Braga and K. P. Travis, J. Chem. Phys., 2005, 123, 134101 CrossRef PubMed.
- Y. K. Zhang and W. T. Yang, Phys. Rev. Lett., 1998, 80, 890 CrossRef CAS.
- J. P. Perdew, K. Burke and M. Ernzerhof, Phys. Rev. Lett., 1996, 77, 3865–3868 CrossRef CAS PubMed.
- S. Grimme, J. Antony, S. Ehrlich and H. Krieg, J. Chem. Phys., 2010, 132, 154104 CrossRef PubMed.
- M. F. Peintinger, D. V. Oliveira and T. Bredow, J. Comput. Chem., 2013, 34, 451–459 CrossRef CAS PubMed.
- M. Krack, Theor. Chem. Acc., 2005, 114, 145–152 Search PubMed.
- M. Bocus and L. Vanduyfhuys, ThermoLIB -- A Python Library for Constructing and Post-Processing Free Energy Surfaces to Extract Thermodynamic and Kinetic Properties, arXiv, 2026, preprint, arXiv:2601.23071, DOI:10.48550/arXiv.2601.23071.
- W. Chen, J. Sobalska, W. Fu, K. A. Tarach, M. Bocus, T. Tang, K. Góra-Marek and V. Van Speybroeck, J. Am. Chem. Soc., 2025, 147, 24719–24733 CrossRef CAS PubMed.
- I. Batatia, D. P. Kovacs, G. Simm, C. Ortner and G. Csányi, Adv. Neural Inf. Process. Syst., 2022, 35, 11423–11436 Search PubMed.
- S. Vandenhaute, M. Cools-Ceuppens, S. DeKeyser, T. Verstraelen and V. Van Speybroeck, npj Comput. Mater., 2023, 9, 19 CrossRef.
- M. Ceriotti, J. More and D. E. Manolopoulos, Comput. Phys. Commun., 2014, 185, 1019–1026 CrossRef CAS.
- V. Kapil, M. Rossi, O. Marsalek, R. Petraglia, Y. Litman, T. Spura, B. Q. Cheng, A. Cuzzocrea, R. H. Meissner, D. M. Wilkins, B. A. Helfrecht, P. Juda, S. P. Bienvenue, W. Fang, J. Kessler, I. Poltavsky, S. Vandenbrande, J. Wieme, C. Corminboeuf, T. D. Kuhne, D. E. Manolopoulos, T. E. Markland, J. O. Richardson, A. Tkatchenko, G. A. Tribello, V. Van Speybroeck and M. Ceriotti, Comput. Phys. Commun., 2019, 236, 214–223 CrossRef CAS.
- J. VandeVondele and J. Hutter, J. Chem. Phys., 2007, 127, 114105 CrossRef PubMed.
- J. Hutter, M. Iannuzzi, F. Schiffmann and J. VandeVondele, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2014, 4, 15–25 CAS.
- M. Bonomi, G. Bussi, C. Camilloni, G. A. Tribello, P. Banas, A. Barducci, M. Bernetti, P. G. Bolhuis, S. Bottaro, D. Branduardi, R. Capelli, P. Carloni, M. Ceriotti, A. Cesari, H. C. Chen, W. Chen, F. Colizzi, S. De, M. De La Pierre, D. Donadio, V. Drobot, B. Ensing, A. L. Ferguson, M. Filizola, J. S. Fraser, H. H. Fu, P. Gasparotto, F. L. Gervasio, F. Giberti, A. Gil-Ley, T. Giorgino, G. T. Heller, G. M. Hocky, M. Iannuzzi, M. Invernizzi, K. E. Jelfs, A. Jussupow, E. Kirilin, A. Laio, V. Limongelli, K. Lindorff-Larsen, T. Lohr, F. Marinelli, L. Martin-Samos, M. Masetti, R. Meyer, A. Michaelides, C. Molteni, T. Morishita, M. Nava, C. Paissoni, E. Papaleo, M. Parrinello, J. Pfaendtner, P. Piaggi, G. Piccini, A. Pietropaolo, F. Pietrucci, S. Pipolo, D. Provasi, D. Quigley, P. Raiteri, S. Raniolo, J. Rydzewski, M. Salvalaglio, G. C. Sosso, V. Spiwok, J. Sponer, D. W. H. Swenson, P. Tiwary, O. Valsson, M. Vendruscolo, G. A. Voth and A. White, Nat. Methods, 2019, 16, 670–673 CrossRef PubMed.
- I. Batatia, P. Benner, Y. Chiang, A. M. Elena, D. P. Kovács, J. Riebesell, X. R. Advincula, M. Asta, M. Avaylon, W. J. Baldwin, F. Berger, N. Bernstein, A. Bhowmik, F. Bigi, S. M. Blau, V. Cărare, M. Ceriotti, S. Chong, J. P. Darby, S. De, F. Della Pia, V. L. Deringer, R. Elijošius, Z. El-Machachi, E. Fako, F. Falcioni, A. C. Ferrari, J. L. A. Gardner, M. J. Gawkowski, A. Genreith-Schriever, J. George, R. E. A. Goodall, J. Grandel, C. P. Grey, P. Grigorev, S. Han, W. Handley, H. H. Heenen, K. Hermansson, C. H. Ho, S. Hofmann, C. Holm, J. Jaafar, K. S. Jakob, H. Jung, V. Kapil, A. D. Kaplan, N. Karimitari, J. R. Kermode, P. Kourtis, N. Kroupa, J. Kullgren, M. C. Kuner, D. Kuryla, G. Liepuoniute, C. Lin, J. T. Margraf, I.-B. Magdău, A. Michaelides, J. H. Moore, A. A. Naik, S. P. Niblett, S. W. Norwood, N. O’Neill, C. Ortner, K. A. Persson, K. Reuter, A. S. Rosen, L. A. M. Rosset, L. L. Schaaf, C. Schran, B. X. Shi, E. Sivonxay, T. K. Stenczel, C. Sutton, V. Svahn, T. D. Swinburne, J. Tilly, C. van der Oord, S. Vargas, E. Varga-Umbrich, T. Vegge, M. Vondrák, Y. Wang, W. C. Witt, T. Wolf, F. Zills and G. Csányi, J. Chem. Phys., 2025, 163, 184110 CrossRef CAS PubMed.
- W. Chen and P. Cnudde, Chin. J. Struct. Chem., 2024, 43, 100412 CAS.
- S. Bailleul, K. Dedecker, P. Cnudde, L. Vanduyfhuys, M. Waroquier and V. Van Speybroeck, J. Catal., 2020, 388, 38–51 CrossRef CAS.
- I. Lezcano-Gonzalez, E. Campbell, A. E. J. Hoffman, M. Bocus, I. V. Sazanovich, M. Towrie, M. Agote-Aran, E. K. Gibson, A. Greenaway, K. De Wispelaere, V. Van Speybroeck and A. M. Beale, Nat. Mater., 2020, 19, 1081–1087 CrossRef CAS PubMed.
- D. Dubbeldam, E. Beerdsen, S. Calero and B. Smit, J. Phys. Chem. B, 2006, 110, 3164–3172 CrossRef CAS PubMed.
- D. Dubbeldam, E. Beerdsen, T. J. H. Vlugt and B. Smit, J. Chem. Phys., 2005, 122, 224712 CrossRef CAS PubMed.
- B. Ensing, A. Laio, M. Parrinello and M. L. Klein, J. Phys. Chem. B, 2005, 109, 6676–6687 CrossRef CAS PubMed.
- G. M. Torrie and J. P. Valleau, J. Chem. Phys., 1977, 66, 1402–1408 CrossRef CAS.
- G. M. Torrie and J. P. Valleau, Chem. Phys. Lett., 1974, 28, 578–581 CrossRef CAS.
- D. Chandler, J. Chem. Phys., 1978, 68, 2959–2970 CrossRef CAS.
- C. H. Bennett, in Algorithms for Chemical Computations, American Chemical Society, 1977, ch. 4, vol. 46, pp. 63–97 Search PubMed.
- B. Peters, in Reaction Rate Theory and Rare Events Simulations, ed. B. Peters, Elsevier, Amsterdam, 2017, pp. 335–362, DOI:10.1016/B978-0-44-456349-1.00013-1.
- T. S. van Erp, Europhys. Lett., 2023, 143, 30001 CrossRef CAS.
- P. G. Bolhuis and D. W. H. Swenson, Adv. Theory Simul., 2021, 4, 2000237 CrossRef.
- T. S. van Erp and P. G. Bolhuis, J. Comput. Phys., 2005, 205, 157–181 CrossRef.
- T. S. van Erp, D. Moroni and P. G. Bolhuis, J. Chem. Phys., 2003, 118, 7762–7774 CrossRef CAS.
- D. Moroni, P. G. Bolhuis and T. S. van Erp, J. Chem. Phys., 2004, 120, 4055–4065 CrossRef CAS PubMed.
- R. Cabriolu, K. M. S. Refsnes, P. G. Bolhuis and T. S. van Erp, J. Chem. Phys., 2017, 147, 17 CrossRef PubMed.
- S. Lin, Y. Zhi, Z. Liu, J. Yuan, W. Liu, W. Zhang, Z. Xu, A. Zheng, Y. Wei and Z. Liu, Natl. Sci. Rev., 2022, 9, nwac151 CrossRef CAS PubMed.
- A. N. Alexandrova and P. Christopher, Matter, 2025, 8, 102209 CrossRef CAS.
- L. Sutto, S. Marsili and F. L. Gervasio, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2012, 2, 771–779 CAS.
- O. Valsson, P. Tiwary and M. Parrinello, Annu. Rev. Phys. Chem., 2016, 67, 159–184 CrossRef CAS PubMed.
- E. D. Goodman, A. C. Johnston-Peck, E. M. Dietze, C. J. Wrasman, A. S. Hoffman, F. Abild-Pedersen, S. R. Bare, P. N. Plessow and M. Cargnello, Nat. Catal., 2019, 2, 748–755 CrossRef CAS PubMed.
- G. M. Torrie and J. P. Valleau, J. Comput. Phys., 1977, 23, 187–199 CrossRef.
- P. G. Bolhuis, D. Chandler, C. Dellago and P. L. Geissler, Annu. Rev. Phys. Chem., 2002, 53, 291–318 CrossRef CAS PubMed.
- B. Peters, in Reaction Rate Theory and Rare Events Simulations, ed. B. Peters, Elsevier, Amsterdam, 2017, pp. 507–537, DOI:10.1016/B978-0-44-456349-1.00019-2.
- A. Urakawa and A. Baiker, Top. Catal., 2009, 52, 1312–1322 CrossRef CAS.
- A. Bruix, J. T. Margraf, M. Andersen and K. Reuter, Nat. Catal., 2019, 2, 659–670 CrossRef CAS.
- G. B. Marin, V. V. Galvita and G. S. Yablonsky, J. Catal., 2021, 404, 745–759 CrossRef CAS.
- M. Stamatakis and D. G. Vlachos, ACS Catal., 2012, 2, 2648–2663 CrossRef CAS.
- A. H. Motagamwala and J. A. Dumesic, Chem. Rev., 2021, 121, 1049–1076 CrossRef CAS PubMed.
- B. W. J. Chen, L. Xu and M. Mavrikakis, Chem. Rev., 2021, 121, 1007–1048 CrossRef CAS PubMed.
- B. D. Vandegehuchte, I. R. Choudhury, J. W. Thybaut, J. A. Martens and G. B. Marin, J. Phys. Chem. C, 2014, 118, 22053–22068 CrossRef CAS.
- M.-O. Coppens, T. Weissenberger, Q. Zhang and G. Ye, Adv. Mater. Interfaces, 2021, 8, 2001409 CrossRef CAS.
- T. Omojola, AIChE J., 2025, 71, e18865 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |