Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Unravelling nanoscale chemistries in complex biological systems using photoinduced force microscopy (PiFM)
Josh A. Davies-Jones *a,
Jacob K. Pattem
*a,
Jacob K. Pattem *b,
Liana R. Azizova
*b,
Liana R. Azizova c,
Reem Al-Luaibi
c,
Reem Al-Luaibi d,
David Morgan
d,
David Morgan de,
Phil Stephens
de,
Phil Stephens b,
Bo Hou
b,
Bo Hou e,
Aled Clayton
e,
Aled Clayton fg,
Wayne Nishio
fg,
Wayne Nishio b and
Philip R. Davies
b and
Philip R. Davies de
de
aGeorgia Institute of Technology, North Avenue, Atlanta, GA 30332, USA
bCardiff School of Dentistry, Heath Park, Cardiff University, CF14 4XY, UK. E-mail: PattemJ@cardiff.ac.uk
cSchool of Life and Medical Sciences, University of Hertfordshire, Hatfield, UK
dCardiff Catalysis Institute, School of Chemistry, Cardiff University, CF24 4HQ, UK
eHarwellXPS, Research Complex at Harwell, Rutherford Appleton Laboratory, Harwell Science and Innovation Campus, OX11 0FA, UK
fSchool of Physics, Cardiff University, CF24 4HQ, UK
gSchool of Medicine, Tenovus Building, Room GF16, University Hospital of Wales, Heath Park, Cardiff, CF14 4XN, UK
First published on 21st April 2026
Abstract
Direct interrogation of nanoscale chemical features on and within biological structures remains a major frontier challenge in biophysical and biomedical research. These nanoscale features govern molecular organization, structural dynamics, and cellular function, yet conventional non-invasive techniques such as Fourier-transform infrared spectroscopy (FTIR) are fundamentally limited by optical diffraction. Although hybrid approaches, including scattering-type scanning near-field optical microscopy (s-SNOM) and atomic force microscopy infrared spectroscopy (AFM-IR), have advanced spatial resolution, they remain insufficient to resolve individual macromolecular assemblies. Furthermore, precise control over the depth of analysis within biological architectures, where critical molecular information underpinning intra- and inter-cellular communication resides, has yet to be fully achieved. Here, we employ photo-induced force microscopy (PiFM), an atomic force microscopy (AFM) based technique that directly measures forces arising from light-induced polarization in the near-field region. These forces, typically on the order of piconewtons, are localized perpendicular to the sample surface. This localization enables a theoretical spatial resolution approaching 5 nm, with depth sensitivity spanning approximately 2–200 nm. Crucially, PiFM can operate under ambient and environmentally controlled conditions, preserving physiologically relevant architectures in vitro. Our findings demonstrate that aldehyde-based fixing, including formalin treatment, causes substantial chemical modifications and spectral overlap within the nuclear envelopes of oral mucosa lamina propria progenitor cells (OMLP-PCs). These effects highlight the necessity for rigorous validation of sample-preparation protocols in nano-spectroscopy. In contrast, live-cell PiFM imaging under controlled humidity conditions enables visualisation of native biomolecular states and dynamic cellular processes in OMLP-PCs. Our approach captured whole-cell and membrane-level phenomena, including extracellular vesicle (EV) biogenesis and nuclear stress responses. PiFM mapping of isolated human bone marrow stromal cell (hBMSC) EVs further uncovers nanoscale compositional heterogeneity at the single-EV level. This work demonstrates the application of PiFM as a transformative nano-spectroscopic tool for probing the structural and spatial chemical information of biological matter, potentially down to 5 nm resolution. By bridging physical chemistry and biophysics, PiFM enables direct visualisation of compositional heterogeneity under near-physiological conditions, offering a non-invasive and in situ pathway for nanoscale characterisation and mechanistic understanding of complex biological systems.
Introduction
Nanoscale infrared (IR) spectroscopy provides a unique window into the chemical organization of biological materials by probing molecular vibrations with nanometre-scale precision. Conventional Fourier-transform infrared (FTIR) microscopy has long been used to characterise cells and tissues based on their rich vibrational fingerprints, yet its spatial resolution is fundamentally limited by diffraction to several micrometres. The advent of scattering-type scanning near-field optical microscopy (s-SNOM) and atomic force microscopy-infrared spectroscopy (AFM-IR) has enabled chemical imaging below this limit,1–4 revealing spatially heterogeneous distributions of proteins, lipids, and nucleic acids within single cells. In these hybrid techniques, an AFM probe tip acts as a local optical antenna: in AFM-IR, infrared absorption induces a photothermal expansion detected by the cantilever, while in Photoinduced Force Microscopy (PiFM), the tip senses the gradient forces arising from infrared-induced dipole interactions.5 PiFM, however, offers enhanced spatial resolution down to ∼5 nm and operates under ambient or controlled environmental conditions without the need for external optical detection, making it especially promising for in situ bioanalysis.6–8These sophisticated tools allow new insights into questions that have previously been very difficult to address. For example, most IR studies on biological specimens rely on chemical fixing, typically by using formaldehyde, paraformaldehyde, or glutaraldehyde to immobilize whole cell and sub-cellular structures before measurement. These aldehyde-based fixatives stabilize proteins and nucleic acids by forming methylene bridges and crosslinks between amine, thiol, and hydroxyl groups. While fixing preserves cell morphology,9 it fundamentally alters the chemical structure and, in turn, vibrational signature of biomolecules, the very parameters that IR spectroscopy seeks to probe. Formaldehyde, for instance, can shift or suppress amide I/II bands by crosslinking lysine and arginine residues, while glutaraldehyde contributes its own strong C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O and C–N stretching bands, complicating spectral interpretation.10 Furthermore, fixing halts enzymatic and metabolic processes, removes labile metabolites, and disrupts lipid–protein interactions. When combined with subsequent dehydration, which changes hydrogen bonding and protein secondary structure, these steps can dramatically distort the molecular fingerprint of the cell's chemical composition. Thus, fixing protocols, though convenient for stability during AFM imaging, are incompatible with the goal of observing original in situ cell states at the nanoscale.
O and C–N stretching bands, complicating spectral interpretation.10 Furthermore, fixing halts enzymatic and metabolic processes, removes labile metabolites, and disrupts lipid–protein interactions. When combined with subsequent dehydration, which changes hydrogen bonding and protein secondary structure, these steps can dramatically distort the molecular fingerprint of the cell's chemical composition. Thus, fixing protocols, though convenient for stability during AFM imaging, are incompatible with the goal of observing original in situ cell states at the nanoscale.
Recent advances have sought to overcome these limitations by imaging hydrated or even living cells using infrared s-SNOM. The most significant demonstration to date is that of Veber et al.11 who achieved nano-infrared imaging and spectroscopy of animal cells in a liquid environment. Their approach employed synchrotron-based broadband s-SNOM through ultrathin silicon carbide (SiC) membranes. Mouse fibroblasts were cultured on 30 nm SiC windows, allowing the near-field probe to interrogate the cell–membrane interface without direct contact. Broadband synchrotron radiation enabled collection of IR spectra across 2100–1050 cm−1, revealing sub-micrometre chemical heterogeneity in proteins, lipids, and DNA. While this represents a major advance, achieving in-liquid nanoscale spectroscopy of viable cells, it also underscores the technical and conceptual limitations of current approaches. The requirement for a synchrotron light source, complex window engineering, and non-standard tip–sample mechanics restricts accessibility and reproducibility. The effective probing depth is limited by the window thickness and refractive losses, while the water absorption cutoff (∼1050 cm−1) excludes key biochemical bands. Moreover, although mouse fibroblasts were examined under buffer and culture medium, the technique probed mainly the SiC window-adjacent region, not the true cellular interior, and depended on fixed reference conditions. The study also confirmed that even paraformaldehyde fixing, employed in part of the experiment to circumvent fibroblast detachment, produced measurable spectral distortions in amide and phosphate bands, consistent with chemical crosslinking effects observed in traditional FTIR studies.
These findings illustrate both the potential and the persistent challenges of nanoscale IR imaging in biological contexts. True in situ chemical mapping of live, unfixed cells remains elusive, owing to the twin difficulties of maintaining physiological hydration and achieving morphological and mechanical stability at the nanometre scale. To preserve native chemical states, it is essential to maintain a controlled humidity environment that supports cellular integrity without introducing bulk water absorption or water–membrane interfaces. This intermediate regime, between desiccation and immersion, has been largely unexplored in IR nano-spectroscopy.
In this study, we use a PiFM system coupled with a precision humidity control module allowing stable operation at a defined water vapour content while minimizing condensation. For the first time, we demonstrate PiFM based nano-vibrational spectroscopy on living stem cells and extracellular vesicles stabilised on cleaved mica surfaces. By avoiding aldehyde fixing entirely, this configuration preserves endogenous protein, lipid, and nucleic acid signatures, enabling authentic nanoscale chemical mapping of living cells and their sub-structures.
Methods
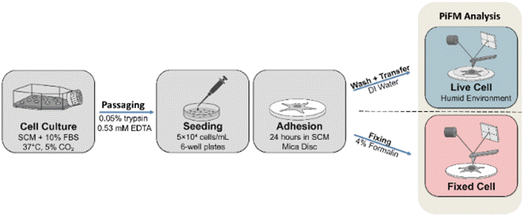
Oral mucosa lamina propria progenitor cells (OMLP-PCs) were cultured according to the protocol described by Knight et al.12 Cells were maintained in serum-containing medium (SCM) composed of Dulbecco's Modified Eagle's Medium (DMEM) supplemented with 2 mM L-glutamine, antibiotics/antimycotics (100 U per mL penicillin G, 100 µg per mL streptomycin sulfate, and 0.25 µg per mL amphotericin-B), and 10% fetal bovine serum (FBS). Cultures were incubated at 37 °C in a humidified atmosphere containing 5% CO2. For all experimental procedures, OMLP-PCs were passaged using 0.05% trypsin/0.53 mM ethylenediaminetetraacetic acid (EDTA) and counted using a hemocytometer. Cells were seeded onto cleaved mica discs placed in 6-well plates at a density of 5 × 104 cells per mL in 2 mL of SCM and allowed to adhere for 24 hours. Following OMLP-PC adhesion, half the mica disks were fixed with 4% (v/v) formalin before analysis, whilst live-cell samples were maintained in the growth medium immediately before analysis. Mica discs with adherent living OMLP-PCs were gently washed with deionised water and transferred to the humid environment for PiFM analysis. Fig. 1 shows the workflow of OMLP-PC culture, seeding and adherence to mica prior to PiFM analysis in either fixed or live cell states.OMLP-PCs were selected because of their widespread use in biological research, largely due to the relative ease of isolation, preparation, and culture. In addition to their practicality, these OMLP-PCs possess well-documented antimicrobial properties and have been investigated for their potential application as therapeutic stem cells. OMLP-PCs typically exhibit a fibroblast-like morphology, with characteristic dimensions ranging from approximately 20–60 µm. They are well characterised in the literature and are known to produce extracellular vesicles (EVs) with diameters in the range of 30–200 nm.13,14 These EVs fall within the optimal size range for analysis in the present study, particularly if generated in situ during imaging.
PiFM analysis
Imaging and spectral data were acquired using a Vista One nano-IR microscope and spectrometer (Molecular Vista Inc., USA), equipped with a Bloc 1930–780 cm−1 quantum-cascade laser (QCL) and Vistascan Version 28 software (Molecular Vista Inc., USA). Measurements utilized non-contact high-resolution platinum–iridium alloy (PtIr) coated Non-Contact High resonance Reflective coating (NCHR) cantilevers (Molecular Vista Inc., USA), calibrated by the manufacturer to a spring constant of 45 N m−1 and a resonance frequency of 335 kHz. Imaging and spectroscopy of live OMLP-PCs were conducted via PiFM on an anti-vibrational stage within an environmental enclosure coupled to a humidifier that maintained a relative humidity (RH) of 68–72%.PiFM can be operated in various dynamic AFM modes, with homodyne (direct detection) and heterodyne (sideband detection) being the modes most used.5 These two modes emphasize different components of the photoinduced force: sideband detection mode exclusively probes the gradient force, which is highly localized and minimizes far-field background contributions. This maximization of the steeply decaying localized force defines the characteristic high spatial resolution (as low as several nanometres) and restricts the measurement volume to the immediate vicinity of the tip, thus inherently limiting the practical depth probed to an estimated 20–30 nm. Direct detection on the other hand exhibits a much shallower dependence on the tip–sample distance and is estimated to sample a depth of ∼200 nm. We refer to the two different modes as “surface” and “bulk” respectively.
Experimental setup
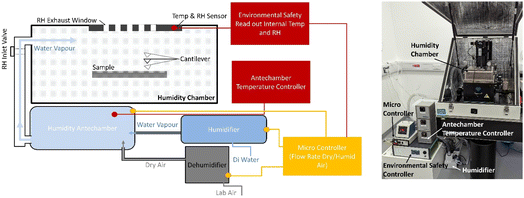
Live-cell PiFM imaging of OMLP-PCs was performed under controlled humidity conditions using an Electro-Tech Systems (ETS) humidity control system, Fig. 2. Setup included a Self-Regenerating Dehumidifier (Model 5478), Humidifier (Model 5482), and Microcontroller (Model 5100), integrated via an Anfatec Environmental & Safety Control Unit. Compressed air (80–100 PSI) was supplied to the dehumidifier and humidifier to generate dry and humidified air streams, which were mixed in the relative humidity (RH) control antechamber. This chamber, equipped with an internal mixing fan and temperature-controlled heater, ensured uniform humid air delivery to the specimen through the RH gate valve. The exhaust was vented through the relief valve on the opposite side of the environmental enclosure.The target RH was set to 70 ± 2%, maintained by automated feedback from the chamber humidity and temperature sensors. The antechamber temperature was held 1 °C below the microscope enclosure temperature to prevent condensation. The system was equilibrated for at least 15 min prior to imaging to ensure environmental stability with cantilever tuning conducted under these conditions. Throughout the imaging, no condensation was observed on the cantilever, optical path, or chamber walls. Following each experiment, the chamber was flushed with dry air for 10 min to restore ambient RH conditions before removal of samples.
Preparation of small extracellular vesicles (sEVs)
Briefly, gold-patterned silicon wafers were fabricated via photoresist patterning and electron-beam deposition to amplify PiFM signal strength.15,16 Surfaces were functionalised with 0.1% 3-aminopropyl triethoxysilane (APTES) in de-ionised water to immobilise the targets. Small extracellular vesicles (sEVs) from human bone marrow stromal cells (hBMSCs) were isolated by ultracentrifugation17 and confirmed to be sEVs using nanoparticle tracking analysis, cryo-electron microscopy, flow cytometry and western blotting. sEVs were incubated on modified and functionalised surfaces for 1 hour in air at room temperature before being rinsed three times with ultrapure water before analysis.Results and discussion
PiFM imaging and spectroscopy of live and fixed cells
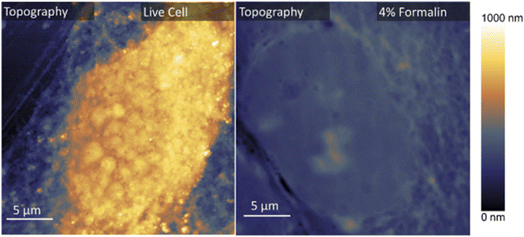
Fig. 3 presents the topography reported by AFM of live (left) and fixed (right) OMLP-PCs. Within the fixed cell, structures which may be attributed to the cell nucleus can be distinguished in the topography caused by the collapse of the cell wall onto the nucleus. Similar effects are not seen in the live cell suggesting such a collapse has not happened as is also indicated by the height difference between the two situations.Quantitative analysis of the AFM topography in Fig. 3 reveals distinct morphological differences between live and fixed cells. The cell wall over the nucleus measured 18.3 µm in width for live OMLP-PC cells, whereas the corresponding width in formalin-fixed cells was 15.6 µm. This represents an approximate 15% reduction following fixing. This is consistent with previous reports showing that aldehyde fixing induces compaction of intracellular components and reduces nuclear dimensions due to covalent cross-linking of proteins and dehydration effects.18 Formaldehyde-induced cross-linking, coupled with enhanced water loss relative to ambient conditions, leads to stiffening of the cytoskeletal network. This process constrains both nuclear and cytoplasmic volume.19
Surface roughness root mean square (RMS) measurement revealed a substantial decrease from 238 nm in the live OMLP-PC nucleus to 60 nm following fixing, indicating a ∼75% loss of the natural membrane topography. Similar alterations in surface complexity have been observed in AFM studies of whole human corneal, endometrial epithelial and rabbit corneal fibroblasts, where live-cell imaging showed well-defined membrane protrusions and ruffling that disappeared after being fixed.20,21 The smoother surface profile observed here post-fixing therefore reflects the collapse of dynamic cytoskeletal protrusions and the suppression of nuclear membrane undulations, as fixing restricts molecular mobility and eliminates inherent structural fluctuations.19,22
Similarly, the height ranges from the mica substrate to the OMLP-PC nucleus apex decreased markedly from ∼1110 nm (live) to ∼496 nm (fixed), reflecting the combined effects of dehydration and cross-linking, flattening the nuclear roughness and form. Previous AFM analyses of living mesenchymal stem cells have shown numerous protrusions of 20–300 nm in height that are largely absent or collapsed after fixing, confirming that chemical processing leads to volumetric reduction and diminished height variance in cellular substructures.22 In addition, AFM studies of fixed human endometrial cancer cells imaged in air have reported similar apparent reductions in whole cell height, which were attributed to dehydration and loss of turgor pressure.23 While this was addressed by capturing replica impressions of cells using a bioimprint technique, this approach prevents direct interrogation of cell surface and subsurface chemical features.
Collectively, these results demonstrate that fixing cells with formalin induces pronounced nanoscale and microscale morphological alterations, including nuclear contraction, loss of membrane roughness/patterning, and overall flattening of the cell body. These changes arise from aldehyde-mediated cross-linking of proteins, complete dehydration-induced collapse, and suppression of dynamic membrane activity. Consequently, fixed-cell topography measurements are likely to underestimate the true structural complexity of living OMLP-PCs, emphasizing the importance of live-cell AFM imaging for accurate morphological, chemical and even mechanical characterisation.
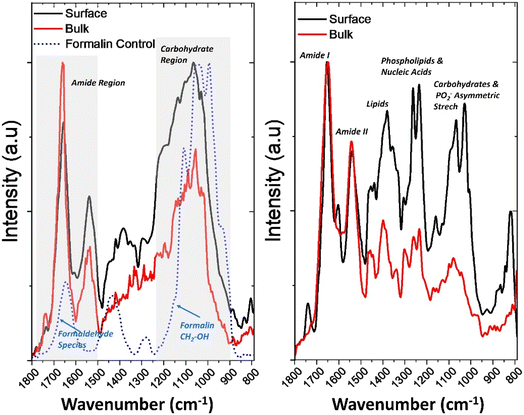
Fig. 4 presents the corresponding bulk and surface spectra of the OMLP-PC nucleus for formalin-fixed (left) and live (right) cells together with a reference spectrum for formalin (blue dotted line24). The fixed OMLP-PC nucleus spectrum shows a broad band near 1630–1650 cm−1 corresponding to C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching coupled with C–O and O–H deformation modes of hydrated formaldehyde species such as methylene glycol. Bands at 1470–1380 cm−1 correspond to CH2 bending vibrations within polymerized and cross-linked formaldehyde structures. The region between 1200 cm−1 and 1100 cm−1 contains C–O and C–O–H stretching vibrations from methylene glycol and polyoxymethylene linkages, while weaker C–O–C and CH2 rocking modes appear below 1000 cm−1. These features indicate the presence of methylene (–CH2–) and hydroxymethyl (–CH2OH) functional groups responsible for aldehyde crosslinking during fixing. The figure shows that in both the fixed and living cell nuclei, distinct vibrational bands corresponding to amide I (∼1650 cm−1) and amide II (∼1550 cm−1) were detected, representing the characteristic protein backbone vibrations. However, in the formalin-fixed cell nucleus, both the surface and bulk spectra's amide I region overlaps with the C
O stretching coupled with C–O and O–H deformation modes of hydrated formaldehyde species such as methylene glycol. Bands at 1470–1380 cm−1 correspond to CH2 bending vibrations within polymerized and cross-linked formaldehyde structures. The region between 1200 cm−1 and 1100 cm−1 contains C–O and C–O–H stretching vibrations from methylene glycol and polyoxymethylene linkages, while weaker C–O–C and CH2 rocking modes appear below 1000 cm−1. These features indicate the presence of methylene (–CH2–) and hydroxymethyl (–CH2OH) functional groups responsible for aldehyde crosslinking during fixing. The figure shows that in both the fixed and living cell nuclei, distinct vibrational bands corresponding to amide I (∼1650 cm−1) and amide II (∼1550 cm−1) were detected, representing the characteristic protein backbone vibrations. However, in the formalin-fixed cell nucleus, both the surface and bulk spectra's amide I region overlaps with the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O of the formaldehyde resulting in a slight broadening and shift of the spectra. There is also a shift to lower wavenumbers of the amide I, not seen in the live cell nucleus which could be attributed to a greater contribution of formalin during surface fixing. The formalin-fixed spectra also show a loss of signal in the 1500–1200 cm−1 region. These bands are typically associated with amide II (N–H bending and C–N stretching of proteins), CH2 and CH3 deformation vibrations from lipids and proteins, symmetric COO− stretching of amino acid side chains, amide III and PO2− asymmetric stretching linked to proteins and nucleic acids. Perhaps the most significantly impacted region of the spectrum is where features appearing between 1200–1100 cm−1 overlap with the formalin FTIR reference spectrum. These are thought to originate from the methylene bridge (–CH2–) vibrations arising from fixing with formalin.24 These overlap with C–O and C–O–H stretching vibrations of carbohydrates and the nucleic acid ribose, as well as PO2− symmetric and asymmetric stretching modes of DNA and RNA, making it difficult to distinguish native cellular phosphate and carbohydrate signals from fixative-derived contributions. Table 1 summarises the spectral overlap between organic samples and formalin. Further research is needed to investigate the impact of biological fixing agents commonly used in traditional imaging when applied to spectral imaging, as preserving only topography and morphology is not sufficient for these emerging techniques.
O of the formaldehyde resulting in a slight broadening and shift of the spectra. There is also a shift to lower wavenumbers of the amide I, not seen in the live cell nucleus which could be attributed to a greater contribution of formalin during surface fixing. The formalin-fixed spectra also show a loss of signal in the 1500–1200 cm−1 region. These bands are typically associated with amide II (N–H bending and C–N stretching of proteins), CH2 and CH3 deformation vibrations from lipids and proteins, symmetric COO− stretching of amino acid side chains, amide III and PO2− asymmetric stretching linked to proteins and nucleic acids. Perhaps the most significantly impacted region of the spectrum is where features appearing between 1200–1100 cm−1 overlap with the formalin FTIR reference spectrum. These are thought to originate from the methylene bridge (–CH2–) vibrations arising from fixing with formalin.24 These overlap with C–O and C–O–H stretching vibrations of carbohydrates and the nucleic acid ribose, as well as PO2− symmetric and asymmetric stretching modes of DNA and RNA, making it difficult to distinguish native cellular phosphate and carbohydrate signals from fixative-derived contributions. Table 1 summarises the spectral overlap between organic samples and formalin. Further research is needed to investigate the impact of biological fixing agents commonly used in traditional imaging when applied to spectral imaging, as preserving only topography and morphology is not sufficient for these emerging techniques.
| Wavenumber/cm−1 | Formalin peak | Overlapping band/cm−1 | Biochemical origin |
|---|---|---|---|
| 1745–1700 | Weak/shoulder carbonyl features from hydrated/formyl/poly-oxymethylene species (depending on formulation) | 1745–1700 | Ester C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching (phospholipids, fatty-acid esters) O stretching (phospholipids, fatty-acid esters) |
| 1685–1630 | Broad band from hydrated formaldehyde species (coupled C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O/C–O/O–H modes) O/C–O/O–H modes) |
∼1650 | Amide I (protein C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, secondary structure) O, secondary structure) |
| 1470–1380 | –CH2– bending/deformation from formaldehyde polymers & methylene bridges | 1452–1388 | CH2/CH3 deformations (lipids, proteins) |
| 1330–1280 | C–O/C–O–H deformation with CH2 wagging (hydrated formaldehyde species) | 1330–1284 | Amide III (protein) |
| 1240–1220 | C–O–C/C–O stretching of methylene-glycol/polyoxymethylene linkages | 1244–1230 | PO2− asymmetric stretch (nucleic acids; some phospholipid contribution) |
| 1156–1120 | C–O–C antisymmetric stretching (formaldehyde polymers) | 1152–1124 | CO–O–C antisymmetric stretch (glycogen; nucleic-acid ribose) |
| 1100–1080 | C–O stretching/CH2 rocking (polyoxymethylene) | 1090–1084 | PO2− symmetric stretch (DNA/RNA) |
| 1060–1040 | C–O stretching of –CH2OH groups | 1060–1045 | C–O stretching (carbohydrates; deoxyribose/ribose) |
The live-OMLP-PC nucleus PiFM spectra closely resemble those reported for whole stem cells in the literature, particularly FTIR profiles described by Aksoy et al.25 The main protein (amide I and II), phosphate, and carbohydrate bands occur at comparable positions, confirming that both techniques capture key biochemical signatures of live cells. However, notable differences are evident in relative band intensities, most prominently within the surface spectra, where lipid- and carbohydrate-associated peaks (∼1735, 1465, and 1030 cm−1) appear enhanced compared with the broader, protein-dominated FTIR profile. These variations primarily arise from the contrasting spatial resolution and sampling volume between the two methods: FTIR probes micrometre-scale regions, effectively averaging signal from the entire cell, whereas PiFM confines its interaction volume to a few tens of nanometres. The result is a spectrum with sharper, higher-contrast features that reflect local surface chemistry rather than whole cell composition. Consequently, PiFM reveals nanoscale heterogeneity that is intrinsically masked in conventional FTIR measurements. The surface spectra in Fig. 4 show enhanced bands at ∼1735 cm−1 (ester C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), ∼1465 cm−1 (CH2 scissoring), and ∼1030 cm−1 (C–O stretching), consistent with lipid- and glycoprotein-rich plasma membrane and glycocalyx. In contrast, the bulk spectra display stronger amide I and II bands (∼1650 cm−1, ∼1545 cm−1) and pronounced phosphate vibrations (∼1240 cm−1, ∼1080 cm−1), reflecting a greater contribution from intracellular proteins and nucleic acids within the near-surface cytoplasm. Comprehensive analysis of the full IR-spectrum enables detailed characterisation of local heterogeneity and facilitates the isolation of specific wavenumbers or characteristic peaks relevant to micro-to-nanoscale surface mapping. Overlaying topographical maps with selected wavenumber combinations further enhances visualisation of the spatial distribution of chemical heterogeneity shown in Fig. 5 with corresponding IR spectra of areas of interest (P1–P3) shown in Fig. 6.
O), ∼1465 cm−1 (CH2 scissoring), and ∼1030 cm−1 (C–O stretching), consistent with lipid- and glycoprotein-rich plasma membrane and glycocalyx. In contrast, the bulk spectra display stronger amide I and II bands (∼1650 cm−1, ∼1545 cm−1) and pronounced phosphate vibrations (∼1240 cm−1, ∼1080 cm−1), reflecting a greater contribution from intracellular proteins and nucleic acids within the near-surface cytoplasm. Comprehensive analysis of the full IR-spectrum enables detailed characterisation of local heterogeneity and facilitates the isolation of specific wavenumbers or characteristic peaks relevant to micro-to-nanoscale surface mapping. Overlaying topographical maps with selected wavenumber combinations further enhances visualisation of the spatial distribution of chemical heterogeneity shown in Fig. 5 with corresponding IR spectra of areas of interest (P1–P3) shown in Fig. 6.
 | ||
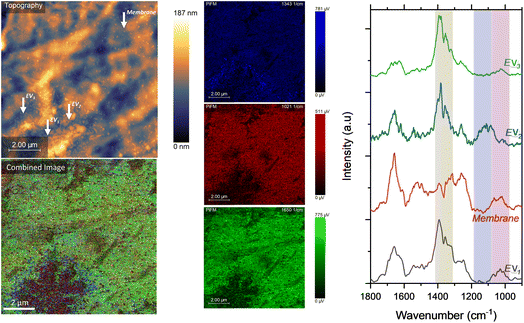
| Fig. 5 Combined spectral maps (left) showing the spatial distribution of key molecular vibrations acquired by photoinduced force microscopy (PiFM) using sideband detection. Individual channels correspond to amide II (1542 cm−1, red), amide I (1653 cm−1, green), and CH2/CH3 deformation + COO− stretching modes (1389 cm−1, blue). Spectra recorded in positions labelled as P1, P2, and P3 are shown in Fig. 6. | ||
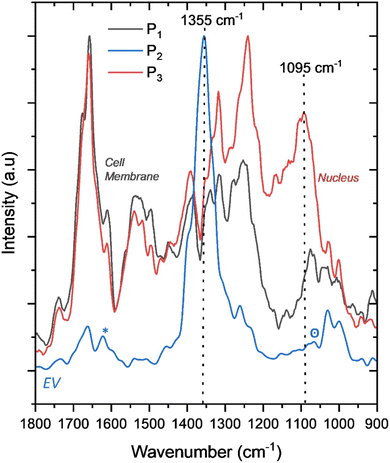 | ||
| Fig. 6 Representative PiFM spectra extracted from the locations indicated in Fig. 5 (P1–P3), corresponding to the cell membrane (P1, grey), nuclear region (P3, red), and extracellular vesicle (EV) (P2, blue). | ||
Prolonged PiFM imaging under QCL laser excitation proved challenging at higher laser powers, as localised heating and photothermal stress often led to cell death and the loss of tip–sample interactions. The hyperspectral imaging shown in Fig. 5, represents one of the most stable long-term live-cell acquisitions, performed at 1% QCL laser intensity under controlled humidity to minimise dehydration effects. Throughout hyperspectral imaging across multiple wavenumbers of interest, the OMLP-PC nucleus exhibited little to no morphological change.
The surface initially remained consistently smooth, suggesting a metabolically inactive or stressed state rather than active motion or proliferation. One notable feature is the prominent protrusion denoted as P3 in the lower half of the OMLP-PC nucleus in Fig. 5. This may be due to the cell nucleus being positioned unusually close to the surface, which can occur during ambient air exposure rather than under physiological liquid conditions. Such positioning can lead to cellular flattening and/or cytoskeletal reorganization, commonly associated with reduced metabolic activity. Some evidence to support this hypothesis comes from the spectroscopy discussed below.
As scanning progressed, several small features (∼150 nm in diameter and ∼15 nm in height) began to develop; seen here in region P2, Fig. 5. Since the cell is under considerable stress under these analysis conditions, it's possible that these features are extracellular vesicles (EVs). The production of EV's is known to increase under environmental and physical stress. Stress-induced EVs appear to function as a protective cellular response, transporting molecular cargo capable of modulating repair pathways, promoting cell survival, and influencing signalling processes in neighbouring cells.26 Typically, they contain membrane proteins (e.g., tetraspanins like CD9, CD63, CD81), cytosolic proteins, lipids and nucleic acids, especially small RNAs. They can also carry markers of cellular damage or stress and reflect the physiological state of the cell they originate from.
The surface-band PiFM spectra at positions P1, P2, and P3, Fig. 5, reveal distinct biochemical variations across different cellular regions. The spectra from P1, an area lacking distinct topographical features and spectroscopically like many other surface sites, which we interpret as the OMLP-PCs collapsed outer cell membrane, and P3, the OMLP-PC nucleus, are the easiest to interpret, showing pronounced differences in the 1200–1070 cm−1 region in Fig. 6. The broad increase in intensity observed in this range for the proposed nuclear spectrum (P3) is likely to correspond to phosphate (PO2−) stretching vibrations and carbohydrate-associated CO–O–C modes. Specifically, in Fig. 6, bands near 1244–1230 cm−1 can be attributed to antisymmetric PO2− stretching from nucleic acids, with minimal phospholipid contribution, while features around 1090–1084 cm−1 arise from symmetric PO2− stretching in RNA and DNA backbones. Additional contributions near 1147–1124 cm−1 are consistent with CO–O–C antisymmetric stretching vibrations of glycogen and nucleic acid ribose units. These combined signals reflect the nucleic acid-rich environment of the nucleus (P3) compared with the more lipid-dominated membrane region (P1), which exhibits comparatively weaker phosphate features and stronger CH2 deformation modes.
The spectrum at P2 is very different from P1 and P3 as it is dominated by a strong peak at 1355 cm−1. After subtracting the large peak at 1355 cm−1, the resulting profile closely resembles those of P1 (OMLP-PC membrane) and P3 (nucleus), displaying the characteristic amide I (∼1650 cm−1) and amide II (∼1550 cm−1) bands. Notably, a shoulder near 1625 cm−1 (marked by *) indicates subtle protein secondary structure rearrangements, possibly reflecting changes in β-sheet or aggregated conformations.
In contrast, P2 in Fig. 6, shows a marked reduction in intensity between 1075–1100 cm−1 (marked by °) compared to P1 and P3, suggesting a decreased contribution from PO2− symmetric stretching vibrations associated with nucleic acids (RNA/DNA) and from C–O stretching of carbohydrate moieties near 1060–1050 cm−1. The distinct peak at 1355 cm−1 corresponds to CH3 symmetric deformation and CH2 wagging vibrations of proteins and lipids, particularly from aliphatic side chains and membrane phospholipids. The pronounced intensity of this peak indicates a local accumulation of lipid and protein species, consistent with the membrane-rich composition characteristic of cell wall-derived extracellular vesicles.
The band position, located marginally below the characteristic CH3 bending maximum (1380–1370 cm−1), probably arises from overlapping C–N stretching modes of peptide backbones or minor spectral shifts due to differences in lipid chain packing. This is consistent with the presence of biological cargo encapsulated within the EVs. Closer examination of the nuclear region was carried out by collecting PiFM spectra across both the membrane and the nucleus, in surface and bulk modes, shown in Fig. 7. Surprisingly, the bulk spectra showed only minimal differences between the two regions. This limited contrast is unexpected, as the biochemical composition of the nucleus, rich in nucleic acids, nuclear proteins, and chromatin-associated phosphates, should, in principle, produce clear spectral distinctions. The most notable bulk-level difference was a more pronounced 1735 cm−1 shoulder over the nucleus, which is typically associated with ester C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching of lipids or lipid-bound proteins. This may reflect the presence of the nuclear envelope, which contains lipid bilayers distinct from the surrounding plasma membrane and can contribute strongly to this band.
O stretching of lipids or lipid-bound proteins. This may reflect the presence of the nuclear envelope, which contains lipid bilayers distinct from the surrounding plasma membrane and can contribute strongly to this band.
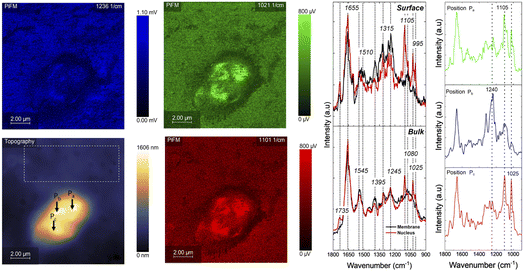 | ||
| Fig. 7 PiFM chemical mapping and spectra collected over the nuclear region (P3) of a live OMLP cell. Topographic and PiFM maps at 1236 cm−1, 1021 cm−1, and 1101 cm−1 show the spatial distribution of molecular signals across the nucleus. Spectra extracted from the nuclear and membrane regions reveal the expected amide I (1655 cm−1) and amide II (1545 cm−1) protein bands, along with features at 1395 cm−1 (CH deformation), 1245–1080 cm−1 (PO2− stretching from nucleic acids), and 1025–995 cm−1 (C–O stretching). Point spectra from positions Pa, Pb & Pc highlight local chemical differences within the nuclear region, with dominant peaks at 1105 cm−1, 1240 cm−1, and 1025 cm−1, indicating variations in phosphate and carbohydrate-related vibrational modes. The dashed box in the topography image identifies the area expanded in Fig. 8. | ||
In contrast, the surface-mode spectra, Fig. 7, reveal much clearer differences between the nucleus and membrane. The nuclear region shows enhanced or uniquely expressed bands at 1105 cm−1, 1025 cm−1, and 995 cm−1, all of which fall within the phosphate and C–O stretching region. These can be associated with DNA/RNA backbones, ribose vibrations, and chromatin-associated sugars. These same features are present in the bulk spectra albeit with reduced prominence, consistent with the smaller sampling volume of the surface mode and the fact that chromatin is located only a few nanometres beneath the nuclear envelope. The membrane, by comparison, exhibits a higher relative signal at 1080 cm−1, attributable to PO2− symmetric stretching from phospholipids and contributions from the glycocalyx, which is rich in carbohydrate-related C–O modes. The surface spectra also show higher relative intensity at 1315 cm−1 and 1395 cm−1 over the membrane compared with the nucleus. These bands correspond to CH2/CH3 deformation and C–N stretching associated with lipid chains and membrane-associated proteins shown in Table 1, consistent with the dense aliphatic environment of the plasma membrane. Their reduced intensity over the nucleus likely reflects the lower abundance of lipid-rich material at the nuclear surface.
Fig. 7 also shows maps of the PiFM signal intensity at 1236 cm−1, 1021 cm−1, and 1101 cm−1 in the nuclear region, revealing discrete hotspots of signal intensity within the nucleus labelled Pa, Pb and Pc. These are likely to correspond to areas of nuclear lobulations, containing chromocenters or highly compacted chromatin domains, caused by stresses induced on the nucleus itself.27 Nuclear lobulations are known to contain elevated concentrations of DNA, RNA, and DNA-binding proteins. Spectra extracted from Pa, Pb and Pc show broadly similar character (Fig. 7, right), supporting the interpretation that they represent chromatin-dense regions. However, small but consistent variations occur within the amide II region (∼1545–1510 cm−1), suggesting local differences in protein secondary structure or chromatin-associated histone content. The most pronounced local variation is observed at Pb, which displays a strong peak at 1240 cm−1. This band corresponds to PO2− antisymmetric stretching, a well-established marker of nucleic acids, and is typically enhanced in regions enriched in transcriptionally active chromatin, ribonucleoproteins, or nascent RNA. Thus, the spectral prominence at 1240 cm−1 may indicate a locally elevated nucleic-acid density or a region of increased chromatin activity within the nucleus.
Fig. 8 is an expansion of the boxed area adjacent to the nuclear region in the topography image in Fig. 7, and shows distinct membrane depressions. Cross sectional line-profiling shows that these are approximately 500–800 nm in width and 20–70 nm in depth. Although it is difficult to assign these structures unambiguously, several possibilities can be considered. One is that they could arise from local membrane instability or deformation induced by experimental conditions, such as mechanical loading from tapping-mode AFM and/or photothermal stress from the PiFM QCL illumination. Alternatively, these depressions bear a superficial resemblance to the pit–depression architectures described for fusion pores (porosomes) at the plasma membrane of secretory cells.28 However, their lateral dimensions are substantially larger than the ∼100–200 nm depressions typically associated with individual fusion pores, making a direct one-to-one correspondence unlikely. Consistent with this, PiFM spectra acquired from these selected areas, and shown in Fig. 7, do not reveal marked chemical differences compared with the surrounding membrane suggesting that, if these features are related to exocytotic machinery, they may represent areas of larger-scale membrane invagination or topographical perturbations rather than discrete, compositionally distinct fusion pore complexes.
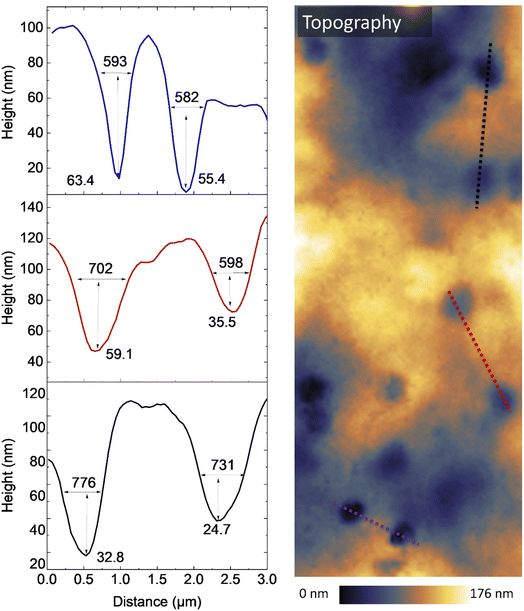 | ||
| Fig. 8 Topography and line-profile analysis of perinuclear membrane features. Topographical map (right) showing depressions adjacent to the nucleus, with dashed lines indicating the locations of extracted line profiles. The corresponding height profiles (left). The imaged region corresponds to the dotted box in Fig. 7. | ||
Fig. 9 focuses on the P2 region from Fig. 5 with a higher resolution scan. The spectral data reveals clear biochemical differences between the cell membrane and the EVs (EV1–EV3) located nearby, as well as notable variations among the EVs themselves. The cell membrane spectrum (red) shows broad amide I (1650 cm−1) and amide II (1550 cm−1) bands accompanied by strong intensity in the 1080–1020 cm−1 region. These can be associated with PO2− stretching vibrations from nucleic acids and phospholipids, as well as C–O–C modes from carbohydrates. This combination reflects the chemically diverse composition of the plasma membrane and its glycocalyx, where proteins, lipids, and surface polysaccharides coexist. In contrast, PiFM spectra from all three EVs display sharper and more localized features, highlighting their distinct biochemical makeup. The 1355 cm−1 band, assigned to CH3/CH2 deformation and C–N stretching vibrations of lipids and protein side chains, is markedly stronger in the EV spectra than in the membrane, confirming the lipid- and protein-rich nature of these vesicles. The EV3 spectrum shows the most intense 1355 cm−1 peak and a dominant amide I band, suggesting a highly ordered, protein- and lipid-dense composition with minimal phosphate or carbohydrate contribution. EV2 exhibits a similar overall profile but with a more pronounced shoulder near 1090 cm−1, indicative of PO2− symmetric stretching, suggesting partial enrichment in nucleic-acid or glycoprotein-associated material. EV1, by contrast, shows comparatively weaker amide I and CH deformation bands but enhanced intensity around 1020–1000 cm−1, consistent with C–O stretching vibrations from carbohydrate or glycan residues.
Collectively, these differences demonstrate that while all extracellular vesicles share a common protein–lipid spectral signature, their relative compositions vary, with EV3 being more protein- and lipid-dominant, EV2 showing mixed protein and phosphate/glycoprotein features, and EV1 enriched in carbohydrate-associated vibrations. Compared with the parent membrane, the EVs as a group display sharper spectral features and reduced nucleic-acid signatures, reflecting their chemically specialized and compositionally distinct microenvironments.
PiFM imaging and spectroscopy of single EVs
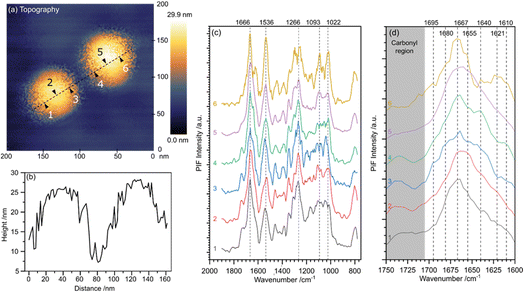
The imaging and spectroscopic characterisation of EVs developing on live cells presents significant challenges, owing to the dynamic nature of the cellular environment and interference from background cellular signals. Nevertheless, recent studies have demonstrated that nano-vibrational spectroscopy of isolated EVs can yield substantially enhanced molecular insights. Ramer et al. investigated exosomes derived from colorectal cancer cells immobilised on zinc selenide substrates,29 whereas our recent work on single small EVs (sEVs) derived from human bone marrow mesenchymal stromal cells (hBMSCs) and DU145 prostate cancer cells reveals30 that PiFM affords detailed analysis of protein secondary structures within individual vesicles.Fig. 10 presents PiFM imaging of two sEVs ∼100 nm in diameter, isolated from hBMSCs and which exhibited expression of the tetraspanin surface markers CD81, CD63, and CD9, as confirmed by flow cytometry. The sEVs were deposited on APTES-modified, gold-patterned substrates to amplify the PiFM signal through the Surface-Enhanced Infrared Absorption (SEIRA) effect, achieving a PiFM signal intensity enhancement of ∼50%. Spectra recorded from the specific locations indicated are shown in Fig. 10(c), with the protein spectral region expanded in Fig. 10(d).
Across all the collected spectra, characteristic bands can be attributed to lipids, phospholipids, and triglycerides within the 1800–1700 cm−1 range, and to amide I and II vibrations between 1700–1500 cm−1. Additional peaks between 1400–1200 cm−1 correspond to methylene bending modes centred near 1260 cm−1, while phosphate bending vibrations from the DNA/RNA backbone are observed at approximately 1240 cm−1 and 1085 cm−1.
Fig. 10(d), expands the protein spectral region, revealing significant variation in the ratio of the amide I (1700–1600 cm−1) to amide II (1600–1500 cm−1) bands, particularly between positions 5 and 6. Key components are listed in Table 2, and compared with values previously reported in the literature.3 This variation indicates heterogeneity in protein structure and function within individual sEVs. Further differences in secondary structure are revealed through deconvolution of the amide I region. For example, the spectrum at position 5 shows a dominant feature at 1655–1650 cm−1, consistent with α-helical conformations, whereas position 4 from the same vesicle exhibits a shoulder at 1640–1620 cm−1, characteristic of β-sheet structures. These structural variations underpin functional diversity and may also serve as potential diagnostic or disease-related biomarkers.
| Secondary structure | PiFM peaks/cm−1 | Literature range/cm−1 |
|---|---|---|
| β-sheet (H-bonded) | 1610, 1620 | 1605–1640 |
| Disordered/random coil | 1640 | 1635–1650 |
| α-helix | 1655 | 1648–1662 |
| β-turns | 1665 | 1660–1690 |
| β-sheet (antiparallel) | 1680, 1695 | 1680–1700 |
Conclusions
Here we have demonstrated the capability of PiFM for nanoscale chemical mapping of complex biological systems, bridging the gap between molecular specificity and spatial resolution. Our findings highlight two specific aspects: first, fixing with aldehyde-based agents such as formalin significantly alters cellular chemistry, introducing spectral overlaps that compromise accurate interpretation. This underscores the need for rigorous evaluation of fixing protocols before widespread adoption of nano-vibrational spectroscopy of fixed specimens. Second, live-cell PiFM imaging under controlled humidity enables authentic characterisation of native biomolecular states and dynamic processes, including EV formation. Furthermore, PiFM analysis of isolated EVs reveals compositional heterogeneity at the single-vesicle level, offering a powerful route to build biochemical libraries and identify disease-related biomarkers.Looking to the future, systematic studies are needed on the chemical impact of different fixing agents to establish best practices for nanoscale vibrational imaging. This will ensure accurate interpretation when fixing biological specimens is unavoidable. In addition, advancing live-cell PiFM requires improved environmental control and low-power imaging strategies to minimize photothermal stress, enabling longer acquisition times and dynamic process monitoring. Furthermore, expanding PiFM-based extracellular vesicle (EV) profiling into large-scale libraries has the potential to uncover compositional signatures linked to disease states, paving the way for its use in diagnostic and therapeutic applications. This advance is likely to benefit from integration with machine learning approaches for spectral classification and correlation with omics data. Finally, combining PiFM with complementary techniques such as AFM-IR or tip enhanced Raman spectroscopy would deliver multimodal insights into cellular architecture and function, ensuring the adoption of nano-vibrational spectroscopy as a cornerstone tool in nanobiology, biomedical research and diagnostics.
Ethical considerations
Local research ethical committee approval had been previously obtained for the isolation of the oral cells utilised in this study. Donors provided informed consent for the use of their materials. Small extracellular vesicles arising from bone marrow stem cells were purchased from commercially available sources.Author contributions
Davies-Jones (conceptualization, investigation, methodology, formal analysis, writing); Pattem (formal analysis, writing); Azizova (investigation, methodology, formal analysis, writing); Al-Luaibi (investigation, methodology); Morgan (writing); Stephens (methodology, resources); Bo Hou (resources, methodology, supervision, writing); Clayton (resources, methodology); Nishio (resources, methodology, supervision, writing); Davies (data curation, funding acquisition, project administration, resources, supervision, writing).Conflicts of interest
There are no conflicts to declare.Data availability
The dataset supporting the findings of this study is available on Figshare at https://doi.org/10.6084/m9.figshare.30600569.Acknowledgements
The PiFM spectrometer was acquired with the EPSRC grant EP/V05399X/1. Small extracellular vesicles were acquired with the BBSRC grant BB/X004449/1. The European Regional Development Fund (ERDF) and the Welsh European Funding Office (WEFO) part-funded the Cardiff Catalysis Institute Microscopy facility. This manuscript was produced with the assistance of a large language model (LLM).Notes and references
- A. Centrone, Annu. Rev. Anal. Chem., 2015, 8, 101–126 CrossRef CAS PubMed.
- D. Kurouski, A. Dazzi, R. Zenobi and A. Centrone, Chem. Soc. Rev., 2020, 49, 3315–3347 RSC.
- S. Banerjee, B. Holcombe, S. Ringold, A. Foes, T. Naik, D. Baghel and A. Ghosh, J. Phys. Chem. B, 2022, 126, 5832–5841 CrossRef CAS PubMed.
- A. C. V. D. dos Santos, N. Hondl, V. Ramos-Garcia, J. Kuligowski, B. Lendl and G. Ramer, ACS Meas. Sci. Au, 2023, 3, 301–314 CrossRef PubMed.
- M. R. Shcherbakov, E. O. Potma, Y. Sugawara, D. Nowak, M. Stepanova, P. R. Davies, J. Davies-Jones and H. K. Wickramasinghe, Nat. Rev. Methods Primer, 2025, 5, 34 CrossRef CAS.
- A. A. Sifat, J. Jahng and E. O. Potma, Chem. Soc. Rev., 2022, 51, 4208–4222 RSC.
- J. A. Davies-Jones and P. R. Davies, Mater. Chem. Front., 2022, 6, 1552–1573 RSC.
- L. M. Otter, M. W. Förster, E. Belousova, P. O'Reilly, D. Nowak, S. Park, S. Clark, S. F. Foley and D. E. Jacob, Geostand. Geoanal. Res., 2021, 45, 5–27 CrossRef CAS.
- Y. Chao and T. Zhang, Appl. Microbiol. Biotechnol., 2011, 92, 381–392 CrossRef CAS PubMed.
- R. Li and H. C. Allen, Sci. Rep., 2025, 15, 22944 CrossRef PubMed.
- A. Veber, C. Spedalieri and J. Kneipp, Small, 2025, 21, e07097 CrossRef CAS PubMed.
- R. Knight, E. Board-Davies, H. Brown, A. Clayton, T. Davis, B. Karatas, J. Burston, Z. Tabi, J. M. Falcon-Perez, S. Paisey and P. Stephens, Stem Cells Transl. Med., 2022, 11, 861–875 CrossRef CAS PubMed.
- B. M. Borowiec, M. Dyszkiewicz-Konwińska, D. Bukowska, M. Nowicki and J. Budna-Tukan, Cells, 2024, 13, 1514 CrossRef CAS PubMed.
- Y. Yang, R. Knight, P. Stephens and Y. Zhang, J. Oral Pathol. Med., 2020, 49, 342–349 CrossRef CAS PubMed.
- S. Zhan, et al., Phys. Status Solidi A, 2024, 217, 1900832 CrossRef.
- T. Chen, S. Zhan, B. Li, B. Hou and H. Zhou, Adv. Opt. Mater., 2020, 2302451 Search PubMed.
- C. Théry, S. Amigorena, G. Raposo and A. Clayton, Curr. Protoc. Cell Biol., 2006, 30, 3.22.1–3.22.29 Search PubMed.
- P. Nienałtowski, M. Baczewska and M. Kujawińska, Photonics Lett. Pol., 2020, 12, 13–15 CrossRef.
- K. W. Kim, Appl. Microsc., 2020, 50, 10 CrossRef CAS PubMed.
- K. Sinniah, J. Paauw and J. Ubels, Curr. Eye Res., 2002, 25, 61–68 CrossRef PubMed.
- L. W. Francis, D. Gonzalez, T. Ryder, K. Baer, M. Rees, J. O. White, R. S. Conlan and C. J. Wright, J. Microsc., 2010, 240, 111–121 CrossRef CAS PubMed.
- S. Casado, M. d. V. T. Lobo and C. L. Paíno, Sci. Rep., 2017, 7, 6767 CrossRef PubMed.
- J. Muys, M. Alkaisi, D. Melville, J. Nagase, P. Sykes, G. Parguez and J. Evans, J. Nanobiotechnol., 2006, 4, 1 CrossRef CAS PubMed.
- E. Gazi, J. Dwyer, N. P. Lockyer, J. Miyan, P. Gardner, C. Hart, M. Brown and N. W. Clarke, Biopolymers, 2005, 77, 18–30 CrossRef CAS PubMed.
- C. Aksoy and F. Severcan, Infrared Spectroscopy and Imaging in Stem Cells and Aging Research, Springer New York, New York, NY, 2019, vol. 2045 Search PubMed.
- J. Hahm, J. Kim and J. Park, Tissue Eng. Regener. Med., 2021, 18, 513–524 CrossRef CAS PubMed.
- J. T. Butler, L. L. Hall, K. P. Smith and J. B. Lawrence, J. Cell. Biochem., 2009, 107, 609–621 CrossRef CAS PubMed.
- B. P. Jena, J. Cell. Mol. Med., 2004, 8, 1–21 CrossRef CAS PubMed.
- N. Hondl, L. Neubauer, V. Ramos-Garcia, J. Kuligowski, M. Bishara, E. Sevcsik, B. Lendl and G. Ramer, ACS Meas. Sci. Au, 2025, 5, 469–476 CrossRef CAS PubMed.
- L. Azizova, J. Davies-Jones, L. Dribika, D. Othman, U. Borucu, I. Honeyborne, D. Morgan, B. Hou, A. Clayton, P. R. Davies and W. Nishio, Prep.
| This journal is © The Royal Society of Chemistry 2026 |