Open Access Article
Open Access ArticleCarbon dioxide hydrogenation on copper and nickel catalysts via a conformal sampling approach
Thantip Roongcharoen* and
Alessandro Fortunelli *
*
CNR-ICCOM, Consiglio Nazionale delle Ricerche, via Giuseppe Moruzzi 1, Pisa 56124, Italy. E-mail: thantip.roongcharoen@cnr.it; alessandro.fortunelli@cnr.it
First published on 18th March 2026
Abstract
We present a computational workflow, the Conformal Sampling of Catalytic Processes (CSCP) approach, and its application to the case of heterogeneous hydrogenation/reduction of carbon dioxide (CO2) on copper and nickel catalysts. CO2 activation is of critical importance for a sustainable global future and one of the major reactions for which sustainable routes must be found urgently. We use the fcc(100) facet of Cu as a worked-out case, and exhaustively derive at the DFT level its reaction mechanisms. We then apply CSCP to first derive a Machine Learning Interatomic Potential (MLIP) for this initial system, and then carry over the knowledge derived on Cu(100) to a pure monometallic facet, Ni(100), to rapidly derive a MLIP for this different system, so as to test the transferability of the approach. The accuracy of the so-derived MLIPs is excellent, predicting both reaction energies and energy barriers for all the mechanistic steps of this complex reaction diagram with consistent and uniform accuracy, with a maximum discrepancy of 0.05 eV for Cu and 0.03 eV for Ni. In the exception cases in which this discrepancy is larger, we show and rationalize that this is due to a change in the reaction mechanism, where the MLIP simulations explore pathways different from the reference DFT ones, without however prejudicing their absolute accuracy. The CSCP-MLIPs are thus shown to be able to assure a transferability en par with the best physics-based models, provide alternative atomistic mechanisms of catalytic processes, and offer themselves as a tool for catalyst rational design on a process societally relevant and exhibiting significant catalytic complexity.
1. Introduction
One major challenge in computational materials discovery is how to exploit existing knowledge accumulated on worked-out systems. This comes with a two-fold critical purpose: (i) avoid reconsidering previous negative outcomes, (ii) use available information to orient, thus accelerate, the search. This need is keenly felt in the search for new catalysts for sustainable chemical processes,1 because: (a) the ensemble of potential candidates (growing exponentially with the set of elements allowed in the search chemical space) is practically unlimited, while (b) the number of actual solutions is minute (as proven by the fact that none of the mass-scale catalytic processes that are now in place are sustainable). The recent developments in Artificial Intelligence promise to contribute decisively to address this need with their ability to process huge datasets, once AI tools are properly adjusted to the specifics of the field. Large Language Models and Machine Learning are two major avenues that AI offers to chemical catalysis. In particular, a sub-topic in atomistic modeling is represented by Machine Learning Interatomic Potentials (MLIPs),2–10 that, by interpolating potentially predictive but computationally demanding (e.g., first-principles) potential energy surfaces (PES) with analytic functions, enable one to dramatically accelerate the modelling and computational prediction of catalytic processes (energy barriers, rates of reaction mechanisms, etc.).Deriving accurate MLIPs for heterogeneous catalysis is the theme of the present contribution, grounded on our recently developed Conformal Sampling of Catalytic Processes (CSCP) approach.8,11 CSCP proposes a practical route to exploit accumulated information on catalytic mechanisms of worked-out systems to accelerate sampling of the same or similar mechanisms of new systems (note that the reaction mechanisms need not to be identical, since CSCP seems to be able to cover all chemically equivalent paths, en par with the best physics-based interatomic potentials11). A rigorous theorem12 is rephrased within the context to state that structures (atomistic geometries) are invariant up to 2nd order in a perturbation theory expansion to a change in the catalyst stoichiometry. This implies that the “catalytic funnel” along which a reaction occurs for a given catalyst can be conformally transferred into the catalytic funnel of a different (novel, unknown) catalyst. The structural database derived to train the reactive MLIP for the worked-out system can thus be carried over to the new system as a first robust approximation (modulus affine transformation to account for distance rescaling8,11), and then quickly refined with a few steps of active learning (1–2 are sufficient in the cases treated so far), to achieve an absolute accuracy of ∼0.03 eV on barrier prediction with respect to the generating first-principles approach. Such sub-chemical accuracy corresponds to the level of accuracy targeted to ensure that the MLIP surrogate model can be employed with confidence in operational catalyst screening. Additionally, since the training structural database is available a priori, the single-point calculations, generating energies and forces over the database geometries as needed to train the MLIP pertaining to the new system, are fully disconnected, thus perfectly parallelizable on HPC machines, maximizing computational throughput8,11,13,14 to an estimated ∼1 hour for screening a new system on a properly equipped HPC system. Importantly, CSCP has been extended with uniform performance to bimetallic systems,10 a necessary step toward the design of multi-metallic catalysts.
The practical details of the CSCP approach, including the latest advances, such as the use of extrapolation techniques derived from the theory of convergence of numerical series and the adjustments for its extension to bimetallic catalysts, can be found in the original ref. 8, 10 and 11 (selected information can be found in Sections 2 and 5). In our original work, we investigated methanol (CH3OH) decomposition on Pt fcc(100) and fcc(111) facets as worked-out catalysis, and then conformally transferred its reaction paths onto the same facets of eight different catalysts (Fe, Co, Ni, Cu, Pd, Ag, Au, and Pt itself at a different adsorbate coverage). In the present work, we test and demonstrate the CSCP potentialities focusing on a different heterogeneous catalytic process, the thermal hydrogenation/reduction of carbon dioxide (CO2), or t-CO2RR, of critical importance for a sustainable global future. We will use the fcc(100) facet of Cu as a worked-out case, and exhaustively derive at the DFT level its reaction mechanisms. We will then apply CSCP to carry over the knowledge derived on Cu(100) to a pure monometallic facet, Ni(100), to test the transferability of the approach. We thus go beyond our original work in terms of catalytic complexity, and demonstrate that the CSCP approach works equally well on such a societally relevant process.
The article is organized as follows. In Section 2, we provide computational details. In Section 3, we derived the reaction energy diagram for CO2RR on the (100) facet of fcc Cu, and then discuss how to apply CSCP to transfer the so-derived information to a transition metal (TM = Ni). In Section 3, conclusions are summarized.
2. Methodology
The conformal sampling of catalytic processes (CSCP) is a computational workflow for building machine-learning interatomic potentials (MLIPs) to explore the potential energy landscape of complex catalytic reactions, such as CO2 hydrogenation on metal catalysts, with the same accuracy as first-principles approaches, but at a much reduced computational cost, and with a high throughput. CSCP provides a strategy to construct the training database for MLIPs by leveraging knowledge from previously studied cases (e.g., previous DFT computational investigations), enabling a rapid generation of the database for new catalytic systems. The approach is based on a theorem12 stating that, in a system governed by coulombic interactions, geometries (not energies) are invariant up to second order in a perturbation expansion upon changing the system. CSCP exploits this theorem by defining the configurations in the initial database for a new system via proper transformations of the configurations previously investigated for the known system (in detail, we use rescaling or affine transformations). In other words, CSCP conformally transfers the “catalytic funnel” along which a reaction occurs on a given catalyst into the catalytic funnel of a novel catalyst, see Fig. 1 for a depiction of representative configurations along the catalytic funnel of the present reaction. Moreover, CSCP exploits the theory of series convergence (and the fact that most regularization methods are engineered to achieve monotonic convergence) to derive an optimal sampling of available computational data from both geometry relaxations and reaction path (NEB) simulations via extraction and extrapolation techniques. Due to the fact that the database is pre-generated, CSCP can optimally exploit parallelization and can thus achieve very high throughput. Due to the fact that the database is grounded on physical principles, the workflow achieves directly usable MLIPs with very few embedded active-learning cycles (only two in the examples studied so far), also maximizing throughput, and achieves energy predictions of high accuracy. The step-by-step details of the workflow are described in the following subsections and in Section 5.2.1 CSCP applied to the CO2 hydrogenation reaction
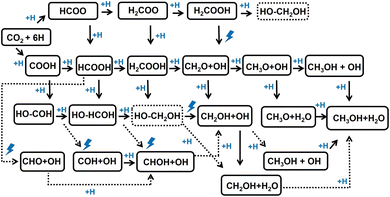
In this work, we applied the CSCP workflow to CO2 hydrogenation to CH3OH and H2O [CO2 + 3H2 → CH3OH + H2O] on metal surfaces. As shown in Scheme 1, the reaction proceeds through a series of hydrogenation steps followed by dissociation of CHxOy species once the intermediates become saturated. CO2 hydrogenation is a complex reaction with multiple competitive pathways that feature bond-cleavage and dissociation events. Metal surfaces offer multiple adsorption sites for intermediates (CO2, COOH, HCOO, CH3OH, CH2OH–OH, CHOH–OH, H2O, OH, H, and others), such as top, bridge, and hollow positions (see Fig. S1), which illustrates the complexity of the reaction. The idea of CSCP is to first generate a comprehensive database describing complex chemical reactions on a worked-out metal catalyst, and then transfer the knowledge from this system to create a new dataset and MLIP potentials for other catalysts, accelerating the construction of reaction energy diagram for the new system using MLIPs. Cu is a well-known catalyst for CO2 hydrogenation to CH3OH.15–17 Several previous DFT works examined the reaction mechanism and the performance of Cu catalysts on a bare Cu surface. Each reaction step foresees a H atom adding to a single partially hydrogenated Hx–COy species. In this work, we used the (100) facet of Cu as a worked-out case: this clearly represents a proof-of-principle case, since the role of the oxide support in determining the performance of the industrial catalyst has been shown to be critical18–24 (see Section 3.1 for further discussion).We then examined the reaction mechanism and energetics of the reaction by placing six H atoms and CO2 on a Cu surface, and allowing the reaction to proceed from CO2 + 6H to CH3OH + H2O. We note that in our previous studies8,11 we focused on CH3OH decomposition, a key pathway in the CH3OH aqueous-phase reforming (APR) reaction for hydrogen production25 [CH3OH + H2O → CO2 + 3H2]. The CSCP scheme was evaluated along the C–H cleavage path [CH3OH → CH2OH + H → CHOH + 2H → COH + 3H → CO + 4H], as well as the C–O and O–H cleavage paths. This motivated us to apply the CSCP procedure to CO2 hydrogenation [CO2 + 3H2 → CH3OH + H2O], the reverse process of CH3OH-APR, to provide a comprehensive assessment of CSCP performance for both C–H and C–O bond cleavage and hydrogenation reactions.
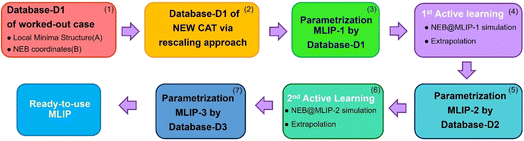
The CSCP workflow in Fig. 2 illustrates the sequential steps for constructing the database-DX and MLIP-X potentials for CO2 hydrogenation. As mentioned, we select the Cu-fcc(100) facet as the worked-out system, for which we also derive a database and a MLIP using the CSCP strategy as a test of the quality and accuracy of the approach, and the Ni-fcc(100) facet as the newly investigated system to evaluate the performance of the CSCP protocol to transfer information from Cu and generate a MLIP for an unknown system. The first step, shown in the pink block in Fig. 2, involves constructing the first database (Database-D1) for the Cu worked-out case using DFT calculations. As customary, we perform two types of DFT simulations: (1) optimization of intermediate species and (2) Nudged Elastic Band (NEB) simulations for transition-state searches. Details of the database-D1 for Cu-fcc(100) are provided in Section 5, Computational details. In the next step, shown in the yellow block in Fig. 2, the CSCP workflow leverages knowledge from the worked-out case, transferring information as a set of coordinates along the reaction pathways tuned via a rescaling approach to build an initial database for the new Ni catalytic system. The rescaled structures are then divided into two sets. The first set (Database A) contains rescaled coordinates of optimized intermediate species [e.g., CO2, COOH, CHOH, HCOO, CH2OH–OH, OH] (see Scheme 1). These coordinates are then DFT-optimized to determine the local minima for Ni. The second dataset (Database B) contains rescaled coordinates along the elementary reaction steps of CO2 hydrogenation, extracted from NEB@DFT calculations performed on the Cu worked-out case. NEB@DFT is highly demanding in terms of computational and human resources, a step which we bypass for the Ni system by exploiting knowledge from the Cu case. The resulting rescaled NEB coordinates are processed through single-point DFT calculations to obtain energies and forces. Database A and Database B are then combined to form Database D1 for Ni, and fed into the parametrization of MLIP-1.
The CSCP employs an equivariant message-passing neural network (MPNN) in the form of the machine-learning MACE MLIP model26 that is trained on the given database, which consists of the XYZ reaction coordinates together with their forces and energies. Details of the MACE parameters are provided in Table S1. After the first MLIP parametrization (producing MLIP-1), the procedure proceeds through two steps of Active Learning (in the magenta, light and dark blue and dark green blocks in Fig. 2) to generate new databases and retrain more accurate MLIP-X potentials, increasing reliability and accuracy. The sampling techniques to produce newly relevant coordinates in the active-learning steps are: (1) running NEB@MLIP-X (X = 1 or 2) for all elementary reaction steps (see Scheme 1) and properly extracting the coordinates from the simulations, and (2) performing extrapolation techniques to generate new coordinates along the reaction pathway. The extrapolation operates on two sets of XYZ coordinates via the equation: N = B + (B − A) = 2B − A, where N are the extrapolated coordinates, and A and B are XYZ coordinates generated from different NEB@MLIP-X iterations. We applied two types of extrapolations: vertical extrapolation, which operates on NEB coordinates obtained at different iterations using the same MLIP-X potential, and horizontal extrapolation, which operates on NEB coordinates generated from different MLIP-X potentials (X = 1 or 2) at the same iteration. Details of the extrapolation technique are described in our previous work.11 We applied extrapolation across all NEB@MLIP-X steps. We note here that the databases (D1, D2, D3) built within CSCP are incremental: new datasets are gathered and merged into the datasets from the earlier stages. After two Active Learning steps and the parametrizations of MLIP-2 and MLIP-3, we assess the performance of the final MLIP-3 for the Cu and Ni catalysts in terms of both internal technical accuracy (RMSE, internal test) and energy prediction with respect to the generating first-principles (DFT) model (external test). The results are presented in Section 3.
Note that we define a threshold of 0.03 eV as our targeted accuracy. This definition corresponds to sub-chemical accuracy (<1 kcal mol−1), or ≈kBT at room temperature, or a factor of ≈3 in kinetic rates at room temperature (a factor <2 at 600 K), and is below the usual accuracy of DFT (around 2 kcal mol−1) as well as typically also of post-Hartree-Fock or post-DFT computational methods. Empirically, we found that in the cases so far investigated (methanol decomposition and CO2 reduction) two active learning steps are sufficient to achieve the targeted accuracy. This is consistent with the principle that the initial, conformally derived database (D1) already contains the essential information on the reaction path on the new catalyst, so that the number of AL steps is reduced to a minimum. In the CSCP protocol in productions runs, control parity plots are performed to test that the quality of the CSCP MLIPs does not degrade for the given catalytic process.
3. Results and discussion
Having two cases for which we derived a sequence of database-DX sets and MLIP-X functions – Cu-fcc(100) and Ni-fcc(100) – we test and provide information for assessing the accuracy of the CSCP approach for each of them separately. We first evaluate the root-mean-square errors (RMSE) of energy (ERMSE) and forces (FRMSE) of the MLIP models with respect to the DFT reference data to assess the MLIP-3 performance. During the parametrization or fitting steps, the Cu and Ni datasets are randomly split into two groups (as customary): a training set (95%) and a validation set (5%). We list in Table 1 the ERMSE and FRMSE for training and validation sets obtained for Cu and Ni. The results indicate excellent RMSE for forces and energies for both datasets on the Cu and Ni systems. The ERMSE values of Cu and Ni for the validation set are 0.7 and 0.6 meV per atom, respectively, while FRMSE of Cu and Ni for the validation set are 10.3 and 10.0 meV per Å per atom, respectively. We observe basically no difference in the RMSE values between the training and validation sets, implying that the dataset is not overfitted. We next used MLIP-3 to perform NEB calculations (NEB@MLIP-3) for all the reaction steps. Three NEB@MLIP-3 calculations were carried out for each reaction step using 12, 24, and 48 intermediate images to assess the trend in energy barriers (Ea) predicted by the NEB@MLIP-3 simulations. The results of NEB@MLIP-3 for the Cu and Ni systems are reported in Tables 2 and 3, respectively. Since the CSCP approach is still in a proof-of-principle stage, we assessed the accuracy of the MLIP Ea predictions by comparing them with actual calculations of the DFT barrier benchmarks for all reaction steps on both Cu and Ni catalysts (naturally, in actual production runs we check MLIP accuracy only via selected parity plots: in operative studies, the comprehensive validation we perform here can be omitted or performed only on a limited number of representative reaction steps). The variation in energy prediction was quantified using ΔE = |ECI-NEB@DFT − ENEB@MLIP| where the ECI-NEB@DFT and ENEB@MLIP are the energy barriers predicted from Climbing Image (CI)-NEB@DFT and NEB@MLIP-3 calculations, respectively. Note that, to greatly speed up the first-principles calculations on Ni(100), we used the coordinates generated from the NEB@MLIP-3 simulations as starting coordinates for the NEB@DFT calculations. Indeed, the use of intermediate images from NEB@MLIP-3 in the NEB@DFT simulations proved very useful, as both NEB@DFT and CI-NEB@DFT starting from them converged quickly.| Cu (100) | Ni (100) | |||
|---|---|---|---|---|
| ERMSE | FRMSE | ERMSE | FRMSE | |
| MLIP-3 | Train: 0.5 | Train: 9.1 | Train: 0.5 | Train: 9.4 |
| Valid: 0.7 | Valid: 10.3 | Valid: 0.6 | Valid: 10.0 | |
| Step | Reaction step | 12-DFT | 12-MLIP-3 | 24-MLIP-3 | 48-MLIP-3 |
|---|---|---|---|---|---|
| a Exception cases where ΔEa > 0.05 eV, for which parity plots were further analyzed. | |||||
| (a) | CO2 + H → HCOO | 0.803 | 0.859 (0.06) | 0.915 (0.11) | 0.820 (0.02) |
| (b) | HCOO + H → H2COO | 1.096 | 1.155 (0.06) | 1.148 (0.05) | 1.137 (0.04) |
| (c) | H2COO + H → H2COOH | 0.810 | 0.870 (0.06) | 0.832 (0.02) | 0.854 (0.04) |
| (d) | H2COOH + H → HO–H2C–OH | 0.917 | 0.880 (0.04) | 0.832 (−0.09) | 0.801 (−0.12)a |
| (e) | CO2 + H → COOH | 1.936 | 2.089 (0.15)a | 1.788 (−0.15)a | 1.863 (0.07)a |
| (f) | COOH + H → HCOOH | 0.521 | 0.539 (0.02) | 0.531 (0.01) | 0.510 (0.01) |
| (g) | HCOOH + H → H2COOH | 0.565 | 0.739 (0.17) | 0.658 (0.09) | 0.575 (0.01) |
| (h) | H2COOH → CH2O + OH | 0.738 | 0.750 (0.01) | 0.736 (0.00) | 0.714 (0.02) |
| (i) | CH2O + OH → CH3O + OH | 0.414 | 0.321 (0.09)a | 0.340 (0.07)a | 0.340 (0.07)a |
| (j) | CH3O+2H + OH → CH3OH + H + OH | 0.670 | 0.704 (0.03) | 0.655 (0.02) | 0.656 (0.01) |
| (k) | HCOO + H → HCOOH | 0.846 | 0.827 (0.02) | 0.814 (0.03) | 0.810 (0.04) |
| (l) | HCOOH + H → H2COOH | 1.033 | 1.024 (0.01) | 1.038 (0.01) | 1.035 (0.00) |
| (m) | HO–CH2OH → OH + CH2OH | 1.403 | 1.482 (0.08) | 1.414 (0.01) | 1.314 (0.09)a |
| (n) | CH3OH + OH + H → CH3OH + H2O | 0.823 | 0.768 (0.06)a | 0.729 (0.09)a | 0.705 (0.12)a |
| (o) | COOH + H → HO–COH | 0.716 | 0.794 (0.08) | 0.788 (0.07) | 0.761 (0.05) |
| (p) | HO–COH + H → HO–CHOH | 0.667 | 0.764 (0.10) | 0.698 (0.03) | 0.710 (0.04) |
| (q) | HCOOH + H → HO–CHOH | 1.160 | 1.071 (0.09) | 1.276 (0.12) | 1.179 (0.02) |
| (r) | H2COOH + H → HO–CH2OH | 0.974 | 0.860 (0.11) | 0.926 (0.05) | 0.916 (0.06) |
| (s) | HO + CH2O + H → CH2OH + OH | 0.605 | 0.568 (0.04) | 0.616 (0.01) | 0.604 (0.00) |
| (t) | HO–COH → HO + COH | 1.313 | 1.397 (0.08) | 1.347 (0.03) | 1.305 (0.01) |
| (u) | HO + COH → OH + CHOH | 0.344 | 0.311 (0.03) | 0.316 (0.03) | 0.306 (0.04) |
| (v) | OH + CHOH + H → OH + CH2OH | 0.150 | 0.154 (0.00) | 0.172 (0.02) | 0.157 (0.01) |
| (w) | OH–CHOH → OH + CHOH | 0.305 | 0.320 (0.02) | 0.298 (0.01) | 0.298 (0.01) |
| (x) | HO–CH2OH + H→ H2O + CH2OH | 1.327 | 1.346 (0.02) | 1.211 (−0.12) | 1.116 (−0.16)a |
| (y) | HCOOH → HCO + OH | 1.063 | 1.094 (0.03) | 1.036 (0.03) | 1.008 (0.05) |
| (z) | HCO + OH → HCOH + OH | 0.605 | 0.507 (0.01) | 0.553 (0.05) | 0.574 (0.03) |
| (aa) | CH2OH + OH → CH2OH + H2O | 0.687 | 0.698 (0.01) | 0.672 (0.02) | 0.675 (0.01) |
| (bb) | CH2OH + H2O → CH3OH + H2O | 0.501 | 0.738 (0.24) | 0.608 (0.11) | 0.545 (0.04) |
| (cc) | CH3O + OH → CH3O + H2O | 0.869 | 0.661 (0.21) | 0.787 (0.08) | 0.835 (0.03) |
| (dd) | H2O + CH3O + H → CH3OH + H2O | 0.798 | 0.731 (0.07) | 0.767 (0.03) | 0.739 (0.06) |
| Step | Reaction step | 12-DFT | 12-MLIP-3 | 24-MLIP-3 | 48-MLIP-3 |
|---|---|---|---|---|---|
| a Exception cases where ΔEa > 0.05 eV, for which parity plots were further analyzed. | |||||
| (a) | CO2 + H → HCOO | 0.928 | 1.099 (0.17) | 1.036 (0.11)a | 0.979 (0.05)a |
| (b) | HCOO + H → H2COO | 1.206 | 1.231 (0.02) | 1.234 (0.03) | 1.231 (0.02) |
| (c) | H2COO + H → H2COOH | 1.443 | 1.438 (0.00) | 1.417 (0.03) | 1.431 (0.01) |
| (d) | H2COOH + H → HO–H2C–OH | 1.126 | 1.186 (0.06) | 1.120 (0.01) | 1.117 (0.01) |
| (e) | CO2 + H → COOH | 1.504 | 1.420 (0.08) | 1.494 (0.01) | 1.475 (0.03) |
| (f) | COOH + H → HCOOH | 0.00 | 0.00 | 0.00 | 0.00 |
| (g) | HCOOH + H → H2COOH | 0.940 | 1.082 (0.14) | 0.974 (0.03) | 0.811 (0.13)a |
| (h) | H2COOH→CH2O + OH | 0.824 | 0.843 (0.02) | 0.814 (0.01) | 0.814 (0.01) |
| (i) | CH2O + OH → CH3O + OH | 0.561 | 0.557 (0.00) | 0.573 (0.01) | 0.583 (0.02) |
| (j) | CH3O+2H + OH → CH3OH + H + OH | 0.922 | 0.907 (0.01) | 0.941 (0.02) | 1.109 (0.19)a |
| (k) | HCOO + H → HCOOH | 0.939 | 0.949 (0.01) | 0.952 (0.01) | 0.956 (0.02) |
| (l) | HCOOH + H → H2COOH | 1.018 | 1.086 (0.01) | 1.105 (0.01) | 1.114 (0.02) |
| (m) | HO–CH2OH → OH + CH2OH | 1.010 | 0.989 (0.02) | 0.974 (0.04) | 0.964 (0.05) |
| (n) | CH3OH + OH + H → CH3OH + H2O | 1.014 | 1.059 (0.05) | 1.063 (0.05) | 1.045 (0.03) |
| (o) | COOH + H → HO–COH | 1.230 | 1.190 (0.04) | 1.180 (0.05) | 1.199 (0.03) |
| (p) | HO–COH + H → HO–CHOH | 1.019 | 1.082 (0.06) | 1.043 (0.02) | 0.973 (0.05) |
| (q) | HCOOH + H → HO–CHOH | 1.219 | 1.239 (0.08) | 1.295 (0.02) | 1.337 (0.02) |
| (r) | H2COOH + H → HO–CH2OH | 1.050 | 0.992 (0.06) | 1.076 (0.03) | 1.073 (0.02) |
| (s) | HO + CH2O + H → CH2OH + OH | 0.750 | 0.816 (0.07) | 0.781 (0.03) | 0.801 (0.05) |
| (t) | HO–COH → HO + COH | 0.705 | 0.741 (0.04) | 0.680 (0.02) | 0.663 (0.04) |
| (u) | HO + COH → OH + CHOH | 1.516 | 1.514 (0.00) | 1.503 (0.01) | 1.477 (0.04) |
| (v) | OH + CHOH + H → OH + CH2OH | 0.581 | 0.612 (0.03) | 0.613 (0.03) | 0.617 (0.04) |
| (w) | OH–CHOH → OH + CHOH | 0.113 | 0.111 (0.00) | 0.086 (0.03) | 0.071 (0.04) |
| (x) | HO–CH2OH + H → H2O + CH2OH | 1.529 | 1.722 (0.19) | 1.359 (0.16) | 1.300 (0.23) |
| (y) | HCOOH → HCO + OH | 1.231 | 1.166 (0.07) | 1.177 (0.05) | 1.202 (0.03) |
| (z) | HCO + OH → HCOH + OH | 1.059 | 1.011 (0.05) | 1.074 (0.02) | 1.077 (0.02) |
| (aa) | CH2OH + OH → CH2OH + H2O | 0.926 | 0.965 (0.04) | 0.950 (0.02) | 0.939 (0.01) |
| (bb) | CH2OH + H2O → CH3OH + H2O | 0.591 | 0.565 (0.03) | 0.620 (0.03) | 0.624 (0.03) |
| (cc) | CH3O + OH → CH3O + H2O | 1.023 | 1.154 (0.13) | 1.138 (0.12) | 1.092 (0.07) |
| (dd) | H2O + CH3O + H → CH3OH + H2O | 0.893 | 0.804 (0.09) | 0.898 (0.00) | 0.866 (0.03) |
We first assess the MLIP performance for the Cu worked-out case. The energy differences in Ea predicted by NEB@DFT and NEB@MLIP (ΔE) for each reaction step are listed in parentheses in Table 2. The Ea simulated by CI-NEB@DFT for each reaction step is listed in the third column (NEB@DFT). The Ea values predicted by NEB@MLIP using 12, 24, and 48 intermediate images are listed in the columns titled 12-MLIP-3, 24-MLIP-3, and 48-MLIP-3, respectively. The NEB@MLIP-3 simulations for the Cu worked-out case show excellent performance, with ΔE values below 0.05 eV for most steps.
Let us now discuss the exception steps, where ΔE exceeds 0.05 eV. The exception steps occur at steps (d), (e), (i), (m), (n) and (x); the corresponding reaction formulas are provided in Table 2.
We can distinguish two classes of discrepancies:
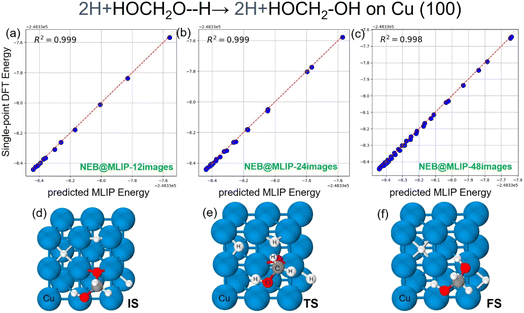
In the first class, we observe that NEB@MLIP predicts lower barriers than NEB@DFT. To further examine the accuracy of the approach, in these cases we produced further test data as reported in Fig. 3. In particular, we performed single-point DFT calculations on the reaction coordinates generated from the NEB@MLIP-3 simulations and generated parity plots. Taking reaction step (d) as an example, this step corresponds to the hydrogen protonation of the H2COOH species [H2COOH + H→ HO–H2C–OH]. The parity plots comparing the energies obtained from NEB@DFT and NEB@MLIP are shown in Fig. 3a–c. These plots exhibit clear linear trends between the two sets of energies, with R2 values of 0.998–0.999 for the three NEB@MLIP simulations using 12, 24, and 48 images (see Fig. 3a–c), respectively, indicating high accuracy in the energy predictions of coordinates along the reaction pathways. The optimized structures along the reaction pathway, as shown in Fig. 3d and e, illustrate that this reaction step involves both the bending and migration of the bulky H2COOH species across the adsorption sites of the Cu surface (from bridge to hollow), together with the migration of a single hydrogen atom that subsequently forms a bond with the O atom of the H2COOH species. Given the efficiency of NEB@MLIP simulations, using 48 images is very fast and provides a more precise optimization of the energy profile along the reaction pathway, with a clear advantage over NEB@DFT, which relies on a constrained number of images due to computational resource limitations. For the other exception cases (e, i, m, n, x), their parity plots are shown in Fig. S2–S7 of the SI. We always found linear correlation values R2 of 0.999–0.998 between MLIP and DFT energies for all NEB simulations, regardless of the number of intermediate images used (12, 24, or 48), indicating the high accuracy of MLIP in energy predictions.
To gain deeper insight into the exceptional cases in the MLIP-3 accuracy analysis, we illustrate in Fig. S8 the reaction energy profile of CH3OH + OH + H → CH3OH + H2O on Cu(100) as a function of reaction coordinates. We found that the water formation step on Cu(100), in the presence of adsorbed CH3OH, corresponds to a composite reaction pathway. First, CH3OH migrates on the surface, induced by nearby OH species, due to hydrogen bonding between the H atom of CH3OH and the OH species (see Fig. S8(d)–(f)). The OH species then reacts with a single H atom adsorbed on the surface to form H2O, while CH3OH acts as a spectator throughout the reaction. Such composite reaction paths often occur on metal surfaces, especially at high coverage. Note the energies in the range −9.1 to −8.8 eV in Fig. S4b–c may appear redundant, or highly correlated, with a risk of data leakage. However, the configurations with similar energies along the reaction pathway reflect the migration of species on the surface and are structurally distinct, in keeping with our rigorous construction of the fitting database. The energy profiles shown in Fig. S8a–c indicate that the final MLIP-3 potential for Cu accurately reproduces the DFT energies, regardless of the number of intermediate images used in the calculations. We did not find examples of the second class of discrepancies for Cu, so we discuss them in the next paragraphs about Ni.
We next evaluate the performance of MLIP-3 for the Ni catalyst. Table 3 lists the Ea values predicted by NEB@MLIP-3 for all the reaction steps, with ΔE values in parentheses representing the energy differences between the NEB@DFT and NEB@MLIP-3 calculations. The results demonstrate that MLIP-3, generated within the CSCP scheme, exhibits outstanding performance in NEB@MLIP calculations and shows excellent agreement with the DFT energy benchmark, with ΔE maximum values as low as 0.03 eV.
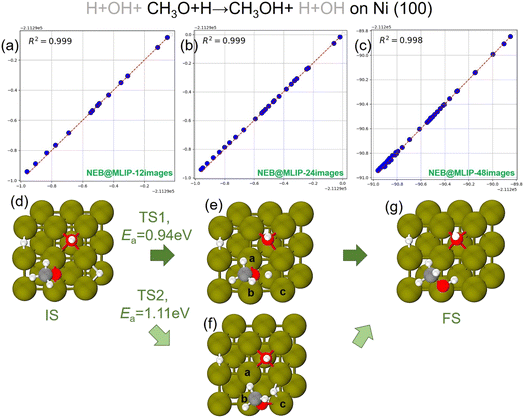
The four exception cases are the reaction steps (a, g, j, and x). The most interesting case is the reaction step j [CH3O + 2H + OH → CH3OH + H + OH], which is an example of the second class of discrepancies we mentioned above, that we now discuss. We thus produced further test data as reported in Fig. 4. In particular, in Fig. 4a–c we report the parity plot for the reaction step j in Table 3. The ΔE values in columns 12-MLIP and 24-MLIP are 0.01 and 0.02 eV, respectively; however, a difference of +0.19 eV in the energy barrier is observed when NEB@MLIP is performed using 48 intermediate images. In other words, the energy barrier found by NEB@MLIP using 48 images is higher than the energy barrier found by NEB@DFT using 12 images or NEB@MLIP using 12 or 24 images. Examination of the parity plots indicates that all three cases exhibit R2 of 0.998–0.999, implying that MLIP accurately reproduces the DFT energies of the coordinates along the reaction pathway for the hydrogen protonation of CH3O to generate CH3OH on Cu with OH and H residues adsorbed on the surface. Further inspection of the reaction profiles and coordinates reveals that NEB@MLIP using 48 intermediate images gives a different reaction path compared with NEB@DFT (whereas NEB@MLIP simulations using 12 and 24 images produce reaction energy pathways that closely resemble those of NEB@DFT). In the case of NEB@MLIP with 48 images, at transition state 2 (TS2), the CH3O species first moves from the bridge site of (a and b) to the adjacent bridge site (b and c) in Fig. 4, while the activated hydrogen atom approaches the migrated CH3O species. The reaction thus occurs at a different active site on the Cu surface compared with TS1 from NEB@DFT, where only the hydrogen atom migrates toward the CH3O species at the site of (a and b) in Fig. 4. Fig. 5 compares the energy profile along the two alternative NEB paths, showing that they are clearly different. In our previous work on CH3OH decomposition,8,11 we observed in some cases a change in the reaction mechanism with a lower energy barrier when employing the final MLIP with a larger number of intermediate images than the NEB@DFT simulations, i.e., as in the Cu case above we only found examples of the first class of discrepancies. In contrast, the present example, reaction j [CH3O+2H + OH → CH3OH + H + OH], shows that using NEB@MLIP simulations with a larger number of intermediate images leads MLIP to explore alternative pathways with higher energy barriers. The MLIP is accurate, but the NEB simulation starting with a larger number of images (we remind that they are initially interpolated linearly from initial and final configurations) may converge to a different reaction path, usually with a lower energy barrier, but sometimes (counterintuitively) with a higher barrier if the initial and final configurations are too distant in the space of atomic coordinates. This discrepancy is thus not connected to the accuracy of the MLIP, but rather to the need to improve the NEB simulations using configurations with a lower number of images as the starting point of NEBs with a larger number of images, or better by performing more systematic reaction path searches than simple NEB simulations, e.g. using Reactive Global Optimization (RGO) approaches.27 Note in this connection that recent advances in Hessian estimates28 enable a very significant speed-up of reaction path search approaches such as RGO.
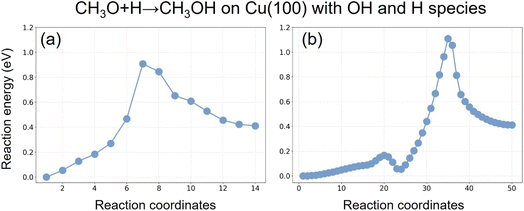 | ||
| Fig. 5 Reaction energy profiles for the hydrogen protonation of CH3O on Cu(100) with OH and H species on the metal surface, obtained using NEB@MLIP with (a) 12 and (b) 24 intermediate images. | ||
To further assess the MLIP-3, we performed ramped-temperature molecular dynamics (MD) simulations in the canonical (NVT) ensemble to evaluate the performance of MLIP-3 as a function of temperature. The adsorption configuration of HO–CH2OH + 2H on the Ni(100) surface was selected as a representative case. The MD simulation began with a heating phase, during which the temperature was gradually increased from 0 to 200 K over 80 ps. This was followed by an equilibration phase at a constant temperature of 200 K for an additional 50 ps. Subsequently, additional heating cycles were performed, in which the temperature was further increased in increments of 50 K over 80 ps, each followed by a 50 ps equilibration phase at the corresponding constant temperature reached in the preceding heating step. This procedure was repeated up to the final temperature of 350 K, as shown in Fig. S9. The two bottom layers of the Ni(100) slab were frozen, while the top three layers were allowed to move freely, consistent with the setup used in the NEB@MLIP-3 and DFT calculations. The input parameters used for the MD simulations are listed in Section S3 in the SI. The results demonstrate that the potential energy of the HO–CH2OH + 2H complex on the Ni(100) surface gradually and steadily increased as the temperature rose to 350 K, indicating the physical stability of the system and consistent energy behavior. From the trajectories sampled every 8000 steps during the heating phase and every 5000 steps during the equilibration phase, we observed that the adsorbate species moved physically on the catalytic surface. The H atoms, having a small mass, migrated faster, while the heavier HO–CH2OH species moved more slowly. The metal surface atoms also exhibited physical motion but remained within the plane without a significant distortion. The results suggest that MLIP-3 can serve as a reliable initial potential to study the effects of temperature on the energetics and catalytic behavior of the CO2 hydrogenation reaction under different temperature conditions.
Finally, we evaluate the reaction energies (Ereaction) using Ereaction = EFS − EIS where Ereaction is the reaction energy, and EFS and EIS are the total energies of the final and initial states in the NEB calculations, respectively. We list Ereaction for all elementary steps of CO2 hydrogenation on Ni(100) in Table S2. The results show that MLIP-3 accurately predicts the reaction energies, with differences in reaction energies (ΔE), less than 0.03 eV when compared with the values computed from NEB@DFT calculations.
3.1 Perspectives: validation and comparison with experiment and previous literature
In the present stage of development, the CSCP approach is still in a proof-of-principle phase. What is missing for it to become a fully operative tool for catalyst predictions to be compared with and potentially triggering experiments are two main points: (i) high-coverage effects; (ii) nanostructured or non-adiabatic catalyst configurations.Point (i) is related to the actual “resting state” of the catalytic surface under the given conditions. To make an example, under conditions of aqueous phase reforming of oxygenates (such as methanol), Pt-bases catalyst surfaces are typically covered by hydrogen adatoms.25 Similar phenomena have been reported in electrocatalysis, e.g., CO coverage in CO2 electro-reduction (CO2RR).29 Under these conditions, not the bare surfaces but surfaces exhibiting a full coverage of, e.g., hydrogen species should therefore be considered as the configurations in the worked-out case or in the predicted reaction energy diagrams to enable a proper, quantitative comparison with experiment. This is due to two reasons: first, there will be an energy price associated with making empty sites for incoming reactants different from the dominant adsorbate species; second, the presence of other adsorbates, even if they are “chemically spectators”, will modify the energetics of the system via many-body effects. The inclusion of this point within CSCP should not present excessive difficulties. One shall only need to generate worked-out or rescaled configurations in a proper way, but there is no intrinsic reason why the conformal principle should fail. Indeed, in our original work,8 we purposely used, as a worked-out case, configurations for aqueous phase reforming of methanol on Pt(100) and Pt(111) under conditions of high coverage of hydrogen, i.e., facets fully covered with hydrogen adatoms on all top sites except two for Pt(111) or on all bridge sites except two for Pt(100). Then, we conformally translated these into configurations for methanol decomposition at low coverage (no spectator hydrogens) on (111) and (100) facets of Au, Ag, Pd, Cu, Ni, Co, Fe, and Pt itself by simply removing the spectator hydrogens. In other words, we used a worked-out case of a different reaction and under completely different coverage conditions within the CSCP strategy and we obtained excellent results.
Clearly, care should be taken to account for high-coverage many-body effects, but in principle without any insurmountable obstacles. Indeed, within this same present study we found a case in point that rationalizes one example of discrepancy between DFT and MLIP-3 larger than 0.05 eV. For the C–O bond dissociation step (HO–CH2OH + H → HO + CH2OH + H) on Cu(100), the difference in energy barrier between DFT and MLIP-3 reads 0.16 eV. Analysis of parity plots shows that MLIP-3 does not reproduce the DFT energies accurately in the high-energy region, and we found that this is due to a many-body coverage effect. One hydrogen is in fact close to the HO–CH2OH adsorbate, and this interaction produces a difference in the energetics of the reaction as predicted by MLIP-3 (in whose parametrization high-coverage effects were not included) with respect to DFT (which obviously includes high-coverage effects), generating a discrepancy especially notable in the energy barrier. This is unwanted because in this phase of the CSCP development we would like to limit ourselves to the low-coverage case. To explore whether CSCP can reproduce this effect, we have parametrized a new MACE@MLIP-3′ using a database composed of the Database-D3 plus the newly generated reaction coordinates for the step HO–CH2OH + H→HO + CH2OH + H obtained via the NEB@MLIP-3. Now, although in doing so we have included in the database a minimal number of configurations, without performing the full extraction and extrapolation procedures foreseen by the CSCP protocol, the parity plot of the new NEB@MLIP -3′ shows a much more homogeneous agreement with respect to DFT, particularly in the higher-energy region, as shown in Fig. S3 of the SI. Moreover, the energy barrier is now predicted to be 1.29 eV, which compares very well with the DFT barrier of 1.33 eV for this step.
A more delicate situation occurs when high-coverage species significantly perturb the catalyst surface, as in the case of subsurface interstitials: this case when the high-coverage perturbation drastically modifies the catalyst is better seen under point (ii). Point (ii) is related to a basic assumption of the CSCP approach in its present version, that is, the use of single-crystal facets as structural models of the catalyst, which excludes less conventional materials or phenomena, such as nanostructured configurations or non-adiabatic processes. Both of these non-conventional factors are likely to play a role in CO2 hydrogenation on the industrial catalyst (CO2-Hydr). Indeed, it is known that the ZnO support drastically promotes the catalytic activity of Cu nanoparticles in CO2-Hydr, such that the catalytically active sites are thought to lie at the interface with the support, whence an ensemble of non-classical configurations have been proposed to account for and rationalize experimental observations,16,18–21,24 including partial ZnO reduction to Zn and successive alloying with Cu.30 It is possible to include these exotic configurations as worked-out systems in a “nanostructured” CSCP approach and compare with experiment (we are indeed working along these lines), but clearly the energetics of nanostructured systems will be different from that of the regular ones considered in the present work. Along another perspective, non-adiabatic thermal phenomena have also been argued to play a role in high-temperature reactions, e.g., in ethylene oxidation on Cu catalysts as monitored via operando transmission electron microscopy.31 These phenomena may play a role also in CO2-Hydr. Using an argument by analogy, switching to the electrochemical (non-thermal) CO2RR, several studies indicated that non-adiabatic bias-induced phenomena give rise to under-coordinated32–34 or oxygen-containing35–37 Cu surfaces, see a recent detailed investigation of CO2RR reaction intermediates.38 These undercoordinated or oxygen-containing sites are critical to favor CO2RR with respect to HER on regular Cu facets, which per se exhibit a very low faradaic efficiency. The robustness of the CSCP MLIPs in MD simulations as reported above gives us confidence that non-adiabatic phenomena can be included in our approach using appropriate configurations to generate the MLIP training database, but the assumption of regular and static facets held so far in CSCP precludes a direct comparison with experiment at this stage.
An indirect validation comes from comparison with previous computational simulations. Apart from technical details, our DFT@PBE-plus-dispersion approach is equivalent (and also quantitatively very close) to that employed in previous theoretical work32,34,38 which, after inclusion of implicit solvation and bias effects, compared accurately with experimental observations. The DFT@PBE-plus-dispersion approach (after including dispersion and implicit solvent) has thus been shown to give reasonably accurate results in comparison with experiment.39–41 Additionally, the same DFT approach has been employed in simulations including explicit water molecules, and used to predict reaction mechanisms42,43 and reaction intermediates,44–46 then validated by in situ experimental vibrational spectroscopy.38,47
Despite these present limitations of the approach, one important question is whether we can nevertheless derive some experimental implications or suggestions or testable predictions for the new (Ni) system. In Fig. 6 we compare the reaction energy diagrams for the most favorable path of CO2-Hydr for Cu(100) derived from our DFT search and for Ni(100) derived from the CSCP approach. Note that the diagrams of Fig. 6 are divided into two sections, where on the left before the dashed vertical line we consider CO2 adsorption on a metal surface with only two hydrogen adatoms and two H2 molecules in the gas phase (CO2 + H + H + 2H2 → HCOO + H + 2H2), whereas on the right after the dashed vertical line all the hydrogen molecules are dissociated and the surface contains a number of H adatoms ranging from zero (CH3OH + H2O) to 5 (HCOO + 5H). Clearly, the diagrams for Cu and Ni are quite different. First, it is apparent that the energy of the states on the Ni(100) catalyst increases much more quickly than on the Cu(100) system. This is due to the fact that Ni interacts much more strongly with hydrogen. This explains why we have a drop of 1.18 eV when we adsorb 4 hydrogens on the Ni surface, accounted for by a break between the HCOO states before and after the dashed vertical lines. This means that Ni(100) must escape from a thermodynamic dip to hydrogenate CO2, thus being a worse catalyst than Cu(100). Note also that in Fig. 6 we report energies, to be converted to free energies under the actual conditions, so naturally high H2 pressures and high temperatures would decrease substantially the thermodynamic penalty associated with first generating the H2COOH intermediate and then dissociating it to CH2O + OH on Cu(100). The stronger interaction of Ni with formaldehyde with respect to the methoxy and methanol intermediates instead contributes to making Ni(100) a worse catalyst than Cu(100). Finally, to the best of our knowledge, there are no experimental results that could be compared with a possible Ni–Cu alloyed catalyst for thermal CO2-Hydr (that we could derive as a straightforward follow-up of the present study10), although Ni–Cu systems have been demonstrated as efficient dual atom catalysts in electrochemical CO2RR.48
4. Conclusions
The computational discovery of new catalysts via high-throughput screening appears as a promising path for solving the daunting societal problems related to the sustainable chemical transformations in a global economy. In this respect, CO2 hydrogenation stands out as one of the major reactions for which sustainable routes must be found urgently. In the present work we extend our recently developed approach to high-throughput catalyst screening, the Conformal Sampling of Catalytic Processes (CSCP) approach, to CO2 thermal hydrogenation. We start from the Cu-fcc(100) surface as a worked-out catalytic facet, and we derive appropriate databases and associated Machine Learning Interatomic Potentials (MLIPs) for both the original Cu-fcc(100) worked-out case and for a new Ni-fcc(100) catalytic surface. The accuracy of the MLIPs thus derived is excellent: we predict both reaction energies and energy barriers for all the mechanistic steps of this complex reaction diagram with consistent and uniform accuracy, with a maximum discrepancy of 0.05 eV for Cu and 0.03 eV for Ni. Also, we show that in the exception cases in which this discrepancy is larger, this is due to a change in the reaction mechanisms, that bring the NEB@MLIP-3 simulations using a larger number of intermediate images (e.g., 48) to explore different pathways than the reference NEB@DFT simulations that use a more limited sampling and a smaller number of intermediate images (12). This change usually lowers the energy barrier, exploring a different saddle point at lower energy, but can also increase it, when the NEB simulation, initialized via a linear interpolation of initial and final states that are too distant in terms of atomistic configurations, converges to a different, higher-barrier pathway (composite reaction paths). We can anyway conclude that the CSCP approach is so validated on a significantly wider set of chemical transformations, thus offering itself as a candidate tool in high-throughput catalyst discovery, at the same time providing alternative atomistic mechanisms of catalytic processes and supporting catalyst rational design.Naturally, as discussed in Section 3.1, the present contribution represents a proof-of-principle study. The industrial working catalyst for CO2 thermal hydrogenation is made of Cu nanoparticles on a ZnO support, where the intimate interaction between the metal and the oxide components has been shown to play a critical role,18–24 a catalyst/support phenomenon that we do not consider here. We believe however that this type of phenomena are within the realm of an appropriately modified CSCP approach, which we defer to future studies. We leave for a future investigation also the introduction of solvation and bias effects, as needed to study CO2 electrocatalytic hydrogenation, for which the Cu fcc(100) surface is a well studied model of a catalytic facet. The structural database we have derived here should be a robust starting point for such an investigation, once the physics of reactant/electrode interactions49 is appropriately included within the CSCP framework.
5. Computational details
5.1 Database construction and DFT parameters
We employed density functional theory (DFT) calculations implemented in the Quantum Espresso (QE) package50,51 to investigate the Cu worked-out case and the Ni catalyst. The concept of database construction in the CSCP scheme for MLIP parametrization involves building two groups of datasets. The first group focuses on generating structures around local-minima regions. The second group aims to produce structures along reaction pathways through transition-state searches performed with Nudged Elastic Band (NEB) calculations.Spin-unrestricted calculations were employed in both DFT optimizations and NEB@DFT simulations. The face-centered cubic (fcc) bulk structures of Cu and Ni were optimized using vc-relax calculations in the QE suite to obtain the lattice parameters of Cu and Ni in their bulk forms. The calculations were performed using GBRV ultra-soft pseudopotentials52 together with the Perdew–Burke–Ernzerhof (PBE) exchange-correlation functional.53 A plane-wave basis set with a cut-off energy of 40 Ry was used for the wave functions, while the charge density was expanded with a cut-off of 200 Ry. A 3 × 3 × 1 k-point mesh was used for both structural optimizations and NEB calculations. An electronic energy convergence threshold of 10−6 Ry was applied.
The Cu(100) and Ni(100) surfaces were modeled using five-layer slabs composed of 45 metal atoms, with a 15 Å vacuum space applied along the z-direction to eliminate interactions between periodic images. The initial magnetization parameter (starting_magnetization) was optimized and set to 0.01 for Cu(100) and 0.3 for Ni(100). Cu and Ni atoms in the top three layers of the surface were allowed to move freely, while the bottom two layers were fixed during the optimization. Details of the input parameters for structural optimization and NEB calculations are provided in Section S1 of the SI.
We first carried out structural optimization of the intermediates (e.g., COOH, HCOO, CHOH, CH2OH, OH, CH2OH–OH, and CHOH–OH; see Scheme 1) on several adsorption sites of the Cu(100) surface including top, bridge, and 4-fold sites to identify the most stable adsorption configurations. These optimized structures were then used as the initial and final states for the subsequent NEB calculations. At this step, we exploit information obtained from several iterations of the DFT optimization by extracting XYZ coordinates together with their forces and energies through an algorithmic sampling technique. Based on the convergence series of the optimization, the structures change markedly during the very first iterations. Our CSCP approach targets this period when sampling for database construction, capturing coordinates around the local-minima basins rather than only the final optimized structures. Coordinates with their corresponding forces and energies in the neighborhood of the local minima are valuable for training the MLIPs, as they may guide the models to navigate these regions correctly. The algorithmic sampling scripts select 20 XYZ coordinates, primarily from the initial iterations, as well as the final structure obtained from the optimization. All extracted coordinates from the optimization tasks are gathered and combined with the NEB coordinates (details provided in the next following section) to form Database 1 of the Cu worked-out case.
We applied NEB and Climbing-Image Nudged Elastic Band (CI-NEB) calculations54,55 to identify the transition-state structure for each reaction step in Scheme 1 and to determine the corresponding energy barriers, enabling construction of the overall energy profile of CO2 hydrogenation on the Cu (100) surface. We first carried out the TS search with 12 intermediate images in the NEB calculation to estimate the activation energy (Ea) and obtain approximate reaction coordinates along the reaction pathway. It is worth noting that CO2 hydrogenation involves bulky steric fragments such as CHOH–OH, CH2OH–OH, and CH3OH + H2O on the metal surfaces, so in several cases the number of intermediate images was increased to 18 to achieve convergence in the NEB search. We then performed the CI-NEB calculation using the reaction coordinates from the initial NEB runs to accurately locate the transition structure and its energy barrier. The energy barrier (Ea, eV) was computed using:
| Ea = ETS − Ereactant |
5.2 MLIPs parameters
The CSCP employs an equivariant message-passing neural network (MPNN) as MACE model26 for parametrizing the database and providing potentials for NEB simulations. The MACE input parameters used for parametrizing MLIP-x for Cu and Ni are listed in Table S1. A total of 300 training steps were applied for the parametrization of the Cu- and Ni-MLIPs. Within the CSCP workflow, three successive rounds of MLIP parametrization were completed for both metal systems. The technical performance of MLIP-x (x = 1–3), evaluated by the RMSE of forces and energies from the MACE model, is summarized in Table S1.After obtaining MLIP-3 for both the Cu and Ni systems, we assessed their performance by performing NEB calculations using the MLIP potentials (NEB@MLIP). For technical implementation, the MACE model, as implemented in the LAMMPS software,56 was interfaced with the OPTIM code.57 We applied MLIP-3 as the force field in conjunction with the OPTIM code to perform NEB calculations. We note that the input parameters at the MLIP level, such as the energy threshold, supercell dimensions, 15 Å vacuum spacing along the z-direction, and fixation of the two bottom layers (see details of the metal slab models), were kept consistent with those used in the DFT optimizations and NEB calculations to ensure technical accuracy and reliability. The input parameters for NEB@MLIP simulations were listed in Section S2 in the SI.
In each reaction step, we carried out three NEB@MLIP simulations using different numbers of intermediate images (12, 24, and 48). The accuracy of the MLIP-predicted energy barriers was evaluated using the equation as ΔE = ECI-NEB@DFT − ENEB@MLIP where ECI-NEB@DFT is the barrier obtained from CI-NEB@DFT, ENEB@MLIP is the barrier predicted by NEB@MLIP-3. The resulting trends in barrier height and convergence behavior are summarized in Table 2 for Cu and Table 3 for the Ni catalyst.
The NEB@MLIP simulations were also employed to generate new datasets during the active-learning stages of the CSCP framework. They serve two purposes: (1) performing NEB calculations and extracting the final set of coordinates to be added to the database, and (2) providing input structures for extrapolation to generate additional datasets (see Methodology section for details). NEB@MLIP-x simulations (x = 1 for active-learning cycle 1 and x = 2 for cycle 2) were performed for all reaction steps using 48 intermediate images. Employing a large number of images helps capture coordinates that reflect not only bond breaking and forming but also rotations and reorientations of adsorbed species, which is valuable for extending the accuracy range of the MLIP model. Note that we do not perform fine-tuning of a previous MACE MLIP for a new system. For example, we did not fine-tune the MLIP of a Ni system using the previously developed MLIP for Cu. Instead, we generate an initial MLIP (MLIP-1) for Ni by training it from scratch on a database for Ni which is constructed by properly rescaling configurations from the Cu database. The validity range of MLIP-1 for Ni systems is then expanded through two active-learning iterations, resulting in MLIP-2 and MLIP-3. Additional database entries from the active-learning cycles, obtained by (1) extracting structures from geometry relaxation and NEB@MLIP simulations and (2) applying extrapolation techniques on NEB@MLIP coordinates, are incorporated into the dataset on which we train MLIP-2 and MLIP-3. To complete the database, on the so-generated database configurations, we perform the single-point calculations using the Quantum ESPRESSO suite to obtain their energies and forces at the DFT level.
Conflicts of interest
There are no conflicts to declare.Data availability
The data that support the major findings of this study are available in the main text, in the supplementary information (SI), or via Zenodo at https://zenodo.org/records/17762707. We report: (i) the output files of MLIP-3′ training of Cu and Ni (100), (ii) the output data from NEB@MLIP-3′ runs including reaction energies, energy barriers, and coordinates with different numbers of intermediate images, (iii) the NEB@DFT energies on the validation results, including 30 reaction steps of CO2 hydrogenation to produce H2O and CH3OH on the investigated Cu and Ni systems, available at https://zenodo.org/records/17762707. Further data are available from the corresponding authors upon reasonable request. The software associated with the CSCP modeling package will be made freely available in the near future.Supplementary information: Table S1 lists the input parameters used for MACE parametrization of MLIP. Table S2 lists the reaction energy simulated from NEB@DFT and NEB@MLIP. Fig. S1 shows the possible adsorption sites for intermediates on the metal (100) surface. Fig. S2–S8 present parity plots comparing single-point DFT energies with NEB@MLIP-3 energies for coordinates along the reaction steps: (e), (i), (m), (n), (x) for the Cu case and (g), (x) for the Ni case (see the full reaction paths in Tables 2 or 3). Section S1 describes the DFT calculation parameters, and Section S2 provides the NEB@MLIP calculation parameters. See DOI: https://doi.org/10.1039/d5fd00132c.
Acknowledgements
We thank the Italian CINECA supercomputing center for computational resources within the ISCRA program. We acknowledge financial support from ICSC, the Italian Centro Nazionale di Ricerca in High Performance Computing, Big Data and Quantum Computing, funded by the European Union (Next Generation EU, grant number CN00000013) – PNRR, Mission 4 Component 2 Investment 1.4, and from the Italian Ministry of Environment and Energy Security POR H2 AdP MMES/ENEA project, funded by the European Union (Next Generation EU) – PNRR, Mission 2, Component 2, Investment 3.5 ″Research and development on hydrogen”. Networking within the COST Action CA21101 “Confined molecular systems: from a new generation of materials to the stars” (COSY) supported by COST (European Cooperation in Science and Technology) and within the International Research Network IRN nanoalloys is also acknowledged.References
- E. Keinan and I. Schechter, Chemistry for the 21st Century, John Wiley & Sons, 2001 Search PubMed.
- B. Wander, M. Shuaibi, J. R. Kitchin, Z. W. Ulissi and C. L. Zitnick, ACS Catal., 2025, 15, 5283–5294 CrossRef CAS.
- A. Omranpour, J. Elsner, K. N. Lausch and J. Behler, ACS Catal., 2025, 15, 1616–1634 CrossRef CAS.
- Y. Hong, B. Hou, H. Jiang and J. Zhang, Wiley Interdiscip. Rev.:Comput. Mol. Sci., 2020, 10, e1450 CAS.
- Y. Xu, Y. Jin, J. S. García Sánchez, G. R. Pérez-Lemus, P. F. Zubieta Rico, M. Delferro and J. J. de Pablo, J. Phys. Chem. Lett., 2024, 15, 9852–9862 CrossRef CAS PubMed.
- S. S. Y. Kim, Presented in part at the Extended Abstracts of the CHI Conference on Human Factors in Computing Systems, Honolulu, HI, USA, 2024 Search PubMed.
- R. Jacobs, D. Morgan, S. Attarian, J. Meng, C. Shen, Z. Wu, C. Y. Xie, J. H. Yang, N. Artrith, B. Blaiszik, G. Ceder, K. Choudhary, G. Csanyi, E. D. Cubuk, B. Deng, R. Drautz, X. Fu, J. Godwin, V. Honavar, O. Isayev, A. Johansson, B. Kozinsky, S. Martiniani, S. P. Ong, I. Poltavsky, K. J. Schmidt, S. Takamoto, A. P. Thompson, J. Westermayr and B. M. Wood, Curr. Opin. Solid State Mater. Sci., 2025, 35, 101214 CrossRef CAS.
- T. Roongcharoen, G. Conter, L. Sementa, G. Melani and A. Fortunelli, J. Chem. Theory Comput., 2024, 20, 9580–9591 CrossRef CAS PubMed.
- S. Xu, J. Wu, Y. Guo, Q. Zhang, X. Zhong, J. Li and W. Ren, Chem. Phys. Rev., 2025, 6, 011309 CrossRef.
- G. Melani, T. Roongcharoen, G. Conter, L. Sementa and A. Fortunelli, J. Phys. Chem. C, 2025, 129, 17472–17483 CAS.
- T. Roongcharoen, G. Conter, G. Melani, L. Sementa and A. Fortunelli, J. Chem. Theory Comput., 2025, 21, 11164–11178 CrossRef CAS PubMed.
- M. Levy and J. P. Perdew, J. Chem. Phys., 1986, 84, 4519–4523 CrossRef CAS.
- S. Curtarolo, G. L. W. Hart, M. B. Nardelli, N. Mingo, S. Sanvito and O. Levy, Nat. Mater., 2013, 12, 191–201 CrossRef CAS PubMed.
- D. Kuryla, G. Csányi, A. C. T. van Duin and A. Michaelides, J. Chem. Phys., 2025, 162, 114122 CrossRef CAS PubMed.
- L. C. Grabow and M. Mavrikakis, ACS Catal., 2011, 1, 365–384 CrossRef CAS.
- F. Studt, M. Behrens, E. L. Kunkes, N. Thomas, S. Zander, A. Tarasov, J. Schumann, E. Frei, J. B. Varley, F. Abild-Pedersen, J. K. Nørskov and R. Schlögl, ChemCatChem, 2015, 7, 1105–1111 CrossRef CAS.
- M. Qiu, H. Tao, R. Li, Y. Li, X. Huang, W. Chen, W. Su and Y. Zhang, J. Chem. Phys., 2016, 145, 134701 CrossRef PubMed.
- M. Behrens, F. Studt, I. Kasatkin, S. Kühl, M. Hävecker, F. Abild-Pedersen, S. Zander, F. Girgsdies, P. Kurr, B.-L. Kniep, M. Tovar, R. W. Fischer, J. K. Nørskov and R. Schlögl, Science, 2012, 336, 893–897 CrossRef CAS PubMed.
- T. Lunkenbein, J. Schumann, M. Behrens, R. Schlögl and M. G. Willinger, Angew. Chem., 2015, 127, 4627–4631 CrossRef.
- S. Kuld, M. Thorhauge, H. Falsig, C. F. Elkjær, S. Helveg, I. Chorkendorff and J. Sehested, Science, 2016, 352, 969–974 CrossRef CAS PubMed.
- S. Kattel, P. J. Ramírez, J. G. Chen, J. A. Rodriguez and P. Liu, Science, 2017, 355, 1296–1299 CrossRef CAS PubMed.
- F. Scholten, K.-L. C. Nguyen, J. P. Bruce, M. Heyde and B. Roldan Cuenya, Angew. Chem., Int. Ed., 2021, 60, 19169–19175 CrossRef CAS PubMed.
- M. Rüscher, A. Herzog, J. Timoshenko, H. S. Jeon, W. Frandsen, S. Kühl and B. Roldan Cuenya, Catal. Sci. Technol., 2022, 12, 3028–3043 RSC.
- K. Mondal, Megha, A. Banerjee, A. Fortunelli, M. Walter and M. Moseler, J. Phys. Chem. C, 2022, 126, 764–771 CrossRef CAS.
- T. Roongcharoen, L. Sementa and A. Fortunelli, J. Phys. Chem. C, 2025, 129, 4032–4042 CrossRef CAS.
- I. Batatia, D. P. Kovacs, G. Simm, C. Ortner and G. Csányi, Adv. Neural Inf. Process. Syst., 2022, 35, 11423–11436 Search PubMed.
- F. R. Negreiros, E. Aprà, G. Barcaro, L. Sementa, S. Vajda and A. Fortunelli, Nanoscale, 2012, 4, 1208–1219 RSC.
- A. Burger, L. Thiede, N. Rønne, V. Bernales, N. Vijaykumar, T. Vegge, A. Bhowmik and A. Aspuru-Guzik, arXiv, 2025, preprint, arXiv:2509.21624, DOI:10.48550/arXiv.2509.21624.
- C. Zhan, F. Dattila, C. Rettenmaier, A. Bergmann, S. Kühl, R. García-Muelas, N. López and B. R. Cuenya, ACS Catal., 2021, 11, 7694–7701 CrossRef CAS PubMed.
- N. J. Divins, D. Kordus, J. Timoshenko, I. Sinev, I. Zegkinoglou, A. Bergmann, S. W. Chee, S. Widrinna, O. Karslıoğlu, H. Mistry, M. Lopez Luna, J. Q. Zhong, A. S. Hoffman, A. Boubnov, J. A. Boscoboinik, M. Heggen, R. E. Dunin-Borkowski, S. R. Bare and B. R. Cuenya, Nat. Commun., 2021, 12, 1435 CrossRef CAS PubMed.
- W. Yu, S. Yue, M. Yang, M. Hashimoto, P. Liu, L. Zhu, W. Xie, T. Jones, M. Willinger and X. Huang, Nat. Commun., 2025, 16, 2029 CrossRef CAS PubMed.
- T. Cheng, H. Xiao and W. A. Goddard, J. Am. Chem. Soc., 2017, 139, 11642–11645 CrossRef CAS PubMed.
- D. Gao, R. M. Arán-Ais, H. S. Jeon and B. Roldan Cuenya, Nat. Catal., 2019, 2, 198–210 CrossRef CAS.
- C. Choi, S. Kwon, T. Cheng, M. Xu, P. Tieu, C. Lee, J. Cai, H. M. Lee, X. Pan, X. Duan, W. A. Goddard and Y. Huang, Nat. Catal., 2020, 3, 804–812 CrossRef CAS.
- M. He, C. Li, H. Zhang, X. Chang, J. G. Chen, W. A. Goddard, M.-j. Cheng, B. Xu and Q. Lu, Nat. Commun., 2020, 11, 3844 CrossRef CAS PubMed.
- M. He, X. Chang, T.-H. Chao, C. Li, W. A. Goddard III, M.-J. Cheng, B. Xu and Q. Lu, ACS Catal., 2022, 12, 6036–6046 CrossRef CAS.
- M. Favaro, H. Xiao, T. Cheng, W. A. Goddard, J. Yano and E. J. Crumlin, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 6706–6711 CrossRef CAS PubMed.
- C. Zhan, F. Dattila, C. Rettenmaier, A. Herzog, M. Herran, T. Wagner, F. Scholten, A. Bergmann, N. López and B. Roldan Cuenya, Nat. Energy, 2024, 9, 1485–1496 CrossRef CAS PubMed.
- H. Xiao, T. Cheng, W. A. Goddard III and R. Sundararaman, J. Am. Chem. Soc., 2016, 138, 483–486 CrossRef CAS PubMed.
- H. Xiao, T. Cheng and W. A. Goddard III, J. Am. Chem. Soc., 2017, 139, 130–136 CrossRef CAS PubMed.
- H. Zhang, W. A. Goddard, Q. Lu and M.-J. Cheng, Phys. Chem. Chem. Phys., 2018, 20, 2549–2557 RSC.
- T. Cheng, H. Xiao and W. A. Goddard III, J. Am. Chem. Soc., 2016, 138, 13802–13805 CrossRef CAS PubMed.
- T. Cheng, H. Xiao and W. A. Goddard, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 1795–1800 CrossRef CAS PubMed.
- T. Cheng, A. Fortunelli and W. A. Goddard, Proc. Natl. Acad. Sci. U. S. A., 2019, 116, 7718–7722 CrossRef CAS PubMed.
- H. Yang, F. R. Negreiros, Q. Sun, M. Xie, L. Sementa, M. Stener, Y. Ye, A. Fortunelli, W. A. Goddard III and T. Cheng, ACS Appl. Mater. Interfaces, 2021, 13, 31554–31560 CrossRef CAS PubMed.
- Q. Sun, Y. Xiang, Y. Liu, L. Xu, T. Leng, Y. Ye, A. Fortunelli, W. A. Goddard III and T. Cheng, J. Phys. Chem. Lett., 2022, 13, 8047–8054 CrossRef CAS PubMed.
- F. Shao, J. K. Wong, Q. H. Low, M. Iannuzzi, J. Li and J. Lan, Proc. Natl. Acad. Sci. U. S. A., 2022, 119, e2118166119 CrossRef CAS PubMed.
- D. Yao, C. Tang, X. Zhi, B. Johannessen, A. Slattery, S. Chern and S.-Z. Qiao, Adv. Mater., 2023, 35, 2209386 CrossRef CAS PubMed.
- R. Jinnouchi and S. Minami, ACS Nano, 2025, 19, 22600–22644 CrossRef CAS PubMed.
- P. Giannozzi, S. Baroni, N. Bonini, M. Calandra, R. Car, C. Cavazzoni, D. Ceresoli, G. L. Chiarotti, M. Cococcioni, I. Dabo, A. Dal Corso, S. de Gironcoli, S. Fabris, G. Fratesi, R. Gebauer, U. Gerstmann, C. Gougoussis, A. Kokalj, M. Lazzeri, L. Martin-Samos, N. Marzari, F. Mauri, R. Mazzarello, S. Paolini, A. Pasquarello, L. Paulatto, C. Sbraccia, S. Scandolo, G. Sclauzero, A. P. Seitsonen, A. Smogunov, P. Umari and R. M. Wentzcovitch, J. Phys.: Condens. Matter, 2009, 21, 395502 CrossRef PubMed.
- P. Giannozzi, O. Andreussi, T. Brumme, O. Bunau, M. Buongiorno Nardelli, M. Calandra, R. Car, C. Cavazzoni, D. Ceresoli, M. Cococcioni, N. Colonna, I. Carnimeo, A. Dal Corso, S. de Gironcoli, P. Delugas, R. A. DiStasio, A. Ferretti, A. Floris, G. Fratesi, G. Fugallo, R. Gebauer, U. Gerstmann, F. Giustino, T. Gorni, J. Jia, M. Kawamura, H. Y. Ko, A. Kokalj, E. Küçükbenli, M. Lazzeri, M. Marsili, N. Marzari, F. Mauri, N. L. Nguyen, H. V. Nguyen, A. Otero-de-la-Roza, L. Paulatto, S. Poncé, D. Rocca, R. Sabatini, B. Santra, M. Schlipf, A. P. Seitsonen, A. Smogunov, I. Timrov, T. Thonhauser, P. Umari, N. Vast, X. Wu and S. Baroni, J. Phys.: Condens. Matter, 2017, 29, 465901 CrossRef CAS PubMed.
- K. F. Garrity, J. W. Bennett, K. M. Rabe and D. Vanderbilt, Comput. Mater. Sci., 2014, 81, 446–452 CrossRef CAS.
- J. P. Perdew, K. Burke and M. Ernzerhof, Phys. Rev. Lett., 1996, 77, 3865–3868 CrossRef CAS PubMed.
- R. Peverati and D. G. Truhlar, J. Phys. Chem. Lett., 2012, 3, 117–124 CrossRef CAS.
- G. Henkelman, B. P. Uberuaga and H. Jónsson, J. Chem. Phys., 2000, 113, 9901–9904 CrossRef CAS.
- A. P. Thompson, H. M. Aktulga, R. Berger, D. S. Bolintineanu, W. M. Brown, P. S. Crozier, P. J. in ’t Veld, A. Kohlmeyer, S. G. Moore, T. D. Nguyen, R. Shan, M. J. Stevens, J. Tranchida, C. Trott and S. J. Plimpton, Comput. Phys. Commun., 2022, 271, 108171 CrossRef CAS.
- D. Wales, Energy Landscapes with Applications to Clusters, Biomolecules and Glasse, Cambridge University Press, 2003 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |