 Open Access Article
Open Access ArticleAtmospheric aerosol nucleation: a methodological review of theoretical calculations and molecular simulation
Yongjian Lian†
 ,
Xurong Bai†
,
Xurong Bai† ,
Ruoying Yuan†
,
Ruoying Yuan† ,
Tingyu Wei
,
Tingyu Wei ,
Hongjun Mao
,
Hongjun Mao ,
Jianfei Peng
,
Jianfei Peng and
Shuai Jiang
and
Shuai Jiang *
*
Tianjin Key Laboratory of Urban Transport Emission Research, State Environmental Protection Key Laboratory of Urban Ambient Air Particulate Matter Pollution Prevention and Control, College of Environmental Science and Engineering, Nankai University, Tianjin 300071, China. E-mail: shuaijiang@nankai.edu.cn
First published on 1st May 2026
Abstract
Atmospheric aerosols play a crucial role in large-scale precipitation, global climate change, and Earth's radiative balance, with new particle formation (NPF) constituting a major source of aerosol particles. NPF is a gas-to-particle phase transition process that involves both the formation of critical clusters—commonly referred to as nucleation—and the subsequent growth of these clusters into larger particles. Theoretical and computational approaches, including quantum chemistry calculations and molecular dynamics simulations, enable investigations of NPF at the microscopic molecular level and provide fundamental insights into the mechanisms governing cluster formation. When combined with cluster dynamics models, these methods further allow quantitative assessments of the atmospheric relevance of proposed nucleation mechanisms. In recent years, machine learning techniques have also emerged as powerful tools for accelerating and optimizing these workflows. In this review, we summarize recent theoretical and computational studies on aerosol nucleation mechanisms from the perspective of molecular clusters. Particular emphasis is placed on classical and representative applications of theoretical methods in nucleation research, including configurational sampling, thermodynamics, and cluster dynamics. We aim for this review to provide a comprehensive overview of the current progress in theoretical and computational nucleation studies, while also highlighting emerging challenges and future research directions in the field.
Environmental significanceAtmospheric new particle formation is a major source of aerosols that influence air quality, cloud properties, and climate forcing, yet its representation in environmental models remains highly uncertain. This review clarifies how modern theoretical and computational methods—ranging from quantum chemistry and molecular simulations to cluster dynamics and machine learning—contribute to a mechanistic understanding of aerosol nucleation at the molecular scale. By synthesizing methodological advances and linking microscopic cluster properties to macroscopic particle formation rates, this work provides a coherent framework for interpreting laboratory observations and improving process-level descriptions in atmospheric models. The review highlights key limitations of current approaches and identifies pathways toward more accurate, scalable, and environmentally relevant predictions of aerosol formation under realistic atmospheric conditions. |
1 Introduction
Atmospheric aerosols significantly impact Earth's radiative balance and human health.1,2 Notably, approximately 60–90% of the aerosol number concentration and 70–95% aerosol mass are formed through gas-to-particle conversion processes,3 a phenomenon known as new particle formation (NPF).4 In addition, the contribution of new particle formation (NPF) to atmospheric cloud condensation nuclei (CCN) is 50–60%,5,6 making it a major contributor to the substantial uncertainties in global climate models and large-scale precipitation models.7 Generally, the NPF process consists of an initial nucleation stage followed by a subsequent growth stage.8 During the nucleation stage, gas-phase molecules form stable molecular clusters (i.e., critical nuclei) through competing collision and evaporation processes, with characteristic sizes of approximately 1.5 nm. During the subsequent growth stage, these newly formed particles grow to larger sizes (typically ≥3 nm) via condensation, coagulation, and related processes. Obtaining key kinetic parameters and a detailed mechanistic understanding of both the nucleation and growth stages is therefore essential for elucidating NPF events.Field observations and laboratory simulations have substantially advanced our understanding of atmospheric NPF under diverse conditions through systematic investigations of its underlying mechanisms.9–12 The development and widespread application of high-resolution mass spectrometers and broad-range particle spectrometers have partially overcome a long-standing experimental barrier by enabling the detection of the composition and number concentrations of molecular clusters and particles smaller than 3 nm. Nevertheless, existing experimental techniques still lack the capability to accurately and quantitatively characterize the chemical composition of clusters and sub-3 nm particles. As a result, the nucleation and early growth mechanisms of these particles remain incompletely understood.
Theoretical approaches, including quantum chemical (QC) calculations, molecular dynamics (MD) simulations, and cluster dynamic simulations, provide access to structural, thermodynamic, and kinetic information at the molecular level and therefore constitute indispensable tools for elucidating the formation processes of sub-3 nm particles.13–15 In recent years, theoretical investigations of NPF have emerged as a major focus in nucleation research.16,17 Kinetic nucleation models which solve the cluster birth–death equation,12,18–20 macroscopic nucleation kinetics and cluster concentration distributions can be derived from microscopic cluster structures and their associated thermodynamic properties. However, atmospheric nucleation is strongly modulated by environmental factors such as temperature, humidity, and precursor concentrations, leading to highly complex cluster structures and thermodynamic landscapes that pose substantial challenges for theoretical modeling. Fortunately, these challenges are being progressively addressed through advances in QC calculations, MD simulations, and artificial intelligence-based approaches.21–24
It is worth noting that several excellent reviews have focused on the application of existing theoretical methods to specific nucleation chemical systems.13–15,25 Therefore, the present article does not address individual systems or their atmospheric implications. Instead, this review summarizes recent progress in theoretical methods for nucleation research and highlights how these developments have addressed key challenges in the field. As illustrated in the Fig. 1, this review is organized into three main sections, which closely link the microscopic properties of clusters with the macroscopic properties of particulate matter. Section 2 discusses configurational sampling techniques, which are essential for identifying global minimum-energy cluster structures and thus determine the reliability of subsequent QC-based thermodynamic and kinetic analyses, including MD simulations, optimization algorithms, and specialized tools. Section 3 presents the principal theoretical frameworks for calculating cluster thermodynamics. Section 4 focuses on solving birth–death equations and its integration with other molecular-scale models or air quality models. Through this integrated perspective, the review not only synthesizes advances in computational approaches but also identifies remaining knowledge gaps and outlines directions for future research.
2 Configurational sampling techniques
This section provides a concise overview of configurational search methods used to identify the global minimum structures of clusters. Configurational search aim to explore the cluster potential energy surface (PES) and determine the global minimum structures that serve as the foundation for subsequent static QC calculations. The general strategy typically involves two key stages: first, generating a diverse set of initial configurations to explore the rough PES landscape using sampling techniques; second, refining these configurations through QC calculations to progressively locate minima on the accurate PES until the global minimum is identified. With increasing cluster size in atmospheric aerosol nucleation, the number of possible configurations grows exponentially. This rapid expansion renders exhaustive enumeration based solely on chemical intuition impractical and necessitates a greater reliance on advanced sampling algorithms. In nucleation research, commonly employed sampling techniques include MD simulations and metadynamics, as well as intelligent optimization algorithms such as Basin Hopping (BH), Artificial Bee Colony (ABC) and Genetic Algorithm (GA).2.1 Molecular dynamics
MD simulations address the sampling challenge by employing force fields to describe intermolecular interactions and by integrating Newton's equations of motion to sample a diverse ensemble of cluster configurations. For instance, Loukonen et al.26 investigated the interactions of sulfuric acid (SA) with ammonia (AM) and dimethylamine (DMA) under hydrated conditions using a combination of MD sampling and QC calculations. For previously unknown cluster structures, the configurational space was expanded via MD simulations followed by a three-step simulated annealing protocol at 1500 K, 200 K, and 0.1 K, where the relaxed low-temperature structures retained as additional initial candidates. Subsequent QC calculations revealed that DMA enhances SA addition to clusters more effectively than AM when the number of water (W) molecules is either zero or greater than two. Similarly, Temelso et al.27 have employed MD simulations to explore the configurational landscape of H2SO4(H2O)n (n = 4–6) clusters. Their global configurational search comprises two stages: a heating phase and a production phase. The heating phase aims to sufficiently relax the cluster structure by gradually increasing the temperature from 5 K to a final temperature (Tf), thereby promoting adequate sampling in the subsequent production phase. Notably, the choice of Tf should be size-dependent. During the production phase, a 10 ns simulation is performed at Tf, from which 200 structures are extracted using the ptraj module for subsequent QC calculations.However, conventional MD simulations are prone to becoming trapped in local low-energy minima, making it difficult to cross potential energy barriers and explore the global minimum structure. Although high-temperatures MD simulations can enhance configurational sampling, they cannot guarantee that all relevant structures are adequately accessed. Metadynamics addresses this limitation by introducing a history-dependent bias potential into MD simulations, thereby facilitating the exploration of high-dimensional potential energy surfaces (PES).28 The external (‘metadynamics’) potential acting on the system at time t is given by:
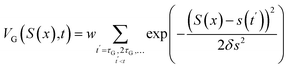 | (1) |
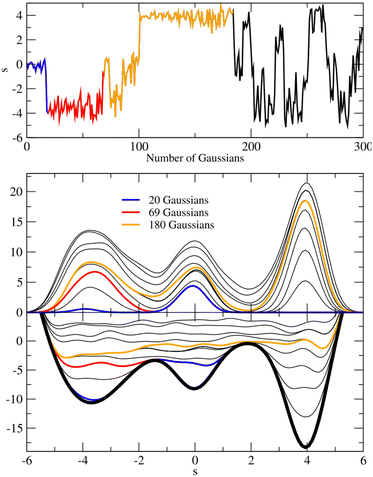 | ||
| Fig. 2 (Upper panel) Trajectory of a one-dimensional system evolved by a Langevin equation on the 3-minima potential represented in the lower panel. The dynamics is biased with a metadynamics potential VG as defined by eqn (1). The parameters are δs = 0.4, w = 0.3 and τG = 300. (Middle panel) time evolution of the metadynamics bias potential VG. Blue line: VG as when the first minimum is filled and the system ‘escapes’ to the second minimum; red line: VG as when also the second minimum is filled; orange line: VG when the entire profile is filled and the dynamics becomes diffusive. (Lower panel) Time evolution of the sum of the metadynamics potential VG and of the external potential, represented as a thick black line. Adapted from Laio et al., 2008.28 Adapted from ref. 28 with permission from Elsevier, copyright 2008. Reproduced from ref. 28 from © IOP Publishing, copyright 2008. Reproduced with permission. All rights reserved. | ||
2.2 Optimization algorithm
Basin-Hopping (BH) method is a global optimization algorithm designed to escape local minima and identify global minimum-energy structures through a combination of random perturbations and local minimizations. It relies on Monte Carlo (MC) simulations, which employ probabilistic sampling based on statistical mechanics. Jiang et al.31 applied the BH method to investigate the structural and energetic properties of Cl−(H2O)n (n = 1–4) clusters and emphasized the critical influence of temperature on the efficiency of the global search. In BH algorithm, diverse candidate structures are generated through successive MC steps initiated from a given structure. Each MC step involves perturbations of cluster geometry via translational moves (with a maximum distance displacement of 2 Å) and rotational (with a maximum angular displacement of 90°) moves, followed by local energy minimization. The effectiveness of the global search is strongly influenced by several key parameters, including the number of independent BH runs, the number of MC steps per run, and the simulation temperature. In practice, the number of independent runs generally increases with cluster size to ensure adequate sampling of the configurational space. First, many initial hydration arrangements are generated. At each step, candidate structures are produced by randomly changing the positions of water molecules and the orientations of hydrogen bonds. Each candidate is then directly relaxed to a stable configuration via DFT geometry optimization, and the optimized energy is used for comparison. Lower-energy structures are accepted, while higher-energy ones are also accepted with a certain probability to avoid being trapped in local minima. A distinctive feature of this work is that every hop is evaluated at the DFT level rather than with an empirical force field, and a consistent global search is carried out for n = 1–4, thereby systematically revealing low-energy isomers and how hydrogen-bonding motifs evolve with hydration number. Conventional Monte Carlo simulations often suffer from trapping in local energy minima, which limits efficient sampling of low-energy cluster configurations. Basin Paving Monte Carlo (BPMC), an extension of Basin Hopping Monte Carlo, overcomes this limitation by incorporating an energy-histogram-based bias on a transformed potential energy surface, thereby enhancing global exploration of stable structures.32 Using BPMC, Xu et al.33 demonstrated that hydration promotes proton transfer from succinic acid (SUA) to dimethylamine (DMA) while simultaneously weakening SA–DMA interactions. Notably, stable aminium–carboxylate ion pairs form only in the presence of more than three water molecules, with hydrated DMA clusters adopting ring- or cage-like hydrogen-bonded structures that stabilize the ionic products.ABC algorithm is inspired by the collective foraging and nest-building behaviors of natural bee swarms and is designed to identify optimal solutions in high-dimensional search spaces. In 2015, Zhang et al. developed the ABCluster program based on the ABC algorithm34,35 and systematically evaluated its performance in locating global and local minimum-energy structures across a wide range of cluster systems. In the context of cluster structure searches, low-energy structures are treated as “optimal food sources”. As illustrated in Fig. 3, ABC algorithm employs three types of agents—employed bees (EM), onlooker bees (OL), and scout bees (SC)—that cooperatively explore the potential energy surface to identify the global minimum (GM). ABCluster is also applicable to rigid molecular clusters such as water, methanol, non-polar molecules.35 The performance of ABCluster is governed by several key parameters, including the population size (SN), scout limit, and maximum number of generations, all of which strongly influence both the efficiency of the configurational search and the associated computational cost. The selection of these parameters was benchmarked by Wu et al.36 They applied optimized parameters (SN = 1280, number of generations gen = 320, scout limit sc = 4, and saving the 1000 lowest-energy structures) to search for the global minimum structures of (SA)n–(DMA)n clusters (n = 0–6), where SA and DMA represent sulfuric acid and dimethylamine, respectively. Using these settings, ten independent parallel runs were found to yield a structural ensemble that comprehensively captured the global minimum configurations.
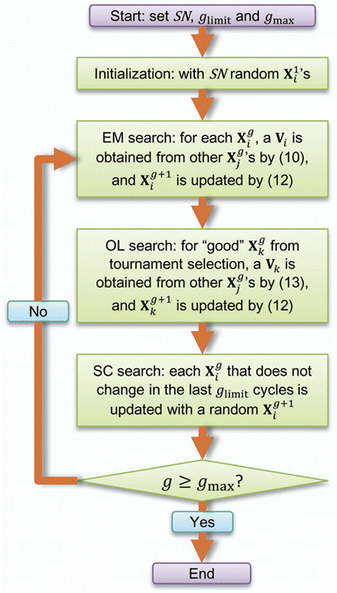 | ||
| Fig. 3 The ABC algorithm for searching the GM of clusters used in ABCluster. EM: employed bees; OL: onlooker bees; and SC: scout bees. Adapted from Zhang et al., 2015.34 Adapted from ref. 34 with permission from the Royal Society of Chemistry, copyright 2015. Permission conveyed through Copyright Clearance Center. | ||
In addition, GA algorithm, which mimics biological evolutionary processes such as selection and crossover to navigate complex configurational landscapes, is particularly effective for exploring intricate potential energy surfaces. Temelso et al.37 employed GA to investigate the influence of mixed bases (ammonia, methylamine, dimethylamine, trimethylamine) on sulfate aerosol formation. GA algorithm is implemented in OGOLEM and CLUSTER packages. The GA workflow comprises three main steps: (1) population initialization, in which 250–1000 initial cluster structures are generated based on atomic coordinates, bond lengths, or angular constraints; (2) crossover optimization and semiempirical refinement, where crossover operations are combined with semiempirical methods (PM7, SCC-DFTB, and EFP2) for efficient energy evaluation; and (3) structure selection, in which low-energy configurations are identified after approximately 5000–30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 crossover cycles.
000 crossover cycles.
Additionally, Kildgaard et al.38 introduced a systematic sampling methodology for large hydrated sulfuric acid clusters, SA-(H2O)n (n = 1–15). Their approach consists of four sequential stages: (1) geometric arrangement, in which sampling points are generated on a Fibonacci sphere surrounding all atoms; (2) orientation enumeration, whereby a water molecule is placed at each sampling point in nine distinct orientations. (3) hydrogen-bond (H-bond) insertion, which generates additional configurations by perturbing water molecules along vectors extrapolated from existing H-bond donor and acceptor sites; and (4) structure deduplication, in which root-mean-square deviations (RMSDs) are computed for all generated configurations, and structures with RMSD < 0.38 Å are considered identical and merged. However, this exhaustive enumeration approach may fail to accurately account for the steric effects induced by alkyl groups in organic nucleation precursors, leading to substantial computational waste due to the evaluation of numerous unnecessary structures.
2.3 Specialized tools
The QC refinement and screening procedures following configurational sampling vary substantially among different studies. This lack of standardization can result in time-consuming workflows and, in some cases, inaccurate identification of the true global minimum structures. To address this issue, Kubečka et al.39 conducted rigorous evaluations and proposed a robust, standardized automated workflow termed Jammy Key for Configurational Sampling (JKCS). The JKCS framework has been expanded into distinct modules: JKCS, JKQC, JKML and JKTS. Moreover, JKCS integrates seamlessly with widely used computational tools such as ABCluster, CREST, Gaussian, ORCA, xTB, and interfaces directly with the Slurm high-performance computing workload manager, enabling highly automated and user-friendly execution on large-scale computing platforms. As illustrated in Fig. 4, transitions between stages can be performed using single-line commands, highlighting the exceptional usability of the framework. Moreover, Temelso et al.40 developed a tool employing the Kuhn–Munkres algorithm to efficiently compute RMSD between isomers with arbitrary atom orderings, enabling rapid alignment of diverse molecular clusters; it is also integrated into JKCS. Over the past several years, JKCS has become a vital tool in atmospheric nucleation research and continues to evolve through the incorporation of new computational techniques. | ||
| Fig. 4 JKCS workflow proposed by Kubečka et al., 2023.39 Adapted from ref. 39 with permission from the American Chemical Society, copyright 2023. | ||
3 Cluster thermodynamics
The thermodynamic properties of nucleating clusters constitute the microscopic foundation of their macroscopic atmospheric behavior. By integrating these properties with cluster dynamics and subsequent atmospheric model simulations, it is possible to elucidate the roles and environmental significance of such clusters in the real atmosphere. This section briefly summarizes recent advances in theoretical approaches for determining key cluster thermodynamic properties.3.1 QC methods benchmarking
For the weak hydrogen-bonded complex, Bork et al. evaluated several DFT methods (PW91, M06-2X, B3LYP-D3, and ωB97X-D) using FTIR spectroscopy for the acetonitrile–HCl complex.43 They showed that only when CC2 is used for single-point corrections do the computed structures and Gibbs free energies agree with experiment, whereas other methods (DFT, MP2, F12, and CCSD(T)) tend to overestimate binding energies. Hansen et al. further assessed B3LYP, ωB97XD and M06-2X for harmonic vibrational frequencies of alcohol–amine complexes and found that B3LYP/aug-cc-pVTZ provides the best agreement with experiment.44 A systematic review by Gadre et al. highlighted that, although DFT is suitable for large clusters, its accuracy is limited by BSSE and functional dependence, and no universal “gold standard” functional has yet been established for dispersion-dominated systems.45 In addition, Hansen et al. showed that B3LYP-D3/aug-cc-pVTZ yields the most accurate Gibbs free energy for the MeOH·DMA complex.46 More recently, Knattrup et al. demonstrated that combining ωB97X-D3BJ/ma-def2-SVP or B97-3c with anharmonic frequency scaling, multi-conformer entropy corrections, and high-level single-point energies at Normal LNO-CCSD(T)/CBS(aug-3, aug-4) achieves sub-1 kcal mol−1 accuracy for experimental data of 11 hydrogen-bonded systems.47
It is worth noting that Nadykto et al. and Kupiainen-Määttäet et al. conducted theoretical studies on the H2SO4–H2O nucleation system involving DMA using PW91PW91/6-311++G (3df, 3pd) and B3LYP//RI-CC2, respectively.49,50 Their results were contradictory. The RI-CC2 method, which has a higher theoretical level, shows poorer agreement with experimental values than functionals such as PW91, highlighting the importance of experimental thermodynamic data for nucleation and the necessity of benchmarking against experimental data.
Elm et al.51 systematically evaluated the performance of 22 density-functional theory (DFT) functionals paired with aug-cc-pVDZ, aug-cc-pVTZ, aug-cc-pV(D+d)Z, aug-cc-pV(T+d)Z, and the 6-311++G(3df,3pd) basis sets for predicting the structures and reaction free energies of atmospheric prenucleation clusters formed by sulfuric acid, water and ammonia. The computed results were benchmarked against experimental data and reference calculations at the gold-standard CCSD/CBS level. Among the tested combinations, the M06-2X functional paired with the 6-311++G(3df,3pd) basis set exhibited consistently high accuracy across all evaluated properties. However, the M06-2X/6-311++G(3df,3pd) combination remains computationally demanding, which limits its direct applicability in large-scale configurational searches. To address this limitation, Elm et al.52 further explored computationally efficient combination of basis sets and DFT methods by analyzing Gibbs free energy thermal contributions and single-point energy performances of generalized density functional theory methods (M06-2X, PW91, ωB97X-D) together with various basis sets (6-311++G(3df,3pd), 6-31++G(d,p), etc.) on a test set of 205 atmospheric clusters. The 6-31++G(d,p) basis set was found to yield the smallest errors in both Gibbs free energy and single-point energy calculations while maintaining high computational efficiency. Notably, the performance of the DFT functionals exhibited a clear dependence on cluster composition: M06-2X showed relatively larger errors for purely inorganic clusters but delivered the highest accuracy for systems containing organic species. These findings indicate that the optimal choice of DFT functional may be system dependent, underscoring the need for further comprehensive benchmarking across broader and more diverse cluster datasets. Although the M06-2X/6-311++G(3df,3pd) approach discussed above provides relatively good accuracy in predicting Gibbs free energy changes for prenucleation reactions, its performance in estimating binding energies for larger nucleation systems can deviate by as much as 1.8 kcal mol−1.53 High-level ab initio methods such as MP2 and CCSD(T) are widely regarded as benchmarks for energetic accuracy; however, their application to larger systems is hampered by basis set incompleteness error (BSIE) when finite basis sets are employed. Explicitly correlated F12 methods address this limitation by incorporating r12 operator into the wavefunction, thereby improving the convergence of the correlation energy and substantially reducing BSIE. The efficacy of F12 approaches for nucleation-related energy calculations has been demonstrated in previous work,53 which showed that inclusion of F12 corrections significantly improves basis set convergence, reduces reliance on very large basis sets, and enables high-precision binding energy calculations for strongly hydrogen-bonded clusters.
Nevertheless, even with the incorporation of F12 corrections that allow MP2 and CCSD(T) methods to approach the complete basis set (CBS) limit, their computational cost remains prohibitively high for large cluster systems. Domain-based local pair natural orbital (DLPNO) methods provide an effective solution by substantially reducing computational cost through sparsification of the orbital space—such as compression of the virtual orbital space using pair natural orbitals (PNOs)—while treating the system as a whole and maintaining high accuracy. The performance of DLPNO method combined with MP2, CCSD(T0), and F12 corrections has been systematically evaluated for the binding energies of 45 representative atmospheric cluster systems.54 The DLPNO-CCSD(T0) method combined with triple-zeta basis set was shown to closely reproduce CCSD(T)/CBS results. Importantly the computational time of the DLPNO-CCSD(T0)/double-zeta combination is comparable to that of commonly used DFT approaches such as M06-2X/aug-cc-pVTZ and ωB97X-D(BJ)/aug-cc-pVTZ. Although DLPNO-CCSD(T0)/aug-cc-pVTZ has been widely adopted in nucleation studies, its high computational cost still limits its practical application to clusters containing on the order of ∼20 molecules.55 More recently, Knattrup et al.56 demonstrated that the local natural orbital (LNO) method, when extrapolated to its Local Approximation Free (LAF) limit, can achieve accuracy comparable to CCSD(T)/CBS. They benchmarked both LNO-CCSD(T) and DLPNO-CCSD(T0) methods—together with different basis sets and extrapolation schemes (CBS, LAF, CPS)—on a set of 218 atmospheric dimer clusters. LNO-CCSD(T) exhibited a superior accuracy-to-cost ratio relative to DLPNO-CCSD(T0) and proved sufficiently efficient to enable high-accuracy binding energy calculations for large systems, such as (SA)15(TMA)15 comprising approximately 300 atoms. In order to find a calculation method that simultaneously balances high precision and low power consumption, Engsvang et al.57 further assessed the accuracy of the semi-empirical methods (GFN1-xTB, GFN2-xTB) for optimizing SA-AM cluster structures, as well as the performance of B97-3c and PBEh-3c in predicting the binding energy of (SA)20(AM)20 cluster. GFN1-xTB was found to perform well for cluster structure optimization, yielding geometries closest to those obtained with DFT methods, but exhibited poor accuracy in binding energy predictions. In contrast, B97-3c achieved binding energy accuracies comparable to DLPNO-CCSD(T0)/aug-cc-pVTZ and aug-cc-pVDZ, and even outperformed ωB97X-D/6-31++G(d,p).
In recent years, Machine Learning (ML) techniques have advanced rapidly and have been widely applied in theoretical studies of nucleation. However, the generation of high-quality training datasets—comprising diverse cluster conformations with accurately computed energies—remains computationally demanding, underscoring the continued need for electronic structure methods that balance high accuracy with low computational cost. In this context, semiempirical, DFT-3c, and DFT methods have been systematically evaluated in terms of energy prediction accuracy, computational efficiency, and their suitability for training Δ-ML models using a large dataset of 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 749 cluster conformations.58 GFN1-xTB was found to be particularly effective for rapid configurational screening, whereas r2SCAN-3c delivered the highest accuracy in direct energy predictions. Moreover, Δ-ML models trained on high-level datasets labeled at the r2SCAN-3c level achieved the best overall performance. These results demonstrate that r2SCAN-3c provides an optimal balance between accuracy and computational efficiency and is therefore especially suited for generating training data for ML-based nucleation studies.
749 cluster conformations.58 GFN1-xTB was found to be particularly effective for rapid configurational screening, whereas r2SCAN-3c delivered the highest accuracy in direct energy predictions. Moreover, Δ-ML models trained on high-level datasets labeled at the r2SCAN-3c level achieved the best overall performance. These results demonstrate that r2SCAN-3c provides an optimal balance between accuracy and computational efficiency and is therefore especially suited for generating training data for ML-based nucleation studies.
The accuracy of Gibbs free energy (G) calculations is inherently sensitive to the underlying molecular geometries. Variations in optimized geometries obtained using different electronic structure methods can thus introduce non-negligible errors into G, yet the practical implications of such errors for cluster thermodynamics remain incompletely understood. Jensen et al. systematically assessed the performance of semi-empirical, DFT, and DFT-3c methods for geometry optimization across a dataset of 1283 acid–base clusters.24 In these benchmarks, r2SCAN-3c, ωB97X-3c, and ωB97X-D3BJ exhibited the highest accuracy among DFT-based approaches, while AMC-xTB performed best among the semiempirical methods. Importantly, DFT-3c methods were found to be substantially faster than both DF-MP2 and ωB97X-D3BJ, underscoring their favorable balance between accuracy and computational efficiency for large-scale configurational screening.
3.2 ML acceleration
ML can reconstruct PES of clusters by learning from datasets that contain accurate structural coordinates and corresponding energies, provided that the configurational space is sampled sufficiently and comprehensively. This capability enables ML to predict cluster thermodynamic properties with high accuracy at computational speeds that far exceed those of conventional QC calculations. As such, ML represents a powerful and promising approach for future nucleation research. In addition, Δ-ML technique, which correct lower-level predictions using higher-level reference data, have been explored in the context of nucleation studies and have been shown to further enhance the accuracy of ML-based models.
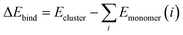 | (2) |
| ΔΔEbind = ΔEDFTbind − ΔEPM7bind | (3) |
The core idea of Δ-ML is to learn the discrepancy in binding energy predictions between DFT and semi-empirical method. In this way, DFT-level binding energies can be efficiently approximated using semi-empirical calculations as baseline, leading to substantial improvements in both computational efficiency and predictive accuracy. Kubečka et al. employed kernel ridge regression (KRR) in combination with Δ-ML to train a ML model on a dataset generated at the semi-empirical level for the sulfuric acid hydration systems, and systematically evaluated the model's extrapolation capability.22 As shown in Fig. 5, GFN1-xTB provided the highest accuracy among the tested semiempirical methods when used as the baseline for Δ-ML training. Moreover, the trained model achieved binding energy predictions with MAE below 0.5 kcal mol−1 for smaller clusters. For larger clusters (e.g., (SA)7(H2O)10), the extrapolation error increased slightly (>1.0 kcal mol−1) but remained significantly lower than that obtained from the underlying semi-empirical method.
 | ||
| Fig. 5 ML curves for four different learning models. The equilibrium cluster structure database were used (compare the direct-ML and ΔPM7-ML curves). Note the logarithmic axes. Adapted from Kubečka et al., 2022.59 Reproduced from ref. 59 with permission from the American Chemical Society, copyright 2022. | ||
They further extended ML applications to a wide range of nucleation systems and cluster sizes using Δ-ML in combination with neural network approach.22 In particular, the Polarizable Atomic Interaction Neural Network (PaiNN),60 a message-passing network specifically designed for three-dimensional molecular structures, was adopted. The NN model achieved accuracy comparable to that of KRR while exhibiting superior computational efficiency for large clusters. As shown in Fig. 6, the NN model also demonstrated excellent generalization capability, achieving an accuracy of approximately 0.6 kcal mol−1 for much larger cluster such as (SA)15(AM)15. Unlike conventional “black box” ML methods such as neural networks and KRR, the k-nearest neighbor (k-NN) model generates predictions based on similarities to existing data instances, thereby offering enhanced interpretability and reliable uncertainty estimates while maintaining an accuracy of approximately 1 kcal mol−1.61 Notably, the Metric Learning for Kernel Regression (MLKR)-based k-NN model reduced computational cost to one-third of that required by KRR. For example, inference over 26![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 775 test points required approximately 57
775 test points required approximately 57![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 CPU seconds using k-NN, compared with 162
000 CPU seconds using k-NN, compared with 162![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 CPU seconds for KRR. These encouraging results strongly indicate that ML methodologies have the potential to drive transformative advances in future nucleation studies.
000 CPU seconds for KRR. These encouraging results strongly indicate that ML methodologies have the potential to drive transformative advances in future nucleation studies.
 | ||
| Fig. 6 Absolute error distributions of the modelled energies and force components for the SAnAMn clusters trained at the B97-3c level. The error bars show the standard deviation. The error bars show the standard deviation. Adapted from Kubečka et al., 2024.22 Adapted from ref. 22 with permission from the Royal Society of Chemistry, copyright 2024. Permission conveyed through Copyright Clearance Center. | ||
Recently, the concept of a universal potential, or foundation model, has attracted increasing attention. ANI-2x, the extended version of ANI-1x, is applicable to the molecules containing C, H, O, N S, F, and Cl elements and has demonstrated good performance for organic molecules, reaction pathways, and noncovalent interactions. Jiang et al.30 benchmarked the ANI-2x neural network potential for predicting the thermodynamic properties of atmospheric clusters. For relative energy prediction and geometry optimization of both low- and high-energy isomers, ANI-2x generally outperformed the PM7 semiempirical method, with the exception of the (SA)1(DMA)1 and (SA)2 systems. In force predictions, ANI-2x exhibited higher root-mean-square errors than PM7 but showed stronger linear correlations with DFT reference data (R2 > 0.90). Overall, although ANI-2x provides improved accuracy relative to conventional semiempirical methods, it has not yet achieved chemical accuracy (∼1 kcal mol−1) and remains limited in force prediction accuracy and general applicability across diverse nucleation systems.
The real atmosphere contains an extremely complex mixture of molecular species, and even with the aid of QC calculations and ML techniques, it remains challenging to comprehensively evaluate the nucleation potential and cluster thermodynamics of all relevant compounds. Consequently, considerable research efforts have been devoted to screening effective nucleating precursor molecules using QC-based methods in combination with quantitative structure–activity relationship (QSAR) analysis. For example, Xie et al.,62 combined QC calculations with a QSAR model to evaluate the free energy (ΔG) of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 dimer formation between 50 amines and methanesulfonic acid (MSA). Based on the resulting QSAR model, they further predicted ΔG values for an additional 145 atmospherically amines and identified candidates with strong binding affinities. As illustrated in Fig. 7, guanidine was identified as the most effective organic amine, and the MSA–guanidine system was predicted to exhibit a higher nucleation rate than the previously proposed MSA–MEA system.
1 dimer formation between 50 amines and methanesulfonic acid (MSA). Based on the resulting QSAR model, they further predicted ΔG values for an additional 145 atmospherically amines and identified candidates with strong binding affinities. As illustrated in Fig. 7, guanidine was identified as the most effective organic amine, and the MSA–guanidine system was predicted to exhibit a higher nucleation rate than the previously proposed MSA–MEA system.
 | ||
| Fig. 7 Variation of simulated steady-state MSA dimer concentration ∑[(MSA)2] (molecules per cm3) (A) and cluster formation rate J (cm−3 s−1) (B) with monomer concentration at 278.15 K and CoagS = 2.6 × 10−3 s−1. Adapted from Liu et al., 2022.62 Reproduced from ref. 62 with permission from the American Chemical Society, copyright 2022. | ||
Besides, Pedersen et al.63 screened promising nucleation precursors by correlating molecular functional groups with cluster stability, with particular emphasis on strong hydrogen-bonding interactions within clusters. Their results demonstrated that clusters containing multiple carboxyl (–COOH) groups are substantially more stable than those bearing other functional groups. This enhanced stability arises from the dual hydrogen-bonding capability of –COOH moiety, which can act as both hydrogen-bond donors (via –OH) and acceptors (via C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O). In addition, a hypothetical Oxygenated Organic Molecule (OOM) structure incorporating multiple –COOH groups was designed, and its formation rate potential (Jpotential) was evaluated based on the thermodynamic properties of sulfuric acid–base-OOM clusters. The calculated Jpotential increased with the concentration of polycarboxylated OOMs, indicating that such OOMs can further stabilize initial SA-base clusters by forming dense hydrogen-bonding networks and may facilitate continued particle growth.
O). In addition, a hypothetical Oxygenated Organic Molecule (OOM) structure incorporating multiple –COOH groups was designed, and its formation rate potential (Jpotential) was evaluated based on the thermodynamic properties of sulfuric acid–base-OOM clusters. The calculated Jpotential increased with the concentration of polycarboxylated OOMs, indicating that such OOMs can further stabilize initial SA-base clusters by forming dense hydrogen-bonding networks and may facilitate continued particle growth.
3.3 Anharmonicity effect
QC calculations constitute a cornerstone of theoretical nucleation studies; however, the widely adopted harmonic approximation is inherently unable to capture vibrational anharmonicity in cluster dynamics. Such anharmonicity arises from both local effects, associated with intramolecular and intermolecular vibrations within a single conformation, and global effects, stemming from the coexistence of multiple low-energy conformers. Despite their potential importance, anharmonic contributions are largely neglected in current studies of atmospheric clusters. Nevertheless, early work by Kathmann et al.64 demonstrated that an explicit treatment of anharmonicity is essential for obtaining accurate thermodynamic properties, particularly for aqueous ionic clusters. Moreover, Partanen et al.65 reported high-accuracy thermodynamic properties for the (SA)1(H2O)1 system using CCSD(T)-F12b/VQZ-F12//DF-MP2-F12/VDZ-F12 computational protocols and systematically examined the impact of vibrational anharmonicity. Their results showed that both local and global anharmonicity exert non-negligible effects on the reaction free energy (ΔG), enthalpy (ΔH), and entropy (ΔS) associated with sulfuric acid–water complex formation. Notably, these contributions were found to be highly sensitive to the underlying electronic-structure treatment and basis-set quality, as summarized in Table 1. Local anharmonicity within a single conformation can be treated using Morse potentials or vibrational perturbation approaches such as VPT2 or HDCPT2, whereas global anharmonicity, arising from the presence multiple conformations, must be captured through statistical-mechanical averaging.| JV(T+d)Z | A'V(T+d)Z | |||||
|---|---|---|---|---|---|---|
| Harm | Anharm | Anharm + AD | Harm | Anharm | Anharm + AD | |
| a For the anharm and anharm + AD calculations, the vibrational partition functions were calculated with the SPT approximation using HDCPT2 with MP2 for all and the passive vibrational degrees of freedom, respectively. In the anharm + AD calculations involving anharmonic domains, the PESs were calculated at the CCSD(T)-F12a/VDZ-F12 level of theory with the geometry optimization at theDF-MP2-F12/VDZ-F12 level of theory. | ||||||
| Local anharmonicity | ||||||
| ΔrG°/kJ mol−1 | −10.8 | −13.7 | — | −11.5 | −15.7 | — |
| ΔrH°/kJ mol−1 | −51.9 | −50.8 | — | −52.0 | −51.2 | — |
| ΔrS°/J K−1 mol−1 | −137.8 | −124.5 | — | −135.9 | −118.8 | — |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||||
| Local + global anharmonicity | ||||||
| ΔrG°/kJ mol−1 | −10.8 | −12.1 | −10.5 | −11.1 | −14.6 | −10.5 |
| ΔrH°/kJ mol−1 | −48.5 | −47.3 | −45.0 | −48.5 | −48.8 | −45.0 |
| ΔrS°/J K−1 mol−1 | −126.4 | −118.2 | −115.8 | −125.3 | −114.9 | −115.9 |
Recently, Halonen66 proposed an improved statistical model that incorporates vibrational anharmonicity through a novel partition function requiring only one additional system-specific parameter. Anharmonicity is also implicitly accounted for through the treatment of the configuration space. This statistical model reproduces key thermodynamic properties including formation free energies and potential energies, for typical clusters with an accuracy of approximately 2kBT. Very recently, Kubečka et al.67 combined umbrella sampling (US) with ML to more effectively capture entropic effects and improve the accuracy of free-energy calculations, thereby addressing limitations of traditional QC methods in treating anharmonic and configurationally complex systems. They found that the potential of mean force (PMF) obtained from US and QC calculations agreed when entropy contributions were negligible. However, for systems such as water dimers and SA–W clusters, substantial discrepancies emerged due to entropy effects neglected in QC. After applying the entropy correction derived from US, the QC-based binding energies showed much improved agreement with experimental values.
3.4 Effect of electric field on clusters thermodynamics
In the real atmosphere, the potential difference between thunderstorm clouds and the ground generates electric field of an appreciable strength, and detection of clusters under vacuum conditions—such as in mass spectrometry—also involves external electric fields. Daub et al.68 investigated the effect of electric fields of varying strengths on the binding energies of water dimer, SA–W, SA–AM, and SA–DMA clusters using QC calculations under applied low (≤0.02 V Å−1) and high (>0.02 V Å−1) electric fields. At low fields strengths, the shift in binding energy were small (<0.5 kcal mol−1) and correlated with the dipole moments, with the sign of the change depending on the specific cluster geometry. In contrast, high electric fields induced stronger binding through geometric distortion and dipole enhancement, and in some cases even triggered proton transfer (e.g., in SA–AM) or ion dissociation when the relevant energy barrier dropped below the neutral binding energies.Moreover, ionization sources such as galactic cosmic rays and lightning ionize atmospheric N2 and O2, producing charged species (e.g., N2+ and O2−). These ions subsequently react with gas-phase nucleation precursors such as H2SO4 and H2O to form ions like HSO4− and H3O+, which can further collide with other precursor molecules to generate molecular clusters.18 The electric fields surrounding these ions also influence gas-phase collision-evaporation processes and the thermodynamic properties of clusters. Nadykto et al. showed that interactions between neutral polar molecules and the electric fields of charged particles or clusters—i.e., dipole–charge interactions—can significantly alter the relationship between theoretical particle diameters and the corresponding mean ion mobilities.69 They also explained the discrepancy between diameters predicted by classical Kelvin–Thomson theory and equivalent diameters derived from experimental mobilities using the Millikan–Fuchs equation, and introduced dipole–charge corrections to the classical Kelvin–Thomson framework. Because classical ion-induced nucleation (IIN) theory considers only the Thomson effect (electrostatic energy), it severely underestimates nucleation rates of polar vapors, fails to explain the distinct responses of polar versus nonpolar vapors to ionization, and shows systematic deviations from experiments. To address this, the dipole moment of condensing molecules was incorporated, establishing a dipole–charge interaction mechanism.70 It was found that larger dipole moments lead to stronger ion-enhanced nucleation. For example, highly polar atmospheric precursors such as sulfuric acid, ammonia, water, and alcohols exhibit strong enhancement, whereas nonpolar molecules (e.g., hexane) are largely unaffected. The revised model successfully reproduces experimental critical supersaturation and temperature dependencies. Furthermore, the classical Kelvin–Thomson (CKT) theory, which accounts only for electrostatic energy, overestimates the enthalpy (by up to ∼20 kcal mol−1) and entropy (by ∼5–20 cal mol−1 K−1) changes for clusters such as H+–(H2O)n, H+–(NH3)n, and H+–(CH3OH)n. To improve this, Yu et al. developed a modified Kelvin–Thomson (MKT) equation by incorporating induced dipole energy, permanent dipole orientation, and the dependence of electric field strength on distance. This approach enables accurate predictions of enthalpy and entropy changes for these clusters, substantially reducing discrepancies with experimental results.71
4 Cluster dynamics
Cluster dynamics translate cluster-level thermodynamic properties, together with ambient environmental conditions, into key kinetic parameters such as growth flux, steady state concentration and cluster formation rates. With continued advances in model formulation, parameterization strategies, technical corrections, and expanding application domains, cluster dynamics has evolved into a robust and versatile framework for quantitatively simulating atmospheric NPF.4.1 Birth–death equations
Early on, Yu et al. developed the Advanced Particle Microphysics (APM) model based on a kinetic framework to investigate the fundamental processes of new particle formation (NPF) by explicitly accounting for interactions among neutral and charged clusters and particles.18 The APM model adopts a more rigorous theory that includes charge trapping and three-body trapping effects,72 instead of the hard-sphere collision hypothesis. This approach was further extended to full-size clusters by considering multiple collisions within an attractive potential field, yielding physically consistent, temperature- and pressure-dependent collision kernel parameters. Using this framework, they demonstrated that ion-induced nucleation can be a dominant source of new particles in the troposphere. Yu et al. subsequently proposed a second-generation ion-mediated nucleation model (IMN), which incorporates dipole–charge interactions to calculate the evaporation free energy of SA molecules from ions-cluster systems, consistent with experimental results and derives the corresponding evaporation rates.19 The model also includes a condensation module for low-volatility organic compounds. Under favorable boundary layer conditions, IMN predicts substantial nanoparticle formation with charge distributions in good agreement with field observations. To further investigate the distinct effects of ammonia on neutral, positively charged, and negatively charged clusters, Yu et al. extended IMN to the ternary ion-mediated nucleation (TIMN) model.12 TIMN couples quantum chemistry calculations to obtain thermodynamic data, enabling accurate estimation of evaporation rates and simulation of the H2SO4–H2O–NH3 ion-mediated nucleation mechanism. The results show that ammonia reduces nucleation barriers differently for each type of cluster and successfully reproduces the observed dependencies of nucleation rates on [NH3], [H2SO4], temperature, relative humidity, and ionization rate in CLOUD experiments.Similarly, the Dynamical Atmospheric Cluster Model (DACM)73 describes the dynamic behavior of nanoscale molecular clusters using a kinetic approach rather than classical theory, consistent with Yu et al.18 However, DACM employs a different evaporation scheme and does not explicitly account for specific interactions between ions and neutral molecules. It systematically quantifies the influence of key parameters, such as evaporation rates and saturation vapor pressures, on cluster concentrations. DACM highlights that realistic evaporation rates are essential for reliable predictions and that quantum chemistry calculations provide a robust foundation. Building upon DACM, the Atmospheric Cluster Dynamics Code (ACDC) includes all relevant evaporation and collision processes involving monomers and clusters, enabling the calculation of evaporation rates, concentrations, formation rates, and growth pathways, and thereby allowing detailed simulations of early-stage cluster dynamics during atmospheric NPF.20 It is worth mentioning that the ACDC and ion induced nucleation model (TIMN) adopt different parameterization schemes for collision processes involving ions or ion clusters. TIMN applies the theory of Hoppel and Frick et al.72 to account for electrostatic interactions of charged nanoclusters through corrections to coagulation rates, whereas ACDC uses the parameterization proposed by Su and Chesnovich et al.,74 which accounts for dipole moments and polarizability of neutral molecular clusters in collision rate coefficients calculations. By integrating thermodynamic data with MATLAB's codes solvers, ACDC effectively handles stiff systems of differential equations and automates equation generation through Perl scripts, facilitating extensions to multi-component systems such as sulfuric acid–amine–water. Sensitivity analyses have shown that key parameters, including temperature, boundary conditions, and condensation sink strength, exert a strong influence on steady-state cluster concentrations and particle formation rates. Moreover, under certain atmospheric conditions, non-monomer collisions and ionic effects can contribute substantially to cluster growth and stabilization.
The format of BDE is as follows:
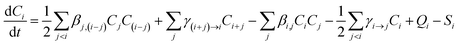 | (4) |
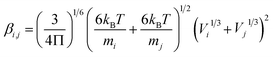 | (5) |
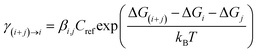 | (6) |
Subsequent developments of ACDC incorporated additional auxiliary parameters to more realistically describe the dynamics of atmospheric aerosol clusters. To bridge the gap between thermodynamic properties obtained from QC calculations and those relevant under real atmospheric conditions, Olenius et al.75 introduced the concept of the actual Gibbs free energy (ΔGactual) in their investigation of SA–AM/DMA cluster growth. In this approach, the reference free energy ΔGref (defined at 1 atm) is converted to values applicable to atmospheric conditions by introducing a pressure-correction term that accounts for the prevailing partial pressures of the participating species (Fig. 8). This correction accurately captures entropy effect, whereby lower partial pressures increase the entropy contribution and consequently raise ΔGactual. By incorporating this adjustment, the method effectively links QC-derived thermodynamic data with realistic atmospheric conditions, providing physically consistent input parameters for predicting nucleation rates. The expression for ΔGactual is as follows:
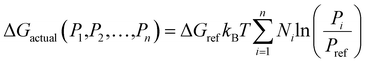 | (7) |
 | ||
| Fig. 8 Main clustering pathways and Gibbs free energies of formation of the clusters in the electrically neutral sulfuric acid–DMA system at T = 5 °C, [SA] = 106 cm−3 and [DMA] = 1 ppt, (Panel (a)) maior routes leading out of the simulated system. For figure clarity, the arrows that fall on top of each other are colored with different shades. (Panel (b)) Formation free energies of a cluster in the system. (Panel (c)) Formation free energy of the growing cluster as a function of growth step. Solid and dashed lines: correspond to the major and minor fluxes, respectively. Adapted from Olenius et al., 2013.75 Reproduced from ref. 75 with permission from the AIP Publishing, copyright 2013. | ||
Furthermore, Elm et al. introduced the concept of cluster formation potential (Jpotential) studies to evaluate the likelihood that small clusters can grow into large clusters.76 As depicted in Fig. 9, Jpotential is defined as the theoretical flux (cm−3 s−1) of small clusters growing into larger clusters in a given systems under specific conditions, typically simulated for (acid)0–2(base)0–2 systems. This metric provides a quantitative criterion for screening systems with high nucleation potential, thereby avoiding redundant calculations. Based on Jpotential, they demonstrated the synergistic enhancement of nucleation arising from the combined effects of base strength and concentration.
 | ||
| Fig. 9 Clusters on the acid–base grid that are allowed to contribute to the potential cluster formation rate, Jpotential. Adapted from Elm et al., 2013.76 Adapted from ref. 76 with permission from the American Chemical Society, copyright 2013. | ||
Classical Nucleation Theory (CNT) treats nucleation as a balance between bulk free-energy gain and interfacial energy cost under a continuum assumption, and has been widely used to describe macroscopic phase transitions such as condensation. However, it fails for atmospheric nucleation because critical clusters are small and discrete, with stability governed by specific interactions (e.g., hydrogen bonding, proton transfer, and electrostatics) that cannot be captured by macroscopic parameters like surface tension. Consequently, CNT shows large systematic errors, including incorrect temperature dependence and critical supersaturation, with nucleation rates often deviating from experiments by several orders of magnitude. To address these limitations, Nadykto et al. developed a corrected CNT (CCNT) by introducing a kinetic consistency term (1/S), a self-consistent correction and an additional empirical term (exp(γA1/kT)/S2).77 CCNT accurately predicts supersaturation and nucleation rates for eight systems—including H2O, D2O, butanol, pentanol, hexanol, dodecane, hexadecane and octadecane—over a wide temperature range (190–340 K). Notably, CCNT remains reliable even when critical clusters contain only a few molecules, whereas classical CNT completely fails in this regime. Besides, a fundamental inconsistency has hindered the direct incorporation of QC data into classical nucleation theory (CNT). Conventional QC free energies for multicomponent clusters do not satisfy the law of mass action and assign nonzero formation free energies to monomers, rendering them incompatible with the nucleation rate exponent intrinsic to CNT. The self-consistent correction mind is also used to solve this problem. Halonen et al.78 proposed a thermodynamically consistent definition of cluster formation free energies based on the law of mass action and a general equilibrium cluster distribution function. Their formulation reconciles QC thermodynamics with CNT and shows excellent agreement with numerical simulations performed using the ACDC. Notably, this approach accurately captures changes in the slope of nucleation rates arising from transitions in the composition of the critical cluster and reproduces the experimentally observed dependencies on temperature and vapor pressure.
Atmospheric H2SO4–H2O nucleation is a key source of new particles, but classical binary homogeneous nucleation (BHN) suffers from major deficiencies, including systematic overestimation of nucleation rates (by 103–107), violation of the law of mass action in cluster distributions, and inconsistent monomer concentrations. To address this, Yu et al. simplified BHN into a quasi-unary H2SO4 nucleation framework by assuming that clusters remain in equilibrium with water, with water content determined solely by temperature and relative humidity, while nucleation is driven only by H2SO4 condensation and evaporation.79 This model reproduces experimental results across a wide range of humidity (4.6–52%) and temperature conditions. To further resolve the key limitations of BHN—rate overestimation, parameter uncertainty, and the breakdown of small-cluster assumptions—they improved the quasi-unary model by constraining sulfuric acid hydration and abandoning the capillarity approximation.80 Similarly, the quasi-univariate approximation is also used to simplify chemical complexity and reduce the difficulty of model computation and parameterization while maintaining accuracy. Olenius et al.81 proposed two simplification strategies within ACDC framework to facilitate efficient prediction of formation rates and populations, as illustrated in Fig. 10. For nucleation rate calculations, two approaches were tested: the “non-interacting additive pathway”, where the total rate equals the sum of rates from each binary system, and “similar-species merging”, in which bases with comparable chemical properties are treated as a single species with superimposed concentrations. The merging method was found to perform well for bases with similar clustering efficiencies, such as dimethylamine and ethylenediamine, where similarity can be assessed based on formation rates or acid–base heterodimer free energies. For the quasi-univariate approximation, the cluster models are simplified by incorporating two key assumptions: the “equilibrium hypothesis”, which posits instantaneous equilibrium between clusters and alkaline species, and the “most stable composition hypothesis”, where clusters are represented by their most thermodynamically favorable composition. This approximation is applicable only under specific conditions—such as systems with strong clustering and low evaporation rates (e.g., SA–DMA) and in the presence of excess stabilizers. In weakly interacting systems, such as SA–AM, however, it tends to overestimate cluster concentrations by 1–2 orders of magnitude and overpredict particle survival probability.
 | ||
| Fig. 10 Schematic presentation of the approaches to simplify multi-component chemistries for (a) assessments of particle formation rate in the presence of multiple species, and (b) modeling of cluster growth by a quasi-unary framework. The axes depict cluster compositions as numbers of molecules of different species. Adapted from Olenius et al., 2023.81 Adapted from ref. 81 with permission from the Royal Society of Chemistry, copyright 2023. Permission conveyed through Copyright Clearance Center. | ||
4.2 Beyond hard-sphere collision models
To address the potential underestimation of collision rate coefficients for polar molecules by conventional hard-sphere models—which can propagate significant errors in ACDC-based simulations—several improved approaches for calculating collision rate coefficients have been proposed using MD simulations. Loukonen et al.82 employed first-principles MD simulations to elucidate the dynamical details of cluster formation in (SA)1(DMA)1 and (SA)1(DMA)1(W)n system. Proton transfer was observed in all simulated collisions, leading to the formation of stable ion-pair clusters. The presence of water molecules did not impede cluster formation; instead, hydration enhanced cluster stability by dissipating excess kinetic energy. Importantly, these simulations highlight the limitations of static QC calculations in capturing nonequilibrium structural transformations and anharmonic vibrational motions during cluster formation.Moreover, Halonen et al.83 employed classical MD to investigate collisions between SA molecules in combination with a Langevin capture framework. By constructing a Langmuir-type capture model based on the attractive region of the PMF, they quantified the collision enhancement arising from long-range intermolecular interactions. Their results showed that, at 300 K, the actual collision rate coefficient is approximately 2.2 times higher than that predicted by conventional hard-sphere collision theory. In subsequent work, they also investigated the atmospheric ion–dipole collisions and calculated rate coefficients of ion–dipole collisions in the gas phase via classical MD and central field methods based on PMF.84 The central field model showed excellent agreement with MD simulations in predicting collision cross-sections and rate coefficients, with errors below 10% across all eight atmospheric ion–dipole systems. It is worth noting that Yu et al. have conducted a series of important studies on ion–dipole collisions in ion-induced nucleation by incorporating dipole–charge interactions into the Kelvin–Thomson equation.69–71
Recently, a new analytical model termed the interacting hard-sphere model has been proposed.85 This model treats clusters as spherical objects and calculates the effective attraction between a monomer and a cluster by integrating long-range Lennard-Jones potentials between individual monomers. When combined with a hard-sphere collision criterion, the model enables efficient estimation of collision cross-sections and rate coefficients. Benchmarking against MD simulations for acid–base cluster systems spanning 1–32 acid–base pairs and temperatures from 200 to 400 K demonstrated that the interacting hard-sphere model substantially outperforms the traditional hard-sphere approximation for small clusters (radii < 2 nm), predicting collision rate coefficients that are 2–3 times higher and agreeing with MD results to within 8%. By contrast, the conventional hard-sphere model underestimates collision rate coefficients for small clusters by a similar factor, potentially propagating order-of-magnitude errors in predicted particle formation rates when applied directly in atmospheric models.
Jiang et al.86 developed a reactive deep neural network force field (DNN-FF) by integrating metadynamics with an active learning strategy to efficiently sample high-energy isomers. By coupling collision rate coefficients derived from Poisson statistics with evaporation rates obtained from QC calculations, they demonstrated that acid–base nucleation rates in polluted environments have likely been underestimated by 1–2 orders of magnitude (Fig. 11). In related work, Liu et al.87 employed machine-learning approaches to investigate collision-driven nucleation in the sulfuric acid–dimethylamine (SA–DMA) system, identifying four distinct nucleation pathways consistent with cloud-chamber observations. Active learning offers a fully iterative and adaptive framework for constructing high-quality training datasets, which is essential for the development of accurate machine-learning models. Along similar lines, Lee et al.88 introduced an active transfer-learning scheme to efficiently sample SA–DMA cluster configurations at the hybrid-DFT level, enabling the training of reliable machine-learning potentials (MLPs) at substantially reduced computational cost.
 | ||
| Fig. 11 A general workflow towards fully ab initio simulation of atmospheric aerosol nucleation. It includes the steps to prepare the data set for training a deep neural network-based force field (DNN-FF), to apply DNN-FF by molecular dynamics (MD), to derive collision rate coefficients from MD, and to couple collision rate coefficients with cluster dynamics model for studying atmospheric aerosol nucleation. (a and b) Shows the metadynamics and active learning techniques used to prepare a data set for the deep neural network, respectively. (c) DNN-FF-driven MD. (d) Cluster dynamics simulation based on MD-derived collision rate coefficients and static quantum chemistry (QC) calculation-derived evaporation rates. Adapted from Jiang et al., 2022.21 Adapted from ref. 21 with permission from the Springer Nature, copyright 2022. Permission conveyed through Copyright Clearance Center. | ||
Sulfuric acid is a key precursor in atmospheric new particle formation (NPF), yet the collision and evaporation kinetics of small clusters remain highly uncertain due to significant errors in quantum chemical (QC) free energy calculations, which can lead to evaporation rate deviations of several orders of magnitude. Additionally, fragmentation during mass spectrometric detection further hinders accurate determination of cluster concentrations, preventing direct reflection of their true distributions. To address this, Kupiainen-Määttä et al.89 developed a Markov chain Monte Carlo (MCMC) method combined with CLOUD experiment related parameters to improve the understanding of cluster evaporation process. They introduced the Metropolis algorithm and the Differential Evolution-Markov Chain algorithm to optimize parameter and overcome multiple local maxima. Including fragmentation dramatically improved the fit, yielding strong agreement with most experimental data and revealing the multiple impacts of ammonia concentration on evaporation rates. For example, the ammonia evaporation rate from HSO4−·(H2SO4)3·NH3 is about 200 s−1 at 20 ppt NH3 but must be below s−1 at 1 ppt NH3 based on cluster concentration distribution obtained from CLOUD experiments and the results of the MCMC simulations.
4.3 Model integration and atmospheric validation
A series of recent studies have leveraged cluster dynamics simulations to advance the understanding of atmospheric nucleation across theoretical, computational, and modeling scales. Chee et al.90 developed a predictive framework for the formation rate of salt nanoparticle clusters (denoted J4×4) based on the stability of acid–base heterodimers comprising one acid and one base molecule, thereby substantially reducing the complexity of nucleation rate estimation. Among the descriptors evaluated, gas-phase acidity (GA) emerged as the most influential predictor: lower GA values, indicative of more facile proton transfer, correlate strongly with enhanced heterodimer stability. The model integrates heterodimer free energy, temperature, and monomer concentration into a normalized heterodimer concentration (Φ), yielding markedly improved predictive performance. Validation against CLOUD data for the SA–AM system demonstrated agreement within two orders of magnitude, underscoring the model's robustness for predicting atmospheric NPF under realistic conditions.ACDC is frequently integrated with complementary theoretical and computational methodologies to achieve a more comprehensive understanding of atmospheric nucleation mechanisms. Shi et al. evaluated the nucleation potential of various aldehydes and their reaction pathways—including hydration, aldol condensation, and polymerization products—in combination with key nucleation precursors such as SA, DMA, and W, using high-level QC calculations coupled with the ACDC.91 Similarly, Li et al. proposed that under conditions of high ammonia pollution and low humidity, the autocatalytic reaction pathway between SO3 and NH3 represents a significant sink for SO3, and revealed that its product, sulfamic acid (SFA), can participate in SA–DMA nucleation, increasing the nucleation rate by approximately twofold.92 Furthermore, Kumar et al.93 demonstrated—through a combination of QC calculations, ACDC, and ab initio molecular dynamics simulations—that nitric acid (HNO3) and ammonia (NH3)/amines rapidly form ion pairs stabilized at the gas–water interface, uncovering a new pathway for atmospheric gas–particle conversion.
Through model iteration, parameter innovation, and integration of multiple computational techniques, cluster dynamics simulations have progressively achieved accurate mapping from idealized systems to complex atmospheric environments. They provide a core methodological foundation for elucidating NPF mechanisms and optimizing climate model parameterization schemes. For instance, Baranizadeh et al. employ the ACDC model to calculate the nucleation rate of H2SO4–H2O–NH3 ternary system, embedding PMCAMx-UF in a lookup table without the need for empirical scaling.94 Yu et al. also generate nucleation rate lookup tables for H2SO4–H2O binary homogeneous nucleation (BHN), H2SO4–H2O–NH3 ternary homogeneous nucleation (THN), H2SO4–H2O ion binary ion mediated nucleation (BIMN), and H2SO4–H2O–NH3 ion ternary ion mediated nucleation (TIMN).95 Their lookup table covers a wide range of key parameters that control binary, ternary, and ion mediated nucleation in the Earth's atmosphere, providing an economically efficient solution for multidimensional modeling. Moreover, Shen and Zhao et al. adapted the ACDC cluster dynamics model, four-dimensional lookup tables were generated and embedded into WRF-Chem, and the most accurate SA–DMA nucleation parameterization was selected to improve simulations for Beijing in both winter and summer. Multiple nucleation mechanisms were further integrated into the E3SM global climate model and WRF-Chem to map their spatial distributions over China and globally, and to quantify the contribution of NPF to CCN.96–98
5 Conclusion and perspective
To date, the nucleation mechanism in NPF has remained a central focus in atmospheric chemistry. Notably, although the application of these technologies to specific chemical systems has generated an extensive and diverse body of studies, such works are outside the scope of this review. Rather, this paper focuses on tracing the initial implementation of these technologies within nucleation research and highlighting the methodological refinements developed specifically for the nucleation field.In the configuration sampling section, the field has evolved from an early reliance on chemical intuition to the adoption of advanced methods such as Basin Hopping (BH), Molecular Dynamics (MD), Genetic Algorithms (GA), metadynamics, and Artificial Bee Colony (ABC) algorithms. These advances have significantly improved our ability to obtain complex cluster structure information, which is critical for the nucleation process. Notably, the rigorous yet efficient JKCS workflow has greatly enhanced the capability to search for global minima in cluster structures. Currently, the ABC method has become a mainstream approach for configuration sampling due to its open-source nature and ease of use, with ABCluster being one of the most widely used implementations. However, several limitations remain. In ABCluster, initial structures are typically generated randomly, which is suitable for small, highly disordered clusters. In contrast, nanoscale clusters often exhibit more ordered, particle-like structures, and purely random initialization may reduce search efficiency. To address this, Zhang et al. improved the ABC algorithm by introducing adaptive learning during the search process.99 The resulting program, NWPEsSe, has been open-sourced, although its applicability to atmospheric nanoscale clusters remains unclear. Another key limitation of algorithms such as ABCluster and NWPEsSe is the rigid-body approximation for molecules. Small molecular units are typically treated as rigid, with internal degrees of freedom—such as bending, torsion, cis–trans isomerization, and variations in bond and dihedral angles—frozen. Only the positions and orientations of these units are optimized. While this assumption improves computational efficiency, it can fail for flexible molecules such as long-chain alkanes, glutaric acid, and adipic acid. These molecules can adopt multiple conformations (e.g., all-trans, hairpin, or twisted), often with large energy differences. As a result, the rigid approximation may miss the true lowest-energy structures. In many cases, the internal conformation directly determines the global minimum of the entire cluster. Without sampling internal degrees of freedom, the true global minimum cannot be reliably identified. A common workaround is to perform conformational search (CS) first, followed by cluster global optimization (GO). However, this sequential approach still does not fully capture the coupling between internal and external degrees of freedom, and its applicability to atmospheric nucleation remains uncertain.100
To elucidate the physical and chemical properties of clusters at the micro molecular level, researchers have utilized rigorous quantum chemistry calculation methods, benchmarking, and Machine Learning (ML) to rapidly obtain highly accurate thermodynamic and spectroscopic data. Tons of work have been conducted on QC calculation methods and basis set for atmospheric clusters, focusing on the challenging thermodynamic acquisition of large-sized clusters. In recent years, ML has been widely applied to predict thermodynamic properties, significantly reducing computational cost and time. However, most existing ML models have limited generalization capability and are typically restricted to one or a few specific systems, making it difficult to cover diverse nucleation pathways. As a result, retraining is often required for new systems. This process depends on large, high-quality QC datasets, which is time-consuming and does not fundamentally resolve the scalability issue. The emergence of general large-atom models and fine-tuning or distillation-based ML frameworks has significantly improved model transferability and generalization ability.101–103 Nevertheless, their application in atmospheric nucleation remains largely unexplored. Moreover, as cluster sizes continue to increase, there is still a strong need for more cost-effective and scalable computational approaches.
Cluster dynamics serves as a bridge linking microscopic structure to macroscopic formation rate and concentration distribution. Atmospheric Cluster Dynamics Code (ACDC) has become the standard tool in this field. Many scholars continue to enhance ACDC using ML and MD, for example, by incorporating collision corrections. Furthermore, the introduction of actual Gibbs free energy and J-potential concepts is pushing ACDC toward nucleation kinetics simulations that more closely resemble real atmospheric conditions. However, evaporation rate calculations in ACDC still rely heavily on cluster binding energies obtained from quantum chemistry (QC). Due to methodological limitations, including finite accuracy and the harmonic approximation, these binding energies carry significant uncertainties. Moreover, molecular dynamics simulations indicate that gas-phase clusters are not static but dynamically interconvert among multiple structures.82 In contrast, QC calculations typically consider only the lowest-energy structure. Although some studies have proposed incorporating multiple conformations, this approach is still limited by the completeness of conformational sampling, i.e., whether all relevant local minima can be identified.104–106 Therefore, applying MD and enhanced sampling techniques to directly simulate cluster evaporation processes and obtain statistically robust evaporation rates with high physical fidelity could effectively improve current QC-based estimates. Loukonen et al. reported that the sticking (or accommodation) coefficient is close to unity for collisions between H2SO4–H2O clusters and dimethylamine, with no observed post-collision dissociation.82 However, it remains unclear whether this assumption holds for larger or more complex clusters. Key open questions include whether the sticking coefficient remains unity, whether post-collision dissociation occurs, and how these effects influence cluster dynamics predictions.
In summary, driven by the challenges in configuration sampling, property characterization, and dynamics data acquisition, a comprehensive theoretical framework centered on quantum chemical calculations has emerged, accompanied by many automated tools. However, high-precision methods like quantum chemistry and Ab Initio Molecular Dynamics (AIMD) are computationally resource-intensive, making it difficult to simulate large-sized clusters or formation processes under complex atmospheric conditions. As a rapidly developing field, Machine Learning holds promise for addressing these bottlenecks by accurately predicting thermodynamic information and simulating collision correction via Machine Learning Force Fields (MLFFs). While MLFFs are currently limited to specific nucleation systems and sizes, the development of a universal MLFF for nucleation has the potential to overcome current limitations. Finally, although direct simulation of cluster evaporation could provide critical microscopic insights and reliable macroscopic evaporation rates, this area currently lacks sufficient theoretical frameworks and effective simulation methods.
Author contributions
Conceptualization: Shuai Jiang; methodology: Yongjian Lian, Xurong Bai, Ruoying Yuan, Shuai Jiang; investigation: Yongjian Lian, Xurong Bai, Ruoying Yuan, Shuai Jiang; formal analysis: Yongjian Lian; data curation: Yongjian Lian, Xurong Bai, Ruoying Yuan, Tingyu Wei; resources: Yongjian Lian, Xurong Bai, Ruoying Yuan, Tingyu Wei; visualization: Yongjian Lian, Xurong Bai, Ruoying Yuan, Tingyu Wei; writing – original draft: Yongjian Lian, Xurong Bai, Ruoying Yuan; writing – review & editing: Yongjian Lian, Shuai Jiang, Tingyu Wei; supervision: Shuai Jiang; funding acquisition: Shuai Jiang, Jianfei Peng, Hongjun Mao.Conflicts of interest
There are no conflicts to declare.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.Acknowledgements
This research was funded by National Natural Science Foundation of China (42477111), the Fundamental Research Funds for the Central Universities of China (63253201, 63251191, 63243126), Natural Science Foundation of Tianjin Municipality (24JCYBJC01700) and Tianjin Key Research and Development Project (24YFXTHZ00070).References
- C. Rose, Q. Zha, L. Dada, C. Yan, K. Lehtipalo, H. Junninen, S. B. Mazon, T. Jokinen, N. Sarnela and M. Sipilä, Observations of biogenic ion-induced cluster formation in the atmosphere, Sci. Adv., 2018, 4, eaar5218 CrossRef PubMed.
- Y. H. Wang, Z. R. Liu, J. K. Zhang, B. Hu, D. S. Ji, Y. C. Yu and Y. S. Wang, Aerosol physicochemical properties and implications for visibility during an intense haze episode during winter in Beijing, Atmos. Chem. Phys., 2015, 15, 3205–3215 CrossRef CAS.
- M. Kulmala, Importance of New Particle Formation for Climate and Air Quality, ACS ES&T Air, 2025, 2, 710–712 Search PubMed.
- S.-H. Lee, H. Gordon, H. Yu, K. Lehtipalo, R. Haley, Y. Li and R. Zhang, New Particle Formation in the Atmosphere: From Molecular Clusters to Global Climate, J. Geophys. Res. Atmos., 2019, 124, 7098–7146 CrossRef CAS.
- F. Yu and G. Luo, Simulation of particle size distribution with a global aerosol model: contribution of nucleation to aerosol and CCN number concentrations, Atmos. Chem. Phys., 2009, 9, 7691–7710 CrossRef CAS.
- J. Merikanto, D. V. Spracklen, G. W. Mann, S. J. Pickering and K. S. Carslaw, Impact of nucleation on global CCN, Atmos. Chem. Phys., 2009, 9, 8601–8616 CrossRef CAS.
- P. Stier, S. C. van den Heever, M. W. Christensen, E. Gryspeerdt, G. Dagan, S. M. Saleeby, M. Bollasina, L. Donner, K. Emanuel, A. M. L. Ekman, G. Feingold, P. Field, P. Forster, J. Haywood, R. Kahn, I. Koren, C. Kummerow, T. L'Ecuyer, U. Lohmann, Y. Ming, G. Myhre, J. Quaas, D. Rosenfeld, B. Samset, A. Seifert, G. Stephens and W.-K. Tao, Multifaceted aerosol effects on precipitation, Nat. Geosci., 2024, 17, 719–732 CrossRef CAS.
- M. Kulmala, How Particles Nucleate and Grow, Science, 2003, 302, 1000–1001 CrossRef CAS PubMed.
- S. Saarikoski, H. Hellén, A. P. Praplan, S. Schallhart, P. Clusius, J. V. Niemi, A. Kousa, T. Tykkä, R. Kouznetsov, M. Aurela, L. Salo, T. Rönkkö, L. M. F. Barreira, L. Pirjola and H. Timonen, Characterization of volatile organic compounds and submicron organic aerosol in a traffic environment, Atmos. Chem. Phys., 2023, 23, 2963–2982 CrossRef CAS.
- J. Kirkby, J. Curtius, J. Almeida, E. Dunne, J. Duplissy, S. Ehrhart, A. Franchin, S. Gagné, L. Ickes, A. Kürten, A. Kupc, A. Metzger, F. Riccobono, L. Rondo, S. Schobesberger, G. Tsagkogeorgas, D. Wimmer, A. Amorim, F. Bianchi, M. Breitenlechner, A. David, J. Dommen, A. Downard, M. Ehn, R. C. Flagan, S. Haider, A. Hansel, D. Hauser, W. Jud, H. Junninen, F. Kreissl, A. Kvashin, A. Laaksonen, K. Lehtipalo, J. Lima, E. R. Lovejoy, V. Makhmutov, S. Mathot, J. Mikkilä, P. Minginette, S. Mogo, T. Nieminen, A. Onnela, P. Pereira, T. Petäjä, R. Schnitzhofer, J. H. Seinfeld, M. Sipilä, Y. Stozhkov, F. Stratmann, A. Tomé, J. Vanhanen, Y. Viisanen, A. Vrtala, P. E. Wagner, H. Walther, E. Weingartner, H. Wex, P. M. Winkler, K. S. Carslaw, D. R. Worsnop, U. Baltensperger and M. Kulmala, Role of sulphuric acid, ammonia and galactic cosmic rays in atmospheric aerosol nucleation, Nature, 2011, 476, 429–433 CrossRef CAS PubMed.
- Y. Liu, W. Nie, X. Qi, Y. Li, T. Xu, C. Liu, D. Ge, L. Chen, G. Niu, J. Wang, L. Yang, L. Wang, C. Zhu, J. Wang, Y. Zhang, T. Liu, Q. Zha, C. Yan, C. Ye, G. Zhang, R. Hu, R.-J. Huang, X. Chi, T. Zhu and A. Ding, The Pivotal Role of Heavy Terpenes and Anthropogenic Interactions in New Particle Formation on the Southeastern Qinghai-Tibet Plateau, Environ. Sci. Technol., 2024, 58, 19748–19761 CrossRef PubMed.
- F. Yu, A. B. Nadykto, J. Herb, G. Luo, K. M. Nazarenko and L. A. Uvarova, H2SO4–H2O–NH3 ternary ion-mediated nucleation (TIMN): kinetic-based model and comparison with CLOUD measurements, Atmos. Chem. Phys., 2018, 18, 17451–17474 CrossRef CAS.
- X. Zhang, S. Tan, X. Chen and S. Yin, Computational chemistry of cluster: Understanding the mechanism of atmospheric new particle formation at the molecular level, Chemosphere, 2022, 308, 136109 CrossRef CAS PubMed.
- J. Elm, D. Ayoubi, M. Engsvang, A. B. Jensen, Y. Knattrup, J. Kubečka, C. J. Bready, V. R. Fowler, S. E. Harold, O. M. Longsworth and G. C. Shields, Quantum chemical modeling of organic enhanced atmospheric nucleation: A critical review, WIREs Comput. Mol. Sci., 2023, 13, e1662 CrossRef CAS.
- C. Li and R. Signorell, Understanding vapor nucleation on the molecular level: A review, J. Aerosol Sci., 2021, 153, 105676 CrossRef CAS.
- X.-C. He, Y. J. Tham, L. Dada, M. Wang, H. Finkenzeller, D. Stolzenburg, S. Iyer, M. Simon, A. Kürten and J. Shen, Role of iodine oxoacids in atmospheric aerosol nucleation, Science, 2021, 371, 589–595 CrossRef CAS PubMed.
- L. Liu, F. Yu, L. Du, Z. Yang, J. S. Francisco and X. Zhang, Rapid sulfuric acid–dimethylamine nucleation enhanced by nitric acid in polluted regions, Proc. Natl. Acad. Sci. U. S. A., 2021, 118, e2108384118 CrossRef CAS PubMed.
- F. Yu and R. P. Turco, From molecular clusters to nanoparticles: Role of ambient ionization in tropospheric aerosol formation, J. Geophys. Res. Atmos., 2001, 106, 4797–4814 CrossRef CAS.
- F. Yu, From molecular clusters to nanoparticles: second-generation ion-mediated nucleation model, Atmos. Chem. Phys., 2006, 6, 5193–5211 CrossRef CAS.
- M. J. McGrath, T. Olenius, I. K. Ortega, V. Loukonen, P. Paasonen, T. Kurtén, M. Kulmala and H. Vehkamäki, Atmospheric Cluster Dynamics Code: a flexible method for solution of the birth-death equations, Atmos. Chem. Phys., 2012, 12, 2345–2355 CrossRef CAS.
- S. Jiang, Y.-R. Liu, T. Huang, Y.-J. Feng, C.-Y. Wang, Z.-Q. Wang, B.-J. Ge, Q.-S. Liu, W.-R. Guang and W. Huang, Towards fully ab initio simulation of atmospheric aerosol nucleation, Nat. Commun., 2022, 13, 6067 CrossRef CAS PubMed.
- J. Kubečka, D. Ayoubi, Z. Tang, Y. Knattrup, M. Engsvang, H. Wu and J. Elm, Accurate modeling of the potential energy surface of atmospheric molecular clusters boosted by neural networks, Environ. Sci.: Adv., 2024, 3, 1438–1451 Search PubMed.
- C. D. Daub, T. Kurtén and M. Rissanen, Molecular dynamics simulations of atmospherically relevant molecular clusters: a case study of nitrate ion complexes, Phys. Chem. Chem. Phys., 2025, 27, 10804–10814 RSC.
- A. B. Jensen and J. Elm, Massive Assessment of the Geometries of Atmospheric Molecular Clusters, J. Chem. Theory Comput., 2024, 20, 8549–8558 CrossRef CAS PubMed.
- M. Engsvang, H. Wu, Y. Knattrup, J. Kubečka, A. B. Jensen and J. Elm, Quantum chemical modeling of atmospheric molecular clusters involving inorganic acids and methanesulfonic acid, Chem. Phys. Rev., 2023, 4, 031311 CrossRef CAS.
- V. Loukonen, T. Kurtén, I. K. Ortega, H. Vehkamäki, A. A. H. Pádua, K. Sellegri and M. Kulmala, Enhancing effect of dimethylamine in sulfuric acid nucleation in the presence of water – a computational study, Atmos. Chem. Phys., 2010, 10, 4961–4974 CrossRef CAS.
- B. Temelso, T. E. Morrell, R. M. Shields, M. A. Allodi, E. K. Wood, K. N. Kirschner, T. C. Castonguay, K. A. Archer and G. C. Shields, Quantum Mechanical Study of Sulfuric Acid Hydration: Atmospheric Implications, J. Phys. Chem. A, 2012, 116, 2209–2224 CrossRef CAS PubMed.
- A. Laio and F. L. Gervasio, Metadynamics: a method to simulate rare events and reconstruct the free energy in biophysics, chemistry and material science, Rep. Prog. Phys., 2008, 71, 126601 CrossRef.
- J. E. Herr, K. Yao, R. McIntyre, D. W. Toth and J. Parkhill, Metadynamics for training neural network model chemistries: A competitive assessment, J. Chem. Phys., 2018, 148, 241710 CrossRef PubMed.
- S. Jiang, Y.-R. Liu, C.-Y. Wang and T. Huang, Benchmarking general neural network potential ANI-2x on aerosol nucleation molecular clusters, Int. J. Quantum Chem., 2023, 123, e27087 CrossRef CAS.
- S. Jiang, Y.-R. Liu, T. Huang, H. Wen, K.-M. Xu, W.-X. Zhao, W.-J. Zhang and W. Huang, Study of Cl−(H2O)n (n = 1–4) using basin-hopping method coupled with density functional theory, J. Comput. Chem., 2014, 35, 159–165 CrossRef CAS PubMed.
- D. J. Wales and H. A. Scheraga, Global optimization of clusters, crystals, and biomolecules, Science, 1999, 285, 1368–1372 CrossRef CAS PubMed.
- W. Xu and R. Zhang, A theoretical study of hydrated molecular clusters of amines and dicarboxylic acids, J. Chem. Phys., 2013, 139, 064312 CrossRef PubMed.
- J. Zhang and M. Dolg, ABCluster: the artificial bee colony algorithm for cluster global optimization, Phys. Chem. Chem. Phys., 2015, 17, 24173–24181 RSC.
- J. Zhang and M. Dolg, Global optimization of clusters of rigid molecules using the artificial bee colony algorithm, Phys. Chem. Chem. Phys., 2016, 18, 3003–3010 RSC.
- H. Wu, M. Engsvang, Y. Knattrup, J. Kubečka and J. Elm, Improved Configurational Sampling Protocol for Large Atmospheric Molecular Clusters, ACS Omega, 2023, 8, 45065–45077 CrossRef CAS PubMed.
- B. Temelso, E. F. Morrison, D. L. Speer, B. C. Cao, N. Appiah-Padi, G. Kim and G. C. Shields, Effect of Mixing Ammonia and Alkylamines on Sulfate Aerosol Formation, J. Phys. Chem. A, 2018, 122, 1612–1622 CrossRef CAS PubMed.
- J. V. Kildgaard, K. V. Mikkelsen, M. Bilde and J. Elm, Hydration of Atmospheric Molecular Clusters: A New Method for Systematic Configurational Sampling, J. Phys. Chem. A, 2018, 122, 5026–5036 CrossRef CAS PubMed.
- J. Kubečka, V. Besel, I. Neefjes, Y. Knattrup, T. Kurtén, H. Vehkamäki and J. Elm, Computational Tools for Handling Molecular Clusters: Configurational Sampling, Storage, Analysis, and Machine Learning, ACS Omega, 2023, 8, 45115–45128 CrossRef PubMed.
- B. Temelso, J. M. Mabey, T. Kubota, N. Appiah-Padi and G. C. Shields, ArbAlign: A Tool for Optimal Alignment of Arbitrarily Ordered Isomers Using the Kuhn–Munkres Algorithm, J. Chem. Inf. Model., 2017, 57, 1045–1054 CrossRef CAS PubMed.
- A. B. Nadykto, F. Yu and J. Herb, Theoretical analysis of the gas-phase hydration of common atmospheric pre-nucleation (HSO4-)(H2O)n and (H3O+)(H2SO4)(H2O)n cluster ions, Chem. Phys., 2009, 360, 67–73 CrossRef CAS.
- J. Herb, Y. Xu, F. Yu and A. B. Nadykto, Large Hydrogen-Bonded Pre-nucleation (HSO4–)(H2SO4)m(H2O)kand (HSO4–)(NH3)(H2SO4)m(H2O)k Clusters in the Earth's Atmosphere, J. Phys. Chem. A, 2013, 117, 133–152 CrossRef CAS PubMed.
- N. Bork, L. Du, H. Reiman, T. Kurtén and H. G. Kjaergaard, Benchmarking Ab Initio Binding Energies of Hydrogen-Bonded Molecular Clusters Based on FTIR Spectroscopy, J. Phys. Chem. A, 2014, 118, 5316–5322 CrossRef CAS PubMed.
- A. S. Hansen, L. Du and H. G. Kjaergaard, The effect of fluorine substitution in alcohol–amine complexes, Phys. Chem. Chem. Phys., 2014, 16, 22882–22891 RSC.
- S. R. Gadre, S. D. Yeole and N. Sahu, Quantum Chemical Investigations on Molecular Clusters, Chem. Rev., 2014, 114, 12132–12173 CrossRef CAS PubMed.
- A. S. Hansen, E. Vogt and H. G. Kjaergaard, Gibbs energy of complex formation – combining infrared spectroscopy and vibrational theory, Int. Rev. Phys. Chem., 2019, 38, 115–148 Search PubMed.
- Y. Knattrup, A. Jensen and J. Elm, Coupled Cluster Free Energies for Atmospheric Molecular Clusters: Benchmark and Matching Experimental Free Energies, ChemRxiv, preprint, 2026, 1–40, DOI:10.26434/chemrxiv-2026-0jz1q.
- H. Henschel, J. C. A. Navarro, T. Yli-Juuti, O. Kupiainen-Määttä, T. Olenius, I. K. Ortega, S. L. Clegg, T. Kurtén, I. Riipinen and H. Vehkamäki, Hydration of Atmospherically Relevant Molecular Clusters: Computational Chemistry and Classical Thermodynamics, J. Phys. Chem. A, 2014, 118, 2599–2611 CrossRef CAS PubMed.
- A. B. Nadykto, J. Herb, F. Yu and Y. Xu, Enhancement in the production of nucleating clusters due to dimethylamine and large uncertainties in the thermochemistry of amine-enhanced nucleation, Chem. Phys. Lett., 2014, 609, 42–49 CrossRef CAS.
- O. Kupiainen-Määttä, H. Henschel, T. Kurtén, V. Loukonen, T. Olenius, P. Paasonen and H. Vehkamäki, Comment on ‘Enhancement in the production of nucleating clusters due to dimethylamine and large uncertainties in the thermochemistry of amine-enhanced nucleation’ by Nadykto et al., Chem. Phys. Lett. 609 (2014) 42–49, Chem. Phys. Lett., 2015, 624, 107–110 CrossRef.
- J. Elm, M. Bilde and K. V. Mikkelsen, Assessment of Density Functional Theory in Predicting Structures and Free Energies of Reaction of Atmospheric Prenucleation Clusters, J. Chem. Theory Comput., 2012, 8, 2071–2077 CrossRef CAS PubMed.
- J. Elm and K. V. Mikkelsen, Computational approaches for efficiently modelling of small atmospheric clusters, Chem. Phys. Lett., 2014, 615, 26–29 CrossRef CAS.
- J. Elm and K. Kristensen, Basis set convergence of the binding energies of strongly hydrogen-bonded atmospheric clusters, Phys. Chem. Chem. Phys., 2017, 19, 1122–1133 RSC.
- G. Schmitz and J. Elm, Assessment of the DLPNO Binding Energies of Strongly Noncovalent Bonded Atmospheric Molecular Clusters, ACS Omega, 2020, 5, 7601–7612 CrossRef CAS PubMed.
- M. Engsvang, J. Kubečka and J. Elm, Toward Modeling the Growth of Large Atmospheric Sulfuric Acid–Ammonia Clusters, ACS Omega, 2023, 8, 34597–34609 CrossRef CAS PubMed.
- Y. Knattrup and J. Elm, Extrapolating Local Coupled Cluster Calculations toward CCSD(T)/CBS Binding Energies of Atmospheric Molecular Clusters, ACS Omega, 2025, 10, 46794–46808 CrossRef CAS PubMed.
- M. Engsvang and J. Elm, Modeling the Binding Free Energy of Large Atmospheric Sulfuric Acid–Ammonia Clusters, ACS Omega, 2022, 7, 8077–8083 CrossRef CAS PubMed.
- A. B. Jensen, J. Kubečka, G. Schmitz, O. Christiansen and J. Elm, Massive Assessment of the Binding Energies of Atmospheric Molecular Clusters, J. Chem. Theory Comput., 2022, 18, 7373–7383 CrossRef CAS PubMed.
- J. Kubečka, A. S. Christensen, F. R. Rasmussen and J. Elm, Quantum Machine Learning Approach for Studying Atmospheric Cluster Formation, Environ. Sci. Technol. Lett., 2022, 9, 239–244 CrossRef.
- K. Schütt, O. Unke and M. Gastegger, Equivariant message passing for the prediction of tensorial properties and molecular spectra, arXiv, 2021, preprint, arXiv:2102.03150, DOI:10.48550/arXiv.2102.03150.
- L. Seppäläinen, J. Kubečka, J. Elm and K. R. Puolamäki, Fast and Interpretable Machine Learning Modeling of Atmospheric Molecular Clusters, J. Phys. Chem. A, 2026, 130, 902–913 CrossRef PubMed.
- Y. Liu, H.-B. Xie, F. Ma, J. Chen and J. Elm, Amine-Enhanced Methanesulfonic Acid-Driven Nucleation: Predictive Model and Cluster Formation Mechanism, Environ. Sci. Technol., 2022, 56, 7751–7760 CrossRef CAS PubMed.
- A. N. Pedersen, Y. Knattrup and J. Elm, A cluster-of-functional-groups approach for studying organic enhanced atmospheric cluster formation, Aerosol Air Qual. Res., 2024, 2, 123–134 CrossRef.
- S. M. Kathmann, G. K. Schenter and B. C. Garrett, Multicomponent dynamical nucleation theory and sensitivity analysis, J. Chem. Phys., 2004, 120, 9133–9141 CrossRef CAS PubMed.
- L. Partanen, V. Hänninen and L. Halonen, Effects of Global and Local Anharmonicities on the Thermodynamic Properties of Sulfuric Acid Monohydrate, J. Chem. Theory Comput., 2016, 12, 5511–5524 CrossRef CAS PubMed.
- R. Halonen, Assessment of Anharmonicities in Clusters: Developing and Validating a Minimum-Information Partition Function, J. Chem. Theory Comput., 2024, 20, 4099–4114 CrossRef CAS PubMed.
- J. Kubečka, Y. Knattrup, G. B. Trolle, B. Reischl, A. S. Lykke-Møller, J. Elm and I. Neefjes, Thermodynamics of Molecular Binding and Clustering in the Atmosphere Revealed through Conventional and ML-Enhanced Umbrella Sampling, ACS Omega, 2025, 10, 39148–39161 CrossRef PubMed.
- C. D. Daub and T. Kurtén, Effect of an Electric Field on the Structure and Stability of Atmospheric Clusters, J. Phys. Chem. A, 2024, 128, 646–655 CrossRef CAS PubMed.
- A. B. Nadykto, J. M. Mäkelä, F. Yu, M. Kulmala and A. Laaksonen, Comparison of the experimental mobility equivalent diameter for small cluster ions with theoretical particle diameter corrected by effect of vapour polarity, Chem. Phys. Lett., 2003, 382, 6–11 CrossRef CAS.
- F. Yu, Dipole Moment of Condensing Monomers: A New Parameter Controlling the IonInduced Nucleation, Phys. Rev. Lett., 2004, 93, 016101 CrossRef.
- F. Yu, Modified Kelvin–Thomson equation considering ion-dipole interaction: Comparison with observed ion-clustering enthalpies and entropies, J. Chem. Phys., 2005, 122, 084503 CrossRef PubMed.
- W. A. Hoppel and G. M. Frick, Ion—Aerosol Attachment Coefficients and the Steady-State Charge Distribution on Aerosols in a Bipolar Ion Environment, Aerosol Sci. Technol., 1986, 5, 1–21 CrossRef CAS.
- M. Kulmala, Dynamical atmospheric cluster model, Atmos. Res., 2010, 98, 201–206 CrossRef CAS.
- T. Su and W. J. Chesnavich, Parametrization of the ion–polar molecule collision rate constant by trajectory calculations, J. Chem. Phys., 1982, 76, 5183–5185 CrossRef CAS.
- T. Olenius, O. Kupiainen-Määttä, I. K. Ortega, T. Kurtén and H. Vehkamäki, Free energy barrier in the growth of sulfuric acid–ammonia and sulfuric acid–dimethylamine clusters, J. Chem. Phys., 2013, 139, 084312 CrossRef CAS PubMed.
- J. Elm, Clusteromics I: Principles, Protocols, and Applications to Sulfuric Acid-Base Cluster Formation, ACS Omega, 2021, 6, 7804–7814 CrossRef CAS PubMed.
- A. B. Nadykto and F. Yu, Simple correction to the classical theory of homogeneous nucleation, J. Chem. Phys., 2005, 122, 104511 CrossRef PubMed.
- R. Halonen, A consistent formation free energy definition for multicomponent clusters in quantum thermochemistry, J. Aerosol Sci., 2022, 162, 105974 CrossRef CAS.
- F. Yu, Quasi-unary homogeneous nucleation of H2SO4-H2O, J. Chem. Phys., 2005, 122, 074501 CrossRef PubMed.
- F. Yu, Improved quasi-unary nucleation model for binary H2SO4–H2O homogeneous nucleation, J. Chem. Phys., 2007, 127, 054301 CrossRef PubMed.
- T. Olenius, R. Bergström, J. Kubečka, N. Myllys and J. Elm, Reducing chemical complexity in representation of new-particle formation: evaluation of simplification approaches, Environ. Sci.: Atmos., 2023, 3, 552–567 CAS.
- V. Loukonen, N. Bork and H. Vehkamäki, From collisions to clusters: first steps of sulphuric acid nanocluster formation dynamics, Mol. Phys., 2014, 112, 1979–1986 CrossRef CAS.
- R. Halonen, E. Zapadinsky, T. Kurtén, H. Vehkamäki and B. Reischl, Rate enhancement in collisions of sulfuric acid molecules due to long-range intermolecular forces, Atmos. Chem. Phys., 2019, 19, 13355–13366 CrossRef CAS.
- I. Neefjes, R. Halonen, H. Vehkamäki and B. Reischl, Modeling approaches for atmospheric ion–dipole collisions: all-atom trajectory simulations and central field methods, Atmos. Chem. Phys., 2022, 22, 11155–11172 CrossRef CAS.
- Q. Liang, C. Zhu and J. Yang, Water Charge Transfer Accelerates Criegee Intermediate Reaction with H2O– Radical Anion at the Aqueous Interface, J. Am. Chem. Soc., 2023, 145, 10159–10166 CrossRef CAS PubMed.
- S. Jiang, Y. R. Liu, T. Huang, Y. J. Feng, C. Y. Wang, Z. Q. Wang, B. J. Ge, Q. S. Liu, W. R. Guang and W. Huang, Towards fully ab initio simulation of atmospheric aerosol nucleation, Nat. Commun., 2022, 13, 6067 CrossRef CAS PubMed.
- Y.-R. Liu and Y. Jiang, Predicting Composition Evolution for a Sulfuric Acid-Dimethylamine System from Monomer to Nanoparticle Using Machine Learning, J. Phys. Chem. A, 2025, 129, 222–231 CrossRef CAS PubMed.
- S. Kang, R. Cai, D. S. Yang, D. J. Ham, M. Kulmala, J. H. Seinfeld, J. Jiang and H. C. Lee, Efficient Configuration Sampling for Hybrid Functional DFT Calculations to Train Machine-Learning Potentials: Application to Atmospheric Chemistry, Small Methods, 2026, 10, 2500547 CrossRef CAS PubMed.
- O. Kupiainen-Määttä, A Monte Carlo approach for determining cluster evaporation rates from concentration measurements, Atmos. Chem. Phys., 2016, 16, 14585–14598 CrossRef.
- S. Chee, K. Barsanti, J. N. Smith and N. Myllys, A predictive model for salt nanoparticle formation using heterodimer stability calculations, Atmos. Chem. Phys., 2021, 21, 11637–11654 CrossRef CAS.
- X. Shi, R. Zhang, Y. Sun, F. Xu, Q. Zhang and W. Wang, A density functional theory study of aldehydes and their atmospheric products participating in nucleation, Phys. Chem. Chem. Phys., 2018, 20, 1005–1011 RSC.
- H. Li, J. Zhong, H. Vehkamaki, T. Kurten, W. Wang, M. Ge, S. Zhang, Z. Li, X. Zhang, J. S. Francisco and X. C. Zeng, Self-Catalytic Reaction of SO3 and NH3 To Produce Sulfamic Acid and Its Implication to Atmospheric Particle Formation, J. Am. Chem. Soc., 2018, 140, 11020–11028 CrossRef CAS PubMed.
- M. Kumar, H. Li, X. Zhang, X. C. Zeng and J. S. Francisco, Nitric Acid–Amine Chemistry in the Gas Phase and at the Air–Water Interface, J. Am. Chem. Soc., 2018, 140, 6456–6466 CrossRef CAS PubMed.
- E. Baranizadeh, B. N. Murphy, J. Julin, S. Falahat, C. L. Reddington, A. Arola, L. Ahlm, S. Mikkonen, C. Fountoukis, D. Patoulias, A. Minikin, T. Hamburger, A. Laaksonen, S. N. Pandis, H. Vehkamäki, K. E. J. Lehtinen and I. Riipinen, Implementation of state-of-the-art ternary new-particle formation scheme to the regional chemical transport model PMCAMx-UF in Europe, Geosci. Model Dev., 2016, 9, 2741–2754 CrossRef CAS.
- F. Yu, A. B. Nadykto, G. Luo and J. Herb, H2SO4–H2O binary and H2SO4–H2O–NH3 ternary homogeneous and ion-mediated nucleation: lookup tables version 1.0 for 3-D modeling application, Geosci. Model Dev., 2020, 13, 2663–2670 CrossRef CAS.
- B. Zhao, N. M. Donahue, K. Zhang, L. Mao, M. Shrivastava, P.-L. Ma, J. Shen, S. Wang, J. Sun, H. Gordon, S. Tang, J. Fast, M. Wang, Y. Gao, C. Yan, B. Singh, Z. Li, L. Huang, S. Lou, G. Lin, H. Wang, J. Jiang, A. Ding, W. Nie, X. Qi, X. Chi and L. Wang, Global variability in atmospheric new particle formation mechanisms, Nature, 2024, 631, 98–105 CrossRef CAS PubMed.
- J. Shen, B. Zhao, A. Ning, W. Nie, C. Yan, Y. Li, R. Cai, A. Saiz-Lopez, M. Shrivastava, B. Chu, D. Gao, N. Myllys, D. Yin, H. Zhang, Y. Gao, Y. Liu, X. Chi, X. Qi, Y. Zhang, Y. Liu, J. Chen, L. Wang, A. Ding, J. Jiang, X. Zhang, M. Kulmala, H. He and S. Wang, Comprehensive understanding of new particle formation in China through advanced modeling, Sci. Bull., 2026, 71, 2083–2093 CrossRef CAS PubMed.
- J. Shen, B. Zhao, S. Wang, A. Ning, Y. Li, R. Cai, D. Gao, B. Chu, Y. Gao, M. Shrivastava, J. Jiang, X. Zhang and H. He, Cluster-dynamics-based parameterization for sulfuric acid–dimethylamine nucleation: comparison and selection through box and three-dimensional modeling, Atmos. Chem. Phys., 2024, 24, 10261–10278 CrossRef CAS.
- J. Zhang, V.-A. Glezakou, R. Rousseau and M.-T. Nguyen, NWPEsSe: An Adaptive-Learning Global Optimization Algorithm for Nanosized Cluster Systems, J. Chem. Theory Comput., 2020, 16, 3947–3958 CrossRef CAS PubMed.
- J. Zhang and V.-A. Glezakou, Global optimization of chemical cluster structures: Methods, applications, and challenges, Int. J. Quantum Chem., 2021, 121, e26553 CrossRef CAS.
- Y. Chen and P. O. Dral, Aiqm2: Organic reaction simulations beyond dft, Chem. Sci., 2025, 16, 15901–15912 RSC.
- R. Benjamin, V. Sander and Š. Vaidotas, Orb-v3: atomistic simulation at scale, arXiv, preprint, 2025, 24, 1833-1847, DOI:10.48550/arXiv.2504.06231.
- M. W. Brandon, D. Misko and F. Xiang, UMA: A Family of Universal Models for Atoms, arXiv, preprint, 2025, 118, 2599-2611, DOI:10.48550/arXiv.2506.23971.
- J. Elm, J. Kubečka, V. Besel, M. J. Jääskeläinen, R. Halonen, T. Kurtén and H. Vehkamäki, Modeling the formation and growth of atmospheric molecular clusters: A review, J. Aerosol Sci., 2020, 149, 105621 CrossRef CAS.
- L. Partanen, H. Vehkamäki, K. Hansen, J. Elm, H. Henschel, T. Kurtén, R. Halonen and E. Zapadinsky, Effect of Conformers on Free Energies of Atmospheric Complexes, J. Phys. Chem. A, 2016, 120, 8613–8624 CrossRef CAS PubMed.
- L. Partanen, V. Hänninen and L. Halonen, Effects of Global and Local Anharmonicities on the Thermodynamic Properties of Sulfuric Acid Monohydrate, J. Chem. Theory Comput., 2016, 12, 5511–5524 CrossRef CAS PubMed.
Footnote |
| † Yongjian Lian, Xurong Bai, Ruoying Yuan contributed to the work equally. |
| This journal is © The Royal Society of Chemistry 2026 |

