Open Access Article
Open Access ArticleOptimization of probiotic beverage made from Sohiong fruit using response surface methodology and artificial neural network–genetic algorithm hybrid model
Roshiya Nongmaithema,
Raju Sasikumar *a,
Irengbam Barun Mangang
*a,
Irengbam Barun Mangang b,
Selva Kumar T
b,
Selva Kumar T ac,
Kambhampati Vivekd and
Amit K. Jaiswal
ac,
Kambhampati Vivekd and
Amit K. Jaiswal *ef
*ef
aDepartment of Agribusiness Management and Food Technology, North-Eastern Hill University, Tura Campus, Chasingre 794002, Tura, West Garo Hills, Meghalaya, India. E-mail: sashibiofoodster@gmail.com
bCollege of Food Technology, Lamphelpat, Central Agricultural University, Imphal-795004, India
cVel Tech Rangarajan Dr. Sagunthala R&D Institute of Science and Technology, Chennai 600062, India
dDepartment of Food Process Engineering, National Institute of Technology, Rourkela, India
eSchool of Food Science and Environmental Health, Faculty of Sciences and Health, Technological University Dublin, City Campus, Central Quad, Grangegorman, Dublin, D07 ADY7, Ireland. E-mail: amit.jaiswal@tudublin.ie
fCentre for Sustainable Packaging and Bioproducts (CSPB), Technological University Dublin, City Campus, Grangegorman, Dublin D07 H6K8, Ireland
First published on 13th April 2026
Abstract
In this study, a probiotic beverage from Sohiong (Prunus nepalensis), an underutilized wild edible fruit rich in phenolics and anthocyanins, was optimized using both Response Surface Methodology (RSM) and an Artificial Neural Network–Genetic Algorithm (ANN–GA) hybrid model. Plackett–Burman screening identified temperature, strain, and inoculum size as significant variables influencing antioxidant response. Subsequent optimization using a Box–Behnken design yielded maximum DPPH scavenging activity of 71.55%, with enhanced total phenolic content (TPC) of 160.00 mg GAE per g, total anthocyanin content (TAC) of 227.85 mg C3GE per 100 mL, and total flavonoid content (TFC) of 183.96 mg QE per 100 mL. The RSM model showed good predictive capacity (R2 = 0.9467; RMSE = 1.98; AAD = 3.21%), but the ANN–GA model outperformed it with higher accuracy (R2 = 0.9988; RMSE = 0.63; AAD = 1.46%). Validation under ANN–GA-optimized conditions closely matched the predicted values, with only a 0.04% deviation in DPPH. These results confirm the superior predictive fidelity of ANN–GA for nonlinear fermentation systems. The optimized Sohiong probiotic beverage demonstrates significant antioxidant activity and is a promising functional beverage. This study highlights the potential of integrating traditional RSM with modern AI tools like ANN–GA in functional food formulation and bioactive compound enrichment.
Sustainability spotlightThis study presents a sustainable bioprocessing strategy to convert Sohiong (Prunus nepalensis), an underutilised wild fruit rich in phenolics and anthocyanins, into a high-value probiotic-fermented functional beverage. By integrating response surface methodology with artificial neural network–genetic algorithm modelling, the process achieved superior antioxidant activity and bioactive retention while minimising trial-and-error experimentation, thereby reducing material, energy, and experimental resource use. The approach promotes the development of clean-label, functional products using locally available biodiversity and aligns with the principles of the circular bioeconomy. Valorisation of indigenous fruits through precision fermentation contributes to SDG 3 (Good Health and Well-being), SDG 9 (Industry, Innovation and Infrastructure), SDG 12 (Responsible Consumption and Production), and SDG 13 (Climate Action). This work offers a scalable, data-driven model for sustainable, eco-innovative food bioprocessing that supports climate-smart nutrition, rural enterprise, and functional-beverage innovation, advancing sustainable food systems and promoting regional agricultural resilience. |
1 Introduction
Prunus nepalensis, commonly known as Sohiong, represents an indigenous fruit species prevalent to the northeastern Himalayan foothills of India, with significant populations concentrated in Meghalaya state. This fruit exhibits distinctive organoleptic characteristics, including its characteristic dark purple pigmentation and complex flavor profile.1,2The biochemical composition of Sohiong (P. nepalensis) demonstrates remarkable therapeutic potential, containing high concentrations of anthocyanins (∼984 mg C3G per 100 g dry mass), essential micronutrients, and bioactive phytochemicals that contribute to its pronounced antioxidant capacity, as shown by DPPH and FRAP assays.3 These constituents have also been applied in product development, such as yoghurt, syrups, and hard candies, where anthocyanin stability was maintained over 14–90 days.4 However, commercial processing remains severely constrained due to an ephemeral harvesting season (August–November) and rapid post-harvest deterioration (shelf life of only 4–5 days under ambient conditions), limiting its accessibility to broader markets.5
The development of innovative processing technologies, particularly the formulation of probiotic-enriched products such as fermented beverages and dehydrated powders, presents a strategic approach to overcoming these inherent limitations. Such value-added initiatives preserve the functional bioactive components and extend the product's commercial viability, therefore addressing the growing consumer preference for nutraceutical products with demonstrated health benefits. Fermentative bioprocessing utilizing probiotic microorganisms represents a well-established biotechnological approach for enhancing the functional properties of food matrices. The strategic application of beneficial microorganisms, including Lactobacillus species and Bifidobacterium strains, facilitates the development of functional foods with demonstrated clinical efficacy in promoting intestinal health, immunomodulation, and optimizing nutrient bioavailability.6,7
Response surface methodology is a collection of statistical and mathematical methods designed to elucidate relationships among multiple independent variables and their corresponding response parameters,8 establishing it as the predominant optimization approach in fermentation process development.9 Contemporary research has witnessed the emergence of alternative algorithmic approaches for fermentation optimization applications. Advanced extraction methodologies have been refined through integration of response surface analysis with reverse propagation neural network–genetic algorithm frameworks. Dragoi et al.10 developed and validated an enhanced adaptive differential evolution algorithm incorporating neural modelling capable of optimizing aerobic fermentation processes using precise computational modelling.
ANN constitutes sophisticated deep learning methodology capable of modelling intricate nonlinear interdependencies among process variables.9 GA represent computationally effective optimization techniques which emulate biological evolutionary mechanisms, including selection pressure, genetic crossover, and mutation operations. Throughout the preceding decade, the synergistic application of ANN–GA hybrid systems has gained prominence in fermentation process optimization and extraction parameter determination, demonstrating superior efficiency in identifying optimal operational conditions with reduced experimental burden. Representative applications include pectin recovery from sunflower seed matrices, polyphenolic compound extraction from green tea substrates, bioactive compound isolation from pitaya peel waste, and ellagitannin recovery from black raspberry seed materials.11–13 The superior performance of ANN systems in modelling complex nonlinear variable relationships often surpasses conventional fitting methodologies such as traditional RSM approaches.14
Although several studies highlight the effectiveness of statistical and AI-based optimization across various food systems, none have explored Sohiong, an underutilized, anthocyanin-rich wild fruit. For the first time, this research integrates RSM with ANN–GA hybrid modeling to optimize Sohiong probiotic beverage fermentation, thereby demonstrating both the functional potential and the superior predictive accuracy of ANN–GA in fermentation systems. The present investigation establishes the following research objectives: (I) implementation of the Plackett–Burman experimental design to identify statistically significant process variables, (II) development and validation of both ANN–GA and RSM predictive models, (III) comparative evaluation of RSM and ANN–GA methodologies to determine the optimal modelling approach and derive optimal fermentation parameters. The research outcomes will provide essential technical foundation for industrial production scaling while contributing valuable insights for fermentation optimization methodologies in similar bioprocessing applications.
2 Methodology
2.1 Sample preparation
Fresh, fully mature Sohiong fruits displaying deep reddish coloration were harvested from Mairang, Meghalaya, India (coordinates: 25°33′41.94″ N, 91°38′9.67″ E). After thorough washing to remove the dust and impurities, fruits were carefully selected based on their intact surface and soft consistency, excluding any damaged specimens. Seeds were manually extracted, the resulting fruit pulp was collected and homogenized using a Philips mixer (India) to prepare samples for subsequent experimental procedures.The probiotic juice was prepared by inoculating a culture (either Lactobacillus plantarum MCC 2974 or Bifidobacterium BB-12) into the pasteurized juice at a concentration range of 4 × 106 to 8 × 106 CFU mL−1. The mixture was incubated at range of 35 to 37 °C for 24 to 72 hours to allow fermentation, thereby enhancing the probiotic content and flavor. The fermented juice was stored at 4 °C until further use.
2.2 DPPH
DPPH scavenging activity was evaluated by mixing 2 mL of 0.1 mM DPPH solution with 1 mL of probiotic Sohiong extracted using methanol. A spectrophotometer was used to measure the absorbance at 517 nm after the mixture was incubated in the dark for 30 minutes. The percentage of radical scavenging activity was calculated to assess the antioxidant potential of the encapsulated Sohiong juice.152.3 TPC
TPC was determined using the Folin–Ciocalteu technique. Folin–Ciocalteu reagent and sodium carbonate solution were mixed with a methanolic extract of the probiotic juice. The absorbance 765 nm was measured following a 30 minutes incubation period at room temperature. Using a gallic acid calibration curve, the results were represented as milligrams of gallic acid equivalents (GAE per g) of powder, with gallic acid serving as the standard.162.4 TAC
TAC was measured using the pH differential method. The probiotic juice was dissolved in buffers at pH 1.0 and pH 4.5. Absorbance was measured at 520 nm and 700 nm, and the anthocyanin content was calculated using the molar extinction coefficient of cyanidin-3-glucoside. Results were expressed as mg of cyanidin-3-glucoside equivalents per gram of powder, reflecting the color-retaining and antioxidant properties of the encapsulated Sohiong juice.172.5 TFC
TFC was determined through the colorimetric technique as per Singh and Banu,18 with slight modification. The extracted sample (50 µL) of each developmental stage was mixed with a 30 µl solution of 5% sodium nitrite and 60 µL of AlCl3 (10%). A 200 µL of 1 M NaOH solution was added to the mixture and adjusted to 1 mL, followed by 5 minutes of mixing using a vortex. The absorbance of the mixture was measured at 760 nm and the result was expressed as mg of rutin equivalents (RE) per 100 g.2.6 One-factor experimental design
The factors influencing the quality of probiotic beverages from Sohiong were initially identified. The factors include incubation temperature, probiotic strains, inoculum size (cells per mL), and inoculation period. A one-factor-at-a-time (OFAT) approach is employed for determining the optimal level of each factor for subsequent experiments. DPPH scavenging activity was used as the primary quality index because it reflects antioxidant functionality, while TPC, TAC, and TFC were used as supporting reference indices. Considering the results of the one-factor experiments, the Plackett–Burman design was used to further screen and analyse the significance of the four selected factors.Based on the findings of the PB experiment, the factors with significant response values were chosen. Six gradients, each with three parallel lines, were established, and the regression model and experimental experience determined the step size and direction. To determine the range and centre points of relevant factors in the ensuing tests, the indexes were carried out in accordance with the one-factor optimal conditions.
Based on the elements influencing importance, the BB experimental design was carried out in accordance with the findings of the steepest-climbing experiment and the PB experiment. The values of the associated factors for the group with the highest DPPH scavenging activity in the steepest-ascending experiment served as the focal point.
2.7 Artificial neural network coupled with genetic algorithm (ANN–GA)
ANN was constructed to model DPPH scavenging activity (%) from three independent variables: inoculum size, inoculation period, and temperature. A feed-forward backpropagation ANN with a 3-10-1 topology (three inputs, one hidden layer with 10 neurons, and one output) was implemented using MATLAB R2021b. Secondary antioxidant parameters (TPC, TAC, and TFC) were evaluated under the optimized conditions for comparative analysis.The dataset size of 25 was split into training (70%), validation (15%), and testing (15%) subsets. The validation set was essential for implementing early stopping and ensuring that both ANN and ANN–GA models generalize beyond the training data. Unlike RSM, which estimates only a limited number of regression coefficients, ANN models contain many adjustable parameters and are therefore prone to overfitting, particularly when trained on small datasets. This risk is further amplified when GA is used to optimize weights and biases. Incorporating cross-validation helps prevent overfitting and ensures that the ANN and ANN–GA models generalize reliably. A logsig–logsig–purelin transfer function configuration was adopted for the input, hidden, and output layers, respectively. The Levenberg–Marquardt algorithm was used to minimize mean squared error (MSE).
Although ANN models are traditionally associated with large datasets, they are increasingly applied to small, well-designed experimental datasets such as those generated by Box–Behnken or central composite designs because these structured designs efficiently capture the underlying nonlinearity with a minimal number of experimental points. In this context, the ANN does not require thousands of data points; instead, it benefits from the high information content and orthogonality of the design matrix. Unlike RSM, which is constrained to quadratic polynomial relationships, the ANN can model complex, nonlinear interactions among variables even when the dataset contains only 15–30 observations. Therefore, the use of ANN is justified not by dataset size alone but by its superior ability to learn nonlinear response patterns inherent to fermentation processes.
To enhance prediction accuracy, a GA was employed to optimize the ANN's initial weights and biases. The GA was run for 100 generations with a population size of 50, using a crossover fraction of 0.8 and mutation rate of 0.01. The fitness function minimized the squared difference between the network output and the target antioxidant values. Overall, the experiment has done by screening the factors, optimizing by RSM, modelling by ANN–GA and the obtained results were validated.
2.8 Statistical analysis
The statistical significance was determined using Analysis of Variance (ANOVA) conducted through Design-Expert software (Version 6.0.8, Stat-Ease Inc., Minneapolis, MN, USA), with a p-value <0.05 considered significant. Both Plackett–Burman and Box–Behnken designs, as well as 3D response surface modeling, were implemented using Design-Expert. ANN modeling and GA-based optimization were performed using MATLAB R2023a (MathWorks, Natick, MA, USA), including data processing and visualization.3 Results and discussions
3.1 Plackett–Burman screening
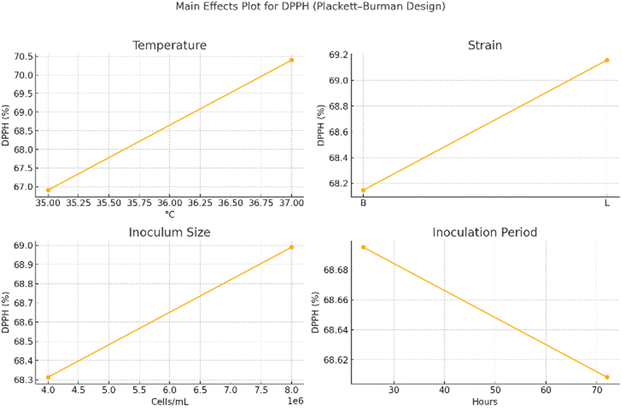
A 12-run design was used to evaluate four real factors (temperature, strain, inoculation period, and inoculum size) and seven dummies, using DPPH as the primary response. Regression analysis identified temperature (p = 0.000003), strain (p = 0.006), and inoculum size (p = 0.035) as statistically significant contributors to DPPH scavenging activity. In contrast, inoculation period was not significant (p = 0.748). These results support the prioritization of the three significant variables for optimization in the steepest-ascent and RSM phases.Antioxidant activity increased as the inoculum size was raised to the optimal level. This is possibly because higher viable cell counts accelerate metabolite production. It was found that a 24-hour inoculation time was ideal. Longer fermentation times reduced antioxidant content by degrading phenolics and anthocyanins, whereas shorter fermentation times led to incomplete fermentation. Optimal probiotic growth and metabolic release were supported at the ideal temperature of 37 °C. Higher temperatures increased stress and reduced the stability of bioactive compounds, while lower temperatures hindered microbial activity.
Optimisation process of fermentation commonly begins with a Plackett–Burman (P–B) screening to identify key factors, followed by RSM for refined modelling. For example, Bajpai et al.19 used P–B design to screen significant medium components for CoQ10 production, and then optimized their levels via RSM. This yielded a substantial increase in CoQ10 titer (from 10.8 to 18.57 mg L−1) under the RSM-optimized conditions. Similarly, Hu et al.16 optimized coffee pulp wine fermentation by first pinpointing critical factors (material-to-liquid ratio, pH, sugar, yeast inoculum) through P–B screening, then applying RSM (central composite design) to model the fermentation response. In both cases, RSM proved effective in navigating multifactor interactions to improve fermentation metrics. For instance, the sugarcane–papaya wine optimization by Patil et al.20 achieved significant enhancement in product yields and antioxidant levels using a comparable RSM-based approach. These findings align well with the current study, where a P–B design likely identified influential variables (e.g., temperature, time, inoculum size) for subsequent RSM modelling. The use of RSM in the Sohiong study is in line with the literature. It provides a quadratic model capturing factor interactions, which is crucial since fermentation outcomes depend on a synergy of conditions. Notably, RSM optimization in fruit-based fermentations has led to improved product qualities. Deshaware et al.21 optimized bitter gourd fermentation to boost nutrients, and Yuan et al.22 optimized jujube wine fermentation for maximal antioxidant content. These precedents support the approach used in the Sohiong juice study and provide a benchmark for expected improvements. The main effects plot (Fig. 1) illustrates a steep upward trend in DPPH with increasing temperature, corroborating the strong coefficient observed in the model. Strain L (Lactobacillus plantarum MCC 2974) outperformed strain B (Bifidobacterium BB-12), validating its selection for final formulation. Similarly, increased inoculum size positively influenced antioxidant activity.
The regression equation based on coded variables is:
| Y(DPPH) = 4.90 + 1.745X1 + 0.505X2 + (1.69 × 10−7)X4 − 0.0018X3 |
3.2 Steepest-ascent experiment
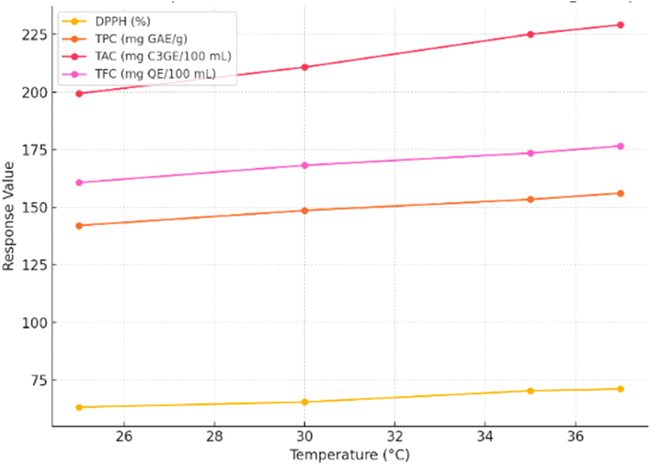
Steepest-ascent experiments were conducted to approach the region of maximum DPPH activity by adjusting the most influential factors identified from the Plackett–Burman design: temperature, strain, and inoculum size. In this phase, Lactobacillus plantarum MCC 2974 was used as the sole strain, while the inoculation period and inoculum size were held constant (24 hours and 4 × 106 CFU mL−1, respectively) to focus on the effect of temperature.The results demonstrated a consistent rise in DPPH scavenging activity with increasing temperature (Fig. 2). The maximum DPPH of 71.11% was achieved at 37 °C, establishing this point as the centre point for the subsequent Box–Behnken design. Corresponding improvements in TPC (156.10 mg GAE per g), TAC (229.23 mg C3GE per 100 mL), and TFC (176.56 mg QE per 100 mL) were also observed at this optimal temperature, indicating co-enhancement of secondary antioxidant parameters.
3.3 Box–Behnken/central composite design and RSM modelling
A three-factor Box–Behnken design was developed using inoculum size (A), inoculation period (B), and temperature (C) as independent variables, with DPPH (%) as the response. The model was fitted to a quadratic polynomial using Design-Expert v6.0.8.The final regression equation in terms of coded factors was:
| Y = 68.06 + 0.014A − 0.011B + 2.52C + 0.32A2 + 0.15B2 + 0.23C2 + 0.16AB − 0.045AC − 0.065BC |
ANOVA results showed that the model was statistically significant (p < 0.001) with R2 = 0.9467, adjusted R2 = 0.8782, predicted R2 = 0.5193, lack-of-fit F = 1.42 (p = 0.3597; not significant), adequate precision = 10.698 (detailed table is provided in the SI). A non-significant lack-of-fit implies that the model adequately fits the data, with no unexplained variation beyond random error. This supports the use of the model for process prediction and optimization.
The response surface plots showed elliptical and curved profiles, confirming the presence of significant interaction effects among the selected factors (Fig. 3). The plots revealed that increasing temperature had a consistent positive impact on DPPH activity, especially when combined with lower inoculum size and shorter fermentation periods.
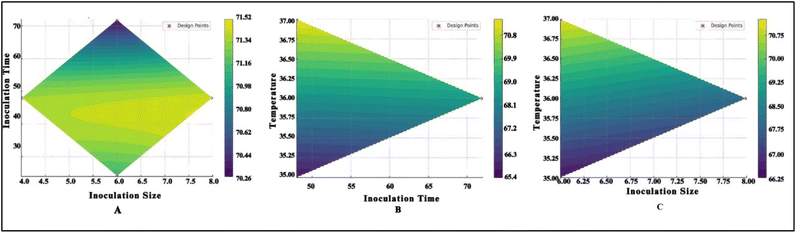 | ||
| Fig. 3 Response-surface and contour plots illustrating the relationships among different process variables for DPPH. | ||
The predicted optimum conditions for DPPH maximization were inoculum size = 4.00 × 106 CFU mL−1; inoculation period = 24 hours; temperature = 37 °C, and predicted DPPH = 71.55%. These conditions were selected for validation and further modelling using ANN.
3.4 Artificial neural network coupled with genetic algorithm (ANN–GA)
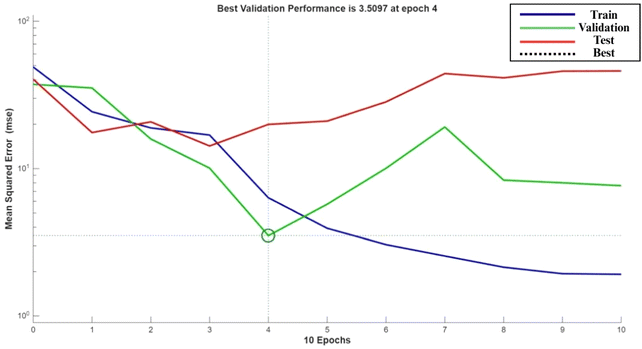
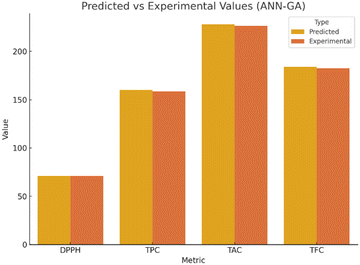
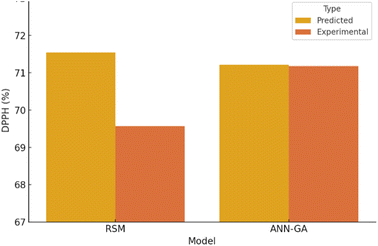
A 3-10-1 feed-forward ANN (logsig–logsig–purelin) was trained on the same design matrix (70% train/15% validate/15% test). After GA optimisation of initial weights and thresholds (population = 50, generations = 100), convergence was reached in 100 generations with minimal fitness deviation. The best model performance was observed at epoch 4, with a validation MSE of 3.5097, as shown in Fig. 4. Regression analysis yielded an overall R2 of 0.9988, confirming an excellent fit between experimental and predicted values. The final optimal input conditions from ANN–GA predicted DPPH = 71.22%, which closely matched the experimentally obtained value of 71.18% (0.04% deviation).The optimized ANN–GA predicted maximum DPPH under the following conditions: inoculum size (8.82 × 106 CFU mL−1); inoculation time (24 h); strain (L. plantarum MCC 2974) predicted DPPH as 71.22%; TPC as 160.00 mg GAE per g; TAC as 227.85 mg C3GE per 100 mL; and TFC as 183.96 mg QE per 100 mL as represented in Fig. 5. These outputs confirm the ANN–GA's ability to achieve or slightly exceed the predicted maxima from the RSM approach, demonstrating its superior predictive fidelity and multi-response optimization capacity.
An important insight across recent studies is that ANN models coupled with GA (ANN–GA) often outperform RSM in capturing nonlinear fermentation dynamics. In Bajpai et al.19 CoQ10 work, an ANN–GA model trained on the same data achieved an R2 of ∼0.999 with a negligible mean squared error (0.0059), compared to R2 ≈ 0.99 for the RSM quadratic model.
The ANN thus explained variance slightly better, reflecting its ability to learn complex, non-linear relationships beyond the polynomial form of RSM. This trend is echoed by Hu et al.16 who reported the ANN–GA model for coffee pulp wine had a higher explained variance (R2 = 0.914) than the RSM model (R2 = 0.890) and a lower root-mean-square error (RMSE = 0.0896 vs. 0.0968). In other words, the ANN–GA provided a tighter fit to the fermentation data, improving predictive accuracy. Anastácio et al.23 observed the same pattern in a phenolic extraction study: the ANN model for DPPH antioxidant response showed higher R2 and lower RMSE than the corresponding RSM model. Even when RSM models are statistically significant, ANNs can capture residual patterns or interactions that RSM misses. In this study, a similar outcome is evident: both modelling approaches likely achieved high R2 (indicative of good fit), but the ANN (especially when optimized via GA) would show lower prediction errors (e.g. RMSE or AAD) than RSM. This aligns with the previous study conducted by Muthusamy et al.11 and Mukherjee et al.24 to generalize better (with lower average absolute deviation, AAD) than RSM in bioprocess optimizations. In summary, the literature consistently shows (and the Sohiong juice results confirm) that ANN–GA modelling can provide more accurate predictions of fermentation outcomes, reflected in higher R2 and smaller error metrics, even though RSM remains valuable for its statistical interpretability.
3.5 Model comparison and validation
ANN–GA shows improved agreement between predicted and actual values compared to RSM (Fig. 6). The ANN–GA model exhibited superior predictive accuracy, as indicated by a higher R2 value (0.9812 vs. 0.9988), lower RMSE (1.12 vs. 0.63), and reduced AAD (1.46% vs. 3.21%) (Table 1). Experimental validation under ANN–GA optimized conditions yielded a DPPH value of 71.18%, differing by only 0.04% from the predicted value of 71.22%, demonstrating excellent model reliability and robustness.| Metric | RSM | ANN–GA |
|---|---|---|
| R2 (unitless) | 0.9467 | 0.9988 |
| RMSE (% DPPH) | 1.98 | 0.63 |
| AAD (%) | 3.21 | 1.46 |
A crucial aspect of optimization studies is how well the models predict actual outcomes (i.e. the accuracy of optimization). The literature suggests that when models are properly trained, their optimum predictions are very reliable. In Bajpai et al.19 study, after ANN–GA optimization predicted an improved CoQ10 yield (≈27.9 mg L−1), experimental validation under those conditions gave 27.04 mg L−1, only about a 2% deviation from the prediction. This tight agreement exemplifies high optimization accuracy. In the coffee pulp wine study (Hu et al. 2025),16 the authors report that the ANN–GA optimal conditions – material ratio ∼4.25%, initial pH 6.92, sugar ∼22.25%, yeast 1.98% were validated experimentally, yielding 10.248 mg L−1 of the target response versus a predicted 10.255 mg L−1. The error here was virtually negligible (≪1%), again highlighting excellent predictive fidelity. By comparison, RSM in that study, while slightly less precise, also provided a solid prediction (the RSM-optimal outcome was only marginally off the ANN's, with R2 ∼0.89). In the bitter gourd-grape fermentation, RSM models were validated by additional experiments and an ANN, which confirmed the RSM-optimised level as near-optimal (e.g., the chosen 35% bitter gourd juice level was deemed ideal and produced the intended low-alcohol, high sensory-acceptance profile). These examples are consistent with the current study, in which the optimized probiotic juice presumably met the targeted quality metrics very close to the model's predictions.
The model fitting parameters reported in the Sohiong manuscript (e.g. high R2 values, low RMSE and AAD) reflect the strong correlation between predicted and observed values at the optimum. This is corroborated by other recent optimizations: for instance, Lau et al.25 optimized a lipase fermentation using RSM and ANN–GA and achieved ∼1.6-fold higher enzyme titer with both methods, with the ANN–GA slightly edging RSM in accuracy. They note that both approaches predicted similar optima, but the ANN–GA model exhibited an improved predictive match to the experimental results (owing to lower error metrics). Likewise, in an ultrasound-assisted extraction study,26 the ANN–GA optimum predictions for yield and antioxidant endpoints showed <5% deviation from experimental values, whereas RSM's optimum had a bit larger gap. Across these cases, a common thread is that optimization accuracy is very high (often within a few percent) when adequate modelling (especially using ANN–GA or a hybrid approach) is employed. The Sohiong juice optimization appears to follow this trend, as the manuscript reports strong agreement between model-predicted and actual values for DPPH, TPC, etc., post-optimization. This high degree of accuracy is critical in validating the chosen approach. It demonstrates that the ANN–GA and RSM models were not only statistically sound but also practically reliable in guiding the formulation of the probiotic juice with maximal functional benefits.
3.6 Secondary quality indices (TPC, TAC, and TFC)
Under the validated optimal conditions, the antioxidant indices showed marked improvement compared with the unfermented control. TPC increased by 22.6% (from 130.5 to 160.0 mg GAE per g); TAC increased by 17.1% (from 194.6 to 227.85 mg C3GE per 100 mL); and TFC increased by 21.7% (from 151.1 to 183.96 mg QE per 100 mL). These enhancements reflect enhanced polyphenolic liberation and biotransformation through controlled fermentation by L. plantarum. The results are consistent with similar trends observed in fermented fruit systems reported in the literature.One of the main goals in functional beverage fermentation is to enhance both the bioactive compounds as well as the antioxidant activity. Multiple studies over the past decade have documented significant improvements in DPPH scavenging activity, TPC, TFC, and TAC following optimization of fermentation conditions. Liao et al.27 provide a striking example: by optimizing the co-fermentation of blueberry juice with mixed probiotics (using a simplex mixture design and GA optimization), they achieved an 82.2% increase in TPC and dramatic rises in specific polyphenols (e.g., rutin up 79%) in the fermented juice compared to the unfermented control.
This coincided with a sharp increment in antioxidant activity, as evidenced by the higher DPPH scavenging capacity in the fermented blueberry juice. In a similar study, Yuan et al.28 observed that fermenting an apple–tomato pulp with lactic cultures led to TPC and TFC increases of ∼21–22%, accompanied by a 40.9% improvement in DPPH free-radical scavenging ability (and ∼22% increase in ABTS activity) relative to the unfermented pulp.
These enhancements are attributed to microbial biotransformation of phytochemicals for instance, Lactobacillus strains can release bound phenolics from the fruit matrix or biosynthesize antioxidant metabolites. The Sohiong-based probiotic juice, rich in anthocyanins (Sohiong is a dark purple fruit), likely showed analogous trends: fermentation optimization would increase TAC and other phenolics, thereby boosting DPPH activity. Indeed, probiotic fermentation is known to elevate anthocyanin content and antioxidant power in fruit substrates. Sun et al.29 and others have reported enhanced anthocyanin stability and antioxidant profiles in fermented berry wines, which aligns with Section 3.5 of the Sohiong study. Additionally, Maselesele et al.30 demonstrated that optimizing a bitter gourd-grape fermentation not only controlled alcohol content but also retained functional components; their RSM-optimized low-alcohol beverage maintained high phenolic content and antioxidant activity. By comparison, the Sohiong juice optimization appears to have succeeded in maximizing these health-related metrics. Any reported gains in DPPH, TPC, TAC, and TFC in the Sohiong study are well within the spectrum of improvements seen in the aforementioned studies.
4 Conclusion
ANN–GA hybrids consistently outperform RSM in predictive accuracy, which matches the Sohiong study's observation of slightly better performance by the ANN model (as indicated by lower RMSE/AAD in section 3.4). Finally, the optimization accuracy in the Sohiong study, evidenced by minimal differences between predicted optimum values and experimental results, is well supported by analogous successes in the literature. In conclusion, the Sohiong-based probiotic juice optimization is corroborated by numerous contemporary studies: all demonstrate that combining statistical designs (P–B, RSM) with machine learning optimization (ANN–GA) yields strong models and maximizes functional properties of fermented beverages with a high degree of confidence in the results. This comparative analysis reinforces the validity of the approaches and outcomes reported in the manuscript, placing them within the broader trend in fermentation process optimization over the last decade. Although the ANN–GA model showed better prediction reliability, the comparatively limited dataset size is still a limitation. Larger datasets (>100) in future research will allow for stronger model training, which will improve the applicability of AI-based optimisation in functional food systems. The marked improvement in antioxidant metrics corroborates the efficacy of probiotic fermentation for valorising Prunus nepalensis pulp. It is acknowledged that the viable cell count was not measured after fermentation, as the focus was on optimizing bioactive content rather than viability. However, future work will include determining the viable cell count at the end of fermentation and during storage. Further research should explore scale-up kinetics, shelf-life stability, and sensory acceptability to translate these laboratory insights into commercial applications.Author contributions
Roshiya Nongmaithem: writing – original draft preparation, review & editing; Raju Sasikumar: conceptualization, investigation, data curation; Irengbam Barun Mangang: writing – original draft preparation; Selva Kumar T: investigation, data curation; Kambhampati Vivek: writing – original draft preparation, review & editing; Amit K. Jaiswal: validation, review & editing, conceptualization.Conflicts of interest
There are no conflicts to declare.Data availability
The data is made available on request.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5fb00940e.
References
- H. Rymbai, V. K. Verma, H. Talang, S. R. Assumi, M. B. Devi, Vanlalruati, R. H. Sangma, K. P. Biam, L. J. Chanu, B. Makdoh, A. R. Singh, J. Mawlein, S. Hazarika and V. K. Mishra, Front. Nutr., 2023, 10, 1039965 CrossRef PubMed.
- A. Thakur, Pharma Innov., 2023, 12, 1252–1256 CrossRef CAS.
- T. L. Swer, K. Chauhan, P. K. Paul and C. Mukhim, Int. J. Biol. Macromol., 2016, 92, 867–871 CrossRef CAS PubMed.
- T. L. Swer, K. Chauhan, C. Mukhim and P. K. Paul, LWT, 2019, 114, 108360 CrossRef CAS.
- C. Deka, S. Roy and K. Pandey, Journal of North East India Studies, 2014, 4, 70–80 Search PubMed.
- C. Mazziotta, M. Tognon, F. Martini, E. Torreggiani and J. C. Rotondo, Cells, 2023, 12, 184 CrossRef CAS PubMed.
- S. Boro, G. Brahma, D. Ray, A. Das and S. Das, Futuristic Trends in Agriculture Engineering & Food Sciences, Iterative International Publishers (IIP), India, 5th edn, 2024, ch. 19, pp. 1–22 Search PubMed.
- R. Sasikumar, H. Chutia and S. C. Deka, J. Microbiol. Biotechnol. Food Sci., 2019, 9, 228 CrossRef CAS.
- R. Sasikumar, K. Vivek, G. Kadirvel, A. K. Jaiswal and J. Agricul, Food Res., 2024, 17, 101224 CAS.
- E. N. Dragoi, S. Curteanu, A. I. Galactiun and D. Cascaval, Appl. Soft Comput., 2013, 13, 222–238 CrossRef.
- S. Muthusamy, L. P. Manickam, V. Murugesan, C. Muthukumaran and A. Pugazhendhi, Int. J. Biol. Macromol., 2019, 124, 750–758 CrossRef CAS PubMed.
- J. Xi, Y. Xue, Y. Xu and Y. Shen, Food Chem., 2013, 141, 320–326 CrossRef CAS.
- G. E. Lee, R. H. Kim, T. Lim, J. Kim, S. Kim, H. G. Kim and K. T. Hwang, Food Chem., 2022, 396, 133712 CrossRef CAS PubMed.
- R. Sasikumar, I. B. Mangang, K. Vivek and A. K. Jaiswal, J. Food Process. Eng., 2023, 46, e14468 CrossRef CAS.
- S. Baliyan, R. Mukherjee, A. Priyadarshini, A. Vibhuti, A. Gupta, R. P. Pandey and C. M. Chang, Molecules, 2022, 27, 1326 CrossRef CAS PubMed.
- R. Hu, F. Xu, L. Zhao and W. Dong, Sci. Rep., 2025, 15, 16684 CrossRef CAS PubMed.
- E. Azarpazhooh, P. Sharayei, S. Zomorodi and H. S. Ramaswamy, Food Bioprocess Technol., 2019, 12, 199–210 CrossRef CAS.
- D. R. Singh and V. S. Banu, Int. J. Innov. Hortic., 2014, 3, 150–153 Search PubMed.
- S. Bajpai, S. Singh, R. Sinha and P. Srivastava, Int. J. Res. Pharm. Sci., 2015, 6, 100–107 CAS.
- P. S. Patil, U. B. Deshannavar, M. Ramasamy and S. Emani, Environ. Technol. Innov., 2021, 21, 101290 CrossRef CAS.
- S. Deshaware, S. Gupta, R. S. Singhal and P. S. Variyar, LWT, 2017, 86, 514–522 CrossRef CAS.
- L. Yuan, G. Li, N. Yan, J. Wu and J. Due, J. Food Sci. Technol., 2022, 59, 288–299 CrossRef CAS PubMed.
- A. Anastácio, R. Silva and I. S. Carvalho, J. Food Sci. Technol., 2016, 53, 4117–4125 CrossRef PubMed.
- R. Mukherjee, R. Chakraborty and A. Dutta, J. Food Process. Eng., 2019, 42, e13124 CrossRef.
- H. L. Lau, F. W. F. Wong, R. N. Zaliha, M. S. Mohamed, A. B. Ariff and S. L. Hii, Biocatal. Agric. Biotechnol., 2023, 50, 102696 CrossRef CAS.
- S. Shekhar, P. Prakash, P. Singha, K. Prasad and S. K. Singh, Foods, 2023, 12, 1925 CrossRef CAS PubMed.
- W. Liao, J. Shen, S. Manickam, S. Li, Y. Tao, D. Li, D. Liu and Y. Han, Food Chem., 2023, 405, 134982 CrossRef CAS PubMed.
- J. Yuan, H. Zhang, C. Zeng, J. Song, Y. Mu and S. Kang, Molecules, 2023, 28, 4363 CrossRef CAS PubMed.
- X. Sun, Z. Yan, J. Zhu, Y. Wang, B. Li and X. Meng, Food Chem., 2019, 279, 63–69 CrossRef CAS.
- T. L. Maselesele, T. B. J Molelekoa, S. Gbashi and O. A. Adebo, Plants, 2023, 12, 3473 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |