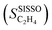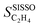Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Rethinking catalysis: interpretable AI and description of real-world conditions via materials genes
Lucas Foppa *ab and
Matthias Scheffler
*ab and
Matthias Scheffler a
a
aThe NOMAD Laboratory at the Fritz Haber Institute of the Max Planck Society, Berlin, Germany. E-mail: foppa@fhi-berlin.mpg.de
bMolecular Simulations from First Principles e.V., Berlin, Germany
First published on 6th February 2026
Abstract
Descriptors link basic physicochemical parameters that characterize the materials and the environment to the catalytic performance. Traditionally, descriptors are rooted in mechanistic understanding of elementary surface reactions gained from surface science and atomistic simulations on well-defined surfaces and under vacuum. However, real-world catalysis operates under elevated pressures and temperatures, where an intricate interplay of multiple physical processes, including significant materials' restructuring and transport phenomena, governs performance. To bridge this gap, we introduced an interpretable artificial intelligence (AI) approach that identifies key physicochemical parameters correlated with the measured catalytic performance. Analogous to genes in biology and medicine, these “materials genes” provide a statistical description of catalysis without requiring the explicit atomistic description of the underlying physical processes. Here, we combine the sure-independence-screening-and-sparsifying-operator (SISSO) symbolic-regression AI approach with a sensitivity analysis based on partial derivatives to determine the most influential genes needed to describe the selectivity of supported palladium-based metal alloy nanoparticles in the hydrogenation of concentrated acetylene streams. The identified genes include the calculated average d-band center and the measured average particle diameter, indicating the crucial role of adsorption and structure sensitivity on the formation of ethylene.
1 Introduction
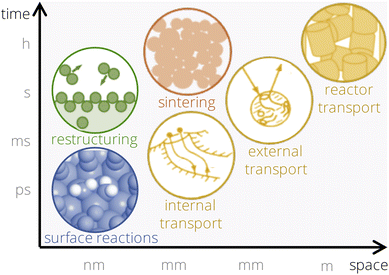
Progress in catalyst design depends on identifying basic physicochemical parameters that characterize materials and correlate with their catalytic performance,1,2 a task complicated by the extensive range of variables that can be tuned for creating a new material. These parameters, or functions thereof, establish quantitative relationships with catalytic-performance metrics such as activity or selectivity. We refer to the functions of these physicochemical parameters as descriptors hereafter. Traditionally, descriptors are rooted in fundamental understanding of the physical processes governing catalysis. Surface science3–8 and ab initio atomistic simulations9–11 provide detailed information on adsorption and elementary reactions occurring on well-defined model surfaces under vacuum and low temperatures. This knowledge led to descriptors based on adsorption energies of surface intermediates involved in important steps of the catalytic cycle.12 Adsorption energies offer an intuitive picture of heterogeneous catalysis, as they embody the Sabatier principle13 of optimal binding strength. They also relate to activation energy barriers through Brønsted–Evans–Polanyi relationships14,15 and to basic electronic properties of the material through models such as the d-band theory.16–18 The use of adsorption-energy-based descriptors is convenient for screening new materials,19 since such quantities can be computed using density functional theory (DFT) calculations. Machine-learning interatomic potentials trained on DFT data further accelerated the screening of materials based on adsorption-energy descriptors.20 However, descriptors derived from idealized model systems such as adsorption energies are often insufficient to describe the catalytic performance observed under realistic, high-performance operating conditions, characterized by high pressures and temperatures.21 This is because under such realistic conditions, the catalytic performance is governed by an intricate interplay of many physical processes (Fig. 1).7,8,22,23 These include not only large and interconnected networks of surface elementary reactions, but also the catalyst's dynamic restructuring and changes in the surface composition during the reaction, mass and heat transport, particle sintering, to name a few. Notably, these physical processes operate across very different time and length scales and are often entangled. Therefore, explicitly modeling the full catalytic progression in order to design a new material is deemed inappropriate. Instead, our goal is to describe the statistical correlation of key physicochemical parameters with the experimental catalyst performance.Artificial intelligence (AI) offers the strategy for identifying correlations between physicochemical parameters and the measured catalytic performance.12,24–28 However, the utilization of experimental data in AI frameworks is complicated by the lack of consistent and well-characterized datasets, i.e., datasets created according to rigorous and standardized procedures.29–32 To address these limitations, we have developed an interpretable AI approach designed to identify physically meaningful descriptors using clean experimental data.2,29 The term “clean data” refers to the fact that the materials considered in the AI analysis are carefully synthesized, characterized, and tested according to standardized and well annotated experimental procedures. Indeed, the properties of the material and its catalytic performance are often sensitive to details of the experimental workflow, such as the sequence of synthesis steps and the pretreatment utilized prior to performance testing. Thus, all relevant steps of the workflow should be recorded in detail in order to guarantee the reproducibility of the experiment as well as the consistency of the resulting data.29–32 Unlike traditional physics-based models and atomistic simulations, this approach focuses on discovering statistical correlations rather than explicitly simulating all the physical processes, thereby leaving it open what structure or composition the working catalyst has and capturing the complexity of catalysis more effectively. Within this approach, a large set of experimentally measured or calculated candidate descriptive parameters, termed primary features, is first chosen. The primary features characterize the materials as well as the reaction environment, e.g., the temperature, pressure, or chemical potential of the phase in which reactants and products are contained. They correlate with physical processes that might be relevant for the system under consideration. Then, AI is utilized to identify a smaller number of key primary features correlated with the measured catalytic performance, from all initially offered primary features. Crucially, we note that AI identifies potentially nonlinear relationships among multiple primary features. In analogy with biology and medicine, these key primary features are termed materials genes of the catalytic function of interest, e.g., the activity or selectivity.2 Thus, the descriptor is a (nonlinear) function of the materials genes. The materials genes correlate with crucial physical processes that trigger, facilitate, or hinder the catalytic function of interest. In view of the high intricacy of these processes, the microscopic relationship between these genes and the catalytic function of interest might remain unknown. This concept has been applied to model the catalytic performance in alkane oxidation,2,33,34 CO oxidation,35 CO2 hydrogenation36 and selective hydrogenation of concentrated acetylene streams.37
The Sure Independence Screening and Sparsifying Operator (SISSO) symbolic-regression method38,39 gained prominence as a systematic AI approach to identify descriptors in materials science and catalysis.40–44 SISSO is well suited for the identification of materials genes in consistent experimental datasets, which often contain a rather small number of materials, e.g., <102, compared to the number of data points typically used to train AI and machine-learning models, e.g., >104. The descriptors identified by SISSO are interpretable in the sense that they depend on a (small) subset of physically meaningful genes, selected from many offered primary features. Additionally, the mathematical relationship between the genes and the target is explicit. Indeed, symbolic-regression methods identify analytical expressions describing the correlations in data.45–47 Such interpretability facilitates the extraction of physical insights beyond mere prediction.
In this work, we complement the SISSO approach with a gradient-based partial-effects (PEs)48,49 sensitivity analysis to identify descriptors for heterogeneous catalysis and quantify the influence of genes selected in the descriptor expressions (Fig. 2). The approach is illustrated for supported metal nanoparticles (NPs) applied in the selective hydrogenation of concentrated acetylene streams.37 The experimental data utilized here were created and analyzed by a focused SISSO approach in a previous publication.37 Here, we utilize a SISSO model for time-dependent selectivity towards ethylene identified in this previous study in order to demonstrate how the PE sensitivity analysis identifies the most influential genes of a SISSO model thus providing detailed physical insight per material.
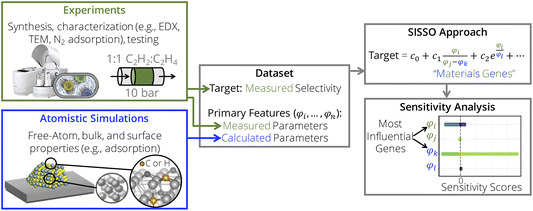 | ||
| Fig. 2 Interpretable AI approach for the identification of descriptors for heterogeneous catalysis based on data measured experimentally and calculated by atomistic simulations. By using consistent experiments and atomistic simulations on model systems, we create a dataset containing a target material’s function of interest (e.g., selectivity) and many candidate descriptive parameters (also called primary features) that characterize the materials and possibly relevant underlying physical processes. Then the sure-independence screening and sparsifying operator (SISSO) symbolic-regression approach identifies analytical expressions correlated with the target and depending on only a few of the initially offered primary features. These selected primary features are termed materials genes. Finally, the partial effects (PEs) sensitivity analysis pinpoints the genes that most influence the SISSO model. Parts of the figure are adapted from ref. 37. | ||
2 Methods
2.1 Dataset
The dataset analyzed in this publication can be found in ref. 37. We analyze metal NPs supported on high-surface-area-α-Al2O3. These catalysts were synthesized via mechanochemistry,50–53 which is a promising, more atom-efficient and environmental friendly alternative to wet synthesis methods.54 Nine palladium-based bimetallic alloy NPs are considered: Pd1Ag1, Pd1Ag5, Pd1Ag9, Pd1Au1, Pd1Au5, Pd1Au9, Pd1Cu1, Pd1Cu5, Pd1Cu9. These materials were characterized prior to the reaction by energy dispersive X-ray analysis in a scanning electron microscope (EDX-SEM), transmission-electron microscopy (TEM), and N2 physisorption. They were tested in the selective hydrogenation of highly concentrated acetylene streams. The reaction was performed in a steel-lined continuous-flow fixed-bed reactor at 10 bar and 150 °C. The applied feed contained a C2H2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) C2H4
C2H4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2 ratio of 1
H2 ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 at a weight hourly space velocity of 90
5 at a weight hourly space velocity of 90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cm3 gcat−1 h−1. The feed contained equimolar acetylene–ethylene mixtures, which would result from a hypothetical electric plasma-assisted methane-to-ethylene process. This plasma process can enable the production of ethylene from natural gas, biogas, or hydrogenated CO2 using the short-term surpluses in electricity from renewable sources.55–57 In this process, acetylene is formed as a by-product in equimolar concentrations and it needs to be selectively converted to ethylene in a dedicated, separate downstream process.58 Note that the selective hydrogenation of concentrated acetylene streams (>14 vol%) is a significantly different catalytic process compared to the selective hydrogenation of acetylene traces (between 0.1 and 2 vol%), which has been investigated much more extensively.55,59–61 Detailed descriptions of the procedures for materials synthesis, characterization, and testing are available elsewhere.37
000 cm3 gcat−1 h−1. The feed contained equimolar acetylene–ethylene mixtures, which would result from a hypothetical electric plasma-assisted methane-to-ethylene process. This plasma process can enable the production of ethylene from natural gas, biogas, or hydrogenated CO2 using the short-term surpluses in electricity from renewable sources.55–57 In this process, acetylene is formed as a by-product in equimolar concentrations and it needs to be selectively converted to ethylene in a dedicated, separate downstream process.58 Note that the selective hydrogenation of concentrated acetylene streams (>14 vol%) is a significantly different catalytic process compared to the selective hydrogenation of acetylene traces (between 0.1 and 2 vol%), which has been investigated much more extensively.55,59–61 Detailed descriptions of the procedures for materials synthesis, characterization, and testing are available elsewhere.37
As an example of target catalytic performance, we analyze the selectivity towards ethylene, denoted SC2H4. This selectivity was monitored during time on stream (tOS). The dataset contains 539 measurements of SC2H4, corresponding to 9 materials measured at multiple tOS's between 0 and 400 minutes (min). We note that the measured SC2H4 changes significantly with tOS for some of the materials (see below). The SISSO modelling aims at describing SC2H4 for different materials as well as its evolution with tOS during the initial stages of the catalytic process. All SC2H4 values correspond to materials and reaction conditions resulting in full (100%) conversion of the reactant acetylene. Thus, the selectivity values can be compared, as they correspond to a fixed conversion of acetylene.
As primary features, we utilized a combination of experimentally measured parameters with parameters calculated using atomistic simulations (Table 1). Four primary features were obtained from the experimental characterization of the materials:53,62 the metal loading (wmetal), the mean and standard deviation of the particle-diameter distribution (Dµ and Dσ), and the specific surface area (sBET). These primary features correspond to the entire material (NP + support) and they relate metal–support interactions and mesoscale properties of the materials. Noteworthy, these materials' properties are measured before the material is exposed to the reaction environment. Under the reaction, it is possible or even likely that properties, e.g., those related to particle-size distribution, change. In addition to the primary features extracted from the characterization of the materials, we included 3 experimental elemental free-atom properties and 3 experimental bulk properties as primary features reflecting the chemistry of the NP bulk, such as the ionization potential (IP) and the closest interatomic distance (dclosest). Those primary features were obtained from tabulated data.63–65 tOS is also offered as a primary feature in order to capture the time-on-stream dependent behavior of SC2H4.
| Type | Name | Symbol | Unit | Method |
|---|---|---|---|---|
| a Composition averages.b Measured prior to applying the material in the reaction. | ||||
| Reaction condition | Time on stream | tOS | min | — |
| NP + support | Total metal loading (weight fraction)b | wmetal | — | EDX-SEM53,62 |
| NP + support | Mean value of particle-diameter distributionb | Dµ | Å | TEM53,62 |
| NP + support | Standard deviation of particle-diameter distributionb | Dσ | Å | TEM53,62 |
| NP + support | Specific surface areab | sBET | m2 g−1 | N2 adsorption53,62 |
| NP free-atom | Ionization potentiala | 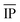 |
eV | Experimental63 |
| NP free-atom | Electron affinitya | 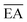 |
eV | Experimental64 |
| NP free-atom | Pauling electronegativitya | 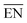 |
— | Experimental64 |
| NP bulka | Closest interatomic distancea |  |
Å | Experimental65 |
| NP bulka | Cohesive energya | 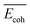 |
eV per atom | Experimental65 |
| NP bulka | Bulk modulusa | 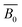 |
GPa | Experimental65 |
| NP clean surfaceb | Energy of the d-band centera | 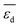 |
eV | DFT-GGA66 |
| NP surface with Cb | Critical surface carbon chemical potentiala | 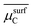 |
eV | DFT-GGA67 |
| NP surface with Cb | Critical subsurface carbon chemical potentiala | 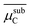 |
eV | DFT-GGA67 |
| NP surface with Cb | Distance expansion due to subsurface carbona | 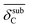 |
Å | DFT-GGA67 |
| NP surface with Cb | Surface deformation energy due to subsurface carbona | 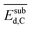 |
eV | DFT-GGA67 |
| NP surface with Cb | Subsurface carbon binding energya | 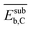 |
eV | DFT-GGA67 |
| NP surface with Hb | Surface binding energya | 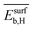 |
eV | DFT-GGA66 |
| NP surface with Hb | Subsurface binding energya | 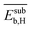 |
eV | DFT-GGA66 |
| NP surface with Hb | Work-function change due to hydrogen adsorptiona | 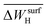 |
eV | DFT-GGA66 |
The 9 primary features calculated with atomistic simulations reflect the properties of the pristine NP surfaces and the interaction of carbon and hydrogen with the surface and subsurface. These primary features were calculated by DFT with the generalized gradient approximation for electron exchange and correlation (DFT-GGA) considering well-defined low-index model surfaces of previous works.66,67 The face-centered-cubic crystal structure and its (111) surface was adopted for palladium, silver, and gold. The properties associated with the most stable surface adsorption sites, i.e., the adsorption site corresponding to the highest binding strength, were considered. Examples of calculated primary features are the energy of the d-band center (εd), and the binding energy of subsurface hydrogen and carbon (Esubb,H and Esubb,C, respectively).
The NP elemental, bulk, and surface properties were converted into materials-specific primary features by taking the composition average, indicated by the bar in 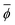 , where ϕ is an elemental, bulk, or surface parameter:
, where ϕ is an elemental, bulk, or surface parameter:
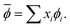 | (1) |
2.2 The SISSO approach
Starting from the primary features, the SISSO approach creates a large pool of analytic expressions, e.g., containing millions of elements, by iteratively applying a predefined set of mathematical operators such as addition, multiplication, logarithm. Then, compressed sensing68,69 is used to identify the few analytical functions, that combined by weighting coefficients provide the best correlation between the selected primary features (genes) and the target property, here the ethylene selectivity SC2H4. The SISSO model has the form
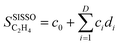 | (2) |
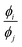 , ϕi−1, exp(ϕi), ϕi2, ϕi3, ln(ϕi), ϕi6, ϕi1/2, ϕi1/3, and exp(−ϕi). The units of the primary features are respected so that, for instance, the operators addition and subtraction are only allowed between features with the same unit. We used a nonlinear optimization during the generation of expressions to include scale and bias terms for the mathematical operations logarithm and exponential.39
, ϕi−1, exp(ϕi), ϕi2, ϕi3, ln(ϕi), ϕi6, ϕi1/2, ϕi1/3, and exp(−ϕi). The units of the primary features are respected so that, for instance, the operators addition and subtraction are only allowed between features with the same unit. We used a nonlinear optimization during the generation of expressions to include scale and bias terms for the mathematical operations logarithm and exponential.39
A nested five-fold cross validation was used in order to determine the hyperparameters of the SISSO model (R and D) and to estimate the predictive performance. In the outer loop of this cross-validation scheme, the dataset is randomly split into five folds with equal size. Each fold is then used as the test set once, while the remaining four folds are used to train a model. In the inner loop, a new five-fold split is done using the training sets obtained from the first split. Each fold obtained from this second split is used as a validation set once, while the remaining four folds are used as the training sets. The validation sets are used to determine the hyperparameters of the SISSO model. The hyperparameters are chosen such as the root-mean-squared errors (RMSEs) evaluated on the validation sets are minimized. For the modelling of SC2H4, we identify R = 2 and D = 3 as appropriate hyperparameters. The test sets are used to estimate the predictive performance. The distribution of errors evaluated on the test sets indicate a good predictive performance for the modelling of SC2H4. The mean test error value, for instance, is equal to 0.105. This value is below 25% of the standard deviation of the distribution of SC2H4 values across the entire dataset. Further details on the application of the SISSO approach to model SC2H4 (ref. 37) and on the nested cross validation71 are available elsewhere.
2.3 Sensitivity analysis
The models identified by SISSO generally only depend on a few of the initially offered primary features. However, the relative importance of these different genes that appear in a SISSO model might differ significantly. Sensitivity analyses identify the most influential input variables of a model.70,72–75 Thus, they facilitate the extraction of physical insight and help determining the most promising materials. Sensitivity analyses are referred to as local when they provide sensitivity scores per data point (e.g., per material) or global when they provide average sensitivity scores for specific data space, e.g., the entire population of materials. Additionally, the sensitivity analysis can be model specific or model agnostic. Model-agnostic sensitivity analyses examine how changes in an input variable affect the target (output) by systematically modifying the values of the input variables and recording the changes in the model output. The Sobol method, for instance, is a popular model-agnostic global sensitivity analysis70,73,76 that decomposes the variance of the model output into contributions from individual input variables and their interactions.Here, we use the gradient-based partial-effects (PE) sensitivity analysis to assess the impact of genes identified by the SISSO approach. The PE method48,49,77 quantifies the impact of a given gene in the model's output when the remaining genes are fixed by means of the partial derivative of the model with respect to this gene.49,78 This model-specific approach exploits the analytical expressions identified by SISSO and provides local sensitivity scores. The local sensitivity analysis is insightful when the physical processes governing the target are significantly different for different groups of materials. In such cases, the contribution of a specific gene might be different for different materials. PEs are less computationally demanding than widely used analyses such as Sobol, since the partial derivative can be obtained analytically.
Let us now apply this concept to 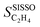 . The partial derivative with respect to gene ϕj is:
. The partial derivative with respect to gene ϕj is:
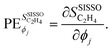 | (3) |
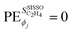 if an initially offered primary feature ϕj was not selected in the SISSO model. The unit of
if an initially offered primary feature ϕj was not selected in the SISSO model. The unit of 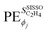 is the unit of the target divided by the unit of the gene ϕj. Thus,
is the unit of the target divided by the unit of the gene ϕj. Thus, 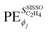 values associated to genes having different units are not directly comparable. This is inconvenient when a quantitative comparison among PEs corresponding to different genes that appear in a given model is desired. In order to obtain PEs that can be directly compared across genes which might have different units and scales, we define scaled partial effects (SPEs) as
values associated to genes having different units are not directly comparable. This is inconvenient when a quantitative comparison among PEs corresponding to different genes that appear in a given model is desired. In order to obtain PEs that can be directly compared across genes which might have different units and scales, we define scaled partial effects (SPEs) as
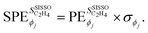 | (4) |
 has the unit of the target. Provided that the SISSO model expression
has the unit of the target. Provided that the SISSO model expression 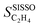 is differentiable, the quantities defined in eqn (3) and (4) are analytical expressions obtained by analytical differentiation. The values of these expressions can be evaluated for each gene or, equivalently, for each data point, providing local sensitivity scores. Global sensitivity scores can then be derived by computing the SPEs for all data points and averaging the results:
is differentiable, the quantities defined in eqn (3) and (4) are analytical expressions obtained by analytical differentiation. The values of these expressions can be evaluated for each gene or, equivalently, for each data point, providing local sensitivity scores. Global sensitivity scores can then be derived by computing the SPEs for all data points and averaging the results:
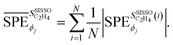 | (5) |
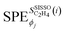 are the values of the SPEs evaluated for the i-th entry of the training data set and the sum runs over the N entries of this data set. The derivatives of SISSO models were evaluated using the SymPy package.79 This paper introduces the PE-based sensitivity approach applied to SISSO models. More details about this PE approach and other applications in materials science will be discussed in an upcoming study.80
are the values of the SPEs evaluated for the i-th entry of the training data set and the sum runs over the N entries of this data set. The derivatives of SISSO models were evaluated using the SymPy package.79 This paper introduces the PE-based sensitivity approach applied to SISSO models. More details about this PE approach and other applications in materials science will be discussed in an upcoming study.80
3 Results and discussion
This section summarizes the results for the nine mentioned NP catalysts. We analyze the evolution of ethylene selectivity SC2H4 with time on stream tOS during the hydrogenation of concentrated acetylene streams at 150 °C. Modeling the evolution of the catalytic performance with tOS allows obtaining insights on the modifications that the materials experience at early stages of the reaction and it allows estimating the time span during which the material can be effectively utilized.3.1 Measured ethylene selectivity
The values of SC2H4 range from −1 to 1, as ethylene is both a product of the acetylene selective hydrogenation reaction and part of the reaction feed. Negative SC2H4 indicates that more ethylene from the feed is consumed than formed from acetylene. Positive SC2H4 indicates selective materials and conditions. The desired behavior is SC2H4 = 1 or, equivalently, 100% conversion. Diverse performance scenarios ranging from unselective to highly selective are observed among the considered materials and tOS (Fig. 3A). The different catalysts show even qualitatively different performance. The highly selective situations present SC2H4 values of up to 0.75. In general, the materials containing palladium–silver NPs are the most selective ones, followed by palladium–gold, and palladium–copper. Additionally, different profiles for the evolution of SC2H4 with tOS are observed depending on the material. Most of the materials based on the palladium–silver alloys are selective from the start of the reaction, i.e., from tOS = 0. For these materials, the selectivity reaches a stable level and then slightly decreases at longer tOS. In contrast, materials containing palladium–gold and, in particular, palladium–copper NPs present constantly increasing SC2H4 with tOS. The initial selectivity of Pd1Cu9, e.g., is SC2H4 = −0.89 but it increases with tOS up to SC2H4 = 0.47 after 365 min. This represents a drastic shift from a completely unselective condition to a highly selective one.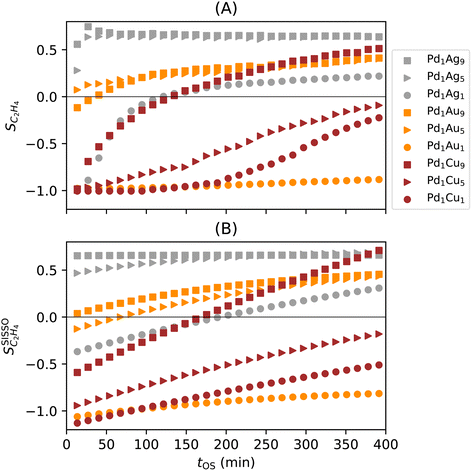 | ||
Fig. 3 Experimental results and SISSO modelling for the ethylene selectivity in the selective hydrogenation of concentrated acetylene streams catalyzed by palladium-based alloys supported on alumina. (A) Measured SC2H4 with time on stream. (B) Fit of the SISSO model of eqn (6) 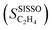 to the data. Figure adapted from ref. 37. to the data. Figure adapted from ref. 37. | ||
3.2 SISSO model for ethylene selectivity
SISSO was applied to the training dataset containing 539 data points, associated to the 9 materials measured at multiple tOS's. This dataset contains the measured target SC2H4 and 20 primary features, including tOS as well as a wide range of measured and calculated parameters characterizing the entire materials set and the chemistry of the bulk and surface of the NPs (Table 1). The choice of some of the offered primary features reflects relevant physical processes identified by previous surface science studies and atomistic simulations on model systems. The selectivity towards ethylene depends on the competition between ethylene desorption and its hydrogenation to ethane. Thus, the d-band center17 is included as a primary feature, as it correlates with the adsorption of ethylene as well as other intermediates of the acetylene hydrogenation reaction such as vinylidene and ethyl.81,82 Additionally, we offer primary features that capture the interaction of NP surfaces with hydrogen, since the availability of surface and subsurface hydrogen impacts the rate of ethylene hydrogenation to ethane.83 Primary features reflecting the interaction of metal surfaces with carbon are also included, as the formation of surface carbides has been shown to be a crucial factor determining the selectivity in palladium systems,57,84–90 as it limits the hydrogen availability. Further details on the dataset and SISSO approach are given in the Methods section.The good quality of the fit provided by the SISSO model for ethylene selectivity identified in ref. 37, denoted 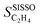 , is shown in Fig. 3B. This model is able to describe the selectivity of the different materials and its evolution with tOS. The expression of the model is
, is shown in Fig. 3B. This model is able to describe the selectivity of the different materials and its evolution with tOS. The expression of the model is
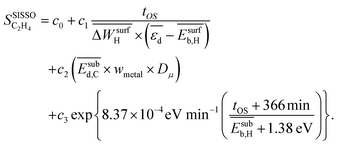 | (6) |
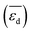 , the average deformation energy of subsurface carbon
, the average deformation energy of subsurface carbon 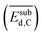 , the average work-function shift with hydrogen adsorption
, the average work-function shift with hydrogen adsorption 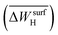 , the average binding energy of hydrogen
, the average binding energy of hydrogen 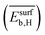 , and the average binding energy of subsurface hydrogen
, and the average binding energy of subsurface hydrogen 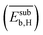 . From the initially offered 20 primary features, SISSO selected 8 relevant genes to describe SC2H4 (eqn (6)). However, the genes that enter in the model expression can impact the model to different extents. In order to identify the most influential genes of eqn (6), the PE analysis is applied.
. From the initially offered 20 primary features, SISSO selected 8 relevant genes to describe SC2H4 (eqn (6)). However, the genes that enter in the model expression can impact the model to different extents. In order to identify the most influential genes of eqn (6), the PE analysis is applied.
3.3 Sensitivity analysis of the SISSO model for ethylene selectivity
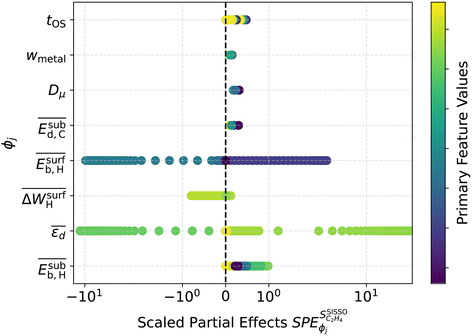
The ’s for the 8 genes that were selected by SISSO in the expression of the model
’s for the 8 genes that were selected by SISSO in the expression of the model 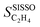 are shown in Fig. 4. In this figure, each line corresponds to one gene and each circle corresponds to the
are shown in Fig. 4. In this figure, each line corresponds to one gene and each circle corresponds to the  evaluated for one data point of the training dataset. The circles are colored according to the values of genes. The scale of the x axis is logarithm for a better visualization, as the SPEs have different magnitudes. For primary features that were not selected by SISSO in eqn (6),
evaluated for one data point of the training dataset. The circles are colored according to the values of genes. The scale of the x axis is logarithm for a better visualization, as the SPEs have different magnitudes. For primary features that were not selected by SISSO in eqn (6), 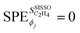 . Details of the PE analysis are available in the Methods section.
. Details of the PE analysis are available in the Methods section.
In order to assess the overall sensitivity of the 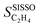 model with respect to the genes, we evaluated the mean absolute values of SPEs
model with respect to the genes, we evaluated the mean absolute values of SPEs 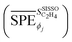 for each gene as a global sensitivity score. The
for each gene as a global sensitivity score. The  values are 0.15, 0.12, 0.23, 0.16, 0.75, 0.092, 2.81, and 0.42 for tOS, wmetal, Dµ,
values are 0.15, 0.12, 0.23, 0.16, 0.75, 0.092, 2.81, and 0.42 for tOS, wmetal, Dµ, 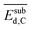 ,
, 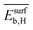 ,
, 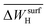 ,
, 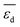 , and
, and 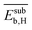 , respectively. Therefore, the relative global influence of the genes to the model decreases as
, respectively. Therefore, the relative global influence of the genes to the model decreases as 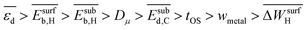 . The most influential gene in the
. The most influential gene in the 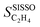 model (eqn (6)) is the average d-band center
model (eqn (6)) is the average d-band center 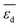 . This can be related to the key role of physical processes related to the adsorption of reaction intermediates on the NP surface. As shown previously, the alloying of palladium with a second metal downshifts the d-band center, weakening the adsorption of π-bonded species such as ethylene, for instance.81,82 Thus, ethylene desorption is favored over its hydrogenation to ethane. The second and third most influential genes are
. This can be related to the key role of physical processes related to the adsorption of reaction intermediates on the NP surface. As shown previously, the alloying of palladium with a second metal downshifts the d-band center, weakening the adsorption of π-bonded species such as ethylene, for instance.81,82 Thus, ethylene desorption is favored over its hydrogenation to ethane. The second and third most influential genes are 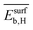 and
and 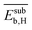 . They reflect the bond strength between the surface and subsurface with hydrogen. Thus, their high relevance can be related to the availability of surface and subsurface hydrogen and its impact on the rate of hydrogenation of ethylene to ethane, which hinders the formation of the selective-hydrogenation product ethylene.91 We note that the values of
. They reflect the bond strength between the surface and subsurface with hydrogen. Thus, their high relevance can be related to the availability of surface and subsurface hydrogen and its impact on the rate of hydrogenation of ethylene to ethane, which hinders the formation of the selective-hydrogenation product ethylene.91 We note that the values of 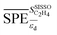 and
and 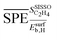 can be positive or negative depending on the material and tOS (Fig. 4), indicating that the relationships between
can be positive or negative depending on the material and tOS (Fig. 4), indicating that the relationships between 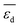 or
or 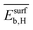 and SC2H4 can be directly or inversely proportional. Thus, the adsorption properties can either favor or hinder the selectivity. Finally, the fourth most influential gene Dµ could be related to a particle size effect on the ethylene selectivity. Indeed, the size of palladium or copper NPs can significantly affect their selectivity in the hydrogenation of unsaturated bonds by controlling the relative amount of corner and edge sites, which are less coordinated and can thus provide stronger adsorption than the surface sites on closely packed surfaces.87,92,93 The most influential genes identified by the sensitivity analysis confirm the crucial role of the adsorption of π-bonded species and hydrogen in modulating the ethylene selectivity, as discussed in previous works addressing the hydrogenation of diluted acetylene streams.81,82 Additionally, the PEs identify a structure-sensitivity effect, which was only discussed in detail for the hydrogenation of dienes.87,92,93 The
and SC2H4 can be directly or inversely proportional. Thus, the adsorption properties can either favor or hinder the selectivity. Finally, the fourth most influential gene Dµ could be related to a particle size effect on the ethylene selectivity. Indeed, the size of palladium or copper NPs can significantly affect their selectivity in the hydrogenation of unsaturated bonds by controlling the relative amount of corner and edge sites, which are less coordinated and can thus provide stronger adsorption than the surface sites on closely packed surfaces.87,92,93 The most influential genes identified by the sensitivity analysis confirm the crucial role of the adsorption of π-bonded species and hydrogen in modulating the ethylene selectivity, as discussed in previous works addressing the hydrogenation of diluted acetylene streams.81,82 Additionally, the PEs identify a structure-sensitivity effect, which was only discussed in detail for the hydrogenation of dienes.87,92,93 The 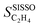 model is less sensitive to the genes
model is less sensitive to the genes 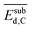 , tOS, wmetal, and
, tOS, wmetal, and 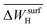 . Thus, the PE analysis indicates a low influence of (sub)surface carbon on the ethylene selectivity for the systems studied here. This indicates that the formation of surface carbides discussed previously in the context of the hydrogenation of diluted acetylene streams might be less relevant to determine the selectivity for the systems considered here.57,84–90
. Thus, the PE analysis indicates a low influence of (sub)surface carbon on the ethylene selectivity for the systems studied here. This indicates that the formation of surface carbides discussed previously in the context of the hydrogenation of diluted acetylene streams might be less relevant to determine the selectivity for the systems considered here.57,84–90
Fig. 5 shows the SPEs for the 4 most influential genes for each of the materials as a function of tOS. The magnitude of the SPE scores associated to the genes 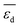 and
and 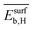 is higher for the material based on Pd1Cu9 compared to the remaining materials. This difference increases with tOS. This suggests that the impact of physical processes related to adsorption on SC2H4 is particularly important for this material, especially at long tOS. It can be related to the fact that the Pd1Cu9 alloy presents the most drastic change in SC2H4 with time, as it is a total hydrogenation catalyst at tOS = 0 min, but it becomes selective at longer reaction times (Fig. 3). Interestingly, whereas
is higher for the material based on Pd1Cu9 compared to the remaining materials. This difference increases with tOS. This suggests that the impact of physical processes related to adsorption on SC2H4 is particularly important for this material, especially at long tOS. It can be related to the fact that the Pd1Cu9 alloy presents the most drastic change in SC2H4 with time, as it is a total hydrogenation catalyst at tOS = 0 min, but it becomes selective at longer reaction times (Fig. 3). Interestingly, whereas  is positive and thus an increase in
is positive and thus an increase in 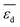 favors selectivity for Pd1Cu9,
favors selectivity for Pd1Cu9, 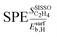 is negative and an increase in
is negative and an increase in 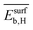 hinders the selectivity. The SPEs related to the gene
hinders the selectivity. The SPEs related to the gene 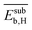 are relatively insensitive to tOS for most materials. However, the
are relatively insensitive to tOS for most materials. However, the 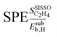 values decrease significantly with tOS for the materials based on Pd1Au9, Pd1Au5, and Pd1Ag5, indicating that physical processes related to subsurface hydrogen are only relevant at the beginning of the induction period for these materials. Finally, the analysis of per-material
values decrease significantly with tOS for the materials based on Pd1Au9, Pd1Au5, and Pd1Ag5, indicating that physical processes related to subsurface hydrogen are only relevant at the beginning of the induction period for these materials. Finally, the analysis of per-material  scores shows that the particle-size distribution impacts the copper-based materials, in particular the materials based on Pd1Cu9 and Pd1Cu5, to a greater extent compared to the remaining materials. This indicates a higher influence of structure-sensitivity for copper-based catalysts. This analysis illustrates how per-data-point, e.g., per-material, SPE scores can be used to obtain insights on the most relevant underlying physical processes for specific data points. In addition to providing global and local, per-data-point physical insights via the identification of the most influential genes in the SISSO model, the sensitivity analysis can also be used to design high-throughput materials-screening protocols.70 In these protocols, the values of the most influential genes are used to determine rules indicating new materials likely presenting desired target property values, e.g., high values.
scores shows that the particle-size distribution impacts the copper-based materials, in particular the materials based on Pd1Cu9 and Pd1Cu5, to a greater extent compared to the remaining materials. This indicates a higher influence of structure-sensitivity for copper-based catalysts. This analysis illustrates how per-data-point, e.g., per-material, SPE scores can be used to obtain insights on the most relevant underlying physical processes for specific data points. In addition to providing global and local, per-data-point physical insights via the identification of the most influential genes in the SISSO model, the sensitivity analysis can also be used to design high-throughput materials-screening protocols.70 In these protocols, the values of the most influential genes are used to determine rules indicating new materials likely presenting desired target property values, e.g., high values.
 | ||
Fig. 5 Evolution of the scaled partial effects  with time on stream (tOS) for the four most influential genes selected by SISSO in the model of eqn (6) with time on stream (tOS) for the four most influential genes selected by SISSO in the model of eqn (6) 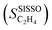 : : 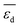 (A), (A), 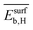 (B), (B), 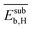 (C), Dµ (D). (C), Dµ (D). | ||
The three most influential genes identified by the sensitivity analysis 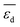 ,
, 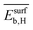 , and
, and 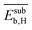 are parameters calculated via DFT-GGA calculations for model systems and they highlight the importance of the adsorption properties for catalysis. However, the relevance of the experimental parameter Dµ stresses the utility of measured primary features to capture aspects that can hardly be included in the theoretical description. Indeed, the distribution of NP sizes is influenced by the properties of the metal alloy as well as by metal–support interactions and synthesis conditions. Additionally, this distribution might be also modified under the reaction conditions. Modelling all such physical processes by theory is unfeasible. The utilization of materials properties measured under conditions close to reaction conditions, e.g., using in situ and operando spectroscopy, as primary features was also shown to be crucial in order to capture the catalyst restructuring effect on the catalytic performance.2,34 Therefore, the combination of experimental and theoretical (calculated) primary features is a promising avenue for the identification of descriptors that capture the complexity of heterogeneous catalysis.36 As systematic experimental and theoretical data become available,20,94–96 the materials-genes concept offers a framework for integrating surface science, atomistic modelling, and operando characterization towards efficient catalyst design.
are parameters calculated via DFT-GGA calculations for model systems and they highlight the importance of the adsorption properties for catalysis. However, the relevance of the experimental parameter Dµ stresses the utility of measured primary features to capture aspects that can hardly be included in the theoretical description. Indeed, the distribution of NP sizes is influenced by the properties of the metal alloy as well as by metal–support interactions and synthesis conditions. Additionally, this distribution might be also modified under the reaction conditions. Modelling all such physical processes by theory is unfeasible. The utilization of materials properties measured under conditions close to reaction conditions, e.g., using in situ and operando spectroscopy, as primary features was also shown to be crucial in order to capture the catalyst restructuring effect on the catalytic performance.2,34 Therefore, the combination of experimental and theoretical (calculated) primary features is a promising avenue for the identification of descriptors that capture the complexity of heterogeneous catalysis.36 As systematic experimental and theoretical data become available,20,94–96 the materials-genes concept offers a framework for integrating surface science, atomistic modelling, and operando characterization towards efficient catalyst design.
Finally, we stress that SISSO and the materials genes will only provide reliable descriptions for materials and reaction conditions governed by the same underlying physical processes that govern the performance of the situations in the training set. The reliability of the SISSO models can be assessed and used in order to systematically acquire new data to improve the description for portions of the data space that are not sufficiently covered by the training dataset.97 This aspect is discussed in detail in the contribution by Nair et al.
4 Conclusions
In this contribution, we propose rethinking the description of heterogeneous catalysis by focusing on statistical correlations identified in experimental and theoretical data generated systematically rather than explicitly modeling all the underlying physicochemical processes and their intricate interplay via atomistic simulations. This approach can capture the intricacy of catalysis more effectively than previous computational approaches, as it does not assume a single underlying physical model. By applying the SISSO approach with the gradient-based PE sensitivity analysis, we identified the most influential basic physicochemical parameters correlated with the selectivity of metal NPs supported on alumina and applied in the hydrogenation of concentrated acetylene streams. These parameters indicate that the adsorption of hydrogen and of molecules containing π bonds modulate the ethylene selectivity. Additionally, a structure-sensitivity effect on the selectivity is captured by the experimentally measured NP size. In addition to the chemical insights, the materials-genes concept enables the design of improved materials and reaction conditions for catalysis.37 In particular, the most influential physicochemical parameters determined by sensitivity analyses can be used to efficiently screen new candidate materials.70 The integration of materials genes with automated experiments and large-scale atomistic simulations offers a promising path towards more predictive and generalizable catalyst design.Conflicts of interest
The authors declare no competing interests.Data availability
The dataset is available in the Excel supplementary information file of ref. 37. The SISSO analysis is available at https://github.com/lfoppa/Focused-AI-SGD-SISSO-acetylene-hydrogenation.Acknowledgements
This work was funded by the ERC Advanced Grant TEC1p (European Research Council, Grant Agreement No. 740233). Open Access funding provided by the Max Planck Society.Notes and references
- J. K. Nørskov and T. Bligaard, Angew. Chem., Int. Ed., 2013, 52, 776–777 CrossRef PubMed.
- L. Foppa, L. M. Ghiringhelli, F. Girgsdies, M. Hashagen, P. Kube, M. Hävecker, S. J. Carey, A. Tarasov, P. Kraus, F. Rosowski, R. Schlögl, A. Trunschke and M. Scheffler, MRS Bull., 2021, 46, 1016–1026 CrossRef CAS PubMed.
- T. Engel and G. Ertl, Elementary Steps in the Catalytic Oxidation of Carbon Monoxide on Platinum Metals, Advances in Catalysis, Academic Press, 1979, vol. 28, pp. 1–78 Search PubMed.
- J. W. Couves, J. M. Thomas, D. Waller, R. H. Jones, A. J. Dent, G. E. Derbyshire and G. N. Greaves, Nature, 1991, 354, 465–468 CrossRef CAS.
- C. T. Campbell, Surf. Sci. Rep., 1997, 27, 1–111 CrossRef CAS.
- G. Ertl, Angew. Chem., Int. Ed., 2008, 47, 3524–3535 CrossRef CAS PubMed.
- G. A. Somorjai and J. Y. Park, Angew. Chem., Int. Ed., 2008, 47, 9212–9228 CrossRef CAS PubMed.
- J. M. Thomas, J. Chem. Phys., 2008, 128, 182502 CrossRef PubMed.
- K. Reuter, C. Stampf and M. Scheffler, Handbook of Materials Modeling: Methods, 2005, pp. 149–194 Search PubMed.
- J. K. Nørskov, F. Abild-Pedersen, F. Studt and T. Bligaard, Proc. Natl. Acad. Sci. U. S. A., 2011, 108, 937–943 CrossRef PubMed.
- A. Bruix, J. T. Margraf, M. Andersen and K. Reuter, Nat. Catal., 2019, 2, 659–670 CrossRef CAS.
- A. J. Medford, A. Vojvodic, J. S. Hummelshøj, J. Voss, F. Abild-Pedersen, F. Studt, T. Bligaard, A. Nilsson and J. K. Nørskov, J. Catal., 2015, 328, 36–42 CrossRef CAS.
- P. Sabatier, La catalyse en chimie organique, C. Béranger, 1920 Search PubMed.
- J. Bronsted, Chem. Rev., 1928, 5, 231–338 CrossRef CAS.
- M. G. Evans and M. Polanyi, Trans. Faraday Soc., 1938, 34, 11–24 RSC.
- B. Hammer and J. Nørskov, Surf. Sci., 1995, 343, 211–220 CrossRef CAS.
- B. Hammer and J. K. Nørskov, Adv. Catal., 2000, 45, 71–129 CAS.
- M. Scheffler and C. Stampfl, Theory of Adsorption on Metal Substrates, in Handbook of Surface Science, ed. K. Horn, and M. Scheffler, Elsevier, Amsterdam, 2000, vol. 2, pp. 286–356 Search PubMed.
- J. K. Nørskov, T. Bligaard, J. Rossmeisl and C. H. Christensen, Nat. Chem., 2009, 1, 37–46 CrossRef PubMed.
- L. Chanussot, A. Das, S. Goyal, T. Lavril, M. Shuaibi, M. Riviere, K. Tran, J. Heras-Domingo, C. Ho, W. Hu, A. Palizhati, A. Sriram, B. Wood, J. Yoon, D. Parikh, C. L. Zitnick and Z. Ulissi, ACS Catal., 2021, 11, 6059–6072 CrossRef CAS.
- J. Abed, J. Kim, M. Shuaibi, B. Wander, B. Duijf, S. Mahesh, H. Lee, V. Gharakhanyan, S. Hoogland, E. Irtem, J. Lan, N. Schouten, A. U. Vijayakumar, J. Hattrick-Simpers, J. R. Kitchin, Z. W. Ulissi, A. van Vugt, E. H. Sargent, D. Sinton and C. L. Zitnick, Open Catalyst Experiments 2024 (OCx24): Bridging Experiments and Computational Models, arXiv, 2024, preprint, arXiv:2411.11783, DOI:10.48550/arXiv.2411.11783.
- H.-J. Freund, G. Meijer, M. Scheffler, R. Schlögl and M. Wolf, Angew. Chem., Int. Ed., 2011, 50, 10064–10094 CrossRef CAS PubMed.
- R. Schlögl, Angew. Chem., Int. Ed., 2015, 54, 3465–3520 CrossRef PubMed.
- L.-H. Mou, T. Han, P. E. S. Smith, E. Sharman and J. Jiang, Adv. Sci., 2023, 10, 2301020 CrossRef CAS PubMed.
- Z.-J. Zhao and J. Gong, Nat. Nanotechnol., 2022, 17, 563–564 CrossRef PubMed.
- K. Takahashi, L. Takahashi, S. D. Le, T. Kinoshita, S. Nishimura and J. Ohyama, J. Am. Chem. Soc., 2022, 144, 15735–15744 CrossRef CAS PubMed.
- T. Taniike, A. Fujiwara, S. Nakanowatari, F. García-Escobar and K. Takahashi, Commun. Chem., 2024, 7, 11 CrossRef PubMed.
- J. A. Esterhuizen, B. R. Goldsmith and S. Linic, Nat. Catal., 2022, 5, 175–184 CrossRef.
- A. Trunschke, G. Bellini, M. Boniface, S. J. Carey, J. Dong, E. Erdem, L. Foppa, W. Frandsen, M. Geske and L. M. Ghiringhelli, et al., Top. Catal., 2020, 63, 1683–1699 CrossRef CAS.
- A. Moshantaf, M. Wesemann, S. Beinlich, H. Junkes, J. Schumann, B. Alkan, P. Kube, C. P. Marshall, N. Pfister and A. Trunschke, Catal. Sci. Technol., 2024, 14, 6186–6197 RSC.
- D. W. Flaherty and A. Bhan, J. Catal., 2024, 431, 115408 CrossRef CAS.
- A. C. Alba-Rubio, P. Christopher, M. L. Personick and K. J. Stowers, J. Catal., 2024, 429, 115259 CrossRef CAS.
- L. Foppa, C. Sutton, L. M. Ghiringhelli, S. De, P. Löser, S. A. Schunk, A. Schäfer and M. Scheffler, ACS Catal., 2022, 12, 2223 CrossRef CAS PubMed.
- L. Foppa, F. Ruther, M. Geske, G. Koch, F. Girgsdies, P. Kube, S. J. Carey, M. Havecker, O. Timpe and A. V. Tarasov, et al., J. Am. Chem. Soc., 2023, 145, 3427–3442 CrossRef CAS PubMed.
- G. Bellini, G. Koch, F. Girgsdies, J. Dong, S. J. Carey, O. Timpe, G. Auffermann, M. Scheffler, R. Schlögl, L. Foppa and A. Trunschke, Angew. Chem., Int. Ed., 2025, 64, e202417812 CrossRef CAS PubMed.
- R. Miyazaki, K. S. Belthle, H. Tüysüz, L. Foppa and M. Scheffler, J. Am. Chem. Soc., 2024, 146, 5433–5444 CrossRef CAS PubMed.
- J. M. Mauß, K. S. Kley, R. Khobragade, N.-K. Tran, J. de Bellis, F. Schüth, M. Scheffler and L. Foppa, ACS Catal., 2025, 15, 12652–12665 CrossRef PubMed.
- R. Ouyang, S. Curtarolo, E. Ahmetcik, M. Scheffler and L. M. Ghiringhelli, Phys. Rev. Mater., 2018, 2, 083802 CrossRef CAS.
- T. A. R. Purcell, M. Scheffler and L. M. Ghiringhelli, J. Chem. Phys., 2023, 159, 114110 CrossRef CAS PubMed.
- C. J. Bartel, S. L. Millican, A. M. Deml, J. R. Rumptz, W. Tumas, A. W. Weimer, S. Lany, V. Stevanović, C. B. Musgrave and A. M. Holder, Nat. Commun., 2018, 9, 4168 CrossRef PubMed.
- C. J. Bartel, C. Sutton, B. R. Goldsmith, R. Ouyang, C. B. Musgrave, L. M. Ghiringhelli and M. Scheffler, Sci. Adv., 2019, 5, eaav0693 CrossRef CAS PubMed.
- S. R. Xie, G. R. Stewart, J. J. Hamlin, P. J. Hirschfeld and R. G. Hennig, Phys. Rev. B, 2019, 100, 174513 CrossRef CAS.
- R. Ouyang, E. Ahmetcik, C. Carbogno, M. Scheffler and L. M. Ghiringhelli, J. Phys.: Mater., 2019, 2, 024002 Search PubMed.
- T. Wang, J. Hu, R. Ouyang, Y. Wang, Y. Huang, S. Hu and W.-X. Li, Science, 2024, 386, 915–920 CrossRef CAS PubMed.
- M. Schmidt and H. Lipson, Science, 2009, 324, 81 CrossRef CAS PubMed.
- L. Wang, F. Li, Y. Chen and J. Chen, J. Energy Chem., 2019, 28(2), 40–49 CrossRef.
- P. Orzechowski, W. La Cava and J. H. Moore, Proceedings of the Genetic and Evolutionary Computation Conference, New York, NY, USA, 2018, pp. 1183–1190 Search PubMed.
- E. Onukwugha, J. Bergtold and R. Jain, PharmacoEconomics, 2015, 33, 25–30 CrossRef PubMed.
- G. S. I. Aldeia and F. O. de França, Proceedings of the Genetic and Evolutionary Computation Conference, New York, NY, USA, 2021, pp. 750–758 Search PubMed.
- H. Schreyer, R. Eckert, S. Immohr, J. deBellis, M. Felderhoff and F. Schüth, Angew. Chem., Int. Ed., 2019, 58, 11262–11265 CrossRef CAS PubMed.
- A. P. Amrute, J. De Bellis, M. Felderhoff and F. Schüth, Chem.–Eur. J., 2021, 27, 6819–6847 CrossRef CAS PubMed.
- J. De Bellis, M. Felderhoff and F. Schüth, Chem. Mater., 2021, 33, 2037–2045 CrossRef CAS.
- K. S. Kley, J. De Bellis and F. Schüth, Catal. Sci. Technol., 2023, 13, 119–131 RSC.
- K. J. Ardila-Fierro and J. G. Hernández, ChemSusChem, 2021, 14, 2145–2162 CrossRef CAS PubMed.
- I.-T. Trotuş, T. Zimmermann and F. Schüth, Chem. Rev., 2014, 114, 1761–1782 CrossRef PubMed.
- M. Bowker, S. DeBeer, N. F. Dummer, G. J. Hutchings, M. Scheffler, F. Schüth, S. H. Taylor and H. Tüysüz, Angew. Chem., Int. Ed., 2022, 61, e202209016 CrossRef CAS PubMed.
- Z. Li, E. Öztuna, K. Skorupska, O. V. Vinogradova, A. Jamshaid, A. Steigert, C. Rohner, M. Dimitrakopoulou, M. J. Prieto, C. Kunkel, M. Stredansky, P. Kube, M. Götte, A. M. Dudzinski, F. Girgsdies, S. Wrabetz, W. Frandsen, R. Blume, P. Zeller, M. Muske, D. Delgado, S. Jiang, F.-P. Schmidt, T. Köhler, M. Arztmann, A. Efimenko, J. Frisch, T. M. Kokumai, R. Garcia-Diez, M. Bär, A. Hammud, J. Kröhnert, A. Trunschke, C. Scheurer, T. Schmidt, T. Lunkenbein, D. Amkreutz, H. Kuhlenbeck, V. J. Bukas, A. Knop-Gericke, R. Schlatmann, K. Reuter, B. R. Cuenya and R. Schlögl, Nat. Commun., 2024, 15, 10660 CrossRef CAS PubMed.
- E. Delikonstantis, E. Igos, M. Augustinus, E. Benetto and G. D. Stefanidis, Sustainable Energy Fuels, 2020, 4, 1351–1362 RSC.
- X. Cao, B. W.-L. Jang, J. Hu, L. Wang and S. Zhang, Molecules, 2023, 28, 2572 CrossRef CAS PubMed.
- M. Salciccioli, Y. Chen and D. G. Vlachos, Ind. Eng. Chem. Res., 2011, 50, 28–40 CrossRef CAS.
- A. Dasgupta, H. He, R. Gong, S.-L. Shang, E. K. Zimmerer, R. J. Meyer, Z.-K. Liu, M. J. Janik and R. M. Rioux, Nat. Chem., 2022, 14, 523–529 CrossRef CAS PubMed.
- K. K. Kley, PhD thesis, Ruhr-Universität Bochum, 2022.
- NIST Chemistry WebBook, NIST Standard Reference Database Number 69, ed. S. Lias, P. J. Linstrom and W. G. Mallard, Gaithersburg, USA, 2005 Search PubMed.
- WebElements, https://www.webelements.com/, accessed: 2024-10-10 Search PubMed.
- P. Janthon, S. A. Luo, S. M. Kozlov, F. Viñes, J. Limtrakul, D. G. Truhlar and F. Illas, J. Chem. Theory Comput., 2014, 10, 3832–3839 CrossRef CAS PubMed.
- J. Greeley and M. Mavrikakis, J. Phys. Chem. B, 2005, 109, 3460–3471 CrossRef CAS PubMed.
- P. Sautet and F. Cinquini, ChemCatChem, 2010, 2, 636–639 CrossRef CAS.
- E. J. Candes and M. B. Wakin, IEEE Signal Process. Mag., 2008, 25, 21–30 Search PubMed.
- L. J. Nelson, G. L. W. Hart, F. Zhou and V. Ozoliņš, Phys. Rev. B, 2013, 87, 035125 CrossRef.
- T. A. R. Purcell, M. Scheffler, C. Carbogno and L. M. Ghiringhelli, J. Open Source Softw., 2022, 7, 3960 CrossRef.
- L. Foppa, T. A. R. Purcell, S. V. Levchenko, M. Scheffler and L. M. Ghiringhelli, Phys. Rev. Lett., 2022, 129, 055301 CrossRef CAS PubMed.
- M. D. Morris, Technometrics, 1991, 33, 161–174 CrossRef.
- I. M. Sobol, Mathematical Modelling and Computational Experiment, 1993, 1, 407–414 Search PubMed.
- M. Affenzeller, S. M. Winkler, G. Kronberger, M. Kommenda, B. Burlacu and S. Wagner, in Gaining Deeper Insights in Symbolic Regression, ed. R. Riolo, J. H. Moore and M. Kotanchek, Springer New York, New York, NY, 2014, pp. 175–190 Search PubMed.
- R. M. Filho, A. Lacerda and G. L. Pappa, Explaining Symbolic Regression Predictions, 2020 IEEE Congress on Evolutionary Computation (CEC), 2020, pp. 1–8 Search PubMed.
- S. Kucherenko, S. Tarantola and P. Annoni, Comput. Phys. Commun., 2012, 183, 937–946 CrossRef CAS.
- E. C. Norton, B. E. Dowd and M. L. Maciejewski, JAMA, 2019, 321, 1304–1305 CrossRef PubMed.
- G. S. I. Aldeia and F. O. de França, Genet. Program. Evolvable Mach., 2022, 23, 309–349 CrossRef.
- A. Meurer, C. P. Smith, M. Paprocki, O. Čertík, S. B. Kirpichev, M. Rocklin, A. Kumar, S. Ivanov, J. K. Moore, S. Singh, T. Rathnayake, S. Vig, B. E. Granger, R. P. Muller, F. Bonazzi, H. Gupta, S. Vats, F. Johansson, F. Pedregosa, M. J. Curry, A. R. Terrel, Š. Roučka, A. Saboo, I. Fernando, S. Kulal, R. Cimrman and A. Scopatz, PeerJ Comput. Sci., 2017, 3, e103 CrossRef.
- L. Foppa and M. Scheffler, arXiv, 2026, preprint arXiv:2604.08122 Search PubMed.
- P. A. Sheth, M. Neurock and C. M. Smith, J. Phys. Chem. B, 2005, 109, 12449–12466 CrossRef CAS PubMed.
- T. Yang, Y. Feng, R. Ma, Q. Li, H. Yan, Y. Liu, Y. He, J. T. Miller and D. Li, ACS Appl. Mater. Interfaces, 2021, 13, 706–716 CrossRef CAS PubMed.
- W. Ludwig, A. Savara, K. H. Dostert, and S. Schauermann, Subsurface Hydrogen Diffusion into Pd Nanoparticles: Role ofLow-Coordinated Surface Sites and Facilitation by Carbon, January 31, 2012 (Web: Jan 3, 2012), DOI:10.1021/jp209033s.
- D. Teschner, J. Borsodi, A. Wootsch, Z. Révay, M. Hävecker, A. Knop-Gericke, S. D. Jackson and R. Schlögl, Science, 2008, 320, 86–89 CrossRef CAS PubMed.
- D. Teschner, Z. Révay, J. Borsodi, M. Hävecker, A. Knop-Gericke, R. Schlögl, D. Milroy, S. D. Jackson, D. Torres and P. Sautet, Angew. Chem., Int. Ed., 2008, 47, 9274–9278 CrossRef CAS PubMed.
- F. Studt, F. Abild-Pedersen, T. Bligaard, R. Z. Sørensen, C. H. Christensen and J. K. Nørskov, Angew. Chem., Int. Ed., 2008, 47, 9299–9302 CrossRef CAS PubMed.
- B. Yang, R. Burch, C. Hardacre, G. Headdock and P. Hu, J. Catal., 2013, 305, 264–276 CrossRef CAS.
- D. Torres, F. Cinquini and P. Sautet, J. Phys. Chem. C, 2013, 117, 11059–11065 CrossRef CAS.
- B. Yang, R. Burch, C. Hardacre, P. Hu and P. Hughes, Surf. Sci., 2016, 646, 45–49 CrossRef CAS.
- Y. Liu, F. Fu, A. McCue, W. Jones, D. Rao, J. Feng, Y. He and D. Li, ACS Catal., 2020, 10, 15048–15059 CrossRef CAS.
- W. Dong, V. Ledentu, P. Sautet, A. Eichler and J. Hafner, Surf. Sci., 1998, 411, 123–136 CrossRef CAS.
- O. E. Brandt Corstius, J. van der Hoeven, G. Sunley and P. de Jongh, J. Catal., 2023, 427, 115103 CrossRef CAS.
- G. Totarella, J. W. de Rijk, L. Delannoy and P. E. de Jongh, ChemCatChem, 2022, 14, e202200348 CrossRef CAS.
- S. Curtarolo, W. Setyawan, G. L. Hart, M. Jahnatek, R. V. Chepulskii, R. H. Taylor, S. Wang, J. Xue, K. Yang, O. Levy, M. J. Mehl, H. T. Stokes, D. O. Demchenko and D. Morgan, Comput. Mater. Sci., 2012, 58, 218–226 CrossRef CAS.
- M. Scheffler, M. Aeschlimann, M. Albrecht, T. Bereau, H.-J. Bungartz, C. Felser, M. Greiner, A. Groß, C. T. Koch, K. Kremer, W. E. Nagel, M. Scheidgen, C. Wöll and C. Draxl, Nature, 2022, 604, 635–642 CrossRef CAS PubMed.
- L. Barroso-Luque, M. Shuaibi, X. Fu, B. M. Wood, M. Dzamba, M. Gao, A. Rizvi, C. L. Zitnick and Z. W. Ulissi, Open Materials 2024 (OMat24) Inorganic Materials Dataset and Models, arXiv, 2024, preprint, arXiv.2410.12771, DOI:10.48550/arXiv.2410.12771.
- A. S. Nair, L. Foppa and M. Scheffler, npj Comput. Mater., 2025, 11, 150 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |