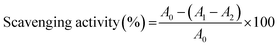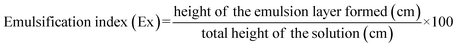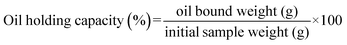Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Exopolysaccharides and conjugated linoleic acids produced by selected lactic acid bacteria: physicochemical attributes, antimicrobial activity, and techno-functional properties of exopolysaccharides
Md. Sadman Tahmid *,
Md. Abunaser,
A. K. M. Masum,
Mohammad Ashiqul Islam
*,
Md. Abunaser,
A. K. M. Masum,
Mohammad Ashiqul Islam and
Md. Harun-ur-Rashid
and
Md. Harun-ur-Rashid *
*
Department of Dairy Science, Bangladesh Agricultural University, Mymensingh-2202, Bangladesh. E-mail: sadmantahmid33@gmail.com; harunds@bau.edu.bd
First published on 9th March 2026
Abstract
Lactic acid bacteria (LAB) exopolysaccharides (EPS) are high-value biopolymers that have predominant applications in the food sector owing to their thickening, gelling, stabilizing, texturizing, emulsifying, and flocculating properties, while conjugated linoleic acids (CLAs) have potential health benefits for humans. This study aims to screen and isolate bacterial strains based on their EPS and CLA production as well as to characterize the carbohydrate and protein content, water and oil holding capacity, emulsification, antimicrobial activity, and hydroxyl radical scavenging activity of EPS. In this study, 50 out of 300 bacterial strains stocked in the laboratory were screened using agar media, and 18 strains were selected based on their EPS production. Afterward, the CLA production and EPS techno-functional properties of the 10 bacterial strains with the highest EPS-producing capacity (Lactobacillus acidophilus, Enterococcus faecium, Lactiplantibacillus plantarum, Streptococcus thermophilus, Lactococcus lactis, Lacticaseibacillus casei, Lactobacillus delbrueckii subsp. bulgaricus, Pediococcus pentosaceus, Lactobacillus paraplantarum, and Lactobacillus delbrueckii subspp. lactis) were evaluated. The EPS production varied significantly, from 39.02 mg L−1 in Lactococcus raffinolactis B425 to 2210.18 mg L−1 in Lactobacillus acidophilus LDMB01. For CLA production, Lactobacillus del. spp. bulgaricus M240 exhibited the highest yield (30.60 µg mL−1). In terms of techno-functional properties, Enterococcus faecium D325 had the highest carbohydrate content (749.89 µg mL−1) while Lactobacillus delbrueckii subsp. bulgaricus M240 exhibited the highest protein content (194.87 µg mL−1). Lacticaseibacillus casei LDMB03 demonstrated the highest water-holding (204.28%) and oil-holding (309.59%) capacity. The 10% EPS exhibited the highest antimicrobial activity against the tested pathogenic bacteria, even at relatively low concentrations. Pediococcus pentosaceus B225 had the strongest emulsification index while the highest hydroxyl radical scavenging activity (94.40%) was recorded for Lactococcus lactis LDMB10. These findings highlight the diverse functional properties of LAB-derived EPS, underscoring their potential for the development of functional foods, pharmaceuticals, and cosmetic products.
Sustainability spotlightThis work promotes sustainable food technology through the use of lactic acid bacteria (LAB) to produce bioactive exopolysaccharides (EPS) and conjugated linoleic acids (CLAs) via natural fermentation. These microbe-derived compounds offer eco-friendly alternatives to synthetic additives, enhancing food functionality, safety, and sustainability. |
1 Introduction
Lactic acid bacteria (LAB) are generally recognized as safe (GRAS), facultatively anaerobic or microaerophilic, non-motile, Gram-positive, catalase-negative, and non-spore-forming bacterial strains that produce organic acids, bacteriocins, and exopolysaccharides during their metabolic activities.1 LAB metabolic activities vary significantly across genera and even strains within the same species owing to their genetic makeup and enzyme repertoires.2 LABs are capable of producing microbial exopolysaccharides (EPS), which are non toxic antioxidants and high molecular weight biodegradable natural carbohydrate polymers. In addition to bacteria, yeasts, fungi, algae, and both eukaryotic and prokaryotic organisms with diverse biological activities have the potential to secrete EPS.3–5 On the basis of its monosaccharide composition, EPS can be classified into homopolysaccharides and heteropolysaccharides.6 According to Dueholm et al.,7 EPS is mostly produced intracellularly, and different pathways are involved in its secretion. It is synthesized via any of the following four biosynthesis pathways: (1). the ATP-binding cassette (ABC) transporter-dependent, (2). Wzx/Wzy-dependent, (3). extracellular, and (4). synthase-dependent pathways.8–10 The biosynthetic pathway involved in EPS production by LAB significantly influences the functional and structural properties of EPS. Structural (chemical and biomolecular) modifications of EPS can improve its yield and also enhance the desired physicochemical and biological activities.6,10,11The molecules of Leuconostoc mesenteroides, Limosilactobacillus fermentum, Limosilactobacillus reuteri, and Streptococcus salivarius produce alternan, dextran, reuteran, and mutan types of EPS via extracellular biosynthetic pathways; however, the properties of these compounds are significantly different. Similarly, the properties of commercial EPS, including that produced by Xanthomonas campestris, Lactobacillus delbrueckii subsp. bulgaricus, Lactobacillus acidophilus, and Lactobacillus plantarum, are distinct, though these bacteria use the Wzx/Wzy-dependent pathway. Stingele et al.12 mentioned that the ST11 and YS4 strains of Str. thermophilus is non-ropy while the Str. thermophilus Sfi6 strain is ropy. Differences exist even between Str. thermophilus Sfi6 and Str. thermophilus Sfi39, and the variation in their gene level is evident from the studies by Stingele et al.12 and Germond et al.13 In addition to the bacterial strain, exopolysaccharide production, structural characterization, biosynthesis, biological properties, physicochemical properties, and effective applications depend on the composition of the fermentation medium, including carbon, nitrogen, salts, and the condition of the culture, such as pH, time, and temperature.14,15 The Lactobacillus helveticus MB2-1 strain, under its optimum growing conditions, produced 340 mg L−1 of EPS at an optimal temperature of 37 °C.16 In contrast, under similar conditions (25% (w/w) dipotassium hydrogen phosphate, 30% (w/w) ethanol, 2% (w/w) 1-butyl-3-methylimidazolium octyl sulfate, and a pH of 10), the EPS yields of Lactobacillus acidophilus and Limosilactobacillus reuteri were found to be 63.30 µg mL−1 and 146.48 µg mL−1, respectively.17
The industrial EPS production is costly, with 40% of expenses slated for microbial growth substrates.18 The extraction of EPS from LAB with distinctive bioactivities is becoming progressively attractive, especially for industrial applications. They are increasingly used in agriculture, medicine, cosmetics, and food because of their beneficial properties, such as stability, solubility, viscosity, elasticity, water holding capacity, oil holding capacity, emulsification property, biocompatibility, gelation, and thickening agent.15,19 During fermentation, the produced EPS enhances texture and flavor and provides health benefits.14 Streptococcus thermophilus EPS is valuable in dairy, improving texture stability, handling tolerance, and melting property while reducing chewiness and hardness in low-fat Mozzarella cheese.20 EPS-producing probiotics significantly enhance food safety, quality, shelf life, and functionality. This is because EPS enhances the rheological properties and water-holding capacity of food matrices, improves emulsion and foam stability, facilitates the formation of protective biofilms that suppress spoilage and pathogenic microorganisms, and functions as prebiotic and immunomodulatory compounds that confer additional health benefits.19,21 LAB EPS regulates gut health, thereby influencing microbiome composition and short-chain fatty acid production, which impacts intestinal function and energy metabolism.22 EPS also offers biological effects, including anti-cholesterol, antibacterial, anticancer, antidiabetic, anti-inflammatory, antimicrobial, antiviral, antioxidant, hypoglycemic, immunomodulatory, and anti-Helicobacter pylori activities.23–25 Lactobacillus plantarum KX041 EPS favors molecular mechanisms against inflammation and obesity26 and Lactiplantibacillus plantarum LCC-605 increases the stability of fermented yoghurt, as well as antioxidant and cholesterol-reducing abilities.27
Like EPS, conjugated linoleic acid (CLA) has received significant attention as a health-promoting compound that prevents atherosclerosis, regulates gut microbiota and immunity, reduces fat or lipid deposition, promotes bone formation, and exerts anti-cardiovascular, antioxidant, anti-obesity, anti-diabetic, and anti-carcinogenic effects.28–31 According to Wu et al.,32 CLA can enhance insulin signalling, fatty acid oxidation, and glucose transport in adipose tissue or the liver. Moreover, EPS-producing LAB have the ability to convert free linoleic acid (LA) into conjugated linoleic acid.33,34 Among LAB, it has been found that many food-grade LAB strains, including Lactobacillus lactis, Lactobacillus casei, Lactobacillus acidophilus and Lactiplantibacillus plantarum can produce CLA in yogurt, cheese, and milk products.34
Given the variability among LAB strains and their diverse metabolic end products, it was hypothesized that the LAB selected for this study would exhibit different potentials for EPS production with distinct physicochemical and techno-functional properties of the produced EPS. Similarly, the selected LAB strains were expected to differ in their CLA production capabilities, which was considered an additional health and well-being-promoting feature of EPS-producing LAB. Therefore, the aim of the study was to screen EPS-producing LAB strains from the selected LAB lot, evaluate the produced EPS, and assess the CLA production potential of the selected EPS-LAB. The set objectives were to quantify the EPS and CLA produced by the screened LAB strains, followed by monitoring, recording, and assessing the physicochemical and techno-functional properties of the produced EPS, viz., total carbohydrate and protein content, hydroxyl radical scavenging activity, antimicrobial activity, emulsification index, oil-holding capacity, water-holding capacity, and solidification ability of EPS. It is anticipated that the outcome of this study will provide valuable insights into the techno-functional properties of EPS produced by bacterial strains and their potential applications in the food industry, especially with regard to product development and the well-being of consumers.
2 Materials and methods
2.1. Selection of bacteria and processing
Bacterial strains were obtained from the Laboratory of Dairy Microbiology and Biotechnology (LDMB), which comprised 266 freeze-dried lactic acid bacteria (LAB) that were isolated from traditionally fermented Dahi in Bangladesh35,36 and 34 LAB strains isolated from raw goat milk and eventually characterized.37 From the 300 LAB strains, 50 LAB strains were chosen. Stock cultures were made in MRS broth (de Man, Rogosa and Sharpe; HiMedia, India), followed by subculturing and eventual storage at −20 °C for further use.2.2. Screening of EPS-producing strains
The ability of the isolates to produce EPS was assessed using the string test, Aniline Blue (AB) agar assay, and Congo Red (CR) agar assay.38 For the string test, the strains were cultured on Luria Bertani (LB) agar (HiMedia, India) plates supplemented with 5% (w/v) sucrose at 37 °C for 48 h. A colony was lifted using an inoculation loop, and those forming a ≥5 mm string were selected. In the AB agar assay, nutrient agar (HiMedia, India) was supplemented with 5% (w/v) sucrose and 0.005% (w/v) aniline blue (SRL, India). EPS-producing isolates showed slightly blue colonies on the AB agar plate. Brain-heart infusion agar (HiMedia, India) was supplemented with 5% (w/v) sucrose and added to 0.08% (w/v) Congo red (HiMedia, India) to make the CR agar. Light to dark black colonies indicate potential EPS-producing strains. Both the AB and CR plates were incubated at 37 °C for ∼48 h. Strains that passed at least any one of the aforementioned assays (a total of 18 strains passed) were considered for subsequent steps.2.3. Production, extraction, and quantification of EPS
The screened EPS-producing isolates were incubated for 72 h at 37 °C in MRS broth supplemented with 5% (w/v) sucrose on a constant orbital shaker (SCI Finetech, Korea) at 120 rpm. During this period, optical density (OD)600 was measured every 6 h using an i9 Hanon UV-VIS double beam spectrophotometer at 600 nm.38,39 After incubation, the samples (50 mL) were heated in a water bath (Daihan Scientific Bath, Korea) at ∼100 °C for 50 min and then cooled to 25 °C. They were then centrifuged at 5500×g (Neya 16R, Remi, India) for 20 min at 4 °C, and the supernatants were collected. Ice-cold ethanol was added to the supernatants (2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1), and the mixtures were incubated overnight at −20 °C. Then, crude EPS precipitation was collected after centrifuging the supernatant–ethanol mixtures at 5500×g for 20 min at 4 °C. Eventually, they were dried overnight at 40 °C (BOD incubators, MRC Lab, Holon, Israel). The dry EPS weight and yield were recorded and calculated, respectively. Crude EPS was purified by redissolving in distilled water, dialyzing with changing water, and lyophilizing.
1), and the mixtures were incubated overnight at −20 °C. Then, crude EPS precipitation was collected after centrifuging the supernatant–ethanol mixtures at 5500×g for 20 min at 4 °C. Eventually, they were dried overnight at 40 °C (BOD incubators, MRC Lab, Holon, Israel). The dry EPS weight and yield were recorded and calculated, respectively. Crude EPS was purified by redissolving in distilled water, dialyzing with changing water, and lyophilizing.
2.4. Functional properties of EPS
To estimate the total protein content, the Bradford micro-assay protocol was used. The reaction solution was made of 0.5 mL of EPS solution and 0.5 mL of the Bradford protein assay reagent (Bio-Rad Quick Start™, USA). After incubation for 5 min at room temperature, absorbance was recorded at 595 nm. Distilled water and bovine serum albumin (BSA) were used as the blank and standard (0–100 µg mL−1; 20 µg mL−1 intervals), respectively.
where A1 = absorbance of the EPS sample solution, A0 = absorbance of the blank, and A2 = absorbance of the control (distilled water was used instead of H2O2).
An optical microscope was used to analyze the particle-size distribution of the emulsions that were formed, as previously described by Kowsalya et al.41 Briefly, 60 µL of each emulsion was placed on a cavity slide and left to stand undisturbed for 5 min. The distribution of oil droplet sizes was observed under a light microscope at 10× magnification.
2.5. Milk coagulation test of EPS strains
The strain Lb. acidophilus LDMB01, with the highest EPS production, was introduced into a sterile skim milk solution (w/v; 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10). The inoculum used was 1% (v/v), and different amounts of sucrose (0%, 3%, 6%, 9%, or 12% w/v) were added. A static culture was created for 24, 36, and 48 h at 30 °C. Subsequently, the morphology of the skim milk was observed at 12 h intervals starting from 24 h of incubation. A negative control was prepared using skim milk containing 0% sugar.5
10). The inoculum used was 1% (v/v), and different amounts of sucrose (0%, 3%, 6%, 9%, or 12% w/v) were added. A static culture was created for 24, 36, and 48 h at 30 °C. Subsequently, the morphology of the skim milk was observed at 12 h intervals starting from 24 h of incubation. A negative control was prepared using skim milk containing 0% sugar.5
2.6. Physiochemical properties of selected EPS-LAB strains
For the oxidase test, the organisms were grown on a glucose agar plate (1% glucose), which was incubated at 37 °C for 24 h. 1% tetramethyl-p-phenylenediamine dihydrochloride was dissolved in distilled water. A few drops of this solution were placed on filter paper (Whatman filter paper, No.1), and a fresh colony was transferred onto it using a cotton bud. A color change within 10–30 seconds indicated a positive result; no change means negative. For the catalase test, a sterile loop was used to transfer a 24 hours culture onto a microscopic slide, and one drop of 30% H2O2 was added to it. Immediate bubble formation indicated a positive result, showing oxygen release.44
For pectin hydrolysis, a medium was prepared using the method of Ortega-Villar et al., with a few modifications.45 At first, a 100 mM phosphate buffer (pH = 7.0) was prepared with 0.4% (w/v) KH2PO4, 0.6% (w/v) NaH2PO4, and Na2HPO4·7H2O, and 0.5% pectin, 0.2% (NH4)2SO4, 0.2% yeast extract, 0.2% MgSO4·7H2O, 0.0001% CaCl2, and 0.05% NaCl were added. This was followed by pH adjustment (to 7.0), inoculation of organisms, and incubation at 37 °C for 72 h. A halo around the colony was interpreted as positive for pectin hydrolysis.
2.7. Screening of LAB for conjugated linoleic acid production
The bacterial strains with high EPS-producing capacity were assessed for conjugated linoleic acid (CLA) production using a spectrophotometric detection method, following a study by Barrett et al.,47 with a few modifications. Briefly, the isolates were activated by successive sub-culturing in MRS broth and grown at 37 °C (BOD incubators, MRC Lab, Holon, Israel). Afterward, the strains were incubated in MRS broth containing free linoleic acid (LA; 0.5 mg mL−1; Sigma-Aldrich L1376-5G) and 2% (w/v) Tween 80 (Merck, Germany). After incubation (at 37 °C for 48 h), 1 mL of the culture was centrifuged at 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800×g for 1 min. The pellet was discarded, and the supernatant was mixed with 2 mL of isopropanol (Merck, Germany) by vortexing and allowed to stand for 3 min. Then, 1.5 mL of hexane (Merck, Germany) was added and vortexed further, and the mixture was allowed to stand for another 3 minutes. The CLA in the culture supernatant was assayed by dispensing 230 µL of the fat-soluble hexane layer into a UV-transparent 96-well plate (Multiskan SkyHigh Spectrophotometer, Thermo Scientific, USA), and absorbance was recorded at 233 nm. The concentration was quantified using a standard curve of conjugated linoleic acid (CLA; Sigma-Aldrich, 05507), consisting of a mixture of cis- and trans-isomers of 9,11- and 10,12-octadecadienoic acids.
800×g for 1 min. The pellet was discarded, and the supernatant was mixed with 2 mL of isopropanol (Merck, Germany) by vortexing and allowed to stand for 3 min. Then, 1.5 mL of hexane (Merck, Germany) was added and vortexed further, and the mixture was allowed to stand for another 3 minutes. The CLA in the culture supernatant was assayed by dispensing 230 µL of the fat-soluble hexane layer into a UV-transparent 96-well plate (Multiskan SkyHigh Spectrophotometer, Thermo Scientific, USA), and absorbance was recorded at 233 nm. The concentration was quantified using a standard curve of conjugated linoleic acid (CLA; Sigma-Aldrich, 05507), consisting of a mixture of cis- and trans-isomers of 9,11- and 10,12-octadecadienoic acids.
2.8. Statistical analysis
All experimental data were tested in triplicate, and the values are expressed as mean ± standard deviation. Statistical analysis was performed using IBM SPSS statistical software (V27.0). A one-way analysis of variance (ANOVA) was performed to test significance, and Duncan's multiple range test (DMRT) was carried out to rank the results while significant differences were observed. A probability of less than 0.05 was considered statistically significant.3 Results and discussion
3.1. EPS screening and production
Lactic acid bacteria genera, such as Lactobacillus, Lactococcus, Leuconostoc, Pediococcus, Weissella, and Enterococcus, can naturally synthesize diverse forms of glycans or polysaccharides. LAB account for the largest EPS-producing bacteria.21 The preliminary screening in this study was based on their potential for EPS production. Despite numerous strains yielding positive results in the string test, the inconsistency of the ropy features rendered the assay only presumptive. Consequently, the CR and AB agar plate procedures were used for confirmation, leading to the selection of 50 strains. The EPS-producing strains exhibited mucoid or ropy colonies on LB agar media that were used for the string test, light to dark black colonies on CR agar, and an intense blue color appearance on AB agar (Table 1). The formation of a string longer than 5 mm was regarded as an initial indication of EPS production (Fig. 1). After the preliminary screening phase, EPS underwent a secondary screening phase called quantification. Eighteen strains, namely Lactobacillus acidophilus LDMB01, Enterococcus faecium D325, Lactiplantibacillus plantarum LDMB05, Streptococcus thermophilus M540-5, Lactococcus lactis LDMB10, Lacticaseibacillus casei LDMB03, Lactobacillus delbrueckii subsp. bulgaricus M240-5, Pediococcus pentosaceus B225, Lactobacillus paraplantarum LDMB11, and Lactobacillus delbrueckii subsp. lactis D340, Streptococcus bovis biotype I D125-4, Leuconostoc mesenteroides subsp.mesenteroides D625-3, Lactobacillus agilis D640-4, Lactococcus raffinolactis B425-4, Lactococcus lactis subsp. lactis D425-4, Streptococcus bovis biotype II J225-1, Leuconostoc mesenteroides subsp. dextranicum M725-1 and Lactobacillus fermentum M240-4, were selected based on their high EPS production.| Isolates | String test | Aniline blue (AB) assay | Congo red (CR) assay |
|---|---|---|---|
| a Strong positive: + + +; moderate positive: + +; weak positive: +; and negative: −.b LDMB01 = Lb. acidophilus LDMB01; D325 = Ent. faecium D325; LDMB05 = Lpb. plantarum LDMB05; M540 = Str. thermophilus M540-5; LDMB10 = Lc. lactis LDMB10; LDMB03 = Lb. casei LDMB03; M240 = Lb. delbrueckii subsp. bulgaricus M240; B225 = P. pentosaceus B225; LDMB11 = Lb. paraplantarum LDMB11; D340 = Lb. delbrueckii subsp. lactis D340; D425-4 = Lc. lactis subsp. lactis D425-4; D625-3 = Leu. mesenteroides spp. mesenteroides D625-3; M240-4 = Lb. fermentum M240-4; J225-1 = Str. bovis biotype II J225-1; D125-4 = Str. bovis biotype I D125-4; M725-1 = Leu. mesenteroides subsp. dextranicum M725-1; D640-4 = Lb. agilis D640-4; and B425-4 = Lc. raffinolactis B425-4. | |||
| Str. bovis biotype I D125-4 | — | — | + |
| Ent. faecium D325-2 | + + | — | + + |
| Lb. delbrueckii subsp. lactis D340-2 | — | — | + |
| P. pentosaceus B225-2 | + + | + | + + + |
| Leu. mesenteroides subsp. mesenteroides D625-3 | — | — | + |
| Lb. agilis D640-4 | — | + | — |
| Lc. raffinolactis B425-4 | + | + | + + |
| Lc. lactis subsp. lactis D425-4 | + | + + + | + |
| Lb. fermentum M240-4 | + + + | — | — |
| Lb. delbrueckii subsp. bulgaricus M240-5 | + + + | + + + | + + + |
| Str. thermophilus M540-5 | — | — | + |
| Str. bovis biotype II J225-1 | + + + | — | + |
| Lc. lactis LDMB10 | — | — | + + + |
| Lb. casei subsp. casei LDMB03 | + + | + + | + + + |
| Lpb plantarum LDMB05 | + + + | — | + + + |
| Lb. paraplantarum LDMB11 | — | + | + + + |
| Lb. acidophilus LDMB01 | + + + | + + + | + + |
| Leu. mesenteroides subsp. dextranicum M725-1 | + + + | + + + | + + |
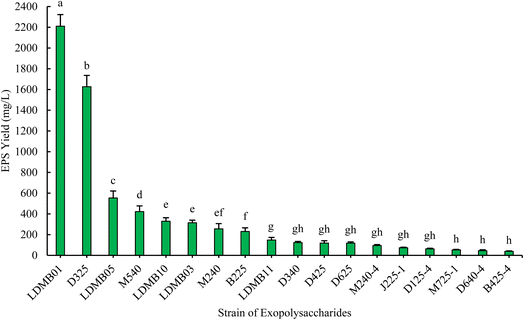
The EPS production of the 18 selected strains varied from 39.02 ± 4.25 mg L−1 to 2210.18 ± 112.14 mg L−1, where the LDMB01 strain showed a maximum EPS-production capacity (Fig. 2). Liu et al.48 reported that Lb. acidophilus produced around 2920 mg EPS L−1, while in the present study, Lb. acidophilus LDMB01 generated 2210.18 mg L−1. The possible cause of this variation may be attributed to the variation in the bacterial strain and/or carbon sources used (glucose or lactose and sucrose, respectively). Liu et al.48 used Lb. acidophilus ATCC mutan, which was obtained from the Microbiology Institute of Chinese Academy (Beijing, China), but we used Lb. acidophilus LDMB01 isolated from raw goat milk. In another study, Hernández-Rosas et al.49 stated that the carbon source is one of the predisposing factors in EPS synthesis by LAB. This was also evident in a report by Padmanabhan et al.50 who found that S. thermophilus ASCC 1275 produced approximately 430 mg L−1 of EPS when grown in a medium supplemented with sucrose, which was higher than the EPS yields with glucose or lactose. Afreen et al.51 used the Plackett–Burman design in their study and identified lactose, yeast extract, tryptone, and CaCl2 as factors for higher EPS production. D325-2 produced the second-highest EPS (1626.51 mg L−1), followed by LDMB05 (553.80 mg L−1), M540-5 (422.34 mg L−1), and LDMB10 (329.48 mg L−1). D425 and LDMB03 produced 118.53 and 314.13 mg L−1 of EPS, respectively. These findings are also supported by Prete et al.,52 who reported that Lc. lactis spp. and Lac. casei demonstrated EPS production levels ranging from 30 to 600 mg L−1. Moreover, EPS production levels of 94.76 and 56.48 mg L−1 were observed in Lb. fermentum M240-4 and S. bovis biotype II J225-1, respectively.
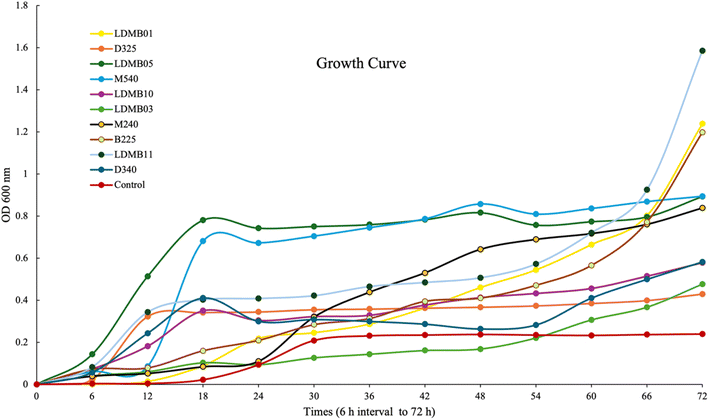
The growth curves of selected EPS strains are presented in Fig. 3. Starting from 18 h up to 66 h, the growth performances of LDMB05 and M540 were higher than those of the other strains. However, after 66 h, the LDMB11, LDMB01, and B225 strains exhibited higher growth. From 60 h onward, all selected strains had higher growth compared to the control, which could have been due to the carbon source in the growth media and orbital shaking during incubation. According to Bates et al.,53 orbital shaking exerts effects on microbial growth through impacts on the temperature and aeration control, the two major variables in microbial growth kinetics. All strains were found to show their maximum growth potential at 72 h. EPS production ability depends on various critical elements affecting bacterial growth, such as relative environment, incubation time, temperature, pH, oxygen rate, optimized culture conditions, and carbon and nitrogen sources, with the quantity of EPS produced being predominantly strain-specific.54 So, orbital shaking may enhance nutrient distribution, create a homogeneous environment, prevent biofilm formation, and enhance growth rates by promoting optimal cell proliferation and metabolic activity.
3.2. Biochemical properties of selective strains
It is worth noting that the isolates were selected based on their stock strain identity, observing the standard biochemical and morphological characteristics and their fermentation with the 14 sugars (Table 2). All the strains were Gram-positive, non-motile, oxidase, catalase, and pectin hydrolysis test-negative. Proteolytic activity was negative in M540. Similar to our findings, positive proteolytic activity had been reported previously by Dong et al.55 in Lb. plantarum, Lb. delbrueckii subsp. bulgaricus, Lc. Lactis, Lb. casei, Lb. acidophilus, P. pentosaceus, and Lb. rhamnosus strains. All the screened EPS-producing LABs were able to utilize glucose, dextrose, galactose, maltose and fructose, except M240. None of the LABs were able to utilize sorbitol, rhamnose, and starch, even LDMB01, the highest EPS producer. Afreen et al.51 screened Lactiplantibacillus paraplantarum NCCP 962 as a high EPS producer from the doi samples and reported their ability to utilize D-galactose, D-lactose, D-sucrose, D-mannose, D-maltose, D-fructose, and D-arabinose, but it was not able to utilize D-sorbitol, D-ribose, and D-xylose as carbon sources.| Parameters | D340 | M240 | LDMB11 | LDMB05 | LDMB10 | LDMB03 | B225 | M540 | D325 | LDMB01 |
|---|---|---|---|---|---|---|---|---|---|---|
| a Positive (+); negative (−); variable (±); and weak positive (+w).b LDMB01 = Lb. acidophilus LDMB01; D325 = Ent. faecium D325; LDMB05 = Lpb. plantarum LDMB05; M540 = Str. thermophilus M540; LDMB10 = Lc. lactis LDMB10; LDMB03 = Lb. casei LDMB03; M240 = Lb. delbrueckii subsp. bulgaricus M240; B225 = P. pentosaceus B225; LDMB11 = Lb. paraplantarum LDMB11; and D340 = Lb. delbrueckii subsp. lactis D340. | ||||||||||
| Gram staining | + | + | + | + | + | + | + | + | + | + |
| Shape | Rod | Rod | Rod | Rod | Cocci | Rod | Cocci | Cocci | Cocci | Rod |
| Motility test | — | — | — | — | — | — | — | — | — | — |
| Glucose agar test | + | + | + | + | + | + | + | + | + | + |
| Proteolytic activity | ± | + | +w | +w | + | ± | +w | — | + | + |
| Lipolytic activity | ± | +w | +w | +w | +w | ± | — | — | — | — |
| Methyl red test | — | — | ± | + | — | — | — | — | ± | +w |
| Oxidase test | — | — | — | — | — | — | — | — | — | — |
| Catalase test | — | — | — | — | — | — | — | — | — | — |
| Pectin hydrolysis | — | — | — | — | — | — | — | — | — | — |
| Fermentation | ||||||||||
| Cellobiose | — | — | + | + | + | + | + | — | + | + |
| Dextrose | + | + | + | + | + | + | + | + | + | + |
| Fructose | + | + | + | + | + | + | + | + | + | + |
| Galactose | + | — | + | + | + | + | + | + | + | + |
| Glucose | + | + | + | + | + | + | + | + | + | + |
| Lactose | + | + | — | + | + | + | — | + | + | + |
| Maltose | + | — | + | + | + | + | + | + | + | + |
| Raffinose | — | — | + | + | — | + | — | — | ± | — |
| Rhamnose | — | — | — | — | — | — | — | — | — | — |
| Sorbitol | — | — | — | — | — | — | — | — | — | — |
| Starch | — | — | — | — | — | — | — | — | — | — |
| Sucrose | + | — | + | + | — | + | — | + | + | + |
| Trehalose | — | — | — | + | +w | — | + | — | + | + |
| Xylose | — | — | + | — | +w | + | ± | — | + | + |
3.3. Functional properties of EPS
The scavenging efficacy of EPS revealed concentration-dependent hydroxyl radicals ranging from 0.5 mg mL−1 to 2 mg mL−1, as shown in Fig. 5. Increasing the EPS concentration from 0.5 to 2.0 mg mL−1 increased the hydroxyl radical-scavenging ability. This effect is probably due to the hydroxyl groups in the EPS donating more active hydrogen atoms.54 However, from 1.5 to 2.0 mg mL−1, no remarkable changes were observed for all the selected strains. For instance, EPS LDMB10 exhibited maximum scavenging activity (90.04 ± 3.70%) at 1.5 mg mL−1, which was 91.21 ± 1.31% at 2.0 mg mL−1. Other studies also confirm the potential antioxidant or scavenging ability of EPS, similar to Lb. rhamnosus, Lcb. paracasei ssp. paracasei, Lb. delbrueckii subsp. bulgaricus, Lb. helveticus, Lpb. plantarum, Lb. acidophilus, and Xanthomonas campestris.54,65 However, among the tested EPS, B225 had the lowest scavenging activity of 75.61% at a 2.0 mg mL−1 concentration. Overall, the findings indicate a concentration-dependent increase in hydroxyl radical-scavenging activity across all the EPS tested, with LDMB10, M240, and LDMB01 EPS showing consistently high activity, suggesting that the EPS from these strains possess potential antioxidant properties. However, Limisolactobacillus fermentum NCDC400 EPS showed only 63.92% scavenging of hydroxyl groups at a 3 mg mL−1 concentration,66 and L. plantarum R301 EPS can remove 96.52% hydroxyl groups at a 2 mg mL−1 concentration.67 However, in another study, Tarannum et al.38 reported scavenging activities of 90.49% and 88.30% at concentrations of 0.755 and 0.738 µg mL−1 for Lacticaseibacillus sp. ME17 and Lactococcus sp. ME7, respectively. All these findings indicate this efficacy as a multifactorial phenomenon. In the current study, Lb. lactis LDMB10, Lb. delbrueckii spp. bulgaricus M240, Lb. acidophilus LDMB01, Lpb. plantarum LDMB05, S. thermophilus M540-4 showed potentials for radical scavenging activity and revealed their suitability as food additives with naturally produced antioxidants.
| Gram-positive bacteria | Gram-negative bacteria | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| L. monocytogenes | S. aureus | B. cereus | E. coli | S. typhi | K. pneumoniae | |||||||
| EPS | Zone of inhibition (mm) in 0.5 mg mL−1 | Zone of inhibition (mm) in 1.0 mg mL−1 | Zone of inhibition (mm) in 0.5 mg mL−1 | Zone of inhibition (mm) in 1.0 mg mL−1 | Zone of inhibition (mm) in 0.5 mg mL−1 | Zone of inhibition (mm) in 1.0 mg mL−1 | Zone of inhibition (mm) in 0.5 mg mL−1 | Zone of inhibition (mm) in 1.0 mg mL−1 | Zone of inhibition (mm) in 0.5 mg mL−1 | Zone of inhibition (mm) in 1.0 mg mL−1 | Zone of inhibition (mm) in 0.5 mg mL−1 | Zone of inhibition (mm) in 1.0 mg mL−1 |
| a Strong inhibition (+++) = >15 mm; moderate inhibition (++) = 10–15 mm; weak inhibition (+) = 5–10 mm; and no inhibition (−) = <5 mm.b LDMB01 = Lb. acidophilus LDMB01; D325 = Ent. faecium D325; LDMB05 = Lpb. plantarum LDMB05; M540 = Str. thermophilus M540-5; LDMB10 = Lc. lactis LDMB10; LDMB03 = Lb. casei LDMB03; M240 = Lb. delbrueckii subsp. bulgaricus M240; B225 = P. pentosaceus B225; LDMB11 = Lb. paraplantarum LDMB11; and D340 = Lb. delbrueckii subsp. lactis D340. | ||||||||||||
| LDMB01 | +++ | +++ | ++ | +++ | + | + | — | — | ++ | +++ | ++ | +++ |
| D325 | + | ++ | +++ | +++ | — | — | + | ++ | + | ++ | — | — |
| LDMB05 | ++ | ++ | ++ | ++ | + | ++ | — | + | ++ | ++ | + | ++ |
| M540 | ++ | ++ | ++ | +++ | + | ++ | + | ++ | + | + | ++ | +++ |
| LDMB10 | +++ | +++ | ++ | +++ | + | + | — | — | +++ | +++ | + | + |
| LDMB03 | ++ | ++ | + | + | + | + | ++ | +++ | +++ | +++ | — | — |
| M240 | ++ | +++ | ++ | ++ | ++ | +++ | + | ++ | ++ | +++ | ++ | ++ |
| B225 | + | + | +++ | +++ | + | + | ++ | +++ | + | ++ | — | — |
| LDMB11 | ++ | ++ | ++ | ++ | + | +++ | — | — | ++ | +++ | + | ++ |
| D340 | ++ | +++ | ++ | ++ | ++ | ++ | ++ | ++ | + | ++ | ++ | +++ |
In addition, the present findings indicate that the antimicrobial activity of LAB EPS is mostly strain-specific. For instance, Fig. 6 shows that EPS D340 showed the highest inhibitory effect against L. monocytogenes ATCC 7644, and LDMB01 exhibited broad-spectrum activity, showing notable inhibition zones against both S. aureus ATCC 12600 and K. pneumoniae ATCC 8047. Similarly, EPS M240 displayed superior efficacy against B. cereus ATCC 11778, while EPS B225 was found to be most effective against E. coli ATCC 25922. In addition, EPS LDMB03 showed the strongest antimicrobial activity against S. typhi ATCC 14028. EPS chelates metals, forms barriers, restricts essential nutrients, disrupts cell walls and cytoplasmic membranes, inhibits mRNA and protein synthesis, disrupts cell proliferation, inactivates enzymes, generates H2O2, and accelerates DNA destruction.57,69,71,72 Evidence of the structural conformation and/or presence and arrangements of different functional groups in EPS from the present study could have been advantageous in explaining the mechanism of their antibacterial activity, which was not covered. The use of antimicrobial compounds added directly to food or its packaging is one practical approach, indicating the industrial significance of EPS.
| Olive oil | E24% (Mean ± SD) | E48% (Mean ± SD) | E96% (Mean ± SD) |
|---|---|---|---|
| a Means of different superscripts significantly differ within row and column (p < 0.05).b LDMB01 = Lb. acidophilus LDMB01; D325 = Ent. faecium D325; LDMB05 = Lpb. plantarum LDMB05; M540 = Str. thermophilus M540-5; LDMB10 = Lc. lactis LDMB10; LDMB03 = Lb. casei LDMB03; M240 = Lb. delbrueckii subsp. bulgaricus M240; B225 = P. pentosaceus B225; LDMB11 = Lb. paraplantarum LDMB11; and D340 = Lb. delbrueckii subsp. lactis D340. | |||
| LDMB01 | 69.77bc ± 6.63 | 65.00de ± 5.76 | 64.50de ± 4.84 |
| D325 | 60.93ef ± 0.93 | 52.00ij ± 1.90 | 46.50l ± 1.70 |
| LDMB05 | 61.87e ± 3.70 | 54.00ghi ± 1.85 | 52.50ghij ± 0.89 |
| M540 | 60.20ef ± 5.17 | 57.00 fg ± 0.98 | 49.00jkl ± 2.11 |
| LDMB10 | 47.50 kl ± 1.44 | 39.00m ± 1.31 | 36.20m ± 4.39 |
| LDMB03 | 63.57de ± 0.57 | 63.57de ± 1.01 | 63.53de ± 0.59 |
| M240 | 63.53de ± 1.63 | 64.00de ± 1.14 | 64.30de ± 1.45 |
| B225 | 84.17a ± 1.99 | 72.90b ± 1.04 | 68.23 cd ± 1.78 |
| LDMB11 | 67.03 cd ± 1.59 | 64.00de ± 0.96 | 63.53de ± 0.97 |
| D340 | 56.50fgh ± 2.10 | 51.00ijkl ± 2.36 | 48.50jkl ± 2.01 |
| Soybean oil | E24% (Mean ± SD) | E48% (Mean ± SD) | E96% (Mean ± SD) |
|---|---|---|---|
| a Means of different superscripts significantly differ within row and column (p < 0.05).b LDMB01 = Lb. acidophilus LDMB01; D325 = Ent. faecium D325; LDMB05 = Lpb. plantarum LDMB05; M540 = Str. thermophilus M540-5; LDMB10 = Lc. lactis LDMB10; LDMB03 = Lb. casei LDMB03; M240 = Lb. delbrueckii subsp. bulgaricus M240; B225 = P. pentosaceus B225; LDMB11 = Lb. paraplantarum LDMB11; and D340 = Lb. delbrueckii subsp. lactis D340. | |||
| LDMB01 | 59.07de ± 2.68 | 50.00klmn ± 2.59 | 44.50q ± 1.50 |
| D325 | 57.23efg ± 1.40 | 49.00lmno ± 1.44 | 46.00opq ± 1.35 |
| LDMB05 | 61.47 cd ± 2.40 | 57.00efg ± 1.35 | 50.00klmn ± 2.07 |
| M540 | 58.53de ± 2.44 | 54.50ghi ± 1.80 | 48.00mnop ± 1.13 |
| LDMB10 | 53.00hijk ± 1.67 | 45.50pq ± 1.90 | 45.00pq ± 1.35 |
| LDMB03 | 62.47c ± 1.52 | 53.50hij ± 2.41 | 47.17nopq ± 1.85 |
| M240 | 58.50de ± 2.36 | 54.93fgh ± 1.19 | 50.50klmn ± 1.41 |
| B225 | 70.83a ± 2.24 | 66.43b ± 1.92 | 57.30efg ± 0.89 |
| LDMB11 | 63.50bc ± 0.66 | 54.47ghi ± 0.93 | 51.00jklm ± 2.21 |
| D340 | 58.00ef ± 1.37 | 51.50ijkl ± 1.50 | 40.00r ± 3.12 |
| Mustard oil | E24% (Mean ± SD) | E48% (Mean ± SD) | E96% (Mean ± SD) |
|---|---|---|---|
| a Means of different superscripts significantly differ within row and column (p < 0.05).b LDMB01 = Lb. acidophilus LDMB01; D325 = Ent. faecium D325; LDMB05 = Lpb. plantarum LDMB05; M540 = Str. thermophilus M540; LDMB10 = Lc. lactis LDMB10; LDMB03 = Lb. casei LDMB03; M240 = Lb. delbrueckii subsp. bulgaricus M240; B225 = P. pentosaceus B225; LDMB11 = Lb. paraplantarum LDMB11; and D340 = Lb. delbrueckii subsp. lactis D340. | |||
| LDMB01 | 57.30 cd ± 2.67 | 53.00ef ± 1.83 | 50.00fghi ± 1.64 |
| D325 | 52.53efg ± 0.92 | 46.50jk ± 1.35 | 41.50lm ± 1.91 |
| LDMB05 | 66.83b ± 2.40 | 59.00c ± 1.15 | 54.50de ± 3.40 |
| M540 | 54.10e ± 2.13 | 49.53ghij ± 1.59 | 44.50 kl ± 1.67 |
| LDMB10 | 47.00ijk ± 1.39 | 39.50mn ± 1.41 | 38.50n ± 1.95 |
| LDMB03 | 59.57c ± 1.76 | 58.50c ± 1.37 | 58.63c ± 1.40 |
| M240 | 54.53de ± 2.40 | 57.50 cd ± 1.65 | 58.90c ± 0.95 |
| B225 | 57.47 cd ± 0.70 | 50.57fgh ± 1.12 | 41.77lm ± 1.44 |
| LDMB11 | 70.00 ± 1.71 | 50.50fgh ± 1.57 | 49.00hij ± 1.44 |
| D340 | 54.00e ± 1.67 | 47.00ijk ± 1.35 | 42.50lm ± 1.37 |
| Black cumin oil | E24% (Mean ± SD) | E48% (Mean ± SD) | E96% (Mean ± SD) |
|---|---|---|---|
| a Means of different superscripts significantly differ within row and column (p < 0.05).b LDMB01 = Lb. acidophilus LDMB01; D325 = Ent. faecium D325; LDMB05 = Lpb. plantarum LDMB05; M540 = Str. thermophilus M540-5; LDMB10 = Lc. lactis LDMB10; LDMB03 = Lb. casei LDMB03; M240 = Lb. delbrueckii subsp. bulgaricus M240; B225 = P. pentosaceus B225; LDMB11 = Lb. paraplantarum LDMB11; and D340 = Lb. delbrueckii subsp. lactis D340. | |||
| LDMB01 | 69.07 cd ± 2.72 | 64.00e ± 2.09 | 64.00e ± 1.11 |
| D325 | 59.13f ± 2.70 | 58.00 fg ± 1.51 | 57.47fgh ± 1.59 |
| LDMB05 | 80.30b ± 2.70 | 84.00a ± 1.44 | 84.50a ± 1.37 |
| M540 | 59.50f ± 2.55 | 53.50i ± 1.73 | 50.50j ± 2.62 |
| LDMB10 | 55.50ghi ± 0.79 | 54.50hi ± 1.57 | 50.00j ± 2.26 |
| LDMB03 | 68.40 cd ± 1.47 | 70.60 cd ± 1.44 | 71.50c ± 1.47 |
| M240 | 67.53d ± 0.68 | 68.20d ± 1.74 | 69.50 cd ± 1.57 |
| B225 | 57.00fgh ± 0.96 | 59.60f ± 2.36 | 60.07f ± 1.40 |
| LDMB11 | 79.00b ± 1.41 | 81.00b ± 0.70 | 80.50b ± 0.70 |
| D340 | 59.00f ± 1.25 | 58.50 fg ± 1.70 | 48.00j ± 1.14 |
| Almond oil | E24% (Mean ± SD) | E48% (Mean ± SD) | E96% (Mean ± SD) |
|---|---|---|---|
| a Means of different superscripts significantly differ within row and column (p < 0.05).b LDMB01 = Lb. acidophilus LDMB01; D325 = Ent. faecium D325; LDMB05 = Lpb. plantarum LDMB05; M540 = Str. thermophilus M540-5; LDMB10 = Lc. lactis LDMB10; LDMB03 = Lb. casei LDMB03; M240 = Lb. delbrueckii subsp. bulgaricus M240; B225 = P. pentosaceus B225; LDMB11 = Lb. paraplantarum LDMB11; and D340 = Lb. delbrueckii subsp. lactis D340. | |||
| LDMB01 | 57.17bcde ± 2.75 | 48.00 kl ± 2.55 | 41.00o ± 2.78 |
| D325 | 54.57efg ± 1.32 | 51.03hijk ± 1.12 | 51.00hijk ± 1.28 |
| LDMB05 | 63.83a ± 3.48 | 58.50bc ± 1.47 | 53.50fgh ± 1.44 |
| M540 | 50.67hijkl ± 2.80 | 47.50lm ± 1.14 | 44.50mn ± 1.01 |
| LDMB10 | 50.00ijkl ± 1.11 | 47.50lm ± 0.44 | 43.00no ± 1.93 |
| LDMB03 | 58.00bcd ± 2.02 | 55.57cdef ± 1.48 | 49.70jkl ± 1.15 |
| M240 | 57.10bcde ± 2.02 | 53.10fghi ± 1.76 | 47.40lm ± 1.56 |
| B225 | 63.43a ± 1.62 | 54.97defg ± 1.12 | 49.43jkl ± 0.71 |
| LDMB11 | 65.50a ± 0.46 | 59.50b ± 0.66 | 52.00ghij ± 2.96 |
| D340 | 56.00cdef ± 1.25 | 54.00efgh ± 1.11 | 49.50jkl ± 1.80 |
| n-Hexane | E24% (Mean ± SD) | E48% (Mean ± SD) | E96% (Mean ± SD) |
|---|---|---|---|
| a Means of different superscripts significantly differ within row and column (p < 0.05).b LDMB01 = Lb. acidophilus LDMB01; D325 = Ent. faecium D325; LDMB05 = Lpb. plantarum LDMB05; M540 = Str. thermophilus M540-5; LDMB10 = Lc. lactis LDMB10; LDMB03 = Lb. casei LDMB03; M240 = Lb. delbrueckii subsp. bulgaricus M240; B225 = P. pentosaceus B225; LDMB11 = Lb. paraplantarum LDMB11; and D340 = Lb. delbrueckii subsp. lactis D340. | |||
| LDMB01 | 45.43bc ± 5.75 | 32.10ef ± 3.01 | 26.37ij ± 3.47 |
| D325 | 49.60ef ± 4.11 | 39.00ij ± 0.95 | 33.00lm ± 2.07 |
| LDMB05 | 63.80a ± 2.60 | 54.50 cd ± 2.36 | 50.00ef ± 1.28 |
| M540 | 57.63bc ± 2.06 | 56.50bc ± 0.85 | 47.50 fg ± 1.11 |
| LDMB10 | 39.47ij ± 1.59 | 37.00jk ± 2.25 | 30.00mn ± 2.25 |
| LDMB03 | 58.03bc ± 2.90 | 56.70bc ± 1.71 | 55.10c ± 0.82 |
| M240 | 51.47de ± 2.56 | 48.27ef ± 0.70 | 44.57gh ± 2.10 |
| B225 | 42.53hi ± 1.66 | 34.93 kl ± 2.90 | 27.60n ± 1.11 |
| LDMB11 | 60.00b ± 1.91 | 56.00c ± 1.97 | 51.00def ± 1.85 |
| D340 | 42.50hi ± 1.30 | 38.47jk ± 2.55 | 31.83lm ± 1.84 |
| Kerosene | E24% (Mean ± SD) | E48% (Mean ± SD) | E96% (Mean ± SD) |
|---|---|---|---|
| a Means of different superscripts significantly differ within row and column (p < 0.05).b LDMB01 = Lb. acidophilus LDMB01; D325 = Ent. faecium D325; LDMB05 = Lpb. plantarum LDMB05; M540 = Str. thermophilus M540-5; LDMB10 = Lc. lactis LDMB10; LDMB03 = Lb. casei LDMB03; M240 = Lb. delbrueckii subsp. bulgaricus M240; B225 = P. pentosaceus B225; LDMB11 = Lb. paraplantarum LDMB11; and D340 = Lb. delbrueckii subsp. lactis D340. | |||
| LDMB01 | 59.60a ± 1.49 | 45.00gh ± 1.67 | 39.00j ± 1.51 |
| D325 | 48.63ef ± 5.36 | 38.00jk ± 1.51 | 35.00 kl ± 1.01 |
| LDMB05 | 56.60 ab ± 2.50 | 49.50ef ± 1.57 | 49.43ef ± 1.66 |
| M540 | 33.00l ± 3.66 | 28.50m ± 2.44 | 20.50n ± 1.67 |
| LDMB10 | 54.00bcd ± 1.25 | 51.60de ± 0.92 | 43.90hi ± 0.95 |
| LDMB03 | 55.50bc ± 3.54 | 52.43cde ± 2.60 | 50.17ef ± 1.35 |
| M240 | 48.90ef ± 1.41 | 41.00ij ± 1.06 | 38.00jk ± 0.87 |
| B225 | 50.10ef ± 1.10 | 39.23j ± 1.91 | 26.40m ± 1.65 |
| LDMB11 | 57.50 ab ± 2.56 | 43.50hi ± 1.41 | 40.50ij ± 1.08 |
| D340 | 47.50 fg ± 1.37 | 44.00hi ± 0.92 | 39.50j ± 1.37 |
| EPS | Olive oil | Soybean oil | Mustard oil | Black cumin oil | Almond oil | n-Hexane | Kerosene |
|---|---|---|---|---|---|---|---|
| a Values with different lowercase letters within a column indicate significant differences (p < 0.05).b LDMB01 = Lb. acidophilus LDMB01; D325 = Ent. faecium D325; LDMB05 = Lpb. plantarum LDMB05; M540 = Str. thermophilus M540-5; LDMB10 = Lc. lactis LDMB10; LDMB03 = Lb. casei LDMB03; M240 = Lb. delbrueckii subsp. bulgaricus M240; B225 = P. pentosaceus B225; LDMB11 = Lb. paraplantarum LDMB11; and D340 = Lb. delbrueckii subsp. lactis D340. | |||||||
| LDMB01 | 66.42b | 51.19e | 53.43c | 65.69e | 48.72e | 48.73c | 47.87c |
| D325 | 53.14ef | 50.74e | 46.84f | 58.20f | 52.20d | 40.53d | 40.54e |
| LDMB05 | 56.12d | 56.16bc | 60.11a | 82.93a | 58.61a | 56.10a | 51.84a |
| M540 | 55.40de | 53.68d | 49.38de | 54.50gh | 47.56ef | 53.88b | 27.33g |
| LDMB10 | 41.61g | 47.83f | 41.67g | 53.33h | 46.83f | 35.49f | 49.83b |
| LDMB03 | 63.56c | 54.38cd | 58.90a | 70.17c | 54.42bc | 56.61a | 52.70a |
| M240 | 63.94bc | 54.64bcd | 56.98b | 68.41d | 52.53d | 48.10c | 42.63d |
| B225 | 75.10a | 64.86a | 49.93d | 58.89f | 55.94b | 35.02f | 38.58f |
| LDMB11 | 64.86bc | 56.32b | 56.50b | 80.17b | 59.00a | 55.67ab | 47.17c |
| D340 | 52.00f | 49.83e | 47.83ef | 55.17g | 53.17cd | 37.60e | 43.67d |
| p Value | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 |
| Time | Olive oil | Soybean oil | Mustard oil | Black cumin oil | Almond oil | n-hexane | Kerosene |
|---|---|---|---|---|---|---|---|
| a Values with different lowercase letters within a column indicate significant differences (p < 0.05).b E24 = Emulsification index at 24 h; E48 = Emulsification index at 48 h; and E96 = Emulsification index at 96 h. | |||||||
| E24 | 63.51a | 60.26a | 57.33a | 65.44a | 57.62a | 52.32a | 51.13a |
| E48 | 58.24b | 53.68b | 51.16b | 65.19a | 52.97b | 46.98b | 43.28b |
| E96 | 55.89c | 47.94c | 47.98c | 63.60b | 48.10c | 41.01c | 38.24c |
| p Value | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 |
The gross particle size of the emulsions (P. pentosaceus B225 EPS in different oils) is depicted in Fig. 7. This experiment revealed that the particle size of emulsified droplets plays a critical role in determining the physical stability, including the creaming rate and flocculation, of different emulsions. Larger and scattered droplets in the emulsion indicate low efficiency of stability formation (Fig. 7C and D). On the contrary, the particle distribution showed mostly smaller, densely packed, and uniform droplets that provided a higher surface area for the EPS to adsorb, resulting in it working as an effective stabilizer and resisting coalescence and sedimentation. This positive trend was also observed by Balyan et al.81 in their work on isolating Lactobacillus EPS as an emulsifier to ensure the extended stability of eugenol encapsulation.
3.4. Coagulation effect of EPS strain in skim milk
The coagulation/solidification effect of the superior EPS strain Lb. acidophilus LDMB01 with different levels of sugar (sucrose) in skim milk is depicted in Fig. 9. After 24 h of fermentation, all the samples remained liquid, and even at 36 h, the sample with 0% sucrose exhibited a liquid appearance, while the samples with 3%, 6%, 9%, and 12% sucrose exhibited an initiation of curd formation. However, after 48 h, the sample with 0% sucrose remained unchanged, while other samples exhibited semi-solid and solid structures. Notably, the sample with 12% sucrose had a more pronounced solidification than all other samples and exhibited a complete curdling characteristic. The outcome of the present study is similar to the findings of Ning et al.5 for Gluconobacter frateurii HDC-08 EPS, Yu et al.92 for Lactiplantibacillus plantarum HDC-01, and Afreen et al.51 for Lactiplantibacillus paraplantarum NCCP 962 EPS, where higher sucrose concentrations and longer fermentation periods resulted in more pronounced curd formation. The coagulation effect is intricately linked to the structure and molecular weight of EPS, the kind of protein present in the milk, as well as the interaction between the EPS and skim milk protein.93 The findings indicated that the LAB might be capable of synthesizing EPS from sucrose, which promotes milk solidification properties in dairy products.3.5. Conjugated linoleic acid production
Food-derived LAB strains are considered to be the most efficient conjugated linoleic acid (CLA) producers. This study selected the LAB strains with the highest EPS-producing capacity for CLA production. The standard curve exhibited a strong linear relationship (R2 = 0.9949). The linear correlation between the peak area and concentration was strong within the concentration range of 0 to 100 µg mL−1. The CLA concentration across the strains varied, with values ranging from 9.56 ± 1.24 µg mL−1 to 30.60 ± 1.27 µg mL−1 when supplied with 0.5 mg mL−1 LA. Among all the strains tested (Fig. 10), the microorganism M240 exhibited the highest CLA conversion/production, reaching 30.60 µg mL−1, followed closely by LDMR01 and LDMB11 with CLA yields of 29.72 and 28.46 µg mL−1, respectively. Wei et al.94 obtained a close result for the Lpb. plantarum Lp-01 strain, which achieved 33.47 µg mL−1 of CLA when cultured in a medium supplemented with pine nut oil, indicating its capacity to convert linoleic acid present in the oil into CLA. The Lb. casei LDMB03 strain exhibited 23.12 µg mL−1 of CLA; however, in another study, using an initial linoleic acid concentration of 0.1% and incubating at 37 °C for 72 h, soymilk fermented by Lactobacillus casei synthesized approximately 839 µg g−1 of CLA.95 Lactobacillus reuteri was reported to have 0.108 mg mL−1 of CLA when incubated under aerobic conditions at 10 °C for 30 h in a broth medium with 20 mg L−1 free LA. It was observed that L. reuteri cells could grow 5.5 times faster CLA isomers (c9, t11 and t10, c12) than free-washed cells.31 Jena and Choudhury96 found that the yogurt culture of Lb. del. subsp. bulgaricus LB340 and Bifidobacterium animalis subsp. lactis BLO4, BB12, B94, and HN019 co-cultured with the S. thermophilus TA040 strain predominantly enhanced CLA content in yogurt during the manufacture. Baliyan et al.97 observed variable CLA production by EPS-producing probiotic LABs isolated from raw cow milk. Chen et al.28 also reported 1.28 to 54.28 µg mL−1 CLA production by different LABs.The variation in CLA production highlights strain-specific capabilities, which could be exhibited by the distinct metabolic pathways and enzymatic activities within each LAB species. These findings suggest that certain LAB EPS strains, particularly Lb. delbrueckii subsp. bulgaricus M240, Lb. acidophilus LDMB01, and Lb. paraplantarum LDMB11 could be promising for applications in CLA-enriched probiotic formulation. Therefore, EPS and CLA were evaluated in this study because they represent two of the most functionally significant metabolites produced by lactic acid bacteria with direct applications in dairy foods. In a study by Amiri et al.,98 Lactobacillus acidophilus LA5 and Bifidobacterium animalis subsp. lactis BB12 were shown to co-produce two metabolites of EPS and CLA under the same culture conditions. This co-production suggests a potential for selecting strains that can simultaneously deliver both texture stability through EPS and health benefits via CLA to fermented products.
4 Conclusions
This study provides comprehensive information about the screening, production and quantification of exopolysaccharides from selected lactic acid bacterial strains, as well as the functional properties of exopolysaccharides and the conjugated linoleic acid producing property of strains with high exopolysaccharide-production capacity. The highest exopolysaccharide production was observed in Lactobacillus acidophilus LDMB01 EPS, and the lowest was found in Lactococcus raffinolactis B425. Lactobacillus delbrueckii subsp. bulgaricus M240 produced the highest amount of conjugated linoleic acid. For the functional properties, the exopolysaccharide obtained from Lacticaseibacillus casei LDMB03 exhibited the highest water- and oil-holding capacity. Owing to the milk coagulation ability of the exopolysaccharide-producing strains, they have potential application in fermented dairy products. Increasing the EPS concentration enhanced antimicrobial activity; however, approximately 11.67% of the tested samples exhibited no antimicrobial effect even at higher concentrations. Accumulated antimicrobial activity, hydroxy radical scavenging activity, emulsification index, water-holding capacity, oil-holding capacity, together with the solidifying efficiency of exopolysaccharides, imply numerous EPS applications in the food industry. Regarding exopolysaccharides and conjugated linoleic acid production, as well as the functional properties of exopolysaccharides, Lactobacillus delbrueckii subsp. bulgaricus M240, Lactiplantibacillus plantarum LDMB05, and Lactobacillus acidophilus LDMB01 can be recommended as suitable lactic acid bacterial strains. Furthermore, producing exopolysaccharides in functional food manufacturing can be highly interesting because of their role not only as health-promoting agents but also as a key factor in enhancing the quality of food products. The future scope of this study should include spectroscopic analyses like Fourier transform infrared (FTIR) spectroscopy, nuclear magnetic resonance (NMR), and advanced microscopy techniques like scanning electron microscopy (SEM), which can be used to examine the produced exopolysaccharides with more precise characteristics important for industrial applications.Author contributions
Md. Sadman Tahmid: conceptualization, data curation, formal analysis, investigation, methodology, software, supervision, validation, visualization, writing – original draft, and writing – review and editing. Md. Abunaser: conceptualization and writing – review and editing. A. K. M. Masum: conceptualization, investigation, supervision, validation, visualization, and writing – review and editing. Mohammad Ashiqul Islam: conceptualization, formal analysis, resources, validation, and writing – review and editing. Md. Harun-ur-Rashid: conceptualization, funding acquisition, project administration, resources, supervision, validation, and writing – review and editing.Conflicts of interest
The authors declare no conflicts of interest.Data availability
The data are available from the corresponding author upon reasonable request.Acknowledgements
This work was supported by the Ministry of Education, GoB (MoE, Project ID: LS-20232873, ref. 2023/15/MoE). The authors acknowledge the Laboratory of Animal Science and Biotechnology, BAU, for the support in providing access to the facility for the 96-well plate spectrophotometer analysis.References
- J. F. Ceron-Cordoba, L. C. Muñoz-Domínguez, J. U. S. Valencia and D. A. Restrepo-Molina, J. Dairy Sci., 2025, 108(4), 3151–3161 CrossRef CAS PubMed.
- N. S. Jaffar, R. Jawan and K. P. Chong, Front. Plant Sci., 2023, 13, 1047945 CrossRef PubMed.
- A. I. Netrusov, E. V. Liyaskina, I. V. Kurgaeva, A. U. Liyaskina, G. Yang and V. V. Revin, Microorganisms, 2023, 11, 1541 CrossRef CAS PubMed.
- F. Salimi and P. Farrokh, World J. Microbiol. Biotechnol., 2023, 39, 213 CrossRef CAS PubMed.
- Y. Ning, H. Cao, S. Zhao, D. Gao and D. Zhao, Polymers, 2024, 16, 1004 CrossRef CAS PubMed.
- Z. Q. Xiong, L.-H. Kong, P. F.-H. Lai, Y.-J. Xia, J.-C. Liu, Q.-Y. Li and L.-Z. Ai, J. Dairy Sci., 2019, 102, 4925–4934 CrossRef CAS PubMed.
- M. K. D. Dueholm, M. Besteman, E. J. Zeuner, M. Riisgaard-Jensen, M. E. Nielsen, S. Z. Vestergaard, S. Heidelbach, N. S. Bekker and P. H. Nielsen, Water Res., 2023, 229, 119485 CrossRef CAS PubMed.
- N. Kaur and P. Dey, Res. Microbiol., 2023, 174, 104024 CrossRef CAS PubMed.
- A. Ponzio, A. Rebecchi, R. Zivoli and L. Morelli, Foods, 2024, 13, 752 CrossRef CAS PubMed.
- J. Wu, X. Han, M. Ye, Y. Li, X. Wang and Q. Zhong, Crit. Rev. Food Sci. Nutr., 2023, 63, 7043–7064 CrossRef CAS PubMed.
- K. Zhang, S. Liu, S. Liang, F. Xiang, X. Wang, H. Lian, B. Li and F. Liu, Int. J. Biol. Macromol., 2024, 257, 128733 CrossRef CAS PubMed.
- F. Stingele, J. R. Neeser and B. Mollet, J. Bacteriol., 1996, 178, 1680–1690 CrossRef CAS PubMed.
- J.-E. Germond, M. Delley, N. D'Amico and S. J. F. Vincent, Eur. J. Biochem., 2001, 268, 5149–5156 CrossRef CAS PubMed.
- D. Kavitake, P. B. Devi, C. Delattre, G. B. Reddy and P. H. Shetty, Int. J. Biol. Macromol., 2023, 226, 111–120 CrossRef CAS PubMed.
- M. Xiao, L. Tang, X. Fu, X. Ren, J. Bi, J. Wang, D. Li, Q. Kong, H. Mou and C. Zhu, Food Hydrocolloids, 2024, 151, 109850 CrossRef CAS.
- Z. Ge, D. Wang, F. Azi, W. Zhao, P. Wang, M. Dong, J. Wang, Y. Zhao and X. Zhao, Int. Dairy J., 2024, 155, 105969 CrossRef CAS.
- A. S. Raj, G. Anup, R. Sivalingam and P. Chandran, Biorem. J., 2024, 28, 491–511 CrossRef CAS.
- B. G. Guerrero, K. D. L. Santos, E. S. Kamimura and C. A. F. De Oliveira, Int. J. Food Sci. Technol., 2024, 59, 17–29 CrossRef CAS.
- N. S. Kiran, C. Yashaswini, S. Singh and B. G. Prajapati, 3 Biotech, 2024, 14, 95 CrossRef PubMed.
- P. Li, Y. Bai, Y. Yang and M. Li, Food Biosci., 2023, 55, 102961 CrossRef CAS.
- H. Pourjafar, F. Ansari, A. Sadeghi, S. A. Samakkhah and S. M. Jafari, Crit. Rev. Food Sci. Nutr., 2023, 63, 8194–8225 CrossRef CAS PubMed.
- M. Tarique, A. H. Ali, J. Kizhakkayil, S.-Q. Liu, F. Oz, E. Dertli, A. Kamal-Eldin and M. Ayyash, Food Chem.:X, 2024, 21, 101073 CAS.
- A. A. Al-Nabulsi, Z. W. Jaradat, F. R. Al Qudsi, L. Elsalem, T. M. Osaili, A. N. Olaimat, G. Esposito, S.-Q. Liu and M. M. Ayyash, LWT-Food Sci. Technol., 2022, 167, 113817 CrossRef CAS.
- M. Ayyash, C. Stathopoulos, B. Abu-Jdayil, G. Esposito, M. Baig, M. S. Turner, A. S. Baba, V. Apostolopoulos, A. Al-Nabulsi and T. Osaili, LWT-Food Sci. Technol., 2020, 131, 109741 CrossRef CAS.
- M. Xiao, X. Ren, Y. Yu, W. Gao, C. Zhu, H. Sun, Q. Kong, X. Fu and H. Mou, Food Chem.:X, 2022, 13, 100233 CAS.
- F. Yue, H. Han, J. Xu, X. Yao, Y. Qin, L. Zhang, X. Sun, J. Huang, F. Zhang and X. Lü, Int. J. Biol. Macromol., 2025, 289, 138803 CrossRef CAS PubMed.
- Y. Yang, X. Shi, J. Zhang, H. Xiao and C. Li, Food Chem., 2025, 465, 142068 CrossRef CAS PubMed.
- C. Chen, F. Tong, R. Sun, Y. Zhang, Z. Pang and X. Liu, Foods, 2024, 13, 1830 CrossRef CAS PubMed.
- M. Du, M. Gong, G. Wu, J. Jin, X. Wang and Q. Jin, J. Agric. Food Chem., 2024, 72, 5503–5525 CrossRef CAS PubMed.
- M. Iorizzo, C. Di Martino, F. Letizia, T. W. Crawford and G. Paventi, Foods, 2024, 13, 975 CrossRef CAS PubMed.
- S. S. Li, L. Xu, J. Qing, X. Wu, H. Li, H. Chen and X. Liu, Food Chem., 2023, 409, 135257 CrossRef CAS PubMed.
- L. Wu, S. Ye, X. Deng, Z. Fu, J. Li and C. Yang, Nutrients, 2024, 16, 1133 CrossRef CAS PubMed.
- R. Chourasia, M. M. Abedin, L. Chiring Phukon, D. Sahoo, S. P. Singh and A. K. Rai, Compr. Rev. Food Sci. Food Saf., 2021, 20, 960–979 CrossRef CAS PubMed.
- M. M. Abedin, R. Chourasia, L. C. Phukon, P. Sarkar, R. C. Ray, S. P. Singh and A. K. Rai, Crit. Rev. Food Sci. Nutr., 2024, 64, 10730–10748 CrossRef CAS PubMed.
- M. Rashid, K. Togo, M. Ueda and T. Miyamoto, Pakistan J. Nutr., 2007, 6, 647–652 CrossRef.
- M. Rashid, K. Togo, M. Ueda and T. Miyamoto, World J. Microbiol. Biotechnol., 2007, 23, 125–133 CrossRef.
- Md. Z. Islam, Md. E. Uddin, Md. T. Rahman, M. A. Islam and Md. Harun-ur-Rashid, Small Rumin. Res., 2021, 205, 106532 CrossRef.
- N. Tarannum, T. J. Hossain, F. Ali, T. Das, K. Dhar and I. H. Nafiz, LWT-Food Sci. Technol., 2023, 186, 115263 CrossRef CAS.
- F. Yang-Chen, J. Chen, S. Ye, Z. Liu and Y. Ding, Process Biochem., 2022, 113, 87–96 CrossRef.
- W. Xia, J. Han, S. Zhu, Y. Wang, W. Zhang and Z. Wu, Int. J. Biol. Macromol., 2023, 230, 123177 CrossRef CAS PubMed.
- M. Kowsalya, T. Velmurugan, R. Mythili, W. Kim, K. G. Sudha, S. Ali, B. Kalpana, S. Ramalingam and M. Prasanna Rajeshkumar, Int. J. Biol. Macromol., 2023, 242, 124842 CrossRef CAS PubMed.
- S. A. Qamar, M. Asgher and M. Bilal, Waste Biomass Valor., 2021, 12, 6847–6859 CrossRef CAS.
- L. C. Silva, G. B. Schmidt, L. G. O. Alves, V. S. Oliveira, R. Laureano-Melo, E. Stutz, J. F. P. Martins, B. P. Paula, R. H. Luchese, A. F. Guerra and P. Rodrigues, Food Bioprod. Process., 2020, 124, 408–418 CrossRef CAS.
- J. Homolak, Free Radical Res., 2022, 56, 343–357 CrossRef CAS PubMed.
- R. Ortega-Villar, A. Escalante, F. Astudillo-Melgar, L. Lizárraga-Mendiola, G. A. Vázquez-Rodríguez, M. E. Hidalgo-Lara and C. Coronel-Olivares, Microorganisms, 2024, 12, 1066 CrossRef CAS PubMed.
- G. Rampanti, D. N. Nedelkoska, T. Kalevska, T. Stojanovska, J. Harasym, F. Cardinali, A. Orkusz, V. Milanović, C. Garofalo, A. Bonifazi, L. Aquilanti and A. Osimani, Heliyon, 2024, 10(17), e37548 CrossRef CAS PubMed.
- E. Barrett, R. P. Ross, G. F. Fitzgerald and C. Stanton, Appl. Environ. Microbiol., 2007, 73, 2333–2337 CrossRef CAS PubMed.
- Q. Liu, X. Huang, D. Yang, T. Si, S. Pan and F. Yang, EXCLI J., 2016, 15, 119–133 Search PubMed.
- F. Hernández-Rosas, J. D. Castilla-Marroquín, J. M. Loeza-Corte, M. A. Lizardi-Jiménez and R. I. Hernández-Martínez, Colegio de Postgraduados, Rev. Mex. Ing. Quim., 2021, 20, 1–21 Search PubMed.
- A. Padmanabhan, Y. Tong, Q. Wu, J. Zhang and N. P. Shah, Front. Microbiol., 2018, 9, 1919 CrossRef PubMed.
- A. Afreen, Z. Ahmed, N. Khalid, I. Ferheen and I. Ahmed, Appl. Microbiol. Biotechnol., 2023, 107, 1189–1204 CrossRef CAS PubMed.
- R. Prete, M. K. Alam, G. Perpetuini, C. Perla, P. Pittia and A. Corsetti, Foods, 2021, 10, 1653 CrossRef CAS PubMed.
- M. K. Bates, D. S. Phillips and J. O'Bryan, Thermo Fisher Scientific, ANSHORBGRT 0916, 2016.
- R. Prete, F. Dell'Orco, G. Sabatini, F. Montagano, N. Battista and A. Corsetti, Foods, 2024, 13, 1663 CrossRef CAS PubMed.
- Y. Dong, J. Ronholm, I. Fliss and S. Karboune, Probiotics Antimicrob. Proteins, 2025, 17(5), 3211–3233 CrossRef CAS PubMed.
- M. A. Arayes, M. E. M. Mabrouk, S. A. Sabry and B. Abdella, Biologia, 2023, 78, 229–240 CrossRef CAS.
- M. Srinivash, R. Krishnamoorthi, P. U. Mahalingam and B. Malaikozhundan, Int. J. Biol. Macromol., 2023, 250, 126171 CrossRef CAS PubMed.
- X. Xu, Q. Peng, Y. Zhang, D. Tian, P. Zhang, Y. Huang, L. Ma, Y. Qiao and B. Shi, Food Nutr. Res., 2020, 64 CAS.
- E. S. Khalil, M. Y. Abd Manap, S. Mustafa, A. M. Alhelli and P. Shokryazdan, Molecules, 2018, 23, 398 CrossRef PubMed.
- Q. Q. Li, Y. Chen, X. Liu, Y. Li, J. Xu, T. Li, W. Xiang and A. Li, Front. Mar. Sci., 2023, 9, 1097200 CrossRef.
- X. Yang-Wu, J. Wu, F. An, J. Xu, M. Bat-Ochir, L. Wei, M. Li, M. Bilige and R. Wu, Int. J. Biol. Macromol., 2022, 208, 288–298 CrossRef PubMed.
- Q. Zhong, B. Wei, S. Wang, S. Ke, J. Chen, H. Zhang and H. Wang, Mar. Drugs, 2019, 17, 674 CrossRef CAS PubMed.
- X. Li, Y. Gong, W. Yao, X. Chen, J. Xian, L. You and P. Fardim, Food Chem.:X, 2021, 12, 100157 CAS.
- Q. Wei, G. Fu, K. Wang, Q. Yang, J. Zhao, Y. Wang, K. Ji and S. Song, Pharmaceuticals, 2022, 15, 581 CrossRef CAS PubMed.
- W. Zang, H. Cao, J. Ge and D. Zhao, Int. J. Biol. Macromol., 2024, 263, 130083 CrossRef CAS PubMed.
- M. Kumari, R. Kumari, B. H. Nataraj, P. A. Shelke, S. A. Ali, R. Nagpal and P. V. Behare, Curr. Res. Food Sci., 2023, 6, 100478 CrossRef CAS PubMed.
- J. Wang, J. Zhang, H. Guo, Q. Cheng, Z. Abbas, Y. Tong, T. Yang, Y. Zhou, H. Zhang, X. Wei, D. Si and R. Zhang, Foods, 2023, 12, 2481 CrossRef CAS PubMed.
- A. K. Abdalla, M. M. Ayyash, A. N. Olaimat, T. M. Osaili, A. A. Al-Nabulsi, N. P. Shah and R. Holley, Front. Microbiol., 2021, 12, 664395 CrossRef PubMed.
- P. Rahnama Vosough, M. B. Habibi Najafi, M. R. Edalatian Dovom, A. Javadmanesh and B. Mayo, J. Food Meas. Char., 2021, 15, 5221–5230 CrossRef.
- F. Nehal, M. Sahnoun, S. Smaoui, B. Jaouadi, S. Bejar and S. Mohammed, Microb. Pathog., 2019, 132, 10–19 CrossRef CAS PubMed.
- M. S. R. Rajoka, H. M. Mehwish, H. F. Hayat, N. Hussain, S. Sarwar, H. Aslam, A. Nadeem and J. Shi, Probiotics Antimicrob. Proteins, 2019, 11, 1132–1142 CrossRef CAS PubMed.
- Microbial Exopolysaccharides: Production and Applications, ed. S. K. Bhatia, P. S. Panesar and S. Mehariya, CRC Press, Boca Raton, 2024 Search PubMed.
- L. Sun, Y. Yang, P. Lei, S. Li, H. Xu, R. Wang, Y. Qiu and W. Zhang, Carbohydr. Polym., 2021, 261, 117872 CrossRef CAS PubMed.
- A. Najjari, M. Jabberi, S. F. Chérif, A. Cherif, H. I. Ouzari, J. A. Linares-Pastén and H. Sghaier, Front. Microbiol., 2024, 15, 1440081 CrossRef CAS PubMed.
- L. De Melo Teixeira, É. Da Silva Santos, R. S. Dos Santos, A. V. G. Ramos, D. C. Baldoqui, M. L. Bruschi, J. E. Gonçalves, R. A. C. Gonçalves and A. J. B. De Oliveira, Int. J. Biol. Macromol., 2024, 278, 134400 CrossRef CAS PubMed.
- V. Nemati and R. Mozafarpour, LWT-Food Sci. Technol., 2024, 199, 116116 CrossRef CAS.
- F. Y. Ushikubo and R. L. Cunha, Food Hydrocolloids, 2014, 34, 145–153 CrossRef CAS.
- X. Feng, Y. Sun, H. Tan, L. Ma, H. Dai and Y. Zhang, Food Chem., 2023, 413, 135653 CrossRef CAS PubMed.
- T. Xiao, X. Ma, H. Hu, F. Xiang, X. Zhang, Y. Zheng, H. Dong, B. Adhikari, Q. Wang and A. Shi, Food Chem.:X, 2025, 29, 102792 CAS.
- D. Kavitake, S. Balyan, P. B. Devi and P. H. Shetty, J. Food Sci. Technol., 2020, 57, 1579–1585 CrossRef CAS PubMed.
- S. Balyan, V. Dadwal and B. S. Patil, Food Biosci., 2024, 61, 104632 CrossRef CAS.
- X. Yang-Feng, J. Feng, Q. Zhu, R. Hong and L. Li, Polymers, 2022, 14, 90 CrossRef PubMed.
- Y. Dai, M. Xu, Z. Zhou and Y. Han, Polymers, 2024, 16, 759 CrossRef CAS PubMed.
- J. Zhang, Y. Yao, J. Li, X. Ju and L. Wang, Food Chem., 2023, 425, 136369 CrossRef CAS PubMed.
- C. Li, W. Li, X. Chen, M. Feng, X. Rui, M. Jiang and M. Dong, LWT–Food Sci. Technol., 2014, 57, 477–485 CrossRef CAS.
- L. Wang, Y. Gu and Z. Lv, Int. J. Food Sci. Technol., 2022, 57, 4076–4085 CrossRef CAS.
- Z. Yang, S. Li, X. Zhang, X. Zeng, D. Li, Y. Zhao and J. Zhang, J. Biosci. Bioeng., 2010, 110, 53–57 CrossRef CAS PubMed.
- S. Sarkar, G. Cabrera-Barjas, R. N. Singh, J. P. Fabi, S. J. M. Breig, J. Tapia, R. K. Sani and A. Banerjee, Sci. Rep., 2024, 14, 25058 CrossRef CAS PubMed.
- I. Trabelsi, N. Ktari, M. Triki, I. Bkhairia, S. Ben Slima, S. Sassi Aydi, S. Aydi, A. Abdeslam and R. Ben Salah, Int. J. Biol. Macromol., 2018, 111, 11–18 CrossRef CAS PubMed.
- X. Yang-Ren, Y. Ren and L. Li, LWT-Food Sci. Technol., 2022, 153, 112345 CrossRef.
- N. Srivastava, S. Kumari, S. Kurmi, A. K. Pinnaka and A. R. Choudhury, Arch. Microbiol., 2022, 204, 399 CrossRef CAS PubMed.
- L. Yu, G. Ye, X. Qi, Y. Yang, B. Zhou, Y. Zhang, R. Du, J. Ge and W. Ping, Front. Microbiol., 2023, 14, 1210302 CrossRef PubMed.
- R. Du, L. Yu, N. Yu, W. Ping, G. Song and J. Ge, Int. J. Biol. Macromol., 2022, 217, 303–311 CrossRef CAS PubMed.
- G. Wei, G. Wu, J. Sun, Y. Qi, Q. Zhao, F. Xu, Z. Zhang and Z. Peng, Foods, 2024, 13, 2472 CrossRef CAS PubMed.
- M. Sasi, S. Kumar, M. Hasan, A. S. R., E. Garcia-Gutierrez, S. Kumari, O. Prakash, L. Nain, A. Sachdev and A. Dahuja, Crit. Rev. Food Sci. Nutr., 2023, 63, 9995–10013 CrossRef CAS PubMed.
- R. Jena and P. K. Choudhury, Probiotics Antimicrob. Proteins, 2025, 17(3), 1–22 CrossRef CAS PubMed.
- N. Baliyan, A. K. Maurya, A. Kumar, V. K. Agnihotri and R. Kumar, LWT-Food Sci. Technol., 2023, 176, 114553 CrossRef CAS.
- S. Amiri, M. Rezazadeh Bari, M. Alizadeh Khaledabad, R. Rezaei Mokarram and M. Sowti Khiabani, Chem. Rev. Lett., 2021, 4, 66–76 CAS.
| This journal is © The Royal Society of Chemistry 2026 |