Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
A conserved glutamate orchestrates transitions between catalytic intermediates in [NiFe]-hydrogenase
Armel F. T. Waffo a,
Christian Lorent
a,
Christian Lorent a,
Sagie Katza,
Cornelius C. M. Bernitzky
a,
Sagie Katza,
Cornelius C. M. Bernitzky b,
Janna Schoknechta,
Marius Horch
b,
Janna Schoknechta,
Marius Horch b,
Oliver Lenz
b,
Oliver Lenz a,
Giorgio Caserta
a,
Giorgio Caserta *a and
Ingo Zebger
*a and
Ingo Zebger *a
*a
aInstitut für Chemie, Sekr. PC14, Technische Universität Berlin, D-10623 Berlin, Germany. E-mail: giorgio.caserta@tu-berlin.de; ingo.zebger@tu-berlin.de
bFachbereich Physik, Freie Universität Berlin, D-14195 Berlin, Germany
First published on 9th April 2026
Abstract
[NiFe]-hydrogenases catalyze the reversible cleavage of molecular hydrogen with exceptional efficiency under mild conditions and, therefore, serve as powerful blueprints for the development of sustainable, bioinspired H2-evolving catalysts. While the structure of the NiFe(CN)2CO active site has been extensively characterized, how outer-sphere residues regulate catalytic dynamics and proton-coupled electron transfer remains poorly understood. Here, we examine the functional role of a strictly conserved glutamate in the second coordination sphere of the regulatory [NiFe]-hydrogenase from Cupriavidus necator. Substitution of this glutamate with glutamine results in a dramatic loss (>99%) of catalytic activity. However, comprehensive IR, EPR, and resonance Raman spectroscopic analyses reveal that the residue is not required for the formation or stabilization of the key catalytic intermediates along the Nia–S → Nia–SR → Nia–C → Nia–L1 sequence. Notably, low-temperature IR spectroscopy shows that the transition from Nia–L1 to Nia–L2 is selectively disrupted in the absence of the conserved glutamate. These results identify the Nia–L2 state as a bona fide catalytic intermediate and demonstrate that the glutamate residue initiates critical outer-sphere rearrangements required to advance the catalytic cycle and enable productive proton transfer. Together, these findings elucidate how the protein matrix actively controls active-site reactivity in [NiFe]-hydrogenases and highlight the importance of second-sphere interactions in tuning catalytic efficiency. This work provides mechanistic principles that are directly relevant to the rational design of synthetic and biomimetic hydrogen-evolving catalysts for sustainable energy conversion.
Broader contextHydrogen is considered as a key energy carrier for a low-carbon future, but its large-scale production and use require efficient, sustainable, and affordable catalysts. In nature, hydrogenases catalyze the reversible hydrogen conversion with exceptionally high rates and minimal overpotential using only earth-abundant metals, making them powerful sources of inspiration for clean energy technologies. However, translating these biological principles into usable catalysts has been hindered by an incomplete understanding of how enzymes control chemical reactivity beyond the metal active site. This work addresses a key challenge in enzyme-mediated catalysis: how does the surrounding protein matrix enable efficient hydrogen conversion. By studying a hydrogen-processing model enzyme, we demonstrate how a conserved amino acid near the active site nickel ion, which was already known to be essential for activity, controls catalysis by directing subtle interactions between the second coordination sphere and the metal active site. These findings underscore the importance of the protein environment in modulating the reactivity of transition-metal-based biocatalysts and offer mechanistic guidance for the rational design of synthetic and biomimetic catalysts. More broadly, the principles identified here may extend to other energy-converting enzymes that operate through proton-coupled electron transfer, thereby providing guidance for the development of next-generation (bio)catalysts for the sustainable production of fuels and chemicals. |
1. Introduction
Hydrogenases use earth-abundant transition metals for fast, efficient, and reversible H2 cleavage, with almost no overpotential.1 In the subgroup of [NiFe]-hydrogenases, catalysis takes place at a heterobimetallic [NiFe] cofactor located deep within the large subunit of the heterodimeric functional unit of the enzyme.2 The bimetallic center is coordinated by four highly conserved cysteine residues—two bind exclusively to the nickel ion and two act as bridging ligands between the nickel and iron ions.3,4 The Fe is additionally coordinated by one carbon monoxide (CO) and two cyanide (CN−) strong-field ligands (Fig. 1). An open coordination site is typically present between the two metals, which may be either empty or occupied by e.g., hydroxide or hydride ligands.5,6 The electrons released from the splitting of H2 are conducted to the physiological redox partners via an array of closely spaced [Fe–S] clusters located in the hydrogenase small subunit – in some cases additional prosthetic groups such as FAD (flavin adenine dinucleotide) or FMN (flavin mononucleotide) are involved in electron transfer.7,8 [NiFe]-hydrogenases exhibit remarkable catalytic diversity (in terms of rates and bias), which is essentially determined by the protein scaffold, which, in addition to the electron relay, also provides gas channels for H2 to access/leave the active site as well as a dedicated pathway consisting of amino acids and water molecules to efficiently transport protons.9–11 Decades of research by various groups contributed to a consensus mechanism of reversible H2 cleavage (Fig. 1).12–14 In brief, the catalytic cycle is presumably initiated by the binding of H2 to the Nia–S intermediate (Ni2+–[![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ]–Fe2+),15–17 which is followed by the cleavage of the H2 molecule, yielding the bridging hydride species of the Nia–SR (Ni2+–H–Fe2+) intermediate.18 Subsequent removal of one electron and one proton results in the formation of the Nia–C state (Ni3+–H–Fe2+).19–21 Nia–C then tautomerizes to the Nia–L form through proton translocation from the bridging hydride to one of the terminal cysteine residues, accompanied by reduction of the nickel center to the formal Ni1+ state.17,22,23 Finally, the removal of the cysteine-bound proton and an electron from the nickel ion regenerates the original Nia–S intermediate, completing the catalytic cycle.24,25
]–Fe2+),15–17 which is followed by the cleavage of the H2 molecule, yielding the bridging hydride species of the Nia–SR (Ni2+–H–Fe2+) intermediate.18 Subsequent removal of one electron and one proton results in the formation of the Nia–C state (Ni3+–H–Fe2+).19–21 Nia–C then tautomerizes to the Nia–L form through proton translocation from the bridging hydride to one of the terminal cysteine residues, accompanied by reduction of the nickel center to the formal Ni1+ state.17,22,23 Finally, the removal of the cysteine-bound proton and an electron from the nickel ion regenerates the original Nia–S intermediate, completing the catalytic cycle.24,25
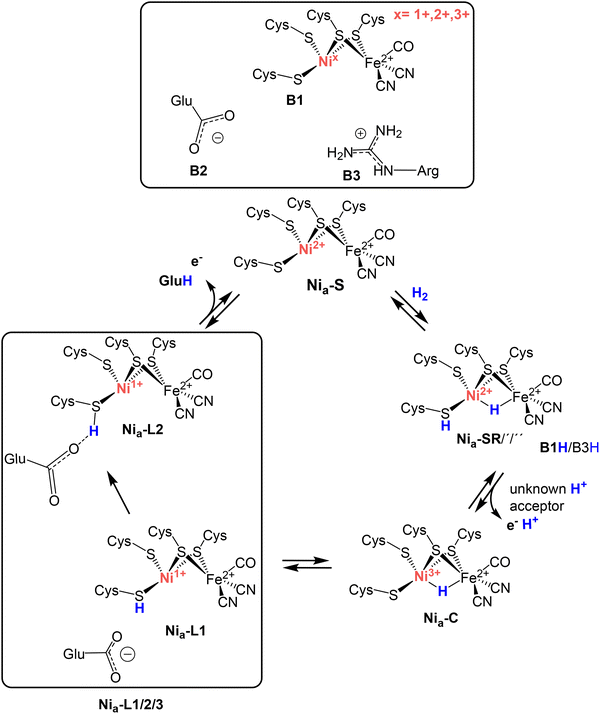 | ||
| Fig. 1 Proposed catalytic cycle of [NiFe]-hydrogenase. Schematic representation of the active site of [NiFe]-hydrogenases (top) and the proposed catalytic cycle comprising the four key intermediates Nia–S, Nia–SR, Nia–C, and Nia–L. Throughout the cycle, only nickel changes its oxidation state (1+, 2+, 3+), while iron remains in the low-spin Fe2+ state. The Nia–SR and Nia–L states each consist of at least three sub-forms. The proposed proton acceptors in the vicinity of the [NiFe] site are a Ni-bound terminal cysteine (Cys479 in RH, B1), a nearby glutamate (Glu13 in RH, B2), and an arginine (Arg411 in RH, B3), which are displayed in the top panel in their protonated/deprotonated forms according to the physiological conditions. A recent study has elucidated the structural details of the Nia–L1 and Nia–L2 species, both of which feature a proton covalently bound to a terminal nickel-coordinating cysteine residue (Cys479 in, RH).29 Nia–L2 is further stabilized by a hydrogen bond between the protonated cysteine (Cys479 in RH) thiolate and a nearby deprotonated glutamate (Glu13 in RH). The catalytic relevance of the Nia–L1 and Nia–L2 intermediates is discussed in detail in this manuscript. | ||
Among the various intermediates, both Nia–SR and Nia–L exist as multiple isoelectronic subforms (Fig. 1).12,13 Although some of these species are thought to be involved in H2 activation, their exact structure and individual roles in catalysis remain experimentally unclear. The O2-tolerant regulatory [NiFe]-hydrogenase from Cupriavidus necator (CnRH) has already been employed in several studies as a model system for probing the electronic and molecular structures of various catalytic intermediates.20,21,26–28
Using IR difference spectroscopy at low temperature, we have recently identified key structural features of two distinct Nia–L subforms.29 The results indicated that Nia–C converts to a metastable Nia–L1 intermediate (observable at T < 130 K) prior to the formation of Nia–L2, with both subforms containing a proton covalently bound to the terminal cysteine coordinated to Ni (Cys479 in RH, Fig. 1). IR difference spectroscopy also revealed structural rearrangements in the vicinity of the active site, including a hydrogen bond between Glu13 and the protonated Cys479 in Nia–L2, supporting a role for Glu13 in proton transfer (PT). Although prior studies have underscored the importance of this conserved glutamate,30–32 the conformational/structural dynamics of the residues surrounding the [NiFe] center remain poorly understood. To assess whether the recently reported outer-sphere spectral features of RH are intrinsically linked to the Nia–C → Nia–L1 → Nia–L2 progression within the catalytic cycle, we substituted Glu13 with glutamine, a structurally conservative yet non-protonatable analogue. Using a combination of low-temperature IR, resonance Raman, and electron paramagnetic resonance (EPR) spectroscopy, we demonstrate that Glu13—strictly conserved across [NiFe]-hydrogenases—is essential for enabling the Nia–L1 → Nia–L2 transition. Beyond confirming the involvement of this conserved glutamate in the proton-transfer network, consistent with observations in other hydrogenase classes, our results reveal how second-sphere residues actively modulate active-site reactivity by driving critical outer-sphere protein rearrangements that promote catalytic turnover.
2. Experimental section
2.1. Construction of the recombinant C. necator strain overproducing RHE13Q
Strain C. necator HF574(pGE567) was used for overproduction and purification of “native” RH, which is a variant of the original RH consisting of two copies of the HoxB and HoxC subunits (HoxB2C2). Truncation of a C-terminal peptide of HoxB and insertion of a Strep-Tag II peptide in its place prevents dimerization and interaction with cognate histidine protein kinase.33 The Strep-Tag II allows easy purification of the catalytically active HoxBC protein (hereinafter referred to as native RH). The E13Q amino acid exchange in HoxC was created by site-directed mutagenesis and Gibson assembly. The mutation was introduced into plasmid pCH1124 (SI, Table S1), carrying the synthetic operon PSH-hoxBstop-Strep-Tag II-hoxC. The plasmid was amplified as two overlapping fragments using the primers 2_#60_fw and 4_#59_rev, carrying the alternative codon (Table S1), together with the primers SFP95 and SFP96. After the Gibson assembly reaction, the resulting plasmid was checked by sequencing, digested with HindIII and SpeI and the hydrogenase-encoding fragment was ligated into the HindIII/SpeI-cut broad-host-range vector pEDY309.34 The resulting plasmid pJS98 was transferred by conjugation from E. coli S17-135 to C. necator strain HF574,36 yielding HF574(pJS98).2.2. Strain cultivation
Recombinant C. necator strains carrying plasmids for overproduction of native RH and its E13Q variant (RHE13Q) were cultivated in a basic mineral medium containing fructose and glycerol as the carbon and energy sources.37 When the bacterial cultures reached an optical density at 436 nm of 11–13, the cells were harvested by centrifugation (11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 × g, 4 °C, 15 min), and the cell pellet was flash frozen in liquid nitrogen and stored at −80 °C until further use.
500 × g, 4 °C, 15 min), and the cell pellet was flash frozen in liquid nitrogen and stored at −80 °C until further use.
2.3. Proteins purification
Native RH and RHE13Q were purified according to the following procedure. Cell pellets of the recombinant strains were resuspended in lysis buffer (5 mL of buffer consisting of 50 mM Tris-HCl, pH 8.0 (at 4 °C), 150 mM NaCl, protease inhibitor cocktail (complete EDTA-free, Roche) and DNase I (Roche) per gram of wet cell paste). The cells were subsequently disrupted at 125 MPa in a French pressure cell (G. Heinemann Ultraschall and Labortechnik, Schwäbisch Gmünd, Germany). Crude extracts were ultracentrifuged for 40 min at 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 × g and 4 °C, and the resulting soluble extract was loaded onto a Strep-Tactin® high-capacity column (IBA, Göttingen, Germany). The column was washed with ten bed volumes of washing buffer (50 mM Tris-HCl, pH 8.0 (at 4 °C), 150 mM NaCl), and the proteins were eluted with 4 bed volumes of washing buffer containing 3 mM D-desthiobiotin. The elution fraction was concentrated by ultrafiltration (4000 × g, 4 °C) using Amicon Ultracel concentrators (Millipore) with a 30 kDa cut-off. The resulting concentrate was diluted 20-fold with washing buffer and again re-concentrated by ultrafiltration. The final protein concentrate was flash-frozen and stored in liquid nitrogen until further use. The protein concentration was determined using a Pierce BCA Protein Assay kit (Thermo Scientific) using bovine serum albumin (BSA) as standard. The purity of the RHE13Q variant was assessed by SDS-PAGE (Fig. S1 in SI).
000 × g and 4 °C, and the resulting soluble extract was loaded onto a Strep-Tactin® high-capacity column (IBA, Göttingen, Germany). The column was washed with ten bed volumes of washing buffer (50 mM Tris-HCl, pH 8.0 (at 4 °C), 150 mM NaCl), and the proteins were eluted with 4 bed volumes of washing buffer containing 3 mM D-desthiobiotin. The elution fraction was concentrated by ultrafiltration (4000 × g, 4 °C) using Amicon Ultracel concentrators (Millipore) with a 30 kDa cut-off. The resulting concentrate was diluted 20-fold with washing buffer and again re-concentrated by ultrafiltration. The final protein concentrate was flash-frozen and stored in liquid nitrogen until further use. The protein concentration was determined using a Pierce BCA Protein Assay kit (Thermo Scientific) using bovine serum albumin (BSA) as standard. The purity of the RHE13Q variant was assessed by SDS-PAGE (Fig. S1 in SI).
2.4. Hydrogenase activity assay
H2-mediated reduction of methylene blue (MB, ε(MB)570nm = 13.1 mM−1 cm−1) by RH was investigated spectrophotometrically using a UV-vis spectrophotometer (Cary 50, Varian, Agilent, Santa Clara, California) as previously described.16 The 2.0-mL cuvette contained a buffer mixture of 50 mM K2HPO4, 100 mM citric acid (pH 7.0) and 0.2 mM MB. The solution was saturated with H2 gas and then enzyme samples were injected using a gas-tight Hamilton syringe. Measurements were performed with three biological replicates. Activities are reported as U mg−1 where 1 U corresponds to 1 µmol of H2 oxidized per minute.2.5. IR spectroscopy
For IR measurements at 298 and 90 K, the RH samples were prepared in 50 mM Tris-HCl (pH 8.0 at 277 K) buffer containing 150 mM NaCl and 25% glycerol and concentrated to ca. 1.2 mM. The glycerol ensures the formation of a transparent glass in the frozen state. The sample was subsequently reduced by exposure to humidified 100% H2 or D2 gas in an anaerobic chamber operating with forming gas (95% N2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5% H2). The samples were transferred into a gas-tight microcuvette for cryogenic measurements consisting of two CaF2 windows with an optical path length of 4 µm. The cell was then transferred into a homemade cryostat cooled with liquid-nitrogen, which was mounted in the sample chamber of a Tensor 27 FTIR spectrometer (Bruker), equipped with a liquid-nitrogen cooled mercury cadmium telluride (MCT) detector. The cell compartment was purged with dried air. Data acquisition, spectral analysis, and Gaussian fitting of the CO/CN bands were performed using Bruker OPUS version 7.8. Spectra with a resolution of 2 cm−1 were recorded by averaging 200 scans. Absorbance spectra were calculated from averaged single channel spectra of the sample using the correspoding buffer spectrum as reference. Light-minus-dark IR difference spectra were calculated accordingly using the corresponding dark single spectra as reference.29 The Nia–C → Nia–L transformation can be induced over a broad temperature range, from 90 K (the lower limit of our liquid N2 cryostat) up to approximately 160–180 K. Above this range, thermal back-conversion to the initial Nia–C state(s) usually takes place. Notably, the Nia–L1 → Nia–L2 transition is extremely slow below 130 K, while at temperatures between 160 and 180 K it proceeds at rates comparable to the light-induced Nia–C → Nia–L1 reaction. This results in an apparent direct conversion of Nia–C to Nia–L2, when illuminating at these temperatures. Therefore, careful adjustment of the experimentally accessible temperature window is essential to selectively resolve the individual active site species.
5% H2). The samples were transferred into a gas-tight microcuvette for cryogenic measurements consisting of two CaF2 windows with an optical path length of 4 µm. The cell was then transferred into a homemade cryostat cooled with liquid-nitrogen, which was mounted in the sample chamber of a Tensor 27 FTIR spectrometer (Bruker), equipped with a liquid-nitrogen cooled mercury cadmium telluride (MCT) detector. The cell compartment was purged with dried air. Data acquisition, spectral analysis, and Gaussian fitting of the CO/CN bands were performed using Bruker OPUS version 7.8. Spectra with a resolution of 2 cm−1 were recorded by averaging 200 scans. Absorbance spectra were calculated from averaged single channel spectra of the sample using the correspoding buffer spectrum as reference. Light-minus-dark IR difference spectra were calculated accordingly using the corresponding dark single spectra as reference.29 The Nia–C → Nia–L transformation can be induced over a broad temperature range, from 90 K (the lower limit of our liquid N2 cryostat) up to approximately 160–180 K. Above this range, thermal back-conversion to the initial Nia–C state(s) usually takes place. Notably, the Nia–L1 → Nia–L2 transition is extremely slow below 130 K, while at temperatures between 160 and 180 K it proceeds at rates comparable to the light-induced Nia–C → Nia–L1 reaction. This results in an apparent direct conversion of Nia–C to Nia–L2, when illuminating at these temperatures. Therefore, careful adjustment of the experimentally accessible temperature window is essential to selectively resolve the individual active site species.
2.6. EPR spectroscopy
RHE13Q samples with a concentration range of 0.2–0.3 mM and a volume of 100 µL were transferred into quartz EPR tubes (4 mm diameter), frozen in cold ethanol (193 K), and stored in liquid nitrogen for further analysis. EPR samples were illuminated during the experiments using the focused light of a collimated 455 nm LED. A Bruker EMXplus spectrometer combined with an ER 4122SHQE resonator, an Oxford EPR 900 helium flow cryostat, and an Oxford ITC4 temperature controller was used for the EPR experiments. Baseline correction of the experimental spectra was done by subtracting a spectrum of buffer solution measured with the same experimental parameters. Broad background fluctuations were additionally corrected by using a polynomial or spline function. If not otherwise noted, the following experimental parameters were used: 1 mW microwave power, 9.29 GHz microwave frequency, 10 G modulation amplitude, and 100 kHz modulation frequency.2.7. Resonance Raman spectroscopy
For resonance Raman (RR) spectroscopic investigations, the 568 line of a Kr+ gas laser (Coherent) was used. Spectra were recorded at 80 K, using a liquid-nitrogen cooled cryostat (Linkam Scientific Instruments), in back-scattering geometry utilizing a confocal setup (Horiba Scientific LabRam). The laser beam was focused by a Nikon 20× objective to the surface of the sample resulting in a spot size of <10 µm. Data acquisition was accomplished with a Peltier-cooled CCD array (213 K, Oxford Instruments Andor). The laser power at the sample was adjusted to 1.5 mW using a neutral density glass filter (Schott AG). All samples were directly transferred from liquid nitrogen to the sample chamber (Linkam THMS600 freezing microscope stage) with minimal exposure to air. Individual spectra were recorded for 180 s, and repetitive scans were accumulated to accomplish overall acquisition times of 1.5–3 h per measurement spot, depending on the quality of the corresponding spectra. To calibrate the probe frequency axis, spectra of toluene and acetonitrile (external standard, marker bands at 522 cm−1 and 2254 cm−1) were recorded before or after individual sample measurements. For each solution sample, at least two spots on different drops were measured and evaluated separately. Data evaluation of the RR spectra was done using the Bruker OPUS software version 6.5 or higher.3. Results and discussion
3.1. Biochemical and basic spectroscopic analysis of RHE13Q
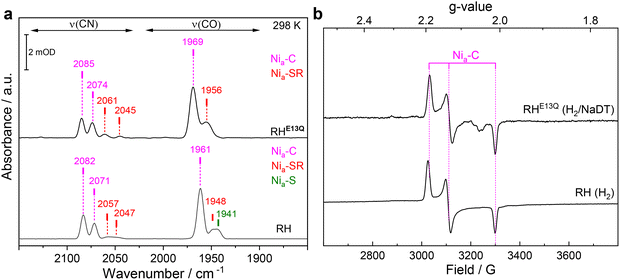
The glutamate-to-glutamine exchange variant of the regulatory [NiFe]-hydrogenase from C. necator (RHE13Q) was generated by site-directed mutagenesis (Table S1, methods section) and purified as described previously (Fig. S1).37 The purified protein showed an H2 oxidation activity of 0.004 ± 0.001 µmol H2 mg−1 min−1, which corresponds to ca. 0.1% of the value of native RH of C. necator. The dramatic effect of the Glu-to-Gln exchange on catalysis is consistent with previous studies on the prototypical [NiFe]-hydrogenase from Solidesulfovibrio fructosivorans (SfH2ase),30 the soluble NAD+-reducing hydrogenase from Hydrogenophilus thermoluteolus (HtSH),38 and the membrane-bound hydrogenases from E. coli Hyd-1 (EcHyd-1) and Hyd-2 (EcHyd-2).31,39 Although this amino acid substitution renders most hydrogenases almost inactive, a few, such as the NADP+-reducing soluble hydrogenase I from Pyrococcus furiosus (PfSH) and the MBH-type Hyn from Thiocapsa roseopersicina BBS,40,41 retain considerable activity. This suggests the presence of alternative H+ transfer pathways and/or rescue mechanisms.To assess whether replacing Glu with Gln in RH influences the incorporation of the active site metals and/or their redox behavior, we first characterized the as-isolated, oxidized C. necator RHE13Q variant using room-temperature IR and native RH as the reference (Fig. 2). The IR spectrum of the as-isolated RHE13Q is dominated by a broad νCO band centered at 1952 cm−1, along with νCN bands at 2075 and 2084 cm−1 (Fig. 2a, top), and a minor species characterized by a νCO band at 1969 cm−1. Native RH, on the other hand, essentially shows only one νCO band at 1943 cm−1 with two νCN bands at 2071 and 2080 cm−1 (Fig. 2a, bottom), corresponding to the Nia–S intermediate of the active site. To facilitate the assignment of the two active site species in as-isolated RHE13Q, we performed complementary EPR and RR spectroscopy. EPR spectroscopic analysis of RHE13Q (Fig. 2b) revealed signals of a paramagnetic state similar to the Nir–B species (Ni3+–OH–Fe2+) of RH, whose g values (gx = 2.20, gy = 2.15, gz = 2.02) resemble those observed for the Nir–B′ species in the oxidized large subunit HoxC of RH and HoxG of MBH.3,42 These observations led to the assignment of the 1969 cm−1 band to the CO stretching vibration of a Nir–B-like state in as-isolated RHE13Q. Based on the weak Nir–B signal, we deduce that most of the active site states of the RHE13Q sample are diamagnetic. This became supported by RR spectroscopy, selectively probing metal–ligand vibrations, such as Fe–CO and Fe–CN stretching and bending modes of the active site, which typically occur in the spectral range of 400–650 cm−1. By comparing the RR data (recorded with an 568 nm excitation line) of as-isolated native RH enriched in the Nia–S intermediate17,23,43 with those from as-isolated RHE13Q, we found that the RR spectroscopic signatures of the two proteins are very similar (Fig. 2c).
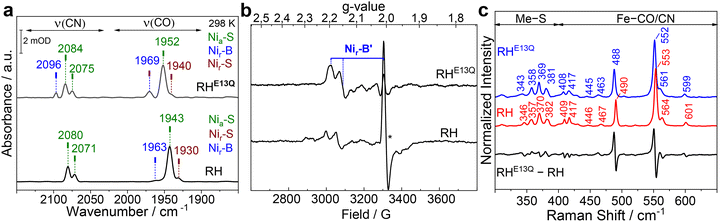 | ||
| Fig. 2 Spectroscopic characterization of as-isolated RHE13Q. (a) IR spectra of the RHE13Q variant (top) and native RH (bottom) recorded at 298 K in the spectral range where the CO/CN-stretching vibrations of the diatomic ligands occur. The νCO and νCN bands are labeled with their corresponding wavenumbers. The IR spectra are dominated by signals attributed to the Nia–S (dark green). Minor contributions of the Nir–B-like (blue) and Nir–S (brown) species were also detected in RHE13Q. (b) EPR spectra of the as-isolated RHE13Q variant and native RH recorded at 80 K. The asterisk marks the weak signal of a [3Fe–4S] cluster most likely resulting from a degraded [4Fe–4S] cluster.16 (c) Solution-phase RR spectra (80 K, excited at 568 nm) of RHE13Q (blue trace) and native RH (red trace). Spectra are normalized with respect to the integral of the Fe–CO/CN marker band centered at 552/553 cm−1. To highlight spectra changes, a difference spectrum RHE13Q–native RH is shown as well (black trace). Spectral regimes dominated by Ni-centered metal–sulfur (Me–S) modes and Fe-centered Fe–CO/CN stretching and bending modes are indicated. Due to the high excitation wavelength, the data is free of [Fe–S] clusters contributions. The data of native RH are reproduced from ref. 44 © 2024 The Authors. Published by Elsevier Inc. | ||
Additionally, we also resolved high-frequency intra-ligand modes of the diatomic CO/CN− ligands at the RH and RHE13Q [NiFe] active sites that allows an unambiguous assignment of the states detected by RR spectroscopy (Fig. S2).44 This in turn suggests that a large portion of the active sites in RHE13Q resides in the Nia–S state, which is diamagnetic and therefore consistent with the EPR analysis. Interestingly, the νCO IR band at 1952 cm−1, which presumably represents the Nia–S intermediate of RHE13Q, is shifted to higher frequencies (ca. 9 cm−1, Fig. 2a), while the corresponding RR Fe–CO/CN absorptions are slightly shifted to lower energies (Fig. 2c) compared to the corresponding bands of native RH. A similar observation has been recently made in an nuclear resonance vibrational spectroscopy (NRVS)/IR spectroscopic study of the RH large subunit HoxC.6 This means that the Fe–CO/CN in the Nia–S species of RHE13Q exhibit weaker metal–ligand bonding (red-shifted RR Fe–CO/CN bands, Fig. 2c), which strengthen the diatomic CO/CN bonds as evidenced by the blue-shifted CO/CN stretching vibrations in IR, Fig. 2a). The increased energies of the CO and CN vibrations of the Nia–S state, which are clearly observable across all observed redox states (see below), probably result from the particular electrostatic interactions with the surrounding atoms in the catalytic center of RHE13Q.31 In particular, while the glutamine side chain is electrically neutral under physiological conditions, our recent data indicate that Glu13 is deprotonated in several active-site intermediates of the native RH enzyme and thus, carries a negative charge.29 The observed shift to higher energies for the CO and CN stretching frequencies in RHE13Q may therefore result from the absence of this negative charge, which likely reduces the electron density around the metal center.
After H2 incubation in the presence of sodium dithionite (NaDT) to prevent rapid reoxidation (Fig. S3), the IR spectrum of the reduced RHE13Q variant exhibits two major CO bands at 1969 and 1956 cm−1 (Fig. 3a, top), which are shifted to higher energies relative to those of the predominant species observed in reduced RH (Fig. 3a, bottom). The corresponding CN absorption frequencies are shown in Fig. 3a and listed in Table S2. Complementary EPR data on reduced RHE13Q revealed, regardless of the specific reduction protocol used (Fig. S4a), the typical spectrum of the Nia–C intermediate observed in native RH (gx = 2.19, gy = 2.14, gz = 2.01, Fig. 3b), which enabled the assignment of the band at 1969 cm−1 to Nia–C (Ni3+–H−–Fe2+). Finally, the broad CO band at 1956 cm−1 likely contains contributions from the diamagnetic bridging-hydride Nia–SR state (Ni2+–H−–Fe2+), which typically accumulates in small amounts in reduced native RH (Fig. 3a, bottom; Fig. S3),27 as well as minor contributions from Nia–S (1952 cm−1, Fig. 2a). Significantly, the enrichment of Nia–C in RHE13Q upon reduction is similar to that in native RH, as previously observed for the corresponding variants of EcHyd1 and SfH2ase.30,31 Thus, the conserved glutamate presumably does not act as H+ acceptor during the Nia–SR → Nia–C transition (Fig. 1).
3.2. Infrared spectroscopic analysis of RHE13Q at low temperatures
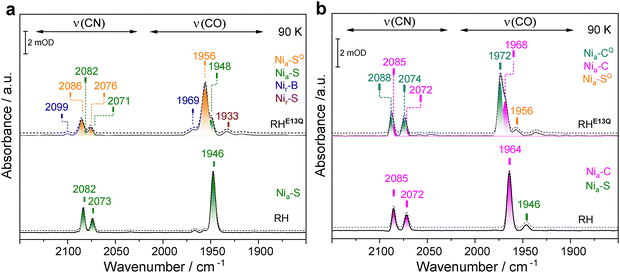
To further examine the impact of the Glu-to-Gln exchange in RH, the protein variant was analyzed in detail using low-temperature IR spectroscopy, which has been demonstrated to provide valuable details about changes in the outer coordination sphere of the [NiFe] active site.22,29,45 As-isolated and H2/NaDT-reduced samples RHE13Q and native RH were rapidly cooled to 90 K using a liquid-nitrogen bath cryostat, and IR spectra were subsequently recorded (Fig. 4). Notably, the IR absorptions of the CO and CN− ligands displayed significantly narrower line shapes at 90 K than at 298 K. This improved resolution facilitated the identification of additional overlapping active-site species that would otherwise be masked by line broadening at room temperature. For example, the broad CO band at 1952 cm−1 (298 K) observed for as-isolated RHE13Q and assigned to the Nia–S state (Fig. 2a) was resolved more clearly at 90 K, revealing two adjacent νCO bands at 1948 and 1956 cm−1. Gaussian fitting of the CO/CN bands suggests the presence of two distinct subforms of the Nia–S state (Fig. 4a). Furthermore, the νCO band at 1969 cm−1 of the reduced RHE13Q sample, assigned to the Nia–C state (Fig. 3a), was also resolved into two distinct bands at 1972 and 1968 cm−1 (Fig. 4b). Interestingly, complementary EPR measurements on the H2/NaDT-reduced protein variant reveal the presence of a single Nia–C species (Fig. 3b). This suggests that the structural differences between the two Nia–C states detected by IR spectroscopy are unlikely to be accompanied by a significant change in the local environment or coordination geometry of the Ni center, resulting in EPR spectra that are indistinguishable for the two species. Our results therefore emphasize an important point: relying on a single spectroscopic technique may not be sufficient to unambiguously assign and distinguish hydrogenase active-site species. Additionally, EPR measurements at 10 K, together with power-dependent saturation analyses at 20 K (Fig. S4b–d), are consistent with the presence of a reduced proximal [4Fe–4S] cluster in H2/NaDT-reduced RHE13Q. This behavior is in agreement with previous observations made for native RH upon incubation with chemical reductants.27 Thus, unlike the PfSH hydrogenase, where distinct Nia–C species have been attributed to differences in the oxidation state of the proximal [4Fe–4S] cluster,14 the two Nia–C species detected in RHE13Q are unlikely to originate from such redox variations. Notably, low-temperature IR spectrum of reduced RHE13Q revealed an almost complete disappearance of the Nia–SR-associated νCO band at 1956 cm−1 (Fig. 3a vs. Fig. 4b). We attribute this behavior to a temperature-dependent change in the redox equilibrium of the Nia–SR and the Nia–C states, as recently described for membrane-bound [NiFe]-hydrogenase from C. necator45 and HtSH.46The IR spectra of the RHE13Q variant differ from those of the native enzyme in two significant ways. First, the dominant IR bands recorded at room temperature appear broader, which is due to the presence of two subforms that can be disentangled at low temperatures. We assume that these subforms differ in some structural element(s) at or near the active site. In support of this interpretation, crystallographic analyses of the analogous E28Q variant of the O2-tolerant EcHyd1 showed that a subset of the active site molecules accommodate an additional H2O or OH− adjacent to the amide headgroup of the glutamine residue.31 Notably, the presence of a sub-population with an anionic species electrostatically equivalent to the deprotonated carboxylate of E13 could account for the IR observations of a minor fraction in RHE13Q characterized by CO/CN stretching signals at lower energies relative to the main fraction. In particular, the bands at 1948 and 1968 cm−1 (Fig. 4a and b) closely match those of the Nia–S and Nia–C states in oxidized and reduced samples of native RH, detected at 1946 and 1964 cm−1, respectively. This strong similarity suggests that the active site sub-states represented by CO bands at 1948 and 1968 cm−1 might reflect a sub-population that bears a hydroxide anion in proximity of the [NiFe] site. This would reinstate the electrostatic effect of the missing negatively charged carboxylate group, thereby, lowering the CO stretching frequencies. These Nia–S and Nia–C subforms account for ∼20–30% of the total absorption integral and are hereafter referred to as the prototypical states of RHE13Q. The second major difference between RHE13Q and RH is that the IR bands belonging to the dominant subforms are blue-shifted by about 6–9 cm−1 both in the as-isolated and reduced RHE13Q samples. Similarly to the IR data recorded at 298 K (Fig. 2a and 3a), the absence of a negative charge near the bimetal center (e.g., deprotonated glutamate or OH−) may strengthen the bonds of the diatomic Fe-ligands, leading to the observed shift of the CO bands to higher energies. The superscript “Q” was incorporated into designation of the more dominant subforms of RHE13Q (1956 and 1972 cm−1), and we refer to them as Nia–SQ and Nia–CQ in the caption of Fig. 4 and the text below. Table S2 summarizes all observed redox states of the active site and their corresponding CO/CN absorptions.
3.3. Photoconversion of active site states in reduced RHE13Q at cryogenic temperature
The Nia–C intermediate,47–49 and, more recently, one Nia–SR subform,46 have been shown to convert into Nia–L species under cryogenic conditions upon exposure to blue light. This approach allows these active-site species to be trapped kinetically and characterized spectroscopically. However, Nia–L species have also been observed at ambient temperature in IR spectroscopic studies on PfSH,14,40 EcHyd1,25,50,51 and isolated catalytic subunits of RH and MBH from C. necator.3,42 Photolysis of the hydride ligand leads to proton translocation to one of the cysteines (Cys479 in C. necator RH) that terminally ligate the Ni ion.22 The two electrons originating from the hydride are transferred to the Ni ion and one of the adjacent [Fe–S] clusters in case of the Nia–SR to Nia–L conversion. During the Nia–C to Nia–L conversion, however, neither protons nor electrons are released from the first coordination sphere, i.e., Nia–C and Nia–L are tautomers. As outlined in the introduction, we have recently elucidated the details of the latter photo-transformation for native RH.29 After illumination of RH in the Nia–C state, photolysis results in the formation of the Nia–L1 subform, which then (in the dark) thermally converts to the thermodynamically more stable Nia–L2 state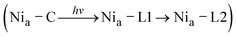 .
.
When the same experiment was performed with H2/NaDT-reduced RHE13Q, illumination with blue LED light (460 nm) at 90 K led to the conversion of the two distinct Nia–C states into two Nia–L states characterized by νCO bands at 1921 and 1915 cm−1 (Fig. 5a).
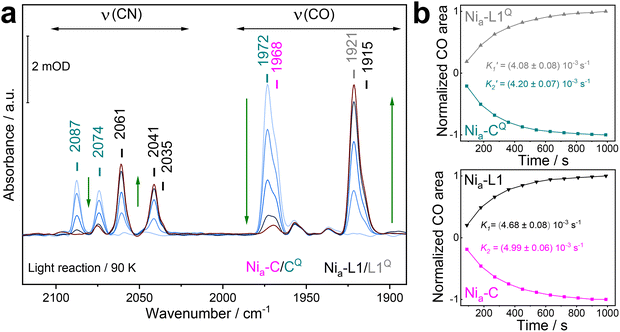 | ||
Fig. 5 IR spectra of as-isolated and H2/NaDT-reduced RHE13Q at 90 K before, during and after illumination. (a) Representative IR absorption spectra of reduced RHE13Q before, during, and after LED-illumination at 460 nm. Early and late spectra are colored light blue and brown, respectively. CO/CN bands of the Nia–C (magenta), Nia–CQ (dark cyan), Nia–L1 (black) and Nia–L1Q (grey) are labeled with corresponding wavenumbers (b) kinetic profiles of the Nia–C/CQ depletion and Nia–L1/L1Q formation at 90 K. The normalized CO areas of Nia–L1 and Nia–C (νCO bands at 1915 and 1968 cm−1) and Nia–L1Q and Nia–CQ (νCO bands at 1921 and 1972 cm−1) were plotted against time. The fitted curves exhibit monoexponential kinetics, indicating two distinct conversions, i.e., Nia–CQ (1972 cm−1) → Nia–L1Q (1921 cm−1) and Nia–C (1968 cm−1) → Nia–L (1915 cm−1). Color code as in (a). The fit parameters are summarized in Table S3. It should be noted that the kinetic rates associated with Nia–C depletion (K2) and Nia–L1 formation (K1) appear to be faster than those estimated for 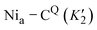 and and 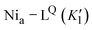 . These differences likely originate from partial back conversion of Nia–L1Q → Nia–CQ, which seems to proceed slightly faster than the corresponding Nia–L1 → Nia–C process. This interpretation is supported by the IR difference spectra shown in Fig. 6a, which indicate enrichment of the Nia–CQ species in RHE13Q after thermal transformation. In this sense, the values for the forward reaction represent apparent rate constants. . These differences likely originate from partial back conversion of Nia–L1Q → Nia–CQ, which seems to proceed slightly faster than the corresponding Nia–L1 → Nia–C process. This interpretation is supported by the IR difference spectra shown in Fig. 6a, which indicate enrichment of the Nia–CQ species in RHE13Q after thermal transformation. In this sense, the values for the forward reaction represent apparent rate constants. | ||
Their relative intensities were similar to those observed for the Nia–C/Nia–CQ subforms. Kinetic analysis of the light-induced reaction indicates that the two Nia–L subforms can be traced back (rate constants K1 ≈ K2 and 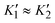 ) to the corresponding Nia–C subforms present prior to illumination (Fig. 5b and Table S3). Accordingly, we designated the more dominant subform at 1921 cm−1 as Nia–L1Q.
) to the corresponding Nia–C subforms present prior to illumination (Fig. 5b and Table S3). Accordingly, we designated the more dominant subform at 1921 cm−1 as Nia–L1Q.
After complete photolysis of the Nia–C/Nia–CQ states and enrichment of the Nia–L1/Nia–L1Q subforms, the illumination was switched off. In native RH, the thermal conversion of the Nia–L1 state to the Nia–L2 state proceeds very slowly at 90 K, and we have previously observed that slightly higher temperatures can accelerate Nia–L2 formation.29 Therefore, the temperature of the RHE13Q sample was first increased to approximately 130 K to accelerate the enrichment of Nia–L2 by thermal transformation and subsequently lowered back to 90 K prior to record the spectral features of the Nia–L2/L2Q states.
Fig. 6a shows an overlay of the IR difference spectra of “after light-minus-dark“ (black traces), where the positive bands highlight enrichment of Nia–L1 before thermal transformation, and “after thermal transformation-minus-dark” (dark yellow and red traces), where the positive bands reflect the enrichment of Nia–L2 at high temperatures. By comparing both RH samples in H2/H2O and D2/D2O, we identified clear spectral differences between native RH and RHE13Q. Both the CO and the CN bands of the Nia–L1 state in native RH (Fig. 6a, upper black and grey traces) undergo a net blue shift in the Nia–L2 state (upper dark yellow and red traces). In particular, the νCO shifts from 1911 to 1914 cm−1 and νCN move from 2037/2056 to 2040/2060 cm−1. These changes were proposed to originate from conformational changes around the [NiFe] site including a new hydrogen bond (H-B) between the deprotonated glutamate and the protonated thiolate of Cys479.29 In contrast, analysis of the CO/CN spectral region of RHE13Q revealed a pronounced similarity between the Nia–L species populated immediately after photolysis (Fig. 6a, lower black trace) and that obtained following thermal transformation (Fig. 6a, lower dark yellow trace). The main CO band exhibits only a slight red shift from 1921 to 1919 cm−1 (marked in wine red), and only the asymmetric CN stretching shows a modest upshift of approximately 2 cm−1 (marked in wine red). These changes in RHE13Q are virtually absent in D2/D2O (Fig. 6a, lower grey and red traces). Complementary EPR measurements on irradiated RHE13Q (90 K, Fig. 6b) yielded signals with g values characteristic of Nia–L1 (predominant, gx = 2.247, gy = 2.090, gz = 2.044) and Nia–L2 (minor, gx = 2.309, gy = 2.076, gz = 2.052) species, closely matching those observed for native RH, similar to the correspondence seen for the parent Nia–C states (Fig. 3b).20,29
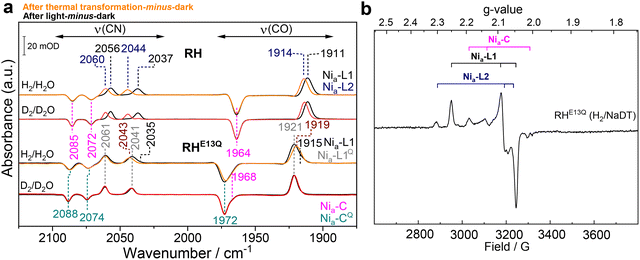 | ||
| Fig. 6 IR and EPR characterization of the Nia–L states of RHE13Q and RH. (a) IR difference spectra at 90 K in the 2150–1850 cm−1 regime for H2/NaDT-reduced native RH (upper traces) and RHE13Q (lower traces) prepared with H2/H2O and D2/D2O. The figure shows two sets of IR difference spectra: the “after light-minus-dark” spectra (black, H2/H2O; grey, D2/D2O) and the “after thermal transformation-minus-dark” spectra (orange, H2/H2O; red, D2/D2O). The spectra of RHE13Q after light exposure show prominent positive absorptions of the active site CO/CN ligands related to the Nia–L1Q at 1921, 2041, and 2061 cm−1 (marked in grey), while negative Nia–CQ-related bands were detected at 1972, 2074, and 2088 cm−1 (dark cyan, Table S2). Minorly populated Nia–L1 (black) and Nia–C (magenta) species are also visible (see also Fig. 5). The “after light-minus-dark” spectra of RH exhibit Nia–L1-related positive νCO/CN bands at 1911, 2037, and 2056 cm−1 (black) while Nia–C-related signals (magenta) were observed at 1964, 2072 and 2085 cm−1. After thermal transformation, native RH is stoichiometrically enriched in the Nia–L2 (νCO/CN bands at 1914, 2044 and 2060 cm−1), while RHE13Q still exhibits predominantly Nia–L1Q-related signals. Minor changes of the CO (1919 cm−1) absorption and the band related to the antisymmetric CN stretching vibration (2043 cm−1) in RHE13Q, which arise from overlapping Nia–L1Q and Nia–L2 CO/CN bands, are marked in wine red (see also Fig. S6). (b) EPR spectrum of the H2/NaDT-reduced RHE13Q recorded at 90 K upon LED-illumination (455 nm) resulting in the formation of the Nia–L1 (gx = 2.247, gy = 2.090, gz = 2.044) and Nia–L2 (gx = 2.309, gy = 2.076, gz = 2.052) species. Trace signals of Nia–C species are also observed. | ||
To obtain detailed information about differences in the second/outer coordination sphere between the active sites of RHE13Q and native RH, we analyzed and compared the IR difference spectra in the ranges from 1450 to 1800 cm−1 (Fig. 7a and b). The IR difference spectra “after light-minus-dark” (Nia–L1/L1Q-minus-Nia–C/CQ) revealed major positive bands at 1705, 1686 and 1622 cm−1 related to the Nia–L1/1Q states, and clear negative bands at 1700, 1675 and 1630 cm−1 deriving from the Nia–C/CQ species. These signals appear in a spectral range that is characteristic for signals from Arg (symmetric, νs, and asymmetric stretching vibrations, νas, of CN3H5+) and Gln residues (νC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) as well as amide I/II bands and water contributions.52,53 Notably, the positive and negative absorption features of the RHE13Q variant closely match those of native RH (Fig. 7a, black traces), suggesting that Glu13 is “dispensable” for the Nia–C → Nia–L1 transition in native RH. In contrast, the analogous glutamate in DvMF hydrogenase has been proposed to undergo significant structural changes during the Nia–C → Nia–L transition (possibly Nia–L2), forming one hydrogen bond (with a cysteine terminally coordinating the Ni) in the Nia–C intermediate and two bonds in the Nia–L(2) state.22 These contrasting results imply that the mechanism of H2 activation in RH and DvMF hydrogenases proceeds with distinct dynamics of outer-sphere residues.
O) as well as amide I/II bands and water contributions.52,53 Notably, the positive and negative absorption features of the RHE13Q variant closely match those of native RH (Fig. 7a, black traces), suggesting that Glu13 is “dispensable” for the Nia–C → Nia–L1 transition in native RH. In contrast, the analogous glutamate in DvMF hydrogenase has been proposed to undergo significant structural changes during the Nia–C → Nia–L transition (possibly Nia–L2), forming one hydrogen bond (with a cysteine terminally coordinating the Ni) in the Nia–C intermediate and two bonds in the Nia–L(2) state.22 These contrasting results imply that the mechanism of H2 activation in RH and DvMF hydrogenases proceeds with distinct dynamics of outer-sphere residues.
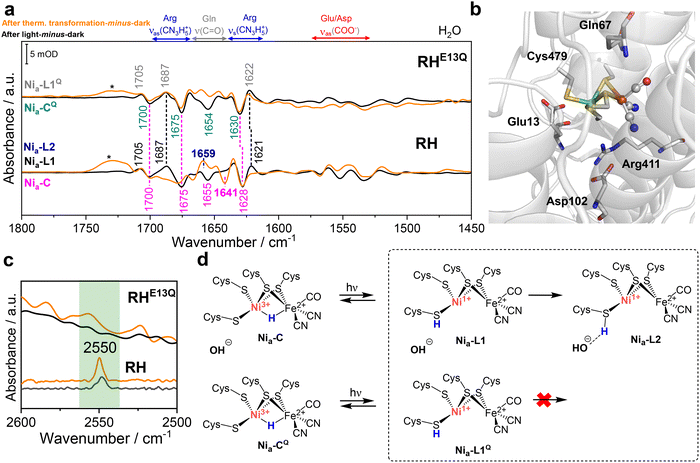 | ||
Fig. 7 IR difference spectra of RHE13Q and native RH revealing important elements of the first and outer coordination sphere of the corresponding [NiFe] sites. (a) “After light-minus-dark” (black traces) and “after thermal transformation-minus-dark” (orange traces) difference spectra of native RH (bottom) and RHE13Q (top) prepared with H2/H2O. The spectra in the range between 1800 and 1450 cm−1 show potential contributions from individual amino acid residues. The spectral regimes characteristic for the main bands of arginine (CN3H5+, νs and νas, blue arrow), glutamine (νC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, grey arrow) and deprotonated aspartate/glutamate (νas COO−, red arrow), are highlighted. Amide I and amide II absorptions, which occur between 1600–1700 cm−1 and 1510–1580 cm−1, respectively, as well as water absorptions (δ(OH) bending, 1635–1670 cm−1) might also contribute to the observed absorptions. We assign the broad band around 1700–1730 cm−1 (marked with *) to an artifact due to slight temperature fluctuations over time. (b) AlphaFill-predicted model of the RH large subunit HoxC including the NiFe(CN)2CO cofactor and selected outer-sphere residues.29 The coordinating cysteines (Cys60, 63, 479, and 482), together with Glu13, Gln67, Arg411, and Asp102, residues are shown in stick representation. Color code: C, grey; O, red; N, blue; S, yellow; Ni, green; Fe, brown. The protein backbone is shown in cartoon representation (grey). (c) Spectral region typical for S–H stretching vibrations for native RH and RHE13Q after light reaction and after thermal transformation. (d) Schematic representation of the [NiFe] active site in RHE13Q variant depicting the likely involved structural elements characterizing Nia–C/CQ, Nia–L1/L1Q and Nia–L2 active site states. For the minorly populated Nia–C, Nia–L1 and Nia–L2 in RHE13Q we propose the presence of an OH− anion in proximity of the [NiFe] site, which can partially compensate the absence of the negatively charged carboxylate side group of the conserved glutamate (see Fig. 1). O, grey arrow) and deprotonated aspartate/glutamate (νas COO−, red arrow), are highlighted. Amide I and amide II absorptions, which occur between 1600–1700 cm−1 and 1510–1580 cm−1, respectively, as well as water absorptions (δ(OH) bending, 1635–1670 cm−1) might also contribute to the observed absorptions. We assign the broad band around 1700–1730 cm−1 (marked with *) to an artifact due to slight temperature fluctuations over time. (b) AlphaFill-predicted model of the RH large subunit HoxC including the NiFe(CN)2CO cofactor and selected outer-sphere residues.29 The coordinating cysteines (Cys60, 63, 479, and 482), together with Glu13, Gln67, Arg411, and Asp102, residues are shown in stick representation. Color code: C, grey; O, red; N, blue; S, yellow; Ni, green; Fe, brown. The protein backbone is shown in cartoon representation (grey). (c) Spectral region typical for S–H stretching vibrations for native RH and RHE13Q after light reaction and after thermal transformation. (d) Schematic representation of the [NiFe] active site in RHE13Q variant depicting the likely involved structural elements characterizing Nia–C/CQ, Nia–L1/L1Q and Nia–L2 active site states. For the minorly populated Nia–C, Nia–L1 and Nia–L2 in RHE13Q we propose the presence of an OH− anion in proximity of the [NiFe] site, which can partially compensate the absence of the negatively charged carboxylate side group of the conserved glutamate (see Fig. 1). | ||
Importantly, the “after thermal transformation-minus-dark” difference spectrum of the RHE13Q variant displays features that are nearly identical to those of the “after light-minus-dark” spectrum (Fig. 7a, top). These observations suggest that the outer coordination sphere of the active site of RHE13Q does not undergo the conformational changes observed in native RH (see, for example, the new set of negative/positive bands at 1641 and 1659 cm−1 in Fig. 7a). Importantly, only weak absorptions are observed in the spectral region associated with deprotonated aspartate/glutamate residues between 1530 and 1570 cm−1. This is particularly evident in experiments conducted in D2/D2O (Fig. S5), where RHE13Q is characterized by nearly stoichiometric Nia–CQ and Nia–L1Q species, consistent with the absence of the conserved glutamate and possibly reflecting an impaired proton-transfer (PT) pathway involving deprotonated carboxylates interacting with E13.29 Based on our previous findings for native RH, where the deprotonated glutamate was shown to form a hydrogen bond with the protonated Ni-bound Cys479 in the Nia–L2 state, we propose that the thermal conversion of Nia–L1/L1Q in RHE13Q (H2/H2O) yields only a minor population of Nia–L2 species. In this variant, the role of the deprotonated glutamate would be mimicked by an electrostatically equivalent hydroxide ligand capable of hydrogen bonding to protonated Cys479 (Fig. 7d), while the predominant Nia–L1Q state remains essentially unchanged.
Guided by these observations, we performed Gaussian fitting of the CO and CN bands of H2/NaDT reduced RHE13Q in H2/H2O (Fig. S6), assuming enrichment of the Nia–L2 state while retaining the Nia–L1Q population. The resulting fit accurately reproduces the experimental CO/CN spectral features of RHE13Q. Accordingly, the bands corresponding to the predominant Nia–L1Q species remain largely unaffected after thermal transformation, and only a small fraction of molecules—represented by the Nia–L1 νCO band at 1915 cm−1—appears to convert into the Nia–L2 state, characterized by a νCO band near 1917 cm−1, partially overlapping with the νCO of the Nia–L1Q at 1921 cm−1. These results suggest that the [NiFe] site in RHE13Q predominantly remains in the outer-sphere Nia–L1Q conformation (Fig. 7a). The only exception is the broad negative band at 1654 cm−1 (associated with Nia–C), which falls within the amide νC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O range and may originate in part from the introduced glutamine residue. Accordingly, this band shows a significantly increased intensity compared to native RH (Fig. 7a, bottom) and exhibits a redshift when the sample is prepared in D2/D2O (Fig. S5, 1651 cm−1), consistent with previously reported solvent isotope effects.53 Furthermore, the observed νC
O range and may originate in part from the introduced glutamine residue. Accordingly, this band shows a significantly increased intensity compared to native RH (Fig. 7a, bottom) and exhibits a redshift when the sample is prepared in D2/D2O (Fig. S5, 1651 cm−1), consistent with previously reported solvent isotope effects.53 Furthermore, the observed νC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O feature vanishes during thermal transformation (Fig. 7a, middle trace), suggesting that the corresponding residue—most likely the newly introduced Gln13—is stabilized after thermal transformation in a conformation resembling that of the Nia–C/CQ states, while all other nearby vibrational features remain unchanged. Finally, whereas native RH displays clear νSH bands for both the Nia–L1 and Nia–L2 states (νSH ≈ 2550 cm−1; Fig. 7c), with the intensity being significantly higher for Nia–L2, where the protonated Cys479 is proposed to form an hydrogen bond with Glu13 (Fig. 1),29 the corresponding IR difference spectra of the RHE13Q variant exhibit no detectable νSH absorptions (Fig. 7c).
O feature vanishes during thermal transformation (Fig. 7a, middle trace), suggesting that the corresponding residue—most likely the newly introduced Gln13—is stabilized after thermal transformation in a conformation resembling that of the Nia–C/CQ states, while all other nearby vibrational features remain unchanged. Finally, whereas native RH displays clear νSH bands for both the Nia–L1 and Nia–L2 states (νSH ≈ 2550 cm−1; Fig. 7c), with the intensity being significantly higher for Nia–L2, where the protonated Cys479 is proposed to form an hydrogen bond with Glu13 (Fig. 1),29 the corresponding IR difference spectra of the RHE13Q variant exhibit no detectable νSH absorptions (Fig. 7c).
However, these observations do not exclude protonation of Cys479 in this variant. We propose that the intrinsic heterogeneity of RHE13Q—specifically, enrichment of Nia–L1/L1Q after light exposure and of Nia–L1Q/L2 after thermal transformation (Fig. 6a and S6)—produces a population of Cys-SH conformers that largely lack the hydrogen bond stabilizing the protonated thiolate in native Nia–L2 (Fig. 7d). Consequently, the associated νSH bands are expected to be substantially broadened, less polarized and therefore exhibit substantially lower intensity, rendering them undetectable in the experimental spectra. Nevertheless, protonation of Cys479 in RHE13Q is supported by the close similarities between the IR (difference) spectra and the EPR signatures of the Nia–L species of RHE13Q and native RH (Fig. 6, 7a, and S6), indicating that the first-sphere and most second-sphere interactions of the [NiFe] active site are largely preserved in the variant.
4. Conclusion
The present study elucidated the dynamics of residues surrounding the [NiFe] active site in the catalytic cycle of hydrogenases by using a variant of the C. necator regulatory hydrogenase in which the conserved proton-accepting glutamate (Glu13) located in the second coordination sphere of the metal center was replaced by glutamine. We performed comprehensive biochemical and spectroscopic analyses, including low-temperature IR, RR, and EPR, and compared the results with those of native RH. According to IR and RR spectroscopy, RHE13Q protein remained in the Nia–S state, as observed for RH.17 Nevertheless, the E13Q variant exhibited only 0.1% of the catalytic activity of native RH, which is consistent with previous studies on other [NiFe]-hydrogenases.30,31,38,39 Exposure of RHE13Q to H2 led to the accumulation of the reduced active site species Nia–C and Nia–SR. Subsequent illumination of the hydrogenase at 90 K induced the photolysis of the bridging hydride of the Nia–C state, resulting in the Nia–L1 state. The corresponding light-minus-dark IR difference spectra for this photoreaction were almost identical for RHE13Q and native RH, indicating very similar structural and conformational changes in both proteins during the Nia–C to Nia–L1 transition. These results suggest that the sequence Nia–S → Nia–SR → Nia–C → Nia–L1, including the first proton release during the Nia–SR to Nia–C transition, proceeds in the same way in the E13Q variant as in native RH. This, in turn, indicates that Glu13 does not play a role in the catalytic cycle from Nia–S to Nia–L1 (Fig. 1).Comparative low-temperature IR spectroscopy of CO and CN− ligands revealed two notable differences: (1) compared to native RH, the variant exhibited subforms for all major catalytic states, and (2) these subforms showed IR shifts of ∼6–8 cm−1 to higher frequencies, consistent with the loss of the negative charge of Glu13. Resonance Raman data supported this with red-shifted Fe–CN/CO vibrations. Interestingly, minor subforms of Nia–S, Nia–C, and Nia–L1 in the variant closely resembled native RH spectra, suggesting the presence of a compensatory hydroxide ligand near the active site, as observed in the corresponding E/Q variant of EcHyd-1.31 This entity could partially substitute for Glu13 in proton transfer, albeit inefficiently. However, the conserved glutamate is necessary for the Nia–L1 to Nia–L2 transition, which is supported by the observation that RHE13Q largely remained in a non-reactive sub-form of Nia–L1 (termed Nia–L1Q) after thermal transformation, while native RH formed the Nia–L2 state that is characterized by defined outer-sphere interactions. These findings differ from previous studies by the Dyer group on PfSH.40 In fact, while the Glu-to-Gln exchange in PfSH has been reported to perturb the Nia–L1/L2 → Nia–S transition, our data on RH suggest that the same exchange disrupts outer-sphere dynamics at an earlier stage, namely at Nia–L1. Based on our results, we assign a new mechanistic role to the conserved glutamate residue. It triggers critical outer-sphere rearrangements, such as the formation of a hydrogen bond with the protonated Ni-bound cysteine, which are essential for an efficient proton-transfer (PT) and thus, for the progression of the catalytic cycle.
An even more comprehensive picture emerges when our findings are combined with the recent preprint by Carr, Ash, Vincent and co-workers, which describes the mobility of the conserved glutamate side chain during the conversion of two Nia–L subforms of EcHyd-1 and EcHyd-2 based on X-ray structural analyses.32 The results emphasize the relevance of the Nia–L2 state and its participation into the hydrogenase catalytic cycle, as shown in Fig. 1. Ultimately, the detailed insights presented here underscore the crucial role of the protein environment in modulating the reactivity of transition metal catalysts—a key factor in the development of synthetic catalysts and biomimetic systems for efficient proton and electron transfer.54–57 This is particularly relevant for (semi)artificial systems such as Ni-substituted rubredoxins and short Ni-binding peptides, which have been proposed as promising hydrogenase mimetics. In several cases, these systems have been suggested to operate via a protonated cysteine thiolate,55,57 although direct experimental evidence remains limited. Our findings highlight the importance of a precisely engineered outer coordination sphere for stabilizing and controlling reactive intermediates. We therefore propose that the deliberate introduction of a tailor-made outer-sphere, increasingly enabled by modern machine-learning-based protein design approaches (e.g., AlphaFold 2/3,58,59 RoseTTAFold,60 RFdiffusion,61 DeepPredict,62 ProteinMPNN,63 and Boltz-264) can significantly enhance catalytic performance and facilitate the stabilization and characterization of key reaction intermediates.
Conflicts of interest
There are no conflicts of interest to declare.Data availability
The authors declare that the data supporting the findings of this study are available within the article and the supplementary information (SI). See DOI: https://doi.org/10.1039/d6ey00004e.Acknowledgements
This work was funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under Germany's Excellence Strategy-EXC 2008-390540038 (“Unifying Systems in Catalysis-UniSysCat”). O. L. and I. Z. acknowledge the support from EU Horizon 2020/McGEA/Proposal ID 101183014 HORIZON-MSCA-2023-SE-01-01.References
- K. A. Vincent, A. Parkin and F. A. Armstrong, Investigating and Exploiting the Electrocatalytic Properties of Hydrogenases, Chem. Rev., 2007, 107, 4366–4413 CrossRef CAS PubMed.
- W. Lubitz, H. Ogata, O. Rüdiger and E. Reijerse, Hydrogenases, Chem. Rev., 2014, 114, 4081–4148 CrossRef CAS PubMed.
- G. Caserta, S. Hartmann, C. Van Stappen, C. Karafoulidi-Retsou, C. Lorent, S. Yelin, M. Keck, J. Schoknecht, I. Sergueev, Y. Yoda, P. Hildebrandt, C. Limberg, S. DeBeer, I. Zebger, S. Frielingsdorf and O. Lenz, Stepwise Assembly of the Active Site of [NiFe]-Hydrogenase, Nat. Chem. Biol., 2023, 19, 498–506 CrossRef CAS PubMed.
- H. Ogata, W. Lubitz and Y. Higuchi, Structure and Function of [NiFe] Hydrogenases, J. Biochem., 2016, 160, 251–258 CrossRef CAS PubMed.
- H. Ogata, T. Krämer, H. Wang, D. Schilter, V. Pelmenschikov, M. Van Gastel, F. Neese, T. B. Rauchfuss, L. B. Gee, A. D. Scott, Y. Yoda, Y. Tanaka, W. Lubitz and S. P. Cramer, Hydride Bridge in [NiFe]-Hydrogenase Observed by Nuclear Resonance Vibrational Spectroscopy, Nat. Commun., 2015, 6, 7890 CrossRef CAS PubMed.
- G. Caserta, V. Pelmenschikov, C. Lorent, A. F. Tadjoung Waffo, S. Katz, L. Lauterbach, J. Schoknecht, H. Wang, Y. Yoda, K. Tamasaku, M. Kaupp, P. Hildebrandt, O. Lenz, S. P. Cramer and I. Zebger, Hydroxy-Bridged Resting States of a [NiFe]-Hydrogenase Unraveled by Cryogenic Vibrational Spectroscopy and DFT Computations, Chem. Sci., 2021, 12, 2189–2197 RSC.
- P. M. Vignais and B. Billoud, Occurrence, Classification, and Biological Function of Hydrogenases: An Overview, Chem. Rev., 2007, 107, 4206–4272 CrossRef CAS PubMed.
- C. Greening, A. Biswas, C. R. Carere, C. J. Jackson, M. C. Taylor, M. B. Stott, G. M. Cook and S. E. Morales, Genomic and Metagenomic Surveys of Hydrogenase Distribution Indicate H2 Is a Widely Utilised Energy Source for Microbial Growth and Survival, ISME J., 2016, 10, 761–777 CrossRef CAS PubMed.
- S. T. Stripp, B. R. Duffus, V. Fourmond, C. Léger, S. Leimkühler, S. Hirota, Y. Hu, A. Jasniewski, H. Ogata and M. W. Ribbe, Second and Outer Coordination Sphere Effects in Nitrogenase, Hydrogenase, Formate Dehydrogenase, and CO Dehydrogenase, Chem. Rev., 2022, 122, 11900–11973 CrossRef CAS PubMed.
- C. Greco, V. Fourmond, C. Baffert, P. Wang, S. Dementin, P. Bertrand, M. Bruschi, J. Blumberger, L. De Gioia and C. Léger, Combining Experimental and Theoretical Methods to Learn About the Reactivity of Gas-Processing Metalloenzymes, Energy Environ. Sci., 2014, 7, 3543–3573 RSC.
- A. Fasano, V. Fourmond and C. Léger, Outer-Sphere Effects on the O2 Sensitivity, Catalytic Bias and Catalytic Reversibility of Hydrogenases, Chem. Sci., 2024, 15, 5418–5433 RSC.
- P. A. Ash, R. Hidalgo and K. A. Vincent, Proton Transfer in the Catalytic Cycle of [NiFe] Hydrogenases: Insight from Vibrational Spectroscopy, ACS Catal., 2017, 7, 2471–2485 CrossRef CAS PubMed.
- H. Tai, Y. Higuchi and S. Hirota, Comprehensive Reaction Mechanisms at and Near the Ni–Fe Active Sites of [NiFe] Hydrogenases, Dalton Trans., 2018, 47, 4408–4423 RSC.
- B. L. Greene, G. E. Vansuch, B. C. Chica, M. W. W. Adams and R. B. Dyer, Applications of Photogating and Time Resolved Spectroscopy to Mechanistic Studies of Hydrogenases, Acc. Chem. Res., 2017, 50, 2718–2726 CrossRef CAS PubMed.
- Y. Ilina, C. Lorent, S. Katz, J. Jeoung, S. Shima, M. Horch, I. Zebger and H. Dobbek, X-ray Crystallography and Vibrational Spectroscopy Reveal the Key Determinants of Biocatalytic Dihydrogen Cycling by [NiFe] Hydrogenases, Angew. Chem., Int. Ed., 2019, 58, 18710–18714 CrossRef CAS PubMed.
- G. Caserta, C. Lorent, V. Pelmenschikov, J. Schoknecht, Y. Yoda, P. Hildebrandt, S. P. Cramer, I. Zebger and O. Lenz, In Vitro Assembly as a Tool to Investigate Catalytic Intermediates of [NiFe]-Hydrogenase, ACS Catal., 2020, 10, 13890–13894 CrossRef CAS PubMed.
- M. Horch, J. Schoknecht, M. A. Mroginski, O. Lenz, P. Hildebrandt and I. Zebger, Resonance Raman Spectroscopy on [NiFe] Hydrogenase Provides Structural Insights into Catalytic Intermediates and Reactions, J. Am. Chem. Soc., 2014, 136, 9870–9873 CrossRef CAS PubMed.
- H. Ogata, K. Nishikawa and W. Lubitz, Hydrogens Detected by Subatomic Resolution Protein Crystallography in a [NiFe] Hydrogenase, Nature, 2015, 520, 571–574 CrossRef CAS PubMed.
- C. Fan, M. Teixeira, J. Moura, I. Moura, H. B. Hanh, J. Le Gall, H. D. Peck and B. M. Hoffman, Detection and Characterization of Exchangeable Protons Bound to the Hydrogen-Activation Nickel Site of Desulfovibrio gigas Hydrogenase: a Proton and Deuterium Q-Band ENDOR Study, J. Am. Chem. Soc., 1991, 113, 20–24 CrossRef CAS.
- M. Brecht, M. Van Gastel, T. Buhrke, B. Friedrich and W. Lubitz, Direct Detection of a Hydrogen Ligand in the [NiFe] Center of the Regulatory H2-Sensing Hydrogenase from Ralstonia eutropha in Its Reduced State by HYSCORE and ENDOR Spectroscopy, J. Am. Chem. Soc., 2003, 125, 13075–13083 CrossRef CAS PubMed.
- G. Caserta, K. Laun, J.-P. Oudsen, I. Sergueev, I. Zebger and O. Lenz, NRVS Spectroscopy Resolves Distinct Bridging Hydride Intermediates in [NiFe]-Hydrogenase, J. Am. Chem. Soc., 2025, 147, 41216–41220 CrossRef CAS PubMed.
- H. Tai, K. Nishikawa, Y. Higuchi, Z. Mao and S. Hirota, Cysteine SH and Glutamate COOH Contributions to [NiFe] Hydrogenase Proton Transfer Revealed by Highly Sensitive FTIR Spectroscopy, Angew. Chem., Int. Ed., 2019, 58, 13285–13290 CrossRef CAS PubMed.
- E. Siebert, M. Horch, Y. Rippers, J. Fritsch, S. Frielingsdorf, O. Lenz, F. Velazquez Escobar, F. Siebert, L. Paasche, U. Kuhlmann, F. Lendzian, M. Mroginski, I. Zebger and P. Hildebrandt, Resonance Raman Spectroscopy as a Tool to Monitor the Active Site of Hydrogenases, Angew. Chem., Int. Ed., 2013, 52, 5162–5165 CrossRef CAS PubMed.
- H. Tai, K. Nishikawa, M. Suzuki, Y. Higuchi and S. Hirota, Control of the Transition between Ni-C and Ni-SIa States by the Redox State of the Proximal Fe–S Cluster in the Catalytic Cycle of [NiFe] Hydrogenase, Angew. Chem., Int. Ed., 2014, 53, 13817–13820 CrossRef CAS PubMed.
- R. Hidalgo, P. A. Ash, A. J. Healy and K. A. Vincent, Infrared Spectroscopy During Electrocatalytic Turnover Reveals the Ni-L Active Site State During H2 Oxidation by a NiFe Hydrogenase, Angew. Chem., Int. Ed., 2015, 54, 7110–7113 CrossRef CAS PubMed.
- C. Lorent, V. Pelmenschikov, S. Frielingsdorf, J. Schoknecht, G. Caserta, Y. Yoda, H. Wang, K. Tamasaku, O. Lenz, S. P. Cramer, M. Horch, L. Lauterbach and I. Zebger, Exploring Structure and Function of Redox Intermediates in [NiFe]-Hydrogenases by an Advanced Experimental Approach for Solvated, Lyophilized and Crystallized Metalloenzymes, Angew. Chem., Int. Ed., 2021, 60, 15854–15862 CrossRef CAS PubMed.
- F. Roncaroli, E. Bill, B. Friedrich, O. Lenz, W. Lubitz and M.-E. Pandelia, Cofactor Composition and Function of a H2 -Sensing Regulatory Hydrogenase as Revealed by Mössbauer and EPR Spectroscopy, Chem. Sci., 2015, 6, 4495–4507 RSC.
- M. Horch, J. Schoknecht, S. L. D. Wrathall, G. M. Greetham, O. Lenz and N. T. Hunt, Understanding the Structure and Dynamics of Hydrogenases by Ultrafast and Two-Dimensional Infrared Spectroscopy, Chem. Sci., 2019, 10, 8981–8989 RSC.
- A. F. T. Waffo, C. Lorent, S. Katz, J. Schoknecht, O. Lenz, I. Zebger and G. Caserta, Structural Determinants of the Catalytic Nia–L Intermediate of [NiFe]-Hydrogenase, J. Am. Chem. Soc., 2023, 145, 13674–13685 CrossRef PubMed.
- S. Dementin, B. Burlat, A. L. De Lacey, A. Pardo, G. Adryanczyk-Perrier, B. Guigliarelli, V. M. Fernandez and M. Rousset, A Glutamate Is the Essential Proton Transfer Gate during the Catalytic Cycle of the [NiFe] Hydrogenase, J. Biol. Chem., 2004, 279, 10508–10513 CrossRef CAS PubMed.
- R. M. Evans, P. A. Ash, S. E. Beaton, E. J. Brooke, K. A. Vincent, S. B. Carr and F. A. Armstrong, Mechanistic Exploitation of a Self-Repairing, Blocked Proton Transfer Pathway in an O2-Tolerant [NiFe]-Hydrogenase, J. Am. Chem. Soc., 2018, 140, 10208–10220 CrossRef PubMed.
- S. B. Carr, W. Li, K. L. Wong, R. M. Evans, S. E. T. Kendall-Price, K. A. Vincent and P. A. Ash, Glutamate “Flick” Enables Proton Tunneling During Fast Redox Biocatalysis, ChemRxiv, 2024, preprint DOI:10.26434/chemrxiv-2024-mn36l.
- T. Buhrke, O. Lenz, A. Porthun and B. Friedrich, The H2-Sensing Complex of Ralstonia eutropha: Interaction Between a Regulatory [NiFe] Hydrogenase and a Histidine Protein Kinase, Mol. Microbiol., 2004, 51, 1677–1689 CrossRef CAS PubMed.
- L. Kleihues, O. Lenz, M. Bernhard, T. Buhrke and B. Friedrich, The H2 Sensor of Ralstonia eutropha Is a Member of the Subclass of Regulatory [NiFe] Hydrogenases, J. Bacteriol., 2000, 182, 2716–2724 CrossRef CAS PubMed.
- R. Simon, U. Priefer and A. Pühler, A Broad Host Range Mobilization System for In Vivo Genetic Engineering: Transposon Mutagenesis in Gram Negative Bacteria, Nat. Biotechnol., 1983, 1, 784–791 CrossRef CAS.
- T. Buhrke, O. Lenz, N. Krauss and B. Friedrich, Oxygen Tolerance of the H2-Sensing [NiFe] Hydrogenase from Ralstonia eutropha H16 Is Based on Limited Access of Oxygen to the Active Site, J. Biol. Chem., 2005, 280, 23791–23796 CrossRef CAS PubMed.
- O. Lenz, L. Lauterbach and S. Frielingsdorf, Methods in Enzymology, Elsevier, 2018, vol. 613, pp. 117–151 Search PubMed.
- C. J. Kulka-Peschke, A.-C. Schulz, C. Lorent, Y. Rippers, S. Wahlefeld, J. Preissler, C. Schulz, C. Wiemann, C. C. M. Bernitzky, C. Karafoulidi-Retsou, S. L. D. Wrathall, B. Procacci, H. Matsuura, G. M. Greetham, C. Teutloff, L. Lauterbach, Y. Higuchi, M. Ishii, N. T. Hunt, O. Lenz, I. Zebger and M. Horch, Reversible Glutamate Coordination to High-Valent Nickel Protects the Active Site of a [NiFe] Hydrogenase from Oxygen, J. Am. Chem. Soc., 2022, 144, 17022–17032 CrossRef CAS PubMed.
- H. Adamson, M. Robinson, J. J. Wright, L. A. Flanagan, J. Walton, D. Elton, D. J. Gavaghan, A. M. Bond, M. M. Roessler and A. Parkin, Retuning the Catalytic Bias and Overpotential of a [NiFe]-Hydrogenase via a Single Amino Acid Exchange at the Electron Entry/Exit Site, J. Am. Chem. Soc., 2017, 139, 10677–10686 CrossRef CAS PubMed.
- B. L. Greene, G. E. Vansuch, C.-H. Wu, M. W. W. Adams and R. B. Dyer, Glutamate Gated Proton-Coupled Electron Transfer Activity of a [NiFe]-Hydrogenase, J. Am. Chem. Soc., 2016, 138, 13013–13021 CrossRef CAS PubMed.
- E. Szőri-Dorogházi, G. Maróti, M. Szőri, A. Nyilasi, G. Rákhely and K. L. Kovács, Analyses of the Large Subunit Histidine-Rich Motif Expose an Alternative Proton Transfer Pathway in [NiFe] Hydrogenases, PLoS One, 2012, 7, e34666 CrossRef PubMed.
- G. Caserta, C. Lorent, A. Ciaccafava, M. Keck, R. Breglia, C. Greco, C. Limberg, P. Hildebrandt, S. P. Cramer, I. Zebger and O. Lenz, The Large Subunit of the Regulatory [NiFe]-Hydrogenase from Ralstonia eutropha – a Minimal Hydrogenase?, Chem. Sci., 2020, 11, 5453–5465 RSC.
- C. M. Silveira, L. Zuccarello, C. Barbosa, G. Caserta, I. Zebger, P. Hildebrandt and S. Todorovic, Molecular Details on Multiple Cofactor Containing Redox Metalloproteins Revealed by Infrared and Resonance Raman Spectroscopies, Molecules, 2021, 26, 4852 CrossRef CAS PubMed.
- C. C. M. Bernitzky, G. Caserta, S. Frielingsdorf, J. Schoknecht, A. Schmidt, P. Scheerer, O. Lenz, P. Hildebrandt, C. Lorent, I. Zebger and M. Horch, Expanding the Scope of Resonance Raman Spectroscopy in Hydrogenase Research: New Observable States and Reporter Vibrations, J. Inorg. Biochem., 2025, 262, 112741 CrossRef CAS PubMed.
- C. Karafoulidi-Retsou, S. Katz, S. Frielingsdorf, O. Lenz, I. Zebger and G. Caserta, A Strong H-Bond Between a Cysteine and the Catalytic Center of a [NiFe]-Hydrogenase, Chem. Commun., 2025, 61, 5778–5781 RSC.
- C. Karafoulidi-Retsou, C. Lorent, S. Katz, Y. Rippers, H. Matsuura, Y. Higuchi, I. Zebger and M. Horch, Light-Induced Electron Transfer in a [NiFe] Hydrogenase Opens a Photochemical Shortcut for Catalytic Dihydrogen Cleavage, Angew. Chem., Int. Ed., 2024, e202409065 CAS.
- M. Pandelia, H. Ogata and W. Lubitz, Intermediates in the Catalytic Cycle of [NiFe] Hydrogenase: Functional Spectroscopy of the Active Site, ChemPhysChem, 2010, 11, 1127–1140 CrossRef CAS PubMed.
- H. Tai, S. Hirota and S. T. Stripp, Proton Transfer Mechanisms in Bimetallic Hydrogenases, Acc. Chem. Res., 2021, 54, 232–241 CrossRef CAS PubMed.
- C. Fichtner, M. Van Gastel and W. Lubitz, Wavelength Dependence of the Photo-Induced Conversion of the Ni–C to the Ni–L Redox State in the [NiFe] Hydrogenase of Desulfovibrio vulgaris Miyazaki F, Phys. Chem. Chem. Phys., 2003, 5, 5507–5513 RSC.
- P. A. Ash, S. E. T. Kendall-Price, R. M. Evans, S. B. Carr, A. R. Brasnett, S. Morra, J. S. Rowbotham, R. Hidalgo, A. J. Healy, G. Cinque, M. D. Frogley, F. A. Armstrong and K. A. Vincent, The Crystalline State as a Dynamic System: IR Microspectroscopy Under Electrochemical Control for a [NiFe] Hydrogenase, Chem. Sci., 2021, 12, 12959–12970 RSC.
- P. A. Ash, S. E. T. Kendall-Price and K. A. Vincent, Unifying Activity, Structure, and Spectroscopy of [NiFe] Hydrogenases: Combining Techniques to Clarify Mechanistic Understanding, Acc. Chem. Res., 2019, 52, 3120–3131 CrossRef CAS PubMed.
- V. A. Lorenz-Fonfria, Infrared Difference Spectroscopy of Proteins: From Bands to Bonds, Chem. Rev., 2020, 120, 3466–3576 CrossRef CAS PubMed.
- A. Barth, Infrared Spectroscopy of Proteins, Biochim. Biophys. Acta, Bioenerg., 2007, 1767, 1073–1101 CrossRef CAS PubMed.
- S. La Gatta, L. Leone, O. Maglio, M. De Fenza, F. Nastri, V. Pavone, M. Chino and A. Lombardi, Unravelling the Structure of the Tetrahedral Metal-Binding Site in METP3 through an Experimental and Computational Approach, Molecules, 2021, 26, 5221 CrossRef CAS PubMed.
- S. Malayam Parambath, A. E. Williams, L. A. Hunt, D. Selvan, N. I. Hammer and S. Chakraborty, A De Novo-Designed Artificial Metallopeptide Hydrogenase: Insights into Photochemical Processes and the Role of Protonated Cys, ChemSusChem, 2021, 14, 2237–2246 CrossRef CAS PubMed.
- J. W. Slater, S. C. Marguet, M. E. Gray, H. A. Monaco, M. Sotomayor and H. S. Shafaat, Power of the Secondary Sphere: Modulating Hydrogenase Activity in Nickel-Substituted Rubredoxin, ACS Catal., 2019, 9, 8928–8942 CrossRef CAS.
- J. Timm, D. H. Pike, J. A. Mancini, A. M. Tyryshkin, S. Poudel, J. A. Siess, P. M. Molinaro, J. J. McCann, K. M. Waldie, R. L. Koder, P. G. Falkowski and V. Nanda, Design of a Minimal Di-Nickel Hydrogenase Peptide, Sci. Adv., 2023, 9, eabq1990 CrossRef CAS PubMed.
- J. Jumper, R. Evans, A. Pritzel, T. Green, M. Figurnov, O. Ronneberger, K. Tunyasuvunakool, R. Bates, A. Žídek, A. Potapenko, A. Bridgland, C. Meyer, S. A. A. Kohl, A. J. Ballard, A. Cowie, B. Romera-Paredes, S. Nikolov, R. Jain, J. Adler, T. Back, S. Petersen, D. Reiman, E. Clancy, M. Zielinski, M. Steinegger, M. Pacholska, T. Berghammer, S. Bodenstein, D. Silver, O. Vinyals, A. W. Senior, K. Kavukcuoglu, P. Kohli and D. Hassabis, Highly Accurate Protein Structure Prediction with AlphaFold, Nature, 2021, 596, 583–589 CrossRef CAS PubMed.
- J. Abramson, J. Adler, J. Dunger, R. Evans, T. Green, A. Pritzel, O. Ronneberger, L. Willmore, A. J. Ballard, J. Bambrick, S. W. Bodenstein, D. A. Evans, C.-C. Hung, M. O’Neill, D. Reiman, K. Tunyasuvunakool, Z. Wu, A. Žemgulytė, E. Arvaniti, C. Beattie, O. Bertolli, A. Bridgland, A. Cherepanov, M. Congreve, A. I. Cowen-Rivers, A. Cowie, M. Figurnov, F. B. Fuchs, H. Gladman, R. Jain, Y. A. Khan, C. M. R. Low, K. Perlin, A. Potapenko, P. Savy, S. Singh, A. Stecula, A. Thillaisundaram, C. Tong, S. Yakneen, E. D. Zhong, M. Zielinski, A. Žídek, V. Bapst, P. Kohli, M. Jaderberg, D. Hassabis and J. M. Jumper, Accurate Structure Prediction of Biomolecular Interactions with AlphaFold 3, Nature, 2024, 630, 493–500 CrossRef CAS PubMed.
- M. Baek, F. DiMaio, I. Anishchenko, J. Dauparas, S. Ovchinnikov, G. R. Lee, J. Wang, Q. Cong, L. N. Kinch, R. D. Schaeffer, C. Millán, H. Park, C. Adams, C. R. Glassman, A. DeGiovanni, J. H. Pereira, A. V. Rodrigues, A. A. Van Dijk, A. C. Ebrecht, D. J. Opperman, T. Sagmeister, C. Buhlheller, T. Pavkov-Keller, M. K. Rathinaswamy, U. Dalwadi, C. K. Yip, J. E. Burke, K. C. Garcia, N. V. Grishin, P. D. Adams, R. J. Read and D. Baker, Accurate Prediction of Protein Structures and Interactions Using a Three-Track Neural Network, Science, 2021, 373, 871–876 CrossRef CAS PubMed.
- J. L. Watson, D. Juergens, N. R. Bennett, B. L. Trippe, J. Yim, H. E. Eisenach, W. Ahern, A. J. Borst, R. J. Ragotte, L. F. Milles, B. I. M. Wicky, N. Hanikel, S. J. Pellock, A. Courbet, W. Sheffler, J. Wang, P. Venkatesh, I. Sappington, S. V. Torres, A. Lauko, V. De Bortoli, E. Mathieu, S. Ovchinnikov, R. Barzilay, T. S. Jaakkola, F. DiMaio, M. Baek and D. Baker, De Novo Design of Protein Structure and Function with RFdiffusion, Nature, 2023, 620, 1089–1100 CrossRef CAS PubMed.
- W. Alanazi, D. Meng and G. Pollastri, DeepPredict: a State-of-the-Art Web Server for Protein Secondary Structure and Relative Solvent Accessibility Prediction, Front. Bioinf., 2025, 5, 1607402 CrossRef PubMed.
- J. Dauparas, I. Anishchenko, N. Bennett, H. Bai, R. J. Ragotte, L. F. Milles, B. I. M. Wicky, A. Courbet, R. J. De Haas, N. Bethel, P. J. Y. Leung, T. F. Huddy, S. Pellock, D. Tischer, F. Chan, B. Koepnick, H. Nguyen, A. Kang, B. Sankaran, A. K. Bera, N. P. King and D. Baker, Robust Deep Learning–Based Protein Sequence Design Using ProteinMPNN, Science, 2022, 378, 49–56 CrossRef CAS PubMed.
- M. Grieswelle, S. Homberg, P. Janssen, J. Kaminski, J. Massa, A.-K. Prinz, S. Georgiev and O. Koch, A New Benchmark for Deep Learning Based Affinity Prediction: Solving the Inter-Protein Scoring Noise Problem, ChemRxiv, 2026, preprint DOI:10.26434/chemrxiv-2025-sf3cs/v2.
| This journal is © The Royal Society of Chemistry 2026 |