Open Access Article
Open Access ArticleCaenorhabditis elegans fed native gut microbiota have altered bioenergetic pathway utilization impacting mitochondrial function and susceptibility to pollutants
Christina M. Bergemann *a,
Laura E. Jamesona,
Isabel W. Kenny-Ganzertb,
Javier Huayta
*a,
Laura E. Jamesona,
Isabel W. Kenny-Ganzertb,
Javier Huayta a,
Pol Castellano-Escuderc,
Arunabh Sarkara,
Olga R. Ilkayevac,
David R. Sherwoodb,
Matthew D. Hirscheyc and
Joel N. Meyer*a
a,
Pol Castellano-Escuderc,
Arunabh Sarkara,
Olga R. Ilkayevac,
David R. Sherwoodb,
Matthew D. Hirscheyc and
Joel N. Meyer*a
aNicholas School of the Environment, Duke University, Durham, NC, USA
bDepartment of Biology, Duke University, Durham, NC, USA
cDuke Molecular Physiology Institute, Duke University School of Medicine, Durham, NC, USA
First published on 12th March 2026
Abstract
The gut microbiome can influence host health by facilitating digestion, immune function, and xenobiotic metabolism. Microbial metabolites can influence mitochondrial function by shifting bioenergetic pathways, potentially altering sensitivity to mitochondrial toxicants. However, mechanisms through which the gut microbiota can alter mitochondrial function and susceptibility to mitochondrial toxicity are not well characterized. We used the model organism Caenorhabditis elegans and the microbiome kit CeMbio, a characterized collection of native gut commensals, to explore the interactions between gut microbiota, mitochondrial function, and chemical susceptibility. C. elegans grown on selected bacterial strains had varying levels of steady-state whole-body ATP, with an ∼3 fold difference between the highest and lowest strains, as well as 2- and 3-fold changes in antioxidant and mitochondrial unfolded protein gene induction. Further, C. elegans grown on selected bacterial strains showed differential sensitivity to short-term exposure to chemicals that inhibit mitochondrial electron transport chain Complexes I, II, and V, and fatty acid oxidation. To test mechanistically how microbiome-mediated sensitivities could result in chemical susceptibility, we carried out follow-up experiments using the Complex I inhibitor rotenone. We found that C. elegans grown on BIGb0170 (Sphingobacterium multivorum) had much higher lethality after 24- and 48-hour exposures than when grown on MYb10 (Acinetobacter guillouiae), MYb11 (Pseudomonas lurida), and OP50 (Escherichia coli) strains. Metabolomic analysis revealed that C. elegans grown on BIGb0170 had lower amounts of triglycerides and acylcarnitines. ATP levels were partially rescued by supplementing BIGb0170 with pyruvate. This work suggests that BIGb0170 can impact mitochondrial function through changes in metabolite abundance, which can increase sensitivity to the Complex I inhibitor rotenone.
Environmental significanceHumans encounter a myriad of chemicals daily, and some individuals are more susceptible to chemical exposure than others, resulting in exacerbated adverse effects or disease. One potential reason for this variability is the gut microbiome, which may influence chemical susceptibility through mechanisms such as altering host stress responses or affecting mitochondrial bioenergetic pathways. The use of synthetic chemicals is increasing, which could mean more impacts on human and environmental health. Changes in bioenergetic pathway utilization can impact susceptibility to mitochondrial toxicants. Understanding how gut microbiota impact our health may contribute to new therapeutic and dietary ways to protect ourselves from the harmful impacts of chemicals and environmental pollution. Further, understanding how chemicals impact gut microbiota can contribute to improved chemical regulation. |
Introduction
The interactions between gut microbiota and mitochondria have profound implications for human health, influencing health outcomes including metabolic,1 neurodegenerative,2 and inflammatory bowel diseases.3,4 Gut microbiota can impact mitochondrial function by producing beneficial compounds such as short-chain fatty acids,4 colanic acid,5 and vitamins6 or releasing potentially detrimental metabolites such as nitric oxide, hydrogen sulfide, and hydrogen peroxide.7–9 These metabolites can significantly alter overall mitochondrial function, affecting adenosine triphosphate (ATP) levels, mitochondrial respiration, biogenesis, regulation of fusion and fission,5 activation of the mitochondrial stress responses,10,11 and redox state alterations.12Differences in mitochondrial function can impact host development, lifespan, and susceptibility to stressors, such as environmental chemicals.6 The total number of synthetic chemicals globally is estimated to be around 350![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000,13 and the United States produces 1500 new chemicals annually, with overall use continuing to rise.14 Understanding their potential toxic effects is necessary to minimize consequences for human and environmental health.15,16 For reasons including their high lipid content, mitochondria are often targeted by chemical pollutants, leading to mitochondrial dysfunction and disruption of mitochondrial homeostasis.17–20 Mitochondrial dysfunction has been linked to several metabolic and neurodegenerative diseases such as obesity, Parkinson's disease, and cancer.21 However, inter-individual variation in mitochondrial function results in a portion of the population being more susceptible to environmental exposures and mitochondrial disease outcomes. An underexplored mechanism that could contribute to interindividual variability in mitochondrial toxicity is the gut microbiome.
000,13 and the United States produces 1500 new chemicals annually, with overall use continuing to rise.14 Understanding their potential toxic effects is necessary to minimize consequences for human and environmental health.15,16 For reasons including their high lipid content, mitochondria are often targeted by chemical pollutants, leading to mitochondrial dysfunction and disruption of mitochondrial homeostasis.17–20 Mitochondrial dysfunction has been linked to several metabolic and neurodegenerative diseases such as obesity, Parkinson's disease, and cancer.21 However, inter-individual variation in mitochondrial function results in a portion of the population being more susceptible to environmental exposures and mitochondrial disease outcomes. An underexplored mechanism that could contribute to interindividual variability in mitochondrial toxicity is the gut microbiome.
Mitochondria generate ATP through oxidative phosphorylation and glycolysis. Oxidative phosphorylation involves the electron transport chain, comprising four complexes that transfer electrons and pump protons to establish a membrane potential, which is utilized by a fifth complex, ATP synthase, for ATP production. In humans, these complexes ultimately rely largely on substrates from the breakdown of dietary components and metabolites produced by the gut microbiota in the colon that feed into the tricarboxylic acid cycle (TCA).22 Thus, gut microbiota can alter bioenergetic pathways by altering metabolite and substrate availability, impacting oxidative phosphorylation and other bioenergetic processes. Different bioenergetic pathways can impact overall ATP levels and potentially alter susceptibility to mitochondrial toxicants.23
Gut microbiota can also activate mitochondrial stress response pathways by releasing bacterial toxins like respiratory chain inhibitors24 or reactive oxygen species.8 The best-studied mitochondrial stress response pathway is the mitochondrial unfolded protein response (mtUPR). Activation of mtUPR can protect against these toxins by upregulating detoxification genes and antimicrobial defenses; however, mitochondrial dysfunction that exceeds the mtUPR's compensatory capacity can lead to irreparable damage.25 A likely reason bacteria release harmful toxins that target mitochondria is mitochondria's role in cell apoptosis26 or to facilitate release of iron, essential for bacterial growth.9,27 Gut microbiota can also alter redox states through the release of pro-oxidant metabolites or antioxidant molecules.12 Changes in redox state can upregulate detoxification pathways to aid in maintaining homeostasis, which can impact susceptibility to pollutants.
In this study, we tested these possibilities using the model organism Caenorhabditis elegans and the collection of native gut microbiota CeMbio.28 C. elegans has many advantages in investigating the role of the microbiome, including ease of creating germ-free larvae and being a bacterivore with well-conserved mitochondrial biology.29–34 The CeMbio collection consists of characterized strains of bacteria isolated from the intestinal tract of wild C. elegans that colonize the intestinal tract, vary substantially in the total number and type of metabolites they produce, and alter the developmental rate in C. elegans.28 Given that these bacterial strains varied in the number and types of metabolites produced, coupled with differences in development, we hypothesized that these strains altered mitochondrial function.35 We selected 6 strains that altered developmental time, encompassing the slowest and fastest strains:28 Sphingobacterium multivorum (BIGb0170), Ochrobactrum vermis (MYb71), Acinetobacter guillouiae (MYb10), Stenotrophomonas indicatrix (JUb19), Pseudomonas berkeleyensis (MSPm1), and Pseudomonas lurida (MYb11). In addition to testing native gut microbiota, we also characterized two common laboratory E. coli strains, OP50 and HT115. We found that the different strains resulted in large variations in bioenergetics, molecular stress responses, and susceptibility to the mitochondrial toxicants. We further characterized the mechanisms underlying the high sensitivity of nematodes grown on BIGb0170 to the Complex I inhibitor rotenone. Overall, this study provides insights into how gut microbiota can modulate mitochondrial function and susceptibility to chemical exposure, highlighting the potential for gut microbiota to influence health outcomes through mitochondrial pathways.
Results
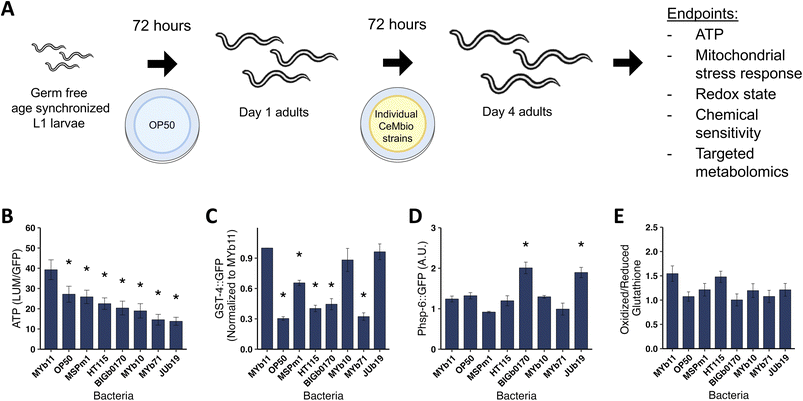
Native gut microbiota impact whole-body ATP levels and stress response pathways in C. elegans
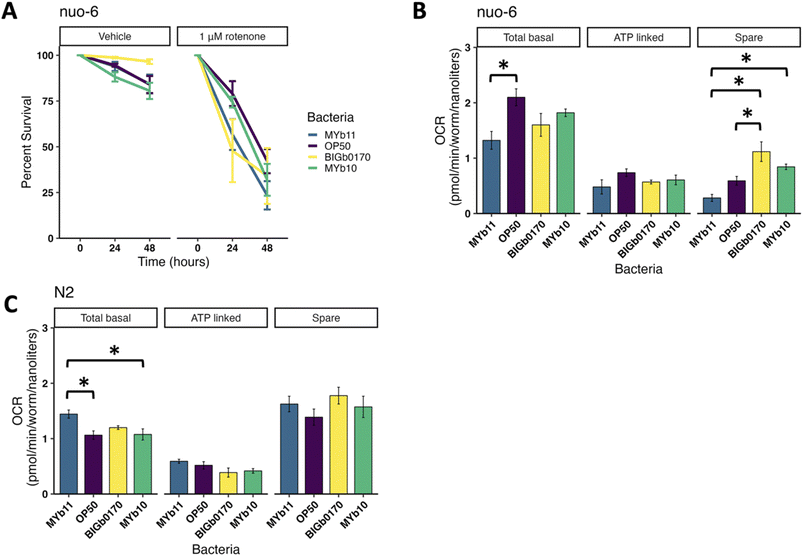
Since mitochondrial function changes dramatically during development,36 we grew C. elegans on selected CeMbio strains after they reached adulthood to avoid developmental delays that could confound interpretation of mitochondrial parameters (Fig. 1A). While C. elegans ingests and digests bacteria as a food source,32 the CeMbio strains used here colonize the C. elegans intestinal tract.28 Interestingly, despite raising all nematodes to adulthood on the E. coli strain OP50, we still saw differences in length after transferring them to the CeMbio strains. We observed a 21% difference between nematodes fed on MYb11 and MYb71 (p = 0.002) and a 16% difference between MYb11 and BIGb0170 (p = 0.017). We saw larger differences in worm volume, with a 45% decrease between MYb11 and MYb71 (p < 0.001) and a 37% difference between MYb11 and BIGb0170 (p < 0.001, SI Fig. 1A and B). Therefore, differences in bacterial diet affect adult as well as larval growth, though to a lesser degree in adulthood than in larval development.28To determine if mitochondrial function was altered in C. elegans fed on these strains, we measured whole-body ATP levels in vivo. We found that ATP levels varied significantly, with the highest levels in nematodes fed MYb11 and lowest in nematodes fed JUb19 (2.8-fold decrease, p < 0.001, Fig. 1B). We considered the possibility that the ATP differences could be explained by differences in mitochondrial abundance or changes in oocyte production; however, we found that differences in ATP levels are not a result of mitochondrial DNA copy number or the presence of the germline (SI Fig. 1C–E).
Next, we measured reporters of a number of stress response pathways,11,37 with a particular focus on mitochondrial stress responses. We assessed induction of glutathione S-transferase (gst-4), a marker of activation of the antioxidant response gene battery38–40 with the C. elegans strain CL2166 and41 observed roughly 3-fold baseline expression differences among the bacterial strains (Fig. 1C). We measured hsp-6 expression, a marker for mitochondrial unfolded protein (mtUPR) activity,42,43 using C. elegans strain SJ4100 after 24 and 72 hours on the bacterial strains and found that strains BIGb0170 and JUb19 increased hsp-6 activity in nematodes at 72 hours, while OP50, BIGb0170, and JUb19 increased hsp-6 at 24 hours (Fig. 1D, 2A SI). We also measured two other mitochondrial stress response pathways in the nematodes grown on the bacterial strains after 24 and 72 hours, namely the mitochondrial MAPK (MAPKmt) and the ethanol and stress response element (ESRE). The MAPKmt pathway was induced by nematodes grown on BIGb0170 after 24 hours and still trending towards an increase at 72 hours, while MSPm1 was trending at 24 hours and significantly induced at 72 hours (SI Fig. 2B–D). None of the bacterial strains significantly induced the ESRE pathway at either 24 hours or 72 hours (SI Fig. 2C–E). Lastly, we looked at the redox state in nematodes using the JV2 strain,44 but did not observe a significant difference across bacterial strains (Fig. 1D), despite differences in H2O2 production (SI Fig. 2F) and gst-4 induction between bacterial strains. Overall, these native gut microbiota cause significant changes to many aspects of C. elegans physiology and stress response biology.
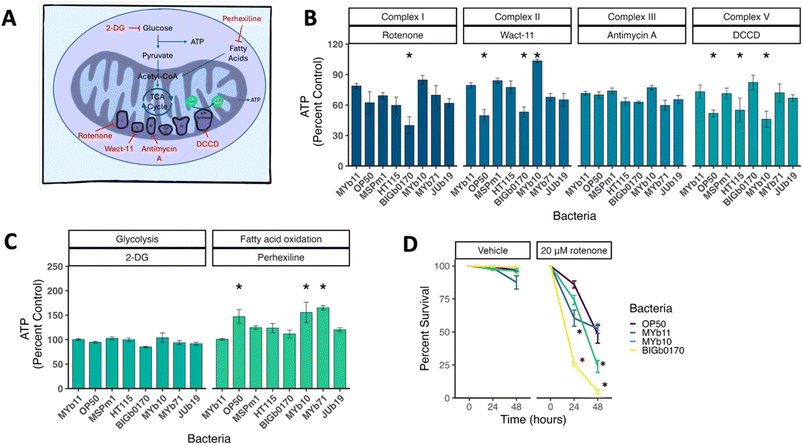
Native gut microbiota impact bioenergetic pathway utilization and sensitivity to rotenone
Given the critical nature of ATP levels for multiple cellular and organismal functions, we sought to understand the bioenergetic changes that contribute to the observed differences in ATP levels. To test this, we measured ATP levels after a short exposure to mitochondrial toxicants that inhibit specific bioenergetic complexes and pathways,45–47 using the ATP reporter strain PE255 (Fig. 2A). The principle behind these assays is that if ATP levels in nematodes on different bacterial diets are differentially altered following pharmacological inhibition of a specific bioenergetic process, we can infer that the bacterial diet alters the utilization of that pathway for ATP production.45 Of the electron transport chain inhibitors tested, we found significant differences in ATP levels in nematodes fed different bacterial strains when exposed to rotenone (Complex I), wact-11 (Complex II), antimycin A (Complex III), and dicyclohexylcarbodiimide (DCCD, Complex V) (Fig. 2B). We also tested two other pathways that can impact mitochondrial function, glycolysis and fatty acid oxidation. We saw no significant differences between nematodes grown on different bacteria with the glycolysis inhibitor 2-deoxy-D-glucose (2-DG), but did find differences with the fatty acid oxidation inhibitor perhexiline (Fig. 2C).To test if microbiota-mediated sensitivities to chemical inhibition of ATP production would result in organismal chemical susceptibility, we carried out follow-up experiments using the Complex I inhibitor rotenone in a subset of four bacterial strains. We chose these bacterial strains because they represented the bacteria that showed the least and most sensitivity to short-term rotenone-induced ATP depletion, as well as the intermediate and commonly used laboratory strain, OP50. We found that C. elegans grown on BIGb0170 exhibited significantly higher lethality after 24- and 48-hour exposures, compared to strains MYb10, MYb11, and OP50 (Fig. 2D).
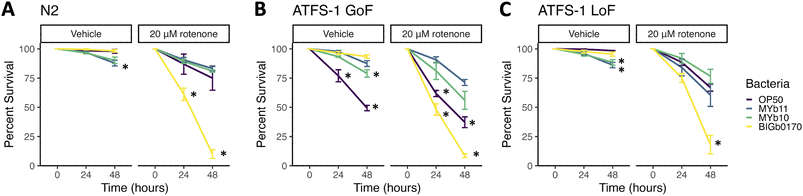
BIGb0170-mediated sensitivity to rotenone does not result from differences in the mtUPR or vulnerability to oxidative stress
Of the four bacteria for which we examined rotenone lethality, only BIGb0170 induced the mtUPR in C. elegans. Therefore, to investigate whether this induction mediates rotenone sensitivity, we tested the survival of nematodes co-exposed to the bacteria and rotenone for 48 hours using a mtUPR gain-of-function (GoF)48 mutant and a knockout mutant (LoF),49 compared to the wild type (N2) (Fig. 3). Interestingly, in N2, rotenone lethality was overall decreased on MYb10, MYb11, and OP50 compared to lethality in the ATP reporter strain, resulting in an even greater relative rotenone sensitivity caused by the BIGb0170 bacteria (Fig. 3A). We found that the mtUPR LoF and GoF nematodes grown on BIGb0170 bacteria still had higher lethality compared to nematodes grown on MYb10, MYb11, or OP50. We also observed higher lethality with the mtUPR GoF mutant in the OP50 DMSO (vehicle) condition, as reported previously,37 likely due to the energy-intensive nature of mtUPR induction. The fact that BIGb0170-fed nematodes still showed higher sensitivity with rotenone than the other strains in both the GoF and LoF strains is inconsistent with the transcriptional activation of mtUPR observed on BIGb0170 explaining rotenone sensitivity.While we didn't see baseline differences in the redox state in nematodes fed different bacteria, we considered the possibility that the different diets might alter sensitivity to secondary prooxidant challenges. To test this, we measured redox state after exposure to rotenone and hydrogen peroxide. We did not observe a significant difference between the bacteria and the treatment in response to hydrogen peroxide (SI Fig. 3A), but we did see a small but statistically significant interaction with exposure to DMSO and rotenone. Nematodes grown on OP50 bacteria initially began in a more-reduced state and became more oxidized after exposure to DMSO and rotenone, compared to the other strains. The only statistical significance we observed between the DMSO and rotenone conditions was in nematodes grown on MSPm1 (SI Fig. 3B). Since BIGb0170-grown nematodes showed no increase in oxidation state after rotenone exposure, it is unlikely that changes in redox state drove rotenone sensitivity.
BIGb0170 exposure causes large metabolomic changes at the baseline and in response to rotenone
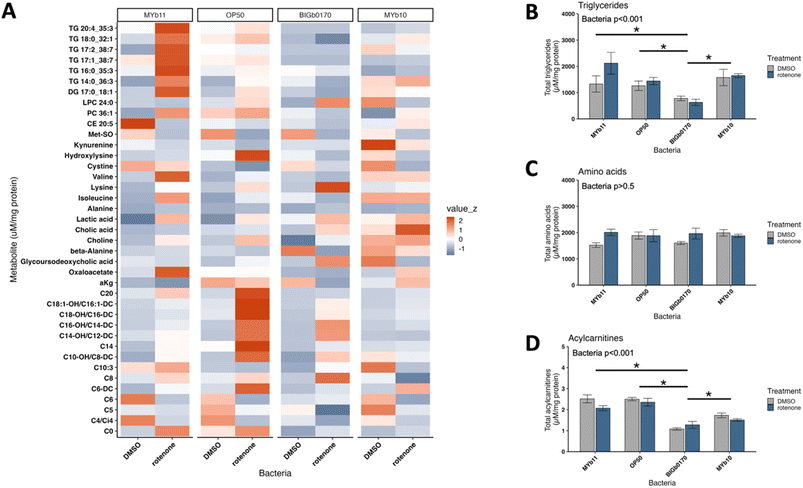
Given that the rotenone sensitivity of C. elegans grown on BIGb0170 was not explained by changes in redox state or mtUPR, we next assessed the differences in metabolites. We used two targeted metabolomics panels, a smaller one that included metabolites specifically involved in mitochondrial metabolism (organic acids, biogenic amines, and acylcarnitines), and a larger one that included metabolites sensitive to gut microbiota influence (Biocrates MxP® Quant 500). Metabolites were measured in C. elegans after 24 hours of co-exposure to bacteria and rotenone, following 72 hours of exposure to bacteria alone. In the Q500 panel, we found that the nematodes grown on BIGb0170 separated from those grown on the other three bacterial strains (PC1 in Fig. 4A). Interestingly, the predicted BIGb0170 bacterial metabolome was also previously reported to be distinct from the other CeMbio metabolomes, based on a principal component analysis of their metabolic profiles.28 This separation occurred regardless of DMSO or rotenone exposure; indeed, there was no clear separation based on rotenone treatment for any of the worm groups using the Q500 panel (Fig. 4A). We also found that nematodes grown on MYb10 bacteria had much higher overall abundance of summed classes of metabolites in the DMSO condition compared to the other bacteria; however, this difference disappeared after rotenone exposure (Fig. 4B). In the targeted mitochondrial panel, we didn't see a separation based on bacteria but did see a separation between the DMSO and rotenone treatments (PC1 in Fig. 4C). PC2 in the targeted mitochondrial panel appeared to be largely driven by the MYb11 rotenone samples. We also didn't see robust differences in the overall abundance of amino acids and biogenic amines in nematodes fed the four different bacteria in the targeted mitochondrial panel (Fig. 4D). Graphs of each metabolite measured in the mitochondrial panel are presented in (SI Fig. 7–9). Interestingly, from our targeted mitochondrial panel, we saw that nematodes grown on BIGb0170 had higher oxidized/reduced glutathione levels compared to nematodes grown on MYb10 and MYb11 bacteria (SI Fig. 4A). Furthermore, we observed that nematodes grown on MYb10 exhibited higher total glutathione levels compared to nematodes grown on MYb11 and BIGb0170 bacteria (SI Fig. 4B).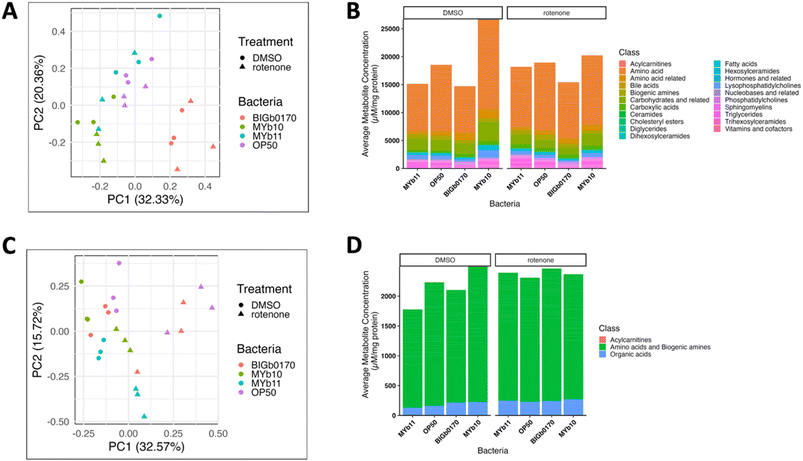 | ||
| Fig. 4 Targeted metabolomics analysis of C. elegans raised on MYb11, OP50, BIGb0170, and MYb10 bacteria. (A) PCA analysis for metabolites measured in the Q500 microbiome sensitive panel and the (B) average metabolite abundance of each class of metabolites measured. (C) PCA analysis for the mitochondrial metabolite panel and the (D) average metabolite abundance of each class measured. Acylcarnitines abundance is not visible in panel D due to low abundance but graphed separately in Fig. 5D. N = 3 biological replicates. | ||
To identify metabolites involved in rotenone sensitivity, we used a linear mixed model to identify metabolites from both panels that showed a statistically significant interaction between the main effects of bacteria and treatment, indicating that they were differentially impacted by rotenone exposure depending on the bacteria. We identified 39 metabolites that exhibited a significant interaction (Fig. 5A). These included 6 triglycerides, 14 acylcarnitines, and 8 amino acids or amino acid-related compounds. We found that nematodes grown on BIGb0170 had lower overall (summed) amounts of triglycerides and acylcarnitines, but not amino acids (Fig. 5B).
Complex I mutant nematodes grown on BIGb0170 do not have sensitivity to rotenone
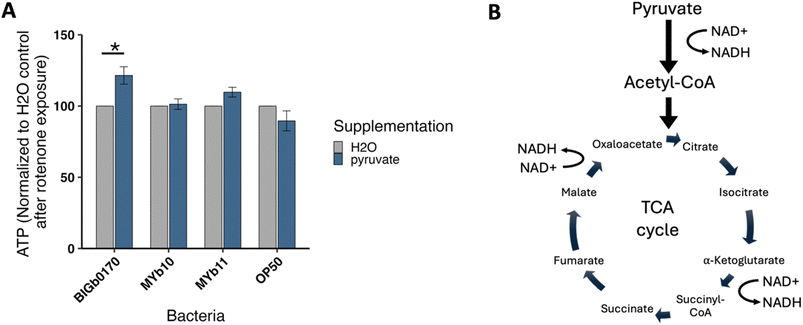
Given the changes in triglycerides and acylcarnitines, we hypothesized that nematodes grown on BIGb0170 bacteria were sensitive to rotenone due to less substrate to fuel the TCA cycle, thereby limiting reducing equivalent availability used to fuel Complex I. The TCA cycle begins with acetyl-CoA which is primarily produced from pyruvate oxidation, branched chain amino acid metabolism, or beta-oxidation in mitochondria. If the effect of BIGb0170 is the result of reduced Complex I substrate availability, BIGb0170 should have less effect on the health of nematodes that are already Complex I activity-deficient and therefore less dependent on Complex I substrate. To test this, we used the Complex I mutant nuo-6(qm200), which carries a partial loss of function resulting in severely reduced Complex I activity compared to wild-type.50 Mutant nematodes were grown on OP50 bacteria until they reached day 1 of adulthood, then plated on the four bacterial strains for 72 hours before being co-exposed to bacteria and rotenone for 48 hours, during which lethality was scored every 24 hours. We no longer saw bacterial sensitivity when nuo-6 nematodes were exposed to rotenone (Fig. 6A), in contrast to the sensitivity of wild-type nematodes fed BIGb0170 exposed to rotenone (Fig. 2D and 3A). Thus, while nuo-6 nematodes are sensitive to rotenone as expected, the sensitizing effect that the BIGb0170 bacteria have in wild-type nematodes did not occur in the context of preexisting Complex I deficiency.To assess whether BIGb0170 bacteria had different effects on oxidative phosphorylation in nuo-6 versus N2, we measured oxygen consumption rates in both strains on all four bacterial diets. Overall, the differences we measured between strains and bacterial diets were small. N2 nematodes had higher spare respiratory capacity than nuo-6 nematodes independent of microbiota. Mitochondrial respiration in nuo-6 nematodes grown on MYb11 was lower than when grown on OP50 (p = 0.03), but we found no bacteria-related differences in mitochondrial basal, non-mitochondrial basal, ATP-linked, or proton leak (SI Fig. 5A). However, nuo-6 nematodes grown on BIGb0170 had higher spare capacity compared to OP50 and MYb11 (Fig. 6B). In wild-type, we observed a significant effect of bacterial food on total basal oxygen consumption, with nematodes grown on MYb11 having 34–36% higher total basal oxygen consumption than those grown on OP50 and MYb10 (Fig. 6C, 5B SI). These results suggest that the compensatory mechanisms that permit nuo-6 nematodes to produce energy and survive despite reduced Complex I activity are robust to the bioenergetic interference caused by BIGb0170 in wild-type nematodes, consistent with BIGb0170 interfering with CI-mediated energy production.
An alternative hypothesis for the sensitivity of nematodes grown on BIGb0170 to rotenone is that exposure to BIGb0170 results in a decrease in the production of Complex I proteins, specifically, although we did not observe a difference in mtDNA copy number. We tested this by measuring the expression levels of two Complex I subunits, using two strains expressing endogenously Crispr-tagged Complex I proteins, NDUF-7 and NDUV-2. We did not see differences between nematodes grown on the four bacterial strains in levels of either subunit (SI Fig. 6A).
BIGb0170 exposure-mediated rotenone sensitivity is worsened by pyruvate dehydrogenase kinase inhibition and partially rescued by pyruvate supplementation
Our targeted metabolomics data and the nuo-6 data are consistent with Complex I substrate being limiting in nematodes grown on BIGb0170. We carried out additional experiments to test this idea.First, we exposed nematodes grown on the four bacterial strains to dichloroacetate (DCA), which inhibits pyruvate dehydrogenase kinase, which inhibits pyruvate dehydrogenase. The pyruvate dehydrogenase complex converts pyruvate to acetyl-CoA for use in the TCA cycle. Thus, DCA increases the rate of conversion of pyruvate to acetyl-CoA. We found that exposure to DCA, for four hours without bacteria present, caused more ATP depletion in BIGb0170-fed nematodes than nematodes fed other bacteria (SI Fig. 6B). This could occur if forcing a rewiring of the bioenergetic pathways adopted by nematodes grown on BIGb0170 by exposing them to DCA results in less-efficient energy production.
We also examined the lifespan of nematodes grown on the four bacteria, starting at day 1 of adulthood. Dietary restriction as well as mitochondrial dysfunction can increase nematode lifespan.51–53 In adult nematodes, dietary restriction has been shown to increase lifespan, while mitochondrial dysfunction lifespan extension occurs mainly during development.53 We found that nematodes grown on BIGb0170 bacteria lived slightly longer than OP50, MYb10, and MYb11 nematodes, also consistent with the possibility that this bacteria has limited substrate51,52 (SI Fig. 6C).
If the Complex I substrate is limiting in nematodes grown on BIGb0170, then sensitivity should be alleviated through supplementation with substrates that increase NADH levels, such as pyruvate.54 We supplemented BIGb0170 bacteria with 2.5 mM sodium pyruvate for 72 hours prior to a one-hour exposure to rotenone and found that pyruvate supplementation partially rescued ATP levels in nematodes after rotenone exposure (Fig. 7, Supplemental Fig. 6D).
Discussion
We found that six strains of native gut microbiota and two commonly used laboratory bacterial strains have distinct and significant impacts on mitochondrial function, metabolite profiles, stress response pathways, and toxicant response in C. elegans. Among these strains, we focused on the response to the mitochondrial inhibitor rotenone in nematodes grown on the BIGb0170 bacteria. These nematodes exhibited the greatest sensitivity to rotenone-mediated ATP depletion and lethality, showed a strong induction of two mitochondrial stress-response genes, and displayed impaired adult growth. We aimed to uncover how this naturally occurring gut bacteria caused such significant effects on mitochondrial function, organismal physiology, and health.Rotenone, by inhibiting Complex I, causes ATP depletion and superoxide anion generation.45,55,56 BIGb0170 bacteria reduced steady-state ATP levels in nematodes, but rotenone sensitivity mediated by BIGb0170 is unlikely to be driven solely by insufficient baseline ATP levels per se, as nematodes grown on other bacterial strains such as MYb10, MYb71, and JUb19 had even lower initial ATP levels but were not sensitive to rotenone-driven ATP depletion. We did not observe significant differences in redox-related endpoints (redox state at baseline or after challenge with H2O2 or rotenone, gst-4 induction) in nematodes grown on BIGb0170. We did see higher oxidized to reduced glutathione levels in nematodes grown on BIGb0170 in our target metabolomics data, but this was only statistically different from nematodes grown on MYb10 and MYb11. Furthermore, while the BIGb1070-mediated differences in induction in tested mitochondrial stress response pathways are interesting in and of themselves, we did not find evidence that they mediated rotenone sensitivity. However, we note that we measured transcriptional activation of the mtUPR pathway, not mtUPR function; we cannot exclude the possibility that BIG0170 bacteria disrupt the functional output of the mtUPR pathway. We also found no change in mitochondrial content as measured by mtDNA copy number or Complex I protein on BIGb0170-fed nematodes, so these do not explain rotenone sensitivity.
While ATP levels themselves are unlikely to explain why C. elegans grown on BIGb0170 were sensitive to rotenone, it appears that bioenergetic alterations are important, potentially by decreasing metabolic flexibility. Our metabolomic data indicated a lower overall abundance of bioenergetic precursor metabolites required to fuel the TCA cycle, and we observed a reduced ability to maintain steady-state ATP levels upon inhibition of either Complex I or Complex II. Pyruvate supplementation partially rescued rotenone-mediated ATP depletion in nematodes grown on the BIGb0170 bacteria, supporting a role for an insufficient supply of TCA cycle substrate. Gut microbiota regulate host lipid metabolism through several mechanisms including altering bile acid metabolism which affects lipid adsorption and by increasing fatty acid oxidation thereby decreasing triglyceride and acylcarnitine levels.57–59 We note that our experimental design did not allow us to distinguish whether the changes in the nematode metabolism were due to the differences in the bacterial metabolites, or physiological response of the worms to the bacteria.
Finally, our Complex I mutant nuo-6 results are consistent specifically with the hypothesis that BIGb0170 bacteria were disrupting Complex I function, since BIGb0170 had no differential effect on the sensitivity of nuo-6 to rotenone. We note that other researchers have also been able to partially rescue BIGb0170-mediated phenotypes with substrate supplementation. Jarkass et al. were able to partially rescue the developmental delay observed in nematodes raised on BIGb0170 by supplementing with linoleic acid.60 Linoleic acid could provide more fatty acids, which could increase TCA cycle substrates and acetyl-CoA levels in nematodes through beta oxidation. However, additional work will be required to demonstrate whether our pyruvate rescue effectively increased acetyl-CoA levels, as we hypothesized, because it may also have other effects. For example, Tauffenberger et al. found that supplementing C. elegans with 10 mM pyruvate (10 mM, versus 2.5 mM which was optimal in these studies) activated glutathione transferase activity (gst-4), and mtUPR (HSP-6 and HSP-60).61 Pyruvate supplementation may rescue ATP levels through alternative mechanisms, particularly redox effects, by consuming NADH and regenerating NAD + via lactate dehydrogenase, thereby supporting glycolysis.
We considered the possibility that BIGb0170 could be producing a metabolite that inhibits Complex I, but consider this unlikely. First, this should result in an increase in the concentrations of bioenergetic substrates (e.g., pyruvate),62–64 since substrate utilization would decrease; however, we did not observe this. Second, inhibition of Complex I should result in higher oxidative stress due to the creation of reactive oxygen species; however, as discussed above, we found no evidence to support this. We also did not observe significant changes in the ESRE pathway, which is normally induced by oxidative stress.65 Caveats to this conclusion are that the (whole-body) reporter strain JV2 that we used for redox state measurements may not be able to detect small or localized changes in redox from the bacterial diet, and we cannot fully exclude the possibility that there was an adaptive response to changes in redox stress since nematodes were on the bacteria for three days prior to imaging. However, if an adaptive response had occurred, we would expect to see increases in antioxidant defenses, such as induction in gst-4 or total glutathione levels, and we did not. Nonetheless, future studies should directly test the possibility of compensatory redox changes in rotenone sensitivity, and assess possible cell-specific, localized differences.
Another potential mechanism by which different microbiota might alter nematode sensitivity to rotenone is differential bacterial metabolism of rotenone. This is unlikely in the case of our ATP measurements, which were after a short one-hour rotenone exposure in the absence of bacteria. However, in our longer-term exposures, it is possible that bacteria could alter the pharmacokinetics of rotenone by altering uptake or depuration processes. Internal chemical concentrations within nematodes could help in assessing this possibility.
Interestingly, we did see an increase in GST-4::GFP in C. elegans grown on MYb11 and MYb10 but not OP50 or BIGb0170. GST-4 aids in detoxification of deleterious compounds and reactive oxygen species,39,66 which could be a potential mechanism behind the partial resistance to rotenone conveyed by nematodes grown on MYb11 and MYb10 bacteria. However, gst-4 can also be induced by the epidermal growth factor, unrelated to oxidative stress, which may explain why nematodes on some bacterial strains that have higher expression of GST-4::GFP but do not show changes in redox state.39 MYb10 also had high baseline total glutathione levels, which may convey resistance. However, it is likely not the sole mechanism since OP50 showed similar lethality with rotenone without changes to gst-4 expression. In general, it is likely that multiple mechanisms contribute to the relative sensitivity and resistance to rotenone in nematodes with different gut microbiota.
More broadly, bacterial effects on mitochondrial function may affect sensitivity to many compounds. Bacteria can modulate sensitivity to environmental pollutants in C. elegans such as aflatoxin B1,67,68 cadmium,69 juglone,6 the pharmaceutical drug metformin70, and antibiotic neomycin71; many of these affect mitochondria, and mitochondrial function is likely to be important in resistance to most forms of cellular toxicity. The tractability of C. elegans, along with resources such as CeMbio, offers a unique system to study host-microbiota-mediated sensitivities in the context of both mitochondrial and other forms of toxicity. Another potential advantage to working with C. elegans is that in this species, the microbiome is also the food source, allowing dissociation of the effects of the microbes themselves from the effects of the microbes on the breakdown of other dietary components. Such studies may help explain interindividual variability in chemical sensitivity, and may suggest therapeutic interventions.
Methods
C. elegans strains and maintenance
The following strains were used in this study: N2, PE255, PE327, QC115, atfs-1(tm4525), JV2, SJ4100, SLR115, ESRE3X, CL2166, MQ1333, NK2845, NK2841. All strains were maintained on K-agar plates with OP50 at 20 °C unless otherwise stated. atfs-1(tm4525) was purchased from NBRP, ESRE3X was gifted from the Kirienko lab, NK2841 (qy170[nduf-7:mNG] I) and NK2845 (qy174[nduv-2:mNG] V) were created and gifted from the Sherwood lab. All other strains were purchased from CGC, which is funded by the NIH Office of Research Infrastructure Programs (P40 OD010440).Bacterial strains and culturing conditions
CeMbio bacterial strains were purchased from CGC. To make stocks of each strain, one colony was picked from plates and cultured in a culture tube (VWR 60818-725) with 10.5 mL of Luria–Bertani broth (LB; 10 g L−1 tryptone, 5 g L−1 yeast extract, 10 g L−1 NaCl) at 25 °C for 24 hours. Stocks were made in 25% glycerol and kept at −80 °C. To culture bacteria, a 10 µL plastic inoculation loop was used to scrape frozen bacteria into a culture tube with 10.5 mL of LB inside a biological cabinet. Once inoculated, Parafilm was wrapped around the cap to ensure sterility and prevent liquid from leaking out. Culture tubes were loaded onto a Boekel Scientific Tube Rotator inside a 25 °C incubator at 15 rpm for 24 hours. After 24 hours, culture tubes were centrifuge at 3000 rcf for 10 min to pellet bacteria. Inside a biological cabinet, 6 mL of supernatant was removed to concentrate bacteria and samples were vortexed to resuspend bacteria prior to seeding plates. 10 cm K-agar plates were seeded with 300 µL of concentrated bacteria and spread evenly using a cell spreader. Seeded plates were maintained at 20 °C for 48–72 hours prior to plating nematodes.Experimental design
To obtain germ free age synchronized eggs, gravid adult nematodes were rinsed off plates with K-medium (51 mM NaCl, 32 mM KCl in ddH20) into a 15 mL centrifuge tube and treated with sodium hydroxide solution (3.5 mL of K-medium, 0.5 mL 5N NaOH, 1 mL 5% sodium hypochlorite) made fresh for 8 minutes then centrifuged for 2 minutes at 2200 rcf to pellet eggs. Supernatant was aspirated, and eggs were rinsed with 15 mL K-medium and centrifuged again. After K-medium was aspirated off, 5 mL of complete K-medium (K-medium, 3 mM calcium chloride, 3 mM magnesium sulfate, 5 µg mL−1 cholesterol) was added to the eggs and transferred to a cell culture flask for 17 hours to allow eggs to hatch and arrest at the L1 stage. Approximately 700 L1 larvae were plated to K-agar plates seeded with OP50 for 72 hours when they reached day 1 of adulthood (Fig. 1A). Nematodes were rinsed off OP50 plates and rinsed 5 times with 15 mL of K-medium to remove remaining bacteria. For each rinse, nematodes were gravity settled to remove bacteria and any eggs that had been laid. Nematodes were then counted and 400 nematodes plated to K-agar plates seeded with individual bacteria for 48 hours where they were again rinsed off to remove progeny and replated to the same bacteria for another 24 hours. After plating to new plates, any remaining progeny were removed.The nuo-6(qm200) (MQ1333) nematodes with a Complex I mutation are slow growing. To account for this, nematodes were raised on OP50 until they reached day 1 of adulthood, which was 5 days before being transferred to the CeMbio strains. Once on the CeMbio strains they were grown as previously stated.
C. elegans length and volume
After nematodes were measured for ATP levels, PE255 nematodes were plated onto unseeded K-agar plates and allowed to dry. Nematodes were imaged using the Keyence BZ-X700 microscope. Images were analyzed using the Wormsizer plugin in Fiji.72In vivo ATP measurements
Synchronized nematodes were obtained as previously described in the experimental design. Nematodes were washed and transferred to a white 96-well plate (VWR 33501-810) at a density of 50 or 75 nematodes/well. Nematodes were measured for luminescence immediately or following a one-hour exposure to rotenone (Sigma R8875), wact-11 (Chembridge Corporation), antimycin A (Enzo ALX-380-075-M010), DCCD (Sigma D80002), and Perhexiline (Tocris 5166) or four-hour exposure to 2-DG (Acros Organics 111![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 980
980![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 010) and DCA (Sigma 347795) according to a previously published protocol.45
010) and DCA (Sigma 347795) according to a previously published protocol.45
Copy number methods
Nuclear and mitochondrial DNA were measured according to the previously published protocol,73 with the addition of using the pCR 2.1 plasmid containing the cox-4 nuclear gene fragment to measure nuclear DNA copy number.Fluorescence imaging and quantification
For redox state using strain JV2, quantification of fluorescence intensity for both 405 and 488 nM wavelengths was performed using a custom-made MATLAB script (Math Works, version 2024b). The script generates a binary mask of an image captured in the 488 nM channel, followed by cleaning of non-worm features in the image using morphological functions. Properties such as area and mean fluorescent intensity of nematodes are extracted from the segmented image by using the “regionprops” function. Similarly, the same properties are extracted from the image captured in the 405 nM channel using the previous mask to ensure that the same nematodes and areas are measured. Background values are subtracted from each worm by calculating the average background intensity of each image (culture medium). Finally, the ratio of oxidized to reduced roGFP is calculated per worm by dividing the fluorescence intensities measured in the 405 nM channel by the 488 nM channel. The script prints a table with a worm label (for tracking purposes), the worm's area, fluorescent intensities at 405 and 488 nM, and the roGFP ratio. Additionally, the script prints the labeled and segmented nematodes overlaid on the original image.For quantification of GFP fluorescent intensity in strains SJ4100, SLR115, WY703, and CL2166, was achieved in a similar manner as described in the previous paragraph. However, because only one imaging channel is used for GFP fluorescence measurements, a modified version of the original script was used that evaluates fluorescent images only in one wavelength channel.
Rotenone lethality
To determine if bacterial diet impacted susceptibility to rotenone, day 4 adult nematodes (144 hours from L1 synchronization) were plated to K-agar plates seeded with bacteria (OD600 = 1) and 1% DMSO (vehicle) or rotenone (20 µM rotenone) for nematodes strains PE255, N2, QC115, and atfs-1(tm4525). For nuo-6(qm200) nematodes, we found that the 20 µM rotenone concentration used in previous experiments was lethal on the four bacterial strains by 24 hours (SI Fig. 10), and the concentration was reduced to 1 µM rotenone. Nematodes were scored every 24 hours for survival. Lethality was determined when no visible movement was present after gently prodding the nematode with a titanium wire.Complex I endogenously-tagged strain and measurements
For full description of strain creation, refer to Kenny-Ganzert et al. 2025.74 Briefly, CRISPR-Cas9 genome editing was used with self-excising hygromycin selection cassette to endogenously tag Complex I subunits NDUV-2 and NDUF-7 with mNeonGreen at the C-terminus. C. elegans were cultured as previously described, and after 72 hours on the selected CeMbio strains, adult nematodes were picked off plates and mounted onto microscope slides with 5% agar pads and anesthetized with 5 mM Levamisole (Millipore Sigma L9756). Z-stack images through the pharynx were captured using a Ziess Axioimager microscope equipped with a Yokogawa CSU-W1 and Hamamatsu ORCA-Quest qCMOS camera controlled by Micromanager Software v2.0.1. The z-stack images were then processed in ImageJ by z-projecting the stacks using sum slices projection type. Mean fluorescence intensity was measured in the terminal pump in the pharynx of the nematode manually using the oval brush selection. Imaging experiments were performed in triplicate.Targeted metabolomics
Nematodes were age synchronized using the same methods as stated above. L1s were grown on 15 cm K-agar plates with 700 µL of 2× OP50 at a density of 2500 nematodes per plate. After 3 days, nematodes were washed off, rinsed 5× with 50 mL K-medium to remove remaining OP50 bacteria. Approximately 1600 nematodes were then plated to K-agar plates seeded with 700 µL of MYb10, MYb11, BIGb0170, or OP50 bacteria at an OD = 2.19. Nematodes were transferred to new plates every 24 hours to ensure they had enough bacteria to not starve, and remove progeny. Plates were seeded two days prior to plating nematodes. After 3 days, nematodes washed and transferred to new 15 cm K-agar plates seeded with bacteria (OD600 = 1) and vehicle (DMSO) or 20 µM rotenone approximately 4 hours before nematodes were plated. After 24 hours, nematodes were washed off plates with K-medium into a 15 mL centrifuge tube and rinsed several times to remove bacteria and progeny via gravity settling. PBS was used for the final rinse. Nematodes were then transferred into a 2 mL Eppendorf tube, centrifuged at 1500 × g at 4C for 15 minutes. All supernatant was removed and 250 µL of 0.6% formic acid (Sigma 695![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 076) was added to the worm pellet and flash frozen in liquid nitrogen before being stored in −80 freezer. When samples were ready to be processed, they were thawed and sonicated on ice. Sonication was performed intermittently with 10 short pulses of 1–2 seconds to avoid overheating the sample. Once sonicated, a 10 µL aliquot was taken for protein quantification, and 240 µL of acetonitrile (VWR International 75-05-8) was added to the suspension and vortexed for one minute. Samples were then centrifuged again at 1500 × g for 15 minutes at 4 °C and supernatant was extracted for metabolite quantification and stored at −80 °C until analyzed.
076) was added to the worm pellet and flash frozen in liquid nitrogen before being stored in −80 freezer. When samples were ready to be processed, they were thawed and sonicated on ice. Sonication was performed intermittently with 10 short pulses of 1–2 seconds to avoid overheating the sample. Once sonicated, a 10 µL aliquot was taken for protein quantification, and 240 µL of acetonitrile (VWR International 75-05-8) was added to the suspension and vortexed for one minute. Samples were then centrifuged again at 1500 × g for 15 minutes at 4 °C and supernatant was extracted for metabolite quantification and stored at −80 °C until analyzed.
Amino acids, biogenic amines, and organic acids were measured by liquid chromatography-tandom mass spectrometry (LC-MS/MS). Acylcarnitines were measured using flow injection electrospray ionization tandem mass spectrometry. For full detailed methods, refer to Supplemental File 3.
Seahorse extracellular flux analyzer
Mitochondrial oxygen consumption rate (OCR) was measured using the Seahorse Xfe96 Extracellular Flux analyzer. Refer to Mello et al. for a detailed protocol.36 Briefly, after nematodes reached day 4 of adulthood, nematodes were rinsed off plates and rinsed thoroughly to remove remaining bacteria.10–20 adult nematodes were loaded into each well of the 96-well plate for measurement. Once OCR measurements were complete, each 96-well plate was imaged for exact counts of nematodes in each well for data normalization.Supplementation with pyruvate
K-agar plates were seeded with bacteria as previously described. 25 mM sodium pyruvate (Sigma P2256) was made fresh each time and added to bacteria prior to seeding 300 µL onto the agar plates at a final concentration of 2.5 mM. Plates were made 48–72 hours prior to nematodes being plated and kept at 20 °C to allow bacterial growth.Statistical analysis
All statistical analyses and graphs were done in RStudio (R version 4.2.1) unless otherwise stated. To test if the bacterial strain had a significant impact on the endpoint, we performed a one-way ANOVA followed by Dunnett's comparison to the MYb11 strain using the multcomp package. We performed a two-way ANOVA to test if there was a significant interaction between the bacterial strain and treatment followed by a Tukey's test using TukeyHSD.Author contributions
Christina M. Bergemann: conceptualization, investigation, data curation, methodology, formal analysis, validation, visualization, writing – original draft, writing – review & editing, Laura E. Jameson: investigation, visualization, writing – review & editing, Isabel W. Kenny-Ganzert: investigation, writing – review & editing, Javier Huayta: software, writing – review & editing, Pol Castellano-Escuder: formal analysis, Arunabh Sarkar: investigation, Olga R. Ilkayeva: investigation, writing – original draft, David R. Sherwood: resources, funding acquisition, writing – review & editing, Matthew D. Hirschey: formal analysis, funding acquisition, writing – review & editing, Joel N. Meyer: conceptualization, funding acquisition, supervision, resources, writing – original draft, writing – review & editing.Conflicts of interest
The authors declare no competing interests.Data availability
Data for this article, including targeted metabolomics data, are available at Duke University Digital Repository under Creative Commons BY Attribution 4.0 International at doi: https://doi.org/10.7924/r4p55v30k.Supplementary information (SI): Fig. S1: nematode length, volume, and mitochondrial to nuclear DNA ratio in C. elegans strain PE255, and whole body ATP and mitochondrial to nuclear DNA ratio in the germline deficient C. elegans strain PE327. Fig. S2: mitochondrial stress response in nematodes grown on CeMbio strains after 24 and 72 hours. Fig. S3: redox state using the JV2 C. elegans strain after exposure to hydrogen peroxide and rotenone. Fig. S4: targeted metabolomics of redox state on a subset of CeMbio strains. Fig. S5: mitochondrial respiration in C. elegans strains N2 and nuo-6. Fig. S6: complex I protein expression, whole-body ATP after DCA exposure, lifespan, and whole-body ATP levels after rotenone with pyruvate dose-response can be found. Fig. S7–S9: targeted metabolomics results for organic acids, acylcarnitines, amino acid and biogenic amine levels. Fig. S10: survival of nuo-6 nematodes after 20 µM rotenone and DMSO control. See DOI: https://doi.org/10.1039/d5em00624d.
Acknowledgements
This study was supported by NIH grants (R01ES034270, P42ES010356, T32ES021432, R35GM118049). We would like to thank Dr. Natasha Kirienko who gifted us the WY703 strain and Dr. Michael Shapira for feedback and edits to the manuscript. We'd also like to thank Duke University School of Medicine for the use of the Proteomics and Metabolomics Core Facility, which provided targeted metabolomics service.References
- T. Vezza, Z. Abad-Jiménez, M. Marti-Cabrera, M. Rocha and V. M. Víctor, Microbiota-mitochondria inter-talk: A potential therapeutic strategy in obesity and type 2 diabetes, Antioxidants, 2020, 9(9), 1–21 CrossRef PubMed.
- F. Borbolis, E. Mytilinaiou and K. Palikaras, The Crosstalk between Microbiome and Mitochondrial Homeostasis in Neurodegeneration, Cells, 2023, 12(3), 429 CrossRef CAS PubMed.
- D. N. Jackson and A. L. Theiss, Gut bacteria signaling to mitochondria in intestinal inflammation and cancer, Gut Microbes, 2020, 11(3), 285–304 CrossRef PubMed.
- Y. Saint-Georges-Chaumet and M. Edeas, Microbiota–mitochondria inter-talk: consequence for microbiota–host interaction, Pathog. Dis., 2016, 74(1), 96 CrossRef PubMed.
- B. Han, P. Sivaramakrishnan, C. C. J. Lin, J. Wang, C. Herman and M. C. Wang Correspondence, et al., Microbial Genetic Composition Tunes Host Longevity In Brief The genetic composition of gut microbes controls the production of metabolites that impact host longevity. Article Microbial Genetic Composition Tunes Host Longevity, Cell, 2017, 169, 1249–1262 CrossRef CAS PubMed.
- A. V. Revovich, R. Lee and N. V. Kirienko, Interplay between mitochondria and diet mediates pathogen and stress resistance in caenorhabditis elegans, PLoS Genet., 2019, 15(3), e1008011 CrossRef PubMed.
- S. F. Erttmann and N. O. Gekara, Hydrogen peroxide release by bacteria suppresses inflammasome-dependent innate immunity, Nat. Commun., 2019, 10(1), 1–13 CAS.
- J. A. Govindan, E. Jayamani, X. Zhang, E. Mylonakis and G. Ruvkun, Dialogue between E. coli free radical pathways and the mitochondria of C. elegans, Proc. Natl. Acad. Sci. U. S. A., 2015, 112(40), 12456–12461 CrossRef CAS PubMed.
- J. Zhang, X. Li, M. Olmedo, A. D. Holdorf, Y. Shang and M. Artal-Sanz, et al., A Delicate Balance between Bacterial Iron and Reactive Oxygen Species Supports Optimal C. elegans Development, Cell Host Microbe, 2019, 26(3), 400–411.e3 CrossRef CAS PubMed.
- B. S. Samuel, H. Rowedder, C. Braendle, M. A. Félix and G. Ruvkun, Caenorhabditis elegans responses to bacteria from its natural habitats, Proc. Natl. Acad. Sci. U. S. A., 2016, 113(27), E3941–E3949 CrossRef CAS PubMed.
- M. W. Pellegrino, A. M. Nargund, N. V. Kirienko, R. Gillis, C. J. Fiorese and C. M. Haynes, Mitochondrial UPR-regulated innate immunity provides resistance to pathogen infection, Nature, 2014, 516(7531), 414 CrossRef CAS PubMed.
- C. Kunst, S. Schmid, M. Michalski, D. Tümen, J. Buttenschön and M. Müller, et al., The Influence of Gut Microbiota on Oxidative Stress and the Immune System, Biomedicines, 2023, 11(5), 1388 CrossRef CAS PubMed.
- Z. Wang, G. W. Walker, D. C. G. Muir and K. Nagatani-Yoshida, Toward a Global Understanding of Chemical Pollution: A First Comprehensive Analysis of National and Regional Chemical Inventories, Environ. Sci. Technol., 2020, 54(5), 2575–2584 CrossRef CAS PubMed.
- R. Naidu, B. Biswas, I. R. Willett, J. Cribb, B. Kumar Singh and N. C. Paul, et al., Chemical pollution: A growing peril and potential catastrophic risk to humanity, Environ. Int., 2021, 156, 106616 CrossRef CAS PubMed.
- P. J. Landrigan, R. Fuller, N. J. R. Acosta, O. Adeyi, R. Arnold and N. Basu, et al., The Lancet Commission on pollution and health, Lancet, 2018, 391(10119), 462–512 CrossRef PubMed.
- E. S. Bernhardt, E. J. Rosi and M. O. Gessner, Synthetic chemicals as agents of global change, Front. Ecol. Environ., 2017, 15(2), 84–90 CrossRef.
- J. N. Meyer, J. H. Hartman and D. F. Mello, Mitochondrial Toxicity, J. Toxicol. Sci., 2018, 162(1), 15–23 CrossRef CAS PubMed.
- M. S. Attene-Ramos, R. Huang, S. Sakamuru, K. L. Witt, G. C. Beeson and L. Shou, et al., Systematic Study of Mitochondrial Toxicity of Environmental Chemicals Using Quantitative High Throughput Screening, Chem. Res. Toxicol., 2013, 26(9), 1323–1332 Search PubMed.
- S. Datta, S. Sahdeo, J. A. Gray, C. Morriseau, B. D. Hammock and G. Cortopassi, A high-throughput screen for mitochondrial function reveals known and novel mitochondrial toxicants in a library of environmental agents, Mitochondrion, 2016, 31, 79–83 CrossRef CAS PubMed.
- M. S. Attene -Ramos, R. Huang, S. Michael, K. L. Witt, A. Richard and R. R. Tice, et al., Profiling of the Tox21 Chemical Collection for Mitochondrial Function to Identify Compounds that Acutely Decrease Mitochondrial Membrane Potential, Environ. Health Perspect., 2015, 123(1), 49–56 CrossRef PubMed.
- Y. Zong, H. Li, P. Liao, L. Chen, Y. Pan and Y. Zheng, et al., Mitochondrial dysfunction: mechanisms and advances in therapy, Signal Transduct. Targeted Ther., 2024, 9(1), 124 CrossRef PubMed.
- D. R. Donohoe, N. Garge, X. Zhang, W. Sun, T. M. O’connell, M. K. Bunger, et al. The Microbiome and Butyrate Regulate Energy Metabolism and Autophagy in the Mammalian Colon. 2011 Search PubMed.
- H. Wen, H. Deng, B. Li, J. Chen, J. Zhu and X. Zhang, et al., Mitochondrial diseases: from molecular mechanisms to therapeutic advances, Signal Transduct. Targeted Ther., 2025, 10(1), 9 CrossRef PubMed.
- Y. Liu, B. S. Samuel, P. C. Breen and G. Ruvkun, Caenorhabditis elegans pathways that surveil and defend mitochondria, Nature, 2014, 508(7496), 406–410 CrossRef CAS PubMed.
- A. M. Schulz and C. M. Haynes, UPRmt-mediated cytoprotection and organismal aging, Biochim. Biophys. Acta, Bioenerg., 2015, 1847(11), 1448–1456 CrossRef CAS PubMed.
- T. Rudel, O. Kepp and V. Kozjak-Pavlovic, Interactions between bacterial pathogens and mitochondrial cell death pathways, Nat. Rev. Microbiol., 2010, 8(10), 693–705 CrossRef CAS PubMed.
- N. V. Kirienko, D. R. Kirienko, J. Larkins-Ford, C. Wählby, G. Ruvkun and F. M. Ausubel, Pseudomonas aeruginosa Disrupts Caenorhabditis elegans Iron Homeostasis, Causing a Hypoxic Response and Death, Cell Host Microbe, 2013, 13(4), 406–416 CrossRef CAS PubMed.
- P. Dirksen, A. Assié, J. Zimmermann, F. Zhang, A. M. Tietje and S. A. Marsh, et al., CeMbio - The Caenorhabditis elegans Microbiome Resource, Genes, Genomes.Genet., 2020, 401309 Search PubMed.
- F. Zhang, M. Berg, K. Dierking, M. A. Félix, M. Shapira and B. S. Samuel, et al., Caenorhabditis elegans as a Model for Microbiome Research, Front. Microbiol., 2017, 8, 485 Search PubMed.
- M. Shapira, Host–microbiota interactions in Caenorhabditis elegans and their significance, Curr. Opin. Microbiol., 2017, 38, 142–147 CrossRef CAS PubMed.
- J. Zhang, A. D. Holdorf and A. J. Walhout, C. elegans and its bacterial diet as a model for systems-level understanding of host–microbiota interactions, Curr. Opin. Biotechnol., 2017, 46, 74–80 CrossRef CAS PubMed.
- F. Cabreiro and D. Gems, Worms need microbes too: microbiota, health and aging in Caenorhabditis elegans, EMBO Mol. Med., 2013, 5(9), 1300–1310 CrossRef CAS PubMed.
- C. Backes, D. Martinez-Martinez, F. Cabreiro and C. elegans, A biosensor for host–microbe interactions, Lab. Anim., 2021, 50(5), 127–135 CrossRef PubMed.
- M. Ezcurra, Dissecting cause and effect in host-microbiome interactions using the combined worm-bug model system, Biogerontology, 2018, 19(6), 567–578 CrossRef CAS PubMed.
- S. L. Rea, N. Ventura and T. E. Johnson, Relationship Between Mitochondrial Electron Transport Chain Dysfunction, Development, and Life Extension in Caenorhabditis elegans, PLoS Biol., 2007, 5(10), 2312–2329 CrossRef CAS PubMed.
- D. F. Mello, L. Perez, C. M. Bergemann, K. S. Morton, I. T. Ryde and J. N. Meyer, Comprehensive characterization of mitochondrial bioenergetics at different larval stages reveals novel insights about the developmental metabolism of Caenorhabditis elegans, PLoS One, 2024, 19(11), e0306849 CrossRef CAS PubMed.
- S. K. Soo, A. Traa, P. D. Rudich, M. Mistry and J. M. Van Raamsdonk, Activation of mitochondrial unfolded protein response protects against multiple exogenous stressors, Life Sci. Alliance, 2021, 4(12), e202101182 CrossRef CAS PubMed.
- N. W. Kahn, S. L. Rea, S. Moyle, A. Kell and T. E. Johnson, Proteasomal dysfunction activates the transcription factor SKN-1 and produces a selective oxidative-stress response in Caenorhabditis elegans, Biochem. J., 2007, 409(1), 205–213 CrossRef PubMed.
- G. Detienne, P. V. de Walle, W. D. Haes, L. Schoofs and L. Temmerman, SKN-1-independent transcriptional activation of glutathione S-transferase 4 (GST-4) by EGF signaling, Worm, 2016, 5(4), e1230585 Search PubMed.
- K. Hasegawa, S. Miwa, K. Isomura, K. Tsutsumiuchi, H. Taniguchi and J. Miwa, Acrylamide-Responsive Genes in the Nematode Caenorhabditis elegans, Toxicol. Sci., 2008, 101(2), 215–225 CrossRef CAS PubMed.
- C. D. Link and C. J. Johnson, in. Reporter Transgenes for Study of Oxidant Stress in Caenorhabditis elegans, ed. C. K. Sen, L. Packer, Methods in Enzymology, Academic Press, 2002, vol. 353, p. 497–505, Redox Cell Biology and Genetics Part B, https://www.sciencedirect.com/science/article/pii/S007668790253072X Search PubMed.
- T. Yoneda, C. Benedetti, F. Urano, S. G. Clark, H. P. Harding and D. Ron, Compartment-specific perturbation of protein handling activates genes encoding mitochondrial chaperones, J. Cell Sci., 2004, 117(18), 4055–4066 CrossRef CAS PubMed.
- C. Benedetti, C. M. Haynes, Y. Yang, H. P. Harding and D. Ron, Ubiquitin-Like Protein 5 Positively Regulates Chaperone Gene Expression in the Mitochondrial Unfolded Protein Response, Genetics, 2006, 174(1), 229–239 CrossRef CAS PubMed.
- P. Back, W. H. De Vos, G. G. Depuydt, F. Matthijssens, J. R. Vanfleteren and B. P. Braeckman, Exploring real-time in vivo redox biology of developing and aging Caenorhabditis elegans, Free Radic. Biol. Med., 2012, 52(5), 850–859 CrossRef CAS PubMed.
- A. L. Luz, C. Lagido, M. D. Hirschey and J. N. Meyer, In vivo Determination of Mitochondrial Function using Luciferase-Expressing Caenorhabditis elegans: Contribution of Oxidative Phosphorylation, Glycolysis, and Fatty Acid Oxidation to Toxicant-Induced Dysfunction, Curr. Protoc. Toxicol., 2016, 69, 25.8.1–25.8.22 Search PubMed.
- A. R. Burns, G. M. Luciani, G. Musso, R. Bagg, M. Yeo and Y. Zhang, et al., Caenorhabditis elegans is a useful model for anthelmintic discovery, Nat. Commun., 2015, 6(1), 7485 CrossRef CAS PubMed.
- C. Lagido, J. Pettitt, A. Flett and L. A. Glover, Bridging the phenotypic gap: Real-time assessment of mitochondrial function and metabolism of the nematode Caenorhabditis elegans, BMC Physiol., 2008, 8(1), 7 CrossRef PubMed.
- M. Rauthan, P. Ranji, N. Aguilera Pradenas, C. Pitot and M. Pilon, The mitochondrial unfolded protein response activator ATFS-1 protects cells from inhibition of the mevalonate pathway, Proc. Natl. Acad. Sci. U. S. A., 2013, 110(15), 5981–5986 CrossRef CAS PubMed.
- A. M. Nargund, M. W. Pellegrino, C. J. Fiorese, B. M. Baker and C. M. Haynes, Mitochondrial Import Efficiency of ATFS-1 Regulates Mitochondrial UPR Activation, Science, 2012, 337(6094), 587–590 CrossRef CAS PubMed.
- W. Yang and S. Hekimi, Two modes of mitochondrial dysfunction lead independently to lifespan extension in Caenorhabditis elegans, Aging Cell, 2010, 9(3), 433–447 CrossRef CAS PubMed.
- E. L. Greer and A. Brunet, Different dietary restriction regimens extend lifespan by both independent and overlapping genetic pathways in C. elegans, Aging Cell, 2009, 8(2), 113–127 CrossRef CAS PubMed.
- G. D. Lee, M. A. Wilson, M. Zhu, C. A. Wolkow, R. de Cabo and D. K. Ingram, et al., Dietary deprivation extends lifespan in Caenorhabditis elegans, Aging Cell, 2006, 5(6), 515–524 CrossRef CAS PubMed.
- A. Dillin, A. L. Hsu, N. Arantes-Oliveira, J. Lehrer-Graiwer, H. Hsin and A. G. Fraser, et al., Rates of Behavior and Aging Specified by Mitochondrial Function During Development, Science, 2002, 298(5602), 2398–2401 CrossRef CAS PubMed.
- J. B. Spinelli and M. C. Haigis, The Multifaceted Contributions of Mitochondria to Cellular Metabolism, Nat. Cell Biol., 2018, 20(7), 745–754 CrossRef CAS PubMed.
- L. H. Sanders and G. J. Timothy, Oxidative damage to macromolecules in human Parkinson disease and the rotenone model, Free Radic. Biol. Med., 2013, 62, 111–120 CrossRef CAS PubMed.
- K. S. Morton, A. J. George and J. N. Meyer, Complex I superoxide anion production is necessary and sufficient for complex I inhibitor-induced dopaminergic neurodegeneration in Caenorhabditis elegans, Redox Biol., 2025, 81, 103538 CrossRef CAS PubMed.
- J. Fu, M. J. Bonder, M. C. Cenit, E. F. Tigchelaar, A. Maatman and J. A. M. Dekens, et al., The Gut Microbiome Contributes to a Substantial Proportion of the Variation in Blood Lipids, Circ. Res., 2015, 117(9), 817–824 CrossRef CAS PubMed.
- G. S. Raza, H. Putaala, A. A. Hibberd, E. Alhoniemi, K. Tiihonen and K. A. Mäkelä, et al., Polydextrose changes the gut microbiome and attenuates fasting triglyceride and cholesterol levels in Western diet fed mice, Sci. Rep., 2017, 7(1), 5294 CrossRef PubMed.
- Y. Ni, L. Qian, S. L. Siliceo, X. Long, E. Nychas and Y. Liu, et al., Resistant starch decreases intrahepatic triglycerides in patients with NAFLD via gut microbiome alterations, Cell Metab., 2023, 35(9), 1530–1547.e8 CrossRef CAS PubMed.
- H. T. E. Jarkass, S. Castelblanco, M. Kaur, Y. C. Wan, A. E. Ellis, R. D. Sheldon, et al., The Caenorhabditis elegans bacterial microbiome influences microsporidia infection through nutrient limitation and inhibiting parasite invasion [Internet]. bioRxiv; 2024, preprint bioRxiv:2024.06.05.597580, DOI:10.1101/2024.06.05.597580.
- A. Tauffenberger, H. Fiumelli, S. Almustafa and P. J. Magistretti, Lactate and pyruvate promote oxidative stress resistance through hormetic ROS signaling, Cell Death Dis., 2019, 10(9), 653 CrossRef PubMed.
- C. P. Gonzalez-Hunt, A. L. Luz, I. T. Ryde, E. A. Turner, O. R. Ilkayeva and D. P. Bhatt, et al., Multiple metabolic changes mediate the response of Caenorhabditis elegans to the complex I inhibitor rotenone, Toxicology, 2021, 447, 152630 CrossRef CAS PubMed.
- A. J. Worth, S. S. Basu, N. W. Snyder, C. Mesaros and I. A. Blair, Inhibition of Neuronal Cell Mitochondrial Complex I with Rotenone Increases Lipid β-Oxidation, Supporting Acetyl-Coenzyme A Levels, J. Biol. Chem., 2014, 289(39), 26895–26903 CrossRef CAS PubMed.
- S. S. Basu and I. A. Blair, Rotenone-Mediated Changes in Intracellular Coenzyme A Thioester Levels: Implications for Mitochondrial Dysfunction, Chem. Res. Toxicol., 2011, 24(10), 1630–1632 Search PubMed.
- E. Tjahjono, A. P. McAnena and N. V. Kirienko, The evolutionarily conserved ESRE stress response network is activated by ROS and mitochondrial damage, BMC Biol., 2020, 18(1), 1–17 CrossRef PubMed.
- Q. Hu, D. R. D'Amora, L. T. Macneil, A. J. M. Walhout and T. J. Kubiseski, The oxidative stress response in caenorhabditis elegans requires the GATA transcription factor ELT-3 and SKN-1/Nrf2, Genetics, 2017, 206(4), 1909–1922 CrossRef CAS PubMed.
- B. Tang, K. S. Xue, J. S. Wang, P. L. Williams and L. Tang, Bacteria pyruvate metabolism modulates AFB1 toxicity in Caenorhabditis elegans, Sci. Total Environ., 2023, 900, 165809 CrossRef CAS PubMed.
- B. Tang, K. S. Xue, J. S. Wang, P. L. Williams and L. Tang, Host-microbiota affects the toxicity of Aflatoxin B1 in Caenorhabditis elegans, Food Chem. Toxicol., 2023, 176, 113804 CrossRef CAS PubMed.
- S. Lee, Y. Kim and J. Choi, Effect of soil microbial feeding on gut microbiome and cadmium toxicity in Caenorhabditis elegans, Ecotoxicol. Environ. Saf., 2020, 187, 109777 CrossRef CAS PubMed.
- F. Cabreiro, C. Au, K. Y. Leung, N. Vergara-Irigaray, H. M. Cochemé and T. Noori, et al., Metformin retards aging in C. elegans by altering microbial folate and methionine metabolism, Cell, 2013, 153(1), 228–239 CrossRef CAS PubMed.
- D. Kim, S. El Khoury, O. M. Pérez-Carrascal, C. DeSousa, D. K. Jung and S. Bohley, et al., Gut microbiome remodeling provides protection from an environmental toxin, iScience, 2025, 28(4), 112209 CrossRef CAS PubMed.
- B. T. Moore, J. M. Jordan and L. R. Baugh, WormSizer: High-throughput Analysis of Nematode Size and Shape, PLoS One, 2013, 8(2), e57142 CrossRef CAS PubMed.
- T. C. Leuthner, J. H. Hartman, I. T. Ryde and J. N. Meyer, PCR-Based Determination of Mitochondrial DNA Copy Number in Multiple Species, Methods Mol. Biol., 2021, 2310, 91–111 CrossRef CAS PubMed.
- I. W. Kenny-Ganzert, L. P. Basta, L. Wang, Q. Chi, C. Su, K. S. Morton, et al., Specialized high-capacity mitochondria fuel cell invasion, bioRxiv, 2025, preprint bioRxiv:2025.05.02.651978, DOI:10.1101/2025.05.02.651978v1.
| This journal is © The Royal Society of Chemistry 2026 |