Open Access Article
Open Access ArticleTargeted modulation of FGFR1 enhances myogenic differentiation for cultured meat production
Somsiri Udompaisarn a,
Prant Chotiphantawanonb,
Sakonwan Kuhaudomlarp
a,
Prant Chotiphantawanonb,
Sakonwan Kuhaudomlarp b,
Ratchakrit Srikuea
b,
Ratchakrit Srikuea c,
Sittiwat Wangwongpaiboonc,
Wasina Watcharanapapanc,
Thananya Soonkum
c,
Sittiwat Wangwongpaiboonc,
Wasina Watcharanapapanc,
Thananya Soonkum d,
Dusit Laohasinnarong
d,
Dusit Laohasinnarong e,
Utid Suriya
e,
Utid Suriya *b and
Tavan Janvilisri
*b and
Tavan Janvilisri *a
*a
aDepartment of Microbiology, Faculty of Science, Chulalongkorn University, Bangkok, Thailand. E-mail: tavan.j@chula.ac.th
bDepartment of Biochemistry, Faculty of Science, Mahidol University, Bangkok, Thailand. E-mail: utid.sur@mahidol.ac.th
cDepartment of Physiology, Faculty of Science, Mahidol University, Bangkok, Thailand
dMahidol University-Frontier Research Facility, Mahidol University, Nakhon Pathom, Thailand
eDepartment of Clinical Sciences and Public Health, Faculty of Veterinary Science, Mahidol University, Nakhon Pathom, Thailand
First published on 11th March 2026
Abstract
The development of cultured meat offers a sustainable and ethical alternative to conventional protein sources, yet its commercial scalability is hindered by the inefficient induction of terminal myogenesis. Porcine muscle stem cells (PMSCs) are a promising cell source, but their effective differentiation into mature muscle fibers remains a significant challenge. The fibroblast growth factor receptor 1 (FGFR1) plays a pivotal role in maintaining the proliferative state of these cells, making its inhibition a compelling strategy to promote differentiation. In this study, we employed a multi-faceted approach, combining in silico and in vitro methods, to identify and validate novel small-molecule FGFR1 modulators. A rigorous virtual screening of 872 compounds against the FGFR1 tyrosine kinase domain, with subsequent filtering against the off-target p38α MAPK, identified methylergometrine as a highly promising candidate alongside the known inhibitor dovitinib. Molecular dynamics simulations confirmed that both compounds form stable complexes with FGFR1, a finding corroborated by saturation transfer difference-nuclear magnetic resonance, which provided direct evidence of their distinct but complementary binding modes. Subsequent in vitro functional assays in C2C12 mouse myogenic cells demonstrated that both compounds enhance myogenic differentiation. Specifically, treatment with dovitinib and methylergometrine led to a robust, dose- and phase-dependent upregulation of key myogenic markers, including myosin heavy chain and myogenin. These findings confirm that dovitinib and methylergometrine effectively promote myotube formation. Similar trends were also observed in PMSCs. Our study introduces a novel strategy for stimulating terminal myogenesis through targeted FGFR1 modulation, with significant implications for improving the efficiency and yield of cell-based cultured meat production.
Sustainability spotlightCellular agriculture offers a path to sustainable protein production, but its scalability is hindered by the high cost of media components needed for muscle cell differentiation. We address this major bottleneck by pioneering a multi-criteria chemical screening to identify methylergometrine, an affordable, clinically-approved compound, as a potent driver for transforming muscle stem cells into mature muscle fibers. This compound drastically reduces the reliance on expensive growth factors, cutting production costs for cultured meat. Our discovery, including a dual-phase chemical strategy to accelerate tissue formation, provides a chemically-defined, cost-effective method to achieve industrial-scale cultivated protein. This work directly supports UN SDG 2 (zero hunger) by making sustainable protein accessible and UN SDG 12 (responsible consumption and production) by offering an efficient, resource-light alternative to conventional meat farming. |
1. Introduction
Dietary protein is a critical nutrient for human growth, development, and healthy aging, supporting physical development during early life and the maintenance of muscle mass, metabolic health, and overall quality of life during aging.1,2 The escalating global demand for sustainable and ethically sourced protein has propelled significant advancements in the development of cultured meat.3,4 Among the diverse cell sources investigated, muscle stem cells, also known as satellite cells, present a compelling platform due to their intrinsic capacity for robust proliferation and efficient differentiation into mature muscle fibers.5,6 However, the commercial scalability of cultured meat production remains hindered by several key challenges including low cell yields, a decline in differentiation efficiency over successive cell passages, and the prevalent use of animal-derived components in current culture media.7 Addressing these challenges necessitates a refined understanding of the molecular mechanisms governing myogenesis, particularly the intricate balance between cell proliferation and terminal differentiation.Myogenic differentiation and myotube formation are widely recognized as fundamental biological processes underlying muscle formation and are critical for the development of muscle-like tissue in cultured meat systems. While large-scale cell expansion is required to generate sufficient biomass, the efficiency of terminal differentiation ultimately defines product quality such as muscle structure, protein composition, and meat texture. Without the transition from mononuclear myoblasts to multinucleated myotubes, the final product lacks the structural integrity of conventional muscle tissue (meat).8,9 Therefore, inefficient differentiation leads to immature and heterogeneous tissue structure and resulting in reducing product consistency, yield, scalability and limiting the feasibility of sustainable cultured meat production. Currently, achieving efficient and consistent myogenic differentiation remains challenging, even when sufficient cell biomass is obtained.10 Consequently, ensuring proper induction and control of myogenic differentiation is essential for the development of scalable and reliable production systems for cultured meat.
Fibroblast growth factor receptor 1 (FGFR1) is a crucial regulator of muscle stem cell fate, acting as a molecular switch that maintains a proliferative, undifferentiated state. Activation of FGFR1 signaling pathway promotes cell cycle progression by inhibiting key cyclin-dependent kinase inhibitors like p21Waf1 and p27Kip1, thereby favoring proliferation over differentiation.11 Conversely, inhibiting or downregulating FGFR1 activity consistently promotes myogenic differentiation. For example, the non-coding RNAs miR-133 and lncR-133a have been shown to facilitate differentiation of C2C12 and goat muscle stem cells by downregulating FGFR1 and attenuating ERK1/2 signaling.12,13 Similarly, leucine-rich repeats and transmembrane domains 1 (LRTM1) negatively regulates FGFR1 by interfering with adaptor protein recruitment, which reduces downstream ERK activation and enhances myotube formation.14 These findings collectively highlight FGFR1 inhibition as a highly promising therapeutic and biotechnological strategy to trigger terminal myogenesis. However, approaches based on non-coding RNA modulation or genetic manipulation of regulatory proteins are difficult to translate into practical culture supplements. To date, defined small-molecule inhibitors suitable for incorporation into muscle cell culture media remain limited. Therefore, in this study, the primary objective of this study was to identify and validate novel pharmacological modulators of FGFR1 that accelerate the transition of muscle stem cells from a proliferative state to terminal myogenesis, thereby optimizing the structural and biochemical maturation of porcine-derived cultured muscle.
To identify novel compounds capable of modulating FGFR1, we employed an in silico approach, leveraging molecular docking and molecular dynamics (MD) simulations to assess the binding affinities and molecular interactions of potential candidates.15–17 The intracellular tyrosine kinase domain of FGFR1, a well-defined target known for its conserved two-lobed kinase fold, was selected for these virtual screening efforts. To validate these in silico predictions and confirm direct protein–compound interactions, we integrated nuclear magnetic resonance (NMR) spectroscopy into our workflow.18 Specifically, saturation transfer difference (STD) NMR was used as a sensitive and powerful tool to rapidly detect ligand binding and map the binding epitope, providing compelling experimental evidence of molecular interaction without the need for protein immobilization or complex labeling.19
In this study, we focused on two small-molecule compounds with the potential to modulate FGFR1 signaling and influence myogenic differentiation. Dovitinib is a well-characterized multi-kinase inhibitor with established anti-proliferative activity in various cancers through its inhibition of FGFR1 signaling.20 While its direct effects on skeletal muscle differentiation have not been previously described, dovitinib's known ability to inhibit FGFR1 and enhance osteoblast differentiation in other cell types suggests its potential to modulate myogenesis.21 In parallel, we investigated methylergometrine, an ergot alkaloid widely used in obstetrics for its uterotonic properties.22 Our preliminary data indicated that methylergometrine also interacts with tyrosine kinases, positioning it as another intriguing candidate for inducing myogenic differentiation. By evaluating both a well-established FGFR1 inhibitor (dovitinib) and a compound with novel, unexplored kinase-binding properties (methylergometrine), we aimed to systematically assess their potential to induce the differentiation of porcine muscle stem cells (PMSCs) and to explore FGFR1 inhibition as a translatable strategy for cultured meat production.
2. Materials and methods
2.1 Molecular docking and virtual screening
To identify compounds with potential affinity for FGFR1, we performed a comprehensive docking-based virtual screening process. The three-dimensional crystal structures of the human FGFR1 tyrosine kinase domain in complex with the inhibitor AZD4547 and the p38α mitogen-activated protein kinase (p38α MAPK) were retrieved from the RCSB Protein Data Bank (PDB IDs: 4V05![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 23 and 3ZSH,24 respectively). These structures served as our protein targets for in silico analysis. A diverse compound library was curated from two primary sources: a commercial nuclear receptor compound library from MedChemExpress and a collection of 20 proprietary in-house compounds, including asiatic acid, ticagrelor, and various alkaloid-like derivatives. All selected ligand structures were geometrically optimized, and their protonation states were meticulously predicted at a physiological pH of 7.4 using the Open Babel software.25 This rigorous preparation ensured the accuracy of subsequent docking simulations. A total of 872 optimized compounds were prepared for virtual screening. The binding site for docking was precisely defined by the coordinates of the co-crystallized ligands within the FGFR1 and p38α MAPK structures. A consistent grid box of 20 Å × 20 Å × 20 Å was centered on these coordinates to encompass the entire active site. The virtual screening was conducted using the AutoDock VinaXB software.26 The simulations were executed on a Linux-based computing system to handle the computational demands of screening such a large library.
23 and 3ZSH,24 respectively). These structures served as our protein targets for in silico analysis. A diverse compound library was curated from two primary sources: a commercial nuclear receptor compound library from MedChemExpress and a collection of 20 proprietary in-house compounds, including asiatic acid, ticagrelor, and various alkaloid-like derivatives. All selected ligand structures were geometrically optimized, and their protonation states were meticulously predicted at a physiological pH of 7.4 using the Open Babel software.25 This rigorous preparation ensured the accuracy of subsequent docking simulations. A total of 872 optimized compounds were prepared for virtual screening. The binding site for docking was precisely defined by the coordinates of the co-crystallized ligands within the FGFR1 and p38α MAPK structures. A consistent grid box of 20 Å × 20 Å × 20 Å was centered on these coordinates to encompass the entire active site. The virtual screening was conducted using the AutoDock VinaXB software.26 The simulations were executed on a Linux-based computing system to handle the computational demands of screening such a large library.
2.2 Molecular dynamics (MD) simulations
To provide a more dynamic and biologically relevant assessment of compound–protein interactions, we performed extensive all-atom MD simulations. The initial complex structures for MD were generated from the top-scoring docking poses identified in the virtual screening. Prior to simulation, the protonation states of ionizable amino acids in the protein were assigned at physiological pH (pH = 7.4) using the PDB2PQR web server.27 The partial atomic charges of the ligand molecules were calculated using the General AMBER Force Field version 2 (GAFF 2) within the Antechamber module of AMBER24. All simulations were performed under periodic boundary conditions using the AMBER ff19SB force field,28 which is optimized for biological macromolecules. The systems were solvated with the explicit TIP3P water model and simulated under constant temperature (310 K) and pressure (1 atm) conditions (NPT ensemble). The entire system, including protein–ligand complex and solvent, was gradually energy-minimized to relieve any steric clashes. Electrostatic interactions were efficiently calculated using the particle mesh Ewald summation method,29 and the SHAKE algorithm was applied to constrain all bonds involving hydrogen atoms, allowing for a larger integration time step. Temperature was regulated using the Langevin thermostat, while pressure was controlled by the Berendsen barostat.30–33 Each production run was conducted for a duration of 200 ns, with trajectories saved every 10 ps for post-dynamic analysis. The resulting trajectories were analyzed using the CPPTRAJ module of AMBER to evaluate structural stability, conformational changes, and key molecular interactions.34 Finally, the binding free energy (ΔGbind) and decomposition (ΔGresiduebind) energy of the ligand–protein complex was estimated using the Molecular Mechanics/Generalized Born Surface Area (MM/GBSA) method using the MMPBSA.py module.352.3 Saturation transfer difference-nuclear magnetic resonance (STD-NMR) spectroscopy
To experimentally validate the direct binding of dovitinib and methylergometrine to FGFR1, we utilized STD-NMR spectroscopy. Recombinant human FGFR1 protein was sourced from Abnova (Cat. No. H00002260-G01). STD-NMR spectra were acquired on a Bruker Avance Ascend 600 MHz Spectrometer, equipped with a 5 mm BBO Prodigy CryoProbe. The probe temperature was maintained at a constant 298 K. Samples were prepared in a 5 mm NMR tube containing 100 nM of FGFR1 protein and a 15 mM concentration of either dovitinib or methylergometrine dissolved in 600 µL of deuterium oxide (D2O). The residual peak (δH 4.79 ppm) was used as the internal chemical shift reference. The spectrometer frequency was set to 600 MHz for both 1H and STD dimensions. Spectral widths for the 1H dimension was set at 9600 Hz with a 30° pulse angle. Each acquisition involved a recycle delay of 2.5 s and 8 scans, collecting a total of 8 complex points for the 1H dimension.2.4 Cell culture and reagents
The mouse myogenic cell line C2C12 (ATCC, CRL-1772) was utilized as an in vitro model system. Cells were maintained in growth medium (GM) composed of Dulbecco's modified Eagle's medium (DMEM; Gibco, 12800-017) supplemented with 10% (v/v) fetal bovine serum (FBS; Gibco, A5256701) and 1% penicillin/streptomycin (Gibco, 15140-122). Cells were cultured at 37 °C in a humidified incubator with a 5% CO2 atmosphere. For experiments with PMSCs, cells were cultured and their myogenic characteristics were verified as previously described.36 All animal procedures and experimental protocols were conducted in accordance with the Animal for Scientific Purposes Act of Thailand, B.E. 2558 (2015), and the protocol was approved by the Animal Ethics Committee of the Faculty of Veterinary Science, Mahidol University (FVS-MU-IACUC: COA No. MUVS-2023-07-40). Briefly, fresh semitendinosus muscles were harvested from 1 week-old piglets. Dovitinib (MedChemExpress, HY-50905) and methylergometrine (Expogin®, 0460228) were the test compounds. Their respective vehicle controls were dimethyl sulfoxide (DMSO) and phosphate-buffered saline (PBS).2.5 Cell viability assay
C2C12 myoblasts were seeded at a density of 2 × 103 cells per well in 96-well plates and cultured in GM for 24 h. Cells were then treated with dovitinib (0.3125, 0.625, 1.25, 2.5, and 5 µM) or methylergometrine (1.25, 2.5, 5, 10, and 20 µM) for 24 or 48 h. Cell viability was evaluated using MTT assay. Briefly, cells were incubated with 0.5% MTT solution (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) (Sigma-Aldrich, M5655) for 2 h at 37 °C to allow the formation of insoluble formazan crystals. Following incubation, the supernatant was carefully aspirated, and the formazan crystals were dissolved in 100% DMSO. Absorbance was measured at 570 nm using a Multiskan GO microplate spectrophotometer (Thermo Scientific, USA).2.6 Myogenic differentiation assessment
To evaluate the effects of the tested compounds on myogenesis, C2C12 cells were used to assess early and late differentiation phases. Cells were cultured in GM until reaching ∼90% confluency and culture medium was replaced with differentiation medium (DM), consisting of DMEM supplemented with 2% (v/v) horse serum (HS; Gibco, 16050-130) and 1% penicillin/streptomycin at day 0. For early differentiation assessment (SI Fig. 1A), cells were plated at 7.5 × 103 cells per well and treated on differentiation day 0. For late differentiation assessment (SI Fig. 1B), cells were plated at 1.2 × 105 cells per well with treatments administered on differentiation days 3–4. Cells were treated with a range of concentrations of dovitinib (0–5 µM) or methylergometrine (0–20 µM). For PMSCs, cells were cultured and treated to assess myogenic differentiation as follows. PMSCs (passage 4) were plated at a density of 1 × 104 cells per well in iMatrix-coated 24-well plates and cultured in DMEM supplemented with 20% FBS, 5 ng mL−1 basic fibroblast growth factor (bFGF), and 20 µM p38 inhibitor SB203580 (13067, Cayman Chemical, USA) and 1% penicillin/streptomycin for 4 days (SI Fig. 1C). After reaching ∼90% confluency, the culture medium was switched to DMEM supplemented with 2% HS to induce differentiation. PMSCs were treated with dovitinib (0–1.25 µM) or methylergometrine (0–10 µM) on days 0, 2, 4, and 6 of the differentiation phases. Culture medium was replaced every other day.2.7 Immunofluorescence staining
Cells were fixed with 4% paraformaldehyde (Sigma-Aldrich, 818715.0100) overnight at 4 °C. Following fixation, cells were permeabilized with 0.1% Triton X-100 (Scharlau, TR0447) and blocked with 5% normal goat serum (Invitrogen, PCN5000). The primary antibody against myosin heavy chain (MHC) (Merck Millipore, 05-716) was incubated for 2 h at room temperature. Subsequently, cells were incubated for 1 h with Alexa Fluor™ 594-conjugated goat anti-mouse Ig cross-adsorbed secondary antibody (Invitrogen, A-11005) in the dark. Nuclei were counterstained for 5 min with 4′,6-diamidino-2-phenylindole (DAPI; Invitrogen, D1306). Images were captured using an inverted fluorescence microscope (Olympus IX83) equipped with an ORCA-Flash 2.8 digital CMOS camera (Hamamatsu Photonics, C11440). Quantitative analysis was performed using OLYMPUS cellSens Dimension Desktop 4.3.1 software. The area fraction (%) of MHC-positive staining was measured as a key metric of myogenic differentiation from 15 randomly captured images. The fusion index was calculated as follows: the percentage of nuclei within multinucleated myotubes (defined as having ≥3 nuclei for early differentiation or ≥10 nuclei for late differentiation) relative to the total number of nuclei in the field. For the calculation of the differentiation index in PMSCs, all nuclei located within MHC-positive myotubes were counted. Fusion and differentiation indices were acquired from the randomization of 6 images.2.8 Western blot analysis
Cellular protein was extracted using RIPA buffer (50 mM Tris–HCl pH 7.5, 150 mM NaCl, 1 mM EDTA, 1% Triton X-100) supplemented with a protease inhibitor cocktail (Sigma-Aldrich, P8340) and a phosphatase inhibitor cocktail (Merck Millipore, 524625). Following centrifugation at 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000×g for 15 min, the supernatant was collected, and protein concentration was determined via a BCA assay (Thermo Scientific). Protein samples (10 µg per lane) were denatured and resolved on a SDS-polyacrylamide gel. The resolved proteins were then transferred to a nitrocellulose membrane (BIO-RAD, 1620112). Membranes were blocked with 5% nonfat milk and incubated overnight at 4 °C with primary antibodies for myogenin (MyoG) (1
000×g for 15 min, the supernatant was collected, and protein concentration was determined via a BCA assay (Thermo Scientific). Protein samples (10 µg per lane) were denatured and resolved on a SDS-polyacrylamide gel. The resolved proteins were then transferred to a nitrocellulose membrane (BIO-RAD, 1620112). Membranes were blocked with 5% nonfat milk and incubated overnight at 4 °C with primary antibodies for myogenin (MyoG) (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200, Santa Cruz Biotechnology, SC-12732) and MHC (1
200, Santa Cruz Biotechnology, SC-12732) and MHC (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2,000, Merck Millipore, 05-716). For loading control, a primary antibody against GAPDH (1
2,000, Merck Millipore, 05-716). For loading control, a primary antibody against GAPDH (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2,000, Cell Signaling Technology, 2118) was used. Secondary horseradish peroxidase (HRP)-conjugated antibodies (Cell Signaling Technology, anti-mouse 7076 and anti-rabbit 7074) were incubated for 1 h at room temperature. Protein bands were visualized using a chemiluminescent HRP detection reagent (Clarity™ Western ECL Substrate, BIO-RAD, 170-5060) and imaged with an Alliance Q9 Advanced Chemiluminescence Imager (UVITEC CAMBRIDGE). Quantification of protein expression levels was performed using ImageJ software (version 1.54g, NIH).
2,000, Cell Signaling Technology, 2118) was used. Secondary horseradish peroxidase (HRP)-conjugated antibodies (Cell Signaling Technology, anti-mouse 7076 and anti-rabbit 7074) were incubated for 1 h at room temperature. Protein bands were visualized using a chemiluminescent HRP detection reagent (Clarity™ Western ECL Substrate, BIO-RAD, 170-5060) and imaged with an Alliance Q9 Advanced Chemiluminescence Imager (UVITEC CAMBRIDGE). Quantification of protein expression levels was performed using ImageJ software (version 1.54g, NIH).
2.9 Statistical analysis
Statistical significance was evaluated using GraphPad Prism version 10. For comparisons involving multiple treatment groups against a single control, a one-way ANOVA was performed, followed by Dunnett's multiple comparisons test. For direct pairwise comparisons between two groups, an unpaired two-tailed Student's t-test with Welch's correction was used, otherwise stated. A p-value of <0.05 was considered statistically significant.3. Results and discussion
3.1 In silico compound screening
To identify novel modulators of FGFR1 tyrosine kinase (FGFR1 TK), we initiated a comprehensive, multi-stage in silico screening campaign. Our primary goal was to find compounds that not only exhibit high binding affinity for FGFR1 but also possess favorable pharmacological properties and are economically viable for large-scale applications such as cultured meat production. The initial step involved a docking-based virtual screen of 872 pre-optimized ligands against the FGFR1 TK domain. Using dovitinib, a known FGFR1 inhibitor with a predicted binding energy (BE) of −8.8 kcal mol−1, as a reference, we prioritized compounds with a BE of −8.8 kcal mol−1 or lower. This first round of screening yielded a substantial pool of 261 compounds with potentially high binding affinity to FGFR1 (Fig. 1A). To mitigate the risk of off-target effects, a crucial consideration for any therapeutic or biotechnological compound, we performed a secondary screening against p38α mitogen-activated protein kinase (p38α MAPK). This kinase is a pivotal regulator of terminal muscle cell differentiation.37 Disruption of p38α MAPK signaling could counteract the desired myogenic effect, even if a compound effectively inhibits FGFR1. Therefore, we sought compounds with a lower binding affinity (i.e., higher BE) for p38α MAPK. This counter-screening strategy, using the known p38α MAPK inhibitor SCIO-469 (BE = −10.4 kcal mol−1) as a benchmark, successfully filtered out compounds with potential off-target binding. This refined approach ultimately narrowed our selection to 20 promising candidates (15 nuclear receptor compounds and 5 in-house compounds) that showed both strong FGFR1 binding and minimal predicted p38α MAPK interaction (BE range: −6.0 to −9.3 kcal mol−1; Fig. 1B).Beyond molecular affinity, practical criteria for a functional compound include solubility and cost. Poor solubility can severely limit a compound's bioavailability and efficacy.38–40 We therefore used the SwissADME web server to predict the solubility of the 20 shortlisted compounds via three distinct models: ESOL, Ali, and SILICOS-IT.41 A compound was considered “soluble” if at least two of the three models concurred, a stringent criterion that further reduced our candidate pool to five compounds: nr848, nr796, nr101, nr12, and inh-1 (Fig. 1C). Finally, we evaluated the cost-effectiveness of these five candidates, a critical factor for the commercial viability of cultured meat. Sourcing pricing information from publicly available databases and, in the case of methylergometrine (inh-1), we found that methylergometrine was by far the most affordable option. This multi-criteria analysis, which considered FGFR1 binding affinity, off-target binding, solubility, and cost, led to the selection of methylergometrine as our primary hit compound. Its 2D chemical structure is depicted in Fig. 1D, and its suitability for further investigation was strongly supported by our comprehensive screening funnel.
3.2 Structural dynamics and binding recognition
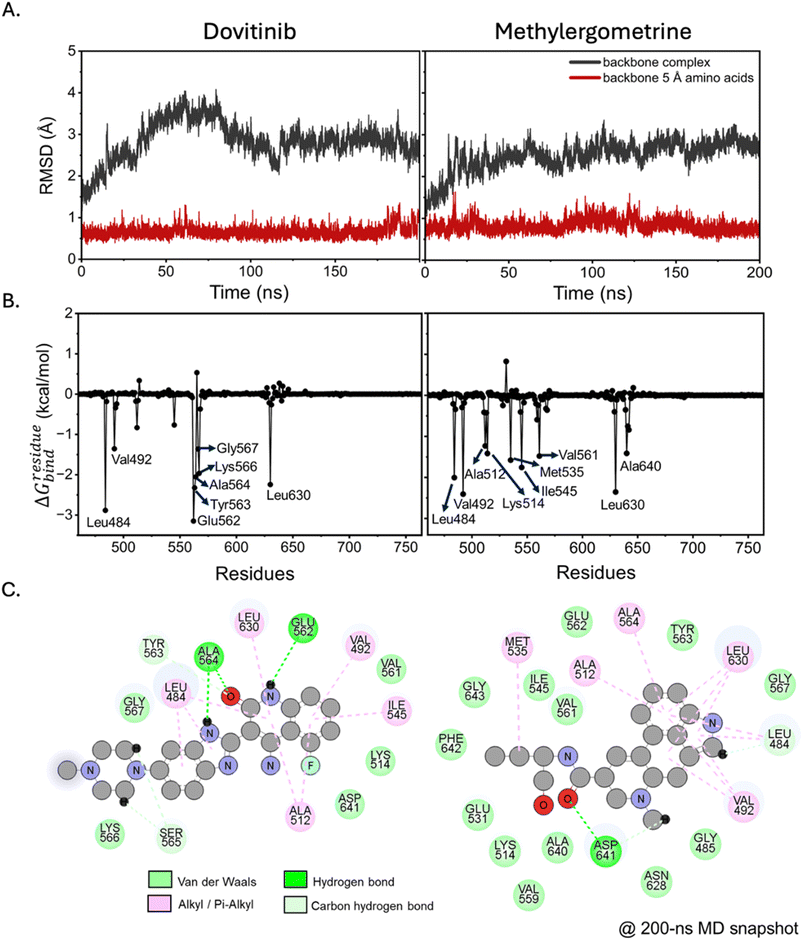
To gain deeper insight into the binding mechanism of our selected compounds, we conducted MD simulations of dovitinib and methylergometrine in complex with FGFR1 TK. We first assessed the stability of the protein–ligand complexes by analyzing the root-mean-square deviation (RMSD) of the protein backbone. As shown in Fig. 2A, the RMSD profiles for both complexes indicated a stable system, with fluctuations primarily originating from distal, flexible regions of the protein rather than the core binding site. The methylergometrine–FGFR1 complex showed remarkable stability with less fluctuation compared to the dovitinib complex, suggesting that methylergometrine forms a robust and stable complex in a dynamic, aqueous environment. To identify the key amino acids involved in binding, we performed a per-residue free energy decomposition analysis on the stable trajectories (180–200 ns). We defined “hotspot” residues as those with a binding free energy decomposition (ΔGresiduebind) of <−1.00 kcal mol−1. This analysis revealed distinct yet overlapping binding patterns. For dovitinib, key residues included Leu484, Val492, Glu562, Tyr563, Ala564, Lys566, Gly567, and Leu630 (Fig. 2B-left). Methylergometrine, on the other hand, interacted with Leu484, Val492, Ala512, Lys514, Met535, Ile545, Val561, Leu630, and Ala640 (Fig. 2B-right). Intriguingly, both compounds shared interactions with Leu484, Val492, and Leu630, suggesting they occupy a similar binding pocket within the FGFR1 active site. These residues have been previously identified as crucial for the binding of other FGFR1 TK inhibitors like AZD4547 and TKI258,23,42,43 highlighting a shared mechanism of inhibition. Methylergometrine's particularly strong interaction with Val492 and Leu630 further underscores its potential as a potent FGFR1 modulator. The visualization of non-covalent interactions from the final MD snapshots (Fig. 2C) confirmed that hydrophobic interactions were the dominant force driving the binding of both compounds. Dovitinib formed hydrophobic bonds with Leu484, Val492, Ala512, Ile545, Leu630, and Met535, while methylergometrine interacted hydrophobically with Ala512, Ala564, Leu630, and Val492. These findings are consistent with the negative van der Waals energy values observed in the MM/GBSA analysis (Table 1), indicating the critical role of these interactions in stabilizing the protein–ligand complexes.| Energy components | Dovitinib | Methylergometrine |
|---|---|---|
| Gas phase (MM) | ||
| ΔEelectrostatic | −24.8905 ± 2.9841 | −22.4979 ± 5.0582 |
| ΔEvdW | −41.2709 ± 3.7583 | −46.4489 ± 3.0806 |
| ΔEgas | −66.1614 ± 5.5764 | −68.9468 ± 4.9408 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||
| Solvation (GBSA) | ||
| ΔGpolar | 30.9189 ± 3.3940 | 26.4121 ± 3.8196 |
| ΔGnonpolar | −4.5782 ± 0.2700 | −6.0067 ± 0.1635 |
| ΔGsolv | 26.3407 ± 3.3494 | 20.4054 ± 3.8826 |
| ΔH | −39.8207 ± 5.2667 | −48.5414 ± 4.9783 |
| −TΔS | 17.9829 ± 6.1009 | 25.4924 ± 4.7232 |
| ΔGbind (MM/GBSA) | −21.8378 ± 8.0597 | −23.0490 ± 6.8624 |
3.3 End-point binding free energy
To quantitatively compare the binding affinities of dovitinib and methylergometrine, we used the end-point MM/GBSA method on the final 20 ns of the MD trajectories. As detailed in Table 1, both compounds exhibited strong, comparable binding affinities, with binding free energy (ΔGbind) values of −21.84 ± 8.06 kcal mol−1 for dovitinib and −23.05 ± 6.86 kcal mol−1 for methylergometrine. The energy decomposition revealed that for both compounds, the binding was primarily driven by the favorable van der Waals interactions (ΔEvdW) in the gas phase. The electrostatic contribution (ΔEelectrostatic) was also favorable but approximately half the magnitude of the van der Waals term, reinforcing the dominance of hydrophobic forces. Interestingly, methylergometrine showed a more favorable solvation energy (ΔGsolv) but also a higher entropic penalty, indicating that while it fits well into the binding pocket, its binding is associated with a greater loss of conformational freedom. Overall, these results from MD simulations are highly consistent with the molecular docking predictions and provide strong theoretical support for the experimental validation.3.4 Experimental validation with STD-NMR spectroscopy
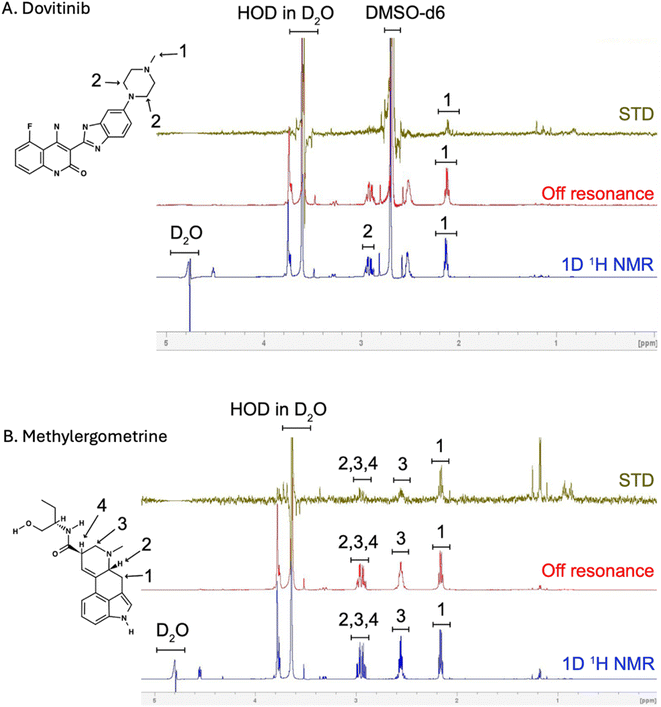
To provide direct experimental evidence of the compound–protein interactions, we performed STD-NMR spectroscopy. For dovitinib, the STD-NMR spectrum showed a clear signal at a single proton position (position 1, Fig. 3A), confirming a direct interaction with the recombinant FGFR1 protein. However, this finding was discordant with our MD simulations, which did not predict this specific proton to be in close proximity to the protein. This discrepancy likely stems from the inherent differences between static computational models and the dynamic, in-solution environment of NMR. The high RMSD fluctuations observed in the dovitinib complex during MD simulations also point to a highly flexible binding mode or potential for multiple transient binding conformations, which could explain the difference between the computational and experimental results. In contrast, the STD-NMR results for methylergometrine were highly consistent with our MD simulations. A strong STD signal was detected at proton position 1 (Fig. 3B), which the MD simulation predicted to be in close contact with the key residue Val492. Weaker, but still detectable, STD signals were also observed at positions 2, 3, and 4. Position 2 was computationally placed near Val492, while positions 3 and 4 were near Asp641, a residue that formed a hydrogen bond with the ligand's neighboring oxygen. The convergence of these experimental and computational findings provides compelling evidence that methylergometrine directly and specifically binds to FGFR1 through multiple contact points, making it a robust candidate for modulating its activity.3.5 Enhancing C2C12 differentiation with dovitinib and methylergometrine
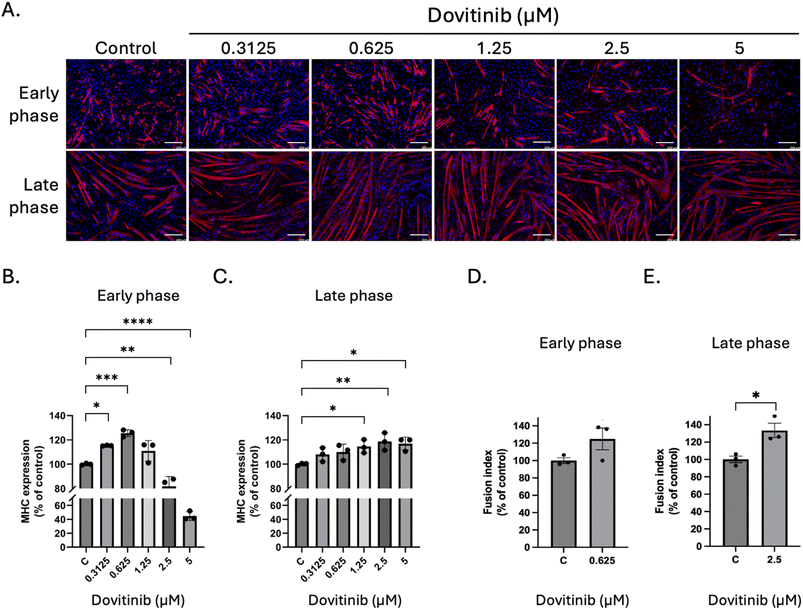
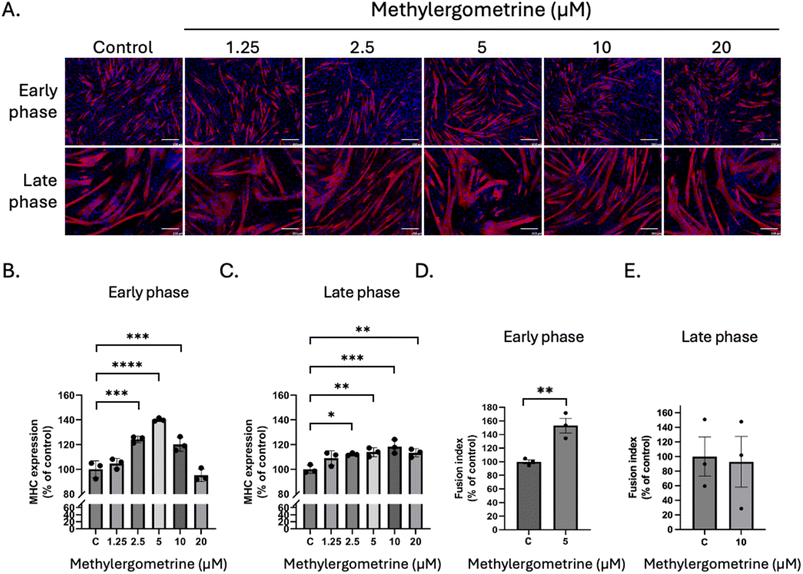
To establish a robust in vitro screening framework, we employed the immortalized C2C12 murine cell line,44 a strategy underpinned by the high evolutionary conservation of the FGFR1 tyrosine kinase domain across human, mouse, and porcine species (SI Fig. 2). Preliminary safety profiling revealed distinct cytotoxic signatures for each candidate. While methylergometrine demonstrated high biocompatibility with IC50 values consistently exceeding 20 µM at both 24 and 48 h, dovitinib exhibited a time-dependent cytotoxic profile, with the IC50 shifting from >5 µM at 24 h to 2.1 µM at 48 h. These pharmacological thresholds informed the selection of sub-toxic concentrations for subsequent functional assays, wherein the pro-myogenic efficacy of both dovitinib and methylergometrine was evaluated across the discrete early and late phases of myogenic differentiation.In the subsequent experiment, the effects of dovitinib and methylergometrine were assessed on C2C12 myogenic differentiation during both early and late myogenic differentiation phases. At low concentrations (0.3125–0.625 µM), dovitinib significantly enhanced the expression of MHC, a key marker of myotube formation, in the early phase (Fig. 4A and B). This effect was most pronounced at 0.625 µM, where MHC expression was ∼123% of the control (p < 0.001). Interestingly, higher concentrations were detrimental, reducing myotube formation. In the late phase, a dose-dependent increase in myotube size was observed (Fig. 4A), and quantitative analysis confirmed a significant increase in MHC expression from 1.25 to 5 µM (Fig. 4C). The fusion index, a measure of myoblast fusion into multinucleated myotubes.45,46 A slight, non-significant increase was observed at 0.625 µM during the early phase (Fig. 4D), whereas a significant increase was detected in the late phase at 2.5 µM (∼134% of control, p < 0.05) (Fig. 4E), suggesting that dovitinib primarily promotes myotube maturation and fusion rather than the initial commitment to differentiation. Next, we investigated methylergometrine's effects. In the early phase, methylergometrine significantly enhanced MHC expression at 5 µM (∼140% of control, Fig. 5A and B), with a notable increase in myotube size and length. In the late phase, MHC expression showed a gradual, dose-dependent increase from 2.5 to 20 µM (Fig. 5C), with peak effects at 20 µM. The fusion index in the early phase at 5 µM (∼150% of control, Fig. 5D) was also significantly increased, indicating a potent ability to drive early fusion events, whereas no significant change was observed in the late phase at 10 µM (Fig. 5E). These immunofluorescence results suggest a phase-specific effect for each compound. Dovitinib appears to be most effective during the later stages of myogenesis, promoting myotube maturation, while methylergometrine is a powerful driver of early myoblast fusion.
In this study, serum-containing medium was utilized as it provides a complex physiological background rich in pro-proliferative growth factors, which naturally suppress the onset of myogenic differentiation. By demonstrating that dovitinib and methylergometrine can effectively trigger terminal differentiation even within this mitogen-heavy environment, we provide a robust proof-of-concept for their ability to physiologically override endogenous proliferative signals. Furthermore, utilizing serum-containing medium baseline minimizes potential confounding variables—such as non-specific cellular stress or metabolic stagnation—that are frequently associated with unoptimized serum-free formulations. This ensures that the accelerated myotube formation observed is a direct result of targeted FGFR1 modulation rather than a secondary response to nutrient deprivation.
3.6 Dovitinib and methylergometrine increase MHC and MyoG expression during myogenic differentiation
To corroborate our immunofluorescence findings, we performed western blot analysis to quantify the expression levels of key myogenic markers: MyoG (a master regulator of myogenic differentiation) and MHC. Consistent with our imaging data, both dovitinib and methylergometrine significantly upregulated MyoG and MHC expression in a dose- and phase-dependent manner. For dovitinib, early-phase treatment at 0.625 µM led to a dramatic ∼4.5-fold increase in MHC expression and a ∼3.4-fold increase in MyoG at 0.3125 µM (Fig. 6A and C). In the late phase, both markers showed robust, dose-dependent increases (Fig. 6B and D), further supporting dovitinib's role in promoting myotube maturation. Similarly, methylergometrine significantly upregulated MyoG and MHC expression. In the early phase, MHC peaked at 5 µM (∼2.6-fold increase) and MyoG at 1.25 µM (∼2.2-fold increase) (Fig. 6E and G). Late-phase treatment also resulted in a significant and dose-dependent increase for both markers (Fig. 6F and H). Collectively, these results from both imaging and western blot analyses paint a clear picture: dovitinib and methylergometrine are potent enhancer of myogenic differentiation, but they appear to act with slightly different temporal profiles. Dovitinib seems to be a more effective molecular switch for priming differentiation and driving terminal maturation, while methylergometrine shows a strong capacity for enhancing myoblast fusion and morphological development, particularly in the early phase. Beyond the induction of well-known myogenic markers, efficient myogenic differentiation for cultured meat applications requires coordinated regulation of cellular metabolism to sustain high levels of protein synthesis and tissue maturation. FGFR1 signaling has been reported to intersect with the PI3K–AKT–mTOR axis, a pathway that promotes cellular growth.47 Therefore, the observed upregulation of MyoG and MHC in this study may reflect enhanced myogenic lineage commitment and an increased capacity for muscle-specific protein synthesis, which is essential for generating sufficient biomass and maintaining structural integrity in a cultured meat system. Myogenic differentiation may also be accompanied by metabolic remodeling, including increased mitochondrial biogenesis and a shift toward oxidative metabolism, to meet the elevated energy demands associated with myotube formation and maturation.48,49 Modulation of FGFR1 signaling may thus indirectly support mitochondrial function, ensuring adequate ATP supply during differentiation. In parallel, coordinated regulation of lipid and fatty acid metabolism has been reported during myogenesis, contributing to membrane expansion, energy homeostasis, and the development of muscle tissue architecture.50 Although mitochondrial activity and lipid metabolism were not directly assessed in the present study, these processes represent important downstream consequences of FGFR1 modulation and are particularly relevant in the context of cultured meat production, where both differentiation efficiency and metabolic robustness critically influence yield, consistency, and product quality.3.7 Translational validation in porcine muscle stem cells (PMSCs)
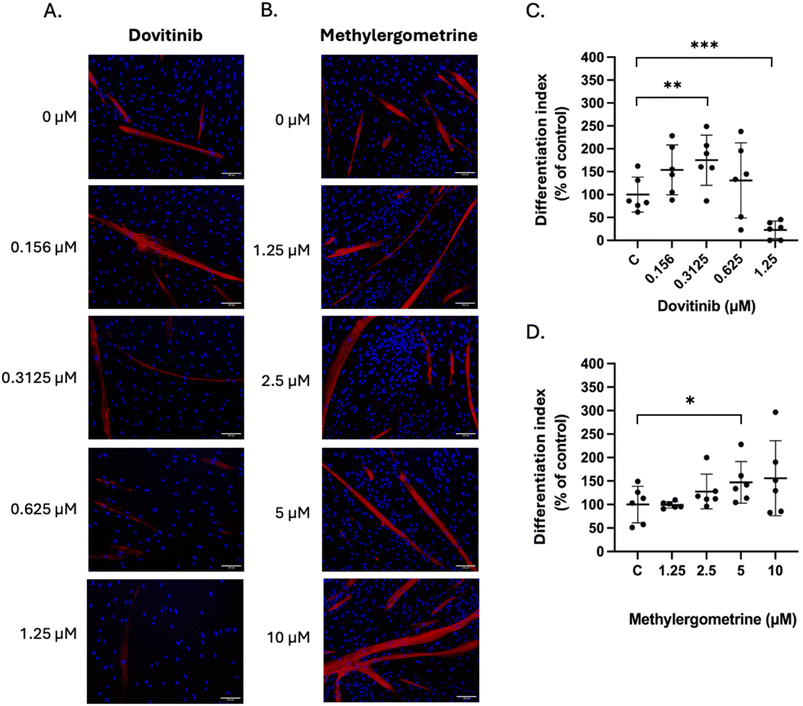
To evaluate the translational viability of our findings for cellular agriculture, we validated the pro-myogenic efficacy of dovitinib and methylergometrine in PMSCs. Immunofluorescence analysis of MHC expression revealed divergent pharmacological profiles for the two compounds (Fig. 7A and B). Dovitinib exhibited a narrow, biphasic therapeutic window; while a low concentration (0.3125 µM) significantly enhanced the differentiation index (Fig. 7C), a dose of 1.25 µM proved cytotoxic, markedly suppressing both MHC expression and total cell viability. In sharp contrast, methylergometrine demonstrated a robust, dose-dependent promotive effect on myotube formation, with 5 µM significantly increasing the differentiation index relative to controls (Fig. 7D). This was further corroborated by linear regression analysis, which identified a significant positive dose–response relationship for methylergometrine (p = 0.017), a correlation notably absent for dovitinib due to its inhibitory effects at higher concentrations. These species-specific results underscore the necessity of validating hits from immortalized models in primary target cells. Given their distinct kinetic profiles, we propose a strategic, sequential application: low-dose dovitinib may serve as an initial molecular trigger for the myogenic program, while methylergometrine drives sustained cellular fusion and structural maturation, providing a dual-action framework for optimizing high-quality muscle tissue formation in cultured meat production.4. Conclusion
This study identifies and characterizes dovitinib and methylergometrine as potent modulators of myogenic differentiation through FGFR1 inhibition. Targeted modulation of FGFR1 signaling enhances myogenic progression by upregulating key differentiation makers, MyoG and MHC, and promoting myotube formation in both C2C12 and PMSCs. This phenomenon can support cellular programs that are essential for muscle tissue formation, including increased protein synthesis capacity and metabolic adaptation during myogenesis. Notably, the phase-dependent effects observed for dovitinib and methylergometrine suggest a potential dual-phase differentiation strategy, in which early and late fusion events of myotube formation can be selectively enhanced. Dovitinib serves as the gold-standard inhibitor to prove the mechanism. Methylergometrine, while a pharmaceutical, provides a scaffold for a more affordable, stable, and generally recognized as safe derivative search. These are lead compounds meant to pave the way for identifying natural, food-grade FGFR1 antagonists such as phytochemicals with similar binding motifs. This concept is particularly relevant to cultured meat production, where efficient differentiation and metabolic stability are critical determinants of tissue quality and production consistency. As FGFR1 modulation likely influences the metabolic partitioning of nutrients into protein versus intramyocellular lipids, future studies may include a full lipidomic profile or total crude protein analysis. Together, these findings support FGFR1 modulation by small molecules as a practical approach to improving muscle tissue formation in scalable cultured meat systems.Author contributions
All authors have full access to the data and accept responsibility for publication. SU, US, and TJ have accessed and verified all data reported in this study. All authors have read, reviewed, and agreed to the final manuscript. US and TJ conceptualised the study. RS, US, TJ designed the experiments. SU, PC, SK, SW, WW, RS, TS, DL and US performed the investigation. SU, SK, RS, US and TJ performed the formal analysis. SU, US, and TJ curated the data. TJ acquired funding and did the project administration. SK, RS, DL, US and TJ provided resources. TJ supervised the project. SU and US wrote the original draft. All authors reviewed and edited the manuscript.Conflicts of interest
There are no conflicts to declare.Abbreviations
| PMSCs | Porcine muscle stem cells |
| FGFR1 | Fibroblast growth factor receptor 1 |
| MD | Molecular dynamics |
| NMR | Nuclear magnetic resonance |
| STD | Saturation transfer difference |
| DMEM | Dulbecco's modified Eagle's medium |
| PBS | Phosphate-buffered saline |
| DM | Differentiation medium |
| HRP | Horseradish peroxidase |
| FGFR1 TK | FGFR1 tyrosine kinase |
| BE | Binding energy |
| MAPK | Mitogen-activated protein kinase |
| RMSD | Root-mean-square deviation |
Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5fb00756a.Acknowledgements
This research project is supported by the NSRF via the Program Management Unit for Human Resources & Institutional Development, Research and Innovation (B05F650014). We acknowledge the Second Century Fund (C2F), Chulalongkorn University for a postdoctoral fellowship to SU.References
- G. Wu, Dietary protein intake and human health, Food Funct., 2016, 7, 1251–1265 RSC.
- J. Bauer, et al., Evidence-based recommendations for optimal dietary protein intake in older people: a position paper from the PROT-AGE Study Group, J. Am. Med. Dir. Assoc., 2013, 14(8), 542–559 CrossRef PubMed.
- M. J. Post, Cultured meat from stem cells: challenges and prospects, Meat Sci., 2012, 92(3), 297–301 CrossRef PubMed.
- C. Bryant and J. Barnett, Consumer acceptance of cultured meat: a systematic review, Meat Sci., 2018, 143, 8–17 CrossRef PubMed.
- J. R. Blanton Jr, et al., Isolation of two populations of myoblasts from porcine skeletal muscle, Muscle Nerve, 1999, 22(1), 43–50 CrossRef CAS PubMed.
- K. J. Wilschut, et al., Isolation and characterization of porcine adult muscle-derived progenitor cells, J. Cell. Biochem., 2008, 105(5), 1228–1239 CrossRef CAS PubMed.
- I. T. Kadim, et al., Cultured meat from muscle stem cells: a review of challenges and prospects, J. Integr. Agric., 2015, 14(2), 222–233 CrossRef CAS.
- X. Guan, et. al., Production of mature myotubes in vitro improves the texture and protein quality of cultured pork, Food Funct., 2023, 14, 3576–3587 RSC.
- T. Ma, et al., Transdifferentiation of fibroblasts into muscle cells to constitute cultured meat with tunable intramuscular fat deposition, Elife, 2024, 13, RP93220 CrossRef CAS PubMed.
- L. Melzener, Optimisation of cell fate determination for cultured muscle differentiation, Commun. Biol., 2024, 7(1), 1493 CrossRef CAS PubMed.
- C. Dombrowski, et al., FGFR1 signaling stimulates proliferation of human mesenchymal stem cells by inhibiting the cyclin-dependent kinase inhibitors p21(Waf1) and p27(Kip1), Stem Cell., 2013, 31(12), 2724–2736 CrossRef CAS PubMed.
- Y. Feng, et al., A feedback circuit between miR-133 and the ERK1/2 pathway involving an exquisite mechanism for regulating myoblast proliferation and differentiation, Cell Death Dis., 2013, 4(11), e934 CrossRef CAS PubMed.
- S. Zhan, et al., LncR-133a Suppresses Myoblast Differentiation by Sponging miR-133a-3p to Activate the FGFR1/ERK1/2 Signaling Pathway in Goats, Genes, 2022, 13(5), 818 CrossRef CAS PubMed.
- H. K. Li, et al., LRTM1 promotes the differentiation of myoblast cells by negatively regulating the FGFR1 signaling pathway, Exp. Cell Res., 2020, 396(1), 112237 CrossRef CAS PubMed.
- D. Liang, et al., Insight into resistance mechanisms of AZD4547 and E3810 to FGFR1 gatekeeper mutation via theoretical study, Drug Des., Dev. Ther., 2017, 11, 451–461 CrossRef CAS PubMed.
- L. Perdios, et al., Conformational transition of FGFR kinase activation revealed by site-specific unnatural amino acid reporter and single molecule FRET, Sci. Rep., 2017, 7, 39841 CrossRef CAS PubMed.
- S. Mahapatra, et al., Molecular dynamics simulations reveal phosphorylation-induced conformational dynamics of the fibroblast growth factor receptor 1 kinase, J. Biomol. Struct. Dyn., 2024, 42(6), 2929–2941 CrossRef CAS PubMed.
- O. Cala, F. Guillière and I. Krimm, NMR-based analysis of protein-ligand interactions, Anal. Bioanal. Chem., 2014, 406(4), 943–956 CrossRef CAS PubMed.
- M. Mayer and B. Meyer, Characterization of Ligand Binding by Saturation Transfer Difference NMR Spectroscopy, Angew Chem. Int. Ed. Engl., 1999, 38(12), 1784–1788 CrossRef CAS PubMed.
- F. André, et al., Targeting FGFR with dovitinib (TKI258): preclinical and clinical data in breast cancer, Clin. Cancer Res., 2013, 19(13), 3693–3702 CrossRef PubMed.
- Y. Lee, et al., A Receptor Tyrosine Kinase Inhibitor, Dovitinib (TKI-258), Enhances BMP-2-Induced Osteoblast Differentiation In Vitro, Mol. Cells, 2016, 39(5), 389–394 CrossRef CAS PubMed.
- B. T. Bateman, et al., Methylergonovine maleate and the risk of myocardial ischemia and infarction, Am. J. Obstet. Gynecol., 2013, 209(5), 459.e1–459.e13 CrossRef CAS PubMed.
- J. A. Tucker, et al., Structural insights into FGFR kinase isoform selectivity: diverse binding modes of AZD4547 and ponatinib in complex with FGFR1 and FGFR4, Structure, 2014, 22(12), 1764–1774 CrossRef CAS PubMed.
- R. Azevedo, et al., X-ray structure of p38α bound to TAK-715: comparison with three classic inhibitors, Acta Crystallogr., Sect. D: Biol. Crystallogr., 2012, 68(Pt 8), 1041–1050 CrossRef CAS PubMed.
- N. M. O'Boyle, et al., Open Babel: an open chemical toolbox, J. Cheminf., 2011, 3, 33 Search PubMed.
- M. R. Koebel, et al., AutoDock VinaXB: implementation of XBSF, new empirical halogen bond scoring function, into AutoDock Vina, J. Cheminf., 2016, 8, 27 Search PubMed.
- E. Jurrus, et al., Improvements to the APBS biomolecular solvation software suite, Protein Sci., 2018, 27(1), 112–128 CrossRef CAS PubMed.
- C. Tian, et al., ff19SB: Amino-Acid-Specific Protein Backbone Parameters Trained against Quantum Mechanics Energy Surfaces in Solution, J. Chem. Theory Comput., 2020, 16(1), 528–552 CrossRef PubMed.
- T. Darden, et al., Particle Mesh Ewald: An N·log(N) Method for Ewald Sums in Large Systems, J. Chem. Phys., 1993, 98, 10089–10092 CrossRef CAS.
- H. J. C. P. Berendsen, et al., Molecular dynamics with coupling to an external bath, J. Chem. Phys., 1984, 81(8), 3684–3690 CrossRef CAS.
- B. P. Uberuaga, M. Anghel and A. F. Voter, Synchronization of trajectories in canonical molecular-dynamics simulations: observation, explanation, and exploitation, J. Chem. Phys., 2004, 120(14), 6363–6374 CrossRef CAS PubMed.
- M. G. Paterlini and D. M. Ferguson, Constant temperature simulations using the Langevin equation with velocity Verlet integration, Chem. Phys., 1998, 236(1), 243–252 CrossRef CAS.
- J.-P. Ryckaert, G. Ciccotti and H. J. C. Berendsen, Numerical integration of the Cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes, J. Comput. Phys., 1977, 23(3), 327–341 CrossRef CAS.
- D. R. Roe and T. E. Cheatham 3rd, PTRAJ and CPPTRAJ: Software for Processing and Analysis of Molecular Dynamics Trajectory Data, J. Chem. Theory Comput., 2013, 9(7), 3084–3095 CrossRef CAS PubMed.
- B. R. Miller 3rd, et al., MMPBSA.py: An Efficient Program for End-State Free Energy Calculations, J. Chem. Theory Comput., 2012, 8(9), 3314–3321 CrossRef PubMed.
- U. Suriya, et al., A diarylheptanoid derivative mediates glycogen synthase kinase 3β to promote the porcine muscle satellite cell proliferation: implications for cultured meat production, Biochem. Biophys. Res. Commun., 2024, 736, 150850 CrossRef CAS PubMed.
- C. Cabane, et al., Regulation of C2C12 myogenic terminal differentiation by MKK3/p38alpha pathway, Am. J. Physiol. Cell Physiol., 2003, 284(3), C658–C666 CrossRef CAS PubMed.
- L. Di, P. V. Fish and T. Mano, Bridging solubility between drug discovery and development, Drug Discov. Today, 2012, 17(9–10), 486–495 CrossRef CAS PubMed.
- J. Alsenz and M. Kansy, High throughput solubility measurement in drug discovery and development, Adv. Drug Delivery Rev., 2007, 59(7), 546–567 CrossRef CAS PubMed.
- T. Kennedy, Managing the drug discovery/development interface, Drug Discovery Today, 1997, 2(10), 436–444 CrossRef.
- A. Daina, O. Michielin and V. Zoete, SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules, Sci. Rep., 2017, 7, 42717 CrossRef PubMed.
- R. Leechaisit, et al., Design, Synthesis, Biological Evaluation and Molecular Modeling Studies of Novel Naphthoquinone-Triazole Hybrids as Potential FGFR1 Tyrosine Kinase Inhibitors, Asian J. Org. Chem., 2025, 14(5), e202400813 CrossRef CAS.
- T. D. Bunney, et al., The Effect of Mutations on Drug Sensitivity and Kinase Activity of Fibroblast Growth Factor Receptors: A Combined Experimental and Theoretical Study, EBioMedicine, 2015, 2(3), 194–204 CrossRef PubMed.
- A. P. Sharples, et al., Modelling in vivo skeletal muscle ageing in vitro using three-dimensional bioengineered constructs, Aging Cell, 2012, 11(6), 986–995 CrossRef CAS PubMed.
- L. Terrie, et al., Enhancing Myoblast Fusion and Myotube Diameter in Human 3D Skeletal Muscle Constructs by Electromagnetic Stimulation, Front. Bioeng. Biotechnol., 2022, 10, 892287 CrossRef PubMed.
- S. Noë, et al., The Myotube Analyzer: how to assess myogenic features in muscle stem cells, Skeletal Muscle, 2022, 12(1), 12 CrossRef PubMed.
- D. Zhang, et al., FGFR1 Induces Acquired Resistance against Gefitinib by Activating AKT/mTOR Pathway in NSCLC, OncoTargets Ther., 2019, 12, 9809–9816 CrossRef CAS PubMed.
- A. Wagatsuma and K. Sakuma, Mitochondria as a potential regulator of myogenesis, Sci. World J., 2013, 2013, 593267 CrossRef PubMed.
- J. Sin, et al., Mitophagy is required for mitochondrial biogenesis and myogenic differentiation of C2C12 myoblasts, Autophagy, 2016, 12(2), 369–380 CrossRef CAS PubMed.
- P. Fabre, et al., Bioactive lipid mediator class switching regulates myogenic cell progression and muscle regeneration, Nat. Commun., 2025, 16(1), 5578 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |