Open Access Article
Open Access ArticleMicrobial-dependent variations in umami compounds during fermentation of prawn by-products
Cia Min Lim ,
Ying Tong Yeo
,
Ying Tong Yeo ,
Cherie Chin
,
Cherie Chin and
Wei Ning Chen
and
Wei Ning Chen *
*
School of Chemistry, Chemical Engineering and Biotechnology, Nanyang Technological University, Singapore 637459, Singapore. E-mail: WNChen@ntu.edu.sg; Fax: +65 67947553; Tel: +65 63162870
First published on 18th April 2026
Abstract
This study investigated the effects of microbial fermentation on the umami taste components of prawn by-products. Six microorganisms — Aspergillus oryzae, Rhizopus oligosporus, Saccharomyces cerevisiae, Bacillus subtilis, Lactiplantibacillus plantarum and Latilactobacillus sakei — were used to ferment prawn by-product juice. The resulting broths were analyzed for umami-related compounds, including free amino acids and flavor nucleotides. Taste activity values (TAVs) and equivalent umami concentrations (EUCs) were calculated to assess umami intensity. Fermentation enhanced acidity and significantly increased the concentrations of umami-active compounds. The EUC values increased markedly from 0.23 g monosodium glutamate (MSG)/100 g at day 0 to 6.36 g MSG/100 g on day 3 in the samples fermented with R. oligosporus. Glutamic acid was identified as the dominant umami taste compound and its TAV exceeded 1 in fermented samples, confirming a perceptible umami contribution. Among the tested strains, R. oligosporus produced the most pronounced umami profile, indicating its potential as an effective starter culture for transforming prawn by-products into value-added, umami-rich ingredients.
Sustainability spotlightThis research introduces a sustainable pathway for the food circular economy by valorizing prawn side-streams, which are a common waste stream from seafood processing, into a high-value food ingredient using fermentation. By reducing reliance on non-renewable resources and transforming waste into functional products, this innovation supports circular economy principles, promotes healthier diets through natural flavors, and significantly reduces food waste, directly addressing UN SDGs 3, 9, and 12 for a more resilient food system. |
1. Introduction
Prawn is one of the most widely traded seafood commodities globally, with international trade volumes exceeding 3.7 million tonnes in recent years.1 Due to their high perishability, fresh prawns are usually processed at low temperatures upon harvest to extend their shelf life. However, such processing generates large quantities of by-products, namely the head, shell, and tail, which can account for approximately 45–60% of the total prawn weight, depending on the species and processing method.2 The global prawn market, valued at approximately USD 74.7 billion in 2025 and projected to reach USD 106.1 billion by 2034, continues to expand, driven largely by the dominance of the Litopenaeus vannamei species, which contributes over 70% of global production.3 This growth is expected to further increase the volume of processing residues, which is currently estimated at around 3.8 million tonnes annually.4 The accumulation of these by-products presents both environmental and economic challenges, highlighting the urgent need for sustainable valorization strategies. These by-products, particularly prawn heads, are rich in free amino acids and nucleotides, making them well-suited for the production of flavor-enhancing ingredients.5Fermentation, a traditional preservation and processing method, has been widely recognized for improving the sensory attributes of foods, particularly flavor and texture. Microorganisms play a pivotal role in this process; through their metabolic activities, they transform food components, imparting unique sensory characteristics and potential health benefits.6 Fermentation has also been applied to seafood products, such as fish sauces, prawn pastes, and fermented fish, not only for preservation but also to enhance flavor and palatability.7
Voidarou et al.8 classified food fermentation processes into three major types, depending on the dominant microorganism and its metabolic activities: lactic acid fermentation, fungal fermentation and alkaline fermentation. The filamentous fungus Aspergillus oryzae, used in soy sauce fermentation, contributes to its characteristic umami flavor through the production of amino acids, nucleotides, and organic acids.9 Similarly, Rhizopus oligosporus, utilized in the production of tempeh, enhances umami intensity through the breakdown of soybean proteins to amino acids.10 Yeasts, in particular Saccharomyces cerevisiae, are commonly employed in the production of fermented beverages like beer and wine, generating esters, organic acids, and other aroma-active products that define the sensory profiles of these fermented beverages.11 In alkaline fermentation, Bacillus subtilis has also been associated with improved sensory quality in pulses and teas by generating peptides and amino acids and reducing undesirable flavor and odor compounds, such as hexanal, a compound responsible for ‘grassy’ and ‘beany’ off-flavors.12,13 Among lactic acid bacteria (LAB), Lactiplantibacillus plantarum (formerly Lactobacillus plantarum) has been widely used due to its versatility and has been found to enhance the umami flavor through the release of amino acids.14 Latilactobacillus sakei (formerly Lactobacillus sakei), another lactic acid bacterium, contributes to flavor development in a diverse range of substrates, such as rice in sake,15 cabbage in kimchi16 and meat in sausages,17,18 through the production of fermentation metabolites.
Flavor is a critical determinant of consumer preference and an important indicator of product quality.19 It encompasses both taste and odor, which arise from the stimulation of gustatory and olfactory receptors by non-volatile and volatile compounds, respectively.20 Umami, one of the five basic taste sensations, plays a critical role in defining the palatability of seafood. In prawns, this taste primarily arises from synergistic interactions among free amino acids, nucleotides, inorganic ions, and organic acids.21
Recent research on prawn by-products has mainly focused on recovering chitin, proteins, peptides, carotenoids, and antioxidants from solid waste such as heads, shells, and tails, often using enzymatic hydrolysis or microbial fermentation.22–27 In industry, prawn waste is typically pressed into liquid and solid fractions to reduce processing energy for the solid fraction. The liquid fraction, accounting for approximately 70% (w/w) of total waste, is often discarded into wastewater streams, increasing treatment costs and resulting in the loss of valuable nutrients, while the solid fraction is utilized for prawn shell meal or chitin and chitosan production.28 To address this inefficiency, Nguyen et al.28 explored the valorization of prawn liquid waste via enzymatic hydrolysis, producing nutrient-rich protein hydrolysates with potential applications in aquaculture. Nevertheless, the potential of fermentation to enhance flavor and umami in prawn by-product juice remains largely unexplored, despite evidence that microbial processes can generate taste-active compounds such as free amino acids and nucleotides in solid substrates.29
As a result, there is a significant gap in the understanding of how microbial fermentation will impact the umami characteristics of prawn by-product juice. Hence, to address this knowledge gap, this present study explores the fermentation of prawn by-product juice using six microorganisms, namely A. oryzae, R. oligosporus, S. cerevisiae, B. subtilis, L. plantarum and L. sakei. The resulting fermented products were analyzed for flavor-related compounds, specifically free amino acids and flavor nucleotides, to identify the most effective microorganism for enhancing umami intensity. This study focused on umami-contributing compounds; however, free amino acids (FAAs) with sweet or bitter attributes are also reported to provide a more comprehensive sensory profile. The findings are expected to provide a theoretical foundation for the valorization of prawn by-products through the development of novel umami-rich fermented ingredients. The growing emphasis on sustainability and the adoption of green technologies to manage food waste further underscore the significance of this work, which aims to advance a zero-waste approach while creating a natural prawn-based flavoring with health benefits.
2. Methods and materials
2.1. Materials and reagents
LC-MS grade acetonitrile (ACN) and formic acid were purchased from Fisher Scientific. Adenosine-5′monophosphate disodium salt (≥99.0%, HPLC), guanosine-5′-monophosphate disodium salt hydrate (from yeast, ≥99%), and inosine-5′-monophosphate disodium salt hydrate (≥99.0%, HPLC), amino acid standards (AAS18), HPLC-grade hexane, and LiChropur-grade reagents for LC-MS analysis, including ammonium acetate, ammonium formate, and 25% ammonia solution were purchased from Sigma-Aldrich (St. Louis, MO, USA). All water used in this research project was ultrapure (Type 1) and obtained from a LaboStar PRO TWF UV ultrapure water system (Evoqua Water Technologies, Günzburg, Germany).Individual nucleotide stock solutions (10 mM) were prepared in ultrapure water from the commercial standards without further purification and stored at −20 °C in the dark. Working standard solutions were freshly prepared daily. The amino acid standard mixture was also prepared freshly on the day of analysis.
2.2. Sample preparation
Fresh prawns were purchased from a local supermarket (NTUC FairPrice, Singapore). The samples were marketed as Pasar Vannamei Prawn (L. vannamei), whole prawn with head and shell, with a size grade of ‘L’ (body length: 15.5 ± 0.4 cm; weight: 23.8 ± 1.05 g). The samples were transported to the laboratory under chilled conditions and were stored at −20 °C upon arrival until further processing. The prawn heads, shells and tails were manually separated from the meat and rinsed three times with ultrapure water. The by-products were blended with water at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (w/v) ratio using a Ninja Foodi Power Blender (Extract Function). The homogenate was filtered through a 100-mesh nylon food sieve to obtain prawn juice, which was subsequently lyophilized and stored at −20 °C until further processing.
1 (w/v) ratio using a Ninja Foodi Power Blender (Extract Function). The homogenate was filtered through a 100-mesh nylon food sieve to obtain prawn juice, which was subsequently lyophilized and stored at −20 °C until further processing.
Prior to fermentation, the lyophilized prawn juice powder was reconstituted with ultrapure water to a concentration of 5% (w/v) and pasteurized in a water bath at 90 °C for 30 min. The absence of viable microorganisms in the reconstituted pasteurized prawn juice was confirmed by plating on a variety of selective and non-selective agar, including potato dextrose agar, Mueller–Hinton agar, Lysogeny Broth agar, plate count agar, Sabouraud dextrose agar with chloramphenicol, and De Man, Rogosa, and Sharpe Agar (data not shown).
2.3. Microorganisms and culture conditions
The microbial strains used in this study were A. oryzae DSM 1147, R. oligosporus DSM 1964, S. cerevisiae BY4741, B. subtilis ATCC 6051, L. plantarum ATCC 14917, and L. sakei subsp. sakei ATCC 15521. Each microorganism was maintained and activated in its corresponding culture medium prior to inoculation (Table S1). The cells or spores of the activated microorganisms were collected by centrifugation at 8000×g for 10 min at 4 °C and washed twice with sterile phosphate buffer (pH 7.0). The resulting cell or spore suspension was used as the inoculum for fermentation experiments.2.4. Fermentation of prawn juice
The reconstituted pasteurized prawn juice substrate was supplemented with 2% (w/v) glucose to support microbial growth. Fermentation was initiated by inoculating with 1% (v/v) of the cell (S. cerevisiae, B. subtilis L. plantarum, and L. sakei) (approx. 108 CFU mL−1) or spore suspension (A. oryzae and R. oligosporus) (approx. 107 spores per mL).Fermentations were conducted for 5 days under the optimal temperature conditions for each microorganism. The samples were collected on days 1, 2, 3, and 5 to monitor the development of flavor compounds. Following sampling, the pH of the fermentation broths was measured with a calibrated pH meter (FiveEasy F20, Mettler-Toledo, Greifensee, Switzerland), and the fermentation broths were stored at −20 °C until further analysis.
2.5. Determination of free amino acids
The samples were diluted with acetonitrile: water (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, v/v) containing 0.1% formic acid. The mixture was centrifuged at 12
1, v/v) containing 0.1% formic acid. The mixture was centrifuged at 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000×g for 10 min at 4 °C. The supernatant was transferred into LC vials for LC-MS analysis. Working standards (0.5–20 µM) were used to construct an external calibration curve (R2 > 0.99) for detection and quantification of the free amino acids.
000×g for 10 min at 4 °C. The supernatant was transferred into LC vials for LC-MS analysis. Working standards (0.5–20 µM) were used to construct an external calibration curve (R2 > 0.99) for detection and quantification of the free amino acids.
Chromatographic separation was performed using an Agilent 1290 Infinity II Series HPLC system (Agilent Technologies, Santa Clara, CA, USA) equipped with a multi-sampler module and a high-speed binary pump. An Agilent Zorbax HILIC Plus column (3.0 × 100 mm, 1.8 µm) coupled with a HILIC-PLUS guard column (3.0 × 5 mm, 1.8 µm) was used. 20 mM ammonium formate in water with 0.1% formic acid was used as mobile phase A, and 20 mM ammonium formate in acetonitrile/water (9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, v/v) with 0.1% formic acid was used as mobile phase B. The flow rate was maintained at 0.4 mL min−1, column temperature at 30 °C and injection volume at 5 µL. The gradient program was initiated at 100% B, followed by a gradient to 89% at 5 min, 88% B at 7 min, and 75% B by 11 min, 70% B by 15 min. Initial conditions of 100% B were re-established at 17 min which was held until 20 min for re-equilibration. Detection was performed using an Agilent 6456 Q-TOF mass spectrometer with an Agilent Jet Stream Dual electrospray ionization (AJS-Dual ESI) interface operating in positive ionization mode. Source conditions were as follows: gas temperature, 150 °C; drying gas flow, 10 L min−1; nebulizer pressure, 20 psi; sheath gas temperature, 400 °C; sheath gas flow, 12 L min−1; capillary voltage (Vcap), 2000 V; nozzle voltage, 0 V; fragmentor voltage, 80 V; skimmer, 65 V; and octupole RF Vpp, 750 V. Data were acquired over m/z 50–1200 at 2.0 spectra per sec.
1, v/v) with 0.1% formic acid was used as mobile phase B. The flow rate was maintained at 0.4 mL min−1, column temperature at 30 °C and injection volume at 5 µL. The gradient program was initiated at 100% B, followed by a gradient to 89% at 5 min, 88% B at 7 min, and 75% B by 11 min, 70% B by 15 min. Initial conditions of 100% B were re-established at 17 min which was held until 20 min for re-equilibration. Detection was performed using an Agilent 6456 Q-TOF mass spectrometer with an Agilent Jet Stream Dual electrospray ionization (AJS-Dual ESI) interface operating in positive ionization mode. Source conditions were as follows: gas temperature, 150 °C; drying gas flow, 10 L min−1; nebulizer pressure, 20 psi; sheath gas temperature, 400 °C; sheath gas flow, 12 L min−1; capillary voltage (Vcap), 2000 V; nozzle voltage, 0 V; fragmentor voltage, 80 V; skimmer, 65 V; and octupole RF Vpp, 750 V. Data were acquired over m/z 50–1200 at 2.0 spectra per sec.
2.6. Determination of free nucleotides
Nucleotides were extracted using a combination of solvent precipitation and liquid–liquid extraction (LLE). Ice-cold solutions of 10 mM ammonium acetate were prepared (i) in water (pH 9) and (ii) in acetonitrile/water (9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, v/v, pH 9). These were added to the samples in a volumetric ratio of 1
1, v/v, pH 9). These were added to the samples in a volumetric ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (aqueous buffer: organic phase: sample), followed by 30 s vortex mixing, and kept at −30 °C for 15 min to precipitate proteins. The samples were centrifuged (24
1 (aqueous buffer: organic phase: sample), followed by 30 s vortex mixing, and kept at −30 °C for 15 min to precipitate proteins. The samples were centrifuged (24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 446×g, 4 °C, 15 min) and the supernatant was mixed with hexane (1
446×g, 4 °C, 15 min) and the supernatant was mixed with hexane (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3, v/v),30 followed by centrifugation under the same conditions. The aqueous layer was subjected to LC-QTOF analysis. Working standards (0.5–50 µM) were used to construct an external calibration curve (R2 > 0.99) for detection and quantification of the free nucleotides.
3, v/v),30 followed by centrifugation under the same conditions. The aqueous layer was subjected to LC-QTOF analysis. Working standards (0.5–50 µM) were used to construct an external calibration curve (R2 > 0.99) for detection and quantification of the free nucleotides.
Separation of AMP, IMP, and GMP was carried out on an Agilent InfinityLab Poroshell 120 HILIC-Z column (100 × 2.1 mm, 2.7 µm) coupled with a HILIC-Z guard column (5 × 2.1 mm, 2.7 µm) maintained at 30 °C. 10 mM ammonium acetate in water, pH 9, was used as mobile phase A and 10 mM ammonium acetate in acetonitrile/water (9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, v/v), pH 9, adjusted by ammonia solution 25% was used as mobile phase B. The elution program was as follows: 90% B initially, decreasing linearly to 50% B over 12 min (held for 1 min), then returning to 90% within 2 min, and equilibrating for 5 min (total run time 20 min). Injection volume was 10 µL.
1, v/v), pH 9, adjusted by ammonia solution 25% was used as mobile phase B. The elution program was as follows: 90% B initially, decreasing linearly to 50% B over 12 min (held for 1 min), then returning to 90% within 2 min, and equilibrating for 5 min (total run time 20 min). Injection volume was 10 µL.
The QTOF-MS parameters followed those reported by Pastor-Belda et al.31 with modifications: negative ionization mode; nebulizer gas pressure, 40 psi; drying gas, 13 L min−1 at 200 °C; sheath gas, 12 L min−1 at 300 °C; capillary voltage, 2500 V; nozzle voltage, 100 V; fragmentor voltage, 350 V; skimmer voltage, 65 V and octupole RF Vpp, 750 V. Data were acquired from m/z 100–1200 at 3 spectra per sec.
2.7. Evaluation of umami taste contribution
The taste activity value (TAV) is used to evaluate the relative contribution of an individual compound to the overall taste perception. It is defined as the ratio of a compound's concentration in the sample to its corresponding taste threshold, indicates that the compound contributes perceptibly to the overall taste profile when exceeding 1:32
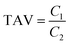 | (1) |
The equivalent umami concentration (EUC) quantifies the synergistic umami intensity effect of specific amino acids, namely L-aspartic acid (Asp) and L-glutamic acid (Glu), in combination with nucleotides such as inosine monophosphate (IMP), guanosine monophosphate (GMP), and adenosine monophosphate (AMP). The EUC expresses the umami intensity as the equivalent concentration of monosodium glutamate (MSG) per 100 g, was calculated as follows:
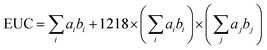 | (2) |
2.8. Data processing and statistical analysis
LC-QTOF data were processed using Agilent MassHunter Quantitative Analysis software (Q-TOF, Quant-My-Way, version 10.1). Compounds were identified based on the retention time and m/z values and quantified using external calibration curves prepared from standard solutions of known concentrations analyzed under identical conditions. Data visualization and statistical analysis were performed using Microsoft Excel 2016 (Microsoft Ltd, WA, USA) and OriginPro 2024 (OriginLab Corporation, MA, USA). One-way ANOVA and two-way ANOVA followed by Tukey's test were applied, with statistical significance set at α = 0.05. The results are reported as mean ± standard deviation (S.D.).3. Results and discussion
3.1. Changes in the pH during fermentation
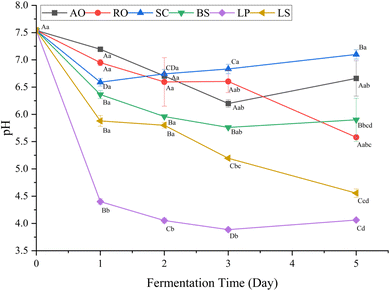
Fig. 1 illustrates the pH changes of the prawn by-product juice fermented with different microbial strains. The pH of the prawn by-product juice decreased progressively during fermentation with all tested microorganisms, consistent with acidification associated with microbial activity.22 The most pronounced acidification occurred in the samples fermented with L. plantarum followed by L. sakei, both of which are lactic acid bacteria known for their efficient carbohydrate utilization and lactic acid formation.34 The reduction in pH may contribute to the suppression of undesirable microorganisms and promote proteolysis, thereby enhancing the release of flavor-related compounds.35 In contrast, fermentation with S. cerevisiae, A. oryzae, R. oligosporus and B. subtilis, resulted in more moderate pH reduction. This milder acidification may be attributed to the high protein and amino-nitrogen content of prawn by-products, which provide buffering capacity against sharp pH changes.36 Also, it is observed that there is slight rebound in the pH in the fermentation of A. oryzae, S. cerevisiae, B. subtilis and L. plantarum. This can be explained by the accumulation of alkaline metabolites during protein degradation in the later stages of fermentation, which may act as a buffer against organic acids, metabolites typically produced in fermentation.37 Overall, the observed pH changes indicate that fermentation modifies the biochemical environment of prawn by-products, thereby facilitating the development of distinct flavor profiles through microorganism-specific metabolic activities.3.2. Effects of fermentation with different microorganisms on free amino acids
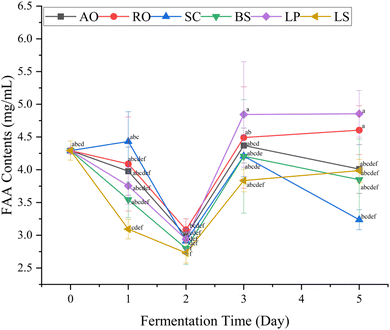
Free amino acids (FAAs) play an important role in flavor development, contributing to the overall sensory characteristics of fermented products. The FAA profile influences taste perception, contributing to umami (glutamic acid and aspartic acid), sweetness (serine, threonine, glycine, alanine and proline), and bitterness (valine, methionine, isoleucine, leucine, phenylalanine, histidine and arginine).38 The content of 17 free amino acids of the fermented prawn by-product juice can be found in Table S2.The changes in FAA concentrations of the prawn by-product juice fermented with different microbial strains throughout the fermentation period are shown in Fig. 2. The total FAA content decreased to its lowest on day 2, suggesting rapid microbial uptake or utilization of the available nitrogen source during the early fermentation stage. A similar reduction in FAAs has been reported in other fermentation systems, such as Pu-erh ripened tea39 and dried longan fermentation.40 The subsequent increase of total FAA content on day 3 likely resulted from enhanced proteolytic activity and peptide hydrolysis during fermentation, indicating active protein degradation by microbial enzymes. The slight decrease observed on day 5 could be attributed to the continued metabolism of amino acids as energy sources or as precursors for secondary metabolite synthesis.41
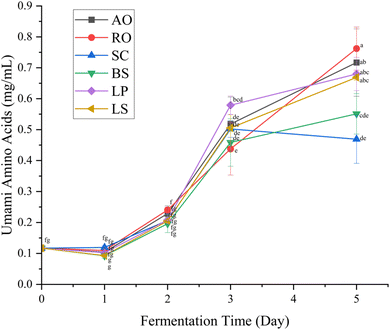
Among the tested microorganisms, L. plantarum and R. oligosporus produced the highest total FAA concentration after five days of fermentation, suggesting their strong proteolytic capacity under the fermentation conditions employed (Table 2). Notably, the concentrations of umami-related amino acids, particularly glutamic acid (Glu) and aspartic acid (Asp), were significantly elevated in all fermentations after day 3 (Fig. 3). This trend is likely due to the high abundance of these amino acids in L. vannamei head and shell proteins, coupled with microbial proteolytic activity during fermentation, which progressively hydrolyzed proteins into free amino acids.42,43 This is in agreement with the findings by Lim et al.,44 where fermentation led to a significant accumulation of Glu and Asp in shrimp paste. These amino acids are key contributors to savory taste perception and serve as precursors for umami enhancement.45 The sweet- and bitter-tasting amino acids follow the trend similar to that of total amino acids (Table 1).
| Amino acids | AO | RO | SC | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| D0 | D1 | D2 | D3 | D5 | D1 | D2 | D3 | D5 | D1 | D2 | D3 | D5 | |
| a N = 3 fermentation trials.b Mean values ± standard deviation with different lowercase superscript letters within the same row are significantly different (two-way ANOVA, Tukey test, and p < 0.05).c Umami = Glu + Asp; Sweet = Thr + Ser + Gly + Ala + Pro; Bitter = Val + Met + Ile + Leu + Phe + His + Arg.d AO – A. oryzae, RO – R. oligosporus, SC – S. cerevisiae, BS – B. subtilis, LP – L. plantarum, LS – L. sakei. | |||||||||||||
| Total | 4.29 ± 0.15abcd | 3.98 ± 0.367abcdef | 2.99 ± 0.07cdef | 4.37 ± 0.27abcd | 4.01 ± 0.37abcdef | 4.09 ± 0.72abcdef | 3.08 ± 0.17bcdef | 4.49 ± 0.77ab | 4.60 ± 0.37a | 4.43 ± 0.46abc | 2.91 ± 0.19def | 4.21 ± 0.31abcde | 3.24 ± 0.15bcdef |
| Umami | 0.12 ± 0.01fg | 0.10 ± 0.003fg | 0.23 ± 0.03fg | 0.52 ± 0.09de | 0.72 ± 0.11ab | 0.11 ± 0.001fg | 0.24 ± 0.01f | 0.44 ± 0.08e | 0.76 ± 0.07a | 0.12 ± 0.005fg | 0.21 ± 0.01fg | 0.50 ± 0.03de | 0.47 ± 0.08de |
| Sweet | 1.08 ± 0.02abc | 1.04 ± 0.04abc | 0.94 ± 0.02abc | 1.27 ± 0.07ab | 0.76 ± 0.27bc | 1.08 ± 0.10abc | 0.94 ± 0.18abc | 1.23 ± 0.36ab | 0.92 ± 0.38abc | 1.18 ± 0.15abc | 0.91 ± 0.07abc | 1.20 ± 0.05ab | 0.66 ± 0.24c |
| Bitter | 1.49 ± 0.03abcde | 1.32 ± 0.22abcdefg | 1.09 ± 0.04efg | 1.40 ± 0.10abcdefg | 1.28 ± 0.19abcdefg | 1.18 ± 0.10defg | 1.15 ± 0.04defg | 1.48 ± 0.21abcde | 1.67 ± 0.42abc | 1.57 ± 0.17abcd | 1.10 ± 0.11defg | 1.35 ± 0.10abcdefg | 1.13 ± 0.09defg |
| Amino acids | BS | LP | LS | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| D1 | D2 | D3 | D5 | D1 | D2 | D3 | D5 | D1 | D2 | D3 | D5 | |
| Total | 3.54 ± 0.27abcdef | 2.80 ± 0.25f | 4.21 ± 0.87abcde | 3.85 ± 0.62abcdef | 3.75 ± 0.28abcdef | 2.94 ± 0.15def | 4.84 ± 0.81a | 4.85 ± 0.36a | 3.09 ± 0.15cdef | 2.73 ± 0.16f | 3.84 ± 0.17abcdef | 3.99 ± 0.17abcdef |
| Umami | 0.09 ± 0.01g | 0.20 ± 0.03fg | 0.46 ± 0.08de | 0.55 ± 0.07cde | 0.10 ± 0.01fg | 0.21 ± 0.001fg | 0.58 ± 0.03bcd | 0.68 ± 0.05abc | 0.09 ± 0.003g | 0.21 ± 0.01fg | 0.51 ± 0.04de | 0.67 ± 0.01abc |
| Sweet | 0.93 ± 0.04abc | 0.80 ± 0.13abc | 1.08 ± 0.38abc | 0.76 ± 0.30bc | 0.98 ± 0.04abc | 0.91 ± 0.06abc | 1.29 ± 0.20a | 1.07 ± 0.06abc | 0.93 ± 0.03abc | 0.90 ± 0.06abc | 1.09 ± 0.07abc | 0.89 ± 0.06abc |
| Bitter | 1.30 ± 0.08abcdefg | 1.15 ± 0.13defg | 1.47 ± 0.37abcdef | 1.33 ± 0.22abcdefg | 1.34 ± 0.06abcdefg | 1.24 ± 0.05bcdefg | 1.70 ± 0.20ab | 1.75 ± 0.10a | 0.97 ± 0.02g | 1.00 ± 0.05fg | 1.21 ± 0.06cdefg | 1.28 ± 0.08abcdefg |
Different microbial strains produce distinct types of proteases, leading to variations in protein breakdown: for instance, bacteria such as Bacillus mainly produce alkaline proteases, while molds like Aspergillus and Rhizopus predominantly produce neutral and acid proteases.46–48 Microbial proteases can be further classified by their catalytic type, such as aspartic endoproteases, cysteine/thiol endoproteases, metalloendoproteases, serine endoproteases, and glutamic acid and threonine endoproteases.49 Once proteins are hydrolyzed, the resulting amino acids can undergo further microbial metabolism through processes such as transamination, deamination, and decarboxylation, generating secondary metabolites.50 This strain-specific diversity in protease type, abundance, and activity helps explain why different microbes produce varying levels of umami amino acids.
3.3. Changes in free nucleotide contents during fermentation
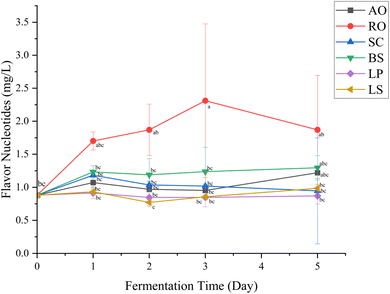
Nucleotides are important flavor-enhancing compounds commonly found in aquatic products, contributing to the umami taste when combined with amino acids.51 Among them, 5′-ribonucleotides—guanosine monophosphate (GMP), inosine monophosphate (IMP), and adenosine monophosphate (AMP)—exhibit strong synergistic effects with L-glutamate, markedly intensifying the umami sensation.52 Differences observed in nucleotide concentrations suggest that the strains varied in their ability to generate or accumulate flavor-active nucleotides during the fermentation of prawn by-product juice (Table 2). Across the fermentation period of five days, all strains except L. plantarum led to a notable increase in the concentration of total umami-active 5′-nucleotides, including IMP, GMP and AMP (Fig. 4). Flavor nucleotides are generated through the metabolic activity of microorganisms, which degrade adenosine triphosphate (ATP) and nucleic acids via enzymes like nucleases (phosphodiesterase) into taste-active 5′-nucleotides.53| AO | RO | SC | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| D0 | D1 | D2 | D3 | D5 | D1 | D2 | D3 | D5 | D1 | D2 | D3 | D5 | |
| a N = 3 fermentation trials.b Mean values ± standard deviation with different lowercase superscript letters within the same row are significantly different (two-way ANOVA, Tukey test, and p < 0.05).c ND (not detected and < limit of quantification).d AO – A. oryzae, RO – R. oligosporus, SC – S. cerevisiae, BS – B. subtilis, LP – L. plantarum, LS – L. sakei. | |||||||||||||
| AMP | 0.36 ± 0.002ab | 0.37 ± 0.02ab | 0.42 ± 0.04ab | 0.41 ± 0.04ab | 0.38 ± 0.04ab | 0.52 ± 0.09ab | 0.49 ± 0.10ab | 0.64 ± 0.26a | 0.55 ± 0.24ab | 0.47 ± 0.01ab | 0.43 ± 0.04ab | 0.40 ± 0.02ab | 0.28 ± 0.22b |
| IMP | 0.40 ± 0.02cd | 0.55 ± 0.04abcd | 0.52 ± 0.05abcd | 0.54 ± 0.06abcd | 0.80 ± 0.09a | 0.54 ± 0.02abcd | 0.53 ± 0.07abcd | 0.57 ± 0.09abcd | 0.76 ± 0.15ab | 0.62 ± 0.02abcd | 0.56 ± 0.03abcd | 0.61 ± 0.01abcd | 0.55 ± 0.45abcd |
| GMP | 0.12 ± 0.04bc | 0.15 ± 0.13bc | 0.04 ± 0.07c | ND | 0.04 ± 0.07c | 0.65 ± 0.05abc | 0.84 ± 0.34ab | 1.10 ± 0.94a | 0.57 ± 0.60abc | 0.09 ± 0.01c | 0.04 ± 0.06c | 0.01 ± 0.01c | 0.11 ± 0.20c |
| Total | 0.88 ± 0.03bc | 1.07 ± 0.08bc | 0.97 ± 0.04bc | 0.95 ± 0.04bc | 1.22 ± 0.09bc | 1.70 ± 0.14abc | 1.87 ± 0.39ab | 2.31 ± 1.16a | 1.87 ± 0.82ab | 1.18 ± 0.02bc | 1.04 ± 0.06bc | 1.02 ± 0.01bc | 0.95 ± 0.80bc |
| BS | LP | LS | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| D1 | D2 | D3 | D5 | D1 | D2 | D3 | D5 | D1 | D2 | D3 | D5 | |
| AMP | 0.45 ± 0.04ab | 0.47 ± 0.05ab | 0.49 ± 0.13ab | 0.43 ± 0.01ab | 0.41 ± 0.03ab | 0.44 ± 0.03ab | 0.43 ± 0.05ab | 0.42 ± 0.08ab | 0.39 ± 0.02ab | 0.40 ± 0.01ab | 0.43 ± 0.03ab | 0.43 ± 0.03ab |
| IMP | 0.55 ± 0.003abcd | 0.51 ± 0.08abcd | 0.53 ± 0.09abcd | 0.63 ± 0.04abc | 0.43 ± 0.07cd | 0.32 ± 0.03cd | 0.31 ± 0.01d | 0.34 ± 0.03cd | 0.45 ± 0.06bcd | 0.36 ± 0.06cd | 0.41 ± 0.04cd | 0.45 ± 0.03bcd |
| GMP | 0.24 ± 0.08bc | 0.20 ± 0.13bc | 0.21 ± 0.18bc | 0.23 ± 0.15bc | 0.08 ± 0.05c | 0.08 ± 0.09c | 0.11 ± 0.11c | 0.12 ± 0.07bc | 0.09 ± 0.08c | 0.01 ± 0.02c | 0.02 ± 0.02c | 0.11 ± 0.08c |
| Total | 1.23 ± 0.09bc | 1.19 ± 0.24bc | 1.24 ± 0.37bc | 1.30 ± 0.18abc | 0.91 ± 0.09bc | 0.85 ± 0.12bc | 0.85 ± 0.14bc | 0.87 ± 0.12bc | 0.93 ± 0.04bc | 0.77 ± 0.07c | 0.86 ± 0.02bc | 0.99 ± 0.04bc |
Among the species tested, R. oligosporus produced the highest total flavor nucleotide content by day 3, suggesting more rapid metabolic turnover during the early stages of fermentation (Fig. 4). A. oryzae, another filamentous fungus known to produce extracellular nucleases, was also among the higher nucleotide producers in this study, although its levels remained below those of R. oligosporus (Table 2).54 This is consistent with previous studies showing that Rhizopus spp. could rapidly colonize substrates and secrete extracellular enzymes during tempeh fermentation.55–57 In addition, R. oligosporus has been reported to produce extracellular ribonucleases, indicating a potential role for RNA degradation in the early accumulation of nucleotides.58 Broader fungal evidence from A. oryzae further supports this, in which S1/P1-type nucleases hydrolyze single-stranded nucleic acids and release mononucleotide products.54 Taken together, these findings suggest that RNA turnover may contribute to the early nucleotide accumulation observed in the fermentation driven by R. oligosporus, although the specific formation pathways of AMP, GMP, and IMP in R. oligosporus have yet to be conclusively established. The elevated levels of IMP and GMP concentration are particularly important, as these compounds interact strongly with umami amino acids, enhancing the overall taste intensity.59
However, the total flavor nucleotides in fermented prawn by-product juice by S. cerevisiae and L. plantarum remained essentially unchanged over the 5-day fermentation period, while those by B. subtilis and L. sakei showed only a small overall increase that is not significantly different from the sample at day 0 (p > 0.05). The limited accumulation of flavor nucleotides in these microbial fermentations may reflect a balance between nucleotide generation and further metabolism, rather than sustained accumulation. In S. cerevisiae, the accumulation of flavor-enhancing nucleotides, is regulated by the purine biosynthetic pathway.60 Ribose-5-phosphate, derived from the pentose phosphate pathway via RKI1, serves as a key precursor for nucleotide formation and is subsequently converted into IMP.61 It can be further metabolized toward GMP or AMP via different enzymatic pathways.62 Similarly, the regulation of IMP, GMP and AMP in B. subtilis is regulated by the purine nucleotide metabolic pathway.63,64 The purine and pyrimidine metabolic pathways not only support the overall synthesis of nucleotides but also enable the conversion between nucleobases, nucleosides, and nucleotides, helping maintain an appropriate balance among these end products.65 In LAB, nucleotide biosynthesis, salvage, and interconversion pathways have been well established, suggesting that released nucleotide-derived compounds may have been rapidly interconverted or taken up by LAB.66 However, the specific mechanisms governing nucleotide accumulation in the present study were not directly measured.
3.4. Umami potential: TAV and EUC analysis
To evaluate the relative contribution of individual compounds to taste perception, the taste activity values (TAVs) and equivalent umami concentrations (EUCs) were calculated (Table 3). The TAVs for 5′-AMP, 5′-IMP and 5′-GMP are not shown as the values are less than 0.01 for all samples. Among the major umami-related compounds, only glutamic acid contributed perceptibly to the overall taste at a concentration above its sensory threshold after fermentation (TAV >1).| EUC (g MSG/100 g) | TAV | |||
|---|---|---|---|---|
| Glu | Asp | |||
| a N = 3 fermentation trials.b Mean values ± standard deviation with different lowercase superscript letters within the same column are significantly different (two-way ANOVA, Tukey test, and p < 0.05).c AO – A. oryzae, RO – R. oligosporus, SC – S. cerevisiae, BS – B. subtilis, LP – L. plantarum, LS – L. sakei. | ||||
| D0 | 0.23 ± 0.04b | 0.12 ± 0.01c | 0.08 ± 0.01h | |
| AO | D1 | 0.27 ± 0.03b | 0.11 ± 0.02c | 0.072 ± 0.003h |
| D2 | 0.66 ± 0.08b | 0.39 ± 0.06c | 0.11 ± 0.01gh | |
| D3 | 1.50 ± 0.31b | 0.94 ± 0.17b | 0.24 ± 0.04bcde | |
| D5 | 2.86 ± 0.69b | 1.33 ± 0.25ab | 0.32 ± 0.04ab | |
| RO | D1 | 0.45 ± 0.06b | 0.10 ± 0.02c | 0.079 ± 0.004h |
| D2 | 1.88 ± 0.50b | 0.40 ± 0.04c | 0.12 ± 0.01fgh | |
| D3 | 6.36 ± 3.71a | 1.24 ± 0.23ab | 0.07 ± 0.02h | |
| D5 | 6.00 ± 2.31a | 1.64 ± 0.52a | 0.27 ± 0.11abcd | |
| SC | D1 | 0.26 ± 0.01b | 0.11 ± 0.01c | 0.09 ± 0.01h |
| D2 | 0.62 ± 0.09b | 0.34 ± 0.02c | 0.10 ± 0.01gh | |
| D3 | 1.59 ± 0.38b | 0.93 ± 0.24b | 0.18 ± 0.02efg | |
| D5 | 2.27 ± 1.74b | 1.14 ± 0.12b | 0.13 ± 0.05fgh | |
| BS | D1 | 0.25 ± 0.02b | 0.090 ± 0.001c | 0.07 ± 0.01h |
| D2 | 0.82 ± 0.35b | 0.35 ± 0.08c | 0.09 ± 0.01h | |
| D3 | 2.48 ± 0.82b | 1.05 ± 0.03b | 0.26 ± 0.03abcd | |
| D5 | 2.80 ± 0.83b | 1.09 ± 0.16b | 0.22 ± 0.02cde | |
| LP | D1 | 0.19 ± 0.03b | 0.097 ± 0.004c | 0.071 ± 0.005h |
| D2 | 0.52 ± 0.11b | 0.33 ± 0.01c | 0.109 ± 0.005gh | |
| D3 | 1.55 ± 0.53b | 0.95 ± 0.17b | 0.22 ± 0.01cde | |
| D5 | 1.95 ± 0.11b | 1.15 ± 0.11b | 0.34 ± 0.02a | |
| LS | D1 | 0.19 ± 0.02b | 0.097 ± 0.004c | 0.064 ± 0.003h |
| D2 | 0.43 ± 0.03b | 0.31 ± 0.02c | 0.112 ± 0.006gh | |
| D3 | 1.46 ± 0.07b | 1.02 ± 0.07b | 0.20 ± 0.01def | |
| D5 | 2.25 ± 0.27b | 1.23 ± 0.04ab | 0.30 ± 0.02abc | |
According to Mau,67 the EUC values can be classified into 4 levels: first level (>10 g MSG per g dry matter), second level (1–10 g MSG per g), third level (0.1–1 g MSG per g), and fourth level (0.1 g MSG per g). The EUC values increased markedly from 0.23 g MSG/100 g at day 0 to 6.36 g MSG/100 g on day 3 in the samples fermented with R. oligosporus. Among the tested microorganisms, fungal fermentations generated higher EUC values; R. oligosporus fermentation yielded the highest EUC value, corresponding to the second-level range, followed by A. oryzae. This corresponds with increases in glutamic acid TAVs, indicating strong synergistic effects between free amino acids and flavor nucleotides in these samples. Notably, the EUC value of the R. oligosporus fermented prawn juice on day 3 was comparable to that reported for fresh whiteleg shrimp meat (6.56 g MSG/100 g),51 but higher than those of Chinese mitten crab (4.2 g MSG/100 g),59 boiled whiteleg shrimp meat (4.11 g MSG/100 g)51 and Chinese shrimp (4.58 g MSG/100 g).68 This suggests that short-term fungal fermentation of prawn heads, a waste stream, can generate ‘meat-like’ levels of umami intensity on par with, or even greater than, those found in fresh seafood.
Phat et al.69 reported a significant positive correlation between EUC values, electronic tongue sensory scores, and human sensory evaluations, supporting the use of EUC as a reliable indicator of umami intensity. The elevated EUC values observed in this study, therefore, demonstrate that microbial fermentation effectively enhances the umami characteristics of prawn by-products, transforming them into flavor-rich substrates.
4. Conclusion
This study demonstrates that microbial fermentation is an effective strategy to enhance the umami characteristics of prawn by-products, contributing to their valorization as sustainable flavoring ingredients. Among the tested microorganisms, R. oligosporus showed the highest efficiency in accumulating umami-related compounds, including glutamic acid, aspartic acid, and flavor nucleotides, resulting in the highest EUC. These findings provide a scientific basis for converting prawn processing residues into high-value, flavor-enhancing ingredients through microbial biotransformation. This can potentially contribute to the circular economy, by harnessing microbial potential for net-zero waste. Nevertheless, this study was conducted under controlled laboratory conditions without sensory validation, which may limit direct translation to industrial applications. In addition, the interactions among flavor-active compounds were inferred from compositional data rather than confirmed through integrated sensory or metabolic analyses. Future work may focus on process optimization, scale-up feasibility, and comprehensive sensory evaluation to support the development of sustainable, prawn-based flavor enhancers for commercial applications.Author contributions
Cia Min Lim: conceptualization, data curation, formal analysis, investigation, methodology, validation, visualization, writing—original draft. Ying Tong Yeo: methodology, validation, writing—review & editing. Cherie Chin: methodology, validation, writing—review & editing. Wei Ning Chen: conceptualization, funding acquisition, project administration, supervision, validation, writing—review & editing.Conflicts of interest
The authors declare that there are no conflicts of interest.Data availability
The manuscript and supplementary information (SI) contain all data necessary to support the findings, statements, and conclusions. Data will be made available on request. Supplementary information is available. See DOI: https://doi.org/10.1039/d5fb00939a.Acknowledgements
We thank Singapore National Research Foundation for funding support (Intra-CREATE programme on Seafood Side-streams Valorization).References
- FAO, GLOBEFISH | Quarterly Shrimp analysis - May 2025, FAO, Rome, Italy, 2025, [cited 2026 Apr 2], Available from: https://openknowledge.fao.org/handle/20.500.14283/cd5727en Search PubMed.
- H. Duppeti, S. Nakkarike Manjabhatta and B. B. Kempaiah, Flavor profile and role of macromolecules in the flavor generation of shrimp meat and valorization of shrimp by-products as a source of flavor compounds: a review, Crit. Rev. Food Sci. Nutr., 2025, 65(1), 123–142, DOI:10.1080/10408398.2023.2268708.
- IMARC, Shrimp Market Size, Share, Growth & Forecast by Environment, Species, Shrimp Size, Distribution Channel, and Region, 2026-2034, International Mining and Resources Conference, 2025 Search PubMed.
- N. Rossi, C. Grosso and C. Delerue-Matos, Shrimp Waste Upcycling: Unveiling the Potential of Polysaccharides, Proteins, Carotenoids, and Fatty Acids with Emphasis on Extraction Techniques and Bioactive Properties, Mar. Drugs, 2024, 22(4), 153, DOI:10.3390/md22040153.
- S. Wu, M. Zhao, S. Gao, Y. Xu, X. Zhao and M. Liu, et al., Change Regularity of Taste and the Performance of Endogenous Proteases in Shrimp (Penaens vannamei) Head during Autolysis, Foods, 2021, 10(5), 5, DOI:10.3390/foods10051020.
- S. S. Sawant, H. Y. Park, E. Y. Sim, H. S. Kim and H. S. Choi, Microbial Fermentation in Food: Impact on Functional Properties and Nutritional Enhancement—A Review of Recent Developments, Fermentation, 2025, 11(1), 15, DOI:10.3390/fermentation11010015.
- J. P. Tamang, P. D. Cotter, A. Endo, N. S. Han, R. Kort and S. Q. Liu, et al., Fermented foods in a global age: East meets West, Compr. Rev. Food Sci. Food Saf., 2020, 19(1), 184–217, DOI:10.1111/1541-4337.12520.
- C. Voidarou, M. Antoniadou, G. Rozos, A. Tzora, I. Skoufos and T. Varzakas, et al., Fermentative Foods: Microbiology, Biochemistry, Potential Human Health Benefits and Public Health Issues, Foods, 2021, 10(1), 69, DOI:10.3390/foods10010069.
- Y. Kong, L. L. Zhang, Y. Y. Zhang, B. G. Sun, Y. Sun and J. Zhao, et al., Evaluation of non-volatile taste components in commercial soy sauces, Int. J. Food Prop., 2018, 21(1), 1854–1866, DOI:10.1080/10942912.2018.1497061.
- M. N. G. Amin, J. Kusnadi, J. L. Hsu, R. J. Doerksen and T. C. Huang, Identification of a novel umami peptide in tempeh (Indonesian fermented soybean) and its binding mechanism to the umami receptor T1R, Food Chem., 2020, 333, 127411, DOI:10.1016/j.foodchem.2020.127411.
- G. M. Walker and G. G. Stewart, Saccharomyces cerevisiae in the Production of Fermented Beverages, Beverages, 2016, 2(4), 30, DOI:10.3390/beverages2040030.
- X. Gao, R. Fan, L. Tan and J. Fu, Recent advances in the applications of Bacillus subtilis in the fermentation of food: Improving product quality and enhancing biological activity, Food Res. Int., 2025, 221, 117427, DOI:10.1016/j.foodres.2025.117427.
- W. S. U. Roland, L. Pouvreau, J. Curran, F. van de Velde and P. M. T. de Kok, Flavor Aspects of Pulse Ingredients, Cereal Chem., 2017, 94(1), 58–65, DOI:10.1094/CCHEM-06-16-0161-FI.
- Y. Lee, Y. J. So, W. H. Jung, T. R. Kim, M. Sohn and Y. J. Jeong, et al., Lactiplantibacillus plantarum LM1001 Improves Digestibility of Branched-Chain Amino Acids in Whey Proteins and Promotes Myogenesis in C2C12 Myotubes, Food Sci. Anim. Resour., 2024, 44(4), 951–965, DOI:10.5851/kosfa.2024.e38.
- A. Tsuji, M. Kozawa, K. Tokuda, T. Enomoto and T. Koyanagi, Robust Domination of Lactobacillus sakei in Microbiota During Traditional Japanese Sake Starter Yamahai-Moto Fermentation and the Accompanying Changes in Metabolites, Curr. Microbiol., 2018, 75(11), 1498–1505, DOI:10.1007/s00284-018-1551-8.
- M. J. Kim, J. Y. Jeong, H. S. Kim, I. M. Hwang, W. Jun and J. H. Lee, Transcriptomic analysis revealed the adaptations of Latilactobacillus sakei to the cold conditions of kimchi fermentation, Food Res. Int., 2025, 217, 116742, DOI:10.1016/j.foodres.2025.116742.
- X. Li, G. Zhao, Y. Zheng, Y. Wang, X. Bai and F. Li, et al., Effects of single fermentation of Lactobacillus sakei and compound fermentation with Staphylococcus carnosus on the metabolomics of beef sausages, Food Chem., 2025, 464, 141728, DOI:10.1016/j.foodchem.2024.141728.
- M. Zagorec and M. C. Champomier-Vergès, Lactobacillus sakei: A Starter for Sausage Fermentation, a Protective Culture for Meat Products, Microorganisms, 2017, 5(3), 56, DOI:10.3390/microorganisms5030056.
- D. Hildebrand, D. Rubin, R. Hadi and T. Kramer, Flavor Fatigue: Cognitive Depletion Influences Consumer Enjoyment of Complex Flavors, J. Consum. Psychol., 2021, 31(1), 103–111, DOI:10.1002/jcpy.1167.
- R. Fondberg, J. N. Lundström, M. Blöchl, M. J. Olsson and J. Seubert, Multisensory flavor perception: The relationship between congruency, pleasantness, and odor referral to the mouth, Appetite, 2018, 125, 244–252, DOI:10.1016/j.appet.2018.02.012.
- J. Luo, D. Frank, A. L. Peterson, B. Nijagal and J. Arcot, Metabolomic characterisation and flavour profiles of prawn, scallop, squid, barramundi, Salmon, snapper, and tuna, Food Chem.:X, 2025, 26, 102284, DOI:10.1016/j.fochx.2025.102284.
- V. Kemsawasd, W. Karnpanit, S. Thangsiri, P. Wongputtisin, A. Kanpiengjai and C. Khanongnuch, et al., Efficient recovery of functional biomolecules from shrimp (Litopenaeus vannamei) processing waste for food and health applications via a successive co-culture fermentation approach, Curr. Res. Food Sci., 2024, 9, 100850, DOI:10.1016/j.crfs.2024.100850.
- J. Tkaczewska, P. Kulawik, E. Jamróz, M. Čagalj, R. F. Matas and V. Šimat, Valorisation of prawn/shrimp shell waste through the production of biologically active components for functional food purposes, J. Sci. Food Agric., 2024, 104(2), 707–715, DOI:10.1002/jsfa.12969.
- J. Chen, H. Lv, J. Li, Q. Zhang, S. Benjakul and Y. Zhang, et al., Upcycling shrimp processing by-products into dual-functional peptides: Insights into bioactivity and flavor, Food Chem., 2025, 493, 145917, DOI:10.1016/j.foodchem.2025.145917.
- J. Ruangwicha, B. Cheirsilp and W. Suyotha, Green biorefinery of shrimp shell waste for α-chitin and high-value co-products through successive fermentation by co-lactic acid bacteria and proteolytic fungus, Bioresour. Technol., 2024, 393, 130106, DOI:10.1016/j.biortech.2023.130106.
- A. M. Sixto-Berrocal, M. Vázquez-Aldana, S. P. Miranda-Castro, M. A. Martínez-Trujillo and M. R. Cruz-Díaz, Chitin/chitosan extraction from shrimp shell waste by a completely biotechnological process, Int. J. Biol. Macromol., 2023, 230, 123204, DOI:10.1016/j.ijbiomac.2023.123204.
- N. Rossi, C. Grosso and C. Delerue-Matos, Shrimp Waste Upcycling: Unveiling the Potential of Polysaccharides, Proteins, Carotenoids, and Fatty Acids with Emphasis on Extraction Techniques and Bioactive Properties, Mar. Drugs, 2024, 22(4), 153, DOI:10.3390/md22040153.
- N. V. Nguyen, P. D. Hai, V. T. My My, D. T. Men, L. D. Trung and H. J. Bavor, Improving Product Added-value from Shrimp (Litopenaeus vannamei) Waste by Using Enzymatic Hydrolysis and Response Surface Methodology, J. Aquat. Food Prod. Technol., 2021, 30(7), 880–892, DOI:10.1080/10498850.2021.1949770.
- J. wei Luo, Li Z. bing, S. Xiao, X. Chen, X. Y. Yang and W. J. hui, Metabolomic analysis of microbial effects on quality and metabolite conversion in rapidly fermented salt-free shrimp by-product sauce, Food Chem.:X, 2025, 32, 103344, DOI:10.1016/j.fochx.2025.103344.
- S. Studzińska and E. Zalesińska, Development of an ultra high performance liquid chromatography method for the separation and determination of nucleotides and nucleosides in extracts from infant milk formulas and human milk samples, J. Food Compos. Anal., 2023, 115, 104949, DOI:10.1016/j.jfca.2022.104949.
- M. Pastor-Belda, I. Fernández-Caballero, N. Campillo, N. Arroyo-Manzanares, M. Hernández-Córdoba and P. Viñas, Hydrophilic interaction liquid chromatography coupled to quadrupole-time-of-flight mass spectrometry for determination of nuclear and cytoplasmatic contents of nucleotides, nucleosides and their nucleobases in food yeasts, Talanta Open, 2021, 4, 100064, DOI:10.1016/j.talo.2021.100064.
- H. Schlichtherle-Cerny and W. Grosch, Evaluation of taste compounds of stewed beef juice, Z. Lebensm. Unters. Forsch. A, 1998, 207(5), 369–376, DOI:10.1007/s002170050347.
- S. Yamaguchi, T. Yoshikawa, S. Ikeda and T. Ninomiya, MEASUREMENT OF THE RELATIVE TASTE INTENSITY OF SOME L-α-AMINO ACIDS AND 5′-NUCLEOTIDES, J. Food Sci., 1971, 36(6), 846–849, DOI:10.1111/j.1365-2621.1971.tb15541.x.
- T. Bintsis, Lactic acid bacteria as starter cultures: An update in their metabolism and genetics, AIMS Microbiol., 2018, 4(4), 665–684, DOI:10.3934/microbiol.2018.4.665.
- E. Erem and M. Kilic-Akyilmaz, The role of fermentation with lactic acid bacteria in quality and health effects of plant-based dairy analogues, Compr. Rev. Food Sci. Food Saf., 2024, 23(4), e13402, DOI:10.1111/1541-4337.13402.
- Y. A. Mennah-Govela, R. P. Singh and G. M. Bornhorst, Buffering capacity of protein-based model food systems in the context of gastric digestion, Food Funct., 2019, 10(9), 6074–6087, 10.1039/C9FO01160A.
- S. Ruiz-Moyano, A. Martín, M. J. Benito, A. Hernández, R. Casquete and M. de Guia Córdoba, Application of Lactobacillus fermentum HL57 and Pediococcus acidilactici SP979 as potential probiotics in the manufacture of traditional Iberian dry-fermented sausages, Food Microbiol., 2011, 28(5), 839–847, DOI:10.1016/j.fm.2011.01.006.
- X. Zhao, Y. Wei, X. Gong, H. Xu and G. Xin, Evaluation of umami taste components of mushroom (Suillus granulatus) of different grades prepared by different drying methods, Food Sci. Hum. Wellness, 2020, 9(2), 192–198, DOI:10.1016/j.fshw.2020.03.003.
- Y. Zhu, Y. Luo, P. Wang, M. Zhao, L. Li and X. Hu, et al., Simultaneous determination of free amino acids in Pu-erh tea and their changes during fermentation, Food Chem., 2016, 194, 643–649, DOI:10.1016/j.foodchem.2015.08.054.
- S. A. Khan, L. Liu, T. Lai, R. Zhang, Z. Wei and J. Xiao, et al., Phenolic profile, free amino acids composition and antioxidant potential of dried longan fermented by lactic acid bacteria, J. Food Sci. Technol., 2018, 55(12), 4782–4791, DOI:10.1007/s13197-018-3411-8.
- D. Wang, F. Cheng, Y. Wang, J. Han, F. Gao and J. Tian, et al., The Changes Occurring in Proteins during Processing and Storage of Fermented Meat Products and Their Regulation by Lactic Acid Bacteria, Foods, 2022, 11(16), 2427, DOI:10.3390/foods11162427.
- Z. Liu, Q. Liu, D. Zhang, S. Wei, Q. Sun and Q. Xia, et al., Comparison of the Proximate Composition and Nutritional Profile of Byproducts and Edible Parts of Five Species of Shrimp, Foods, 2021, 10(11), 2603, DOI:10.3390/foods10112603.
- Z. Y. Ter, L. S. Chang, A. S. Babji, N. A. M. Zaini, S. Fazry and S. R. Sarbini, et al., A review on proteolytic fermentation of dietary protein using lactic acid bacteria for the development of novel proteolytically fermented foods, Int. J. Food Sci. Technol., 2024, 59(3), 1213–1236, DOI:10.1111/ijfs.16888.
- J. Y. Lim, Y. J. Choi, H. Yu, J. Y. Choi, J. H. Yang and Y. B. Chung, et al., Investigation of Metabolite Differences in Salted Shrimp Varieties during Fermentation, ACS Omega, 2023, 8(50), 47735–47745, DOI:10.1021/acsomega.3c06046.
- J. Tong, G. Li, X. Liu and X. Zheng, Insight into the formation mechanism of umami taste in traditional Chinese soybean paste: A review, LWT–Food Sci. Technol., 2025, 224, 117896, DOI:10.1016/j.lwt.2025.117896.
- O. L. Tavano, A. Berenguer-Murcia, F. Secundo and R. Fernandez-Lafuente, Biotechnological Applications of Proteases in Food Technology, Compr. Rev. Food Sci. Food Saf., 2018, 17(2), 412–436, DOI:10.1111/1541-4337.12326.
- A. Usman, S. Mohammed and J. Mamo, Production, Optimization, and Characterization of an Acid Protease from a Filamentous Fungus by Solid-State Fermentation, Int. J. Microbiol., 2021, 2021(1), 6685963, DOI:10.1155/2021/6685963.
- A. Razzaq, S. Shamsi, A. Ali, Q. Ali, M. Sajjad and A. Malik, et al., Microbial Proteases Applications, Front. Bioeng. Biotechnol., 2019, 7, 110, DOI:10.3389/fbioe.2019.00110.
- O. P. Ward, 3.49 - Proteases, in Comprehensive Biotechnology, ed. M. Moo-Young, Oxford, Pergamon, 3rd edn, 2011, pp. 604–615, DOI:10.1016/B978-0-444-64046-8.00187-7..
- S. S. Sawant, H. Y. Park, E. Y. Sim, H. S. Kim and H. S. Choi, Microbial Fermentation in Food: Impact on Functional Properties and Nutritional Enhancement—A Review of Recent Developments, Fermentation, 2025, 11(1), 15, DOI:10.3390/fermentation11010015.
- H. Duppeti, S. N. Manjabhatta, A. Martin and B. Bheemanakere Kempaiah, Effects of different processing methods on the biochemical composition, color and non-volatile taste active compounds of whiteleg shrimp (Litopenaeus vannamei), Food Chem. Adv., 2022, 1, 100118, DOI:10.1016/j.focha.2022.100118.
- S. Yamaguchi and K. Ninomiya, Umami and Food Palatability, J. Nutr., 2000, 130(4), 921S–926S, DOI:10.1093/jn/130.4.921S.
- K. Zhang, T. T. Zhang, R. R. Guo, Q. Ye, H. L. Zhao and X. H. Huang, The regulation of key flavor of traditional fermented food by microbial metabolism: A review, Food Chem.:X, 2023, 19, 100871, DOI:10.1016/j.fochx.2023.100871.
- T. Kovaľ, L. H. Østergaard, J. Lehmbeck, A. Nørgaard, P. Lipovová and J. Dušková, et al., Structural and Catalytic Properties of S1 Nuclease from Aspergillus oryzae Responsible for Substrate Recognition, Cleavage, Non–Specificity, and Inhibition, PLoS One, 2016, 11(12), e0168832, DOI:10.1371/journal.pone.0168832.
- J. C. de Reu, R. Marcel ten Wolde, J. de Groot, M. J. R. Nout, F. M. Rombouts and H. Gruppen, Protein Hydrolysis during Soybean Tempe Fermentation with Rhizopus oligosporus, J. Agric. Food Chem., 1995, 43(8), 2235–2239, DOI:10.1021/jf00056a050.
- L. A. Fernandez Castaneda, J. Auer, M. Langton and G. Zamaratskaia, Rhizopus spp. and the Evolution of Tempeh from Tradition to Modern Uses, in Fungi, ed. P. Navarro Simarro, New York, NY, Springer US, 2026, pp. 277–300, DOI:10.1007/978-1-0716-4989-3_20.
- Y. Zhang, R. Wei, F. Azi, L. Jiao, H. Wang and T. He, et al., Solid-state fermentation with Rhizopus oligosporus RT-3 enhanced the nutritional properties of soybeans, Front. Nutr., 2022, 9 DOI:10.3389/fnut.2022.972860.
- D. G. Glitz and E. A. Woodroof, Purification and characterization of two extracellular ribonucleases from Rhizopus oligosporus, Biochemistry, 1971, 10(9), 1532–1540, DOI:10.1021/bi00785a004.
- D. W. Chen and M. Zhang, Non-volatile taste active compounds in the meat of Chinese mitten crab (Eriocheir sinensis), Food Chem., 2007, 104(3), 1200–1205, DOI:10.1016/j.foodchem.2007.01.042.
- B. Daignan-Fornier and B. Pinson, Yeast to Study Human Purine Metabolism Diseases, Cells, 2019, 8(1), 67, DOI:10.3390/cells8010067.
- S. C. Jung, H. Oh, W. Eom, Y. S. Jin, S. H. Park, K. Park, et al., Scarless Genetic Engineering of Saccharomyces cerevisiae for Enhanced Guanosine Monophosphate Production as a Natural Flavor Enhancer, 35(12), pp. 1–9, DOI:10.4014/jmb.2508.08034.
- M. Escobar-Henriques and B. Daignan-Fornier, Transcriptional Regulation of the Yeast GMP Synthesis Pathway by Its End Products, J. Biol. Chem., 2001, 276(2), 1523–1530, DOI:10.1074/jbc.M007926200.
- A. Deng, Q. Qiu, Q. Sun, Z. Chen, J. Wang and Y. Zhang, et al., In silico-guided metabolic engineering of Bacillus subtilis for efficient biosynthesis of purine nucleosides by blocking the key backflow nodes, Biotechnol. Biofuels Bioprod., 2022, 15(1), 82, DOI:10.1186/s13068-022-02179-x.
- K. Ishii and I. Shiio, Regulation of Purine Nucleotide Biosynthesis in Bacillus subtilis, Agric. Biol. Chem., 1973, 37(2), 287–300, DOI:10.1080/00021369.1973.10860683.
- P. O. Ljungdahl and B. Daignan-Fornier, Regulation of Amino Acid, Nucleotide, and Phosphate Metabolism in Saccharomyces cerevisiae, Genetics, 2012, 190(3), 885–929, DOI:10.1534/genetics.111.133306.
- M. Kilstrup, K. Hammer, P. Ruhdal Jensen and J. Martinussen, Nucleotide metabolism and its control in lactic acid bacteria, FEMS Microbiol. Rev., 2005, 29(3), 555–590, DOI:10.1016/j.fmrre.2005.04.006.
- J. L. Mau, The Umami Taste of Edible and Medicinal Mushrooms, Int. J. Med. Mushrooms, 2005, 7(1 & 2), 119–126, DOI:10.1615/IntJMedMushr.v7.i12.120.
- L. Chen, W. Zeng, Y. Rong and B. Lou, Characterisation of taste-active compositions, umami attributes and aroma compounds in Chinese shrimp, Int. J. Food Sci. Technol., 2021, 56(12), 6311–6321, DOI:10.1111/ijfs.15304.
- C. Phat, B. Moon and C. Lee, Evaluation of umami taste in mushroom extracts by chemical analysis, sensory evaluation, and an electronic tongue system, Food Chem., 2016, 192, 1068–1077, DOI:10.1016/j.foodchem.2015.07.113.
| This journal is © The Royal Society of Chemistry 2026 |