Open Access Article
Open Access ArticleSystematic review on sustainable valorization of food waste by Yarrowia lipolytica
Yunkai Tan†
a,
Huaimin Wang† a,
Changyu Hub,
Jing Fubcde,
Rodrigo Ledesma-Amaro
a,
Changyu Hub,
Jing Fubcde,
Rodrigo Ledesma-Amaro bcde,
Fatwa F. Abdi
bcde,
Fatwa F. Abdi a,
Hsien-Yi Hsua,
Xue Wang
a,
Hsien-Yi Hsua,
Xue Wang a,
Hairong Chengf and
Carol Sze Ki Lin
a,
Hairong Chengf and
Carol Sze Ki Lin *a
*a
aSchool of Energy and Environment, City University of Hong Kong, Tat Chee Ave, Kowloon, Hong Kong. E-mail: carollin@cityu.edu.hk
bDepartment of Bioengineering, Imperial College London, London, SW7 2AZ, UK
cImperial College Centre for Synthetic Biology, Imperial College London, London, SW7 2AZ, UK
dThe Microbial Food Hub, Imperial College London, London, SW7 2AZ, UK
eBezos Centre for Sustainable Protein, Imperial College London, SW7 2AZ, UK
fState Key Laboratory of Microbial Metabolism, School of Life Sciences and Biotechnology, Shanghai Jiao Tong University, Shanghai, China
First published on 28th April 2026
Abstract
The escalating generation of global food waste presents both a profound environmental liability and an untapped resource for the circular bioeconomy. This systematic review evaluates the sustainable valorization of heterogeneous food waste streams using the versatile oleaginous yeast Yarrowia lipolytica. Following PRISMA guidelines, we analyze recent advancements (2006–2025) in feedstock pretreatment, highlighting enzymatic hydrolysis as the optimal strategy for liberating bioavailable monomers without generating the toxic byproducts associated with thermochemical methods. We detail the unique dual-trophic metabolic capacity of Y. lipolytica to simultaneously assimilate hydrophilic (glucose, glycerol) and hydrophobic (fatty acids) substrates, driving the synthesis of high-value bioproducts ranging from microbial lipids and bulk organic acids to rare sugars. Furthermore, the review synthesizes cutting-edge strain optimization techniques—including CRISPR/Cas9, Adaptive Laboratory Evolution (ALE), and emerging Artificial Intelligence (AI) frameworks—that accelerate bioprocess design and enhance inhibitor tolerance. Techno-Economic Analysis (TEA) and Life Cycle Assessment (LCA) confirm the platform's viability, demonstrating significant reductions in global warming potential and land use compared to conventional agricultural and petrochemical sources. Finally, we address current bottlenecks, such as substrate toxicity and scale-up constraints, and propose future directions, including synthetic microbial consortia and multi-product biorefineries, to solidify Y. lipolytica as a cornerstone of zero-waste biomanufacturing.
Sustainability spotlightFood waste remains a global challenge, contributing significantly to environmental degradation and resource loss. This review highlights the sustainable valorization of food waste through enzymatic hydrolysis integrated with Y. lipolytica fermentation, offering a robust pathway to high-value bioproducts. We evaluate cutting-edge strategies to enhance food waste valorization sustainability, including recombinant strain engineering, the design of synthetic microbial consortia, and the integration of AI technology for bioprocess optimization. These advancements directly support UN SDGS 9 (industry, innovation, and infrastructure), 12 (responsible consumption and production), 13 (climate action), and 15 (life on land) by transforming waste streams into a circular bioeconomy, reducing carbon footprints, and improving the economic viability of green manufacturing. |
1. Introduction
The escalating generation of global food waste represents one of the most pressing environmental and economic challenges of the 21st century.1 With approximately 1.3 billion tons of food lost or wasted annually, conventional disposal methods such as landfilling and incineration have proven highly unsustainable, contributing significantly to anthropogenic greenhouse gas emissions and the loss of valuable land resources.2–4 However, within the framework of a circular bioeconomy, this immense volume of organic waste is no longer viewed merely as an environmental liability, but rather as a highly abundant, carbon-rich feedstock.5 Comprising a complex matrix of carbohydrates, proteins, and lipids, municipal and agro-industrial food waste holds immense potential for bioconversion into high-value biochemicals, biofuels, and nutraceuticals.6Despite its potential, the biochemical heterogeneity of food waste presents formidable technical barriers for traditional microbial fermentation.7 The diverse and fluctuating concentrations of starches, triglycerides, and lignocellulosic fractions require sophisticated pretreatment strategies—such as hydrothermal, fungal, or enzymatic hydrolysis—to liberate bioavailable monomers.8 Furthermore, the resulting hydrolysates are often laden with microbial inhibitors, variable carbon-to-nitrogen (C/N) ratios, and extreme osmotic pressures.9,10 Conventional industrial microbes frequently lack the metabolic plasticity required to efficiently co-utilize these complex, mixed hydrophilic and hydrophobic streams, leading to suboptimal carbon recovery and economic inefficiencies.6,10
To overcome these metabolic bottlenecks, the non-conventional, oleaginous yeast Y. lipolytica has emerged as a premier microbial chassis for food waste valorization.11 Recognized for its ‘Generally Recognized as Safe’ (GRAS) status, Y. lipolytica possesses a unique dual-trophic metabolism capable of simultaneously assimilating hexoses, pentoses, glycerol, and free fatty acids.12–14 Its robust secretome, rich in extracellular lipases and proteases, allows it to thrive in non-sterile, lipid-rich environments like waste cooking oils and municipal organic fractions.15 By precisely modulating environmental stressors such as nitrogen or phosphate limitation, the carbon flux within Y. lipolytica can be directed toward the accumulation of single-cell oils (SCOs), the secretion of bulk organic acids (e.g., citric and succinic acid), or the synthesis of specialized polyols and rare sugars.13,16,17
The transition of this ‘Waste-to-Value’ pipeline from laboratory-scale proof-of-concept to industrial reality is being rapidly accelerated by modern biotechnology.18,19 Precision genome-editing tools like CRISPR/Cas9, combined with Adaptive Laboratory Evolution (ALE), are rewiring the yeast to tolerate severe inhibitory conditions and maximize product titers.19–21 Concurrently, the integration of Artificial Intelligence (AI) and machine learning into bioprocess design is shifting metabolic engineering from iterative trial-and-error to highly predictive, data-driven frameworks, optimizing everything from enzyme secretion to real-time bioreactor dynamics.22
Therefore, the objective of this systematic review is to comprehensively evaluate the sustainable valorization of food waste using Y. lipolytica. Guided by the PICOS framework, this review synthesizes recent literature (2006–2025) to critically assess optimal pretreatment strategies, elaborate on the yeast's complex substrate assimilation pathways, and highlight the latest advancements in genetic and AI-driven strain engineering. Finally, by integrating Techno-Economic Analysis (TEA) and Life Cycle Assessment (LCA) metrics, this paper aims to quantify the economic viability and environmental superiority of the Y. lipolytica platform, ultimately charting the course toward integrated, multi-product biorefineries in a zero-waste future.
2. Material and methods
2.1. PICOS-based eligibility criteria
The selection of literature for this systematic review was guided by the PICOS (Population, Intervention, Comparison, Outcome, and Study design) framework to ensure a comprehensive yet focused evaluation of food waste bioconversion.23 Specifically, the Population (P) encompassed diverse food waste streams, ranging from municipal organic fractions to lipid-rich agro-industrial residues. The Intervention (I) was strictly limited to bioprocessing and valorization schemes centered on Y. lipolytica. Where data permitted, Comparisons (C) were made against alternative microbial cell factories or conventional chemical routes. Primary Outcomes (O) included quantitative performance metrics (e.g., titer and productivity) as well as sustainability indicators, including Life Cycle Assessment (LCA) and Techno-Economic Analysis (TEA). Finally, the Study Design (S) restricted the scope to original peer-reviewed research and technical reports published within the 2006–2025 timeframe.2.2. Literature search and study selection
A systematic search was conducted following the PRISMA (Preferred Reporting Items for Systematic Reviews and Meta-Analyses) guidelines to maintain transparency and reproducibility.24,25The primary search was executed via the Scopus database using the boolean string: TITLE-ABS-KEY (‘food waste’) AND TITLE-ABS-KEY (‘Yarrowia’). An initial screening of 109 records was performed (Fig. 1). Titles and abstracts were first vetted against the inclusion criteria, followed by a rigorous full-text evaluation to curate the final database for synthesis.
 | ||
| Fig. 1 The PRISMA process used for screening documents related to food waste valorization by Yarrowia lipolytica. | ||
2.3. Data extraction and synthesis
The data from the chosen studies were organized into a systematic repository. Extracted parameters encompassed: (1) strain and feedstock: Y. lipolytica strain types, feedstock composition (particularly C/N ratio and free amino nitrogen content), and requisite pretreatment procedures; (2) bioreactor configurations (e.g., batch, fed-batch, or continuous) and physicochemical parameters (e.g., volume); (3) target products: production metrics for lipids, organic acids, polyols, and proteins; and (4) sustainability indicators: environmental footprints (greenhouse gas emissions, energy consumption) and economic feasibility criteria.3. Results and discussion
3.1. Sustainable feedstock utilization: diversity and pretreatment
The transformation of food waste into a value-added product typically involves hydrolysis, which functions as the initial stage that converts insoluble organic matter—including starch, proteins, and lipids—into bioavailable monomers like reducing sugars, amino acids, and fatty acids.26,27 This biochemical transformation is essential for disrupting the complex structure inherent in food waste, thereby enhancing nutrient solubility and facilitating more efficient downstream microbial conversion into value-added bioproducts.28 Due to the extreme heterogeneity and chemical diversity of food waste (Table 1), sustainable utilization depends on identifying these specific compositions to devise the downstream manufacturing approach.29 Strategic pretreatment techniques, including hydrothermal and enzymatic methods, are essential for addressing this diversity by surmounting inherent obstacles such as rate limitations.30,31| Feedstocks | Specification | References |
|---|---|---|
| Municipal organic waste | Kitchen waste, bakery waste, canteen waste, restaurant waste, grains/eggs/dairy/meat/vegetables, spoiled dairy, waste bread powder | 46–49 |
| Fish waste | Pangasius (heads/fins/innards/bones), mixed (anchovies/mantis/rose shrimps), waste fish oil | 50–52 |
| Agro-wastes/by-products | Banana/orange/lemon/potato/papaya peels, apple pectin, brewers' spent grain, sugar beet molasses | 47, 53 and 54 |
| Lignocellulosic biomass | Sugarcane bagasse, wheat straw/corn stover hydrolysate, corn plant biomass | 55–57 |
| Waste cooking oils/fatty acids | Waste frying oil, canola/palm/soybean/rapeseed oil, waste fish oil, triolein, oleic acid | 58–60 |
3.1.1.1. Municipal organic waste. A principal category of food waste for hydrolysis identified in these selected papers is mixed organic waste, commonly known as kitchen waste, canteen waste, or the organic fraction of municipal solid waste.32 These feedstocks are intrinsically heterogeneous, generally comprising a complex mixture of carbohydrates (starch, cellulose), proteins, and lipids.33,34 For example, the combinations consisting of different proportions of potatoes, fish, cheese, and fruits necessitate multi-enzyme systems for efficient solubilization into fermentable sugars and free amino acids.35
3.1.1.2. Fruit, vegetable, and agro-waste. A variety of plant-based residues, including fruit, vegetable, and agricultural waste, can function as sustainable feedstocks for hydrolysis.36 For instance, green kiwi peel, pumpkin peels, and melon waste are rich in pectin and lignocellulosic components, requiring hydrothermal or enzymatic hydrolysis to release glucose and other hexoses.37,38 Additionally, agro-waste, such as molasses, okara (soybean residue), and corn steep liquor, is utilized as a concentrated source of carbon and nitrogen.39 Lignocellulosic biomass, including non-food residues like sugarcane bagasse, wheat straw, and corn stover, can be hydrolyzed to generate sugar-rich media that replicate conventional fermentation substrates.40,41
3.1.1.3. Lipid-rich and hydrophobic feedstocks. Lipid-rich feedstocks, such as waste cooking oils, are often identified as principal or auxiliary carbon sources for hydrolysis, which transforms triglycerides into fatty acids and glycerol for biofuels and chemicals.42,43 During enzymatic hydrolysis, lipases facilitate the breakdown of lipids into long-chain fatty acids (LCFA) and glycerol.44 Moreover, hydrophobic feedstocks, including trap greases, rendering fats, and soapstocks, can be hydrolyzed using specialized hydrolase enzymes (e.g., EC 3.1.1.3), which adsorb onto hydrophobic water–oil interfaces and effectively catalyze the decomposition of hydrophobic substrates into fatty acids and glycerol.45
Consequently, food waste exhibits a significant biochemical heterogeneity, which will fundamentally influence the efficiency of downstream valorization processes, such as hydrolysis. Specifically, the chemical variability in food waste, characterized by differing concentrations of carbohydrates, proteins, and lipids, presents a considerable technical challenge in achieving consistent product yields during hydrolysis, as the existing enzymes may not align with the available substrate, resulting in a decline in hydrolysis rates until the microbial population adapts its secretion.61,62 To address these complexities, the application of customized pretreatment strategies (Table 2) acts as a crucial prerequisite for standardizing various feedstock streams. These interventions promote the breakdown of resistant organic structures, thereby improving nutrient bioavailability and optimizing the efficacy of subsequent microbial-based valorization systems.
| Strategy | General process | Glucose production | Glycerol production | Fatty acids production | References |
|---|---|---|---|---|---|
| Hydrothermal pretreatment | Food waste is subjected to high temperatures (160–280 °C) and pressures to induce organic dissolution and macromolecular hydrolysis | Rapid starch/cellulose hydrolysis to glucose occurs at 100–175 °C, though higher temperatures (>200 °C) cause degradation into furan compounds | Lipids are hydrolyzed into glycerol and fatty acids, primarily at temperatures exceeding 280 °C or under supercritical water conditions | Bio-crude yields from HTL can reach 27.5%, with total fatty acid methyl ester content ranging from 15–37 wt% | 81, 86 and 87 |
| Fungal pretreatment | Utilizes filamentous fungi (e.g., Aspergillus spp.) to secrete hydrolytic enzymes in situ, decomposing waste over several days | Efficient monomeric sugar recovery is possible through fungal enzymes, though the process typically requires longer reaction times compared to other methods | Fungal lipases (e.g., from A. oryzae) can release up to 17 g L−1 of glycerol from fat-rich dairy substrates | Fungi facilitate the release of free fatty acids from complex lipids, which can then be assimilated to accumulate 41–57% lipid in fungal biomass | 68 and 88–90 |
| Enzymatic pretreatment | Direct application of commercial or purified hydrolytic enzymes (amylases, lipases, proteases) under mild conditions (50–60 °C) | High recovery efficiency, yielding up to 164 g L−1 of reducing sugars from kitchen waste or up to 95% conversion efficiency in optimized platforms | Lipase-mediated hydrolysis effectively liberates glycerol; two-step platforms can yield a hydrolysate containing 3% glycerol prior to further fermentation | Targeted lipase pretreatment significantly enhances the dissolution of vegetable oils and animal fats, liberating long-chain fatty acids for subsequent bioconversion | 61, 84 and 91 |
3.1.2.1. Hydrothermal pretreatment. Hydrothermal pretreatment (HTP), which is alternatively categorized as hydrothermal carbonization or liquefaction depending on operational severity, represents a versatile thermochemical platform for the valorization of high-moisture food waste into energy-dense carriers and platform chemicals.63 By leveraging pressurized subcritical water as both a solvating medium and a reactive species, HTP facilitates the rapid depolymerization of complex organic matrices.7 This approach is particularly advantageous as it obviates the energy-intensive dehydration steps required by conventional thermal conversion technologies.64
A pivotal biochemical transformation during HTP involves the solvothermal hydrolysis of the lipid fraction, wherein triglycerides undergo nucleophilic attack to yield long-chain fatty acids and glycerol.65 Under subcritical and supercritical conditions, the significantly elevated ionic product of water enhances its autometabolic catalytic capacity, effectively shifting the hydrolysis equilibrium toward maximal glycerol liberation.66 The recovery of this glycerol is of particular interest in a biorefinery context, as it serves as a versatile biosynthetic intermediate for the production of hydrogen, volatile organic acids (e.g., lactic acid), or propanediols via downstream catalytic or hydrothermal pathways.67 Furthermore, integrating HTP with biological valorization—such as anaerobic digestion or microbial fermentation—enhances nutrient bioavailability and reduces substrate viscosity.68 This rheological improvement, coupled with the potential for synergistic co-digestion of food waste with crude glycerol, significantly optimizes subsequent bioenergy yields.
Despite these benefits, the application of HTP is frequently constrained by the formation of refractory byproducts and the inherent thermochemical instability of liberated metabolites.30 High-temperature regimes inevitably trigger Maillard reactions between hydrolyzed carbohydrates and proteinaceous nitrogen, resulting in the formation of complex melanoidins and nitrogenous heterocycles, such as pyrroles and pyridines.7 These recalcitrant compounds act as potent microbial inhibitors, potentially impairing the efficiency of subsequent enzymatic hydrolysis and microbial fermentation stages.
3.1.2.2. Fungal pretreatment. Fungal pretreatment represents a cornerstone of the ‘myco-biorefinery’ framework, leveraging the complex secretomes of filamentous fungi to homogenize and valorize heterogeneous food waste.69 This biological strategy primarily utilizes the secretion of extracellular lipases (triacylglycerol acyl hydrolases), which catalyze the interfacial hydrolysis of ester bonds in triglycerides.70 Unlike conventional chemical catalysis, fungal lipases operate under mild environmental conditions and exhibit high chemo- and regioselectivity.71 These attributes facilitate the efficient liberation of glycerol and long-chain fatty acids from complex lipid-rich streams, such as waste cooking oils and slaughterhouse effluents, providing a refined substrate for subsequent enzymatic or microbial processes.72
Key fungal genera, including Aspergillus, Rhizopus, and Penicillium, are utilized for their robust lipolytic activity and operational stability under varied substrate conditions.70 Through solid-state fermentation, these microorganisms can utilize agro-industrial residues as low-cost matrices to produce crude enzymatic extracts in situ, thereby improving the economic feasibility of the valorization chain.73 For instance, specific strains of Rhizopus oligosporus and Aspergillus niger have demonstrated the capacity to reduce the lipid content of organic waste by over 90%, generating a hydrolysate enriched with glycerol and free fatty acids that is highly bioavailable for downstream yeast fermentation.74
Despite its high specificity, fungal pretreatment is constrained by protracted reaction kinetics, which necessitate significantly longer residence times compared to thermochemical platforms.75 Operationally, the requirement for stringent substrate sterilization to mitigate microbial competition and the inherent susceptibility of fungal cultures to contamination impose substantial energetic and economic burdens.76 Furthermore, the accumulation of metabolic intermediates, such as liberated glycerol and organic acids, can trigger carbon catabolite repression or induce deleterious pH shifts, effectively compromising enzymatic efficiency and net yields.77 Collectively, these operational complexities and the high capital expenditure required for specialized bioreactor infrastructure represent critical techno-economic bottlenecks for large-scale industrial implementation.
3.1.2.3. Enzymatic pretreatment. Enzymatic pretreatment represents a high-precision biochemical platform for the selective recovery of fermentable monomers, specifically glucose, glycerol, and free fatty acids (FFAs), from heterogeneous food waste.48 This approach employs a targeted consortium of hydrolytic enzymes: amylases (notably α-amylase and glucoamylase) for the saccharification of starch fractions into glucose, and lipases for the interfacial hydrolysis of triglycerides into glycerol and long-chain fatty acids.78,79 By operating under mild thermodynamic conditions, enzymatic catalysis facilitates the comprehensive deconstruction of complex organic matrices into a bioavailable, nutrient-dense hydrolysate.48 Empirical studies have demonstrated that a synergistic mixture of carbohydrases, proteases, and lipases can effectively solubilize the recalcitrant organic components of municipal kitchen waste.80 Furthermore, high-efficiency glucose recovery has been achieved using optimized glucoamylase dosages, yielding reducing sugar concentrations as high as 186.2 g L−1.81 Such titers significantly enhance the volumetric productivity of downstream fermentation processes.
Compared to hydrothermal and fungal strategies, enzymatic pretreatment emerges as the superior method for integration with Y. lipolytica-based valorization for the following reasons: (1) unlike hydrothermal pretreatment, which inevitably triggers Maillard reactions and the formation of toxic dehydration products such as furfural and hydroxymethylfurfural, enzymatic pathways operate at temperatures that preclude the generation of these recalcitrant byproducts, ensuring a ‘clean’ substrate, preventing the metabolic repression of Y. lipolytica and maintaining high cell viability;82,83 (2) while fungal pretreatment is frequently hampered by protracted residence times (often requiring several days) and the risk of microbial contamination, enzymatic hydrolysis offers rapid saccharification kinetics, allowing for tighter integration into a continuous or fed-batch biorefinery workflow, thereby significantly increasing the overall space-time yield of the valorization plant;83,84 and (3) enzymatic pretreatment allows for the precise modulation of the carbon-to-nitrogen (C/N) ratio in the resulting hydrolysate through controlled proteolysis.83 Since Y. lipolytica requires specific nitrogen-limited conditions to trigger de novo lipogenesis, the ability to liberate specific amounts of free amino nitrogen alongside glucose and glycerol provides a level of metabolic tunability that is unattainable with the relatively non-selective hydrothermal or slow fungal methods.85 Consequently, the superior selectivity, operational stability, and compatibility of enzymatic pretreatment with the physiological requirements of Y. lipolytica establish it as the optimal homogenization strategy for translating diverse food waste streams into high-value bioproducts.
The strategic integration of enzymatic pretreatment with the non-conventional yeast Y. lipolytica establishes a highly efficient and sustainable framework for the valorization of heterogeneous food waste into high-value bioproducts. As a robust oleaginous host, Y. lipolytica exhibits significant metabolic plasticity, enabling the simultaneous or sequential assimilation of glucose, glycerol, and long-chain fatty acids—the primary monomers liberated during enzymatic saccharification and lipolysis.79,85 This metabolic versatility allows for the comprehensive utilization of carbon sources that would otherwise remain recalcitrant or inhibitory to conventional microbial hosts.85 Furthermore, the ability to modulate the nutrient profile of the resulting hydrolysate ensures that the carbon flux is precisely directed toward target biosynthetic pathways, such as de novo lipogenesis or organic acid secretion.78 Consequently, the following sections will provide a rigorous review of the specific carbon assimilation mechanisms and the downstream metabolic pathways utilized by Y. lipolytica to convert these waste-derived substrates into marketable biocommodities.
Unlike many conventional industrial yeasts, Y. lipolytica possesses the innate capability to thrive on the complex ‘nutrient broth’ resulting from food waste hydrolysis, which typically comprises a mixture of hexoses, pentoses, glycerol, and FFAs.83,94 This section delineates the metabolic pathways through which these hydrolysis products act as carbon and energy precursors, driving the synthesis of high-value-added metabolites such as microbial oils (single-cell oils, SCOs), organic acids, polyols, and specialized nutraceuticals.95,96
3.1.3.1. Metabolism of hydrophilic hydrolysis products. The enzymatic saccharification of the carbohydrate fraction in food waste predominantly yields glucose, alongside significant titers of galactose, mannose, and pentoses like xylose.83 Furthermore, the hydrolysis of the lipid fraction (discussed in Section 3.1.2.3) liberates substantial quantities of glycerol.
3.1.3.1.1. Hexose and pentose. Glucose remains the preferred carbon source, entering the cell via high-affinity hexose transporters and being processed through the Embden–Meyerhof–Parnas (EMP) glycolytic pathway.13 The resulting pyruvate flux is a critical junction; it can be directed toward the tricarboxylic acid (TCA) cycle for biomass accumulation or diverted toward de novo lipogenesis or citrate secretion under nutrient-limited conditions.13,17,95 Notably, while wild-type Y. lipolytica often exhibits suboptimal pentose utilization due to carbon catabolite repression (CCR), the glucose-rich environment of food waste hydrolysates often exceeding 50 g L−1 provides sufficient initial flux to establish high cell densities. In engineered strains, the conversion of xylose from the lignocellulosic components of food waste further enhances carbon efficiency, ensuring that the entirety of the ‘sugar footprint’ is transformed into value-added products.83,84
3.1.3.1.2. Glycerol. The liberation of glycerol during enzymatic or hydrothermal lipolysis provides Y. lipolytica with a highly reduced carbon source. Glycerol enters the central metabolism via the glycerol-3-phosphate (G3P) or dihydroxyacetone (DHA) pathways.21,79 This substrate is particularly advantageous for the production of organic acids, such as citric and isocitric acids, and polyols like erythritol and mannitol.17,93,95 Because glycerol catabolism bypasses several regulatory bottlenecks of glycolysis, it often results in higher yields of acetyl-CoA, the primary precursor for the synthesis of L-ergothioneine and other high-value terpenoids.13,96
3.1.3.2. Metabolism of hydrophobic hydrolysis products. Y. lipolytica is known for its unique ability to use hydrophobic substrates, which are found in large amounts in municipal and industrial food waste.94 Section 3.1.2 showed how pretreatment, like enzymatic lipolysis, mix these lipids together to make bioavailable FFAs and glycerol.79
The FFAs liberated from waste cooking oils or animal fats are internalized through specialized transport mechanisms and activated into fatty acyl-CoAs.12 Y. lipolytica can utilize these through two distinct routes. In the ‘growth-centric’ route, fatty acids undergo β-oxidation within the peroxisomes to generate energy and acetyl-CoA; conversely, in the ‘valorization-centric’ route, the yeast performs ex novo lipid accumulation, where extracellular fatty acids are directly incorporated into intracellular triacylglycerols.13,79 This process is significantly more energy-efficient than de novo synthesis from sugars, as it bypasses the complex cytosolic fatty acid synthase system. This capability allows the yeast to upcycle low-quality waste fats into high-value microbial oils with tailored fatty acid profiles suitable for biolubricants or nutraceutical-grade Omega-3 fatty acids.14,79
Beyond acting as a carbon source, the presence of hydrophobic hydrolysate in the fermentation medium serves as a potent inducer for the secretion of extracellular lipases.78 This creates a self-reinforcing bioconversion loop: the initial hydrolysis products stimulate the yeast to produce more enzymes, which further deconstructs any residual complex lipids in the waste stream.
3.1.3.3. Synergy of hydrolysate components in high-value synthesis. The true power of using food waste hydrolysates as a feedstock lies in the metabolic synergy between hydrophilic and hydrophobic components. For example, the simultaneous presence of glucose and FFAs allows the yeast to maintain high glycolytic flux (providing NADPH and ATP) while simultaneously accumulating lipids.13,79 Recent research demonstrates that engineered Y. lipolytica can convert the glucose and amino acids found in food waste hydrolysates into complex secondary metabolites like betanin (a natural red food colorant).97 In these systems, the FAN liberated during the proteolytic pretreatment of food waste acts as the essential nitrogen source for nitrogen-containing pigments and antioxidants like L-ergothioneine.22 Titers for these products are significantly enhanced when the feedstock is a complex hydrolysate rather than a pure sugar, likely due to the presence of trace minerals and micronutrients inherent in organic waste.98,99
In conclusion, the products of food waste solubilization discussed in Section 3.1.2 are not merely waste derivatives, they are high-density metabolic fuels that prime Y. lipolytica for the synthesis of a diverse portfolio of bioproducts (Table 3). By aligning the yeast's natural ‘hydrophobic–hydrophilic’ dual-nature metabolism with the chemical complexity of food waste, a scalable and sustainable biomanufacturing platform is established (Fig. 2). The subsequent sections will detail how these metabolic fluxes are specifically regulated to optimize the titers and yields of target metabolites.
| Feedstock | Metabolic pathway | Target product | Significance in food waste valorization | References |
|---|---|---|---|---|
| Glucose | Embden–Meyerhof–Parnas (EMP) & pentose phosphate pathway (ppp) | Primary energy source for biomass accumulation; precursor for citric acid and de novo single cell oil | High-titer availability (∼54.2 g L−1) supports rapid onset of high cell density | 17, 83, 84 and 95 |
| Xylose | Pentose metabolism (via xylitol/xylulose) | Auxiliary carbon sink; utilized for secondary metabolite production in late-stage fermentation | Ensures comprehensive carbon recovery from lignocellulosic fractions of vegetable waste | 83, 92 and 94 |
| Glycerol | G3P & Dihydroxyacetone (DHA) pathways | High-yield precursor for polyols (erythritol) and acetyl-CoA-derived compounds | Bypasses glycolytic bottlenecks, enhancing flux toward specialized antioxidants like L-ergothioneine | 13, 17, 21, 93, 95 and 96 |
| Free fatty acids | β-oxidation & ex novo lipid accumulation | Direct incorporation into intracellular triacylglycerols; induction of extracellular lipases | Enables upcycling of waste fats into tailored lipids with industrial-grade fatty acid profiles | 12–14, 78, 79, 94 and 106 |
| Free amino nitrogen | Proteolysis & amino acid catabolism | Regulates the C/N ratio; essential for nitrogenous metabolites (e.g., betalains) | Act as the ‘metabolic trigger’ for the shift from growth to high-value product accumulation | 22 and 95–97 |
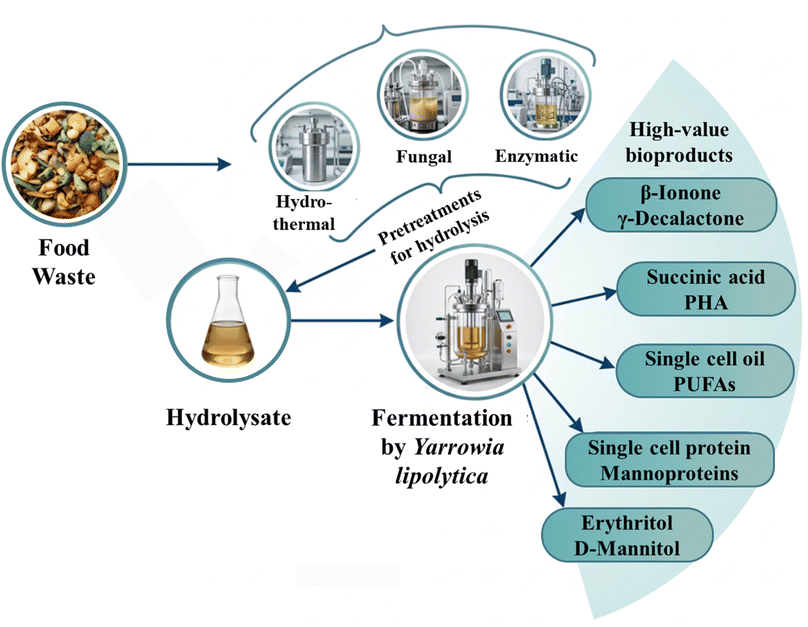 | ||
| Fig. 2 Integrated biorefinery platform for food waste valorization by hydrolysis and Y. lipolytica. Note: images in this figure are from literature.100–105 | ||
3.2. Metabolic pathways and genetic engineering strategies
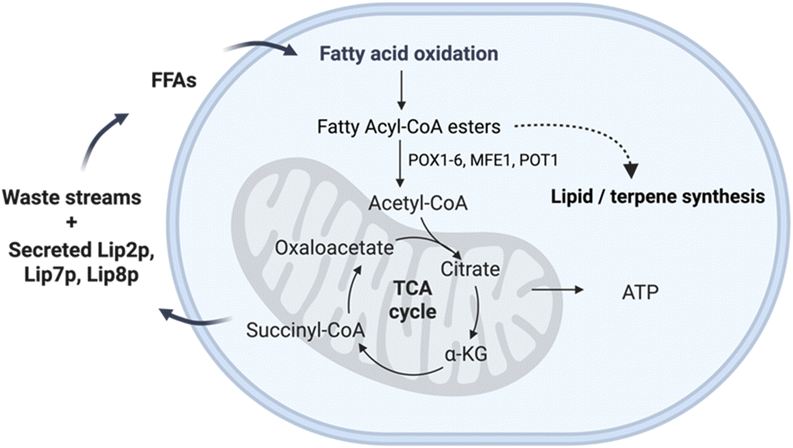
The β-oxidation pathway represents the cornerstone of Y. lipolytica's capacity to valorize lipid-rich waste streams, including waste cooking oil (WCO), crude glycerol from biodiesel production, and fatty acid distillates.107 This multi-compartment process initiates with extracellular lipase secretion—principally Lip2p, Lip7p, and Lip8p—which hydrolyze triglycerides into free fatty acids (FFAs) and glycerol.78 Notably, lipid presence induces substantial upregulation of lipase biosynthesis, with WCO supplementation demonstrating 5-fold higher lipase activity compared to oil-free cultures (Fig. 3).108
Following hydrolysis, FFAs traverse cellular membranes via transport proteins including Pxa1p, Pxa2p, and Ant1p, subsequently undergoing activation to fatty acyl-CoA esters. The peroxisomal β-oxidation spiral then systematically processes these activated fatty acids through the concerted action of six acyl-CoA oxidase paralogs (POX1–POX6), multifunctional enzyme MFE1, and 3-ketoacyl-CoA thiolase POT1.108 Each catalytic cycle excises a two-carbon acetyl-CoA unit, generating (R)-3-hydroxyacyl-CoA intermediates that may be diverted toward product synthesis or further catabolized for energy generation.107
The metabolic fate of β-oxidation-derived acetyl-CoA exhibits substantial flexibility. Under growth-associated conditions, carbon enters the tricarboxylic acid (TCA) cycle to support biomass accumulation. Conversely, pathway engineering through deletion of β-oxidation genes (PEX10, MFE1, POT1) dramatically expands fatty acyl-CoA pools available for redirecting toward lipid biosynthesis, terpenoid production, or other acetyl-CoA-derived products.109 This blockade strategy has proven particularly effective for α-bisabolene production, wherein combined β-oxidation interruption and lipid droplet engineering achieved titers of 1954.3 mg L−1—representing a 96-fold improvement over control strains.
Volatile fatty acids (VFAs) generated through food waste acidogenesis constitute another critical substrate category. Y. lipolytica demonstrates hierarchical consumption patterns, preferentially assimilating acetic acid via direct conversion to acetyl-CoA, followed by sequential utilization of propionic, butyric, and longer-chain VFAs. This metabolic organization enables efficient conversion of anaerobic digestate-derived VFAs into storage lipids, with optimized C![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) N ratios (>200
N ratios (>200![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) and acetic acid enrichment (6
1) and acetic acid enrichment (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 acetic
1 acetic![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) hexanoic) supporting lipid contents exceeding 43% w/w. The propionyl-CoA synthetase SePrpE has been identified as a critical engineering target for odd-chain fatty acid (OCFA) production, with overexpression in PHD1-deficient backgrounds achieving 60.4% OCFA enrichment (Fig. 4).109
hexanoic) supporting lipid contents exceeding 43% w/w. The propionyl-CoA synthetase SePrpE has been identified as a critical engineering target for odd-chain fatty acid (OCFA) production, with overexpression in PHD1-deficient backgrounds achieving 60.4% OCFA enrichment (Fig. 4).109
For carbohydrate-rich waste streams including food waste hydrolysates, lignocellulosic biomass, and expired glucose, Y. lipolytica deploys glycolysis and the pentose phosphate pathway (PPP) with distinctive regulatory characteristics. The yeast's Crabtree-negative nature constitutes a decisive advantage, ensuring minimal ethanol overflow metabolism and enabling efficient co-utilization of mixed sugars without carbon loss to byproduct formation. This trait proves particularly advantageous for food waste hydrolysates containing varied sugar compositions.110
Glucose metabolism proceeds through canonical glycolytic conversion to pyruvate, with subsequent routing toward acetyl-CoA for fatty acid synthesis or oxaloacetate for TCA cycle anaplerosis. Under nitrogen-limiting conditions characteristic of lipid accumulation phases, mitochondrial citrate accumulation triggers secretion to the cytoplasm, where ATP-citrate lyase cleavage generates acetyl-CoA for fatty acid synthase (FAS)-mediated de novo lipid synthesis.109 This nitrogen-responsive regulatory switch enables lipid contents of 30–48% dry cell weight from sugar-based feedstocks (Fig. 5).110,111
Glycerol assimilation exhibits notably faster kinetics than glucose utilization, attributable to higher oxygen uptake rates and the presence of three glycerol transport genes versus a single hexose transporter. Biodiesel-derived crude glycerol serves as an exceptional feedstock, with engineered strains achieving lipid contents up to 38% DCW and erythritol titers of 178 g L−1 following pathway optimization (Fig. 5).112 However, xylose metabolism remains limited by carbon catabolite repression and insufficient pentose pathway flux, necessitating engineering interventions for efficient lignocellulosic valorization.
The metabolic interplay between lipid and sugar substrates enables sophisticated process designs, including two-stage fermentation systems where initial glucose-supported growth transitions to VFAs or oil feeding for product accumulation. This approach circumvents the nitrogen limitation typically required for lipid accumulation while maximizing overall carbon conversion efficiency.
| Tool | Type | Details | References |
|---|---|---|---|
| Adaptive Laboratory Evolution (ALE) | Evolutionary engineering | Cultivating strains under selective pressure over long periods to naturally evolve tolerance and enhanced utilization of complex industrial wastes (e.g., cocoa fatty acid distillate) without targeted genetic modification | 113 |
| Targeted gene deletion/knockout | Genetic engineering | Inactivating specific native pathways (e.g., deleting POX genes to block β-oxidation) to prevent the degradation of accumulated lipids or to redirect carbon flux toward organic acids like succinic acid | 55, 109, 114 and 115 |
| Heterologous pathway assembly | Synthetic biology | Introducing and expressing non-native genes to enable the host to synthesize novel, high-value compounds such as β-ionone aroma, pyomelanin, or odd-chain fatty acids from organic residues | 109 |
| Microbial co-cultivation | Bioprocessing | Utilizing mixed cultures of different Y. lipolytica strains to synergistically enhance the bioconversion of mixed agro-industrial by-products into complex metabolites like 2-phenylethanol | 116 |
| Computational metabolic modeling | In silico/computational | Using computational algorithms to predict rational metabolic pathway designs, balance redox cofactors, and optimize precursor supply before physical strain construction begins | 117 |
The establishment of CRISPR/Cas9 systems for Y. lipolytica—exemplified by the EasyCloneYALI and pCAS1yl platforms—has revolutionized strain construction through unprecedented editing efficiency and versatility.109 These systems leverage direct tRNA-sgRNA fusions to enhance guide RNA processing, enabling precise gene disruptions, integrations, and multiplexed modifications.
Marker-free gene deletions constitute a primary application, utilizing gRNA-assisted homologous recombination to eliminate competing pathways without residual selection markers. Critical deletions include: (a) Yl4HPPD: eliminates pyomelanin formation, enhancing precursor availability for aromatic compound synthesis;118 (b) β-glucosidases (YALI1_B18845g, YALI1_B18887g): prevents betanin degradation, supporting high-titer food colorant production (1271 mg L−1);55 (c) SDH5: blocks succinate oxidation, diverting TCA flux toward succinic acid accumulation (13-fold titer improvement);55,114 (d) PEX10/MFE1/POT1: disrupts peroxisomal β-oxidation, forcing fatty acyl-CoA accumulation for lipid or terpenoid production;119 (e) PHD1: Inactivates 2-methylcitrate dehydratase, sensitizing cells to propionic acid while redirecting carbon toward odd-chain fatty acid synthesis.120
Pathway integration and optimization exploits specific docking platforms (pBR322, AXP, POX4 locus) for stable genomic insertion of heterologous biosynthetic operons. The combinatorial assembly of betanin biosynthesis enzymes (MjDOD, EvTYH, BvSGT2) with multiple copy integration exemplifies this approach, achieving record product titers.97 Similarly, carotenoid cleavage dioxygenase integration at the POX4 locus enabled efficient β-ionone production from organic waste hydrolysates.82
Feedback-resistant allele introduction addresses metabolic bottlenecks through incorporation of deregulated enzyme variants. The shikimate pathway enzymes YlARO4K221L and YlARO7G141S, which are insensitive to aromatic amino acid feedback inhibition, substantially enhance flux toward tyrosine-derived products when combined with appropriate pathway deletions.54
Fine-tuned gene expression regulation through promoter engineering enables metabolic pathway optimization without fundamental genomic reconfiguration. Y. lipolytica benefits from a diverse promoter repertoire spanning constitutive, inducible, and hybrid architectures.56
Strong constitutive promoters drive high-level heterologous expression: (a) hp4d: hybrid promoter carrying four UAS1B elements derived from XPR2; exhibits robust activity without carbon source repression, widely deployed for terpenoid and lipid pathway engineering. (b) PTEF/PTEF1: translation elongation factor promoter providing reliable constitutive expression. (c) PGPD2: glycerol-3-phosphate dehydrogenase promoter effective for phospholipid pathway engineering.56
Inducible and condition-responsive promoters enable dynamic metabolic control: (a) POT1 promoter: oleic acid-inducible system facilitating oxygen-responsive SDH2 regulation for succinic acid production enhancement (4.5-fold improvement).55 (b) synthetic 4UASpTEF: customized hybrid architecture for optimized expression strength.
Translational optimization complements transcriptional control through codon adaptation to Y. lipolytica usage preferences and Kozak sequence incorporation. These modifications have achieved 3.6-fold expression enhancement for PHA synthase and substantial improvements for bacterial genes including tNDPS1 and ldhA.121
Multi-copy integration strategies exploit defective marker systems (ura3d4) and resistance cassettes (LEU, hygromycin) for gene dosage amplification. The pINA1292 vector enables copy number variation through defective URA3 selection, with seven-fold copy number increases correlating with proportional titer improvements for polyhydroxyalkanoate production.56
The most impactful engineering outcomes emerge from coordinated application of multiple tools. The ‘push–pull’ strategy for α-bisabolene production exemplifies this integration: ‘push’ mechanisms overexpress rate-limiting mevalonate pathway genes (HMG1, ABC-G1 transporter) to enhance precursor supply, while ‘pull’ mechanisms expand lipid droplet storage capacity through DGA1 and OLE1 overexpression, creating intracellular reservoirs for lipophilic product accumulation. This synthetic biology approach achieved the highest reported α-bisabolene production in Y. lipolytica.1
Low-pH tolerance evolution addresses the economic burden of neutralizing agent addition during organic acid production. The evolutionary trajectory from PGC01003 through PSA02004 to PSA3.0 demonstrates progressive adaptation through repeated batch fermentation in an in situ fibrous bed bioreactor (isFBB) under gradually decreasing pH. Selection at pH 3.0 yielded a strain exhibiting 4.8-fold increased succinic acid titer under acidic conditions, abolished acetate production through pyruvate decarboxylase and aldehyde dehydrogenase downregulation, 43% reduction in NaOH consumption, stable phenotype maintenance without genetic drift.55
Glucose metabolism restoration in SDH-deficient backgrounds illustrates ALE's capacity to resolve engineered metabolic deficiencies. Immobilized-cell adaptive evolution using absorbent cotton matrices enabled 14 transfers over 21 days, increasing glucose consumption rates from 0.07 to 0.30 g L−1 h−1. The evolved strain PSA02004 achieved 87.9 g L−1 succinic acid from food waste hydrolysate with 85% of theoretical yield—2.3-fold higher productivity than the parental strain.122
Lipid production enhancement from inhibitory feedstocks has been demonstrated through extended cultivation protocols. Strain EVO1, derived from 79 passages on cocoa fatty acid distillate containing growth-inhibitory caffeine (5% w/w), exhibited >2-fold higher biomass and 3–4-fold increased lipid titers (2.3 vs. 0.5 g L−1). Transcriptomic characterization revealed metabolic rewiring: β-oxidation genes (POX2, POX3, POT1) and intracellular lipases were downregulated at early growth phases, while extracellular lipases (LIP2, LIP8) and fatty acid activation genes (AAL9, FAA1) were upregulated—suggesting enhanced exogenous lipid uptake and reduced catabolism.113
Pre-adapted environmental isolates frequently outperform laboratory-evolved strains, highlighting the efficacy of natural selection in harsh waste environments. The V1 strain of Pichia kudriavzevii—isolated from matured vinasse—exemplifies this principle with growth capacity at pH 2.0 and 45 °C, achieving saturated OD values 2.6-fold higher than engineered Y. lipolytica mutants on identical media. Whole-genome sequencing revealed SNP enrichment in lipid metabolism genes (TAG lipases, fatty acid elongases), indicating evolutionary adaptation to stringent conditions.113
Y. lipolytica strain A-10, recovered from oil-contaminated soil, demonstrates robust growth on raw rapeseed oil and crude glycerol without prior laboratory evolution.123 Similarly, marine-derived halotolerant strains tolerate up to 11% NaCl while maintaining high lipase activity. These observations suggest that environmental reservoirs harbor substantial untapped genetic diversity for bioprocess applications.123
While formal ALE protocols remain underrepresented in the literature, studies document adaptive mechanisms that mirror evolutionary outcomes:
Membrane remodeling constitutes a primary tolerance strategy. Fed-batch cultures with molasses hydrolysate induce ergosterol production up to 60.16 mg g−1 oil, enhancing membrane fluidity and osmotic stress resistance.124 Ole1p-mediated unsaturated fatty acid enrichment similarly improves weak acid and alcohol tolerance.
C![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) N ratio manipulation alleviates short-chain fatty acid inhibition through metabolic mode switching. Low C
N ratio manipulation alleviates short-chain fatty acid inhibition through metabolic mode switching. Low C![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) N ratios (∼80
N ratios (∼80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) prioritize nitrogen assimilation toward growth, while high C
1) prioritize nitrogen assimilation toward growth, while high C![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) N conditions (>200
N conditions (>200![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) divert carbon to lipid storage—demonstrating metabolic flexibility that could be selected and enhanced through directed evolution.125
1) divert carbon to lipid storage—demonstrating metabolic flexibility that could be selected and enhanced through directed evolution.125
Nutrient limitation adaptation reveals unexpected evolutionary trajectories. Phosphate limitation (0 g L−1 KH2PO4) proved superior to nitrogen limitation for SCFA-derived lipid production, achieving 0.30 g g−1 yields—2.3-fold higher than phosphate-replete conditions. This suggests phosphate starvation activates evolutionary conserved stress responses that rewire metabolism toward carbon storage.126
Process-parameter optimization can impose selective pressure analogous to ALE. Controlled oxygen transfer coefficient (kLa 16 h−1) maximized lipase activity (12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 U per L) and lipid accumulation (48% w/w) from WCO, while excessive aeration (93 h−1) induced oxidative stress and protease release. This oxygen sensitivity indicates potential for evolution under redox stress.127
000 U per L) and lipid accumulation (48% w/w) from WCO, while excessive aeration (93 h−1) induced oxidative stress and protease release. This oxygen sensitivity indicates potential for evolution under redox stress.127
Explicit ALE protocols for Y. lipolytica remain limited compared to bacterial and S. cerevisiae systems. Priority targets include lignocellulosic inhibitor tolerance: systematic evolution against furfural, 5-hydroxymethylfurfural (5-HMF), and phenolic compounds prevalent in pretreated biomass hydrolysates; high substrate concentration robustness: progressive adaptation to undiluted food waste hydrolysates exceeding 200 g L−1 glucose equivalents; co-culture compatibility: evolution for stable performance in synthetic consortia with bacteria for integrated food waste bioconversion.128
The combination of natural isolation, rational engineering, and directed evolution offers a powerful tripartite strategy for developing industrial biocatalysts capable of thriving in the complex, variable environments characteristic of food waste valorization processes.
In recent years, artificial intelligence (AI) and machine learning (ML) have emerged as powerful tools to accelerate strain engineering, pathway design, and bioprocess optimization.130
By integrating large-scale biological datasets with predictive algorithms, AI-driven strategies enable more efficient identification of metabolic targets, improved enzyme engineering, and real-time optimisation of fermentation processes. These approaches offer promising opportunities to enhance the efficiency and scalability of Y. lipolytica-based food waste valorisation systems (Fig. 6).
3.2.4.1. AI-assisted enzyme and strain engineering. To effectively break down complex polymers found in food waste, such as proteins and fat, Y. lipolytica often requires expression of heterologous enzymes.131 For traditional protein engineering, methods like directed evolution are widely used for selection of enzymes with optimal affinity and stability.132 However, these methods often require extensive screening and multiple rounds of mutagenesis, which highlights the potential value of AI-assisted approaches.
Recent advances in machine learning (ML) have enabled the prediction of enzyme structure–function relationships, thereby accelerating protein engineering efforts. Within the deep learning (DL) framework, convolutional neural networks (CNNs) are widely used architectures that process spatial and structural data.133 CNNs are able to recognize local patterns and hierarchical features, which allows CNNs to predict the impact of mutations.134 These predictive tools allow researchers to prioritise promising mutations before experimental validation, significantly reducing the experimental search space.
In Y. lipolytica, ML approaches have been applied to improve heterologous protein secretion. For example, machine learning has been applied to design N-terminal signal peptides in Y. lipolytica, which play an important role in directing heterologous proteins to different secretory pathways.135 Through integration of directed evolution and ML, researchers achieved approximately a 2.9-fold increase in secretion titers for enzymes such as PET hydroxylase and alpha-amylase.135 These results highlight the potential of AI-assisted protein engineering to accelerate the development of efficient enzymatic systems for bioconversion of food waste.
3.2.4.2. AI-guided metabolic modelling and pathway optimisation. In Y. lipolytica, AI-guided metabolic modelling has the potential to transform bioprocess design from traditional ‘trial-and-error’ engineering to more predictive AI frameworks. These methodologies systematically address the yeast's complex metabolic architecture to maximise the production of high-value compounds from food waste.
Genome-scale metabolic models (GEMs), such as iNL895 and iYali21, provide the mathematical scaffolding for simulating Y. lipolytica metabolism.136 AI can enhance these models by integrating with historical data and ML algorithms. In one study, a model trained on data from more than 100 publications predicted the production titers of various compounds in Y. lipolytica. The model achieved an R2 of 0.87 for predicting titer more than 1 g L−1.137
The integration of AI with multi-omics data also has the potential to revolutionize the metabolic engineering in Y. lipolytica by enabling the training of predictive models using high-dimensional biological datasets. Since the data of multi-omics data are high-dimensional and often non-linear, traditional statistical methods may struggle to extract meaningful relationships.138 AI approaches, particularly DL models, are capable of identifying hidden patterns across multiple biological layers.139
3.2.4.3. AI-enabled fermentation monitoring and process optimisation. The transition from lab-scale production to industrial production can often lead to problems because lab-scale shake flasks often fail to translate to large-scale bioreactors. Consequently, real-time monitoring and control of the fermentation process are critical in the translation process. However, some important indicators such as dry cell weight (DCW), substrate concentration and lipid accumulation remain analytically challenging during fermentation.
To address these limitations, soft sensors driven by Artificial Neural Networks (ANN) have been implemented to estimate these variables. These models utilise real-time data, including fermentation duration, dissolved oxygen (DO) concentration, glucose levels, and pH, to predict complex parameters that are difficult to monitor.140 Furthermore, Reinforcement Learning (RL) agents can be employed for the dynamic regulation of critical process parameters. RL-based control strategies can potentially adjust parameters such as nutrient feeding strategies and environmental conditions during fermentation based on historical process data.141 Such data-driven control strategies may help improve process robustness and reduce batch variability during large-scale fermentation by roughly 60%.142
3.2.4.4. Challenges and technical barriers. Although AI-driven approaches offer distinct competitive advantages over conventional methodologies (Fig. 7), their practical implementation remains constrained by several technical challenges. High-precision model training relies on the availability of large volumes of standardised, high-quality data sets which remains notably sparse within the biomanufacturing sector.143 Additionally, complex architectures like deep neural networks often suffer from a lack of interpretability, effectively operating as ‘black boxes’ models.144 Future research should therefore focus on integrating interpretable AI approaches with high-quality biological datasets to improve the reliability and biological insight of AI-driven metabolic engineering strategies. Consequently, despite the significant promise of AI in metabolic engineering, its practical application has not yet been extensively utilised in Y. lipolytica.
3.3. Value-added products and sustainability contributions
| Product category | Market price (est. USD per kg) | Key scholars | References |
|---|---|---|---|
| a The metrics of selecting key scholars are H-Index >40, titer record >100 g L−1, landmark paper >200 citations, and waste versatility >30% of portfolio. | |||
| Microbial lipids | $1.60–$5.00 | Ratledge, C (Hull); Park, Y & Nicaud, J (INRAE); Papanikolaou, S (AUA) | 166–169 |
| Citric & succinic acids | $0.80–$2.50 | Rymowicz, W (Wroclaw); Nielsen, J (Chalmers); Ji, X (Nanjing Tech) | 170–172 |
| Erythritol & polyols | $3.00–$7.00 | Cheng, H (SJTU); Fickers, P (Liège) | 173–175 |
| Rare sugars (e.g. threitol) | $50.00–$200.00 | Cheng, H (SJTU); Bilal, M; Izumori (Kagawa University) | 176–178 |
| Single-cell protein | $1.00–$2.50 | Amaral, P (UFRJ) | 179 |
| Flavors (β-ionone) | $300.00–$600.00 | Stephanopoulos, G (MIT); Zhou, Y (DICP); Scheper, T (LUH) | 180 and 181 |
Y. lipolytica is uniquely suited for the primary valorization of these streams due to its exceptional metabolic plasticity and ‘Generally Recognized as Safe’ (GRAS) status.146 Its robust secretome, characterized by high-activity extracellular lipases (up to 257.3 U per g DCW) and proteases, allows it to thrive in the complex, non-sterile nutrient environments of food waste hydrolysates.83,108,147 The yeast's ability to simultaneously assimilate hydrophilic substrates (glucose, glycerol, and organic acids) and hydrophobic compounds (long-chain fatty acids and triacylglycerols) is facilitated by a specialized expansion of the CYP52 cytochrome P450 family.94,148 This dual-trophic capability is particularly relevant for valorizing oil-rich kitchen leftovers, where starch-derived glucose and lipid-derived glycerol can be co-fermented to maximize total carbon recovery.78,94
Strategic process control—specifically nitrogen limitation and pH modulation—allows for the precise partitioning of carbon flux toward desired bulk metabolic outputs.149,150 Under nitrogen-limiting conditions at neutral pH, the citrate shuttle is activated, enabling Y. lipolytica to accumulate single-cell oils exceeding 50% of its dry cell weight or to secrete citric acid at concentrations surpassing 100 g L−1.151–153 Furthermore, engineered strains have demonstrated the capacity to produce dicarboxylic acids, such as succinic acid, at titers of 140.6 g L−1, providing renewable precursors for the polymer sector.154 Beyond these bulk acids, the platform effectively synthesizes high-value terpenoids and pigments, such as β-ionone, from crude waste streams.82 Finally, the residual yeast biomass, characterized by a balanced amino acid profile and high protein content (up to 57.9%), serves as a nutrient-dense single-cell protein for the aquaculture and livestock sectors, ensuring a zero-waste bioprocessing loop.49,155
The primary stage of this specialized process involves the assimilation of food waste hydrolysates under conditions where Y. lipolytica exhibits high tolerance to inhibitory compounds like furfural.158 By directing carbon flux from waste-derived glucose toward the erythrose-4-phosphate intermediate, high titers of erythritol are achieved.159,160 Furthermore, the synthesis of specialized polyols like D-threitol has been achieved by leveraging the yeast's native metabolism, effectively turning a low-value waste stream into a functional food ingredient valued for its low-caloric properties and high stability.161
The valorization potential is further amplified when Y. lipolytica is employed as a platform for ‘surface-display’ or ‘membrane-associated’ biocatalysis. Building on the characterization of membrane-bound enzymes—such as the sorbitol dehydrogenases identified in recent SJTU studies—engineered Yarrowia strains can perform regiospecific oxidations of the very polyols they produce.162,163 For example, once the yeast has converted food waste into sorbitol or mannitol, membrane-associated enzymes facilitate the subsequent transformation into rare sugars like L-xylo-3-hexulose.163
To ensure the economic sustainability of this platform, advanced enzyme stabilization techniques are critical. The application of multi-point covalent attachment and the formation of cross-linked enzyme aggregates (CLEAs) allow the specific dehydrogenases required for rare sugar synthesis to maintain activity across multiple batches of food waste hydrolysate.164,165 This stability is essential when dealing with the variable pH and mineral content inherent in municipal organic waste. Ultimately, moving beyond simple disposal to the precision-engineering of high-margin nutraceuticals represents the ‘highest tier’ of the food waste biorefinery.
| Assessment category | Key parameters | Benchmarks | Sustainability contribution | References |
|---|---|---|---|---|
| Techno-Economic (TEA) | Feedstock cost | −$40 to −$100 per ton (tipping fees) | Converts waste liability into a revenue-generating asset; offsets OPEX | 192 and 193 |
| Lipid MSP | $1.60–$2.10 per kg SCO | Achieves price parity with conventional vegetable oils via waste-to-oil conversion | 180 and 194 | |
| Internal Rate of Return (IRR) | 15–25% Increase | Enhanced by multi-product streams (bulk + specialized metabolites) | 178 | |
| Life Cycle (LCA) | Global Warming Potential (GWP) | −0.5 to −1.2 kg CO2-eq per kg waste | Net negative carbon footprint by diverting waste from methane-emitting landfills | 195 |
| Land use efficiency | ∼100× reduction vs. palm oil | Preserves biodiversity (SDG 15) by decoupling oil production from arable land | 173 | |
| Energy demand | 50–60 °C (enzymatic hydrolysis) | Lower ‘cradle-to-gate’ energy compared to thermo-chemical liquefaction | 177 | |
| Social & circularity | Waste diversion | ∼1.3 billion tons (global scale) | Directly addresses SDG 12.3 (food waste reduction) | 196 |
| Resource recovery | 25.1–57.9% protein content | Provides sustainable Single-Cell Protein (SCP) for the aquaculture circular loop | 179, 197 | |
| Process refinement | Green chemistry | 40% toxicity reduction | Transitioning to supercritical CO2 or bio-solvents for downstream processing | 177 |
The economic sustainability of Y. lipolytica bioprocessing is heavily influenced by the raw material cost and the final product's market value.185,186 Conventional fermentation often relies on refined glucose, which can account for up to 60–70% of total operating expenditures (OPEX).186 By utilizing food waste hydrolysates, the feedstock cost is effectively neutralized or even turned into a revenue stream via tipping fees, which in municipal settings can range from $35 to $100 per ton.185
TEA models for single-cell oil production by Y. lipolytica suggest that at an industrial scale (e.g., 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 tons per year), the minimum selling price of microbial lipids can be reduced to approximately $2.10 per kg when integrated with waste-derived nutrients.186 However, the true economic strength of the platform lies in the ‘multi-product biorefinery’ approach discussed in Sections 3.3.1 and 3.3.2. By co-producing bulk citric acid with high-margin specialized metabolites like D-threitol or L-ergothioneine, the internal rate of return of a facility can increase by 15–25%.97,185,186 The recovery of single-cell protein from the residual yeast biomass further offsets costs, providing a balanced amino acid profile for the animal feed market at a competitive price point of $1200 per ton, thereby closing the economic loop.84,186
000 tons per year), the minimum selling price of microbial lipids can be reduced to approximately $2.10 per kg when integrated with waste-derived nutrients.186 However, the true economic strength of the platform lies in the ‘multi-product biorefinery’ approach discussed in Sections 3.3.1 and 3.3.2. By co-producing bulk citric acid with high-margin specialized metabolites like D-threitol or L-ergothioneine, the internal rate of return of a facility can increase by 15–25%.97,185,186 The recovery of single-cell protein from the residual yeast biomass further offsets costs, providing a balanced amino acid profile for the animal feed market at a competitive price point of $1200 per ton, thereby closing the economic loop.84,186
From an environmental perspective, the primary contribution of the Y. lipolytica platform is the significant reduction in Global Warming Potential (GWP) compared to landfilling or incineration.187 Landfilling food waste is a major source of anthropogenic methane (CH4), a greenhouse gas with a warming potential 25 times greater than CO2.84 LCA studies indicate that diverted food waste utilized for yeast fermentation results in a net negative carbon footprint, often displacing 0.5 to 1.2 kg CO2-equivalent per kg of waste processed.184,187
A critical parameter in these LCAs is the energy efficiency of the enzymatic hydrolysis stage. As highlighted in Section 3.1, operating at mild temperatures (50–60 °C) significantly reduces the ‘cradle-to-gate’ energy demand compared to thermo-chemical liquefaction.83 Furthermore, when compared to traditional vegetable oil production (e.g., palm oil), microbial SCO produced from waste by Y. lipolytica circumvents the environmental costs of land-use change and deforestation.188,189 Quantifiable data shows that microbial oil production requires roughly 100 times less land area than palm oil for the same yield, directly contributing to the United Nations Sustainable Development Goal (SDG) 15: Life on Land.186,188,190
Despite the metabolic advantages, the sustainability of the process is often challenged by the energy requirements of downstream recovery.191 Intracellular products like lipids and certain polyols require cell disruption (e.g., high-pressure homogenization or bead milling) and solvent extraction. Current research into ‘green solvents’ and the use of the yeast's native secretome to facilitate autolysis is critical.186 LCA data suggests that shifting from hexane-based extraction to supercritical CO2 or bio-based solvents can reduce the environmental toxicity index of the process by up to 40%.191
The integration of enzymatic hydrolysis and Y. lipolytica fermentation serves as a multi-valent tool for sustainable development: (a) SDG 12 (responsible consumption and production): by transforming 1.3 billion tons of annual global food loss into industrial precursors, the platform exemplifies target 12.3 (reducing food waste);97,183 (b) SDG 9 (industry, innovation, and infrastructure): the development of high-stability biocatalysts, such as the multi-point immobilized enzymes discussed in our research, provides a blueprint for resilient and sustainable industrialization;145,164 and (c) SDG 13 (climate action): the displacement of petroleum-based succinic acid and palm-based lipids provides a low-carbon alternative for the chemical and food industries.150,188
In conclusion, the sustainability of the Y. lipolytica food waste biorefinery is rooted in its ability to decouple industrial growth from resource depletion. While the TEA confirms that high-value specialized metabolites are necessary to ensure financial self-sufficiency, the LCA confirms that the platform is a vital strategy for mitigating the environmental impact of municipal organic waste. Future developments in integrated ‘one-pot’ hydrolysis and fermentation will likely further optimize these parameters, solidifying the role of non-conventional yeasts in a zero-waste future.
3.4. Key limitations and sustainability bottlenecks
Transitioning the oleaginous yeast Y. lipolytica from laboratory-scale experiments to industrial biorefineries represents a cornerstone of the modern circular bioeconomy. While this versatile microorganism excels at converting diverse, low-cost waste streams into high-value bioproducts like lipids, organic acids, and specialty chemicals, realizing its full commercial potential requires overcoming significant hurdles. Looking further to the intricate dynamics of process scale-up, metabolic engineering, and substrate toxicity, we can better understand the current frontier of this research area and the innovative strategies being developed to push it forward.Furthermore, some inhibitory compounds directly inhibit enzymes involved in the synthesis of products of interest, thereby reducing the flux through the pathway and ultimately lowering productivity. In order for yeast cells to cope with and respond to these adverse environmental conditions, sophisticated membrane remodeling strategies were applied. Furan derivatives such as furfural are known to inhibit dehydrogenases and elevate reactive oxygen species, while simultaneously increasing plasma membrane permeability.199 In response, the engineered yeast strains modify their lipidome by increasing the saturation of fatty acids and elevating ergosterol levels to rigidify and stabilize the lipid bilayer.200 Furthermore, integrated phospholipidomics reveals that yeast adjusts the fatty-acyl-chain length of phosphatidylcholine and phosphatidylinositol as a compensatory mechanism to counteract the fluidizing effects of these inhibitors.201,202
Beyond physical pretreatments, the biological breakdown of waste introduces its own chemical hurdles. Anaerobically digested wastes provide abundant VFAs, but their toxicity to the yeast is highly dependent on the specific acid profile. While acetic acid is easily assimilated, higher-chain acids like propionic and butyric acids severely inhibit cellular replication. This delicate dynamic necessitates strict alkaline pH control or complex two-stage feeding strategies to maintain cell viability.203,204 Furthermore, utilizing seawater or fish processing brine to offset freshwater usage introduces extreme osmotic stress.205 This triggers the high osmolarity glycerol (HOG) signaling pathway, which facilitates the intracellular accumulation of glycerol to balance osmotic pressure.206 Structurally, yeast adapts to high salinity by increasing the concentration of complex sphingolipids and very-long-chain fatty acids, which promotes a thicker and less permeable membrane barrier.207 Additionally, the yeast remodels its membrane by increasing the concentration of complex sphingolipids and adjusting the unsaturated/saturated fatty acid (UFA/SFA) ratio to prevent potential-induced depolarization and cellular collapse.208 These multifaceted adaptations are supported by its superior lipid-synthetic machinery underscore the resilience of Y. lipolytica in converting toxic food waste streams into high-value bioproducts.
Compounding this issue is the impairment of primary metabolism. Extensive genetic modifications designed to overproduce specific biochemicals can unintentionally cripple the yeast's native ability to metabolize primary carbon sources like glucose. This massive metabolic burden leads to sluggish growth rates and reduced overall cellular fitness. Consequently, researchers frequently have to perform secondary rounds of engineering simply to restore basic metabolic functions, a requirement that adds immense complexity and time to the strain development pipeline.122
Furthermore, inducing the yeast to accumulate these valuable lipids requires delicate and precise nutrient starvation. While nitrogen limitation is the traditional trigger, recent findings emphasize the equally critical role of phosphate limitation. Balancing these dual starvation parameters in heterogeneous, nutrient-rich waste streams demands highly sophisticated, real-time bioreactor controls. If phosphate starvation is applied too early or too severely, it completely halts cellular replication, making industrial-scale optimization an incredibly delicate balancing act.126
3.5. Strategies to enhance sustainability and future opportunities
The transition of Y. lipolytica from a laboratory-scale model to an industrial cornerstone of the circular bioeconomy necessitates a shift from simplified batch fermentations to integrated, multi-stage bioprocesses.182,211 The inherent heterogeneity of food waste presents significant rheological and metabolic challenges; however, as evidenced by recent advancements, these obstacles provide unique opportunities for engineering biological and mechanical synergies that enhance both techno-economic viability and environmental sustainability.145,212A primary strategy for enhancing sustainability involves the transition toward consolidated bioprocessing (CBP) and synthetic microbial consortia.76,213 Current limitations in food waste valorization often stem from the metabolic specialization of single strains. Future opportunities lie in the design of mutualistic microbial communities—for instance, pairing recombinant Y. lipolytica with cellulolytic or amylolytic species like Bacillus amyloliquefaciens.214 Such systems facilitate the simultaneous liquefaction of complex polysaccharides and the conversion of liberated monomers into high-value fatty acids and lipopeptides, effectively reducing the energy footprint associated with external enzyme supplementation.
Furthermore, the integration of ‘inter-kingdom’ synergies represents a frontier in sustainable waste management. The pairing of Y. lipolytica fermentation with Black Soldier Fly (BSF) larvae rearing offers a compelling solution to the energy-intensive hurdle of downstream biomass recovery.74 By utilizing the yeast to pre-digest complex lipids and enrich the waste stream with high-quality microbial protein, the bioavailability of the substrate for BSF larvae is significantly enhanced.74 This integrated approach bypasses conventional, carbon-heavy separation techniques like high-speed centrifugation, as the larvae act as biological ‘harvesters’ that convert yeast-enriched waste into high-value insect protein and chitin.74,215
On a molecular level, future sustainability is tethered to the advancement of CRISPR-Cas9 and other high-efficiency genetic toolkits to broaden the substrate range of Y. lipolytica.19,216 Engineering strains capable of ‘co-utilization’ (the simultaneous consumption of glucose, xylose, and volatile fatty acids) is essential to minimize residual carbon in the effluent and improve overall process yields.19,216,217 Moreover, the development of robust, ‘inhibitor-tolerant’ phenotypes will reduce the need for intensive detoxifying pretreatments, which are currently a major driver of GWP (Global Warming Potential) in biorefinery life cycles.156,158
Finally, the future of Y. lipolytica sustainability lies in the ‘Multi-Product Biorefinery’ concept.185 Rather than targeting a single metabolite, future facilities must be designed to fractionate the fermentation broth into a portfolio of products—ranging from bulk biofuels and organic acids to high-tier specialized nutraceuticals.97,145,185 By maximizing the value extracted from every kilogram of food waste, the economic ‘tipping point’ for sustainable biomanufacturing can be reached.186,218 This holistic approach, blending advanced genetic engineering with creative bioprocess integration, ensures that Y. lipolytica will fulfill its potential as a primary engine for environmental remediation and sustainable resource recovery.
4. Conclusions and future strategic perspectives
This review has systematically synthesized the multifaceted potential of Y. lipolytica in the valorization of food waste. While laboratory-scale successes in producing lipids, organic acids, and functional metabolites are well-documented, the field must now transcend iterative strain engineering to embrace a more holistic, systems-level integration for industrial viability.To address the chronic challenge of feedstock variability that plagues industrial bioprocessing, future research should move beyond static genetic interventions. Digital-twin-enabled bioprocess control would be the cornerstone of this transition.219,220 By integrating dynamic metabolic models with real-time fermentation telemetry, the system can autonomously reconcile metabolic flux in response to compositional fluctuations, thereby ensuring the batch-to-batch consistency essential for large-scale operations.
From a macro-logistical perspective, the inherent perishability and high transport costs of food waste suggest that modular and decentralized biorefinery units offer significant strategic advantages over centralized facilities. This ‘in-situ valorization’ model facilitates a shorter carbon footprint and aligns more closely with the circular economy's requirement for localized resource recovery, enhancing the overall resilience of the supply chain.
Furthermore, to bolster economic competitiveness, the strategic focus must shift from single-product outputs toward cascading, multi-stream valorization architectures. A ‘zero-waste’ refinery that sequentially recovers high-value proteins, converts residual fractions into specialized lipids via Y. lipolytica, and ultimately upgrades biomass into biochar, will fundamentally redefine the TEA of microbial waste management.
In conclusion, Y. lipolytica should not be viewed merely as a waste-mitigation tool, but as a central hub for high-value biomanufacturing. By bridging the gap between synthetic biology, process engineering, and socio-economic assessments, this yeast platform is poised to spearhead the transformation of food waste from an environmental liability into a strategic asset, driving the global transition toward a carbon-neutral bioeconomy.
Author contributions
Yunkai Tan: conceptualization, methodology, writing – original draft. Huaimin Wang: conceptualization, methodology, writing – original draft. Changyu Hu: writing – original draft. Jing Fu: writing – review & editing. Rodrigo Ledesma-Amaro: writing – review & editing. Fatwa F. Abdi: writing – review & editing. Hsien-Yi Hsu: writing – review & editing. Xue Wang: writing – review & editing. Hairong Cheng: writing – review & editing. Carol Sze Ki Lin: project administration, writing – review & editing, supervision.Conflicts of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.Data availability
Data will be made available on request.Acknowledgements
This publication is fully funded by the Strategic Seed Fund of City University of Hong Kong (Project No. 7020113).References
- W. Jia, S. Li, J. Wang, J. T. Lee, C. S. K. Lin, O. Mašek, H. Zhang and X. Yuan, Green Chem., 2024, 26, 1790–1805 RSC.
- P. K. Sarangi, P. Pal, A. K. Singh, U. K. Sahoo and P. Prus, Resources, 2024, 13, 164 CrossRef.
- M. S. Mia, M. M. Ahmed and W. Zzaman, Discover Food, 2025, 5, 275 CrossRef.
- Y. M. Wang, Z. W. Yuan and Y. Tang, Resour., Environ. Sustainability, 2021, 4, 100023 CrossRef.
- I. Berzina and K. Spalvins, Environ. Clim. Technol., 2023, 27, 639–653 CAS.
- P. Torabi, N. Hamdami, N. Soltanizadeh, O. Farhadian and A. Le-Bail, Cleaner Mater., 2024, 13, 100269 CrossRef CAS.
- X. Zhang, Q. Qin, X. Sun and W. Wang, Front. Nutr., 2022, 9, 986705 CrossRef PubMed.
- G. Wu, J. W. R. Chong, K. S. Khoo, D. Y. Y. Tang and P. L. Show, Biotechnol. Biofuels Bioprod., 2025, 18, 74 CrossRef PubMed.
- J. d. V. Moreira, PhD thesis, Université Paris-Saclay, 2019.
- A. Soni, S. Patil, A. Ghosalkar, S. Nandi and P. Kumbhar, J. Pure Appl. Microbiol., 2024, 18, 2095–2112 CrossRef.
- C. Li, K. L. Ong, Z. Cui, Z. Sang, X. Li, R. D. Patria, Q. Qi, P. Fickers, J. Yan and C. S. K. Lin, J. Hazard. Mater., 2021, 401, 123414 CrossRef CAS PubMed.
- C. M. Bauer, C. Schmitz and M. F. Landell, Biotechnol. Res. Innovation, 2022, 6, e2022001 CrossRef.
- M. Kavscek, G. Bhutada, T. Madl and K. Natter, BMC Syst. Biol., 2015, 9, 72 CrossRef.
- D. Xie, E. N. Jackson and Q. Zhu, Appl. Microbiol. Biotechnol., 2015, 99, 1599–1610 CrossRef CAS PubMed.
- A. F. Abdala, A. P. Gallardo, L. G. Olvera and E. M. Silva, Bioresour. Bioprocess., 2017, 4, 5 CrossRef PubMed.
- J. Cescut, B. Portelli and C. Molina-Jouve, Presented in Part at the European Congress of Chemical Engineering - ECCE 2015, Nice, France, 2015 Search PubMed.
- W. Sabra, R. R. Bommareddy, G. Maheshwari, S. Papanikolaou and A. P. Zeng, Microb. Cell Fact., 2017, 16, 78 CrossRef.
- C. Holkenbrink, J. Beder and M. I. Dam, Presented in Part at the Metabolic Engineering 11, Awaj, Japan, 2016 Search PubMed.
- T. Q. Shi, H. Huang, E. J. Kerkhoven and X. J. Ji, Appl. Microbiol. Biotechnol., 2018, 102, 9541–9548 CrossRef CAS PubMed.
- J. Dahlin, PhD thesis, Technical University of Denmark, 2020.
- A. Tsirigka, E. Theodosiou, S. I. Patsios, A. Tsoureki, A. Andreadelli, E. Papa, A. Aggeli, A. J. Karabelas and A. M. Makris, Microb. Cell Fact., 2023, 22, 62 CrossRef CAS.
- M. E. Jach, T. Baj, M. Juda, R. Swider, B. Mickowska and A. Malm, AMB Express, 2020, 10, 35 CrossRef CAS.
- D. Jobson, G. G. Karunasena, N. Nabi, D. Pearson and E. Dunstan, Sustainability, 2024, 16, 1963 CrossRef.
- Z. Wang, X. Li, Y. Li, H. Liu, C. S. Ki Lin, J. Sun and Q. Wang, Environ. Pollut., 2024, 361, 124901 CrossRef CAS PubMed.
- M. J. Page, J. E. McKenzie, P. M. Bossuyt, I. Boutron, T. C. Hoffmann, C. D. Mulrow, L. Shamseer, J. M. Tetzlaff, E. A. Akl and S. E. Brennan, Br. Med. J., 2021, 372 Search PubMed.
- Z. Y. Liu, T. S. P. de Souza, B. Holland, F. Dunshea, C. Barrow and H. A. R. Suleria, Processes, 2023, 11, 840 CrossRef CAS.
- R. Kumari, A. Singh, R. Sharma and P. Malaviya, Environ. Chem. Lett., 2024, 22, 1759–1790 CrossRef CAS.
- Z. Wang, X. Li, H. Liu, C. S. K. Lin and Q. Wang, Renewable Sustainable Energy Rev., 2025, 224, 116086 CrossRef CAS.
- A. Sarker, R. Ahmmed, S. Ahsan, J. Rana, M. K. Ghosh and R. Nandi, Sustainable Food Technol., 2024, 2, 48–69 RSC.
- N. Dutta, M. Usman, G. Luo and S. C. Zhang, Sustainable Chem., 2022, 3, 35–55 CrossRef CAS.
- J. Chen, X. Ma, M. Liang, Z. Guo, Y. Cai, C. Zhu, Z. Wang, S. Wang, J. Xu and H. Ying, Waste, 2024, 2, 451–473 CrossRef.
- C. F. T. Baptista, R. P. Rodrigues and M. J. Quina, Sustainability, 2024, 16, 4368 CrossRef.
- A. Singh, P. Prajapati, S. Vyas, V. K. Gaur, R. Sindhu, P. Binod, V. Kumar, R. R. Singhania, M. K. Awasthi and Z. Zhang, Bioenergy Res., 2023, 16, 105–122 CrossRef.
- I. V. Dos Anjos, N. Coelho, H. Duarte, D. N. Proenca, M. F. Duarte, R. Barros, S. Raposo, S. Goncalves, A. Romano and B. Medronho, Polymers, 2025, 17, 2251 CrossRef CAS PubMed.
- Y. Y. Yao, Y. Ye, K. Xiong, S. C. Mao, J. W. Jiang, Y. Q. Chen, X. Li, H. B. Liu, L. C. Liu, B. Cai and S. Song, Foods, 2026, 15, 402 CrossRef CAS PubMed.
- S. Plakantonaki, I. Roussis, D. Bilalis and G. Priniotakis, Processes, 2023, 11, 1580 CrossRef CAS.
- R. Venugopalan, A. Negi, P. Balasubramanian, S. Chandra, S. Paramasivam and J. A. Moses, Curr. Res. Nutr. Food Sci., 2025, 14 Search PubMed.
- R. C. Ray, S. S. Behera, O. Awogbemi, B. S. Sooch, H. Thatoi, S. Rath and N. Aguilar-Rivera, AIMS Microbiol., 2025, 11, 462–500 CAS.
- S. Maan, A. Abdelhafez, A. S. Ahmed and E. Hassan, Arab Univ. J. Agric. Sci., 2022, 30, 43–53 Search PubMed.
- P. Kumar, in Biomass-Based Green Technology for Circular Economy, CRC Press, Boca Raton, USA, 1st edn, 2025, pp. 278–294 Search PubMed.
- M. Jahangeer, M. U. Rehman, R. Nelofer, M. Nadeem, B. Munir, W. Smułek, T. Jesionowski and S. A. Qamar, Top. Catal., 2025, 68, 929–950 CrossRef CAS.
- M. Pabba, A. M. Shah, M. H. Salim, S. Mettu, M. M. R. Talukder and S. R. Puniredd, Ind. Eng. Chem. Res., 2026, 65, 3027–3048 CrossRef CAS.
- X. Wang, Y. Wang, L. Zuo, S. Guo, P. Song, W. Kong and B. Shen, Biomass Convers. Biorefin., 2025, 15, 75–97 CrossRef CAS.
- Y. Y. Lee, T. K. Tang, E. S. Chan, E. T. Phuah, O. M. Lai, C. P. Tan, Y. Wang, N. A. Ab Karim, N. H. Mat Dian and J. S. Tan, Crit. Rev. Food Sci. Nutr., 2022, 62, 4169–4185 CrossRef CAS PubMed.
- F. Schermer, PhD thesis, University of Regensburg, 2023.
- Y. Li, H. Zhu, F. Zhou, Y. Zheng, Y. Zhang and Y. Zheng, Sci. Rep., 2025, 15, 16497 CrossRef CAS.
- N. O. Ezeh, T. N. T. Nwagu, O. C. Amadi, B. Okolo, A. Moneke and R. Agu, Biocatal. Agric. Biotechnol., 2023, 47 Search PubMed.
- J. H. Mou, W. Yan, Z. H. Qin, M. A. Haque, Y. H. Miao, F. X. Xin, X. Wang, P. Fickers and C. S. K. Lin, Chem. Eng. J., 2024, 481, 148408 CrossRef CAS.
- W. A. Rasaq, B. Matyjewicz, K. Swiechowski, Z. Lazar, P. Kupaj, T. Janek, M. Valentin and A. Bialowiec, J. Cleaner Prod., 2024, 456, 142385 CrossRef CAS.
- A. Abun, K. Haetami, D. Rusmana and R. F. Ramdhan, Microbiologyopen, 2025, 14, e70040 CrossRef CAS.
- B. Zieniuk, M. Woloszynowska, E. Bialecka-Florjanczyk and A. Fabiszewska, Sustainability, 2020, 12, 1–18 CrossRef.
- D. Gottardi, M. Ciccone, L. Siroli, R. Lanciotti and F. Patrignani, Fermentation, 2022, 8, 708 CrossRef CAS.
- A. Lalić, J. Jagelavičiūtė, T. Rezić, Z. Trivunović, D. Žadeikė and L. Bašinskienė, Sustainability, 2025, 17 Search PubMed.
- S. Mitri, N. Louka, T. Rossignol, R. G. Maroun and M. Koubaa, Fermentation, 2025, 11, 611 CrossRef CAS.
- V. Korka, A. Petropoulos, S. M. Ioannidou, C. S. K. Lin, A. Koutinas and P. Fickers, FEMS Yeast Res., 2025, 25, foaf052 CrossRef CAS PubMed.
- C. Li, W. Lin, K. L. Ong, J. Mou, C. S. K. Lin and P. Fickers, in Synthetic Biology of Yeasts, ed. F. Darvishi Harzevili, Springer International Publishing, Heidelberg, Germany, 2022, ch. 9, pp. 227–250 Search PubMed.
- S. Heppner and Y. D. Livney, Future Foods, 2025, 11, 100523 CrossRef CAS.
- A. Laribi, J. Brys, A. Selmania, A. Ikhlef, I. Btaiche, A. Mouzai, B. Zieniuk and D. N. Bouchedja, Foods, 2025, 14, 3696 CrossRef CAS PubMed.
- C. Ciliberti, A. Biundo, M. Colacicco, G. Agrimi and I. Pisano, Chem. Eng. Trans., 2022, 93, 241–246 Search PubMed.
- J. Malajowicz, A. Fabiszewska, B. Zieniuk, J. Brys, M. Kozlowska and K. Marciniak-Lukasiak, Foods, 2025, 14, 187 CrossRef CAS PubMed.
- S. Baksi, U. Sarkar, R. Villa, D. Basu and D. Sengupta, Sustainable Energy Technol. Assess., 2023, 55, 102963 CrossRef.
- N. Abad, A. Uranga, B. Ayo, J. M. Arrieta, Z. Bana, I. Azua, I. Artolozaga, J. Iriberri, S. J. Gonzalez-Roji and M. Unanue, Environ. Microbiol., 2023, 25, 548–561 CrossRef CAS.
- T. Mkilima, Biofuels, Bioprod. Biorefin., 2025, 19, 2581–2608 CrossRef CAS.
- J. C. Martins-Vieira, P. C. Torres-Mayanga and D. Lachos-Perez, Bioenergy Res., 2023, 16, 1296–1317 CrossRef CAS.
- M. A. Khan, B. H. Hameed, M. R. Siddiqui, Z. A. Alothman and I. H. Alsohaimi, Foods, 2022, 11, 4036 CrossRef CAS PubMed.
- K. Wierzchowska, M. Roszko, D. Derewiaka, K. Szulc, B. Zieniuk, D. Nowak and A. Fabiszewska, Food Biosci., 2024, 62, 105321 CrossRef CAS.
- M. Yang, L. Ding, P. Wang, Y. Wu, C. Areeprasert, M. Wang, X. Chen, F. Wang and G. Yu, Fuel, 2023, 334, 126790 CrossRef CAS.
- Y. X. Yan, A. Z. Liu, Q. Y. Shang, R. P. Singh, D. F. Fu, T. A. Kotsopoulos and I. A. Fotidis, J. Cleaner Prod., 2025, 521, 146270 CrossRef CAS.
- T. Chandukishore, K. Narasimhulu, R. Ledesma-Amaro and A. A. Prabhu, Chem. Eng. J., 2025, 519, 165519 CrossRef CAS.
- D. Correa-Galeote, L. Argiz, A. Mosquera-Corral, A. V. Del Rio, B. Juarez-Jimenez, J. Gonzalez-Lopez and B. Rodelas, New Biotechnol., 2022, 71, 47–55 CrossRef CAS PubMed.
- O. D. Ianieva, Mikrobiol. Zh., 2020, 82, 65–87 CrossRef.
- S. Saraiva, C. Saraiva, T. Lazou, S. Chaintoutis, J. R. Mesquita, A. C. Coelho and P. Poeta, Ger. J. Vet. Res., 2025, 5, 118–127 Search PubMed.
- P. K. Chaurasia, S. L. Bharati, S. Singh, A. M. Sivalingam, S. Shankar and A. Mani, RSC Sustainability, 2025, 3, 1234–1266 RSC.
- H. Lu, Y. Liu, W. Shen, Y. Zhou, X. Ma, S. Sun, X. Dong, F. Ji, H. Tong, J. Xu, G. He and W. Xu, Waste Manage., 2023, 166, 152–162 CrossRef CAS PubMed.
- L. Garnier, F. Valence, A. Pawtowski, L. Auhustsinava-Galerne, N. Frotte, R. Baroncelli, F. Deniel, E. Coton and J. Mounier, Int. J. Food Microbiol., 2017, 241, 191–197 CrossRef CAS PubMed.
- W. Shang, Y. M. Zhang, M. Z. Ding, H. Z. Sun, J. X. He and J. S. Cheng, J. Environ. Manage., 2024, 371, 123177 CrossRef CAS PubMed.
- Z. Zhang, H. H. Wong, P. L. Albertson, M. D. Harrison, W. O. Doherty and I. M. O'Hara, Bioresour. Technol., 2015, 192, 367–373 CrossRef CAS PubMed.
- M. Colacicco, C. Ciliberti, A. Biundo, G. Agrimi and I. Pisano, Chem. Eng. Trans., 2022, 93, 247–252 Search PubMed.
- J. L. Fraga, C. P. L. Souza, A. D. S. Pereira, E. C. G. Aguieiras, L. O. de Silva, A. G. Torres, D. G. Freire and P. F. F. Amaral, 3 Biotech, 2021, 11, 191 CrossRef PubMed.
- J. X. He, Z. J. Hou, B. Qiao, M. Z. Ding, Y. M. Zhang, Q. M. Xu and J. S. Cheng, J. Environ. Chem. Eng., 2024, 12, 113161 CrossRef CAS.
- C. S. Zhang, X. X. Kang, F. H. Wang, Y. F. Tian, T. Liu, Y. Y. Su, T. T. Qian and Y. F. Zhang, Energy, 2020, 208, 118379 CrossRef CAS.
- S. Chen, Y. Lu, W. Wang, Y. Hu, J. Wang, S. Tang, C. S. K. Lin and X. Yang, Front. Microbiol., 2022, 13, 960558 CrossRef PubMed.
- J.-H. Mou, W. Yan, Z.-H. Qin, M. A. Haque, Y.-H. Miao, F.-X. Xin, X. Wang, P. Fickers and C. S. K. Lin, Chem. Eng. J., 2023, 481, 148408 CrossRef.
- E. Carsanba, B. Agirman, S. Papanikolaou, P. Fickers and H. Erten, Fermentation, 2023, 9, 687 CrossRef CAS.
- C. Gao, Q. Qi, C. Madzak and C. S. Lin, J. Ind. Microbiol. Biotechnol., 2015, 42, 1255–1262 CrossRef CAS PubMed.
- H. J. Kim, Y. G. Choi, G. D. Kim, S. H. Kim and T. H. Chung, Water Sci. Technol., 2005, 52, 51–59 CrossRef CAS.
- K. A. Barrera-Rivera and A. Martinez-Richa, Molecules, 2017, 22, 1917 CrossRef PubMed.
- M. Zhang, D. Zhang, Y. Wei, B. Zhou, C. Yan, D. Wang, J. Liang and L. Zhou, Waste Manage., 2022, 151, 1–9 CrossRef CAS PubMed.
- Q. Wang, Z. T. Qi, W. L. Fu, M. Z. Pan, X. D. Ren, X. Zhang and Z. M. Rao, Fermentation, 2024, 10, 648 CrossRef CAS.
- I. Ntaikou, M. Alexandropoulou, M. Kamilari, S. A. Alamri, Y. S. Moustafa, M. Hashem, G. Antonopoulou and G. Lyberatos, Energy, 2023, 281, 128259 CrossRef CAS.
- H. Wang, H. F. Ma, C. Jin, J. X. Ma, X. Li, L. X. Tang and J. Si, Ind. Crops Prod., 2025, 227, 120845 CrossRef CAS.
- M. Larroude, PhD thesis, Université Paris-Saclay, 2020.
- B. Zieniuk and A. Fabiszewska, World J. Microbiol. Biotechnol., 2018, 35, 10 CrossRef PubMed.
- Y. V. Soong, N. Liu, S. Yoon, C. Lawton and D. Xie, Eng. Life Sci., 2019, 19, 423–443 CrossRef CAS PubMed.
- S. S. Khaligh, E. Polat and M. Altinbas, Waste Biomass Valorization, 2023, 14, 2037–2059 CrossRef.
- S. A. van der Hoek, M. Rusnak, I. H. Jacobsen, J. L. Martinez, D. B. Kell and I. Borodina, FEBS Lett., 2022, 596, 1356–1364 CrossRef CAS PubMed.
- P. T. Thomsen, S. Meramo, L. Ninivaggi, E. Pasutto, M. Babaei, P. M. Avila-Neto, M. C. Pastor, P. Sabri, D. Rago, T. U. Parekh, S. Hunding, L. E. J. Christiansen, S. Sukumara and I. Borodina, Nat. Microbiol., 2023, 8, 2290–2303 CrossRef CAS PubMed.
- D. T. H. Do, C. W. Theron and P. Fickers, Microorganisms, 2019, 7, 229 CrossRef CAS PubMed.
- P. Khunnonkwao, S. Thitiprasert, P. Jaiaue, K. Khumrangsee, B. Cheirsilp and N. Thongchul, Heliyon, 2024, 10, e30830 CrossRef CAS PubMed.
- M. P. Diez, E. Barahona, M. A. de la Rubia, A. F. Mohedano and E. Diaz, Int. J. Hydrogen Energy, 2024, 89, 1383–1393 CrossRef CAS.
- G. Kaur, H. M. Wang, M. H. To, S. L. K. W. Roelants, W. Soetaert and C. S. K. Lin, J. Cleaner Prod., 2019, 232, 1–11 CrossRef.
- T. H. Kwan, D. Pleissner, K. Y. Lau, J. Venus, A. Pommeret and C. S. Lin, Bioresour. Technol., 2015, 198, 292–299 CrossRef CAS PubMed.
- C. Li, S. Gao, X. Yang and C. S. K. Lin, Bioresour. Technol., 2018, 249, 612–619 CrossRef CAS PubMed.
- S. Rehman, Y. S. Yang, R. D. Patria, T. Zulfiqar, N. K. Khanzada, R. J. Khan, C. S. K. Lin, D. J. Lee and S. Y. Leu, Bioresour. Technol., 2023, 390, 129858 CrossRef CAS PubMed.
- X. Yang, M. Zhu, X. Huang, C. S. Lin, J. Wang and S. Li, Bioresour. Technol., 2015, 198, 47–54 CrossRef CAS PubMed.
- N. Kuncharoen, S. Techo, A. Savarajara and S. Tanasupawat, Mycology, 2020, 11, 279–286 CrossRef CAS PubMed.
- M. Lopes, S. M. Miranda, J. M. Alves, A. S. Pereira and I. Belo, Eur. J. Lipid Sci. Technol., 2019, 121, 1800188 CrossRef.
- M. Colacicco, C. Ciliberti, G. Agrimi, A. Biundo and I. Pisano, Energies, 2022, 15, 5217 CrossRef CAS.
- M. de Vicente, C. Gonzalez-Fernandez, J. M. Nicaud and E. Tomas-Pejo, Microb. Cell Fact., 2025, 24, 32 CrossRef CAS PubMed.
- R. Yang, Z. Chen, P. Hu, S. Zhang and G. Luo, Bioresour. Technol., 2022, 361, 127677 CrossRef CAS PubMed.
- D. Atzmüller, F. Hawe, D. Sulzenbacher and A. Cristobal-Sarramian, Agron. Res., 2019, 17, 2172–2179 Search PubMed.
- M. Rakicka, B. Rukowicz, A. Rywińska, Z. Lazar and W. Rymowicz, J. Cleaner Prod., 2016, 139, 905–913 CrossRef CAS.
- N. Sofeo, M. G. Toi, E. Q. G. Ee, J. Y. Ng, C. T. Busran, B. R. Lukito, A. Thong, C. Hermansen, E. C. Peterson, R. Glitsos and P. Arumugam, Bioresour. Technol., 2024, 394, 130302 CrossRef CAS PubMed.
- C. Gao, X. Yang, H. Wang, C. P. Rivero, C. Li, Z. Cui, Q. Qi and C. S. K. Lin, Biotechnol. Biofuels, 2016, 9, 179 CrossRef PubMed.
- R. Gao, Z. Li, X. Zhou, W. Bao, S. Cheng and L. Zheng, Biotechnol. Biofuels, 2020, 13, 3 CrossRef CAS PubMed.
- M. K. Jensen and J. D. Keasling, FEMS Yeast Res., 2015, 15, 1–10 CAS.
- B. Simonaviciene, A. Araoyinbo, J. Ali, J. McGowan, D. A. Fitzpatrick, G. Jones, C. Ferreira, A. R. Pitt, C. M. Spickett, V. Postis and C. de Marcos Lousa, Biotechnol. Biofuels Bioprod., 2025, 18, 76 CrossRef CAS PubMed.
- M. Larroude, D. Onesime, O. Rue, J. M. Nicaud and T. Rossignol, Microorganisms, 2021, 9, 838 CrossRef CAS PubMed.
- T. Dulermo and J. M. Nicaud, Metab. Eng., 2011, 13, 482–491 CrossRef CAS PubMed.
- E. U. Kiran and A. Trzcinski, EC Microbiol., 2017, 8, 98–112 Search PubMed.
- A. Botelho, A. Penha, J. Fraga, A. Barros-Timmons, M. A. Coelho, M. Lehocky, K. Stepankova and P. Amaral, Polymers, 2020, 12, 649 CrossRef CAS PubMed.
- X. Yang, H. Wang, C. Li and C. S. K. Lin, J. Agric. Food Chem., 2017, 65, 4133–4139 CrossRef CAS PubMed.
- K. Spalvins, Z. Geiba, Z. Kusnere and D. Blumberga, Environ. Clim. Technol., 2020, 24, 457–469 CAS.
- K. Wierzchowska, D. Derewiaka, B. Zieniuk, D. Nowak and A. Fabiszewska, Eur. Food Res. Technol., 2023, 249, 2675–2688 CrossRef CAS.
- E. Tomas-Pejo, S. Morales-Palomo and C. Gonzalez-Fernandez, Bioengineered, 2023, 14, 2286723 CrossRef PubMed.
- S. Morales-Palomo, E. Tomas-Pejo and C. Gonzalez-Fernandez, Microb. Biotechnol., 2023, 16, 372–380 CrossRef CAS PubMed.
- M. Lopes, S. M. Miranda, J. M. Alves, A. S. Pereira and I. Belo, Eur. J. Lipid Sci. Technol., 2019, 121, 1800188 CrossRef.
- D. R. A. Sanya and D. Onesime, Appl. Microbiol. Biotechnol., 2022, 106, 7397–7416 CrossRef CAS PubMed.
- M. L. Sun, X. Gao, L. Lin, J. Yang, R. Ledesma-Amaro and X. J. Ji, J. Agric. Food Chem., 2024, 72, 94–107 CrossRef CAS PubMed.
- S. Hassoun, F. Jefferson, X. Shi, B. Stucky, J. Wang and E. Rosa, Integr. Comp. Biol., 2022, 61, 2267–2275 CrossRef PubMed.
- M. Carmona-Cabello, I. L. Garcia, D. Leiva-Candia and M. P. Dorado, Curr. Opin. Green Sustainable Chem., 2018, 14, 67–79 CrossRef.
- Y. Wang, P. Xue, M. Cao, T. Yu, S. T. Lane and H. Zhao, Chem. Rev., 2021, 121, 12384–12444 CrossRef CAS PubMed.
- L. Alzubaidi, J. Zhang, A. J. Humaidi, A. Al-Dujaili, Y. Duan, O. Al-Shamma, J. Santamaria, M. A. Fadhel, M. Al-Amidie and L. Farhan, J. Big Data, 2021, 8, 53 CrossRef PubMed.
- A. Dhillon and G. K. Verma, Prog. Artif. Intell., 2019, 9, 85–112 CrossRef.
- Z. Wu, W. Chen, Y. Hong, Y. Wang and P. Xu, Synth. Syst. Biotechnol., 2025, 10, 1275–1283 CrossRef CAS PubMed.
- Y. Guo, L. Su, Q. Liu, Y. Zhu, Z. Dai and Q. Wang, Comput. Struct. Biotechnol. J., 2022, 20, 2503–2511 CrossRef CAS PubMed.
- J. J. Czajka, T. Oyetunde and Y. J. Tang, Metab. Eng., 2021, 67, 227–236 CrossRef CAS PubMed.
- J. E. Flores, D. M. Claborne, Z. D. Weller, B. M. Webb-Robertson, K. M. Waters and L. M. Bramer, Front. Artif. Intell., 2023, 6, 1098308 CrossRef PubMed.
- A. Yetgin, Quant. Biol., 2025, 13, e70002 CrossRef PubMed.
- K. Wang, W. Zhao, L. Lin, T. Wang, P. Wei, R. Ledesma-Amaro, A. H. Zhang and X. J. Ji, Biotechnol. Bioeng., 2023, 120, 1015–1025 CrossRef CAS PubMed.
- S. K. Khanal, A. Tarafdar and S. You, Bioresour. Technol., 2023, 375, 128826 CrossRef CAS PubMed.
- M. C. Coleman and D. E. Block, Biotechnol. Bioeng., 2006, 95, 412–423 CrossRef CAS PubMed.
- B. Nithya, C. P. Jetlin, G. David Raj, A. Allwyn Sundarraj, P. Nehru and P. Selvakumar, in The Role of Artificial Intelligence in Advancing Applied Life Sciences, ed. T. Emara, E. Hassan, T. Trinh, G. Li and A. Saber, IGI Global, Hershey, USA, 2025, ch. 8, pp. 235–264 Search PubMed.
- Z. M. Hein, D. Guruparan, B. Okunsai, C. M. N. Che Mohd Nassir, M. D. C. Ramli and S. Kumar, Biology, 2025, 14, 1453 CrossRef PubMed.
- J. Mou, PhD thesis, City University of Hong Kong, 2023.
- J. Fu, S. Zaghen, H. Lu, O. Konzock, N. Poorinmohammad, A. Kornberg, R. Ledesma-Amaro, D. Koseto, A. Wentzel, F. Di Bartolomeo and E. J. Kerkhoven, Sci. Adv., 2024, 10, eadn0414 CrossRef CAS PubMed.
- K. Wierzchowska, K. Szulc, B. Zieniuk and A. Fabiszewska, Molecules, 2025, 30, 959 CrossRef CAS PubMed.
- M. Spagnuolo, M. Shabbir Hussain, L. Gambill and M. Blenner, Front. Microbiol., 2018, 9, 1077 CrossRef PubMed.
- C. Li, K. L. Ong, X. F. Yang and C. S. K. Lin, Chem. Eng. J., 2019, 371, 804–812 CrossRef CAS.
- E. Stylianou, C. Pateraki, D. Ladakis, C. Damala, A. Vlysidis, M. Latorre-Sánchez, C. Coll, C. S. K. Lin and A. Koutinas, Biochem. Eng. J., 2021, 174, 108099 CrossRef CAS.
- M. E. Kıymacı, D. ŞİMŞEk and N. Altanlar, J. Fac. Pharm. Ankara Univ., 2022, 46, 450–457 Search PubMed.
- C. E. Robles-Rodríguez, R. Muñoz-Tamayo, C. Bideaux, N. Gorret, S. E. Guillouet, C. Molina-Jouve, G. Roux and C. A. Aceves-Lara, Biotechnol. Bioeng., 2017, 115, 1137–1151 CrossRef PubMed.
- D. Sarris, E. Tsouko, A. Photiades, S. S. Tchakouteu, P. Diamantopoulou and S. Papanikolaou, Microorganisms, 2023, 11, 2243 CrossRef CAS PubMed.
- V. Kumar and P. Longhurst, Curr. Opin. Green Sustainable Chem., 2018, 13, 118–122 CrossRef.
- M. Rakicka-Pustulka, J. Miedzianka, D. Jama, S. Kawalec, K. Liman, T. Janek, G. Skaradzinski, W. Rymowicz and Z. Lazar, Microb. Cell Fact., 2021, 20, 195 CrossRef CAS PubMed.
- O. Konzock, S. Zaghen and J. Norbeck, BMC Microbiol., 2021, 21, 77 CrossRef CAS PubMed.
- A. A. Prabhu, D. J. Thomas, R. Ledesma-Amaro, G. A. Leeke, A. Medina, C. Verheecke-Vaessen, F. Coulon, D. Agrawal and V. Kumar, Microb. Cell Fact., 2020, 19, 121 CrossRef CAS PubMed.
- J. Kim, H. F. Son, S. Hwang, G. Gong, J. K. Ko, Y. Um, S. O. Han and S. M. Lee, Int. J. Mol. Sci., 2022, 23, 4761 CrossRef CAS PubMed.
- P. M. Niang, A. Arguelles-Arias, S. Steels, O. Denies, J. M. Nicaud and P. Fickers, Cell Biol. Int., 2020, 44, 651–660 CrossRef CAS PubMed.
- N. Wang, P. Chi, Y. W. Zou, Y. R. Xu, S. Xu, M. Bilal, P. Fickers and H. R. Cheng, Biotechnol. Biofuels, 2020, 13, 176 CrossRef CAS PubMed.
- K. K. Miller and H. S. Alper, Appl. Microbiol. Biotechnol., 2019, 103, 9251–9262 CrossRef CAS PubMed.
- C. Goncalves, C. Ferreira, L. G. Goncalves, D. L. Turner, M. J. Leandro, M. Salema-Oom, H. Santos and P. Goncalves, Front. Microbiol., 2019, 10, 2510 CrossRef PubMed.
- S. Xu, Y. W. Zou, L. Y. Ji, M. Bilal and H. R. Cheng, ACS Sustainable Chem. Eng., 2024, 12, 4041–4050 CrossRef CAS.
- L. Huang, W. Wang, K. Wang, Y. Li, J. Zhou, A. Pang, B. Zhang, Z. Liu and Y. Zheng, Bioprocess Biosyst. Eng., 2024, 47, 1659–1668 CrossRef CAS PubMed.
- J. Liu, J. Li, P. Chen, Y. Zeng, J. Yang and Y. Sun, J. Sci. Food Agric., 2024, 104, 5999–6007 CrossRef CAS PubMed.
- C. Ratledge and S. Hopkins, in Modifying Lipids for Use in Food, Woodhead Publishing, Cambridge, UK, 2006, ch. 5, pp. 80–113 Search PubMed.
- Y. K. Park and J. M. Nicaud, Microorganisms, 2020, 8, 1937 CrossRef CAS PubMed.
- F. Thevenieau and J.-M. Nicaud, Ocl, 2013, 20, D603 CrossRef.
- S. Papanikolaou and G. Aggelis, Eur. J. Lipid Sci. Technol., 2011, 113, 1031–1051 CrossRef CAS.
- W. Rymowicz, A. Rywinska, B. Zarowska and P. Juszczyk, Chem. Pap., 2006, 60, 391–394 CAS.
- T. Sun, Y. Yu, K. Wang, R. Ledesma-Amaro and X. J. Ji, Bioresour. Technol., 2021, 337, 125484 CrossRef CAS PubMed.
- J. M. Otero, D. Cimini, K. R. Patil, S. G. Poulsen, L. Olsson and J. Nielsen, PLoS One, 2013, 8, e54144 CrossRef CAS PubMed.
- P. Fickers, H. Cheng and C. Sze Ki Lin, Microorganisms, 2020, 8, 574 CrossRef CAS PubMed.
- Y. Xu, P. Chi, J. Lv, M. Bilal and H. Cheng, Biochim. Biophys. Acta, Gen. Subj., 2021, 1865, 129740 CrossRef CAS PubMed.
- Y. Xu, L. Ji, S. Xu, M. Bilal, A. Ehrenreich, Z. Deng and H. Cheng, Biochim. Biophys. Acta, Gen. Subj., 2023, 1867, 130289 CrossRef CAS PubMed.
- Y. Xu, P. Chi, M. Bilal and H. Cheng, Appl. Microbiol. Biotechnol., 2019, 103, 5143–5160 CrossRef CAS PubMed.
- M. Bilal, M. Asgher, H. Cheng, Y. Yan and H. M. N. Iqbal, Crit. Rev. Biotechnol., 2019, 39, 202–219 CrossRef CAS PubMed.
- P. Chi, S. Q. Wang, X. M. Ge, M. Bilal, P. Fickers and H. R. Cheng, Biochem. Eng. J., 2019, 149, 107259 CrossRef CAS.
- P. F. F. Amaral, J. M. da Silva, M. Lehocky, A. M. V. Barros-Timmons, M. A. Z. Coelho, I. M. Marrucho and J. A. P. Coutinho, Process Biochem., 2006, 41, 1894–1898 CrossRef CAS.
- G. Stephanopoulos, A. A. Aristidou and J. Nielsen, Metabolic Engineering: Principles and Methodologies, Elsevier, Amsterdam, Netherlands 1998 Search PubMed.
- S. Yang, R. Chen, X. Cao, G. Wang and Y. J. Zhou, Nat. Commun., 2024, 15, 253 CrossRef CAS PubMed.
- L. Murphy and D. J. O’Connell, Fermentation, 2024, 10, 583 CrossRef CAS.
- K. Wierzchowska, B. Zieniuk and A. Fabiszewska, Waste Biomass Valorization, 2022, 13, 757–779 CrossRef CAS.
- N. Elginöz, K. Khatami, I. Owusu-Agyeman and Z. Cetecioglu, Front. Sustainable Food Syst., 2020, 4, 23 CrossRef.
- M. K. Awasthi, S. Harirchi, T. Sar, V. Vs, K. Rajendran, R. Gomez-Garcia, C. Hellwig, P. Binod, R. Sindhu, A. Madhavan, A. N. A. Kumar, V. Kumar, D. Kumar, Z. Zhang and M. J. Taherzadeh, Bioresour. Technol., 2022, 360, 127592 CrossRef CAS PubMed.
- E. E. Karamerou, S. Parsons, M. C. McManus and C. J. Chuck, Biotechnol. Biofuels, 2021, 14, 57 CrossRef CAS PubMed.
- S. Sae-ngae, B. Cheirsilp, Y. Louhasakul, T. T. Suksaroj and P. Intharapat, Sustainable Environ. Res., 2020, 30, 11 CrossRef.
- F. Abeln and C. J. Chuck, Microb. Cell Fact., 2021, 20, 221 CrossRef CAS PubMed.
- P. Shaigani, D. Awad, V. Redai, M. Fuchs, M. Haack, N. Mehlmer and T. Brueck, Microb. Cell Fact., 2021, 20, 220 CrossRef CAS PubMed.
- Z. E. Duman-Ozdamar, V. A. P. Martins Dos Santos, J. Hugenholtz and M. Suarez-Diez, Microb. Cell Fact., 2022, 21, 228 CrossRef PubMed.
- S. Parsons, M. J. Allen, F. Abeln, M. McManus and C. J. Chuck, J. Cleaner Prod., 2019, 232, 1272–1281 CrossRef CAS.
- M. Dourou, D. Aggeli, S. Papanikolaou and G. Aggelis, Appl. Microbiol. Biotechnol., 2018, 102, 2509–2523 CrossRef CAS PubMed.
- W. Rymowicz, A. Rywinska and W. Gladkowski, Chem. Pap., 2008, 62, 239–246 CAS.
- N. Loira, T. Dulermo, J. M. Nicaud and D. J. Sherman, BMC Syst. Biol., 2012, 6, 35 CrossRef PubMed.
- S. Maina, V. Kachrimanidou and A. Koutinas, Curr. Opin. Green Sustainable Chem., 2017, 8, 18–23 CrossRef.
- K. E. d. S. Oliveira and S. R. da Cal Seixas, in Encyclopedia of Sustainability in Higher Education, Springer, 2019, pp. 2007–2013 Search PubMed.
- J. M. Nicaud, Yeast, 2012, 29, 409–418 CrossRef CAS PubMed.
- I. Chiyanzy, M. Brienzo, M. García-Aparicio, R. Agudelo and J. Görgens, J. Chem. Technol. Biotechnol., 2015, 90, 449–458 CrossRef CAS.
- F. Gutmann, C. Jann, F. Pereira, A. Johansson, L. M. Steinmetz and K. R. Patil, Biotechnol. Biofuels, 2021, 14, 41 CrossRef CAS PubMed.
- Y. Jia, Q. Zhang, J. Dai, X. Zheng, X. Meng, R. Zhou, H. Yang, L. Yao and X. Chen, BioResources, 2023, 18, 228 CAS.
- J.-M. Xia and Y.-J. Yuan, J. Agric. Food Chem., 2009, 57, 99–108 CrossRef CAS PubMed.
- J. Yang, M.-Z. Ding, B.-Z. Li, Z. L. Liu, X. Wang and Y.-J. Yuan, OMICS, 2012, 16, 374–386 CrossRef CAS PubMed.
- R. Gao, Z. Li, X. Zhou, W. Bao, S. Cheng and L. Zheng, Biotechnol. Biofuels, 2020, 13, 3 CrossRef CAS PubMed.
- S. Morales-Palomo, C. González-Fernández and E. Tomás-Pejó, J. Environ. Chem. Eng., 2022, 10, 107354 CrossRef CAS.
- A. Dobrowolski, K. Drzymala, D. A. Rzechonek, P. Mitula and A. M. Mironczuk, Front. Microbiol., 2019, 10, 547 CrossRef PubMed.
- S. Zhuang, K. Smart and C. Powell, J. Am. Soc. Brew. Chem., 2017, 75, 244–254 CrossRef CAS.
- L. Ferraz, M. Sauer, M. J. Sousa and P. Branduardi, Front. Microbiol., 2021, 12, 715891 CrossRef PubMed.
- D. Wang, M. Zhang, J. Huang, R. Zhou, Y. Jin and C. Wu, J. Microbiol. Biotechnol., 2019, 30, 62 CrossRef PubMed.
- S. Li, L. Rong, S. Wang, S. Liu, Z. Lu, L. Miao, B. Zhao, C. Zhang, D. Xiao, K. Pushpanathan, A. Wong and A. Yu, Chem. Eng. Sci., 2022, 249, 117342 CrossRef CAS.
- W. C. Vong and S. Q. Liu, J. Sci. Food Agric., 2017, 97, 135–143 CrossRef CAS PubMed.
- K. Wierzchowska, B. Zieniuk and A. Fabiszewska, Waste Biomass Valorization, 2022, 13, 757–779 CrossRef CAS.
- Z. Lazar, N. Liu and G. Stephanopoulos, Trends Biotechnol., 2018, 36, 1157–1170 CrossRef CAS PubMed.
- D. Crespo-Roche, D. González-García, J. A. Méndez-Líter, R. Ledesma-Amaro, A. Prieto and J. Barriuso, bioRxiv, 2025, 12, 695487 Search PubMed.
- S. Bai, B. Qiao, Z. J. Hou, G. R. Gao, C. Y. Cao, J. S. Cheng and Y. J. Yuan, Chemosphere, 2023, 310, 136864 CrossRef CAS PubMed.
- H. R. Su, B. Zhang, R. C. Yang, J. Y. Shi, S. C. He, S. F. Dai, D. W. Wu and Z. Y. Zhao, Recycling, 2026, 11, 8 CrossRef.
- S. D. Yook, J. Kim, G. Gong, J. K. Ko, Y. Um, S. O. Han and S. M. Lee, GCB Bioenergy, 2020, 12, 670–679 CrossRef CAS.
- Y. K. Park, PhD thesis, Université Paris-Saclay, 2020.
- C. Naveira-Pazos, R. Robles-Iglesias, C. Fernández-Blanco, M. C. Veiga and C. Kennes, Rev. Environ. Sci. Biotechnol., 2023, 22, 1131–1158 CrossRef CAS.
- K. U. Chande, S. Pharande and R. Karodi, Biopress J. Comput. Life Sci., 2025, 1, 25–36 Search PubMed.
- H. Abdi, A. Jahanian and H. Nozari, in Dynamic and Safe Economy in the Age of Smart Technologies, IGI Global Scientific Publishing, 2025, pp. 105–120 Search PubMed.
Footnote |
| † These authors contributed equally to this manuscript. |
| This journal is © The Royal Society of Chemistry 2026 |