Comparative machine learning and deep learning frameworks for robust carcinogenicity prediction and activity cliff analysis
Arkaprava
Banerjee
 ,
Vinay
Kumar
,
Vinay
Kumar
 and
Kunal
Roy
and
Kunal
Roy
 *
*
Drug Theoretics and Cheminformatics Laboratory, Department of Pharmaceutical Technology, Jadavpur University, Kolkata 700 032, India. E-mail: kunal.roy@jadavpuruniversity.in; kunalroy_in@yahoo.com
First published on 8th January 2026
Abstract
Many industrial chemicals are recognized as carcinogenic to humans. In this study, we developed predictive models for binary carcinogenicity data in rats that are closely associated with human carcinogenicity. This study involves a range of feature-based and chemical language modeling approaches. After the training-test split and selection of essential structural and physicochemical descriptors, we developed a simple linear discriminant analysis model. Thereafter, we computed similarity- and error-based descriptors, pooled them with structural and physicochemical descriptors, and developed classification read-across structure–activity relationship (c-RASAR) models using a range of machine learning algorithms, including an artificial neural network (ANN). Additionally, the pooled feature matrix was used to compute two ARKA (arithmetic residuals in K-groups analysis) descriptors, and a simple logistic regression model was trained on the two-descriptor feature matrix. Moreover, we adopted the long short-term memory (LSTM) architecture to develop a model based on SMILES strings. The results suggested that the logistic regression RASAR-ARKA model was the best-performing, and it was subsequently used to predict external data efficiently, along with the ANN c-RASAR model. Moreover, the ARKA framework allowed us to identify activity cliffs and explain the reason for mispredictions. In addition to providing an efficient prediction framework, the structure–function analysis suggests that the presence of nitrogen atoms, including in hydrazine derivatives and nitrosamines, and greater branching are responsible for carcinogenicity, while increased molecular size reduces the carcinogenic potency.
Environmental significanceNumerous environmental chemicals pose carcinogenic risks to humans, yet widespread data gaps hinder effective risk assessment and prioritization. Experimental studies are expensive and involve ethical considerations; therefore, in this study, we developed predictive models that efficiently estimate the carcinogenicity of query chemicals. Additionally, the structural features responsible for eliciting carcinogenicity and activity cliffs were identified, thereby providing a clear understanding for chemists. With a reliable predictive system and knowledge of carcinogenic structural features, safer disposal and the development of safer alternatives may be pursued. |
Introduction
Increasing industrialization in modern times has led to the accumulation of a wide array of chemicals in the environment. These chemicals pose significant hazards to various flora and fauna, ultimately disrupting the entire biological ecosystem. Accumulation of such chemicals, especially in higher organisms such as humans, can have significant adverse effects, and they can target vital organs such as the heart, kidneys, liver, and brain.1 The most common adverse effect can be attributed to their carcinogenic potency, for which over 3 million cases are reported annually in the European Union (EU).2,3 This necessitates rapid and efficient gap-filling of data on the carcinogenic potential of these chemicals, which cannot be determined experimentally in human subjects due to time, cost, labor, and ethical constraints. New approach methodologies (NAMs) are therefore currently needed.The main objective of adopting NAMs is to bypass time-consuming experimental approaches and to use novel tools (e.g. in silico methodologies) to rapidly fill toxicity data gaps. This has been endorsed by regulatory bodies like the US Food and Drug Administration (FDA), whose recent press release, dated April 10, 2025, clarifies that the FDA is considering replacing animal testing with NAMs (https://www.fda.gov/news-events/press-announcements/fda-announces-plan-phase-out-animal-testing-requirement-monoclonal-antibodies-and-other-drugs). These progressive steps aim not only to reduce ethical considerations but also to cope with the increasing demand for data gap filling.
The quantitative structure–activity relationship (QSAR) technique is widely adopted in silico NAMs, aimed at efficiently predicting toxicity data gaps. This involves using a set of compounds with known activity–toxicity profiles and then training a mathematical model to identify the important structural and physicochemical features of the molecules.4 This model can then be used to predict unseen chemicals – a data-gap-filling process. Various internal and external validation metrics are used to estimate model robustness, goodness-of-fit, and external predictive performance.5 However, due to statistical constraints, this approach struggles to perform well on limited training data,6,7 which is why non-statistical approaches like read-across have gained significant attention in predictive toxicology.8
Read-across is a simple approach in which a target compound is evaluated by identifying its nearest neighbors within a set of source compounds with known experimental response data.9,10 Higher weights are often assigned to compounds comparatively closer to the target compound.10 This is a rather simpler approach with no statistical limitations that proves beneficial for predicting data-poor endpoints. Depending on the number of close source and target compounds involved, read-across approaches can be classified as one-to-one, one-to-many, many-to-one, or many-to-many, with numerous close source compounds generally leading to greater prediction reliability for the target compound(s).
Although read-across is an effective predictive tool, one key limitation of quantitative read-across is its lack of interpretability of the contributing features. To mitigate this, and to leverage the strengths of QSAR and read-across, similarity- and error-based information from closely related congeners is used to compute descriptors that are employed in a statistical/machine learning modeling framework.11,12 This approach is known as the quantitative/classification read-across structure–activity relationship (q-RASAR/c-RASAR) approach.11,13 The q-RASAR/c-RASAR models use a predefined structural and physicochemical feature space to compute compound similarity. Subsequently, a wide array of similarity- and error-based measures is computed and used as descriptors in RASAR models. In most previous studies, this approach has resulted in improved predictive performance for unseen compounds compared to the corresponding QSAR model, using the same amount of chemical and feature space.12
Dimensionality reduction, from a modeling perspective, is an approach that helps maintain the ratio of the number of training compounds to the number of descriptors by effectively encoding information from a larger number of features into a smaller number of variables. This is especially useful for small dataset modeling where there is a limited number of compounds, and warrants a wide feature space. In this context, supervised dimensionality reduction approaches such as arithmetic residuals in K-groups analysis (ARKA) have proven useful, especially for binary classification-based small dataset modeling.6 The basic idea is that they segregate the training set into active and inactive classes, and then identify the contributions of various descriptors to the active and inactive classes. Thereafter, the descriptors with greater influence in the active and inactive classes are aggregated into two distinct groups. This is followed by assigning weights to each descriptor in a group, and the ARKA descriptors are computed based on the weighted summation of the standardized descriptor matrix.6 This algorithm provides a foundation for a simple, reproducible supervised dimensionality reduction framework that can improve model performance despite the expected loss of chemical information.6
In the past, various research groups have worked on modeling carcinogenicity. The foundational contributions were made in the works of Benigni and their group,14–18 who have developed expert systems like Toxtree (https://toxtree.sourceforge.net/index.html) that provide a module for predicting various toxicological endpoints. Additionally, various research studies were also directed at identifying structural alerts for carcinogenicity.19–26 In the present work, we developed several models to predict the carcinogenicity of industrial chemicals. This involved not only using various machine learning and deep learning algorithms but also different cheminformatics approaches, as discussed above. Initially, we developed QSAR models using different machine learning algorithms. Subsequently, using the selected feature space, we computed RASAR descriptors, fused them with the selected QSAR descriptors, and developed various machine learning c-RASAR models. Moreover, the same feature space was used to develop an artificial neural network (ANN) RASAR model, thereby employing deep learning concepts within the c-RASAR methodology. Again, the same fused feature space was used to compute the ARKA descriptors, and RASAR-ARKA classification models were developed, as reported for the first time. Additionally, we adopted a SMILES-based “feature-independent” chemical language modeling (CLM) algorithm, the long short-term memory (LSTM) network, which can learn the required features from the tokenization of the SMILES strings.27–29 The best model was selected based on balanced performance across internal and external validation metrics, and it was used to predict a set of true external compounds.
Materials and methods
Dataset collection and preparation
The rodent carcinogenicity dataset employed in this study was obtained from the work of Fjodorova et al. (2010)30 and consisted of 805 curated chemical compounds (SI-1), including 421 carcinogenic (1) and 384 non-carcinogenic (0) ones, for classification-based modeling. All the structures were processed using MarvinView (v5.9.4), where aromatization was applied and explicit hydrogens were added to ensure consistency in structural representation.31 Aromatization is a standardization procedure that converts eligible ring systems into their aromatic form, thereby improving the accuracy and uniformity of molecular representations for subsequent descriptor calculation and model development.32 The preparation step ensured reproducibility and suitability of the dataset for predictive analysis.Descriptor calculation, pretreatment, and dataset division
In this section, a comprehensive descriptor set comprising 2D molecular descriptors and MACCS-166 fingerprints was calculated using the alvaDesc software (v3.0) tool.33 This combined approach enabled robust capture of key structural and physicochemical attributes of the compounds while ensuring high reproducibility in descriptor computation, a critical requirement for reliable prediction of true external datasets. Following descriptor calculation, a systematic descriptor-thinning strategy was applied to eliminate variables exhibiting high intercorrelation and low variance, thereby reducing redundancy and enhancing model interpretability. This pretreatment step was performed using the Pretreatment V-WSP v1.2 tool,34,35 which ensured the selection of informative and statistically relevant features. Subsequently, the curated dataset was split into training and test sets at a 75![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 25 ratio using the Dataset Division GUI v1.2 tool36,37 and modified k-medoids clustering implemented through the Modified k-Medoid tool v1.3,35,38 ensuring a representative and balanced partitioning of chemical space. Additionally, an alternative division at a 1
25 ratio using the Dataset Division GUI v1.2 tool36,37 and modified k-medoids clustering implemented through the Modified k-Medoid tool v1.3,35,38 ensuring a representative and balanced partitioning of chemical space. Additionally, an alternative division at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio was also employed.39 The training set was used for model development, while the test set was used for rigorous external validation.
1 ratio was also employed.39 The training set was used for model development, while the test set was used for rigorous external validation.
Selection of the most significant features and model development
In this investigation, a structured and multi-level feature selection strategy was employed to identify the most significant descriptors for reliable QSAR model development. Initially, a most discriminating feature algorithm was applied to screen important variables using the MDF_Identifier-v1.0 tool,40 and descriptors exhibiting an absolute difference value exceeding 0.103 were retained for further analysis. The selection of this 0.103 cutoff was carefully chosen to ensure optimal inclusion of descriptors with meaningful discriminatory power while minimizing the retention of redundant or weakly informative variables. Furthermore, a random forest-based selection approach41 was also employed, where descriptors with an importance value greater than 0.005 were considered statistically relevant and selected for continued evaluation. Finally, the important features obtained from both selection strategies were combined and subjected to SHAP (Shapley additive explanations) analysis42 to assess their individual contribution and influence on model predictions. Based on this integrative evaluation, the most informative set of 33 descriptors was finalized and employed for linear discriminant analysis (LDA)-based QSAR model development.43 This rigorous, hierarchical selection process ensured enhanced model interpretability, improved predictive accuracy, and robust generalizability for the developed QSAR framework.Statistical validation of the LDA model
The LDA QSAR model was developed using the structural and physicochemical descriptors previously identified as the most discriminative. Before model development, descriptor values for both training and test compounds were standardized using the Scale v1.0 tool,44 with scaling based on the mean and standard deviation of the training set descriptors. This step ensured consistency and minimized bias arising from differences in descriptor magnitude, thereby improving model reliability. To achieve robust and statistically sound validation, the LDA model was rigorously evaluated using a comprehensive set of classification-based internal and external performance metrics. These included accuracy, balanced accuracy, specificity, precision, recall, F1-score, Matthews correlation coefficient (MCC), Cohen's κ, and the area under the receiver operating characteristic curve (AUC).39 Collectively, these metrics provided a detailed assessment of model predictability, stability, and overall classification performance.ML-based c-RASAR modeling
In this step, to implement c-RASAR modeling, the RASAR-Desc-Calc-v3.0.3 tool13 was used to calculate RASAR descriptors, with the QSAR training and test set feature matrices as inputs. The default settings of the read-across hyperparameters (Gaussian kernel similarity, σ = 1, and number of close source compounds = 10) associated with the Read-Across-v4.2.2 tool were used to compute RASAR descriptors.10,11 The 15 similarity and error-based RASAR descriptors, thus computed for classification modeling, were then merged with the selected 33 QSAR descriptors to obtain a complete feature pool of 48 descriptors. Thereafter, the training and test sets, with the combined feature pool, were used to develop an array of different machine learning (ML) models, namely random forest classifier,45 support vector classifier,46 linear discriminant analysis,47 logistic regression,48 AdaBoost classifier,49 gradient boosting classifier,50 extreme gradient boosting classifier,51 extreme gradient boosting random forest classifier,51 linear support vector classifier,46 k-nearest neighbors classifier,52 quadratic discriminant analysis,53 Gaussian Naïve Bayes classifier,54 Gaussian process classifier,55 and CatBoost classifier56 models, and also an artificial neural network (ANN) model,57 after standardizing the training and test set feature matrices. While the ML models were developed using the scikit-learn, xgboost, and CatBoost libraries with default hyperparameter settings, the ANN model was developed using the TensorFlow library, and the hyperparameters were optimized using Random Search from Keras-Tuner. All the models were validated internally and externally using a range of validation metrics as stated above.Computation of the ARKA descriptors and development of a RASAR-ARKA model
The arithmetic residuals in K-groups analysis, a.k.a. ARKA, is a supervised dimensionality reduction approach that has been shown to enhance the performance of classification-based models.6,58 In this study, we used the combined 48-descriptor pool to compute two ARKA descriptors (ARKAdesc-v2.1, available at https://sites.google.com/jadavpuruniversity.in/dtc-lab-software/arithmetic-residuals-in-k-groups-analysis-arka) using a supervised dimensionality reduction workflow. This two-descriptor “RASAR-ARKA” model was then used to train a simple logistic regression model,48 which was also validated internally and externally. In a previous publication,3 the RASAR descriptors were computed on a hybrid feature matrix of QSAR and ARKA descriptors, thereby generating “ARKA-RASAR” models; however, the present work generates ARKA descriptors from the hybrid feature space of QSAR and RASAR descriptors, which generated “RASAR-ARKA” models.Development of a long short-term memory network (LSTM) model – a chemical language modeling approach
In addition to the different feature-based modeling approaches (QSAR, c-RASAR, ARKA) employed, we explored the ability of language models like LSTM to generate predictions.59 This model takes the SMILES strings as inputs, performs a tokenization step, automatically extracts relevant features, and develops predictive models. As language models are known to be “data hungry”, the training set data points were oversampled. The LSTM model was trained using the TensorFlow library, and the model was validated using the set of internal and external validation metrics mentioned previously.True external set prediction
To assess the true external predictive performance of the developed model, we conducted a true external validation. A true external dataset with known human carcinogenic status was sourced from Toma et al.60 After data curation and removal of common compounds present in both our training and test sets, we prepared a curated list of 505 “unseen” compounds. This set was subjected to prediction using the best-performing models, and the corresponding validation metrics were computed to estimate its true generalizability.The complete workflow is presented in Fig. 1.
Results and discussion
This section presents the results obtained from the different modeling approaches and explains the important structural and physicochemical components that are responsible for the carcinogenic potency.Statistical results of the LDA QSAR model (feature space containing structural and physicochemical information)
The success of QSAR model development depends on the efficient selection of essential features during training. In this context, after dataset splitting, we adopted a multi-level feature selection strategy. This involved a model-agnostic most discriminating feature selection strategy, where the important descriptors are selected based on their discriminatory ability between the positive and negative classes. Additionally, the important features obtained from the tree-based random forest variable importance analysis were selected. Finally, the lists of important features obtained from both approaches were pooled, and the combined set was subjected to SHAP analysis using four modeling algorithms: linear discriminant analysis, support vector machine, logistic regression, and random forest. The final list of 33 important descriptors was then selected based on their importance in the SHAP plots. Thereafter, the training and test sets, with the selected features, were standardized, and a simple LDA QSAR model was developed. Table 1 shows the internal and external validation statistics of the LDA QSAR model. The QSAR training and test sets have also been made available in the (SI-1). Additionally, the list of 33 QSAR descriptors, along with their description, has been presented in Table S1 of the (SI-2).| Model | Type | Desc. | Acc. | Prec. | F-mea. | Ckappa | MCC | Rec. | Spec. | BA | AUC |
|---|---|---|---|---|---|---|---|---|---|---|---|
| a Desc = descriptors, Acc = accuracy, Prec = precision, F-mea = F-measure, Ckappa = Cohen's k, Rec = recall, Spec = specificity, BA = balanced accuracy. | |||||||||||
| Training set | |||||||||||
| LDA | QSAR | 33 | 0.682 | 0.693 | 0.698 | 0.361 | 0.361 | 0.703 | 0.657 | 0.680 | 0.740 |
| RF | c-RASAR | 48 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 |
| LSTM | CLM | SMILES | 0.977 | 0.966 | 0.978 | 0.953 | 0.954 | 0.991 | 0.962 | 0.976 | 0.998 |
| ANN | c-RASAR | 48 | 0.678 | 0.699 | 0.687 | 0.356 | 0.356 | 0.675 | 0.682 | 0.678 | 0.750 |
| LR | RASAR-ARKA | 2 | 0.650 | 0.672 | 0.659 | 0.300 | 0.300 | 0.647 | 0.654 | 0.650 | 0.707 |
| (CP-ANN)30 | QSAR | 8 MDL | 0.910 | — | — | — | — | 0.960 | 0.860 | — | — |
| (CP-ANN)30 | QSAR | 12 dragon | 0.890 | — | — | — | — | 0.900 | 0.870 | — | — |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||||||||
| Test set | |||||||||||
| LDA | QSAR | 33 | 0.653 | 0.658 | 0.679 | 0.303 | 0.304 | 0.702 | 0.600 | 0.651 | 0.735 |
| RF | c-RASAR | 48 | 0.729 | 0.745 | 0.738 | 0.457 | 0.457 | 0.731 | 0.726 | 0.729 | 0.784 |
| LSTM | CLM | SMILES | 0.583 | 0.602 | 0.599 | 0.164 | 0.165 | 0.596 | 0.568 | 0.582 | 0.621 |
| ANN | c-RASAR | 48 | 0.694 | 0.687 | 0.722 | 0.383 | 0.385 | 0.760 | 0.621 | 0.690 | 0.763 |
| LR | RASAR-ARKA | 2 | 0.719 | 0.718 | 0.738 | 0.434 | 0.435 | 0.760 | 0.674 | 0.717 | 0.765 |
| (CP-ANN)30 | QSAR | 8 MDL | 0.730 | — | — | — | — | 0.750 | 0.690 | — | — |
| (CP-ANN)30 | QSAR | 12 dragon | 0.690 | — | — | — | — | 0.750 | 0.610 | — | — |
Statistical results of the RF and ANN c-RASAR model (feature space containing structural, physicochemical, and similarity-based information)
c-RASAR models encode similarity and error-based information from the read-across hypothesis, which often results in models with enhanced predictivity. In this context, we have computed 15 similarity and error-based RASAR descriptors (Table S1) on the selected QSAR feature pool. These descriptors were then merged with the 33 selected QSAR descriptors to obtain a complete 48 descriptor feature pool. Thereafter, the training and test sets were standardized, and various ML modeling algorithms were employed. The random forest c-RASAR model appeared to be the best-performing model, whose internal and external validation statistics are reported in Table 1. Additionally, we also trained an ANN c-RASAR model, and the corresponding model statistics have also been reported in Table 1. As evident from the model statistics, the ANN c-RASAR model provides balanced performance for the training and test data, whereas the random forest c-RASAR model appears to be overfitted, although generating the highest accuracy for the test set compounds.Statistical results of the LR RASAR-ARKA model (a supervised dimensionality reduced feature space containing structural, physicochemical, and similarity-derived information)
The recent application of the supervised dimensionality reduction framework, the ARKA approach,6 has been shown to improve model statistics, even though dimensionality reduction is associated with the loss of chemical information. In this study, we computed two ARKA descriptors, considering the combined 48 descriptor pool (QSAR + RASAR). The training and test sets, now consisting of just two ARKA descriptors, were then standardized and used to train a logistic regression RASAR-ARKA model. Table 1 shows the various internal and external validation metrics of the LR RASAR-ARKA model.Statistical results of the LSTM deep learning model (a chemical language modeling algorithm)
As the three different previously described modeling approaches are based on some selected features, we decided to explore the performance of models that do not require a set of input features. In this context, we adopted a chemical language modeling algorithm – the long short-term memory network (LSTM) – to check the predictive ability. Table 1 presents the internal and external validation statistics for the developed LSTM model.Discussion of the modeling results and identification of the best model
This analysis is based on the results of the different modeling approaches presented in Table 1. We adopted three different feature-based approaches, namely the QSAR, c-RASAR, and ARKA methods, and developed four different models. Additionally, with the growing acceptance and applications of CLMs, we developed an input feature space-independent LSTM model. As evident from the results, the RF c-RASAR model appears to be overfitted, despite the fact that it generated the highest predictive accuracy. Additionally, the LDA QSAR and ANN c-RASAR models, although having a good balanced performance on the training and test sets, do not generate a sufficiently high predictive accuracy. The language model (LSTM) also appears to be overfitted, with a sub-optimal predictive accuracy. Therefore, considering the external predictive ability, robustness, algorithmic simplicity, statistical compliance, and the ease of reproducibility, the feature-based LR RASAR-ARKA appears to be the best-performing model. Moreover, it is not always the case that the models restricted to a fixed feature space underperform in comparison to language models that can extract features of their own.29 The performance of the optimal model primarily depends on the number of available data points and the important features, and such cases in which the feature-based models outperform CLMs highlight the effectiveness and importance of the multi-level feature selection strategy that we employed.Applicability domain analysis
An analysis of the applicability domain is crucial in a QSAR study. For this, we used the leverage approach61 to identify structural outliers using the freely available Hi_Calculator-v2.0 tool (https://sites.google.com/jadavpuruniversity.in/dtc-lab-software/home), where the QSAR training and test set descriptor matrices were used as inputs. This approach identified fourteen (14) structural outliers from the training set (2.3%) and six (6) “outside AD” compounds from the test set (3%). The compounds, along with their leverage values, AD status, and the threshold leverage, are presented in the (SI-1). This limited number of outliers suggests that our model encodes a wide chemical space.Identification of the essential structural and physicochemical features
This is an important aspect that helps non-cheminformatic researchers understand the structural and physicochemical properties of molecules associated with carcinogenicity. As per the Organisation for Economic Co-operation and Development (OECD) Principle 5, the model should be supported by a mechanistic interpretation if possible.5 Therefore, to understand the relative contributions of structural features, we developed SHAP62 plots for the training and test sets, using the selected QSAR descriptors as the input feature matrix. This enabled us to view the important features and their directionality of contribution towards carcinogenicity. Fig. 2 represents the SHAP analysis plots for the training and test sets for the top 20 descriptors, where the important descriptors appear at the top of the plots.From the SHAP plots, it is clear that the descriptor B04[C–N] exhibits the highest importance in both the training and test sets. This descriptor depicts the presence or absence of C and N atoms at a topological distance 4, thereby signifying the presence of nitrogen atoms. As evident from Fig. 2, this descriptor has a positive contribution towards carcinogenicity, and this can be explained by the fact that nitrogen-containing compounds tend to show carcinogenic properties. This can be exemplified in compounds like salbutamol (#698) and o-nitroanisole (#517), which contain a secondary amino group and a nitro group, respectively, thus showing carcinogenicity. Similarly, the descriptor B01[N–N] depicts the presence or absence of two nitrogen atoms at a topological distance 1, and this descriptor contributes positively towards carcinogenicity. Hydrazine-derived molecules are known to possess carcinogenic and neurotoxic potential by interacting with DNA and eliciting DNA damage.63 This can be exemplified by hydrazines like isoniazid (#403) and hydrazones like acetone[4-(5-nitro-2-furyl)-2-thiazolyl] hydrazone (#5), which show the presence of N–N at a topological distance of 1 and are therefore carcinogenic. Additionally, nitrosamines are well-known carcinogens that cause genotoxicity in humans.64,65 Their importance is also evident from the various available cheminformatic models that focus exclusively on nitrosamines eliciting carcinogenicity.66,67 This can be observed in nitrosamine compounds like diallylnitrosamine (#205), which shows carcinogenicity. The descriptor DBI represents the branching index of molecules and contributes positively to carcinogenicity. The degree of branching in a molecule is related to the steric hindrance towards electrophilic or nucleophilic attack, thus increasing the stability of the molecule and preventing its hydrolysis. This can be exemplified in branched compounds like actinomycin D (#22), which has a higher DBI value and possesses carcinogenic properties. Similarly, piperidine (#657), which does not possess branching, does not exhibit carcinogenicity. The descriptor J_D represents the Balaban J index from the topological distance matrix, and this descriptor contributes negatively towards carcinogenicity. This descriptor is computed considering the number of edges in a molecule and the vertex-distance degrees of adjacent vertices.5 Therefore, a higher J_D value signifies a higher number of edges, which actually indicates an increase in molecular weight and size. As explained by Zhong et al.,68 molecules possessing higher molecular weights fail to get absorbed in significant amounts, and therefore chemicals having high molecular weight and size possess insignificant carcinogenic risk. This can be exemplified by compounds like Malathion (#419), which has a higher J_D value and is non-carcinogenic, whereas compounds like formaldehyde (#333) have a lower J_D value and are carcinogenic.
Explaining mispredictions – analysis of activity cliffs (ACs)
ACs are pairs or groups of compounds that are structurally very similar, but differ significantly in their potency.69 Such compounds violate the similarity principle, which serves as the basis for cheminformatic model development. In this section, we aim to identify and analyze activity cliffs that can misdirect the modeling algorithm and explain why they lead to mispredictions. In this context, the ARKA framework not only provides a supervised dimensionality framework for developing improved models but also efficiently identifies ACs. As per the theory of ARKA, descriptors possessing higher mean values in the active class are grouped in ARKA_1, while the other descriptors showing higher mean values in the inactive class are grouped in ARKA_2.6 As a result, it is expected that active compounds should have a positive ARKA_1 and a negative ARKA_2 value, while inactive compounds should have a negative ARKA_1 and a positive ARKA_2 value. Compounds exhibiting the opposite characteristics, i.e. active compounds having negative ARKA_1 and positive ARKA_2 values or inactive compounds having positive ARKA_1 and negative ARKA_2 values, can be termed ACs.6Fig. 3 depicts the ARKA_2 v/s ARKA_1 plots for the training and test sets, identifying the potential ACs/prediction cliffs. | ||
| Fig. 3 . ARKA_2 v/s ARKA_1 plots for (a) the training set and (b) the test set, identifying ACs and prediction cliffs. | ||
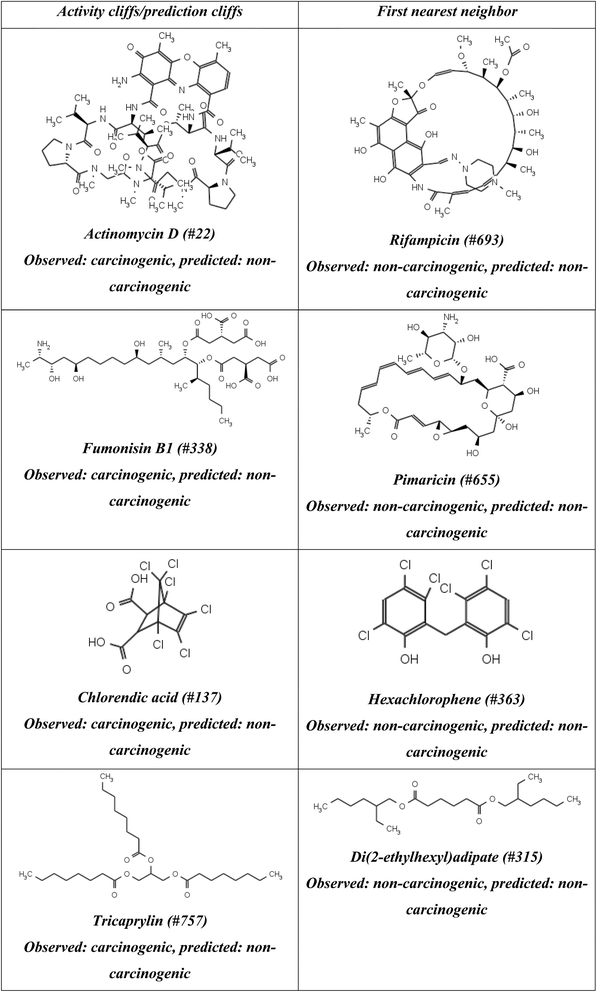
In Fig. 3, we identified the most significant activity cliffs (based on their Euclidean distance from the origin of the ARKA_2 v/s ARKA_1 plots). As evident from Fig. 3a, the compounds actinomycin D and fumonisin b1, although possessing carcinogenic properties experimentally, show similarity to non-carcinogenic compounds, and are therefore located in the second quadrant. If we examine the nearest neighbors of actinomycin D, they appear structurally very different from the training set representatives, with low similarity to them. Additionally, the first nearest neighbor of actinomycin D (a carcinogen) is Rifampicin (#693) – a non-carcinogen (Table 2). Moreover, among the ten nearest neighbors of actinomycin D, only two are carcinogens, while all the other compounds are non-carcinogens. This explains the AC nature of actinomycin D. A similar observation is seen with fumonisin B1, which also shows AC behavior. Among the test set compounds, chlorendic acid and tricaprylin are the only prediction cliffs (PCs). In the case of chlorendic acid, a carcinogen, its first nearest neighbor is hexachlorophene (#363), which is non-carcinogenic (Table 2). Additionally, like the previous cases, chlorendic acid has very low levels of structural similarity with the test set compounds. However, as far as the ten nearest neighbors are concerned, there is an equal distribution of carcinogens and non-carcinogens. These observations collectively suggest that, because the test set lacks similar structures, chlorendic acid considers non-carcinogenic compounds to be the most similar, which explains its PC nature. A similar observation can also be seen in the case of tricaprylin, a carcinogen, whose first nearest neighbor is a non-carcinogen, which has low levels of similarity to the training set compounds, and has a higher number of non-carcinogenic compounds among the ten nearest neighbors. These observations also collectively infer that tricaprylin is a PC. It is interesting to note that all these ACs and PCs have been misclassified as non-carcinogens by the LR RASAR-ARKA model, while their first nearest neighbors, bearing the opposite class labels, have been correctly classified as non-carcinogens (Table 2). These examples also demonstrate how the ARKA framework can be used to explain model mispredictions and identify activity/prediction cliffs.
True external set predictions – a test to check the model's generalizability
As shown in Table 1, the final model (LR RASAR-ARKA) demonstrated balanced performance for the training and test set compounds. However, to further evaluate the model's generalizability, we performed true external set predictions. A true external set of human carcinogenicity data was sourced from the works of Toma et al.60 Curation of chemical structures yielded 505 unique compounds that are not members of either our training or test sets. The predictive accuracy of our LR RASAR-ARKA model on the true external data was 0.574, which is acceptable, considering the fact that the models were developed using rat data while the true external set is human data. Additionally, we used the ANN c-RASAR model for prediction, which resulted in a predictive accuracy of 0.618. To compare the predictive accuracies of our models with established benchmark expert systems, we additionally generated predictions using TOPKAT, available in Biovia Discovery Studio (https://www.3dsbiovia.com/products/collaborative-science/biovia-discovery-studio/), and CAESAR predictions in VEGA,70 and the results are presented in Table 3.| Models/expert systems | Trained on | True external set | Prediction accuracy | Notes |
|---|---|---|---|---|
| a The model that predicts the true external compounds most accurately. | ||||
| LR-RASAR-ARKA (present work) | Rat data | Human data | 0.574 | Two-descriptor model, satisfactory predictivity |
| ANN c-RASAR (present work)a | Rat data | Human data | 0.618 | c-RASAR model with the best predictivity |
| TOPKAT | Rodent data | Human data | 0.598 | Inferior predictive accuracy |
| VEGA (CAESAR)70 | Rat data | Human data | 0.612 | Comparable predictive accuracy with our ANN c-RASAR model |
From Table 3, it is clear that the model with the highest predictive accuracy is the ANN c-RASAR model. This model provided superior true external set performance compared to the TOPKAT expert system. Additionally, it could not be confirmed that our “true external” compounds were genuinely external to the expert system, as the training set information is unknown. Comparing the results of our ANN c-RASAR model with the CAESAR predictions in VEGA, our model had a slightly superior predictive accuracy. Moreover, the two-descriptor LR-RASAR-ARKA model showed competitive predictive accuracy. In view of the above observations, it can be concluded that our models can efficiently be used to predict human carcinogenicity. The list of true external compounds, along with their prediction status, is presented in the (SI-1).
Additionally, using the ARKA approach, we identified 21 prediction cliffs from the list of true external compounds (SI-1). As expected, all these compounds were incorrectly predicted by both the LR RASAR-ARKA and ANN c-RASAR models. Out of the 21 compounds, 20 were carcinogens that were predicted as non-carcinogens, while one was a non-carcinogen that was predicted as a carcinogen. This clear explanation of mispredictions can only be made possible by combining the RASAR and ARKA frameworks.
Comparison with the previously reported models
This section compares the developed model with the previously reported models. The previous work of Fjodorova et al.30 involved developing CP-ANN models on two different feature spaces – 8 MDL descriptors and 12 Dragon descriptors. As evident from their model statistics (also presented here in Table 1), their models suffered from significant overfitting, with a large difference between the training and test set statistics. Additionally, the optimum number of epochs, one of the main hyperparameters of an ANN model, was selected based on the highest test set accuracy.30 This appears to be an incorrect approach considering the theories of machine and deep learning, where the hyperparameters are tuned using only the training set data points. Therefore, their test set compounds are not “unseen” to the model. Conversely, our simple logistic regression RASAR-ARKA model achieves balanced performance on both the training and test sets, and we also report additional important validation metrics that further help assess the class-specific predictive ability and the overall predictive capacity of the model. It is to be noted that our random forest and LSTM models showed superior statistics for the training compounds; however, we have not considered them as our best models, despite the fact that the RF model showed the highest predictive accuracy, because those models suffered from overfitting, like the previously reported models. Moreover, our final model (LR RASAR-ARKA) was built on just two modeling descriptors, while the previous work utilized 8 and 12 descriptors. Comparing the external predictive ability, our model's accuracy is almost the same as that of the best model from Fjodorova et al.,30 whose test set apparently did not strictly serve as “unseen” to the model. Comparing the sensitivity/recall values for the external set, our model showed slightly higher recall, suggesting it can more efficiently identify carcinogenic compounds. Additionally, we have identified ACs and PCs in the dataset and explained the reasons for their mispredictions, which is absent in previous work.Novelty of the present work
Conclusion
The carcinogenicity of molecules is a growing concern, especially given the current bioaccumulation of large amounts of industrial chemicals in ecosystems. These chemicals affect a wide array of living organisms by integrating into food chains. Exposure of humans to such carcinogens affects a wide array of different vital organs, thus interfering with their survival. As experimental determination of carcinogenicity in humans involves a lot of time and strict ethical considerations, predictive modeling can serve as a useful tool to identify carcinogens. In this work, we developed multiple machine learning and deep learning models aimed at developing a robust, predictive, and reliable framework for predicting carcinogenicity. We considered some of the feature-based approaches like QSAR, c-RASAR, and ARKA, and also adopted chemical language models like LSTM. While the QSAR model was trained using simple linear discriminant analysis, the c-RASAR models consisted of a machine learning random forest algorithm and an artificial neural network. The supervised dimensionally reduced ARKA descriptors were computed on the hybrid feature space of the QSAR and c-RASAR descriptors, and a simple logistic regression model was trained. Canonical SMILES strings served as the inputs for the LSTM model, which can automatically extract essential features. The modeling results indicated that although the random forest c-RASAR model had the highest external predictive accuracy, it was overfitted. Conversely, the LR RASAR-ARKA model not only had a significantly high predictive performance but also showed balanced performance for both the training and test sets and was therefore identified as the best model. This model was then used to predict a true external dataset with satisfactory predictive accuracy. Additionally, the ANN c-RASAR model showed the highest true external predictivity in comparison to some of the widely used benchmark predictive tools. Structure–activity relationship (SAR) studies identified that the presence of nitrogen atoms, especially in hydrazine derivatives and nitrosamines, is responsible for the carcinogenic properties. Additionally, a greater degree of branching was identified to be associated with carcinogenicity. Moreover, it was identified that small molecules are potent carcinogens, as larger molecules are less readily absorbed into target tissues. Finally, this study shows that feature-based approaches can potentially exceed the predictive ability of recent state-of-the-art CLM algorithms when an efficient feature selection strategy is used. The only limitation of the RASAR-ARKA approach can be the lack of direct interpretability of descriptors. However, the multiclass-ARKA approach provides response range–specific interpretability that cannot be achieved using a standard QSAR model.Conflicts of interest
None declared.Data availability
The data associated with this work has been provided in supplementary information (SI). Supplementary information: SI-1 contains the data set for modeling and external data set for predictions. SI-2 lists the definition of different molecular and RASAR descriptors. SI-3 contains the code for the LR RASAR-ARKA model. See DOI: https://doi.org/10.1039/d5em01001b.Acknowledgements
The authors thank LSRB (DRDO) for a q-RASAR project.References
- D. O. Carpenter, Human health effects of environmental pollutants: new insights, Environ. Monit. Assess., 1998, 53, 245–258, DOI:10.1023/A:1006013831576.
- F. Madia, A. Worth, R. Corvi, Analysis of Carcinogenicity Testing for Regulatory Purposes in the European Union. EUR 27765, Publications Office of the European Union, Luxembourg (Luxembourg), 2016, JRC100609, DOI:10.2788/547846.
- A. Banerjee and K. Roy, A new approach methodology (NAM) for carcinogenicity prediction of organic chemicals using the multiclass ARKA framework and machine-learning-based stacking regression, J. Hazard. Mater., 2025, 496, 139302, DOI:10.1016/j.jhazmat.2025.139302.
- C. Hansch and T. Fujita, ρ-σ-π analysis. a method for the correlation of biological activity and chemical structure, J. Am. Chem. Soc., 1964, 86, 1616–1626, DOI:10.1021/ja01062a035.
- K. Roy, S. Kar, R. N. Das, Understanding the Basics of QSAR for Applications in Pharmaceutical Sciences and Risk Assessment, Academic Press, NY, 2015, DOI:10.1016/C2014-0-00286-9.
- A. Banerjee and K. Roy, ARKA: a framework of dimensionality reduction for machine-learning classification modeling, risk assessment, and data gap-filling of sparse environmental toxicity data, Environ. Sci.: Processes Impacts, 2024, 26, 991–1007, 10.1039/D4EM00173G.
- A. Gajewicz, What if the number of nanotoxicity data is too small for developing predictive Nano-QSAR models? An alternative read-across based approach for filling data gaps, Nanoscale, 2017, 9, 8435–8448, 10.1039/C7NR02211E.
- G. Patlewicz, M. T. D. Cronin, G. Helman, J. C. Lambert, L. E. Lizarraga and I. Shah, Navigating through the minefield of read-across frameworks: A commentary perspective, Comput. Toxicol., 2018, 6, 39–54, DOI:10.1016/j.comtox.2018.04.002.
- S. Manganelli, E. Benfenati, Use of Read-Across Tools, in Silico Methods for Predicting Drug Toxicity. Methods in Molecular Biology, ed Benfenati, E., Humana Press, New York, NY, 2016, 1425, DOI:10.1007/978-1-4939-3609-0_13.
- M. Chatterjee, A. Banerjee, P. De, A. Gajewicz-Skretna and K. Roy, A novel quantitative read-across tool designed purposefully to fill the existing gaps in nanosafety data, Environ. Sci.: Nano, 2022, 9, 189–203, 10.1039/D1EN00725D.
- A. Banerjee and K. Roy, First report of q-RASAR modeling toward an approach of easy interpretability and efficient transferability, Mol. Diversity, 2022, 26, 2847–2862, DOI:10.1007/s11030-022-10478-6.
- K. Roy, A. Banerjee, Q-RASAR. A Path to Predictive Cheminformatics, Springer, NY, 2024, DOI:10.1007/978-3-031-52057-0.
- A. Banerjee and K. Roy, Prediction-inspired intelligent training for the development of classification read-across structure–activity relationship (c-RASAR) models for organic skin sensitizers: assessment of classification error rate from novel similarity coefficients, Chem. Res. Toxicol., 2023, 36, 1518–1531, DOI:10.1021/acs.chemrestox.3c00155.
- R. Benigni and C. Bossa, Mechanisms of Chemical Carcinogenicity and Mutagenicity: A Review with Implications for Predictive Toxicology, Chem. Rev., 2011, 111, 2507–2536, DOI:10.1021/cr100222q.
- R. Benigni, Predictive toxicology today: the transition from biological knowledge to practicable models, Expert Opin. Drug Metab. Toxicol., 2016, 12, 989–992, DOI:10.1080/17425255.2016.1206889.
- B. Schilter, R. Benigni, A. Boobis, A. Chiodini, A. Cockburn, M. T. D. Cronin, E. L. Piparo, S. Modi, A. Thiel and A. Worth, Establishing the level of safety concern for chemicals in food without the need for toxicity testing. Regulat, Toxicol. Pharmacol., 2014, 68, 275–296, DOI:10.1016/j.yrtph.2013.08.018.
- G. J. Myatt, E. Ahlberg, Y. Akahori, D. Allen, A. Amberg, L. T. Anger, A. Aptula, S. Auerbach, L. Beilke, P. Bellion, R. Benigni, J. Bercu, E. D. Booth, D. Bower, A. Brigo, N. Burden, Z. Cammerer, M. T. D. Cronin, K. P. Cross and L. Custer, et al., In silico toxicology protocols. Regulat, Toxicol. Pharmacol., 2018, 96, 1–17, DOI:10.1016/j.yrtph.2018.04.014.
- R. R. Tice, A. Bassan, A. Amberg, L. T. Anger and M. A. Beal, et al., In silico approaches in carcinogenicity hazard assessment: Current status and future needs, Comput. Toxicol., 2021, 20, 100191, DOI:10.1016/j.comtox.2021.100191.
- A. M. Helguera, M. A. C. Perez, M. P. Gonzalez, R. M. Ruiz and H. G. Diaz, A topological substructural approach applied to the computational prediction of rodent carcinogenicity, Bioorg. Med. Chem., 2005, 13, 2477–2488, DOI:10.1016/j.bmc.2005.01.035.
- F. Li, T. Fan, G. Sun, L. Zhao, R. Zhong and Y. Peng, Systematic QSAR and iQCCR modelling of fused/non-fused aromatic hydrocarbons (FNFAHs) carcinogenicity to rodents: reducing unnecessary chemical synthesis and animal testing, Green Chem., 2022, 24, 5304–5319, 10.1039/D2GC00986B.
- A. M. Helguera, M. P. Gonzalez, M. N. D. S. Cordeiro and M. A. C. Perez, Quantitative structure carcinogenicity relationship for detecting structural alerts in nitroso-compounds, Toxicol. Appl. Pharmacol., 2007, 221, 189–202, DOI:10.1016/j.taap.2007.02.021.
- A. M. Helguera, M. P. Gonzalez, M. N. D. S. Cordeiro and M. A. C. Perez, Quantitative structure−carcinogenicity relationship for detecting structural alerts in nitroso compounds: species, rat; sex, female; route of administration, gavage, Chem. Res. Toxicol., 2008, 21, 633–642, DOI:10.1021/tx700336n.
- A. M. Helguera, G. Perez-Machado, M. N. D. S. Cordeiro and R. D. Combes, Quantitative structure-activity relationship modelling of the carcinogenic risk of nitroso compounds using regression analysis and the TOPS-MODE approach, SAR QSAR Environ. Res., 2010, 21, 277–304, DOI:10.1080/10629361003773930.
- X. Wu, Q. Zhang, H. Wang and J. Hu, Predicting carcinogenicity of organic compounds based on CPDB, Chemosphere, 2015, 139, 81–90, DOI:10.1016/j.chemosphere.2015.05.056.
- H. Zhang, Z.-X. Cao, M. Li, Y.-Z. Li and C. Peng, Novel naïve Bayes classification models for predicting the carcinogenicity of chemicals, Food Chem. Toxicol., 2016, 97, 141–149, DOI:10.1016/j.fct.2016.09.005.
- A. P. Toropova and A. A. Toropov, CORAL: QSAR models for carcinogenicity of organic compounds for male and female rats, Comput. Biol. Chem., 2018, 72, 26–32, DOI:10.1016/j.compbiolchem.2017.12.012.
- F. Grisoni, G. Schneider, De Novo Molecular Design with Chemical Language Models, in Artificial Intelligence in Drug Design. Methods in Molecular Biology, ed Heifetz, A.. Humana, New York, NY, 2022, vol. 2390, DOI:10.1007/978-1-0716-1787-8_9.
- S. Hochreiter and J. Schmidhuber, Long short-term memory, Neural Comput., 1997, 9, 1735–1780, DOI:10.1162/neco.1997.9.8.1735.
- A. Banerjee, S. Kar, K. Roy, G. Patlewicz, I. Shah, P. G. Karamertzanis, G. Gini and E. Benfenati, From Feature-Based Chemical Similarity to Chemical Language Models—A Paradigm Shift in Computer-Aided Molecular Design and Property Predictions, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2025, 15, e70057, DOI:10.1002/wcms.70057.
- N. Fjodorova, M. Vračko, M. Novič, A. Roncaglioni and E. Benfenati, New public QSAR model for carcinogenicity, Chem. Cent. J., 2010, 4, S3, DOI:10.1186/1752-153X-4-S1-S3.
- Chemaxon: MarvinView Software Version 5.9.4. Copyright Ⓒ 1998–2025, Chemaxon Ltd, https://chemaxon.com Search PubMed.
- M. González-Medina, J. J. Naveja, N. Sánchez-Cruz and J. L. Medina-Franco, Open chemoinformatic resources to explore the structure, properties and chemical space of molecules, RSC Adv., 2017, 7, 54153–54163, 10.1039/C7RA11831G.
- A. Mauri alvaDesc: A Tool to Calculate and Analyze Molecular Descriptors and Fingerprints, in Ecotoxicological QSARs. Methods in Pharmacology and Toxicology, ed Roy. K., Humana, New York, NY, 2020, DOI:10.1007/978-1-0716-0150-1_32, https://www.alvascience.com/alvadesc-descriptors/.
- D. Ballabio, V. Consonni, A. Mauri, M. Claeys-Bruno, M. Sergent and R. Todeschini, A novel variable reduction method adapted from space-filling designs, Chemom. Intell. Lab. Syst., 2014, 136, 147–154, DOI:10.1016/j.chemolab.2014.05.010.
- P. Ambure, R. B. Aher, A. Gajewicz, T. Puzyn and K. Roy, NanoBRIDGES software: open access tools to perform QSAR and nano-QSAR modeling, Chemom. Intell. Lab. Syst., 2015, 147, 1–13, DOI:10.1016/j.chemolab.2015.07.007 , https://teqip.jdvu.ac.in/QSAR_Tools/#DPT.
- R. W. Kennard and L. A. Stone, Computer aided design of experiments, Technometrics, 1969, 11, 137–148, DOI:10.1080/00401706.1969.10490666.
- T. M. Martin, P. Harten, D. M. Young, E. N. Muratov, A. Golbraikh, H. Zhu and A. Tropsha, Does rational selection of training and test sets improve the outcome of QSAR modeling?, J. Chem. Inf. Model., 2012, 52, 2570–2578, DOI:10.1021/ci300338w , https://teqip.jdvu.ac.in/QSAR_Tools/#DatasetDiv.
- H.-S. Park and C.-H. Jun, A simple and fast algorithm for K-medoids clustering, Expert Syst. Appl., 2009, 36, 3336–3341, DOI:10.1016/j.eswa.2008.01.039 , https://teqip.jdvu.ac.in/QSAR_Tools/#ModKMedoid.
- V. Kumar, A. Banerjee and K. Roy, Breaking the barriers: Machine-learning-based c-RASAR approach for accurate blood–brain barrier permeability prediction, J. Chem. Inf. Model., 2024, 64, 4298–4309, DOI:10.1021/acs.jcim.4c00433.
- A. Nandy, S. Kar and K. Roy, Development of classification-and regression-based QSAR models and in silico screening of skin sensitisation potential of diverse organic chemicals, Mol. Simul., 2014, 40, 261–274, DOI:10.1080/08927022.2013.801076 , https://sites.google.com/jadavpuruniversity.in/dtc-lab-software/home#h.obtkiouqoqut.
- G. Cano, J. Garcia-Rodriguez, A. Garcia-Garcia, H. Perez-Sanchez, J. A. Benediktsson, A. Thapa and B. A. Alastair, Automatic selection of molecular descriptors using random forest: application to drug discovery, Expert Syst. Appl., 2017, 72, 151–159, DOI:10.1016/j.eswa.2016.12.008.
- D. R. Shin, I. H. Song and S. K. Lee, Interpretable QSAR modelling for immunotoxicity prediction using enhanced fingerprint and SHAP-based feature selection, SAR QSAR Environ. Res., 2025, 36, 955–969, DOI:10.1080/1062936X.2025.2578237.
- F. Pedregosa, G. Varoquaux, A. Gramfort, V. Michel, B. Thirion, O. Grisel, M. Blondel, P. Prettenhofer, R. Weiss, V. Dubourg, J. Vanderplas, A. Passos, D. Cournapeau, M. Brucher, M. Perrot and E. Duchesnay, Scikit-learn: Machine learning in Python, J. Mach. Learn. Res., 2011, 12, 2825–2830 Search PubMed , http://scikit-learn.org/stable/#;Jupyter%20Notebook: https://jupyter.org/try-jupyter/lab/.
- G. W. Snedecor and W. G. Cochran, Statistical Methods, Wiley-Blackwell, 8th edn, 1989 Search PubMed.
- L. Breiman, Random Forests, Mach. Learn., 2001, 45, 5–32, DOI:10.1023/A:1010933404324.
- C. Cortes and V. Vapnik, Support-vector networks, Mach. Learn., 1995, 20, 273–297, DOI:10.1007/BF00994018.
- S. Zhao, B. Zhang, J. Yang, J. Zhou and Y. Xu, Linear discriminant analysis, Nat. Rev. Methods Primers, 2024, 4, 70, DOI:10.1038/s43586-024-00346-y.
- J. C. Stoltzfus, Logistic regression: a brief primer, Acad. Emerg. Med., 2011, 18, 1099–1104, DOI:10.1111/j.1553-2712.2011.01185.x.
- Y. Freund and R. E. Schapire, A decision-theoretic generalization of on-line learning and an application to boosting, J. Comput. Syst. Sci., 1997, 55, 119–139, DOI:10.1006/jcss.1997.1504.
- J. H. Friedman, Greedy Function Approximation: A gradient boosting machine, Ann. Stat., 2001, 29, 1189–1232 Search PubMed . https://www.jstor.org/stable/2699986.
- R. P. Sheridan, W. M. Wang, A. Liaw, J. Ma and E. M. Gifford, Extreme gradient boosting as a method for quantitative structure–activity relationships, J. Chem. Inf. Model., 2016, 56, 2353–2360, DOI:10.1021/acs.jcim.6b00591.
- K. O. K-Nearest Neighbors, in Dimensionality Reduction with Unsupervised Nearest Neighbors. Intelligent Systems Reference Library, Springer, Berlin, Heidelberg, 2013, p. 51, DOI:10.1007/978-3-642-38652-7_2.
- Y. Qin, A review of quadratic discriminant analysis for high-dimensional data, WIREs Comput. Stat., 2018, 10, e1434, DOI:10.1002/wics.1434.
- M. Ontivera-Ortega, A. L. Castellanos, G. Valente, R. Goebel and M. Valdes-Sosa, Fast Gaussian Naïve Bayes for searchlight classification analysis, Neuroimage, 2017, 163, 471–479, DOI:10.1016/j.neuroimage.2017.09.001.
- C. E. Rasmussen, C. K. I. Williams, Gaussian Process for Machine Learning, MIT Press, 2006. https://gaussianprocess.org/gpml/chapters/RW.pdf Search PubMed.
- J. T. Hancock and T. M. Khoshgoftaar, CatBoost for big data: an interdisciplinary review, J. Big Data, 2020, 7, 94, DOI:10.1186/s40537-020-00369-8.
- A. Krogh, What are artificial neural networks?, Nat. Biotechnol., 2008, 26, 195–197, DOI:10.1038/nbt1386.
- A. W. Sobanska, A. Banerjee and K. Roy, organic sunscreens and their products of degradation in biotic and abiotic conditions—in silico studies of drug-likeness and human placental transport, Int. J. Mol. Sci., 2024, 25, 12373, DOI:10.3390/ijms252212373.
- S. Hochreiter and J. Schmidhuber, Long Short-Term Memory, Neural Comput., 1997, 9, 1735–1780, DOI:10.1162/neco.1997.9.8.1735.
- C. Toma, A. Manganaro, G. Raitano, M. Marzo, D. Gadaleta, D. Baderna, A. Roncaglioni, N. Kramer and E. Benfenati, QSAR models for human carcinogenicity: an assessment based on oral and inhalation slope factors, Molecules, 2021, 26, 127, DOI:10.3390/molecules26010127.
- P. Gramatica, Principles of QSAR models validation: internal and external, QSAR Comb. Sci., 2007, 26, 694–701 CrossRef CAS.
- R. Rodriguez-Perez and J. Bajorath, Interpretation of compound activity predictions from complex machine learning models using local approximations and shapley values, J. Med. Chem., 2020, 63, 8761–8777, DOI:10.1021/acs.jmedchem.9b01101.
- P. S. Spencer and G. E. Kisby, Role of hydrazine-related chemicals in cancer and neurodegenerative disease, Chem. Res. Toxicol., 2021, 34, 1953–1969, DOI:10.1021/acs.chemrestox.1c00150.
- A. R. Tricker and R. Preussmann, Carcinogenic N-nitrosamines in the diet: occurrence, formation, mechanisms and carcinogenic potential, Mutat. Res., Genet. Toxicol., 1991, 259, 277–289, DOI:10.1016/0165-1218(91)90123-4.
- K. P. Cross and D. J. Ponting, Developing structure-activity relationships for N-nitrosamine activity, Comput. Toxicol., 2021, 20, 100186, DOI:10.1016/j.comtox.2021.100186.
- S. Schieferdecker and E. Vock, quantum chemical evaluation and QSAR modeling of n-nitrosamine carcinogenicity, Chem. Res. Toxicol., 2025, 38, 325–339, DOI:10.1021/acs.chemrestox.4c00476.
- V. Frecer and S. Miertus, Theoretical QSAR study on carcinogenic potency of N-nitrosamines, Neoplasma, 1988, 35, 525–538 Search PubMed.
- M. Zhong, X. Nie, A. Yan and Q. Yuan, Carcinogenicity prediction of noncongeneric chemicals by a support vector machine, Chem. Res. Toxicol., 2013, 26, 741–749, DOI:10.1021/tx4000182.
- D. Stumpfe, H. Hu and J. Bajorath, Evolving Concept of Activity Cliffs, ACS Omega, 2019, 4, 14360–14368, DOI:10.1021/acsomega.9b02221.
- A. Roncaglioni, A. Lombardo and E. Benfenati, The VEGAHUB Platform: The Philosophy and the Tools. Alt, Lab. Anim., 2022, 50, 121–135, DOI:10.1177/02611929221090530.
| This journal is © The Royal Society of Chemistry 2026 |