Open Access Article
Open Access ArticleChitosan based ecofriendly nanoencapsulation formulation of Clitoria ternatea extract coated bacterial probiotic strain Lactobacillus acidophilus: a sustainable approach to improve probiotic functionality, stability, and tolerance
M. Lavanya and
S. Karthick Raja Namasivayam *
*
Centre for Applied Research, Saveetha School of Engineering, Saveetha Institute of Medical and Technical Sciences (SIMATS), Chennai-602105, Tamil Nadu, India. E-mail: biologiask@gmail.com; Tel: +91 9842441198
First published on 18th February 2026
Abstract
In this study, a sustainable nanoencapsulation formulation of Clitoria ternatea extract-coated bacterial probiotic strain Lactobacillus acidophilus using nanoscale chitosan (CT-CS-LA-NC) was developed to enhance probiotic functionality, stability, and tolerance. The structural and functional characteristics of the nanocapsules were analyzed using scanning electron microscopy (SEM), Fourier transform infrared spectroscopy (FTIR), and thermogravimetric analysis (TGA). These analyses confirmed that the CT-CS-LA-NC nanocapsules were nanoscale in size, highly thermostable, and exhibited strong interactions among the polymer matrix, bacterial cell surface components, and plant extract. The encapsulation efficiency was found to be 98.43%, indicating effective preservation of viable probiotic cells during the encapsulation process. The antibacterial activity of CT-CS-LA-NC demonstrated significant inhibition against Escherichia coli and Bacillus subtilis. Antioxidant assessment revealed a free radical scavenging activity of 48.2 ± 1.32%. Under simulated gastric conditions (pH 2.5 and 3.5), the survivability of free and nanoencapsulated L. acidophilus showed notable differences after 36 hours. The encapsulated cells exhibited a higher survival rate of 60.93 ± 0.68% at pH 2.5, compared to only 21.29 ± 0.13% for free cells. Similarly, under bile tolerance conditions, encapsulated cells showed a survivability of 39.46 ± 0.68% at pH 2.5. Storage stability studies at 4 °C over 28 days revealed a rapid decline in the viability of free L. acidophilus cells, whereas CT-CS-LA-NC encapsulated cells maintained 94% viability after 7 days and 52% after 28 days. Molecular docking analysis using CB-Dock demonstrated strong binding affinities between anthocyanin, lactic acid, and chitosan ligands and gut-associated proteins such as O-GlcNAcase (7K41) and S-layer associated protein (SlpA) (8AE1), showing binding energies of −9.4 and −9.6 kcal mol−1, respectively. Overall, the biocompatible and environmentally safe CT-CS-LA-NC formulation significantly improved probiotic stability during storage, enhanced survivability under harsh gastrointestinal conditions, and prolonged probiotic shelf life. These findings highlight its potential application as a sustainable ingredient for functional foods and nutraceutical formulations.
Sustainability spotlightThis research introduces a sustainable and biocompatible nanoencapsulation strategy using chitosan and Clitoria ternatea extract to enhance the functionality, stability, and survivability of the probiotic strain Lactobacillus acidophilus. The developed CT-CS-LA-NC system integrates natural, renewable materials to improve probiotic tolerance under acidic and bile conditions, strengthen antibacterial and antioxidant properties, and extend storage stability without synthetic additives. By leveraging green biopolymers and plant-derived bioactives, this study contributes to the advancement of environmentally responsible probiotic delivery systems. The approach aligns with sustainable food innovation goals by promoting resource-efficient, safe, and functional probiotic formulations suitable for next-generation functional foods and nutraceuticals. |
1. Introduction
Food spoilage is a significant issue affecting the economy and brand reputation of manufacturers. The food preservation and processing industry also seeks alternatives to conventional preservatives.1 The food industry is utilizing probiotic microorganisms, specifically lactic acid bacteria and bifidobacteria, as bio-preservatives to create healthy and safe food products.2 These probiotics act as bio-preservatives, balancing beneficial and harmful bacteria in plant-based foods.3,4 The FDA and WHO define probiotics as “live microorganisms which when administered in adequate amounts confer a health benefit on the host,” which includes genera like Saccharomyces, Bifidobacterium, Lactobacillus, Enterococcus, Bacillus, Escherichia, and Streptococcus.5,6Probiotics compete with harmful bacteria for adhesion sites, preventing colonization. They strengthen the intestinal barrier, reducing permeability and preventing harmful substances from entering the bloodstream.7 Additionally, probiotics produce antimicrobial substances that inhibit pathogenic bacteria growth and can modulate the immune system to enhance defence mechanisms. Some probiotics also contribute to neurotransmitter synthesis, and influence the gut–brain communication.8 Selecting the right probiotic strain is crucial for specific conditions. The amount consumed, formulation, and storage conditions are crucial for their effectiveness. Probiotics must survive the acidic stomach environment and small intestine bile. Quality control and shelf-life also impact the efficacy of probiotic products. Combining probiotics with prebiotics or other beneficial compounds can enhance their effects.7,9 Among the crucial probiotic characteristics of Lactobacillus acidophilus that have been demonstrated in vitro and belong to the first group are antibiotic production, bile susceptibility, adaptability to low pH, adherence to human colon cells, lactase activity, and stability of the product. The latter group includes the general probiotic advantages observed in feeding studies at the animal level, such as immune response regulation, lowering serum cholesterol, improving lactose metabolism, and preventing or treating infections. When combined with chitosan-based antimicrobial preservatives, these probiotics can significantly enhance shelf life and food safety and maintain the quality of food products, presenting a natural and sustainable alternative to traditional preservation methods.10 Among probiotic bacteria, Lactobacillus acidophilus is one of the more efficient types. Because of its advantageous properties, studies have indicated that it can be added to fermented foods. For Lactobacillus acidophilus to be effective as probiotics, it must adapt to the intestinal environment of the host and be viable and physiologically active at the target site in the host.11
Encapsulated probiotics are used in various products, and they are used to protect cores from degradation, reduce evaporation, improve material handling, and enhance the stability of the active ingredients. These benefits make encapsulated probiotics an essential component in the formulation of dietary supplements and functional foods.12 Encapsulation methods involve trapping or adhering drugs to a matrix or surface, or bonding drugs to the material. Molecular weight and stability play crucial roles in determining their effectiveness and safety. Larger molecules may pose challenges, and interactions between drugs and capsules influence encapsulation. Various capsule production methods have been developed, influenced by factors like particle size, reproducibility, and final product stability.13 Alginate is the most widely used biopolymer for microencapsulation because it is inexpensive, non-toxic, forms gentle matrices with calcium chloride, and easily entraps living microbial cells. It is also a food additive that is widely acknowledged and can be used in food without any harmful effects. Its low stability in the presence of chelating agents and in acidic environments below pH 2 restricts its application.14 Coating alginate beads with chitosan enhances the stability and vitality of encapsulated probiotic microbes. The colon's microbiota breaks down chitosan and solubilizes alginate gel by retaining calcium ions.15
Probiotic foods are also sources of other nutritional compounds, such as antioxidants, fibre, unsaturated fatty acids, minerals, or vitamins and offer synergistic effects to health.16 Clitoria ternatea is a flower rich in antioxidants, flavonoids, and phenolic compounds, with antimicrobial properties. It is used in nanoencapsulation to improve the stability and bioavailability of bioactive compounds, protect sensitive ingredients from oxidation, and enhance controlled nutrient release in food and pharmaceutical applications.17 Compared to synthetic encapsulating agents, C. ternatea offers natural antioxidant protection, higher encapsulation efficiency for anthocyanins, and better bioavailability than conventional microencapsulation techniques.18 Although Clitoria ternatea has been investigated for its antioxidant and antibacterial characteristics, its utilization in chitosan–alginate-based nanoencapsulation for probiotic stabilization and ecotoxicity evaluation is insufficiently reported. In this study, the viability of L. acidophilus with probiotic qualities was examined, along with the isolation and encapsulation of C. ternatea extract coated chitosan-L. acidophilus nanocapsules (CT-CS-LA-NC). The polydispersity index, zeta potential, thermogravimetric analysis (TGA), scanning electron microscopy (SEM), Fourier transform infrared spectroscopy (FTIR), and encapsulation efficiency were used to examine the structural and functional properties followed by the antioxidant properties, gastrointestinal survivability and storage stability of probiotic nanocapsules. The environmental compatibility was evaluated through ecotoxicity profiling and molecular interaction modeling.
2. Materials and methods
2.1 Reagents and chemicals
All reagents and chemicals for nanocapsule preparation and other biological assays including protein profiling study were purchased from Sigma. Hi-Media, India, provided the culture medium used for the microbiological analysis (extra-pure analytical grade). All the chemicals were obtained with high purity and were used without any further purification.2.2 Isolation of Lactobacillus acidophilus from curd
The probiotic strain used in this study was isolated from fresh homemade curd prepared from cow milk obtained from a local milk store in Purasawalkam, Chennai. The sample was collected in a sterile flask, and stored under refrigerated conditions (4 °C). The initial pH of the curd sample was measured to be 4.7. Initially, 10 g of curd sample was homogenised in 90 mL of sterile distilled water to make a 10−1 (w/v) stock suspension. Serial dilutions were performed on this suspension by transferring 1 mL of the preceding dilution into 9 mL of sterile distilled water to generate a range of 10−2 to 10−5. Then, 0.1 mL of each serially diluted sample was poured into already pre-solidified MRS agar and incubated at 37 °C for 24 hours with constant homogeneous shaking under aerobic conditions. The bacterial isolates were purified using the streak plate method and kept for four weeks at 4 °C on agar slants until characterisation. Gram stain and acid production tests were used to further analyse the isolate's cellular morphology and staining properties.192.3 Identification of probiotic strain
The isolated Lactobacillus species is identified using morphological (Gram staining, spore test, and motility test), biochemical (indole production test and catalase test), and molecular characterization (16S rRNA sequencing) for the identification of lactic acid bacteria isolates. Plates with white and creamy colonies (presumptive for LAB) were selected and purified through three successive transfers on MRS medium.20PCR amplicons and a heat cycler were used for the amplification process. The 16S rRNA gene sequence reported in this study was identified with NCBI sequences. After the sequences were aligned, MEGA version 4.0 was used to create a phylogenetic tree.21
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm. The biomass that was obtained from L. acidophilus was centrifuged and rinsed with 0.1 M phosphate-buffered saline. The cells were then resuspended in phosphate buffered saline. The washed cells served as the inoculum source.22
000 rpm. The biomass that was obtained from L. acidophilus was centrifuged and rinsed with 0.1 M phosphate-buffered saline. The cells were then resuspended in phosphate buffered saline. The washed cells served as the inoculum source.222.4 Preparation of C. ternatea extract coated chitosan-L. acidophilus nanocapsules (CT-CS-LA-NC)
C. ternatea (butterfly pea flowers) was purchased from local retail outlets. They were subjected to a pretreatment process by double washing in running tap water and distilled water to remove impurities. Ten grams of butterfly pea flower petals were homogenized and soaked in 100 mL of distilled water. The mixture was then placed on a rotary shaker for 72 hours. After this period, the suspension was gently shaken and filtered through filter paper to remove the solids. The filtered crude extract was concentrated using a rotary evaporator set at 50 °C (TOP-300RE) to a constant weight and the resulting dry mass yield was measured as 5.059 mg mL−1; the concentration was determined by dividing the final mass by the initial volume using a gravimetric method.22 The concentrated extract from the butterfly pea flower petals was collected in screw-cap vials for the formulation of nanocapsules. The qualitative analysis of aqueous C. ternatea flower extract was assessed by the standardized method reported by Chakraborty et al. (2017).23In a typical procedure, nano-encapsulated chitosan and probiotic (Lactobacillus) with plant extract (C. ternatea) was formulated. 25 mL of 1.2% sodium alginate and 1 mL of C. ternatea extract and 0.5 mL of probiotic suspension (CT-CS-LA-NC) were drawn into a 10 mm diameter Pasteur pipette. These were then carefully dispensed drop by drop into a solution containing 100 mL of 10% CaCl2 and 1 mL of Tween 20, forming beads. Droplets were immersed in a calcium chloride solution for 30 minutes. The beads were retrieved, rinsed twice with sterile water, and blotted in a Petri dish to remove any remaining water, resulting in surface-dried beads.24
2.5 Encapsulation efficiency
The encapsulation efficiency (EE) of coating materials refers to their ability to hold or encapsulate core materials within the capsule, and it is a key feature of wall materials. After the CT-CS-LA-NC beads dissolved, the encapsulated probiotic cells were counted directly to determine the encapsulation efficiency (EE). 1 mL of the initial probiotic mixture was plated onto MRS agar after being serially diluted in sterile phosphate-buffered saline (PBS, pH 7.2). After 48 hours of incubation at 37 °C, the initial cell concentration (Ni) was calculated by counting the colony-forming units (CFU) on the plates. For the encapsulated cells, 1 g of CT-CS-LA-NC beads was homogenized using a sterile glass homogenizer and added to 9 mL of sterile PBS, and then 1 mL of the solution was serially diluted in sterile PBS and plated on MRS agar. The colonies were counted after 48 hours of incubation at 37 °C to determine the final cell concentration (Nf).25 The encapsulation efficiency was calculated using the following formula:| EE (%) = (Nf/Ni) × 100 |
2.6 Characterization of CT-CS-LA-NC
The structural and functional properties of the nanocapsule were determined by using appropriate analytical techniques. The findings provided essential insights into the capsule's stability and release mechanisms, facilitating prospective uses in preservation. The data indicate that the nanocapsule may improve efficacy while minimizing side effects.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 with distilled water before analysis. Measurements were conducted in triplicate at a regulated temperature of 25 ± 2 °C, with the detector oriented at a 90° scattering angle. The Z-average diameter (mean particle size) and PDI were determined by averaging the measurements done in triplicate.26
100 with distilled water before analysis. Measurements were conducted in triplicate at a regulated temperature of 25 ± 2 °C, with the detector oriented at a 90° scattering angle. The Z-average diameter (mean particle size) and PDI were determined by averaging the measurements done in triplicate.26![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000×.
000×.2.7 Antioxidant activity
| % DPPH free radical scavenging activity = [(A0 − A1)/(A0)] |
2.8 Antibacterial activity
Antibacterial activity of the nanocapsule was tested against Bacillus subtilis (ATCC 6051) and Escherichia coli (ATCC 25922) adopting a well diffusion assay. All the experiments were carried out with free L. acidophilus, nano-encapsulated L. acidophilus, and secretors of free L. acidophilus and nano-encapsulated L. acidophilus separately. Inocula of respective treatments were prepared in MRS broth incubated at 37 °C for 18 hours. The secretory product of L. acidophilus was obtained from the culture supernatant of the respective treatment group (inoculated MRS broth centrifuged at 8000 rpm for 10 min); the collected supernatant was syringe filtered and used as the source for antibacterial activity. Inocula of the respective bacterial cultures prepared in MRS broth were uniformly spread on the sterile MRS agar media. Wells were made on the swabbed agar using a sterile gel puncture (8 mm). 100 µL of viable inocula of L. acidophilus thus prepared was added to the wells. For 24 hours, the seeded plates were incubated at 37 °C.29 The plates were examined for zones of inhibition following the incubation period.2.9 Evaluation of acid tolerance
The modified acid tolerance test was performed as described previously by Ehrmann et al. (2002).30 Acid stress condition was created by adjusting the pH of the MRS broth to 2.5 using 1 N HCl in addition. The inoculum thus prepared was added to 100 mL of sterile MRS broth in various conical flasks at concentrations (v/v) ranging from 0.5 mL to 2.5 mL, respectively, and incubated at 37 °C under shaking conditions. Additionally, the CT-CS-LA-NC was suspended in the MRS broth at concentrations (w/v) varying from 0.5 g to 2.5 g and incubated for 48 hours. These treatment groups were observed and compared against a control group. The turbidity of the culture media was monitored at several time intervals (12 hours, 24 hours, and 36 hours). After the incubation period, the optical densities (OD 570 nm) were recorded at 12-hour intervals for 48 hours. The negative control, which was only the media, showed no growth. The same procedure was repeated for all the treatment and control groups under the same experimental conditions, respectively. All tests were carried out in triplicate.2.10 Evaluation of bile tolerance
A previously adapted technique by Akmal et al. (2022)31 was used to assess bile tolerance. The simulated bile conditions were prepared by adding 0.3 g in 100 mL (0.3% w/v) of MRS broth and using a 1 M sodium hydroxide solution to adjust the pH to 7.5. After preparation, 100 mL of sterile MRS broth in various conical flasks were filled with the inoculum at concentrations ranging from 0.5% (v/v) to 2.5% (v/v), respectively, and shaken while being incubated at 37 °C. After being dissolved in the MRS broth at concentrations varying from 0.5% (w/v) to 2.5% (w/v), the CT-CS-LA-NC was incubated for 36 hours. These treatment groups were observed and contrasted with the control group. There was no growth in the negative control, which consisted solely of the media. Then, a spectrophotometer was used to measure the track bacterial growth at 560 nm. Every experiment was run in triplicate.2.11 Storage stability
The stability and storage conditions of the free probiotic and CT-CS-LC-NC were comparatively evaluated over 28 days, via a method adapted from Thinkohkaew et al. (2024).32 Both the encapsulated alginate beads and the free probiotic cell suspension were stored in sterile conical tubes containing 0.5% sterile sodium chloride (NaCl) solution at 4 °C. The viability of the probiotics in the free suspension and the beads was assessed at different time periods, including 7, 14, 21, and 28 days. The encapsulated samples were dissolved in NaCl to release cells. After serial dilutions and the dot-plating method, the viable cell count was measured on the MRS agar plates. The plates were incubated at 37 °C for 48 hours before colony-forming units (CFU) were counted. The survival rate for each sample at each time point was then determined and compared to the initial viable cell count to determine the probiotic viability under refrigerated conditions.2.12 Ecotoxicity assessment of CT-CS-LA-NC
The ecotoxic effect of the fabricated nanocapsule was determined by measuring the phytotoxic effect and soil parameters.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio. Protein movement in the gel was observed by adding Bromophenol Blue (BPB) as a tracking dye to the sample buffer. The control and treated samples' crude protein extracts were dissolved in sodium phosphate buffer (pH 7.5) and centrifuged for three minutes at 10
1 ratio. Protein movement in the gel was observed by adding Bromophenol Blue (BPB) as a tracking dye to the sample buffer. The control and treated samples' crude protein extracts were dissolved in sodium phosphate buffer (pH 7.5) and centrifuged for three minutes at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm. Soluble proteins were separated using stacking and linear gradient acrylamide separating gels at a constant current until the tracking dye reached the separating gel's edge. Coomassie Brilliant Blue R-250 (CBBR) staining was applied to the gels. 0.05 M Tris–glycine buffer was used for the electrophoresis, which was carried out at 10 °C (pH 8.3) until the sample bands crossed the stacking layer. The protein bands were visualized.
000 rpm. Soluble proteins were separated using stacking and linear gradient acrylamide separating gels at a constant current until the tracking dye reached the separating gel's edge. Coomassie Brilliant Blue R-250 (CBBR) staining was applied to the gels. 0.05 M Tris–glycine buffer was used for the electrophoresis, which was carried out at 10 °C (pH 8.3) until the sample bands crossed the stacking layer. The protein bands were visualized.2.13 In silico docking
The 3D structures of the proteins were obtained from the RCSB PDB database (https://www.rcsb.org/pdb) using the following PDB IDs: 7K41, 8AE1. The ligand molecules anthocyanin (PubChem CID: 154824275), chitosan (PubChem CID: 441477), and lactic acid (PubChem CID: 612) were obtained from the PubChem website (https://pubchem.ncbi.nlm.nih.gov). The target proteins' optimal protein–ligand interaction sites were found using CB-Dock software, which also computes the target proteins' size and shape using a curvature-based cavity identification technique and performs docking using AutoDock Vina software. Based on Vina scores, CB-Dock evaluates the binding modes and provides an interactive three-dimensional model of them.352.14 Statistical analysis
All data were obtained from experiments carried out at least in duplicate. The results were compared using one-way analysis of variance (ANOVA) followed by Duncan's post hoc (Prism software version 5.0). Results were considered significant when P < 0.05.3. Results and discussion
3.1 Isolation and identification of L. acidophilus
The probiotic strain L. acidophilus used in this study was isolated from the curd by a culture-dependent method. The serially diluted curd sample was spread plated on sterile MRS agar followed by incubation under optimum conditions (identification of the isolate was done by morphological and biochemical properties shown in Table 1). The L. acidophilus was isolated from curd. The selected bacterial strain was identified by morphological and biochemical characterization as L. acidophilus. Morphologically, the single colonies were circular and creamy white. The bacteria were Gram-positive rods and cocci-shaped, and were further observed under a phase contrast microscope as shown in Fig. S1. This study revealed Gram-positive, non-spore-forming rods showing a negative result (red colour): all the isolated strains were found to be catalase-negative (no bubble formation) and indole test-negative bacteria. This is in agreement with the study conducted by Rozos et al. (2018).36 Thus, the results obtained coincided with L. acidophilus strain characteristics.| Characterization | Result |
|---|---|
| Gram staining | Gram positive |
| Spore test | Non-spore forming rods |
| Motility test | Non-motile |
| Indole production test | Negative |
| Catalase test | Negative |
The 16S rRNA gene sequence of isolated strains showed 96% similarity to L. acidophilus. All isolated strains corresponding to Lactobacillus were considered to be the best match based on top hit similarity.37 The sequence has been deposited in the NCBI GenBank database under accession number PX470079. The phylogenetic trees were designed in the bootstrap neighbor-joining (NJ) method (Fig. S2) based on 16S rRNA gene sequencing.
3.2 Preparation of C. ternatea extract coated chitosan-L. acidophilus nanocapsules (CT-CS-LA-NC)
C. ternatea extract-coated chitosan-L. acidophilus nanocapsules were prepared by the simple in situ principle of C. ternatea, which was readily extracted in distilled water under ambient conditions, and the final yield of the metabolite was found to be 1.029 g L−1. The extracted metabolite was stored in screw cap vials and used for further studies. Phytochemical screening of C. ternatea leaves indicated the presence of active components such as alkaloids, flavonoids, tannins, phenols, and saponins, with steroids being absent, as shown in Table 2. In a related study, Neda et al. (2013)38 detailed the phytochemical compounds and nutritional composition found in C. ternatea flowers.| S. no. | Phytochemical analysis | Presence/absence |
|---|---|---|
| 1 | Alkaloids | + |
| 2 | Flavonoids | + |
| 3 | Steroids | − |
| 4 | Phenol | + |
| 5 | Tannin | + |
| 6 | Saponin | − |
A simple in situ dispersion method was employed for the preparation of nanocapsules. Opaque probiotic nanocapsules with a sphere-like structure (Fig. 1) and highly stable characteristics were fabricated using C. ternatea extract, chitosan polymer, and L. acidophilus. Synthesized CT-CS-LA-NC was characterised by techniques like scanning electron microscopy, Fourier transform infrared spectroscopy, and thermogravimetric analysis. Among the various substances, chitosan is an important polymer used for the encapsulation of probiotic-stabilised nanocapsules, revealing enhanced biological activities. Chitosan improves solubility, stability and resistance to degradation, which offers controlled drug delivery to specific sites and has enhanced antimicrobial properties.39,40
3.3 Encapsulation efficiency of CT-CS-LA-NC
The encapsulation efficiency (EE) of the CT-CS-LA-NC beads was found to be 98.43% (Fig. S3). The CT-CS-LA-NC coating material's significant efficacy in preserving the probiotic cells during the encapsulation process is due to its high efficiency. The results of the present study agreed with Afzaal et al. (2020)41 who reported that sodium alginate microbead encapsulating probiotics are 98% more effectively than carrageenan. The alginate matrix entraps more cells than the other material, but both materials entrap the necessary number of cells. Probiotic survival is increased by encapsulation with alginate and whey protein, suggesting that efficient probiotic consumption is necessary for colonization in the colon.3.4 Characterization techniques
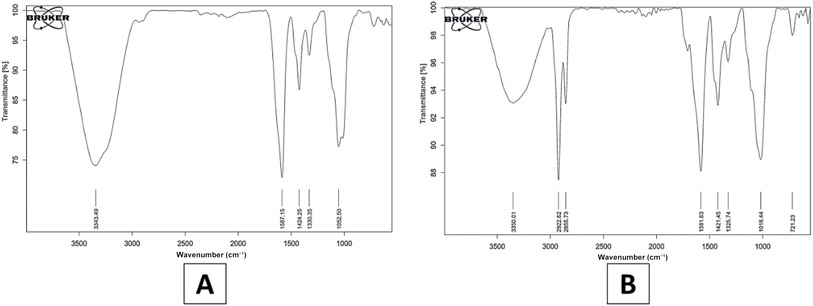
The FTIR spectroscopy results of the synthesized chitosan-probiotic-Clitoria ternatea-nanocapsule (CT-CS-LA-NC) revealed absorption peaks across the range of 720 cm−1 to 3350 cm−1, signifying molecular interactions and the presence of various functional groups (Fig. 3). The peak at 3350 cm−1 might be related to O–H stretching, which indicates that hydroxyl groups are present. Chitosan mostly consists of N–H and O–H stretching, amide bands, C–O–C stretching, and C–H stretching, among several other peaks.46,47 Broad absorption peaks at 2922 cm−1 and 2855 cm−1 suggest C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O carboxyl stretching and typical C–H stretching in alkanes, respectively. A sharp peak at 1582 cm−1 could correspond to C
O carboxyl stretching and typical C–H stretching in alkanes, respectively. A sharp peak at 1582 cm−1 could correspond to C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching vibrations in aromatic compounds.48 The distinct peaks at 1421 cm−1 and 1326 cm−1 are often indicative of C–C stretching (Kumar and Mohideen, 2024).47 Additionally, the absorption at 1018.44 cm−1 might represent C–N stretching vibrations found in ethers or alcohols, signifying the presence of aliphatic amines.49 Lastly, the peak at 721 cm−1 is commonly associated with C–H rocking modes, further confirming the molecular characteristics of the nano-encapsulated structure. Senthil Kumar et al. (2024)50 reported that this is due to glycosidic linkages in chitosan, polysaccharides, and the bacterial cell wall of Lactobacillus fermentum. Chitosan (CS) is the reference peak, and shifts in existing peaks can be attributed to interactions between compounds. This can be explained by the glycosidic bonds present in polysaccharides, present in chitosan as well as the bacterial cell wall of Lactobacillus fermentum.50 With chitosan (CS) as a reference peak, we can confirm the presence of chitosan in the synthesised NPs and mild shifts in existing peaks could be linked to interactions between the bacterial components.51 The FTIR spectra of the CS–NC (Fig. 3A) control and the CT-CS-LA-NC (Fig. 3B) show significant structural modifications. The CT-CS-LA-NC spectrum reveals two bands at 2922 cm−1 and 2855 cm−1, indicating the presence of organic compounds such as lipids, proteins from L. acidophilus, and organic acids from C. ternatea. These peaks are absent in the pure chitosan control. The shift in the N–H bending/amide II peak from 1587.15 cm−1 in the control to 1581.63 cm−1 in the formed sample confirms the encapsulation of the plant extract. The most significant change is the shift of the characteristic chitosan C–O/C–N stretching peak from 1052.50 cm−1 to 1018.44 cm−1. This shift suggests that the encapsulated components and chitosan have strong molecular interactions, which confirms the successful chemical integration and stabilization of the nanocomposite.
C stretching vibrations in aromatic compounds.48 The distinct peaks at 1421 cm−1 and 1326 cm−1 are often indicative of C–C stretching (Kumar and Mohideen, 2024).47 Additionally, the absorption at 1018.44 cm−1 might represent C–N stretching vibrations found in ethers or alcohols, signifying the presence of aliphatic amines.49 Lastly, the peak at 721 cm−1 is commonly associated with C–H rocking modes, further confirming the molecular characteristics of the nano-encapsulated structure. Senthil Kumar et al. (2024)50 reported that this is due to glycosidic linkages in chitosan, polysaccharides, and the bacterial cell wall of Lactobacillus fermentum. Chitosan (CS) is the reference peak, and shifts in existing peaks can be attributed to interactions between compounds. This can be explained by the glycosidic bonds present in polysaccharides, present in chitosan as well as the bacterial cell wall of Lactobacillus fermentum.50 With chitosan (CS) as a reference peak, we can confirm the presence of chitosan in the synthesised NPs and mild shifts in existing peaks could be linked to interactions between the bacterial components.51 The FTIR spectra of the CS–NC (Fig. 3A) control and the CT-CS-LA-NC (Fig. 3B) show significant structural modifications. The CT-CS-LA-NC spectrum reveals two bands at 2922 cm−1 and 2855 cm−1, indicating the presence of organic compounds such as lipids, proteins from L. acidophilus, and organic acids from C. ternatea. These peaks are absent in the pure chitosan control. The shift in the N–H bending/amide II peak from 1587.15 cm−1 in the control to 1581.63 cm−1 in the formed sample confirms the encapsulation of the plant extract. The most significant change is the shift of the characteristic chitosan C–O/C–N stretching peak from 1052.50 cm−1 to 1018.44 cm−1. This shift suggests that the encapsulated components and chitosan have strong molecular interactions, which confirms the successful chemical integration and stabilization of the nanocomposite.
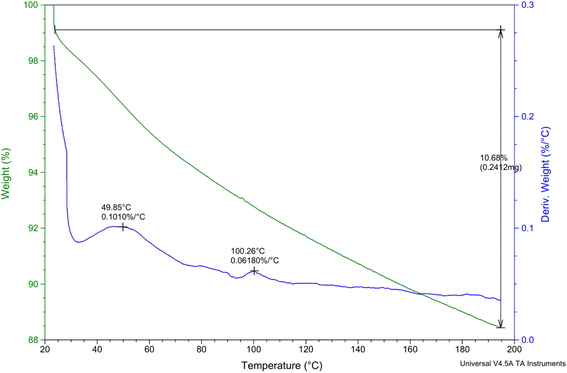
The sample weighed 2.2590 mg initially, and as the temperature increased, the mass changed. At 49.85 °C, a slight weight loss of 0.1010% was noted during the initial decomposition stage. This implies the elimination of any remaining moisture or volatile components, which is common for materials based on chitosan, according to Zhang et al.'s (2021) study.53 This significant loss suggests the breakdown of polymeric structures, such as chitosan and probiotic encapsulation materials, due to thermal degradation. The weight percentage dropped to 0.0618% in the second decomposition stage at 100.26 °C indicating further water loss or slight degradation of low-molecular-weight compounds. At end of the experiment, a final retained weight of 10.68% was recorded, indicating a total weight loss of 0.2412 mg. The results obtained by Hong et al. (2007)54 confirm the existence of several thermal degradation phases and indicate that nanoencapsulation improves thermal stability by postponing degradation relative to pure chitosan.
3.5 Antibacterial activity
Antibacterial activity was tested against E. coli and B. subtilis using a well-diffusion assay. The assay was performed with viable inocula of the respective L. acidophilus treatment groups, including their secretory products. The results showed that all treatments inhibited the growth of both bacterial strains. The nano-encapsulated L. acidophilus and its secretory product exhibited the largest zones of inhibition for Escherichia coli, measuring 14.2 ± 0.26 cm and 14.3 ± 0.27 mm, respectively. The zone of inhibition formed by the secretory product of the free L. acidophilus was 13.1 ± 0.17 cm, whereas the smallest zone was established by the free L. acidophilus itself, measuring 12 ± 0.19 mm for Bacillus subtilis as shown in Table 3. The nano-encapsulated L. acidophilus and its secretory product were similarly quite effective, which exhibited a zone of inhibition of 15.1 ± 0.24 mm and 14.4 ± 0.32 mm. The inhibition of the free L. acidophilus and its secretory product was similar, with zones of 13 ± 0.19 mm and 13.4 ± 0.21 mm, respectively. Similar to the viable inocula, the secretory products from both free and encapsulated formulations showed growth inhibition. These findings are consistent with those of Soltani et al. (2022)55 who reported that the most significant growth inhibition zones were associated with the extracts of L. acidophilus (16 mm) and L. casei (15 mm) strains. The secretome of both Lactobacillus strains exhibited a greater growth inhibition zone.| S. no. | Treatment | Zone of inhibition (mm) | |
|---|---|---|---|
| Escherichia coli (ATCC 25922) | Bacillus subtillis (ATCC 6051) | ||
| a Values are presented as the mean ± standard error of the mean (n = 3). Different letters in the same columns denote statistical significance (P < 0.05) according to Duncan's multiple range test, which was performed following a significant one-way ANOVA. | |||
| 1 | Free L. acidophilus | 12 ± 0.19c | 13 ± 0.19c |
| 2 | Nano-encapsulated L. acidophilus | 14.2 ± 0.26a | 15.1 ± 0.24a |
| 3 | Secretory product of free L. acidophilus | 13.1 ± 0.17b | 13.4 ± 0.21c |
| 4 | Secretory product of nanoencapsulated L. acidophilus | 14.3 ± 0.27a | 14.4 ± 0.32b |
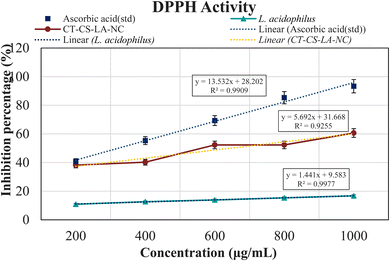
3.6 Antioxidant activity
3.7 Viability of free L. acidophilus and CT-CS-LA-NC under acidic conditions
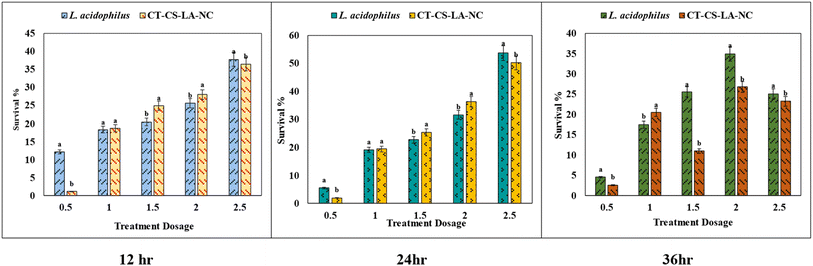
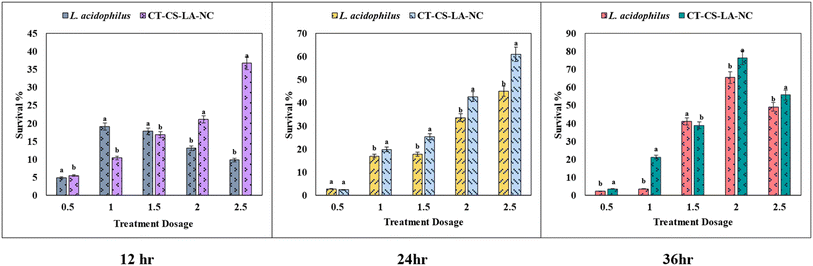
Probiotic bacteria are typically delivered in food systems and must be acid tolerant to survive in the human gastrointestinal tract. The average time from entry to release from the stomach is ninety minutes, though this can vary depending on other digestive processes (Ding and Shah, 2009).14 Encapsulation improves the survival of probiotic bacteria by providing a protective barrier against harsh environmental conditions, such as the stomach's low pH. This barrier protects the cell membrane and cell wall from being destroyed right away, allowing the bacteria to travel through the gastric environment and into the intestine.57In this acid tolerance study, the survival of free Lactobacillus acidophilus and nano-encapsulated CT-CS-LA-NC was assessed under simulated gastric conditions at pH 2.5 and pH 3.5 over 12, 24, and 36 hours is shown in Tables S1 and S2. The concentration of free L. acidophilus and CT-CS-LA-NC was varied from 0.5 mL to 2.5 mL. Initially, at a pH of 2.5, the acid resistance of free L. acidophilus was moderate over 12 hours, with survival rates varying from 12.08 ± 0.10 to 37.80 ± 0.21. The nano-encapsulated CT-CS-LA-NC showed a similar survival rate, ranging from 1.07 ± 0.12 to 36.49 ± 0.34. Free cells showed better viability of 4.51 ± 0.21 to 25.12 ± 0.16 than the nano-encapsulated cells (2.58 ± 0.15 to 23.26 ± 0.16) after 24 hours of acid exposure. The lower total viable count compared to the free cells may result from the probiotic's gradual and prolonged release from the encapsulation. However, the most significant difference was seen after 36 hours (Fig. 6). Encapsulated cells showed a highly significant survivability rate of 60.93 ± 0.68 at 2.5 mL. In contrast, the survival of free L. acidophilus peaked at only 45 ± 0.13.
At pH 3.5, CT-CS-LA-NC maintained similar vitality ranging from 1.93 ± 0.18 to 50.23 ± 0.43 after 12 hours of exposure whereas free L. acidophilus also showed similar vitality (5.47 ± 0.24 to 36.68 ± 0.26). Free cell survivability decreased at 24 hours (4.85 ± 0.02 to 9.80 ± 0.08). After 36 hours, CT-CS-LA-NC showed 76.46 ± 0.60 (2 mL), exceeding free cells (65.42 ± 0.73) and exhibiting enhanced acid tolerance. The survivability of CT-CS-LA-NC was greatly improved confirming the enhanced acid tolerance (60.93 ± 0.68) at 2.5 mL after 36 hours, compared to 45 ± 0.13 for free L. acidophilus, indicating the high efficiency of the encapsulated L. acidophilus (Fig. 7).
These results correspond with prior research. Varela-Pérez et al. (2022)58 indicated that nanoencapsulation improves probiotic survival under acidic conditions. Arratia-Quijada et al. (2024)59 examined persistent acid tolerance in encapsulated strains. Argyri et al. (2012)60 highlighted that resistance to low pH is specific to strains, employing pH 2.5 to identify acid-tolerant Lactobacilli. The results indicate the survival rate at different concentrations and time intervals, showing CT-CS-LA-NC's efficiency in preserving probiotic viability under simulated gastrointestinal conditions. Chitosan acts as a prebiotic, providing nourishment for probiotics such as L. acidophilus in the large intestine.61 The synbiotic effect – the combination of a probiotic and a prebiotic—substantially enhances the colonization and therapeutic advantages of the encapsulated bacteria. The synergistic relationship between the components of the product enhances probiotic viability, stability, and therapeutic potential, providing a robust protective effect against gut pathogens, ensuring successful delivery of L. acidophilus.62
3.8 Viability of free L. acidophilus and CT-CS-LA-NC under bile conditions
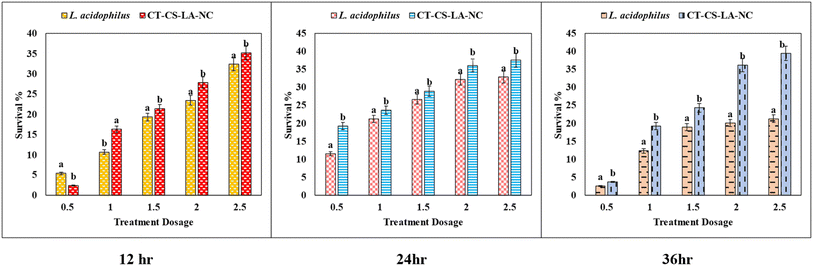
The survival rates of both free and nano-encapsulated L. acidophilus (CT-CS-LA-NC) at two pH values (2.5 and 3.5) over 12, 24, and 36 hours were investigated in this bile tolerance study as shown in Table S3. At a low concentration of 0.5 mL, the tolerance of the free L. acidophilus cells was significantly higher than that of the nano-encapsulated cells (5.39 ± 0.13). This implies that the encapsulation might slowly dissolve or activate. However, the nano-encapsulated CT-CS-LA-NC cells showed a noticeably higher tolerance when the concentrations were raised to a higher level (between 1 and 2.5 mL). The CT-CS-LA-NC maximum tolerance at 2.5 mL was 35.24 ± 0.3, greater than the free cell maximum tolerance of 32.41 ± 0.12.After 24 hours of exposure, the slow-release profile of CT-CS-LA-NC was noticeably superior to that of free L. acidophilus. Across a range of nanoencapsulation concentrations (0.5 mL to 2.5 mL), CT-CS-LA-NC demonstrated significantly higher survival rates. CT-CS-LA-NC exhibited a survival rate of 19.25 ± 0.15 at the lowest concentration of 0.5 mL, whereas free cells only showed an 11.56 ± 0.21 survival rate (Fig. 8). As exposure duration increases, nano-encapsulation significantly enhances protection against bile tolerance, as shown by its consistent performance across all of the various concentrations that were assessed. After 36 hours, the highest survivability rates were found in encapsulated cells. At all concentrations, CT-CS-LA-NC's tolerance was significantly greater than that of free L. acidophilus. The survival of free L. acidophilus was only 21.29 ± 0.13, while the encapsulated cells demonstrated a survivability of 39.46 ± 0.68 at 2.5 mL. This extremely significant difference highlights the high effectiveness of the encapsulated L. acidophilus (CT-CS-LA-NC) in resisting the harsh environment of bile acids, particularly at higher concentrations and for longer periods of time.
3.9 Ecotoxicity assessment of CT-CS-LA-NC
3.10 Storage stability
The probiotic cells encased in an encapsulating matrix must endure severe conditions, including external factors and the gastrointestinal tract, while also preserving their viability during storage. The storage temperature has a major impact on the viability of the cells during the storage time. This study involved the storage of free L. acidophilus cells and encapsulated cells at a refrigerated temperature of 4 °C. Fig. 11 depicts the storage stability of free L. acidophilus and CT-CS-LA-NC probiotics when stored at a refrigeration temperature of 4 °C for a duration of 28 days. The viability of the free cells showed rapid decline, with the survival rate decreasing from an initial 7.98 to 7.8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log10
log10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CFU per mL by the end of 28 days. However, the CT-CS-LA-NC encapsulated cells also had similar higher survival rates, retaining 7.84
CFU per mL by the end of 28 days. However, the CT-CS-LA-NC encapsulated cells also had similar higher survival rates, retaining 7.84![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log10
log10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CFU per mL viability after 7 days and maintaining more than half of their initial population up to 14 days after the 28 days. Probiotic viability and storage durability are enhanced by the steady and gradual release of cells from the coating material, chitosan biopolymer, which traps L. acidophilus inside the alginate. Fareez et al. (2014)65 reported that while decreasing viability loss at 75 and 90 °C, Alg–XG–Ch increased the storage durability of L. plantarum LAB12 at 4 °C, and L. plantarum LAB12, combined in Alg–XG–Ch, might be used as a new functional food ingredient with health benefits.
CFU per mL viability after 7 days and maintaining more than half of their initial population up to 14 days after the 28 days. Probiotic viability and storage durability are enhanced by the steady and gradual release of cells from the coating material, chitosan biopolymer, which traps L. acidophilus inside the alginate. Fareez et al. (2014)65 reported that while decreasing viability loss at 75 and 90 °C, Alg–XG–Ch increased the storage durability of L. plantarum LAB12 at 4 °C, and L. plantarum LAB12, combined in Alg–XG–Ch, might be used as a new functional food ingredient with health benefits.
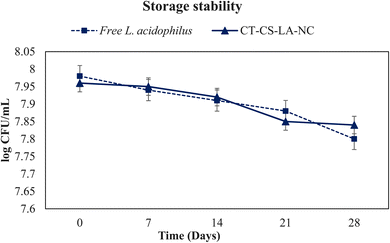 | ||
| Fig. 11 Storage stability of free L. acidophilus and CT-CS-LA-NC at 4 °C for 28 days. The presented values are expressed as means ± SD (n = 3). | ||
3.11 In silico docking
In the present study, the docking analysis focused on identifying the molecular interactions between two proteins associated with probiotics, O-GlcNAcase (7K41) and S-layer associated protein (SlpA) (8AE1), along with three significant compounds from the nanoencapsulated system, such as anthocyanin, lactic acid, and chitosan (Fig. 12). Anthocyanin has a high binding affinity for O-GlcNAcase (7K41) and SlpA (8AE1), with interaction scores of −9.4 kcal mol−1 and −9.6 kcal mol−1, respectively. These interaction scores show that anthocyanin forms stable complexes with proteins, indicating its potential to impact their functions, which is consistent with previous studies on its bioactivity, which is associated with its ability to interact with proteins. According to the study by Hidalgo et al. (2012)66 anthocyanin contributes to the preservation of gastrointestinal health by modulating the microbial composition of the gut and providing antioxidant activity in the large intestine. Chitosan exhibited significant interaction scores of −5.2 kcal mol−1 for O-GlcNAcase and −5.4 kcal mol−1 for SlpA, shown in Table 4, which indicates the distinct binding sites (534 for 7K41 and 2163 for 8AE1) on each protein. Its interaction with SlpA, a key component of the bacterial cell wall, supports its role in protecting the cell membrane. Likewise, lactic acid also had notable interaction scores of −4.1 kcal mol−1 (7K41) and −3.8 kcal mol−1 (8AE1). These in silico findings explain the nano-encapsulated system's synergistic effects and emphasize the molecular interactions between anthocyanin, chitosan, and lactic acid.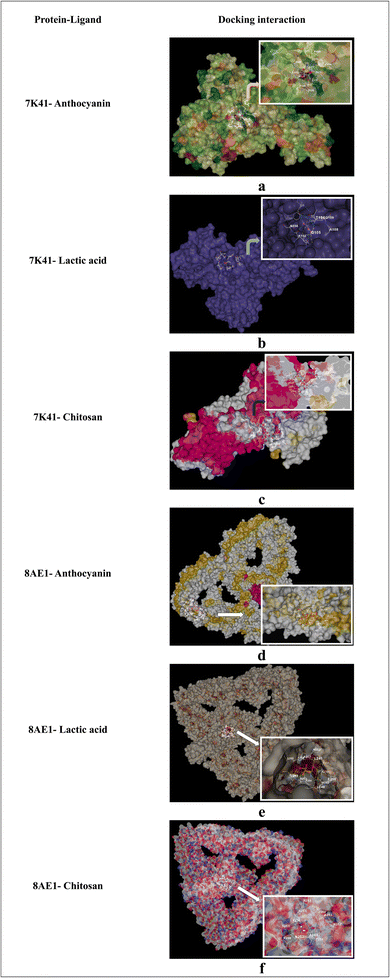 | ||
| Fig. 12 (a–f) In silico docking interaction of proteins (7K41, 8AE1) with ligands (anthocyanin, lactic acid, and chitosan). | ||
| Protein | Ligand | Interaction score (kcal mol−1) | Cavity volume |
|---|---|---|---|
| 7K41 | Anthocyanin | −9.4 | 2633 |
| 7K41 | Lactic acid | −4.1 | 2633 |
| 7K41 | Chitosan | −5.2 | 534 |
| 8AE1 | Anthocyanin | −9.6 | 893 |
| 8AE1 | Lactic acid | −3.8 | 2141 |
| 8AE1 | Chitosan | −5.4 | 2163 |
4. Conclusion
The study successfully demonstrates the nanoencapsulation of Lactobacillus acidophilus using chitosan and Clitoria ternatea extract (CT-CS-LA-NC). The effective encapsulation of Lactobacillus acidophilus using this approach presents new possibilities for its use as a biopreservative in functional foods, drinks, and nutraceuticals. This method's effective encapsulation of Lactobacillus acidophilus creates potential opportunities for its use as a biopreservative in nutraceuticals, functional foods, and drinks. Nanocapsules with improved thermal and acid stability would facilitate the development of novel probiotic-enriched products, such as juices, food products, and dietary supplements, which are traditionally difficult to formulate with probiotics. It may also be able to prolong the shelf life of perishable goods, eliminating the need for artificial preservatives, given its proven antibacterial qualities against common food pathogens like B. subtilis and E. coli. CT-CS-LA-NC provides a versatile and sustainable solution that offers both functional health advantages and enhanced product stability in response to the growing consumer demand for natural products.Consent for publication
We consent to publish in this journal.Author contributions
M. Lavanya: data curation, formal analysis, investigation, methodology, software, original manuscript writing. S. Karthick Raja Namasivayam: conceptualization, supervision, investigation, manuscript review and editing.Conflicts of interest
We declare no conflict of interest.Data availability
The data will be made available on request from the authors.Supplementary information (SI): comprehensive characterization and stability data for Lactobacillus acidophilus and the CT-CS-LA-NC formulation. It includes Gram staining, phylogenetic tree analysis, and physicochemical characterization via encapsulation efficiency, particle size distribution (DLS), and zeta potential measurements. Additionally, the file contains detailed comparative viability data for free L. acidophilus and CT-CS-LA-NC across varying pH levels (2.5, 3.5, and 7.5) and treatment dosages over 12, 24, and 36-hour intervals. See DOI: https://doi.org/10.1039/d5fb00833f.
References
- S. K. Amit, Md. M. Uddin, R. Rahman, S. M. R. Islam and M. S. Khan, Agric. Food Secur., 2017, 6, 51 CrossRef.
- M. Martínez-Ballesta, Á. Gil-Izquierdo, C. García-Viguera and R. Domínguez-Perles, Foods, 2018, 7, 72 CrossRef PubMed.
- T. Didari, S. Solki, S. Mozaffari, S. Nikfar and M. Abdollahi, Expert Opin. Drug Saf., 2014, 13, 227–239 CrossRef PubMed.
- R. R. Watson and V. R. Preedy, Probiotics, Prebiotics, and Synbiotics: Bioactive Foods in Health Promotion, Academic Press, 2015 Search PubMed.
- S. Fijan, Int. J. Environ. Res. Public Health, 2014, 11, 4745–4767 CrossRef PubMed.
- P. A. Mackowiak, Front. Public Health, 2013, 1, 52 CrossRef PubMed.
- A. Latif, A. Shehzad, S. Niazi, A. Zahid, W. Ashraf, M. W. Iqbal, A. Rehman, T. Riaz, R. M. Aadil, I. M. Khan, F. Özogul, J. M. Rocha, T. Esatbeyoglu and S. A. Korma, Front. Microbiol., 2023, 14, 1216674 CrossRef PubMed.
- K. Prajapati, K. Bisani, H. Prajapati, S. Prajapati, D. Agrawal, S. Singh, M. Saraf and D. Goswami, Syst. Microbiol. Biomanuf., 2023, 4, 386–406 CrossRef.
- M. Bernatek, H. Sommermeyer, A. Wojtyła, J. Piątek and S. Lachowicz-Wiśniewska, J. Health Inequal., 2023, 9, 194–200 CrossRef.
- M. L. Marco and S. Tachon, Curr. Opin. Biotechnol., 2012, 24, 207–213 CrossRef PubMed.
- A. Pugazhendhi, M. A. Alshehri, S. Kandasamy, P. K. Sarangi and A. Sharma, Food Chem., 2024, 464, 141762 CrossRef PubMed.
- L. Stasiak-Różańska, A. Berthold-Pluta, A. S. Pluta, K. Dasiewicz and M. Garbowska, Int. J. Environ. Res. Public Health, 2021, 18, 1108 CrossRef PubMed.
- K. V. M. Rameez, P. Santhoshkumar, K. S. Yoha and J. A. Moses, Curr. Res. Nutr. Food Sci. J, 2024, 12, 539–560 Search PubMed.
- W. K. Ding and N. P. Shah, J. Food Sci., 2009, 74, 100–107 Search PubMed.
- M. Chávarri, I. Marañón, R. Ares, F. C. Ibáñez, F. Marzo and M. Del Carmen Villarán, Int. J. Food Microbiol., 2010, 142, 185–189 CrossRef PubMed.
- D. Dahiya and P. S. Nigam, Fermentation, 2022, 8, 303 CrossRef CAS.
- M. Pateiro, B. Gómez, P. E. S. Munekata, F. J. Barba, P. Putnik, D. B. Kovačević and J. M. Lorenzo, Molecules, 2021, 26, 1547 CrossRef CAS PubMed.
- E. J. Jeyaraj, Y. Y. Lim and W. S. Choo, Sci. Rep., 2022, 12, 14890 CrossRef CAS PubMed.
- A. Menconi, G. Kallapura, J. D. Latorre, M. J. Morgan, N. R. Pumford, B. M. Hargis and G. Tellez, Biosci. Microbiota, Food Health, 2014, 33, 25–30 CrossRef PubMed.
- C. A. Okafor and P. E. Ekwueme, Asian Food Sci. J., 2024, 23, 95–104 CrossRef.
- B. Tilahun, A. Tesfaye, D. Muleta, A. Bahiru, Z. Terefework and G. Wessel, Int. J. Food Sci., 2018, 2018, 1–7 CrossRef PubMed.
- G. C. V. Gamage, Y. Y. Lim and W. S. Choo, Front. Plant Sci., 2021, 12, 792303 CrossRef PubMed.
- S. Chakraborty, S. Sahoo, A. Bhagat and S. Dixit, Zenodo (CERN Eur. Organ. Nucl. Res.), 2017, 5, 197–208 Search PubMed.
- H. Choukaife, A. A. Doolaanea and M. Alfatama, Pharmaceuticals, 2020, 13, 335 CrossRef CAS PubMed.
- D. Arepally, R. S. Reddy and T. K. Goswami, Food Funct., 2020, 11, 8694–8706 RSC.
- M. Cassayre, D. T. De Souza, M. Claeys-Bruno, A. Altié, P. Piccerelle and C. Sauzet, Nanomaterials, 2025, 15, 1034 CrossRef CAS PubMed.
- M. S. Blois, Nature, 1958, 181, 1199–1200 CrossRef CAS.
- Md. M. Rahman, Md. B. Islam, M. Biswas and A. H. M. K. Alam, BMC Res. Notes, 2015, 8, 621 CrossRef PubMed.
- N. Soltani, S. Abbasi, S. Baghaeifar, E. Taheri, M. F. S. Jadid, P. Emami, K. Abolhasani and F. Aslanshirzadeh, Biotechnol. Rep., 2022, 36, e00760 CrossRef CAS PubMed.
- M. A. Ehrmann, P. Kurzak, J. Bauer and R. F. Vogel, J. Appl. Microbiol., 2002, 92, 966–975 CrossRef CAS PubMed.
- U. Akmal, I. Ghori, A. M. Elasbali, B. Alharbi, A. Farid, A. S. Alamri, M. Muzammal, S. M. B. Asdaq, M. A. E. Naiel and S. Ghazanfar, Fermentation, 2022, 8, 328 CrossRef CAS.
- K. Thinkohkaew, N. Aumphaiphensiri, T. Tangamornsiri, N. Niamsiri, P. Potiyaraj and I. Suppavorasatit, J. Agric. Food Res., 2024, 18, 101405 CAS.
- Z. Abbas, M. A. Hassan, W. Huang, H. Yu, M. Xu, X. Chang, X. Fang and L. Liu, Influence of magnesium oxide (MgO) nanoparticles on maize (Zea mays L.), Agronomy, 2024, 14(3), 617, DOI:10.3390/agronomy14030617.
- S. P. Fling and D. S. Gregerson, Anal. Biochem., 1986, 155, 83–88 CrossRef CAS PubMed.
- Y. Liu, M. Grimm, W.-T. Dai, M.-C. Hou, Z.-X. Xiao and Y. Cao, Acta Pharmacol. Sin., 2019, 41, 138–144 CrossRef PubMed.
- G. Rozos, C. Voidarou, E. Stavropoulou, I. Skoufos, A. Tzora, A. Alexopoulos and E. Bezirtzoglou, Front. Microbiol., 2018, 9, 517 CrossRef PubMed.
- G. Sumalata and M. Shiva Prakash, J. Adv. Sci. Res., 2023, 14, 40–49 Search PubMed.
- G. D. Neda, M. S. Rabeta and M. T. Ong, Int. Food Res. J., 2013, 20, 1229–1234 CAS.
- P. R. Deshmukh, A. Joshi, C. Vikhar, S. S. Khdabadi and M. Tawar, Syst. Rev. Pharm., 2022, 13, 685–693 Search PubMed.
- Y. Pérez-Pacheco, B. Tylkowski and R. García-Valls, Molecules, 2025, 30, 252 CrossRef PubMed.
- M. Afzaal, F. Saeed, M. Saeed, M. Azam, S. Hussain, A. A. Mohamed, M. S. Alamri and F. M. Anjum, Int. J. Food Prop., 2020, 23, 1899–1912 CrossRef CAS.
- L. Weng, L. Wu, R. Guo, J. Ye, W. Liang, W. Wu, L. Chen and D. Yang, J. Nanobiotechnol., 2022, 20(1), 356, DOI:10.1186/s12951-022-01563-x.
- L. L. Barrera-Necha, Z. N. Correa-Pacheco, S. Bautista-Baños, M. Hernández-López, J. E. M. Jiménez and A. F. M. Mejía, Adv. Microbiol., 2018, 08(04), 286–296, DOI:10.4236/aim.2018.84019.
- M. A. Ahghari, M. R. Ahghari, M. Kamalzare and A. Maleki, Sci. Rep., 2022, 12, 10491 CrossRef CAS PubMed.
- S. Vijayaram, R. Sinha, C. Faggio, E. Ringø and C.-C. Chou, AIMS Microbiol., 2024, 10, 986–1023 CAS.
- J. J. Joseph, D. Sangeetha and T. Gomathi, Int. J. Biol. Macromol., 2015, 82, 952–958 CrossRef PubMed.
- S. S. Kumar and S. S. Mohideen, Sci. Rep., 2024, 14, 21182 CrossRef.
- M. C. Curk, F. Peledan and J. C. Hubert, FEMS Microbiol. Lett., 1994, 123, 241–248 CrossRef CAS.
- R. V. Mathai, J. C. Mitra and S. K. Sar, Rasayan J. Chem., 2021, 14, 1423–1434 CrossRef CAS.
- S. Senthil Kumar and S. S. Mohideen, Sci. Rep., 2024, 14, 21182 CrossRef CAS PubMed.
- R. Shanmugam, T. Munusamy, S. Jayakodi, K. A. Al-Ghanim, M. Nicoletti, N. Sachivkina and M. Govindarajan, Fermentation, 2023, 9, 413 CrossRef CAS.
- S. K. R. Namasivayam, M. Manohar, J. A. Kumar, K. Samrat, A. Kande, R. S. A. Bharani, C. Jayaprakash and S. Lokesh, Environ. Res., 2022, 212, 113386 CrossRef PubMed.
- Y. Zhang, Y. Wu, M. Yang, G. Zhang and H. Ju, Materials, 2021, 14, 5538 CrossRef CAS PubMed.
- P. Hong, S. Li, C. Ou, C. Li, L. Yang and C. Zhang, J. Appl. Polym. Sci., 2007, 105, 547–551 CrossRef CAS.
- N. Soltani, S. Abbasi, S. Baghaeifar, E. Taheri, M. F. S. Jadid, P. Emami, K. Abolhasani and F. Aslanshirzadeh, Biotechnol. Rep., 2022, 36, e00760 CrossRef CAS PubMed.
- G. I. Edo, A. N. Mafe, N. F. Razooqi, E. C. Umelo, T. S. Gaaz, E. F. Isoje, U. A. Igbuku, P. O. Akpoghelie, R. A. Opiti, A. E. A. Essaghah, D. S. Ahmed, H. Umar and D. U. Ozsahin, Des. Monomers Polym., 2024, 28, 1–34 Search PubMed.
- N. Sulistiani, I. Novarina, N. Inawati, A. Dinoto, H. Julistiono, R. Handayani and S. Saputra, IOP Conf. Ser.: Earth Environ. Sci., 2020, 572, 012026 CrossRef.
- A. Varela-Pérez, O. O. Romero-Chapol, A. G. Castillo-Olmos, H. S. García, M. L. Suárez-Quiroz, J. Singh, C. Y. Figueroa-Hernández, R. Viveros-Contreras and C. Cano-Sarmiento, Foods, 2022, 11, 740 CrossRef PubMed.
- J. Arratia-Quijada, K. Nuño, V. Ruíz-Santoyo and B. A. Andrade-Espinoza, J. Funct. Foods, 2024, 116, 106192 CrossRef CAS.
- A. A. Argyri, G. Zoumpopoulou, K.-A. G. Karatzas, E. Tsakalidou, G.-J. E. Nychas, E. Z. Panagou and C. C. Tassou, Food Microbiol., 2012, 33, 282–291 CrossRef PubMed.
- S. S. Kumar and S. S. Mohideen, Sci. Rep., 2024, 14, 21182 CrossRef PubMed.
- S. Nigam, D. V. Ram and A. A. M. Kumar, Int. J. Pharm. Sci. Rev. Res., 2022, 154–160 CrossRef.
- M. A. O. Torio, M. Adachi, R. N. Garcia, K. Prak, N. Maruyama, S. Utsumi and E. M. Tecson-Mendoza, Food Res. Int., 2011, 44, 2984–2990 CrossRef CAS.
- Ma. C. Gamis, L. Y. Uy, A. Laurena, W. Hurtada and M. A. Torio, Appl. Sci., 2020, 10, 8787 CrossRef CAS.
- I. M. Fareez, S. M. Lim, R. K. Mishra and K. Ramasamy, Int. J. Biol. Macromol., 2014, 72, 1419–1428 CrossRef PubMed.
- M. Hidalgo, M. J. Oruna-Concha, S. Kolida, G. E. Walton, S. Kallithraka, J. P. E. Spencer, G. R. Gibson and S. De Pascual-Teresa, J. Agric. Food Chem., 2012, 60, 3882–3890 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |