Open Access Article
Open Access ArticleSuitability of concentration and extraction protocols for genomic applications in wastewater-based epidemiology
Laura A. E.
Van Poelvoorde
 *a,
Bert
Bogaerts
a,
Kevin
Vanneste
a,
Bavo
Verhaegen
b,
Veronik
Hutse
c,
Sigrid C. J.
De Keersmaecker
*a,
Bert
Bogaerts
a,
Kevin
Vanneste
a,
Bavo
Verhaegen
b,
Veronik
Hutse
c,
Sigrid C. J.
De Keersmaecker
 a,
François E.
Dufrasne
d,
Koenraad
Van Hoorde
b and
Nancy
Roosens
*a
a,
François E.
Dufrasne
d,
Koenraad
Van Hoorde
b and
Nancy
Roosens
*a
aTransversal activities in Applied Genomics, Sciensano, Brussels, Belgium. E-mail: laura.vanpoelvoorde@sciensano.be; nancy.roosens@sciensano.be
bFoodborne pathogens, Sciensano, Brussels, Belgium
cEpidemiology of infectious diseases, Sciensano, Brussels, Belgium
dNational Influenza Centre, Department of Infectious Diseases in Human, Laboratory of Viral Diseases, Sciensano, Brussels, Belgium
First published on 25th March 2026
Abstract
Wastewater-based epidemiology (WBE) has emerged as a powerful tool for community monitoring, offering a non-invasive, cost-effective means of assessing public health. PCR-based methods, including digital droplet PCR (ddPCR), combined with reverse transcription (RT) for RNA viruses, are widely used for detection of known pathogens in wastewater. Recently, sequencing approaches have gained importance. Targeted amplicon sequencing enables high-sensitivity detection of specific pathogens, while metagenomic sequencing provides untargeted profiling of microbial and viral communities, supporting surveillance of novel pathogens and antimicrobial resistance (AMR). A major challenge across all genomic approaches remains the efficient concentration and extraction of nucleic acids, particularly for low-abundance targets like respiratory viruses. This study aimed to evaluate the fit-for-purpose of concentration and extraction protocols for wastewater surveillance, focussing on their suitability for genomic downstream applications: (RT-)ddPCR, amplicon sequencing and metagenomics. Both RNA (viral pathogens) and DNA (AMR markers) targets were included and experiments were performed on raw and spiked wastewater. The results revealed substantial variability in method performance. Ultrafiltration with a concentrating pipette (InnovaPrep) consistently yielded lower viral concentrations, whereas precipitation-based methods, magnetic capture and a column-based concentration/extraction kit provided higher yield and better sensitivity. PCR inhibitors reduced targeted sequencing efficiency, while metagenomics produced consistent taxonomic profiles across methods. Overall, the study highlights the importance of protocol optimization to improve sensitivity in WBE. Optimization of protocols, particularly for detecting low abundance pathogens and inhibitor removal, remains essential. Broader validation across diverse wastewater matrices and microbial species will be critical to strengthen the role of WBE in public health surveillance.
Water impactThis study evaluates the suitability of concentration and extraction protocols for genomic applications in wastewater-based epidemiology. Identifying effective methods for recovering pathogens from wastewater is critical for public health surveillance. By improving sensitivity and consistency, these optimized protocols enhance early-warning systems for infectious diseases and antimicrobial resistance. The findings support more reliable wastewater monitoring strategies, enabling communities to respond more rapidly to emerging health threats. |
1 Introduction
Wastewater-based epidemiology (WBE) has emerged as a powerful tool in public health surveillance.1 By providing a non-invasive way to monitor pathogens shed by infected individuals, WBE enables the detection and quantification of community-level pathogen circulation.2 This approach offers a timely and cost-effective complementary method to clinical surveillance, supporting early warning, outbreak preparedness and the assessment of overall community health.3WBE has previously been effective in monitoring enteric viruses such as poliovirus, norovirus and hepatitis A virus,4,5 as well as bacterial pathogens including Salmonella and Escherichia coli,6 and has more recently gained attention for its role in SARS-CoV-2 surveillance.7 PCR-based methods are the most commonly used downstream detection approaches in WBE with digital droplet PCR (ddPCR) and reverse transcription ddPCR (RT-ddPCR) offering high sensitivity and absolute quantification of nucleic acid targets. RT-ddPCR methods have been widely applied for the detection of viral pathogens such as SARS-CoV-2,8–10 respiratory syncytial virus (RSV),11 and influenza viruses,11 while ddPCR methods have been used to detect antimicrobial resistance (AMR) genes.12 In the context of wastewater surveillance, ddPCR is a powerful and widely used platform due to its absolute quantification and its reduction in the biases linked to the PCR inhibitors.13 Recently, also sequencing approaches have become more prominent in WBE. Targeted amplicon sequencing enables high-resolution monitoring of specific circulating pathogens and their variants.14 In contrast, metagenomics offers an untargeted and comprehensive view of microbial and viral communities in wastewater. While metagenomics remains primarily used in research settings, it holds great promise for the detection of novel and emerging pathogens as well as AMR markers their bacterial hosts.15 However, sensitivity remains a major challenge, as sequencing will typically large amounts of background DNA and RNA, while targets of interest may represent only a very small fraction of the total reads.
Despite significant technological advances in downstream genomic analysis, a major bottleneck remains: the efficiency of pathogen concentration and nucleic acid extraction methods. This limitation is particularly critical for detecting low-abundance targets, such as respiratory viruses or single-copy AMR genes within rare bacterial species, within complex matrices like wastewater. Enhancing these upstream processes is essential to improve the sensitivity and reliability of WBE. Indeed, pathogen detection in wastewater presents distinct challenges compared to clinical testing. Whereas clinical samples are collected directly from individuals with a known or suspected infection, and typically contain relatively high viral concentrations, wastewater samples exhibit much lower virus loads, and are further challenged by a complex matrix with numerous PCR inhibitors and diverse background DNA.16 Therefore, effective concentration and extraction methods are necessary to achieve sensitive and reliable detection without concentrating the inhibitors, especially for viruses, that are present at low concentrations in wastewater such as SARS-CoV-2, respiratory syncytial virus (RSV) and influenza viruses,17 and AMR genes that are present at low concentrations.12 Despite the widespread use of various concentration and extraction methods, there is little consensus on which one is the most efficient for specific downstream applications.18–20 Another consideration is whether to analyse the liquid or solid fraction of wastewater, or both, as different fractions may capture different pathogen populations.21,22
To address the current uncertainty regarding optimal concentration and extraction methods for wastewater-based surveillance of pathogens and AMR markers, this study aims to evaluate the fit-for-purpose of several protocols that have been applied in wastewater based epidemiology, with a focus on their suitability for typical genomic downstream applications, namely digital droplet PCR (ddPCR), reverse transcription digital droplet PCR (RT-ddPCR), amplicon sequencing, and metagenomics. In contrast to previous method-comparison studies that typically focus on one analytical approach, this study integrates (RT-)ddPCR, targeted amplicon sequencing, and untargeted metagenomics. Moreover, within the study RNA viral pathogens and DNA AMR markers are evaluated together, thus providing an assessment of concentration and extraction methods across multiple downstream genomic applications, rather than evaluating performance based on a single metric or endpoint. The assessment includes four concentration methods previously described in wastewater application, including PEG/NaCl precipitation,23 magnetic capture (Nanotrap magnetic virus particles),24 ultrafiltration with a concentrating pipette (InnovaPrep),25 and ultracentrifugation,26 in combination with two extraction kits, including QIAamp Viral RNA Mini Kit and QIAamp RNeasy PowerFecal Pro Kit. In addition, the ZymoPURE Water DNA/RNA Kit was included as it represents a commercial and integrated approach combining concentration and extraction into a single workflow. Rather than focussing on direct comparisons between individuals methods, the goal of this study was to evaluate which methods can consistently yield sufficient nucleic acid of adequate purity to support reproducible downstream results across multiple workflows. Within this study, the performance of the methods was assessed in terms of measured target concentrations, by the success of downstream sequencing, and by the consistency of results across multiple sampling dates and locations, rather than by technical replicate reproducibility or absolute nucleic acid yield alone. Both DNA (two AMR targets, namely blaCTX-M and tetM) and RNA (norovirus serotypes GI and GII, SARS-CoV-2, RSV A and B, and influenza A and B) targets were included to ensure a comprehensive evaluation. Given that some RNA targets such as respiratory viruses are typically only present at low concentration in raw wastewater samples, sensitivity and comparability may be compromised. Therefore, these methods were tested not only on raw (non-spiked) wastewater samples, but also on samples spiked with SARS-CoV-2, RSV, and influenza viruses, which are generally close to the limit of detection in raw samples. This dual approach enabled a more robust assessment of method performance under both realistic and controlled conditions.
2 Methods
2.1 Sample collection
Five influent wastewater samples were collected as 24 hour composite samples over a period of 2 months with a 2 week interval in the spring of 2024, from the two Belgian wastewater treatment plants. Once collected, the wastewater samples were transported the same day at 4 °C and the concentration and extraction methods were performed as soon as possible after arrival within 72 hours. For each sample, the pH and the conductivity were measured (Table S1).To evaluate the concentration and extraction methods, two types of samples were used, i.e., 50 mL of raw wastewater; and 50 mL of spiked wastewater. To ensure consistency across workflows, a starting volume of 50 mL wastewater was processed for all concentration methods. The spiked wastewater was prepared by adding at each time point (n = 5) clinical samples that contain influenza A and B virus, RSV A and B virus or SARS-CoV-2 virus to 600 mL of raw wastewater sample to obtain a final concentration of either 50 or 100 cp μL−1 (Table 1). The initial concentration was measured using their respective RT-ddPCR method described below. These samples were obtained through the Belgian National Influenza Centre from the two main surveillance systems in Belgium, ILI for mild infections and SARI for moderate and severe infections. These spiked wastewater samples will serve as positive controls to verify method performance under controlled higher-input conditions for viral targets that are typically present at low concentrations in raw wastewater. These spiked samples didn't undergo any additional pre-treatment steps. The spiked wastewater was used to enable relative, method-to-method comparison of concentration and extraction performance under controlled conditions because calculating the absolute recovery efficiencies will prove challenging due to background levels of the same targets already present in the sample.
| Sample | Pathogen | Final concentration (cp μL−1) |
|---|---|---|
| H3N2_23-IS-00280 | Influenza A H3N2 | 100 |
| H1N1_23-IS-01030 | Influenza A H1N1 | 50 |
| RSVA_A2 | RSV A | 100 |
| RSVB_WV/146 | RSV B | 50 |
| B_VIC_23-IG-00115 | Influenza B (VIC) | 50 |
| SC2_23-IS-00311 | SARS-CoV-2 | 100 |
An overview of sample handling, storage conditions, concentration and extraction workflows, target types, and downstream analytical methods is provided in Table S1.
2.2 Concentration and extraction methods
Various concentration and extraction methods were evaluated on all raw and spiked wastewater samples to determine their suitability for downstream (RT-)ddPCR, targeted sequencing and metagenomics. Concentration methods included the InnovaPrep Concentrating Pipette Select, Nanotrap magnetic virus particles, ultracentrifugation, and PEG/NaCl precipitation. Each method followed specific protocols involving chemical pre-treatment, centrifugation, and resuspension in buffer solutions, with all concentrated eluates stored at −80 °C prior to extraction (see SI Methods). The ZymoPURE Water DNA/RNA Kit was also included as a combined concentration–extraction approach.Two Qiagen kits were tested for the nucleic acid extraction: the QIAamp RNeasy PowerFecal Pro Kit, used with all concentration methods, and the QIAamp Viral RNA Mini Kit, only paired with the PEG/NaCl precipitation concentration method. Both kits were used according to the manufacturers' instructions, with slight modifications to accommodate sample volume (see SI Methods). The QIAamp RNeasy PowerFecal Pro and ZymoPURE Water DNA/RNA kit have inhibitor removal steps included in their protocol.
2.3 (RT-)ddPCR quantification
Quantification of RNA and DNA targets was performed using RT-ddPCR and ddPCR to evaluate the efficiency of concentration and extraction methods. RNA targets included SARS-CoV-2 (ORF1a and RdRp_IP4),8 influenza A/B, RSV A/B, and norovirus GI/GII (Table 2). DNA targets included the antimicrobial resistance genes blaCTX-M and tetM12 (Table 2).| Target | Name | 5′ → 3′ sequence | Ref. |
|---|---|---|---|
| bla CTX-M | bla CTX-M _FW | ACCAAYGATATYGCGGTKAT | 12 |
| bla CTX-M _RV | ACATCGCGRCGGCKYTCT | ||
| bla CTX-M _PR | FAM/TCGTGCGCCGCTG/MGB-Eclipse | ||
| tetM | tetM_FW | GGTTTCTCTTGGATACTTAAATCAATCR | 12 |
| tetM_RV | CCAACCATAYAATCCTTGTTCRC | ||
| tetM_PR | HEX/ATGCAGTTATGGARGGGATACGCTATGGY/IABkFQ | ||
| RSVA | RSVQA1 | GCTCTTAGCAAAGTCAAGTTGAATGA | 27 |
| RSVQA2 | TGCTCCGTTGGATGGTGTATT | ||
| RSVQA_PR | FAM/ACACTCAAC-ZEN-AAAGATCAACTTCTGTCATCCAGC/IABkFQ | ||
| RSVB | RSVQB1 | GATGGCTCTTAGCAAAGTCAAGTTAA | 27 |
| RSVQB2 | TGTCAATATTATCTCCTGTACTACGTTGAA | ||
| RSVQB_PR | HEX/TGATACATT-ZEN-AAATAAGGATCAGCTGCTGTCATCCA/IABkFQ | ||
| FluA | InfA_Forward | GACCRATCCTGTCACCTCTGAC | 28 |
| InfA_Reverse | AGGGCATTYTGGACAAAKCGTCTA | ||
| Infa_Probe | FAM/TGCAGTCCT-ZEN-CGCTCACTGGGCACG/IABkFQ | ||
| FluB | InflB_For | AAATACGGTGGATTAAATAAAAGCAA | 28 |
| InflB Rev | CCAGCAATAGCTCCGAAGAAA | ||
| InflB_Probe | /HEX/CACCCATAT-ZEN-TGGGCAATTTCCTATGGC/3IABkFQ | ||
| ORF1a | ORF1a-F | AGAAGATTGGTTAGATGATGATAGT | 8 |
| ORF1a-R | TTCCATCTCTAATTGAGGTTGAACC | ||
| ORF1a-P | FAM/TCCTCACTG-ZEN-CCGTCTTGTTGACCA/IABkFQ | ||
| RdRp | RdRp_IP4-F | GGTAACTGGTATGATTTCG | 8 |
| RdRp_IP4-R | CTGGTCAAGGTTAATATAGG | ||
| RdRp_IP4-P | HEX/TCATACAAA-ZEN-CCACGCCAGG/IABkFQ |
RT-ddPCR was performed using the One-Step RT-ddPCR Advanced Kit for Probes (Bio-Rad), and ddPCR was conducted using the 2× ddPCR Supermix for Probes (no dUTP, Bio-Rad), following the manufacturer's instructions (see SI Methods). Droplet generation and amplification were performed using the QX200™ Droplet Generator and T100™ Thermal Cycler (Bio-Rad), respectively, and results were read on the QX200™ Droplet Reader. Data analysis was performed using QuantaSoft software v1.7.4.0917 with manual thresholding (see SI Methods). Negative controls consist of no-template controls to monitor for contamination. The limit of detection (LOD) for each (RT-)ddPCR assay is assay specific and has been calculated previously for the SARS-CoV-2 genes9 and AMR genes.12 Although the other assays were not re-estimated, it is assumed to be consistent for the samples across all concentration and extraction methods.
For each target, generalized linear models with a Gaussian error distribution were fitted to compare concentrations obtained with different concentration and extractions methods, using the precipitation method combined with the QIAamp RNeasy PowerFecal Pro Kit as reference. The concentrations were log10-transformed prior to analysis. For spiked targets, models were fitted separately from the raw wastewater. Estimated coefficient were exponentiated to obtain fold changes relative to the reference method, and 95% confidence intervals were calculated. P-values for method comparisons were extracted from the model summaries and adjusted for multiple testing using the false discovery rate method.
2.4 Targeted sequencing
To evaluate whether the routine sequencing protocol remained effective and whether PCR amplification was impacted by the presence of inhibitors, SARS-CoV-2 sequencing was performed on wastewater samples from two time points and from one Belgian WWTP. SARS-CoV-2 was selected for this assessment as it remains one of the most well-established pathogens currently monitored in wastewater surveillance using targeted sequencing. This sequencing protocol, detailed in the SI Methods, was applied on the various samples for which all concentration and extraction methods were used.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000× using Seqtk sample (v1.4),29 based on total read bases versus reference genome length (NC_045512.2). The raw reads were then trimmed using fastp v0.23.4.30 Reads were aligned with BWA-MEM (v0.7.17)31 using −k 19 and −r 1.5. The trimmed paired and unpaired reads were mapped separately and merged with SAMtools merge (v1.17).32 Coverage statistics (median depth and genome coverage) were calculated using SAMtools depth v1.17 with the −a option (see SI Methods). Consensus sequences were generated through an iterative mapping approach, where reads were repeatedly aligned to an updated reference and variants were called and incorporated at each step. Iterations continued until the consensus stabilized or a set maximum was reached. Variants were filtered to ensure sufficient depth, quality, and allele frequency, resulting in high-confidence consensus sequences.
000× using Seqtk sample (v1.4),29 based on total read bases versus reference genome length (NC_045512.2). The raw reads were then trimmed using fastp v0.23.4.30 Reads were aligned with BWA-MEM (v0.7.17)31 using −k 19 and −r 1.5. The trimmed paired and unpaired reads were mapped separately and merged with SAMtools merge (v1.17).32 Coverage statistics (median depth and genome coverage) were calculated using SAMtools depth v1.17 with the −a option (see SI Methods). Consensus sequences were generated through an iterative mapping approach, where reads were repeatedly aligned to an updated reference and variants were called and incorporated at each step. Iterations continued until the consensus stabilized or a set maximum was reached. Variants were filtered to ensure sufficient depth, quality, and allele frequency, resulting in high-confidence consensus sequences.
2.5 Metagenomics
To characterize the communities within the sample and evaluate whether consistent taxonomic patterns were obtained across different methods, unbiased metagenomics was performed on wastewater samples collected at three time points from one Belgian WWTP. The primary objective of this metagenomic analysis was to assess the comparative consistency of taxonomic profiles generated by different upstream methods. This metagenomics protocol, detailed below, was applied on the various samples that were concentrated and extracted with Nanotrap method in combination with the QIAamp RNeasy PowerFecal Pro Kit, the precipitation method in combination with the QIAamp RNeasy PowerFecal Pro Kit and the ZymoPURE Water DNA/RNA Kit. These methods were selected for metagenomics because they yielded the highest overall nucleic acid concentrations in (RT-)ddPCR assays and demonstrated reliable amplification performance in SARS-CoV-2 targeted sequencing, ensuring that metagenomic comparisons were conducted on workflows demonstrated to be suitable for downstream genomic applications.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio of AMPure XP beads to DNA (see SI Methods).
1 ratio of AMPure XP beads to DNA (see SI Methods).
For adapter ligation, 25 μL ligation buffer, 10 μL quick T4 DNA ligase, and 5 μL ligation adapter (ONT Ligation Sequencing Kit V14) were added. After 10 min incubation at room temperature, samples were cleaned with AMPure XP beads and short fragment buffer to ensure DNA fragments from all sizes are retained. Libraries were eluted in 15 μL of elution buffer and quantified with Qubit BR to adjust to 35–50 fmol for sequencing on an ONT R10.4.1 Flow Cell run on a GridION device for 72 h using MinKNOW v24.02.16 (see SI Methods).
Reads were trimmed with SeqKit v2.3.1,34 and taxonomic classification was performed with Kraken2 v2.1.1 (ref. 35) using the in-house ‘full’ Kraken2 database (accessed 13 January 2024).36 Classification results were visualized using Krona,37 and comparisons across samples were visualized to depth level 5 via radar charts created in RStudio (v2022.02.2.485) using the fmsb package (v0.7.6) (see SI Methods).
3 Results
A comparative analysis was performed to evaluate the efficiency and reliability of various concentration and extraction methods for downstream analysis in wastewater-based detection of viral pathogens and AMR markers. First, (RT-)ddPCR was used to quantify recovery of both DNA and RNA targets, providing a measure of yield and sensitivity for AMR genes and viral pathogens, respectively. Second, the suitability of each method was tested for downstream SARS-CoV-2 targeted sequencing, with success defined as sufficient genome coverage and depth without strong evidence of inhibition. Finally, metagenomic sequencing was applied to characterize the broader microbial DNA community, with consistency defined as the similarity of taxonomic profiles across the different methods. This metagenomics analysis provided a comparative assessment of workflow consistency. Each analysis included both raw (with naturally low concentration of respiratory viruses) and spiked (relatively high concentration of respiratory viruses) wastewater samples collected from one Belgian WWTP.3.1 Evaluation based on the (RT-)ddPCR results
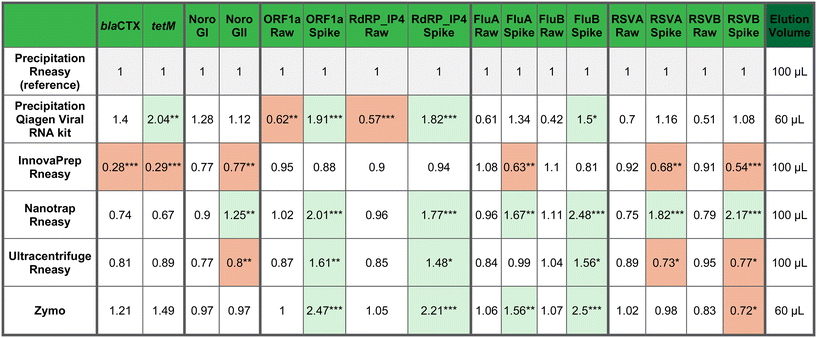
The various concentration and extraction methods were evaluated using (RT-)ddPCR with DNA (AMR targets: blaCTX-M and tetM) and RNA targets (ORF1a, RdRP_IP4, influenza A, influenza B, RSVA, RSVB, norovirus GI, norovirus GII). This evaluation included the following concentration methods: precipitation, Nanotrap magnetic virus particles, InnovaPrep and ultracentrifuge; in combination with the QIAamp RNeasy PowerFecal Pro Kit. Additionally, the precipitation method was also evaluated in combination with the QIAamp Viral RNA Mini Kit and finally, also the commercial kit ZymoPURE Water DNA/RNA Kit was evaluated.For the respiratory viruses, including SARS-CoV-2, RSV A, RSV B, influenza A, and influenza B, the analysis of raw wastewater samples generally showed no significant differences between the concentration or extraction methods (Table 3). This likely reflects the low viral concentrations typically present in these samples, which limits statistical power to distinguish between small differences in recovery efficiency. To address this, wastewater was also spiked with respiratory viruses, which provided a clearer basis for comparing methods, as higher input levels allowed differences in recovery to become detectable.
The comparative evaluation was conducted against a reference method namely the precipitation method paired with the QIAamp RNeasy PowerFecal Pro Kit. Across both raw and spiked samples, method performance was assessed in terms of recovery efficiency, measured as the absolute concentration of each target detected by (RT-)ddPCR. While results varied per target (Table 3), most differences between methods were not statistically significant. Notably, the InnovaPrep method consistently yielded the lowest concentration across all viral targets tested. In contrast, the precipitation, Nanotrap and ZymoPURE Water DNA/RNA methods generally produced higher recoveries indicating greater suitability for downstream applications. Moreover, within the specific concentration–extraction pairings evaluated, (RT-)ddPCR results indicated that both the QIAamp RNeasy PowerFecal Pro Kit and the QIAamp Viral RNA Mini Kit performed similarly when combined with PEG precipitation (Table 3). However, no inference can be made regarding their relative performance when combined with other concentration methods. Additional pairwise comparisons between methods can be derived from the model coefficients present in Table 3 or from the mean concentrations that are included in Table S2.
Moreover, a dilution-based qPCR inhibition assessment was performed by comparing Cq values obtained from undiluted extracts and 5-fold dilutions targeting the tetM gene (Table S6). For most concentration–extraction workflows, the observed Cq shifts following dilution were close to the expected value of 2.32 cycles, indicating limited PCR inhibition. However, pronounced deviations from the expected shift were observed for specific methods, most notably PEG precipitation combined with the QIAamp Viral RNA Mini Kit, where undiluted extracts showed delayed or absent amplification that was partially or fully recovered upon dilution.
3.2 Evaluation based on the SARS-CoV-2 targeted sequencing results
To evaluate whether the routine illumina sequencing protocol for SARS-CoV-2 could still be effectively applied and whether PCR amplification was impacted by PCR inhibitors in the wastewater, SARS-CoV-2 sequencing was applied on the samples collected at two time points from one Belgian WWTP. These samples were processed using all concentration and extraction methods evaluated in this study, and sequencing results were obtained for both raw and spiked wastewater samples.Sequencing results from the raw wastewater samples showed poor genome coverage and low median sequencing depth (Table 4), making it difficult to determine whether the suboptimal results were due to the inefficiency of the methods or simply due to the insufficient SARS-CoV-2 concentration for effective sequencing. Consequently, the spiked wastewater samples, which contained much higher SARS-CoV-2 concentrations, provided a more reliable evaluation of the effectiveness of the different protocols on the sequencing method.
| Date | Concentration method | Extraction method | ORF1a (cp μL−1) | R d R p (cp μL−1) | Median depth | Genome coverage | |
|---|---|---|---|---|---|---|---|
| Spiked wastewater | 18/03/2024 | INNOV | RN | 554.40 | 1243.00 | 8289.50 | 99.91% |
| NANO | RN | 2805.00 | 2651.00 | 10045.00 | 99.98% | ||
| Prec | QIA | 1826.00 | 2508.00 | 557.00 | 95.37% | ||
| Prec | RN | 1002.10 | 1430.00 | 9904.00 | 99.70% | ||
| UltraC | RN | 2134.00 | 2310.00 | 9849.00 | 99.71% | ||
| Zymo | Zymo | 4444.00 | 4400.00 | 10012.50 | 99.87% | ||
| 29/04/2024 | INNOV | RN | 1973.40 | 1870.00 | 10023.00 | 99.71% | |
| NANO | RN | 2301.20 | 2235.20 | 7740.50 | 99.73% | ||
| Prec | QIA | 4967.60 | 6292.00 | 7.00 | 89.84% | ||
| Prec | RN | 1036.20 | 1282.60 | 4318.00 | 99.81% | ||
| UltraC | RN | 2728.00 | 2820.40 | 10013.00 | 99.98% | ||
| Zymo | Zymo | 3256.00 | 3410.00 | 10032.00 | 99.81% |
In the spiked wastewater samples, most concentration and extraction methods yielded high genome coverage and median depth, indicating successful sequencing across workflows. Notably, the combination of PEG precipitation with the QIAamp Viral RNA Mini Kit consistently resulted in lower genome coverage and median depth compared to other methods. These results suggests that this specific pairing may be less optimal for targeted sequencing under the evaluated conditions and likely reflecting the co-concentration of PCR inhibitors, as neither the concentration nor extraction method includes an inhibitor removal step. While no internal amplification control (IAC) was included in the ddPCR assays to directly quantify inhibition, a limitation of this study, these findings provide valuable guidance for selecting concentration–extraction workflows that maximize sequencing success and support more reliable interpretation of viral detection wastewater.
3.3 Evaluation based on the metagenomics results
The characterization of the taxonomic profile in wastewater and the assessment of the consistency of these profiles were evaluated across the different concentration and extraction methods by performing metagenomic sequencing. The primary objective of this analysis was to assess the consistency of taxonomic profiles across selected workflows, rather than to comprehensively characterize viral diversity or benchmark sensitivity. The samples used for this evaluation were samples collected from at three time points from one Belgian WWTP, and nanopore sequencing was conducted on both raw and spiked wastewater samples for a selection of methods. These include the Nanotrap method in combination with the QIAamp RNeasy PowerFecal Pro Kit (red), the precipitation method in combination with the QIAamp RNeasy PowerFecal Pro Kit (blue) and the ZymoPURE Water DNA/RNA Kit. These were selected based on their superior performance in the (RT-)ddPCR results and SARS-CoV-2 sequencing results.The sequencing results (Table S4) show considerable variability in the total number of reads obtained per sample. This variability can likely partly be attributable to technical factors such as the number of active pores per flow cell, but also reflects the inherent heterogeneity of wastewater samples themselves, rather than being strongly associated with a specific method (Table S5). The median read quality and median read length remained consistent across all tested methods and did not show any significant associations with the tested method (Table S5).
When analysing the taxonomic profile of the non-spiked samples (Fig. 1), the Kraken2 results revealed that the community structures were highly similar between the different methods. Across all three sampling dates, bacterial sequences consistently represented the largest fraction of the classified reads. In contrast, very few viral reads were detected in both raw and spiked wastewater samples, preventing the construction of meaningful viral taxonomic profiles. This limited sensitivity can mask potential method-dependent differences, and the metagenomic results should therefore be interpreted as a consistency-base comparison rather than an evaluation of viral detection sensitivity. Fig. S1 provides an overview of the taxonomic profiles of the spiked samples, which also exhibited consistent trends.
4 Conclusions
WBE has become a valuable tool for the monitoring of pathogens and AMR genes at the population level, offering a cost-effective and non-invasive approach to track public health trends. However, it is possible that differences in concentration and extraction methods can directly affect the sensitivity and reliability of genomic downstream applications, including DNA and RNA quantification, targeted sequencing and metagenomics, in wastewater. This study assessed several concentration and extraction protocols, evaluating both DNA and RNA targets through (RT-)ddPCR, targeted sequencing and metagenomics profiling in wastewater samples. To account for the typically low concentrations of respiratory viruses in wastewater, both raw and spiked samples were analysed, thereby enabling evaluation of methods under low and controlled high concentrations.Overall, several protocols demonstrated robust and consistent recovery across both DNA and RNA targets, particularly the Nanotrap method paired with the QIAamp RNeasy PowerFecal Pro Kit and the ZymoPURE Water DNA/RNA Kit. These methods provided reproducible detection across viral (RNA) and AMR (DNA) markers by (RT-)ddPCR and yielded reliable SARS-CoV-2 genome coverage in targeted sequencing, within the specific concentration–extraction pairing evaluated, suggesting their suitability for applications requiring high sensitivity under these tested workflows. In contrast, the InnovaPrep method consistently resulted in lower recovery across both AMR and viral targets, indicating reduced efficiency in capturing nucleic acids. For the spiked samples, recovery efficiencies were not calculated in this study because that the spiked viruses were already present at a certain concentration. Future work should incorporate dedicated process controls to enable quantitative assessment of recovery.
In Belgium, the wastewater surveillance also includes SARS-CoV-2 sequencing at some critical time points, such as the emergence of a new variant. To ensure reliable results, it is important to confirm that the methods remain effective for amplifying SARS-CoV-2. However, targeted sequencing has shown some challenges in practice. Raw wastewater samples often contain very low amounts of virus, especially during periods of low circulation, which can lead to limited genome coverage and shallow sequencing depth. Under these conditions, variant detection and genomic surveillance become less reliable. To help overcome this limitation, spiked wastewater samples were also included and analysed. It was observed that the combination of PEG precipitation and QIAamp Viral RNA Mini Kit resulted in lower genome coverage and sequencing depth. This outcome was consistent in both raw and spiked wastewater samples, suggesting that the reduced performance was not solely due to the low viral concentration. The absence of an inhibitor removal step in this protocol may negatively impact amplification efficiency, particularly within the PEG precipitation–QIAamp Viral RNA Mini Kit workflow evaluated here. In contrast, the other methods that include a PCR inhibitor removal step, performed well, especially for the spiked samples, producing high coverage and depth within their respective concentration–extraction combinations. PCR inhibition is often cited as a concern in wastewater analysis depending on both concentration and extraction methods as well as the molecular target. PEG precipitation, in particular, has been associated with co-concentration of inhibitory compounds leading to an apparent reduced recovery and interference with PCR methods.38 A limited dilution-based qPCR assessment targeting tetM suggested that most methods showed Cq shifts close to the theoretical expectation, indicating no or limited inhibition. In contrast, the PEG precipitation concentration method in combination with the QIAamp Viral RNA Mini Kit displayed strongly aberrant Cq differences or amplification failure consistent with substantial PCR inhibition (Table S6). Although this assessment was not systematic and cannot be used for quantitative comparison, the observations align with previous reports of method- and matrix-dependent inhibition in wastewater and support the importance of inhibitor mitigation strategies when interpreting molecular results.
Metagenomics sequencing was performed only on the methods that had the best performance in the (RT-)ddPCR and targeted sequencing evaluations, namely the Nanotrap method in combination with the QIAamp RNeasy PowerFecal Pro Kit, the precipitation method in combination with the QIAamp RNeasy PowerFecal Pro Kit and the ZymoPURE Water DNA/RNA Kit. Focusing on the top-performing workflows enabled a robust and informative comparison of consistency across methods and provides valuable insights into their suitability for metagenomic applications, while appropriately aligning the analysis with the intended scope of the study rather than positioning it as a sensitivity benchmark. Interestingly, the taxonomic profiles were highly consistent across the selected concentration and extraction methods carried forward to metagenomic sequencing, indicating that once an effective concentration and extraction protocol is established, reliable taxonomic profiling can be achieved, regardless of the specific method used across different methods within this top-performing subset. By focusing on higher taxonomic levels, the analysis emphasizes the stability and reproducibility of the taxonomic profiles, underscoring the robustness of the selected workflows for comparative metagenomic analyses. This consistency is crucial for the interpretation of metagenomic data, as it suggests that reliable community-level insights can be obtained across different protocols. However, the taxonomic profiles were dominated by bacterial reads with little viral reads detected. This reflects a well-recognized limitation of untargeted metagenomics in wastewater, where RNA viruses are typically too scarce to be reliably recovered without enrichment or amplification. Low-abundance species, including most viruses, may therefore remain undetected despite apparent consistency in the bacterial profiles. Future studies could investigate probe-based capture, random amplification, or nanopore adaptive sampling as possible approaches to enhance viral signal recovery.
Our findings align with previous wastewater-based epidemiology studies, which generally report that no single concentration method is universally optimal across all targets and matrices. PEG/NaCl protocols can yield relatively high and reproducible recoveries for SARS-CoV-2 and other viruses, but also often tends to co-concentrate PCR inhibitors, leading to variable assay performance across wastewater matrices.38–41 This observation aligns with our results, where PEG precipitation paired with an extraction workflow lacking inhibitor removal showed reduced sequencing performance despite adequate target detection by RT-ddPCR. Ultrafiltration methods, such as InnovaPrep, are widely applied in wastewater surveillance but have shown variable performance for enveloped viruses and can be sensitive to clogging and sample composition.18,42–47 Magnetic capture approaches, including Nanotrap, have been reported to improve RNA virus recovery and extract purity, and in our study demonstrated robust and reproducible performance across multiple downstream applications.48–50 Importantly, whereas most previous studies evaluate a single target or analytical application, our work provides a unified fit-for purpose comparison of concentration and extraction methods across multiple viruses and AMR targets in RT-ddPCR along with SARS-CoV-2 sequencing and shotgun metagenomics using shared upstream methods. It should also be noted that wastewater physicochemical characteristics, such as solids content, organic load, and conductivity, are known to influence concentration efficiency and PCR inhibition, and may therefore contribute to some of the observed method-dependent differences. Within this study, the pH and conductivity were measured. pH values were consistently near neutral across all samples, while conductivity varied across time points and locations, indicating variability in wastewater composition (Table S1).
While analytical performance is critical for selecting concentration and extraction methods, practical considerations are also important for routine wastewater surveillance. Methods differed substantially in hands-on time, scalability, equipment requirement, and relative consumable costs (Table S7). Integrated or magnetic capture-based workflows offered most favourable balance between throughput, hands-on time and scalability, highlighting their suitability for routine surveillance. In contrast, precipitation- and ultracentrifugation based methods, while cost-efficient in terms of consumables, were more time-intensive and less scalable without significant investment, which may limit their applicability for high-throughput surveillance.
While this study offers valuable insights into methodological performance, several aspects offer opportunities for further refinement. First, although consistent detection across DNA and RNA targets was observed across the methods, some target-specific variability remains and could be further characterized. Second, the study was restricted to two wastewater treatment plants, so additional evaluations across diverse matrices and seasonal conditions would help generalize these findings. Third, while inhibition was not comprehensively assess using IAC, limited dilution-based testing suggested method-dependent inhibitory effects, partially mitigating this limitation. Standardized inhibition controls should be included in future studies to further refine performance comparisons. Furthermore, while our work demonstrates fit-for-purpose comparisons across multiple methods, controlled recovery experiments with technical replicates could provide precise quantification and establish method-specific thresholds under standardized conditions. Finally, only limited physicochemical metadata such as pH and conductivity has been collected. As a result, this may also influence the variability in target recovery. Future studies incorporating should focus on understanding these effects to better generalize method performance across wastewater types.
Taken together, the results offer a strong foundation for the further optimization of these protocols. The findings highlight the importance of PCR inhibitor removal and pathogen and AMR gene concentration optimization, and demonstrate that certain protocol are particularly well-suited for high-sensitivity downstream applications. Furthermore, the study underscores the need for further optimization for RNA virus recovery and for evaluation across a wider diversity of wastewater types and pathogen and AMR genes loads. Addressing these factors will help refine methodologies for WBE, ultimately improving the detection and genomic surveillance of public health threats at the community level.
Author contributions
Conceptualization: NR; project administration: NR; data curation: LVP, BV, KVH; methodology: LVP, FED; formal analysis: LVP; resources: SDK, KV; software: BB, LVP; validation: LVP; investigation: LVP; visualization: LVP; writing – original draft preparation: LVP, NR; writing – review & editing: all authors; funding acquisition: NR, VH; supervision: NR.Conflicts of interest
The authors declare that there are no conflicts of interest.Data availability
The data supporting this article have been included as part of the supplementary information (SI).Supplementary information: Table S1: Overview of sample handling, concentration and extraction workflows, physicochemical parameters, and downstream analytical methods. Table S2: Mean concentrations obtained by (RT-)ddPCR for viral and AMR targets across the evaluated concentration and extraction methods. Table S3: Comparison of concentration and extraction methods using targeted sequencing for SARS-CoV-2 across raw and spiked wastewater samples. Table S4: Overview of sequencing results for metagenomic analysis of wastewater samples collected from the one Belgian WWTP on three time points in 2024. Table S5: Statistical analysis of sequencing metrics, evaluating potential associations between the concentration/extraction methods and sequencing output characteristics. Table S6: Dilution-based qPCR inhibition assessment on selected wastewater samples using the tetM gene. Table S7: Qualitative practical consideration for concentration and extraction methods evaluated in this study in 2024. See DOI: https://doi.org/10.1039/d5ew00930h.
Acknowledgements
This study was financed by Sciensano through COVID-19 special funding by the COVIDDIVER project and by the regional and federal Belgian authorities. The samples provided in this study came from the national wastewater surveillance coordinated by Sciensano, which was funded by the regional (AVIQ, Vivalis and Departement Zorg Vlaanderen) and federal Belgian authorities. The surveillance was conducted thanks to the collaboration with the regional wastewater agencies (Hydria, Aquiris by Veolia, Aquafin and SPGE).References
- Wastewater-based epidemiology: current uses and future opportunities as a public health surveillance tool [Internet], [cited 2024 Jun 17], Available from: https://pubs.ciphi.ca/doi/full/10.5864/d2021-015.
- F. Wu, W. L. Lee, H. Chen, X. Gu, F. Chandra and F. Armas, et al., Making waves: Wastewater surveillance of SARS-CoV-2 in an endemic future, Water Res., 2022, 219, 118535 CrossRef CAS PubMed.
- J. I. Levy, K. G. Andersen, R. Knight and S. Karthikeyan, Wastewater surveillance for public health, Science, 2023, 379(6627), 26–27 CrossRef CAS PubMed.
- L. Chacón, E. Morales, C. Valiente, L. Reyes and K. Barrantes, Wastewater-Based Epidemiology of Enteric Viruses and Surveillance of Acute Gastrointestinal Illness Outbreaks in a Resource-Limited Region, Am. J. Trop. Med. Hyg., 2021, 105(4), 1004–1012 Search PubMed.
- T. Hovi, L. M. Shulman, H. van der Avoort, J. Deshpande, M. Roivainen and E. M. de Gourville, Role of environmental poliovirus surveillance in global polio eradication and beyond, Epidemiol. Infect., 2012, 140(1), 1–13 CrossRef CAS.
- Si. Bonetta, C. Pignata, E. Lorenzi, M. De Ceglia, L. Meucci and Sa. Bonetta, et al., Detection of pathogenic Campylobacter, E. coli O157:H7 and Salmonella spp. in wastewater by PCR assay, Environ. Sci. Pollut. Res., 2016, 23(15), 15302–15309 CrossRef CAS PubMed.
- M. Hamouda, F. Mustafa, M. Maraqa, T. Rizvi and A. A. Hassan, Wastewater surveillance for SARS-CoV-2: Lessons learnt from recent studies to define future applications, Sci. Total Environ., 2021, 759, 143493 CrossRef CAS PubMed.
- L. A. E. Van Poelvoorde, M. Gand, M. A. Fraiture, S. C. J. De Keersmaecker, B. Verhaegen and K. Van Hoorde, et al., Strategy to Develop and Evaluate a Multiplex RT-ddPCR in Response to SARS-CoV-2 Genomic Evolution, Curr. Issues Mol. Biol., 2021, 43(3), 1937–1949 CrossRef CAS PubMed.
- L. A. E. Van Poelvoorde, A. Gobbo, S. J. D. Nauwelaerts, B. Verhaegen, M. Lesenfants and R. Janssens, et al., Development of a reverse transcriptase digital droplet polymerase chain reaction-based approach for SARS-CoV-2 variant surveillance in wastewater, Water Environ. Res., 2024, 96(3), e10999 CrossRef CAS PubMed.
- L. A. E. Van Poelvoorde, C. Picalausa, A. Gobbo, B. Verhaegen, M. Lesenfants and P. Herman, et al., Development of a Droplet Digital PCR to Monitor SARS-CoV-2 Omicron Variant BA.2 in Wastewater Samples, Microorganisms, 2023, 11(3), 729 CrossRef CAS.
- A. Zafeiriadou, L. Kaltsis, N. S. Thomaidis and A. Markou, Simultaneous detection of influenza A, B and respiratory syncytial virus in wastewater samples by one-step multiplex RT-ddPCR assay, Hum. Genomics, 2024, 18(1), 48 CrossRef CAS PubMed.
- A. Gobbo, M. A. Fraiture, L. Van Poelvoorde, S. C. J. De Keersmaecker, C. Garcia-Graells and K. Van Hoorde, et al., Strategy to develop and validate digital droplet PCR methods for global antimicrobial resistance wastewater surveillance, Water Environ. Res., 2024, 96(11), e11145 CrossRef CAS PubMed.
- N. Rački, D. Morisset, I. Gutierrez-Aguirre and M. Ravnikar, One-step RT-droplet digital PCR: a breakthrough in the quantification of waterborne RNA viruses, Anal. Bioanal. Chem., 2014, 406(3), 661–667 CrossRef.
- M. Combe, E. Cherif, T. Deremarque, G. Rivera-Ingraham, F. Seck-Thiam and F. Justy, et al., Wastewater sequencing as a powerful tool to reveal SARS-CoV-2 variant introduction and spread in French Guiana, South America, Sci. Total Environ., 2024, 924, 171645 CrossRef CAS PubMed.
- R. Silvester, W. B. Perry, G. Webster, L. Rushton, A. Baldwin and D. A. Pass, et al., Metagenomics unveils the role of hospitals and wastewater treatment plants on the environmental burden of antibiotic resistance genes and opportunistic pathogens, Sci. Total Environ., 2025, 961, 178403 CrossRef CAS PubMed.
- J. Alhama, J. P. Maestre, M. Á. Martín and C. Michán, Monitoring COVID-19 through SARS-CoV-2 quantification in wastewater: progress, challenges and prospects, Microb. Biotechnol., 2022, 15(6), 1719–1728 CrossRef CAS PubMed.
- E. K. Hayes, M. T. Gouthro, J. J. LeBlanc and G. A. Gagnon, Simultaneous detection of SARS-CoV-2, influenza A, respiratory syncytial virus, and measles in wastewater by multiplex RT-qPCR, Sci. Total Environ., 2023, 889, 164261 CrossRef CAS PubMed.
- W. Ahmed, P. M. Bertsch, A. Bivins, K. Bibby, K. Farkas and A. Gathercole, et al., Comparison of virus concentration methods for the RT-qPCR-based recovery of murine hepatitis virus, a surrogate for SARS-CoV-2 from untreated wastewater, Sci. Total Environ., 2020, 739, 139960 CrossRef CAS PubMed.
- M. Shaffer, D. North and K. Bibby, Evaluating Nanotrap Microbiome Particles as A Wastewater Viral Concentration Method, Food Environ. Virol., 2025, 17(1), 10 CrossRef CAS PubMed.
- S. E. Philo, E. K. Keim, R. Swanstrom, A. Q. W. Ong, E. A. Burnor and A. L. Kossik, et al., A comparison of SARS-CoV-2 wastewater concentration methods for environmental surveillance, Sci. Total Environ., 2021, 760, 144215 CrossRef CAS PubMed.
- V. Fernández-Fernández, M. Ramil, R. Cela and I. Rodríguez, Solid-phase extraction and fractionation of multiclass pollutants from wastewater followed by liquid chromatography tandem-mass spectrometry analysis, Anal. Bioanal. Chem., 2022, 414(14), 4149–4165 CrossRef PubMed.
- A. L. Wang, M. Jiang, A. Nguyen, S. R. Kane, M. K. Borucki and R. S. Kantor, et al., Benchmarking Concentration and Extraction Methods for Wastewater-Based Surveillance of Eight Human Respiratory Viruses: Implications for Rapid Application to Novel Pathogens, Environ. Sci. Technol., 2025, 59(36), 19107–19118 CrossRef CAS PubMed.
- C. Diamanti, L. Nousis, P. Bozidis, M. Koureas, M. Kyritsi and G. Markozannes, et al., Wastewater Surveillance of SARS-CoV-2: A Comparison of Two Concentration Methods, Viruses, 2024, 16(9), 1398 CrossRef CAS PubMed.
- P. Liu, L. Guo, M. Cavallo, C. Cantrell, S. P. Hilton and A. Nguyen, et al., Comparison of Nanotrap® Microbiome A Particles, membrane filtration, and skim milk workflows for SARS-CoV-2 concentration in wastewater, Front. Microbiol., 2023, 14, 1215311 CrossRef PubMed.
- M. S. Angga, B. Malla, S. Raya, M. Kitajima and E. Haramoto, Optimization and performance evaluation of an automated filtration method for the recovery of SARS-CoV-2 and other viruses in wastewater, Sci. Total Environ., 2023, 882, 163487 CrossRef CAS PubMed.
- X. Zheng, Y. Deng, X. Xu, S. Li, Y. Zhang and J. Ding, et al., Comparison of virus concentration methods and RNA extraction methods for SARS-CoV-2 wastewater surveillance, Sci. Total Environ., 2022, 824, 153687 CrossRef CAS PubMed.
- H. J. Luo, X. B. Huang, H. L. Zhong, C. X. Ye, X. Tan and K. Zhou, et al., Epidemiological characteristics and phylogenic analysis of human respiratory syncytial virus in patients with respiratory infections during 2011–2016 in southern China, Int. J. Infect. Dis., 2020, 90, 5–17 CrossRef CAS PubMed.
- Y. Chen, D. Cui, S. Zheng, S. Yang, J. Tong and D. Yang, et al., Simultaneous Detection of Influenza A, Influenza B, and Respiratory Syncytial Viruses and Subtyping of Influenza A H3N2 Virus and H1N1 (2009) Virus by Multiplex Real-Time PCR, J. Clin. Microbiol., 2011, 49(4), 1653–1656 CrossRef CAS PubMed.
- H. Li, Seqtk [Internet], 2023, [cited 2023 Aug 2], Available from: https://github.com/lh3/seqtk Search PubMed.
- S. Chen, fastp 1.0: An ultra-fast all-round tool for FASTQ data quality control and preprocessing, iMeta, 2025, 4(5), e70078 CrossRef CAS PubMed.
- H. Li and R. Durbin, Fast and accurate short read alignment with Burrows-Wheeler transform, Bioinformatics, 2009, 25(14), 1754–1760 CrossRef CAS PubMed.
- H. Li, B. Handsaker, A. Wysoker, T. Fennell, J. Ruan and N. Homer, et al., The Sequence Alignment/Map format and SAMtools, Bioinformatics, 2009, 25(16), 2078–2079 CrossRef CAS PubMed.
- W. De Coster and R. Rademakers, NanoPack2: population-scale evaluation of long-read sequencing data, Bioinformatics, 2023, 39(5), btad311 CrossRef CAS PubMed.
- W. Shen, B. Sipos and L. Zhao, SeqKit2: A Swiss army knife for sequence and alignment processing, iMeta, 2024, 3(3), e191 CrossRef CAS PubMed.
- D. E. Wood, J. Lu and B. Langmead, Improved metagenomic analysis with Kraken 2, Genome Biol., 2019, 20(1), 257 CrossRef CAS PubMed.
- B. Bogaerts, J. Van Braekel, A. Van Uffelen, J. D'aes, M. Godfroid and T. Delcourt, et al., Galaxy @Sciensano: a comprehensive bioinformatics portal for genomics-based microbial typing, characterization, and outbreak detection, BMC Genomics, 2025, 26(1), 20 CrossRef PubMed.
- B. D. Ondov, N. H. Bergman and A. M. Phillippy, Interactive metagenomic visualization in a Web browser, BMC Bioinf., 2011, 12(1), 385 CrossRef PubMed.
- S. A. Sapula, J. J. Whittall, A. J. Pandopulos, C. Gerber and H. Venter, An optimized and robust PEG precipitation method for detection of SARS-CoV-2 in wastewater, Sci. Total Environ., 2021, 785, 147270 CrossRef CAS PubMed.
- M. T. Flood, N. D'Souza, J. B. Rose and T. G. Aw, Methods Evaluation for Rapid Concentration and Quantification of SARS-CoV-2 in Raw Wastewater Using Droplet Digital and Quantitative RT-PCR, Food Environ. Virol., 2021, 13(3), 303–315 CrossRef CAS PubMed.
- P. A. Barril, L. A. Pianciola, M. Mazzeo, M. J. Ousset, M. V. Jaureguiberry and M. Alessandrello, et al., Evaluation of viral concentration methods for SARS-CoV-2 recovery from wastewaters, Sci. Total Environ., 2021, 756, 144105 CrossRef CAS PubMed.
- S. Yang, Y. Jiao, Q. Dong, S. Li, C. Xu and Y. Liu, et al., Evaluating approach uncertainties of quantitative detection of SARS-CoV-2 in wastewater: Concentration, extraction and amplification, Sci. Total Environ., 2024, 951, 175285 CrossRef CAS PubMed.
- W. Ahmed, A. Bivins, S. L. Simpson, W. J. M. Smith, S. Metcalfe and B. McMinn, et al., Comparative analysis of rapid concentration methods for the recovery of SARS-CoV-2 and quantification of human enteric viruses and a sewage-associated marker gene in untreated wastewater, Sci. Total Environ., 2021, 799, 149386 CrossRef CAS PubMed.
- G. Scott, N. P. Evens, J. Porter, D. I. Walker, G. Scott and N. P. Evens, et al., The Impact of Viral Concentration Method on Quantification and Long Amplicon Nanopore Sequencing of SARS-CoV-2 and Noroviruses in Wastewater, Microorganisms, 2025, 13(2) CrossRef CAS PubMed Available from: https://www.mdpi.com/2076-2607/13/2/229.
- K. Farkas, C. Pellett, N. Alex-Sanders, M. T. P. Bridgman, A. Corbishley and J. M. S. Grimsley, et al., Comparative Assessment of Filtration- and Precipitation-Based Methods for the Concentration of SARS-CoV-2 and Other Viruses from Wastewater, Microbiol. Spectrum, 2022, 10(4), e01102–e01122 Search PubMed.
- D. Kaya, D. Niemeier, W. Ahmed and B. V. Kjellerup, Evaluation of multiple analytical methods for SARS-CoV-2 surveillance in wastewater samples, Sci. Total Environ., 2022, 808, 152033 CrossRef CAS PubMed.
- M. H. Jafferali, K. Khatami, M. Atasoy, M. Birgersson, C. Williams and Z. Cetecioglu, Benchmarking virus concentration methods for quantification of SARS-CoV-2 in raw wastewater, Sci. Total Environ., 2021, 755, 142939 CrossRef CAS PubMed.
- M. S. Fonseca, B. A. S. Machado, C. d. A. Rolo, K. V. S. Hodel, E. d. S. Almeida and J. B. de Andrade, Evaluation of SARS-CoV-2 concentrations in wastewater and river water samples, Case Stud. Chem. Environ. Eng., 2022, 6, 100214 CrossRef CAS PubMed.
- M. Jiang, A. L. W. Wang, N. A. Be, N. Mulakken, K. L. Nelson and R. S. Kantor, Evaluation of the Impact of Concentration and Extraction Methods on the Targeted Sequencing of Human Viruses from Wastewater, Environ. Sci. Technol., 2024, 58(19), 8239–8250 CrossRef CAS PubMed.
- N. Cheshomi, A. Alum, M. F. Smith, E. S. Lim, O. Conroy-Ben and M. Abbaszadegan, Viral concentration method biases in the detection of viral profiles in wastewater, Appl. Environ. Microbiol., 2024, 91(1), e01339-24 CrossRef PubMed.
- M. Thornton, G. Eder, F. Amman, A. Pantielieieva, J. Vierheilig and A. Bergthaler, Comparative wastewater virome analysis with different enrichment methods, Water Res., 2025, 285, 123985 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |