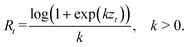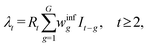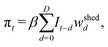Open Access Article
Open Access ArticleUnderstanding transmission and infections of respiratory syncytial virus through wastewater-based epidemiology
Katherine B.
Ensor
 *a,
Jose
Palacio
*a,
Jose
Palacio
 a,
Sallie A.
Keller
b,
Rebecca
Schneider
c,
Kaavya
Domakonda
c,
Loren
Hopkins
d and
Lauren
Stadler
a,
Sallie A.
Keller
b,
Rebecca
Schneider
c,
Kaavya
Domakonda
c,
Loren
Hopkins
d and
Lauren
Stadler
 e
e
aDepartment of Statistics, Rice University, MS 138, Houston, TX, USA. E-mail: ensor@rice.edu; Tel: +1 713 348 4687
bBiocomplexity Institute, University of Virginia, Charlottesville, NC, USA
cHouston Health Department, Houston, TX, USA
dHouston Health Department & Department of Statistics, Rice University, Houston, TX, USA
eDepartment of Civil & Environmental Engineering, Rice University, Houston, TX, USA
First published on 17th March 2026
Abstract
Wastewater-based epidemiology (WBE) is an effective tool for tracking community circulation of respiratory viruses. We address a scientific gap that takes measured wastewater viral load of respiratory syncytial virus (RSV) and estimate the effective reproduction number and the number of infections in the population. We advocate a modular approach to the analysis. We first estimate the trend and current level of the RSV viral load and quantify the uncertainty. These estimates become input for our Bayesian renewal model for both the infection rate and the number of infected individuals. The modular approach simplifies the analysis pipeline while maintaining scientific integrity. Further, the modular approach supports translation to other viruses by using disease-specific models for estimated transmission and cases in the second phase of the analysis.
Water impactWastewater surveillance provides a powerful approach for tracking viral infection dynamics in a community. Monitoring pathogens through sewer systems yields population-level information that informs public health guidance, resource allocation, and risk assessment. RSV wastewater monitoring provides timely estimates of the effective reproduction number and the number of infected individuals. |
1 Introduction
Wastewater-based epidemiology (WBE) has become an essential tool for monitoring infectious diseases at the population level. By analyzing viral genetic material in wastewater, WBE offers a non-invasive and cost-effective way to detect early signs of community spread, regardless of whether individuals seek medical care or undergo clinical testing.1For the respiratory syncytial virus (RSV), a leading cause of severe respiratory illness in infants, older adults, and immunocompromised people, clinical surveillance faces challenges from under reporting and delays in reporting. Many RSV cases are managed at home without hospitalization, so clinical records do not fully reflect community transmission. RSV measured in wastewater provides insight into community disease dynamics.2 In a comprehensive study of 176 sites during the 2022–2023 RSV season, Zulli et al.3 observed that RSV RNA concentrations at state and national levels were linked to infection positivity and hospitalization rates. Allen et al.4 detailed the implementation of a WBE approach in Northern Ireland to track RSV community transmission over the 2021 and 2022 seasons, correlating wastewater RSV levels with clinical cases. Through sequencing and phylogenetic analysis, they compared RSV A and B G-gene sequences from wastewater and clinical samples to elucidate transmission patterns. A study of an active international land border found that wastewater signals peaked in Detroit (MI, USA) for the 2022–2023 RSV season approximately 5 weeks prior to the peak in Windsor (ON, Canada).5 The authors further found a strong positive relationship between wastewater disease concentrations and hospitalization rates in the Canada location.
Building on the success of WBE for the RSV, we address a scientific gap that takes measured wastewater viral load of RSV and estimate the effective reproduction number and the number of infections in the population, advocating a modular approach to the analysis. We first estimate the trend and current level of the RSV viral load and quantify the uncertainty in the trend. These estimates become input for a Bayesian renewal model for both the infection rate and the number of infected individuals. The modular approach simplifies the analysis pipeline while maintaining scientific integrity. This approach further supports translation to other viruses by refinement of the estimate of the trend and using disease-specific models for estimated transmission and cases in the second phase of the analysis.
2 Methods
We estimate the weekly reproduction rate, Rt, and number of infections, It, from RSV RNA concentrations collected each Monday between January 16, 2023, and December 30, 2024. The samples are taken from the largest wastewater treatment plant in Houston, TX, which serves an estimated 551![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 150 residents. The facility's size and regular sampling ensure that it provides one of the most dependable wastewater signals in the city.
150 residents. The facility's size and regular sampling ensure that it provides one of the most dependable wastewater signals in the city.
The temporal relationship between the number of infections, It, is compared to the estimated number of RSV cases for the city during the same time period, proportionally allocated to the population served by the single treatment plant.
2.1 Wastewater measurements
Twenty-four-hour composite samples were collected weekly each Monday between January 16, 2023, and December 30, 2024. Laboratory analysis of the RSV RNA signal is available each Friday.6 Wastewater samples are collected on RSV RNA viral loads expressed in 109 genome copies per day (B gc per day).7,8 Viral loads are constructed by multiplying observed concentrations (gc L−1) by each plant's median daily influent flow (L per day) and rescaling by 109. In this analysis, values below the laboratory limit of detection (LOD) were set to a uniform random number between one-half of the LOD and the LOD.2.2 RSV syndromic data
RSV healthcare encounter data are sourced from the Houston Health Department's syndromic surveillance system, HHD-ESSENCE.9 Health encounter data reflect city-wide counts and include routine, urgent care, emergency department, and hospitalization encounters from healthcare facilities that are on boarded. The city-wide counts are proportionally adjusted based on the percentage of the population served by the wastewater treatment plant.2.3 Model and estimation
We use a two-step modular approach to the data analysis and estimating the temporal trajectory of the reproduction number and number of infections in the community.Following the approach in ref. 10, let xt denote the trend in wastewater RNA viral load, measured on a log10 basis, at time t. The dynamics of the trend are specified by the state equation as
(xt − xt−1) = (xt−1 − xt−2) + wt, wt ∼ N(0,![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) σ2w), σ2w), |
The observation equation of the state-space model is given by
| yt = xt + vt, vt ∼ N(0, σ2v), |
We estimate the parameters of the SSM by maximum likelihood. Parameter estimation is an iterative process of likelihood evaluation using the Kalman filter at a given set of parameters and nonlinear optimization moving us through the parameter space. For a fixed parameterization, the Kalman filter provides efficient calculation of the filtered states or conditional means, and their uncertainties or conditional variances. We implement this process using the MARSS package in R. A nice feature is that this algorithm and software can easily manage missing values in our time series. The SSM framework coupled with the Kalman Filter technology, can also manage irregularly spaced observations as seen in ref. 11 and 12.
The output from stage one of our analysis is the estimated state or filtered value, which we denote by ![[x with combining circumflex]](https://www.rsc.org/images/entities/i_char_0078_0302.gif) t|t = E[xt|y1,…,t] at time t throughout our time series. In other words, the estimated state at time t is the expected value of the trend (on a log10 scale) at time t, given all observations through time t. We also obtain the estimated variances of the state at time t,
t|t = E[xt|y1,…,t] at time t throughout our time series. In other words, the estimated state at time t is the expected value of the trend (on a log10 scale) at time t, given all observations through time t. We also obtain the estimated variances of the state at time t, ![[P with combining circumflex]](https://www.rsc.org/images/entities/i_char_0050_0302.gif) t|t = Var[xt|y1,…,t], or the conditional variance of xt at time t.
t|t = Var[xt|y1,…,t], or the conditional variance of xt at time t.
Importantly, estimation is performed on the log10 scale and subsequently converted to the natural measurement scale (gc L−1), ensuring consistency with the lognormal likelihood adopted in the measurement layer of the renewal model. You can simplify this step by taking the natural log rather than log10; each work equally as well as we ultimately transform back to the original measurement scale.
2.4 Disease transmission dynamics
The effective reproduction number Rt is modeled through a scaled softplus transformation of an unconstrained unobserved variable zt for disease transmission, orThis transformation guarantees positivity of Rt while behaving approximately linear around zero and is numerically stable.
The unobserved variable for disease transmission, namely {zt}, is modeled as a random walk with normally distributed noise. This parameterization provides a parsimonious yet flexible representation of week-to-week changes in disease transmission. Specifically, zt = zt−1 + ε where ε ∼ N(0, σ2ε) and z1 ∼ N(1, σ2ε) with σε > 0. The parameter k controls the curvature of the link between zt and Rt and regulates how fluctuations in the process translate into changes in transmission.
Further, let It denote the number of new infections during week t. Following the renewal formulation,13 the expected number of cases, denoted by λt, is expressed as a convolution of past infections with the transmission-interval distribution:
The weekly temporal weights, winfg, for newly infected individuals are set by segments of a gamma probability density function (pdf) for weeks 1, …, G, and rescaled to sum to one. This construction is standard in renewal-based epidemic models.13–15 The weights are explicitly defined in ref. 16, and demonstrated in the code.
Following ref. 17, we assume a mean of 7.5 days and a standard deviation of 2.1 days for RSV for the gamma pdf to define the infection transmission weights.
There is not a one-to-one relationship between wastewater RNA copies and the number of infections, rather this relationship is inferred through our renewal process. The magnitude of It depends on our assumed initial conditions. For our purposes, we set the initial number to the citywide average weekly RSV health care encounters, scaled by the share of the city's population in the service area.18 This initial setting is considered conservative, as it does not account for individuals who did not seek medical care.
2.5 Linking wastewater to infections
We link the number of infections to RNA virus c per L through the shedding weights {wshedd}Dd=0, which incorporate the timing of viral shedding after infection. The expected wastewater value iswhere β > 0 converts infections into 109 genome copies per day per infection.
Similar to the infection transmission weights, the weekly weights ({wshedd}Dd=0) for the shedding distribution for a given individual are also modeled as a discretized gamma distribution, with maximum history of D weeks, and rescaled to sum to one. Clinical evidence provided in ref. 19 leads us to assume a mean of 4.6 days and a standard deviation of 2.0 days of the gamma pdf to obtain the temporal weights for the duration of viral shedding. Because our data are aggregated weekly, the interpretation of lag zero differs from daily formulation. Specifically, d = 0 corresponds to shedding occurring within the same calendar week as infection, which is biologically plausible given that RSV shedding can begin within the first few days of illness.20,21
The wastewater RNA trend estimate, say ût|t, is provided by step 1, where ût|t follows a lognormal distribution, and the log(ût|t) is normally distributed with mean ![[x with combining circumflex]](https://www.rsc.org/images/entities/i_char_0078_0302.gif) t|t and variance
t|t and variance ![[P with combining circumflex]](https://www.rsc.org/images/entities/i_char_0050_0302.gif) t|t. The latter are the output from the state equation of our SSM, in step 1. This parametrization implies that the expected value of ut is in fact πt, defining our link between the wastewater trend and the infection dynamics.
t|t. The latter are the output from the state equation of our SSM, in step 1. This parametrization implies that the expected value of ut is in fact πt, defining our link between the wastewater trend and the infection dynamics.
For the state-space model, we performed the estimations using maximum likelihood with BFGS optimization through the MARSS package,22 which utilizes Kalman filtering for efficient likelihood evaluation. The uncertainty in the parameter estimates for σv and σw of the SSM is not incorporated in the Bayesian renewal model. We estimate the parameters of the Bayesian renewal model using nimble23via MCMC. Prior distributions for the parameters β, k, and σε include a log-scale prior for β, a half-normal prior for the random-walk volatility σε, and a log-normal prior on k to regularize the curvature of the softplus link. Hyperparameters are chosen to ensure weakly informative priors. The MCMC sampler jointly estimated the unobserved values of Rt and It along with key distribution parameters, producing posterior samples for each. Multiple chains were implemented with warm-up iterations discarded. Convergence of the MCMC algorithm was assessed using the rank-normalized split ![[R with combining circumflex]](https://www.rsc.org/images/entities/i_char_0052_0302.gif) . Mixing was verified through the relative Monte Carlo standard error (relMCSE).
. Mixing was verified through the relative Monte Carlo standard error (relMCSE).
3 Results and discussion
3.1 Fitted model and diagnostics
The SSM defining the trend requires estimation of three parameters, specifically the initial state x0, the variance of the noise for the observation equation σ2v, and the variance of the noise for the state equation σ2w. These unbiased estimated parameters for the SSM defining the trend in the RSV wastewater viral load are![[x with combining circumflex]](https://www.rsc.org/images/entities/i_char_0078_0302.gif) 0 = 0.0074 (SE 0.6783),
0 = 0.0074 (SE 0.6783), 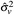 = 0.02145 (SE 0.0034), and
= 0.02145 (SE 0.0034), and 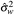 = 0.0005 (SE 0.0002). The important variance parameters are both small, with small standard errors, indicating that the model separates the trend from the sampling and measurement error. The uncertainty in these parameters is not included in the Bayesian renewal model. The estimated WW trend and its variance are conditional on the point estimates for the variance parameters. The accuracy of the estimates of these parameters supports this analysis decision and does not influence the stage 2 outcomes.
= 0.0005 (SE 0.0002). The important variance parameters are both small, with small standard errors, indicating that the model separates the trend from the sampling and measurement error. The uncertainty in these parameters is not included in the Bayesian renewal model. The estimated WW trend and its variance are conditional on the point estimates for the variance parameters. The accuracy of the estimates of these parameters supports this analysis decision and does not influence the stage 2 outcomes.
Additional assessment of the SSM assumptions is performed by examination of the standardized residuals. The standardized residuals of the fitted SSM model do not exhibit any additional autocorrelation over a 4 week history, based on the Ljung–Box test (p-value = 0.33). Further, the Kolmogorov–Smirnov test for normality of the standardized residuals indicate that normality is a reasonable assumption (p-value = 0.09). The SSM model does an excellent job of capturing the WW viral trend.
Table 1 summarizes posterior estimates and convergence diagnostics for the scalar parameters of the Bayesian renewal model. We observe strong convergence with rank-normalized split ![[R with combining circumflex]](https://www.rsc.org/images/entities/i_char_0052_0302.gif) values between 1.005 and 1.018. Further the chains are well mixed as indicated by the small relative Monte Carlo standard errors (relMCSEs) (≈0.01–0.02) across all parameters. The link-curvature parameter k centers around 8, consistent with a moderately sharp softplus mapping from zt to Rt. The innovation standard deviation σε is small (0.089), suggesting gradual week-to-week changes in transmission. For the wastewater, the scaling factor β is 14.865 (SD 3.388 with 95% credible interval [9.465, 22.352]). The wastewater observation dynamic variance is provided through the SSM, in other words, the time-varying
values between 1.005 and 1.018. Further the chains are well mixed as indicated by the small relative Monte Carlo standard errors (relMCSEs) (≈0.01–0.02) across all parameters. The link-curvature parameter k centers around 8, consistent with a moderately sharp softplus mapping from zt to Rt. The innovation standard deviation σε is small (0.089), suggesting gradual week-to-week changes in transmission. For the wastewater, the scaling factor β is 14.865 (SD 3.388 with 95% credible interval [9.465, 22.352]). The wastewater observation dynamic variance is provided through the SSM, in other words, the time-varying ![[P with combining circumflex]](https://www.rsc.org/images/entities/i_char_0050_0302.gif) t.
t.
![[R with combining circumflex]](https://www.rsc.org/images/entities/i_char_0052_0302.gif) /relMCSE) for scalar parameters. The SSM-filtered viral-load inputs with week-specific variances
/relMCSE) for scalar parameters. The SSM-filtered viral-load inputs with week-specific variances ![[P with combining circumflex]](https://www.rsc.org/images/entities/i_char_0050_0302.gif) t are treated as known
t are treated as known
| Parameter | Mean | SD | 95% CI |
![[R with combining circumflex]](https://www.rsc.org/images/entities/i_char_0052_0302.gif) /relMCSE /relMCSE |
|---|---|---|---|---|
| β | 14.737 | 3.969 | [8.591, 23.945] | 1.018/0.021 |
| k | 8.026 | 4.727 | [1.765, 18.600] | 1.005/0.009 |
| σ ε | 0.078 | 0.027 | [0.032, 0.137] | 1.017/0.019 |
3.2 Number of infections and transmission dynamics from wastewater
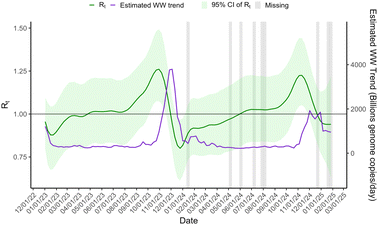
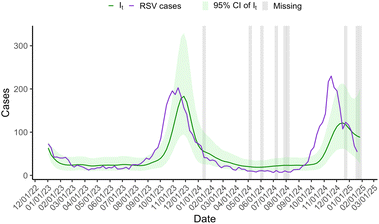
Fig. 1 displays posterior means and 95% credible bands for the estimated number of infections (It). As a reference, the estimated wastewater viral load in 109 RNA copies per day is also included. It rises sharply in late 2023, drops in January 2024, remains low through mid-2024, and shows a moderate uptick in autumn 2024. The peak in the estimated number of cases It occurs approximately one week before the peak in the wastewater trend. Because the absolute scale of It is not identified from wastewater alone, we interpret It in relative terms and anchor its level with an initial starting point of 64 case, estimated from the healthcare encounter data.Fig. 2 depicts the time series of the effective reproduction number (Rt). For the majority of the two-year period, the reproduction number is 1, indicating no change in disease levels. The reproduction number increases in October 2023 and again in October/November 2024, corresponding to the increase of disease measured in WW. Disease transmission quickly returned to stable levels after the rise, as indicated by reproduction numbers less than one in January 2024, and then stabilized at one.
The results shown in Fig. 1 and 2 were driven solely by wastewater trends. The only case information used in modeling was the initial starting value, which was set to 64 cases based on the observed RSV healthcare encounter cases for the week of January 2, 2023. Additionally, our modeling decisions for the transmission and shedding weights were based on scientific references, not estimated from our own data. For comparison, Fig. 3 shows posterior means and 95% credible bands for the estimated number of infections (It) and the observed time series of weekly RSV healthcare encounter cases, proportionally adjusted for the population served by the wastewater treatment plant.
The estimated number of cases closely follows the observed cases in both magnitude and pattern. The observed cases peak two weeks before the estimated cases during both periods of increased disease prevalence. During the second peak, wastewater levels did not rise as much as expected, leading to a lower case estimate, specifically It.
The mismatch in peak levels between the estimated cases and observed cases could be due to several choices in our modeling framework. The first is selecting a random walk for the transmission process. A modification that includes two weeks of history instead of one may be more appropriate. Additionally, the disease transmission weights and shedding weights for the renewal equations were derived from scientific literature rather than being specifically fitted to the RSV case data for this study area. This was an intentional decision for the study. Our aim was to evaluate how well we could model the case counts and reproduction number relying solely on wastewater.
Conclusions
Through this study we have demonstrated a practical and scientifically valid approach to estimate the effective reproduction number and the number of infections in a population where wastewater RNA copies per liter are measured. The focus of our application is the respiratory syncytial virus in a large sewershed serving over 500![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 people residing in Houston, TX. The two-stage approach separates estimating the wastewater viral trend from the downstream estimate of the infection counts and dynamics. This analytic approach simplifies the analysis pipeline, bringing actionable science to real-world applications in a timely fashion.
000 people residing in Houston, TX. The two-stage approach separates estimating the wastewater viral trend from the downstream estimate of the infection counts and dynamics. This analytic approach simplifies the analysis pipeline, bringing actionable science to real-world applications in a timely fashion.
Further, the two-stage approach allows the analyst to focus on two very different modeling challenges. By capturing the trend dynamics in wastewater, the analyst has a path forward for now casting and short-term forecasts of wastewater virus levels. The use of the SSM results in a clear separation of the underlying trend from the observation noise driven by both measurement and sampling uncertainty. We have specified a simple SSM model to track RSV RNA levels for our study area, and it worked well. However, the SSM technology is very flexible and can handle more complexity as needed for a specific situation. Further, the SSM technology supports missing or irregularly sampled wastewater time series. By handling these data issues early, we streamline the next step of the analysis, understanding the disease dynamics.
The stage-two Bayesian renewal model to estimate the number of individuals with the disease and the effective reproduction number can be implemented in an on-line fashion providing realtime actionable public health information. It is simpler than a Bayesian methodology that tries to address both modeling steps one and two simultaneously. Again, the input to the Bayesian renewal system is the estimated wastewater viral trend, and the uncertainty in that estimate.
An additional input to the stage-two Bayesian renewal model is the initial number of infected persons, representing the starting point of the disease. In our study, we used a proportional allocation of the infected population within Houston at the beginning of the study. This approach worked well, allowing us to accurately capture the levels and disease dynamics of RSV. It is important to note that this starting point serves as an anchor for estimating the number of infected individuals with the Bayesian renewal model. Essentially, we define the population with this setting. Since we relied on healthcare encounter data, our estimated number at later times reflects the same population. However, this estimate is lower than the actual number of infected individuals, as it does not include those who are sick but do not seek medical care. If there is reliable scientific data on the degree of under counting, this information could be used to refine the initial estimate of infected individuals. We also implicitly assume that the disease dynamics, for example the shedding distribution, does not change over time. If more scientific knowledge is available about the disease, this information can be incorporated into the modeling. Beyond this initial point, our disease trajectory is driven entirely by the wastewater trend estimate and its uncertainty.
The fact that the estimated number of infections measured from wastewater peaks after the healthcare encounter counts reflects the lag in observed wastewater levels. This scientific feature is worth further study to elevate the public health use of wastewater for understanding RSV spread in the community. The lag may be reflective of infected individuals in the population who are not seeking medical care. Advances in scientific knowledge about the disease and measurement dynamics can be incorporated into the stage two model to improve the public health utility of the wastewater estimated infections present in the community.
Wastewater-based epidemiology (WBE) is a proven effective tool for tracking community circulation of respiratory viruses. We contribute to the scientific gap that takes measured wastewater viral load of respiratory syncytial virus (RSV) and estimate the trajectory of the effective reproduction number and the number of infections in the population. Future statistical research is warranted to understand the mismatch in the timing of peak levels in number of RSV health care encounters. Specific areas to address include the assumption that disease transmission is a random walk, and the weights selected from the scientific literature for both the shedding and transmission distributions. Another hypothesis for the timing mismatch is that clinical data captures the onset of the disease by documenting severe cases, while cases estimated through wastewater reflect the spread of RSV to the larger and potentially less vulnerable population. These are questions that require further scientific investigation to answer.
Author contributions
Ensor, Rice Co-PI of the Houston Wastewater Epidemiology CDC National Wastewater Surveillance System Center of Excellence, is the corresponding author. She led the research and wrote the initial draft of the manuscript. Palacio researched, developed, and implemented the statistical methodology for disease transmission, producing all results in the paper and is lead author on a deeper statistical paper on this same topic. Keller collaborated on the two-stage implementation with specific attention to uncertainty quantification. Schneider oversees the day-to-day implementation of the HHD WBE system and represents the team in national wastewater surveillance data analysis conversations. Domakonda serves as HHD manager for this project and all wastewater surveillance programs for HHD. Hopkins serves as HHD PI for this project, and ensures expert science reaches decision makers and that the team is addressing the right questions. Stadler, Rice Co-PI of this project, oversees all aspects of the laboratory analyses for the Center, and brings the essential scientific leadership and expertise in WBE including quantitative areas. All authors contributed to the scientific discussion and edited the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
Wastewater RSV levels and health care encounter counts used in this manuscript are hosted on the Kinder Institute Urban Data Platform (https://www.kinderudp.org).24 R code for modeling is available through https://github.com/hou-wastewater-epi-org.Acknowledgements
The authors disclose receipt of the following financial support for the research, authorship, and/or publication of this article: this work was supported by the Centers for Disease Control and Prevention (ELC-ED grant no. 6NU50CK000557-01-05 and ELC427 CORE grant no. NU50CK000557). The authors acknowledge the Houston Public Works for their contributions to the HHD WBE system. The authors would like to acknowledge the CDC National Wastewater Surveillance System (NWSS) scientific community. For more information on the Houston Wastewater Epidemiology Center of Excellence, see https://www.hou-wastewater-epi.org.Notes and references
- M. B. Diamond, A. Keshaviah, A. I. Bento, O. Conroy-Ben, E. M. Driver, K. B. Ensor, R. U. Halden, L. P. Hopkins, K. G. Kuhn, C. L. Moe, E. C. Rouchka, T. Smith, B. S. Stevenson, Z. Susswein, J. R. Vogel, M. K. Wolfe, L. B. Stadler and S. V. Scarpino, Wastewater Surveillance of Pathogens Can Inform Public Health Responses, Nat. Med., 2022, 28, 1992–1995 CrossRef CAS PubMed.
- B. Hughes, D. Duong, B. J. White, K. R. Wigginton, E. M. G. Chan, M. K. Wolfe and A. B. Boehm, Respiratory syncytial virus (RSV) RNA in wastewater settled solids reflects RSV clinical positivity rates, medRxiv, 2021, preprint, posted December 5, 2021, DOI:10.1101/2021.12.01.21267014.
- A. Zulli, M. R. Varkila, J. Parsonnet, M. K. Wolfe and A. B. Boehm, Observations of Respiratory Syncytial Virus (RSV) Nucleic Acids inWastewater Solids Across the United States in the 2022–2023 Season: Relationships with RSV Infection Positivity and Hospitalization Rates, ACS ES&T Water, 2020, 4, 1657–1667 Search PubMed.
- D. M. Allen, M. I. Reyne, P. Allingham, A. Levickas, S. H. Bell, J. Lock, J. D. Coey, S. Carson, A. J. Lee, C. McSparron, B. F. Nejad, J. McKenna, M. Shannon, K. Li, T. Curran, L. J. Broadbent, D. G. Downey, U. F. Power, H. E. Groves, J. M. McKinley, J. W. McGrath, C. G. G. Bamford and D. F. Gilpin, Genomic Analysis and Surveillance of Respiratory Syncytial Virus Using Wastewater-Based Epidemiology, J. Infect. Dis., 2024, 230, e895–e904 CrossRef CAS PubMed.
- M. Beach, R. Corchis-Scott, Q. Geng, A. M. Podadera Gonzalez, O. Corchis-Scott, E. Harrop, J. Norton, A. Busch, R. A. Faust, B. Irwin, M. Aloosh, K. K. S. Ng and R. M. McKay, Wastewater-Based Surveillance of Respiratory Syncytial Virus Reveals a Temporal Disconnect in Disease Trajectory across an Active International Land Border, Environ. Health, 2025, 3, 425–435 CAS.
- L. Hopkins, D. Persse, K. Caton, K. Ensor, R. Schneider, C. Mc-Call and L. B. Stadler, Citywide Wastewater SARS-CoV-2 Levels Strongly Correlated with Multiple Disease Surveillance In dicators and Outcomes over Three COVID-19 Waves, Sci. Total Environ., 2023, 855, 158967 CrossRef CAS PubMed.
- J. Wu, K. B. Ensor, L. Hopkins and L. B. Stadler, Assessment and application of GeneXpert rapid testing for respiratory viruses in school wastewater, Environ. Sci.: Water Res. Technol., 2025, 11, 64–76 RSC.
- J. Wu, M. X. Wang, T. J. Treangen, K. Ensor, L. Hopkins and L. B. Stadler, Sensitivity, Throughput, and Cost Analysis of Concentration Methods for Multitarget Pathogen Wastewater Monitoring, ACS ES&T Water, 2025, 5, 5531–5541 Search PubMed.
- Houston Health Department, Respiratory Diseases Surveillance Report, 2024–2025 Influenza Season (MMWR Week 2025-02), Houston health department technical report, 2025 Search PubMed.
- K. B. Ensor, J. C. Schedler, J. Palacio, R. Schneider, K. Domakonda, L. Hopkins and L. B. Stadler, A Nonlinear Hierarchical Time Series Approach to Citywide Trend Assessment of Viruses, Hot Spot Signals, and Right-Sizing System, Data Sci. Sci., 2025, 4(1) DOI:10.1080/26941899.2025.2559690.
- R. H. Shumway and D. S. Stoffer, Time Series Analysis and Its Applications: With R Examples, Springer Nature, 5th edn, 2025 Search PubMed.
- D. Ouyang, L. Chung, M. Osborn, T. W. Schacker and C. R. Doss, Dynamic Linear Models for Wastewater-Based Epidemiology with Missing Values: an Application to Covid-19 Surveillance, Data Sci. Sci., 2025, 4, 2562199 CrossRef.
- C. Fraser, Estimating individual and household reproduction numbers in an emerging epidemic, PLoS One, 2007, 2, e758 CrossRef PubMed.
- J. S. Huisman, J. Scire, L. Caduff, X. Fernandez-Cassi, P. Ganesanandamoorthy, A. Kull, A. Scheidegger, E. Stachler, A. B. Boehm, B. Hughes, A. Knudson, A. Topol, K. R. Wigginton, M. K. Wolfe, T. Kohn, C. Ort, T. Stadler and T. R. Julian, Wastewater-based estimation of the effective reproductive number of SARS-CoV-2, Environ. Health Perspect., 2022, 130, 057011 CrossRef CAS PubMed.
- D. Champredon, I. Papst and W. Yusuf, ern: An R package to estimate the effective reproduction number using clinical and wastewater surveillance data, PLoS One, 2024, 19, e0305550 CrossRef CAS PubMed.
- J. R. Palacio, K. B. Ensor, S. A. Keller, R. Schneider, K. Domakonda, L. Hopkins and L. B. Stadler, Inferring Transmission Dynamics of Respiratory Syncytial Virus from Houston Wastewater, arXiv, 2025, preprint, arXiv:2511.17816, DOI:10.48550/arXiv.2511.17816.
- M. A. Vink, M. C. J. Bootsma and J. Wallinga, Serial intervals of respiratory infectious diseases: A systematic review and analysis, Am. J. Epidemiol., 2014, 180, 865–875 CrossRef PubMed.
- Houston Health Department, Respiratory Diseases Surveillance Report: 2024–2025 Influenza Season, MMWR Week 2025-02, 2025, https://www.houstonhealth.org/media/11526/download?inline=, Accessed 2025-09-30 Search PubMed.
- M. Cevik, P. A. Piedra, V. Demicheli, C. Cantarelli, D. Manno, S. Vergnano, S. Pullan and R. Harwood, Virology, transmission, and clinical features of respiratory syncytial virus in adults, BMJ, 2023, 380, e072292 Search PubMed.
- C. B. Hall, C. E. Long and K. C. Schnabel, Respiratory syncytial virus infections in previously healthy working adults, Clin. Infect. Dis., 2001, 33, 792–796 CrossRef CAS PubMed.
- J. P. DeVincenzo, C. M. El Saleeby and A. J. Bush, Respiratory syncytial virus load predicts disease severity in previously healthy infants, J. Infect. Dis., 2005, 191, 1861–1868 CrossRef PubMed.
- E. E. Holmes, E. J. Ward and K. Wills, MARSS: Multivariate autoregressive state-space models for analyzing time-series data, R J., 2012, 4, 11–19 CrossRef.
- P. de Valpine, D. Turek, C. Paciorek, C. Anderson-Bergman, D. Temple Lang and R. Bodik, Programming with models: writing statistical algorithms for general model structures with NIMBLE, J. Comput. Graph. Stat., 2017, 26, 403–413 CrossRef.
- L. Hopkins, K. Domakonda, K. Caton, L. Schneider, R. abd Stadler and K. Ensor, City of Houston Respiratory Syncytial Virus Wastewater Monitoring and Health Care Encounter Data (Version 1) [Dataset], 2026, https://www.kinderudp.org Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |