Integrating experimental and theoretical approaches to unveil structure–bioactivity relationships in silver(I) N-substituted glycine complexes
Gabriela
Kuzderová
a,
Róbert
Gyepes
 b,
Alan
Liška
c,
Jana
Havlíčková
b,
Mária
Vilková
d,
Simona
Žiláková
e,
Martin
Kello
b,
Alan
Liška
c,
Jana
Havlíčková
b,
Mária
Vilková
d,
Simona
Žiláková
e,
Martin
Kello
 e,
Tomáš
Pagáč
f,
Petra
Olejníková
f,
Eva
Petrovová
g,
Ľudmila
Balažová
g,
Henrieta
Matajová
h,
Mária
Kožurková
h,
Danica
Sabolová
h,
Michaela
Rendošová
a and
Zuzana
Vargová
e,
Tomáš
Pagáč
f,
Petra
Olejníková
f,
Eva
Petrovová
g,
Ľudmila
Balažová
g,
Henrieta
Matajová
h,
Mária
Kožurková
h,
Danica
Sabolová
h,
Michaela
Rendošová
a and
Zuzana
Vargová
 *a
*a
aDepartment of Inorganic Chemistry, Faculty of Science, P. J. Šafárik University, Moyzesova 11, 041 54 Košice, Slovakia. E-mail: zuzana.vargova@upjs.sk
bDepartment of Inorganic Chemistry, Faculty of Science, Charles University, Hlavova 2030, 128 00 Prague, Czechia
cDepartment of Molecular Electrochemistry and Catalysis, J. Heyrovský Institute of Physical Chemistry of the CAS, Dolejškova 3/2155, 182 23 Praha 8, Czechia
dNMR laboratory, Faculty of Science, P. J. Šafárik University, Moyzesova 11, 041 54 Košice, Slovakia
eDepartment of Pharmacology, Faculty of Medicine, P. J. Šafárik University, Trieda SNP 1, 040 11 Košice, Slovakia
fDepartment of Biochemistry and Microbiology, Slovak University of Technology, Radlinskeho 9, 812 37 Bratislava, Slovakia
gDepartment of Morphological Disciplines, University of Veterinary Medicine and Pharmacy, Komenského 73, 041 81 Košice, Slovakia
hDepartment of Biochemistry, Faculty of Science, P. J. Šafárik University, Moyzesova 11, 041 54 Košice, Slovakia
First published on 18th March 2026
Abstract
N-methyl and N-glycyl glycine derivatives – namely (trimethylammonium)acetate (N,N,N-trimethylglycine, betaine, Bet), N,N-dimethylglycine (Dmg), N-methylglycine (sarcosine, Sar), and glycylglycylglycine (GlyGlyGly) – as naturally occurring glycine metabolites, were employed as stabilizing ligands for silver(I) ions, leading to the formation of water-soluble polymeric coordination compounds: {[Ag(HSar)(NO3)]}n (AgSar), {[Ag(HDmg)(NO3)]}n (AgDmg), {[Ag3(HBet)2(NO3)3]}n (AgBet), and [Ag(HGlyGlyGly)(NO3)] (AgGlyGlyGly). The composition and structures of the resulting complexes were unambiguously confirmed using attenuated total reflection Fourier-transform infrared spectroscopy (ATR-FTIR), elemental analysis, thermogravimetric analysis (TGA), and single-crystal X-ray diffraction (SC-XRD). Speciation and stability studies in relevant test and growth media were conducted using a novel multi-technique approach combining potentiometric titrations, electrochemical measurements, 1H NMR and UV-Vis spectroscopy and were additionally supported by theoretical calculations. From a biological evaluation standpoint, all complexes demonstrated significantly enhanced antistaphylococcal activity (including S. aureus L12) compared to silver(I) nitrate (AgNO3), with AgBet and AgGlyGlyGly exhibiting approximately 10-fold, and AgSar and AgDmg approximately 5-fold, increased efficacy. Moreover, AgBet and AgGlyGlyGly were nearly twice as active as silver(I) sulfadiazine (AgSD), a clinically used but poorly water-soluble antimicrobial agent. Remarkably, AgBet and AgGlyGlyGly also showed unusual potent antifungal activity, being 10- to 100-fold more effective than both AgNO3 and AgSD against Rhizopus oryzae, the causative agent of mucormycosis. In cytotoxicity assays, AgSar exhibited the greatest selectivity and sensitivity, being over three times higher than cisplatin (cisPt), against the human breast adenocarcinoma cell line MDA-MB-231. New approaches from the perspectives of structure–activity relationships and bioavailability of the novel silver(I) complexes were explored through human serum albumin (HSA) binding studies and molecular docking calculations, as well as experimental and computational evaluations of lipophilicity and additional Lipinski parameters. Furthermore, the silver(I) complexes’ action mechanism was assessed via β-galactosidase inhibition and PCR amplification inhibition in E. coli, intracellular reactive oxygen species (ROS) production, their effects on cell cycle progression and binding to ctDNA.
Introduction
Silver and its simple compounds, mainly silver(I) nitrate salt, have been used as antibacterial agents for centuries. It is well established that this antimicrobial activity is primarily attributed to silver(I) ions, regardless of whether they are delivered via simple salts, complex compounds, or silver nanoparticles (AgNPs).1 However, identifying a suitable “carrier” for silver(I) ions to ensure adequate bioavailability has posed a long-standing challenge. Achieving an optimal balance of solubility, chemical stability, ligand exchange capacity, and biological effect (for supporting and enhancing the effect) – characterized by high selectivity and low toxicity – remains a demanding task in the development of silver(I)-based therapeutics. Among the various silver-based drug development strategies being investigated, silver nanoparticles are currently the most studied. At the same time, organometallic compounds, particularly those incorporating N-heterocyclic carbene (NHC) ligands, have also shown significant potential. Despite the fact that up to 50% of commercial nanoproducts contain AgNPs, their clinical application is limited by sensitivity to oxygen and a pronounced tendency toward particle aggregation.2 Consequently, recent research has increasingly focused on developing new biomaterials to stabilize metal nanoparticles in a more environmentally and biologically friendly manner. Biosurfactants, commonly used in practice, form stabilizing coatings around nanoparticles, suppressing aggregation and enhancing long-term colloidal stability.3As an alternative to AgNPs, silver(I) complexes with NHC ligands offer promising advantages. These compounds can release silver(I) ions directly at the target site (e.g., the cell membrane), providing sustained activity.1 However, synthesizing bioavailable NHC-based complexes often requires elaborate structural modifications and synthetic routes that may be environmentally unfriendly and economically demanding.4
To date, only a limited number of silver(I)-based agents have been approved for medical use, including simple salts such as AgNO3 and Ag2SO4, coordination compounds like [AgF(NH3)2], silver(I) sulfadiazine (AgSD), Ag–albumin complexes, and silver nanoparticles (colloidal silver, Ag2O).5 Therefore, the development of coordination compounds incorporating biologically and ecologically compatible ligands – particularly those offering economic advantages – represents a promising direction. Amino acids and peptides are especially attractive in this regard, as antimicrobial peptides (AMPs) are naturally involved in the immune defence of living organisms.
Glycine, one of the simplest and most fundamental amino acids, serves as a key structural unit in proteins and exhibits diverse substitution potential.6 This versatility leads to the formation of various biologically active metabolites, including N-methylated forms such as betaine ((trimethylammonium)acetate, Bet), N,N-dimethylglycine (Dmg), and sarcosine (Sar), all of which are involved in transmethylation pathways.6,7 In addition, Sar plays a central role in numerous physiological processes, serving as a major metabolic precursor for molecules such as glutathione, creatine, purines, and serine.6 Similarly, Dmg is an essential component of one-carbon metabolism and is widely used as a dietary supplement. It has also been studied for its potential in treating neurological disorders due to its agonistic activity at the glycine-binding site of the NMDA (N-methyl-D-aspartate) receptor.8 Bet functions as a methyl group donor and osmoprotectant, and its anti-inflammatory properties have been well documented.9
Importantly, glycine and its N-methyl derivatives are physiologically tolerated and would not exhibit known toxic effects when administered as part of a therapeutic formulation. Previous studies,10–14 including our own,15–18 have shown that the combination of silver(I) ions with amino acids or dipeptides produces notable in vitro antimicrobial and antiproliferative effects—often comparable or superior to those observed for clinically used agents such as AgSD and cisPt. Among the complexes we previously investigated, silver(I) complexes with glycine and the dipeptide glycylglycine (GlyGly) were among the most active. Based on these findings, we extended our study to include N-methylated derivatives of glycine and the tripeptide glycylglycylglycine (GlyGlyGly) to examine the impact of methyl or glycyl substitution on the biological activity of their silver(I) complexes. However, a critical factor affecting their therapeutic potential is the stability of these complexes, which directly influences silver(I) ion bioavailability.
Therefore, we conducted a comprehensive study on the coordination behaviour of N-substituted glycine derivatives (Bet, Dmg, Sar, and GlyGlyGly) toward silver(I) ions. Although the solid-state structures of many such complexes are available in the Cambridge Structural Database,19 there remains a significant gap in understanding their speciation in solution under aqueous conditions.20 Accordingly, this study aimed to elucidate the solution speciation of these silver(I) complexes using an unconventional multi-technique approach that combines potentiometric, spectroscopic, electrochemical, and theoretical methods. Investigation of the complexes’ solution behaviour enabled the isolation of new coordination compounds in the solid state, followed by determination of their composition and structures by single-crystal X-ray diffraction, as well as evaluation of their stability in test and culture media commonly employed in in vitro biological assays. Furthermore, the novel biological assessment—integrating multi-technique experimental data ranging from in vitro to in vivo studies with theoretical estimations and predictions—provides a comprehensive overview of the bioavailability, potential therapeutic relevance and mechanism of action of the newly developed complexes.
Results and discussion
Complexes’ formation in aqueous solution
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β210 values correspond to the deprotonation of the carboxylic group (H2L ↔ HL + H+), while log
β210 values correspond to the deprotonation of the carboxylic group (H2L ↔ HL + H+), while log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β110 values describe the deprotonation of the amino group (HL ↔ L− + H+).
β110 values describe the deprotonation of the amino group (HL ↔ L− + H+).
Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β110 β110 |
Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β210 β210 |
Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β111 β111 |
Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β011 β011 |
Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β021 β021 |
Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β−111 β−111 |
|
|---|---|---|---|---|---|---|
| a Ref. 22. b Ref. 23. c Ref. 24. d Ref. 25. e Ref. 20. f Ref. 21. g Ref. 26. | ||||||
| Gly | 9.57a | 11.90a | — | — | — | — |
| 9.78d | 12.03d | |||||
| Ag(I)-Gly | — | — | — | 3.20a | 6.63a | — |
| — | — | — | 3.5(3)f | 7.3(3)f | — | |
| Zn(II)-Gly | — | — | — | 4.96(4)a | 9.19(8)a | — |
| Cu(II)-Gly | — | — | — | 8.19(4)a | 15.1(1)a | — |
| 8.27(1)g | 15.34(1)g | — | ||||
| Cd(II)-Gly | — | — | — | 4.25(4)a | 7.77(8)a | — |
| Fe(II)-Gly | — | — | — | 4.13a | 7.65a | — |
| Sar | 9.77 (2) | 11.82 (4) | — | — | — | — |
| 9.98 (4)a | 12.16 (1)a | — | — | — | ||
| Ag(I)-Sar | — | — | — | 2.78(6) | — | — |
| Zn(II)-Sar | — | — | — | 4.53a | — | — |
| Cu(II)-Sar | — | — | — | 7.68a | — | — |
| Cd(II)-Sar | — | — | — | — | — | — |
| Fe(II)-Sar | — | — | — | 3.52a | — | — |
| Dmg | 9.67 (1) | 11.40 (3) | — | — | — | — |
| 9.77 (4)a | 11.8a | — | — | — | ||
| 9.75b | — | — | — | — | — | |
| Ag(I)-Dmg | — | — | — | 2.69(8) | — | — |
| Zn(II)-Dmg | — | — | — | 3.97a | — | — |
| Cu(II)-Dmg | — | — | — | 7.25a | — | — |
| Cd(II)-Dmg | — | — | — | — | — | — |
| Fe(II)-Dmg | — | — | — | — | — | — |
| GlyGly | 7.94 (1) | 10.90 (2) | — | — | — | — |
| 8.08 (3)c | 11.21c | — | — | — | — | |
| 8.01 (6)d | 11.38 (8)d | — | — | — | — | |
| Ag(I)-GlyGly | — | — | — | 2.66(7) | 5.49 (4) | −6.57(4) |
| — | — | — | 2.72e | 4.98e | — | |
| Zn(II)-GlyGly | — | — | — | 3.38(8)e | 6.22(10)e | — |
| — | — | — | 3.31(9)d | 6.4(3)d | — | |
| Cu(II)-GlyGly | — | — | — | 5.55(5)e | — | — |
| 5.55(5)g | — | — | ||||
| Cd(II)-GlyGly | — | — | — | 2.89(3)e | 5.36(6)e | — |
| Fe(II)-GlyGly | — | — | — | 2.62 | — | — |
| GlyGlyGly | 7.84(1) | 11.05(2) | — | — | — | — |
| 7.89(2)c | 11.10c | — | — | — | — | |
| 7.63(7)g | 10.94(1)g | — | ||||
| Ag(I)-GlyGlyGly | — | — | 9.2(1) | 2.81(6) | 5.71(4) | −6.54(3) |
| Zn(II)-GlyGlyGly | — | — | — | 3.20(2)e | 5.81 | — |
| Cu(II)-GlyGlyGly | — | — | — | 5.05(8)e | — | — |
| 5.12(2)g | — | — | ||||
| Cd(II)-GlyGlyGly | — | — | — | 2.70 | 5.3 | — |
| Fe(II)-GlyGlyGly | — | — | — | — | — | — |
As anticipated, comparison of the N-substituted glycine derivatives acid–base properties reveals that the number of methyl and glycyl substituents significantly affects the acidity/basicity of both the carboxylate and amino groups. N-Methyl substitution leads to an increase in carboxylate group acidity, whereas glycyl substituents have the opposite effect, lowering it. In the case of the betaine ligand, bearing the –N+(CH3)3 group, the increase in carboxyl acidity is particularly pronounced; however, due to experimental limitations, reliable values for the acid–base properties of Bet could not be obtained (the estimated value according to NIST database is 1.7).20 Regarding the amino group, N-methyl substitution results in enhanced basicity, while N-glycyl substitution causes a decrease which is consistent with the known behaviour of primary and secondary amines. The ligands’ speciation is presented in Fig. S1.
Experimental data fitting using the OPIUM27 program gave the most consistent results for Ag![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) L ratios of 1
L ratios of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 and 1
2 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4. In the binary systems Ag(I)-Sar and Ag(I)-Dmg, only a single mononuclear complex species, [Ag(Sar)] and [Ag(Dmg)], respectively, was identified. Fig. 1 illustrates the distribution of these complexes as a function of pH at a 1
4. In the binary systems Ag(I)-Sar and Ag(I)-Dmg, only a single mononuclear complex species, [Ag(Sar)] and [Ag(Dmg)], respectively, was identified. Fig. 1 illustrates the distribution of these complexes as a function of pH at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 molar ratio. Both [Ag(Sar)] and [Ag(Dmg)] species begin to form at pH values above 8, following deprotonation of Sar and Dmg, reaching relative abundances exceeding 50%.
4 molar ratio. Both [Ag(Sar)] and [Ag(Dmg)] species begin to form at pH values above 8, following deprotonation of Sar and Dmg, reaching relative abundances exceeding 50%.
 | ||
Fig. 1 Distribution of Ag+ species in the Ag(I)-Sar/Dmg binary systems in a molar ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 (c(Ag(I)) = 1 mM, Sar/Dmg = 4 mM), I = 0.1 M, 25 °C. 4 (c(Ag(I)) = 1 mM, Sar/Dmg = 4 mM), I = 0.1 M, 25 °C. | ||
In contrast to N-methyl-substituted glycine ligands, the N-glycyl derivatives, in the presence of Ag(I) ions in solution, form not only the [Ag(GlyGly)] and [Ag(GlyGlyGly)] complexes, but also the bis-ligand species [Ag(GlyGly)2]− and [Ag(GlyGlyGly)2]−. Their relative distributions at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 metal-to-ligand ratio are shown in Fig. 2. For the Ag(I)-GlyGly system, the distribution reveals the presence of deprotonated [Ag(GlyGly)] species in the pH range 6–10, in correlation with the [Ag(GlyGly)2]− complex, which dominates with a maximum abundance of about 60%. In the Ag(I)-GlyGlyGly system, [Ag(GlyGlyGly)] species appear at lower abundance, while the [Ag(GlyGlyGly)2]− complex predominates between pH 7.5 and 9.5. Additionally, in both systems, hydroxo complexes [Ag(GlyGly)(OH)]− and [Ag(GlyGlyGly)(OH)]− begin to form at pH ≥ 8, with Ag2O precipitation occurring above pH 10.
4 metal-to-ligand ratio are shown in Fig. 2. For the Ag(I)-GlyGly system, the distribution reveals the presence of deprotonated [Ag(GlyGly)] species in the pH range 6–10, in correlation with the [Ag(GlyGly)2]− complex, which dominates with a maximum abundance of about 60%. In the Ag(I)-GlyGlyGly system, [Ag(GlyGlyGly)] species appear at lower abundance, while the [Ag(GlyGlyGly)2]− complex predominates between pH 7.5 and 9.5. Additionally, in both systems, hydroxo complexes [Ag(GlyGly)(OH)]− and [Ag(GlyGlyGly)(OH)]− begin to form at pH ≥ 8, with Ag2O precipitation occurring above pH 10.
 | ||
Fig. 2 Distribution of Ag+ species in the binary system Ag(I)-GlyGly/GlyGlyGly binary systems in a molar ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 (c(Ag(I)) = 1 mM, GlyGly/GlyGlyGly = 4 mM), I = 0.1 M, 25 °C. 4 (c(Ag(I)) = 1 mM, GlyGly/GlyGlyGly = 4 mM), I = 0.1 M, 25 °C. | ||
Similar to the differences observed in the acid–base properties of N-methyl and N-glycyl glycine derivatives (dissociation constants, Table 1), clear distinctions are also evident in their complexation behavior (stability constants, Table 1). While glycine forms a relatively stable [Ag(Gly)] species,21 the stability of analogous complexes decreases for N-methyl substituted derivatives (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β011). This decrease can be explained by steric effects and the increased electron density around the nitrogen atom of the amino group, resulting from the presence of N-methyl functional groups exhibiting a positive inductive effect. These factors may diminish the coordinating contribution of the nitrogen atom toward the silver(I) ion. Unlike the Ag-Gly system, where [Ag(Gly)2]− species are formed, N-methyl-substituted ligands do not give rise to bis-ligand complexes. On the other hand, N-glycyl substitutions allow not only carboxylate coordination but also the participation of the N-glycyl or N-glycylglycyl groups in the silver(I) ion coordination sphere. As shown previously,25 such interactions (and even possible ligand rearrangements in the basic pH range) enhance complex stability. This additional stabilization of the coordination sphere also contributes to the formation of the bis-ligand complexes [Ag(GlyGly)2]− and [Ag(GlyGlyGly)2]− stabilization (log
β011). This decrease can be explained by steric effects and the increased electron density around the nitrogen atom of the amino group, resulting from the presence of N-methyl functional groups exhibiting a positive inductive effect. These factors may diminish the coordinating contribution of the nitrogen atom toward the silver(I) ion. Unlike the Ag-Gly system, where [Ag(Gly)2]− species are formed, N-methyl-substituted ligands do not give rise to bis-ligand complexes. On the other hand, N-glycyl substitutions allow not only carboxylate coordination but also the participation of the N-glycyl or N-glycylglycyl groups in the silver(I) ion coordination sphere. As shown previously,25 such interactions (and even possible ligand rearrangements in the basic pH range) enhance complex stability. This additional stabilization of the coordination sphere also contributes to the formation of the bis-ligand complexes [Ag(GlyGly)2]− and [Ag(GlyGlyGly)2]− stabilization (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β021).
β021).
When comparing the complexation trends of glycine and its N-substituted derivatives with divalent metal ions (Table 1), analogous patterns are observed in species formation. However, comparison with monovalent metal ion complexes is far more difficult, as stability constant data are scarce or unavailable. To our knowledge, only glycine has documented values in this regard, with the NIST database reporting log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β011 = 1.51 for [Tl(Gly)] species and an estimated value of log
β011 = 1.51 for [Tl(Gly)] species and an estimated value of log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β011 = −0.5 for [Na(Gly)].
β011 = −0.5 for [Na(Gly)].
For the amino acids Sar and Dmg, two distinct singlets are observed, corresponding to the methylene and methyl groups: the –CH2CO2 protons (HA) and CH3NH– (Sar) or (CH3)2N– (Dmg) protons (HB) (Fig. S2 and S3). At low pH values (2.08 and 3.07 for Sar; 2.32 and 3.13 for Dmg), the signals of HA and HB indicate the presence of diprotonated forms, where both the carboxyl and amino groups are protonated. Between pH 3.88 and 7.95, the zwitterionic species dominate. At higher pH (above 7.95 for Sar and 6.89 for Dmg), significant upfield shifts (towards lower ppm values) are observed, corresponding to amino group deprotonation and confirming the predominance of the deprotonated form. These 1H NMR results are fully consistent with the potentiometric data.
The pH-dependent 1H NMR spectra of the Ag(I)-Sar and Ag(I)-Dmg binary systems (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 molar ratio) are shown in Fig. S6 and S7. At higher pH values, however, spectra could not be obtained due to precipitation. In both systems, pronounced changes in ligand signal positions are observed at pH 4.90 (Sar) and 4.83 (Dmg). To emphasize the differences between free and Ag(I)-bound Sar and Dmg, comparative diagrams of proton chemical shifts were constructed for the pH range 2–5 (Fig. 3). Within this range, no significant differences are detected between the δH values of the –CH2–CO2 (HA) and CH3NH–/(CH3)2N– (HB) protons in the complexes and those in the free ligands. This suggests that complexation is negligible between pH 2–5, in agreement with potentiometric titrations, which indicate that complex formation occurs only at higher pH (8–9) (see distribution diagrams, Fig. 1). Nevertheless, slight changes observed at low pH point to the initial involvement of the carboxylate group in Ag(I) binding, while the zwitterionic form becomes dominant at higher pH (a process not detectable under potentiometric conditions). Nevertheless, the changes observed at pH 2 (ΔδHA = 0.16/0.1 ppm Ag(I)-Sar/Dmg) (Fig. 3) indicate that, upon dissociation, the carboxylate group engages in coordination with the Ag(I) ion, a phenomenon that remains undetectable under potentiometric conditions.
4 molar ratio) are shown in Fig. S6 and S7. At higher pH values, however, spectra could not be obtained due to precipitation. In both systems, pronounced changes in ligand signal positions are observed at pH 4.90 (Sar) and 4.83 (Dmg). To emphasize the differences between free and Ag(I)-bound Sar and Dmg, comparative diagrams of proton chemical shifts were constructed for the pH range 2–5 (Fig. 3). Within this range, no significant differences are detected between the δH values of the –CH2–CO2 (HA) and CH3NH–/(CH3)2N– (HB) protons in the complexes and those in the free ligands. This suggests that complexation is negligible between pH 2–5, in agreement with potentiometric titrations, which indicate that complex formation occurs only at higher pH (8–9) (see distribution diagrams, Fig. 1). Nevertheless, slight changes observed at low pH point to the initial involvement of the carboxylate group in Ag(I) binding, while the zwitterionic form becomes dominant at higher pH (a process not detectable under potentiometric conditions). Nevertheless, the changes observed at pH 2 (ΔδHA = 0.16/0.1 ppm Ag(I)-Sar/Dmg) (Fig. 3) indicate that, upon dissociation, the carboxylate group engages in coordination with the Ag(I) ion, a phenomenon that remains undetectable under potentiometric conditions.
 | ||
Fig. 3 Comparison of 1H NMR chemical shifts of free ligands (Sar/Dmg) and the binary systems (Ag(I)-Sar/Dmg) (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4). 4). | ||
Fig. S4 and S5 show the 1H NMR spectra of GlyGly and GlyGlyGly recorded over the pH range 2–11. As previously reported,25 the dipeptide GlyGly displays two methylene signals (Fig. S4), whereas the tripeptide GlyGlyGly exhibits three (Fig. S5). The central methylene group –CH2–C(O)– located between two amide groups (HA), shows the highest chemical shift value due to deshielding by the adjacent carbonyls. This resonance undergoes only minor changes, shifting from δ = 4.07 ppm (d, J 5.94) at pH 2.21 to 4.04 ppm (s) at pH 6.75 and 4.00 ppm (s) at pH 11.12. In contrast, more pronounced shifts are observed for the methylene groups near the terminal carboxylate and amino functionalities (Fig. S5). The –CH2CO2 protons (HB) shift from δ = 4.02 ppm (d, J 5.90) at pH 2.21 to 3.88 ppm (s) at pH 6.75 and 3.79 ppm (s) at pH 11.12, while the –CH2NH2 (HC) shift from δ = 3.91 ppm (s) at pH = 2.21 to δ = 3.79 (d, J 5.94) at pH 6.75 and 3.42 ppm (s) at pH 11.12. These pH-dependent chemical shift variations confirm the stepwise deprotonation of GlyGly and GlyGlyGly, with the fully deprotonated forms predominating above pH 8.
The 1H NMR spectra of the Ag(I)-GlyGly and Ag(I)-GlyGlyGly systems (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 molar ratio) as a function of pH are shown in Fig. S8 and S9. As in the Ag(I)-Sar and Ag(I)-Dmg systems, spectra above pH 7.00 could not be recorded due to precipitation. Pronounced signal shifts are observed in the pH range 2–6. To illustrate these changes, comparative diagrams of chemical shifts for free versus Ag(I)-bound GlyGly and GlyGlyGly are presented in Fig. 4. In the Ag(I)-GlyGly system, the –CH2CO2 (HA) protons exhibit the largest shift relative to free GlyGly at pH 2–3, while the –CH2NH2 (HB) protons show the most significant changes between pH 5–6. Comparable variations in the HB and HC signals are also observed in the spectra of free and complexed GlyGlyGly (Fig. 4).
4 molar ratio) as a function of pH are shown in Fig. S8 and S9. As in the Ag(I)-Sar and Ag(I)-Dmg systems, spectra above pH 7.00 could not be recorded due to precipitation. Pronounced signal shifts are observed in the pH range 2–6. To illustrate these changes, comparative diagrams of chemical shifts for free versus Ag(I)-bound GlyGly and GlyGlyGly are presented in Fig. 4. In the Ag(I)-GlyGly system, the –CH2CO2 (HA) protons exhibit the largest shift relative to free GlyGly at pH 2–3, while the –CH2NH2 (HB) protons show the most significant changes between pH 5–6. Comparable variations in the HB and HC signals are also observed in the spectra of free and complexed GlyGlyGly (Fig. 4).
 | ||
Fig. 4 Comparing hydrogen atoms chemical shifts of the free ligands (GlyGly/GlyGlyGly) and the binary systems (Ag(I)-GlyGly/GlyGlyGly) (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4). 4). | ||
Potentiometric and 1H NMR titrations’ comparison reveals a shift in the onset of complex formation: whereas potentiometric data indicate the appearance of complexes from pH ∼6, NMR titrations show evidence of their formation already at lower pH values. A marked change in chemical shift at pH 2 (HA for Ag-GlyGly and HB for Ag-GlyGlyGly) points to coordination of Ag(I) ions by the carboxylate group, similar to what is observed in the Ag-Sar and Ag-Dmg systems, though here the effect is more pronounced. With increasing pH, additional chemical shift changes are detected for CH2 protons near the terminal amino groups of both the dipeptide and tripeptide, suggesting a possible rearrangement of coordination. This rearrangement likely involves participation of the terminal amino group in an amine–amide chelate mode, consistent with earlier findings.25 Notably, the addition of Ag(I) ions to the ligand solution induces significant changes in ligand proton chemical shifts even under mildly acidic conditions, indicating early onset of complexation.
Theoretical estimation
To assess the coordination mode of N-methyl/glycyl derivatives of glycine in aqueous solution, we used various computational approaches for relevant input structures. From the thermodynamic point of view, the cyclic structures containing both the amino nitrogen as well as one of the carboxylate oxygen donor atoms (Fig. 5) seem to be the most favourable. However, their predicted stability constants (Table 2, column OAgN) are apparently larger than those found experimentally. Moreover, they tend to increase with the number of glycyl moieties, which has also not been observed in the potentiometric measurements. | ||
| Fig. 5 Optimized structures with both amino as well as carboxylic donor groups.28 | ||
| Equilibrium | Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) K K |
||
|---|---|---|---|
| OAgN | OAgO | Exp. | |
| [Ag(H2O)2]+ + Gly− ⇌ [AgGly] + 2H2O | 5.0 | 3.5 | 3.5 |
| [Ag(H2O)2]+ + GlyGly− ⇌ [AgGlyGly] + 2H2O | 6.0 | 3.4 | 2.87 |
| [Ag(H2O)2]+ + GlyGlyGly− ⇌ [AgGlyGlyGly] + 2H2O | 8.4 | 3.0 | 2.84 |
| [Ag(H2O)2]+ + Sar− ⇌ [AgSar] + 2H2O | 5.6 | 3.5 | 2.68 |
| [Ag(H2O)2]+ + Dmg− ⇌ [AgDmg] + 2H2O | 8.0 | 4.0 | 2.78 |
On the other hand, the cyclic structures including only the two carboxylate oxygen donor atoms (Fig. 6) are expected to be less stable (Table 2, column OAgO). However, the calculated stability constants fit better the experimental ones, and additionally, they are monotone within the whole series. Hence it supports the hypothesis that the coordination environment of the silver(I) central atom prefers oxygen atoms to nitrogen atoms, similarly, like in the solid phase.
 | ||
| Fig. 6 Optimized structures with the carboxylic donor group only.28 | ||
Complexes’ isolation in solid state
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 molar ratio of reactants. For AgGlyGlyGly, varying the reactant ratio (1
1 molar ratio of reactants. For AgGlyGlyGly, varying the reactant ratio (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 to 1
1 to 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2, Ag
2, Ag![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ligand) consistently produced solid products of the same composition. All synthesized complexes are air-stable, were stored in the absence of light, and dissolve in polar solvents.
ligand) consistently produced solid products of the same composition. All synthesized complexes are air-stable, were stored in the absence of light, and dissolve in polar solvents.
{[Ag(HSar)(NO3)]}n (AgSar). The crystal structure of coordination compound AgSar is depicted in Fig. 7. The complex crystallizes in the monoclinic lattice with space group P21/n. The asymmetric unit of the complex consists of one silver(I) ion, one molecule of HSar as zwitterion and one coordinated nitrate anion.
 | ||
| Fig. 7 Crystal structure and atomic labelling scheme (i = −x, 1 − y, 1 − z) for AgSar with atomic displacement ellipsoids drawn at 50% probability level. | ||
The crystal structure of the AgSar complex reveals that the Ag1 atom is coordinated by two oxygen atoms O1i (i = −x, 1 − y, 1 − z) and O2 from two different HSar molecules and one oxygen atom O3 from the nitrate anion in a distorted trigonal geometry. Ag(I) ions are bridged through two HSar molecules, and the Ag⋯Ag distance between the two Ag(I) ions is 2.8463(2) Å (Fig. 7, blue dashed bonds). These contacts are shorter than the sum of their van der Waals radii (3.44 Å), indicating the presence of significant argentophilic interactions.29 Similar argentophilic interactions have also been observed in other silver(I) complexes based on glycine, such as [Ag2(HGly)2]n(NO3)2n (AgGly), [Ag3(Gly)2NO3]n and [Ag(GlyGly)]n(NO3)n (AgGlyGly).17,21,30
As shown in Fig. 7, HSar ligands are coordinated to Ag(I) ions through their carboxylate functional group in a syn–syn coordination mode. Selected bond distances and angles are given in Table S2 in SI. The bond distances between the Ag(I) ions and the syn-coordinated oxygen atoms from the carboxylate groups are 2.2770(9) Å (for Ag1–O1i) (i = −x, 1 − y, 1 − z) and 2.2222(11) Å (for Ag1–O2). Based on a search in the CSD,19 similar bond distances were also observed in another metal-sarcosinate complex [Mn(Sar)2(H2O)2]Cl2 with a syn–anti bidentate bridging coordination mode of the HSar ligand.31 The largest angle value in the complex AgSar is observed between the atoms O1i–Ag1–O2 (148.660(39)°).
The bond lengths C1–O1 and C1–O2 (1.2631(19) Å and 1.2485(18) Å) in the structure of the AgSar complex are consistent with bond lengths observed in the free zwitterionic form of the HSar ligand.32 Additionally, in the structure of the AgSar complex, the value of torsion angle C1–C2–N1–C3 62.721(162)° indicates a gauche conformation of HSar ligand, which was also observed in the crystal structure of the [CdCl2(HSar)] complex.33
In the neutral crystal form of the AgSar complex the nitrato ligand including N2 is coordinated to the Ag1 atom through one oxygen atom O3 in monodentate coordination mode with a bond length of 2.4095(13) Å. The same coordination mode of nitrate anion to silver(I) ions was also reported in other silver(I) amino acidate complexes.15,16,18
The 3D crystal structure is stabilized by N–H⋯O and C–H⋯O intermolecular hydrogen bonding interactions (Fig. S10, orange dashed bonds). The list of all possible hydrogen bonds is summarized in Table S3 in SI.
{[Ag(HDmg)(NO3)]}n (AgDmg). The crystal structure of the AgDmg complex is depicted in Fig. 8. Like the AgSar complex, the coordination compound AgDmg crystallizes in a monoclinic lattice, but with the space group P21/c. Moreover, the composition of the asymmetric unit is the same as that of the AgSar complex.
 | ||
| Fig. 8 Crystal structure and atomic labelling scheme (i = 1 − x, 1 − y, 1 − z; ii = 2 − x, 1 − y, 1 − z) for AgDmg with argentophilic interactions (blue dashed lines). | ||
While distorted trigonal geometry around silver(I) ions was observed in the crystal structure of complex AgSar, single-crystal X-ray analysis clearly revealed tetracoordination around silver(I) ions in the complex AgDmg. Atom Ag1 is coordinated by three oxygen atoms O1i, O2 and O2ii (i = 1 − x, 1 − y, 1 − z; ii = 2 − x, 1 − y, 1 − z) from three different HDmg molecules and also by one oxygen atom O3 belonging to a coordinated nitrate anion. Similar to the AgPro and AgHyp complexes16 the calculated value of the parameter τ434 for the Ag1 ion in the AgDmg complex, without consideration of Ag⋯Ag interactions, is 54%, which indicates a distorted seesaw (C2v) geometry.
The coordination mode of carboxylate functional group to silver(I) ions in AgDmg complex is also slightly different from the previous coordination compound AgSar. HDmg molecules are coordinated to silver(I) ions through the carboxylate group in syn–syn–anti bidentate bridging coordination mode resulting in the formation of a 1D polymeric chain propagating along the a crystallographic axis with significant Ag⋯Ag interactions of 2.9056(3) Å (Fig. 8 and Fig. S11 blue dashed bonds).
The Ag–O bond distances of syn- coordinated oxygen atoms are 2.300(1) Å (for Ag1–O1i (i = 1 − x, 1 − y, 1 − z)) and 2.2627(10) Å (for Ag1–O2), which are comparable to the distances in the complex AgSar. On the other hand the bond distance between the Ag(I) ion and anti-coordinated oxygen atom O2ii (ii = 2 − x, 1 − y, 1 − z) is slightly longer (2.4293(11) Å). Moreover, in the case of the carboxylate functional group, the O1–C1–O2 bond angle is 125.05(14)° These values are in accordance with values reported in published structures of complexes.35,36 Selected bond distances and angles are provided in Table S4.
Similarly to the AgSar complex, the nitrate anion is monodentately coordinated to Ag(I) ion through one oxygen atom, with an observed Ag1–O3 bond length of 2.4388(11) Å. The 3D crystal structure is stabilized by N–H⋯O, C–H⋯O and N–H⋯N intermolecular hydrogen bonding interactions (Fig. S12, orange dashed bonds). For detail, the H-bond geometry is listed in Table S5 in SI.
{[Ag3(HBet)2(NO3)3]}n (AgBet). The crystal structure of complex AgBet is depicted in Fig. 9. In contrast to the AgSar and AgDmg complexes, compound crystallizes in an orthorhombic lattice with space group P21212. Moreover, the asymmetric unit consists of three crystallographically independent silver(I) ions, two HBet molecules as zwitterions and three coordinated nitrate anions.
 | ||
| Fig. 9 Crystal structure and atomic labelling scheme (i = x, y, 1 + z, 1 − z; ii = x, y, −1 + z) for AgBet with argentophilic interactions (blue dashed lines). | ||
According to Fig. 9, it is evident that the Ag1 ion is coordinated by three oxygen atoms O2, O3 and O5i (i = x, y, 1 + z) in distorted trigonal geometry. On the other hand, Ag2 and Ag3 ions in the AgBet complex exhibit tetrahedral geometry. The Ag2 ion is tetracoordinated by three oxygen atoms O1, O4 and O2ii (ii = x, y, −1 + z) originating from three different HBet molecules and one oxygen atom O5 belonging to one of the coordinated nitrate anions. Similarly, the Ag3 ion is tetracoordinated by four oxygen atoms O5i (i = x, y, 1 + z), O6, O9 and O11, all of which, however, originate from nitrate anions. As in the case of complex AgDmg, the shape of the coordination polyhedron can be determined using the structural parameter τ4.34 The calculated values of τ4 for the Ag2 and Ag3 ions excluding consideration of Ag⋯Ag interactions are 61% and 66%, respectively, indicating a distorted seesaw (C2v) geometry. Based on the CSD,19 nine silver(I) complexes with betaine are recorded, while tetracoordination around the Ag(I) ion is observed in seven compounds.
Similarly to the AgDmg, AgGly and AgGlyGly complexes, HBet molecules adopt a syn–syn–anti coordination mode to Ag1 and Ag2 atoms through their carboxylate functional group. The Ag–O bond distances of the syn–syn coordinated oxygen atoms are in the range of 2.1950(22)–2.3289(19) Å. In contrast, the Ag–O distances of the anti-coordinated oxygen atoms are elongated, ranging from 2.4137(17) Å to 2.4506(23) Å. The observed Ag–O bond distances are in good agreement with those reported for another silver(I)–betaine complex [{Ag2L′2(H2O)2(NO3)2}n] (L′ = Bet).37 Selected bond distances and angles are given in Table S6 in SI.
As mentioned above, the asymmetric unit of the AgBet complex also contains three coordinated nitrate anions. Similarly to the AgSar and AgDmg complexes, two nitrate anions including N4 and N5 atoms are monodentately coordinated to the Ag3 atom with bond distances 2.4508(24) Å (for Ag3–O9) and 2.5591(28) Å (for Ag3–O11). On the other hand, the third nitrate anion including N3 atom adopts multi-bridging coordination mode to Ag2, Ag3 and Ag3ii atoms via two oxygen atoms O5 and O6 with Ag–O bond distances in the range of 2.4516(25)–2.5883(26) Å. The bridging properties of the mentioned nitrate anion and also HBet ligands contribute to the formation of a 1D polymeric chain propagating along the c axis (Fig. S13, blue dashed bonds). The Ag⋯Ag distance within the polymeric chain is 2.8677(4) Å, which suggests the presence of argentophilic interactions.29 Furthermore, significant argentophilic interactions were observed between silver(I) ions of adjacent layers, resulting in a double-layered structure (Fig. S13, green dashed bonds).
The 3D molecular structure of the AgBet complex is stabilized exclusively by 15 types of C–H⋯O hydrogen bonds. A complete list of all possible hydrogen bonds is provided in Table S7 in the SI, and the possible C–H⋯O hydrogen bonds are illustrated in Fig. S14 (orange dashed bonds).
Complexes’ stability in biological testing and growth media
In parallel, 1H NMR spectra of the corresponding free ligands were recorded under identical conditions. The pH values of the ligand solutions matched those of the complexes, except for the Bet ligand solution, which was adjusted from pH 6 to pH 4 to ensure comparability. Comparison of the free ligand and complex spectra indicates that coordination of the central atom induces only minor proton signal shifts. As discussed previously, coordination predominantly involves the carboxylate group, without additional stabilizing interactions such as multidentate chelation. Consequently, the observed spectral changes resemble those typical of deprotonation. Nevertheless, ligand coordination markedly influences the biological properties of the complexes, as detailed in the bioassay section.
In all cases, the discussed redox processes were controlled by transport (linear limiting current dependence on concentration in DCP or RDE, linear peak current dependence on square root of scan rate in CV).
As it follows from the polarographic data (Table 3 and Fig. S21), the only reduction signal related to the complexes was observed as single one-electron wave at half-wave potentials near to 0.14 (±0.01), 0.16 (±0.01), and 0.18 (±0.01) V versus SCE at concentrations of 0.4, 1.0, and 2.0 mM, respectively. Apart from that, the polarograms sometimes contain another signal at more positive potentials due to the proximity of the anodic mercury potential-window limit. To avoid this complication, analogical measurements using the glassy carbon rotating disc electrode were conducted (Table 4 and Fig. S22). However, the polarization curve shape acquired on the solid (GC) electrode is non-symmetric, and in some cases two distinct waves can be identified. Anyway, in both cases (DCP, RDE), the observed response includes single one-electron reduction corresponding to reduction of silver(I) aqua ion yielding the metallic or amalgamated silver eqn (1).
| Ag+(aq) + e− → Ag(s) | (1) |
| Substance | E 1/2/V vs. SCE | ||
|---|---|---|---|
| c/mM | 0.4 | 1.0 | 2.0 |
| AgNO3 | 0.14 | 0.16 | 0.17 |
| AgGly | 0.15 | 0.17 | 0.18 |
| AgGlyGly | 0.14 | 0.15 | 0.19 |
| AgGlyGlyGly | 0.13 | 0.16 | 0.19 |
| AgSar | 0.14 | 0.15 | 0.19 |
| AgDmg | 0.14 | 0.16 | 0.17 |
| Substance | E 1/2/V vs. SCE | ||||
|---|---|---|---|---|---|
| c/mM | 0.4 | 1.0 | 4.0 | ||
| AgNO3 | 0.14 | 0.26 | 0.26 | 0.30 | |
| AgGly | 0.26 | 0.28 | 0.31 | ||
| AgGlyGly | 0.22 | 0.25 | 0.30 | ||
| AgGlyGlyGly | 0.17 | 0.26 | 0.23 | 0.24 | |
| AgSar | 0.18 | 0.26 | 0.22 | 0.28 | 0.29 |
| AgDmg | 0.15 | 0.27 | 0.23 | 0.28 | 0.30 |
The half-wave potential shift observed in various analyte concentrations can be addressed to the fact that one of the redox forms always has activity of (1) (solid silver, ared), while the other (aqueous silver(I) species, aox) is different in each experiment series (2). With respect to the definition of a Nernstian half-wave potential E1/2, the surface concentration of the analyte c(x=0)Ag+ is half of the corresponding concentration in bulk  (3). Accordingly, the half-wave potentials measured in the 4 mM solutions should be shifted by 0.06 V positively compared to the 0.4 mM concentration. Such an estimate is in satisfactory agreement with the presented results (Tables 3, 4 and Fig. S21, S23).
(3). Accordingly, the half-wave potentials measured in the 4 mM solutions should be shifted by 0.06 V positively compared to the 0.4 mM concentration. Such an estimate is in satisfactory agreement with the presented results (Tables 3, 4 and Fig. S21, S23).
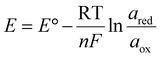 | (2) |
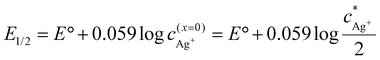 | (3) |
To confirm the reduction mechanism of the silver(I) complexes in solution, CV measurements were performed. The cyclic voltammetry (Fig. S23) shows a distinct cathodic peak (in the range from +0.23 to +0.32 V vs. SCE), followed by an anodic peak (+0.43 to +0.46 V vs. SCE), which is a response typical for the silver(I) aqua ion, i.e. one-electron reduction yielding the metallic silver. Similar electrochemical behaviour and one-electron reduction process was also reported in the case of another silver(I) complex [Ag2(NO3)(tpmc)]NO3·1.7H2O (tpmc = N,N′,N″,N‴-tetrakis(2-pyridylmethyl)-1,4,8,11-tetraazacyclotetradecane)40 and a series of silver(I) camphorimine complexes with the general formula [Ag(NO3)(YL)].41
UV-Vis spectra were subsequently compared for the different systems (growth medium/growth medium + complex/growth medium + ligand/growth medium + AgNO3) (Fig. S25). The results show that absorbance differences are more pronounced in the antimicrobial media (SB, MHB) than in the cytotoxic media (RPMI, RPMI(SA)). Moreover, the spectra of AgSar, AgDmg, AgBet, and AgGlyGlyGly differ distinctly from those of AgNO3 and the corresponding free ligands (Sar, Dmg, Bet, GlyGlyGly). These observations support the biological activity results of the synthesized complexes, as compared to the activities of the free ligands or the silver(I) ion, discussed in the following section.
Complexes’ bioassays
| S. aureus | S. aureus | E. coli | C. parapsilosis | |
|---|---|---|---|---|
| L12 | ||||
| MIC90 | MIC90 | MIC90 | MIC80 | |
| a Ref. 38. b Ref. 30. | ||||
| AgSar | 5s | 5s | 5s | 50s |
| AgDmg | 5s | 5s | 2.5s | 25s |
| AgBet | 2.5s | 2.5s | 2.5s | 10 |
| AgGlyGlyGly | 2.5s | 2.5s | 2.5s | >50 |
| Sar | >1000 | >1000 | >1000 | >1000 |
| Dmg | >1000 | >1000 | >1000 | >1000 |
| Bet | >1000 | >1000 | >1000 | >1000 |
| GlyGlyGly | >1000 | >1000 | >1000 | >1000 |
| AgNO3 | 25 | 25 | 25 | 100 |
| AgSD | 4 | 4 | 2 | 5 |
{[Ag2(HGly)2]}n(NO3)2n![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) a a |
5 | — | 5 | 11 |
{[Ag4(HAla)4(NO3)3]}n(NO3)n![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) a a |
6 | — | 4 | 12 |
{[Ag(HGlyGly)]}n(NO3)n![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) a a |
22 | — | 10 | 30 |
{[Ag3(Gly)2NO3]}n![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) b b |
164 | — | 164 | — |
To quantitatively assess their efficacy, we determined the minimum inhibitory concentration required to suppress 90% of cell growth (MIC90) for each complex and ligands. While all compounds were capable of completely inhibiting bacterial growth (data not shown), the concentrations required to achieve full inhibition varied considerably. That is why we chose to express antibacterial activity using MIC90 values, which provide a more standardized basis for comparison. These values represent the lowest concentrations required to inhibit 90% of bacterial growth relative to the control (1% EtOH; 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 dilution).
1 dilution).
In line with these findings, all silver(I) complexes consistently exhibited a bacteriostatic effect. To further evaluate their antimicrobial potential, we extended our analysis to include antifungal activity. This broader screening aimed to assess whether the observed antibacterial effects could also be replicated in fungal models.
The results of the antifungal assays revealed that most fungal strains were susceptible to the tested silver(I) complexes. Notably, R. oryzae, a representative of lower fungi, displayed susceptibility. For the remaining strains, growth inhibition of up to 80% was observed, and corresponding MIC80 values are presented in Table 5.
A closer look at the individual strains revealed some interesting patterns. As summarized in Table 5, all bacterial strains tested were susceptible to the silver(I) complexes. S. aureus showed notable susceptibility, particularly to AgDmg and AgGlyGlyGly, both of which achieved MIC90 values of 2.5 µM (Table 5 and Fig. S26). On the other hand, it is shown in Table 5 that all the tested complexes could be considered as compounds with potent comparable antistaphylococcal activity. Moreover, both the antibiotic-sensitive Staphylococcus aureus strain and the methicillin-resistant strain MRSA exhibited comparable susceptibility to the tested silver(I) complexes. No major differences in the inhibition profiles were observed between the two strains, indicating that the antimicrobial activity of the complexes is not affected by the presence of classical β-lactam resistance mechanisms. The sensitivity of MRSA to the silver(I) complexes suggests that their mode of action bypasses conventional antibiotic resistance pathways.
A comparable trend was observed in E. coli, a Gram-negative model organism, with AgBet, and AgGlyGlyGly showing MIC90 values of 2.5 µM (Fig. S27). These data indicate that the silver(I) complexes possess broad-spectrum antibacterial activity. To investigate their effects under dynamic conditions, we also assessed bacterial susceptibility when the silver(I) complexes were added during the exponential growth phase. This experimental setup better mimics real-time exposure and allows for a more realistic estimation of compound efficacy. While bacterial growth was effectively stopped after complex addition, the required concentrations were substantially higher – at least 10-fold greater than the MIC90 values obtained in bacteriostatic growth (Fig. S26, S27B, D; S28, S29C, D).
Fungal susceptibility was similarly evaluated. The yeast C. parapsilosis responded comparably to other fungal strains (Table 6), with growth dynamics in the presence of silver(I) complexes illustrated in Fig. S30. Among all tested complexes, highest antifungal activity was found at a concentration of 10 µM. Interestingly, R. oryzae emerged as the most sensitive strain among the tested fungi, with MIC90 values of 10 and 15 µM depending on the complex used (Table 6). The remaining model fungal strains exhibited consistent responses, with MIC80 values ranging from 25 to 50 µM, particularly in the case of AgBet.
| R. oryzae | A. alternata | M. gypseum | |
|---|---|---|---|
| MIC90 | MIC90 | MIC90 | |
| a Ref. 38. | |||
| AgSar | 15 | 25 | 25 |
| AgDmg | 15 | 25 | 25 |
| AgBet | 10 | 50 | 50 |
| AgGlyGlyGly | 10 | >100 | >100 |
| Sar | >1000 | >1000 | >1000 |
| Dmg | >1000 | >1000 | >1000 |
| Bet | >1000 | >1000 | >1000 |
| GlyGlyGly | >1000 | >1000 | >1000 |
| AgNO3 | 100 | 100 | 500 |
| AgSD | 1000 | >1000 | 50 |
{[Ag2(HGly)2]}n(NO3)2n![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) a a |
50s | 10s | — |
{[Ag4(HAla)4(NO3)3]}n(NO3)n![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) a a |
50s | >100 | — |
{[Ag(HGlyGly)]}n(NO3)n![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) a a |
50s | >100 | — |
The AgSar, AgDmg, AgBet and AgGlyGlyGly complexes’ bioeffect comparison, in a first approximation (it is necessary to keep in mind the experimental conditions),38 suggests a higher antimicrobial effect of AgBet and AgGlyGlyGly against S. aureus (Table 5) and of AgDmg, AgBet and AgGlyGlyGly against E. coli than the analogous complexes with glycine, alanine (C-methyl derivative of glycine) and GlyGly-(N-glycyl derivative of glycine). Moreover, all four complexes are more effective against R. oryzae than the above-mentioned published complexes, which expands their potential for use in antifungal therapy (Table 6).
Since the activity of silver(I) complexes in both cases (antimicrobial and antifungal) is very similar, we assume that the ligand is very important for transport into prokaryotes or eukaryotes. The ligands are derivatives of glycine and are probably transported into the cells by the glycine transporter. Considering our previous results of silver(I) complexes with other amino acid or di- or tripeptide ligands, it seems that glycine as the terminal amino acid of the peptide provides an advantage in bioavailability as the amino acid with the highest affinity for the amino acid transporter.38
| HCT116 | MDA-MB-231 | A549 | A2058 | PaTu 8902 | HepG2 | Jurkat | BJ-5ta | |
|---|---|---|---|---|---|---|---|---|
| HCT116 (human colorectal carcinoma), MDA-MB-231 (human mammary gland adenocarcinoma), A549 (human alveolar adenocarcinoma), A2058 (human metastatic melanoma), PaTu 8902 (human pancreatic adenocarcinoma), HepG2 (human hepatocellular carcinoma), Jurkat (human leukaemic T-cell lymphoma) and human dermal fibroblasts (BJ-5ta). | ||||||||
| AgSar | 21.2 ± 4.9 | 7.8 ± 0.1 | 32.2 ± 0.1 | 26.7 ± 3.4 | 36.1 ± 3.0 | 29.9 ± 1.8 | 6.0 ± 0.1 | 33.2 ± 0.9 |
| AgDmg | 33.5 ± 0.2 | 8.4 ± 0.6 | 33.9 ± 1.6 | 23.6 ± 4.5 | 37.1 ± 2.5 | 30.7 ± 1.5 | 6.0 ± 0.1 | 33.6 ± 0.6 |
| AgBet | 33.3 ± 1.9 | 11.7 ± 1.3 | 39.2 ± 6.3 | 30.5 ± 3.3 | 27.7 ± 6.4 | 21.6 ± 5.3 | 6.8 ± 1.0 | 12.7 ± 1.5 |
| AgGlyGlyGly | 32.4 ± 1.2 | 18.4 ± 6.3 | 32.5 ± 3.0 | 32.8 ± 3.6 | 29.2 ± 4.9 | 24.4 ± 4.7 | 10.5 ± 1.5 | 25.2 ± 11.3 |
| Sar | >100 | >100 | >100 | >100 | >100 | >100 | >100 | >100 |
| Dmg | >100 | >100 | >100 | >100 | >100 | >100 | >100 | >100 |
| Bet | >100 | >100 | >100 | >100 | >100 | >100 | >100 | >100 |
| GlyGlyGly | >100 | >100 | >100 | >100 | >100 | >100 | >100 | >100 |
| AgNO3 | 5.6 ± 1.4 | 6.1 ± 0.3 | 7.8 ± 1.8 | 7.4 ± 1.3 | 6.5 ± 0.6 | 5.5 ± 0.1 | 5.7 ± 0.0 | 10.5 ± 2.1 |
| cisPt | 14.5 ± 2.5 | 26.7 ± 5.0 | 17.3 ± 2.2 | 18.8 ± 5.5 | 20.7 ± 3.1 | 14.0 ± 2.8 | 6.2 ± 0.1 | 31.0 ± 0.7 |
A reliable comparison can perhaps be considered with our previous results, where the AgGly complex is more effective against the tested cancer cell lines (HCT116, A2058, PaTu8902, Jurkat; IC50 = 6.3, 4.4, 4.0, 4.3)21 than Ag(I) complexes with N-glycine derivatives (Table 7). On the other hand, AgGlyGly (IC50 = 34.5)17 is similarly effective as complexes with glycine derivatives against HCT116 and less effective against MDA-MB-231 and Jurkat (IC50 = 13.0 and 14.8)17 than AgSar, AgDmg and AgBet complexes (Table 7). Since the effect against MDA-MB-231 can be considered interesting, we compared our data with [Ag3(Gly)2NO3]n, but also with other carboxylates in the work of Cristina N. Banti's group,30 which observed lower IC50 values (2.7 µM for [Ag3(Gly)2NO3]n; 4.9 µM for [Ag(salH)(tpAs)3], salH = salicylic acid, tpAs = triphenylarsine; 3.6 µM for {[Ag(p-HO-bzaH)(tpAs)3](dmf)}, p-HO-bzaH = p-hydroxy-benzoic acid).
Moreover, the complexes' effect on angiogenesis was investigated using the in ovo method on the chorioallantoic membrane, and the results were analysed using software enhanced with artificial intelligence, IKOSA.50 Angiogenesis is the physiological process of developing new blood vessels from pre-existing vasculature. From the physiological point of view, it plays a crucial role in growth, maturation and wound healing.51 This process is very important in diseases such as diabetic retinopathy,52 rheumatoid arthritis,53 cardiac hypertrophy54 or tumorigenesis.55 A tumour with a diameter greater than 4 mm cannot grow without a blood supply. Deprivation of angiogenesis is one of the supported therapeutic strategies for cancer treatment.
The concentration 30 µM was set as described earlier. A previous study on the cytotoxic potential showed the ineffectiveness of the ligands alone, so we decided to test only the final complexes (AgBet, AgDmg, AgGly, AgGlyGly, AgGlyGlyGly). Three parameters were assessed: the number of branching points, the total length of vessels, and the mean thickness of vessels.56
The number of branching points provides information about the development of new vessels from pre-existing vessels.56 After 72 hours, the control group exhibited a significantly higher increase in this parameter compared to groups treated with the tested substances. The substance AgGlyGly had the highest effect angiogenesis, which led to 21% reduction of branching points. It was followed by AgBet (12% reduction), AgDmg and AgGly (both 7%), AgGlyGlyGly (2%). This indicates that the tested substances reduce neovascularization and decrease the amount of vessels compared to the control.
The total length of vessels building the vascular network increases during normal angiogenesis. This increase occurs because of newly formed vessels.57 Such an increase was observed in the case of the control and all tested substances. However, the total length was lower in the case of the tested substances (AgGlyGly, AgBet, AgDmg, AgGly) compared to the control. This indicates that the tested substances impede neovascularization, resulting in reduced vascular density and a lower amount of vessels.
The newly formed vessels are generally thinner, which leads to a smaller total thickness of new vessels developed compared to the initial vessel thickness before the application of tested substances.58 Such a reduction was observed in all groups, including the control. However, the vessels developed after treatment with the tested substances were thicker than unaffected vessels. This suggests that the tested substances negatively affect the quality of vessels.
The reduction in vessel length, mean thickness, and number of branching points, collectively confirm that the tested substances adversely affect both the quality and quantity of vessels. Many biosynthesized and commercial silver nanoparticles also reduce the process of angiogenesis.59 All three tested parameters suggested that silver(I) complexes with amino acids have the potential to reduce neovascularization, which could be beneficial for cancer therapy.
Structure–activity relationship
Regarding the structure–activity relationship of the complexes, which differ in N-methylation (glycylation) of the ligand framework, we employed methods designed to yield both reliable and interpretable results. Specifically, we determined the binding constant (Kb) and binding site of silver(I) complexes with human serum albumin (HSA)—the major blood protein involved in numerous physiological processes. In addition, the n-octanol/water partition coefficient (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P), a key indicator of lipophilicity in drug discovery that strongly influences absorption, distribution, metabolism, excretion, and toxicity, was measured. Further parameters from Lipinski's rule of five (molecular weight, MW < 500; hydrogen bond acceptors, HBA < 10; hydrogen bond donors, HBD < 5; log
P), a key indicator of lipophilicity in drug discovery that strongly influences absorption, distribution, metabolism, excretion, and toxicity, was measured. Further parameters from Lipinski's rule of five (molecular weight, MW < 500; hydrogen bond acceptors, HBA < 10; hydrogen bond donors, HBD < 5; log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P < 5) were also evaluated. Moreover, interesting results from in vitro antimicrobial tests led us to consider the possible mechanism of action of silver(I) complexes. Therefore, we decided to perform several in vivo measurements, such as β-galactosidase activity inhibition and PCR amplification inhibition of E. coli by silver(I) complexes, and intracellular Reactive Oxygen Species (ROS) production induced by silver(I) complexes. Moreover, in view of the more significant cytotoxicity of the Ag(I) complexes against breast cancer compared with the other tested cell lines, we decided to evaluate their effect on cell cycle phases and on ctDNA to explore a possible mechanism of action against the selected breast cancer cell line. Together, these data provide more detailed insights into the bioavailability and potential action mechanism of the studied complexes.60–63
P < 5) were also evaluated. Moreover, interesting results from in vitro antimicrobial tests led us to consider the possible mechanism of action of silver(I) complexes. Therefore, we decided to perform several in vivo measurements, such as β-galactosidase activity inhibition and PCR amplification inhibition of E. coli by silver(I) complexes, and intracellular Reactive Oxygen Species (ROS) production induced by silver(I) complexes. Moreover, in view of the more significant cytotoxicity of the Ag(I) complexes against breast cancer compared with the other tested cell lines, we decided to evaluate their effect on cell cycle phases and on ctDNA to explore a possible mechanism of action against the selected breast cancer cell line. Together, these data provide more detailed insights into the bioavailability and potential action mechanism of the studied complexes.60–63
 | ||
| Fig. 11 Fluorescence spectra of HSA upon the addition of complex AgBet in 10 mM Phosphate-buffered saline (pH = 7.4). Inset: Stern–Volmer plot. | ||
The values of the Stern–Volmer constant (KSV, in M−1) are determined by the slope of the diagram F0/F vs. [Q]. Kq is the bimolecular quenching rate constant, and τ0 is the average lifetime of the molecule without quencher. Since the fluorescence lifetime of protein is 10−8 s, the Kq values were calculated using the formula Kq = KSV/τ0.64,65 The obtained Kb constants for investigated silver(I) complexes AgSar, AgDmg, AgBet and AgGlyGlyGly were determined in the range from 2.60 × 104 M−1 to 7.71 × 104 M−1. All constants are shown in Table 8. The binding constant (Kb) and the number of binding sites is obtained by the double logarithm regression curve (modified Stern–Volmer eqn (3)) in which it is assumed that silver(I) complex binds independently to equivalent sites of the protein macromolecule. The slope and intercept of this logarithmic relationship represent the number of binding sites and Kb values, respectively (Table 8 and Fig. S34). The interaction between HSA and silver(I) complexes is of medium strength and the highest KSV and Kb constant was determined for the AgBet complex. The binding parameter n ≈ 1 designates the existence of only one binding site for the investigated Ag(I) complex in HSA.
| Complex | AgBet | AgGlyGlyGly | AgDmg | AgSar |
|---|---|---|---|---|
| K sv × 103 [M−1] | 11.72 | 2.39 | 5.13 | 3.01 |
| K q × 1011 [M−1 s−1] | 11.72 | 2.39 | 5.13 | 3.01 |
| K b × 104 [M−1] | 7.71 | 3.23 | 5.87 | 2.60 |
| n | 1.08 | 0.96 | 1.04 | 0.94 |
The results of fluorescence measurements show that the fluorescence intensity of HSA was sufficiently reduced by the new synthesized complexes. It follows from these results that the investigated silver drugs are capable of efficiently binding to HSA. The calculated binding constants are slightly higher than those determined for silver(I) complexes with 1,2-bis(4-pyridyl)ethane/ethene66 and thiophene-2-carboxylate ligand67 but are of the same order of magnitude.
HSA is known to contain three homologous structural domains, designated I, II, and III. Each domain is additionally divided into two subdomains A and B. The main drug-binding sites in HSA, known as site I and site II, are positioned in the hydrophobic cavities of subdomains IIA and IIIA.68
We additionally examined the binding site of silver(I) complexes on HSA using the binding site markers warfarin and ibuprofen. These two site marker probes are routinely used to localize the binding site on serum albumin. Most small molecules bind to this protein at Sudlow site I (subdomain IIA, e.g. warfarin) and Sudlow site II (subdomain IIIA, e.g. ibuprofen). The percentage of probe displacement (Fig. 12) was calculated from the plot of F/F0 × 100% versus probe/HSA, according to the method proposed by Sudlow et al.69 As can be seen from Fig. 12 and Fig. S35A, B, we observed a more significant decrease in the fluorescence for the AgSar complex in the presence of warfarin than in the presence of ibuprofen, therefore we suppose that the silver(I) complex AgSar binds to the same site in HSA as warfarin, i.e. subdomain IIA, Sudlow site I in HSA. Similar results were obtained by Y.-Y. Fang et al. for natural bioactive compound Ginkol.70 Furthermore, for AgBet, AgGlyGlyGly and AgDmg complexes, when both site markers were present, the fluorescence decreased approximately equally. Consequently, we assume that these complexes can bind to both sites (site I and site II of HSA) (Fig. S36–38, 39A, B, 41A and B). Similar multiple binding to HSA was found by Atli Thorarensen et al.71 for a fluorescent antibacterial analogue.
 | ||
| Fig. 12 Bar graph for complex AgSar presenting site markers displacement assay by ibuprofen (IBU) and warfarin (WAR). | ||
Moreover, the binding of our silver(I) complexes to Sudlow site I (warfarin site) or Sudlow site II (ibuprofen site) is also reflected in the binding constants Kb determined by using eqn (7) (in Experimental) (Table S9), which fall in the range of 0.76–14.23 × 103 M−1. It is known that the binding constant decreases when the complex binds to the same site as the corresponding marker, due to competition with the marker. All determined binding constants are lower than those reported in the literature for warfarin (6.17 × 105 M−1)72 and for ibuprofen (2 × 106 M−1).73
| Ligand | Affinity/kJ mol−1 (best) | Affinity/kJ mol−1 (worst) |
|---|---|---|
| AgGly | −17.3 | −14.4 |
| AgGlyGly | −21.8 | −19.0 |
| AgGlyGlyGly | −18.9 | −14.7 |
| AgSar | −16.4 | −13.7 |
| AgDmg | −16.6 | −14.2 |
| AgBetNO3 | −23.1 | −16.2 |
| Ibuprofen | −29.4 | −21.6 |
| R-Warfarin | −37.1 | −31.8 |
| S-Warfarin | −37.7 | −30.8 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P (either experimentally76 or theoretically63), but the experimentally determined and theoretically estimated correlation seems to be the more reliable approach. The octanol–water partition coefficients (log
P (either experimentally76 or theoretically63), but the experimentally determined and theoretically estimated correlation seems to be the more reliable approach. The octanol–water partition coefficients (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P) of the silver(I) complexes AgBet, AgDmg, and AgSar were determined using a quantitative 1H NMR-based shake-flask method described by Rücker et al.77 This approach, which directly integrates analyte signals relative to the water peak, enables the estimation of log
P) of the silver(I) complexes AgBet, AgDmg, and AgSar were determined using a quantitative 1H NMR-based shake-flask method described by Rücker et al.77 This approach, which directly integrates analyte signals relative to the water peak, enables the estimation of log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values without needing phase separation or external calibration (Fig. S42).
P values without needing phase separation or external calibration (Fig. S42).
To ensure accurate quantitation, an extended relaxation delay of 20 s (exceeding 5 × T1) was applied between scans. Maintaining many complex data points while keeping the acquisition time constant is crucial for achieving well-defined frequency resolution. A 5 s acquisition time was used to maximise spectral fidelity and ensure consistent, high-quality frequency and amplitude measurements.78
All three analysed complexes exhibited negative log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values, indicating a preference for the aqueous phase over the organic phase (Table 10). The measured values were −2.11 for AgSar, −1.23 for AgBet, and −1.18 for AgDmg, suggesting a trend of increasing lipophilicity in the order: AgSar < AgBet < AgDmg. In addition to these three, AgGlyGly and AgGlyGlyGly complexes were initially included in the study. However, it was not possible to determine their log
P values, indicating a preference for the aqueous phase over the organic phase (Table 10). The measured values were −2.11 for AgSar, −1.23 for AgBet, and −1.18 for AgDmg, suggesting a trend of increasing lipophilicity in the order: AgSar < AgBet < AgDmg. In addition to these three, AgGlyGly and AgGlyGlyGly complexes were initially included in the study. However, it was not possible to determine their log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values using this method because their characteristic 1H NMR signals overlapped with the broad base of the water signal. This spectral interference prevented accurate integration and thus reliable quantification.
P values using this method because their characteristic 1H NMR signals overlapped with the broad base of the water signal. This spectral interference prevented accurate integration and thus reliable quantification.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values
P values
| Complex | RIinitw | RIequilw | Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P P |
|---|---|---|---|
| AgSar | 0.0005168 | 0.000513 | −2.11 |
| AgDmg | 0.0006622 | 0.000621 | −1.18 |
| AgBet | 0.0007417 | 0.0007 | −1.23 |
No comparable literature values were found for these specific complexes, therefore comparison was made with previously measured log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values for other silver(I) complexes: AgGly −2.74,15 AgAla −2.36,15 AgPro −1.46,16 AgHyp −1.16,16 and AgPhe −0.96.15 These values confirm the predominantly hydrophilic nature of the silver(I) complexes, as reflected by their negative partition coefficients. Among the newly analysed complexes, AgSar exhibited the lowest lipophilicity, likely due to its specific ligand characteristics. Compared with the previously studied set, the log
P values for other silver(I) complexes: AgGly −2.74,15 AgAla −2.36,15 AgPro −1.46,16 AgHyp −1.16,16 and AgPhe −0.96.15 These values confirm the predominantly hydrophilic nature of the silver(I) complexes, as reflected by their negative partition coefficients. Among the newly analysed complexes, AgSar exhibited the lowest lipophilicity, likely due to its specific ligand characteristics. Compared with the previously studied set, the log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values of AgSar, AgBet, and AgDmg fall within an intermediate range, suggesting moderate lipophilicity relative to strongly hydrophilic complexes such as AgGly and AgAla. On the other hand, AgPhe remains the most lipophilic complex observed to date in our studies. These results demonstrate that the lipophilicity of silver(I) complexes can be modulated through appropriate ligand selection, which may have implications for their solubility, membrane permeability, and potential biological activity.
P values of AgSar, AgBet, and AgDmg fall within an intermediate range, suggesting moderate lipophilicity relative to strongly hydrophilic complexes such as AgGly and AgAla. On the other hand, AgPhe remains the most lipophilic complex observed to date in our studies. These results demonstrate that the lipophilicity of silver(I) complexes can be modulated through appropriate ligand selection, which may have implications for their solubility, membrane permeability, and potential biological activity.
A clear relationship can be observed between the log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values and the structural characteristics of the coordinating amino acid ligands. Complexes containing small, non-polar ligands such as glycine (AgGly, −2.74) and alanine (AgAla, −2.36) exhibit the lowest log
P values and the structural characteristics of the coordinating amino acid ligands. Complexes containing small, non-polar ligands such as glycine (AgGly, −2.74) and alanine (AgAla, −2.36) exhibit the lowest log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values, reflecting their high hydrophilicity. Their small side chains (hydrogen for glycine and methyl for alanine) offer minimal contribution to lipophilicity.
P values, reflecting their high hydrophilicity. Their small side chains (hydrogen for glycine and methyl for alanine) offer minimal contribution to lipophilicity.
In contrast, complexes with cyclic or bulkier ligands such as proline (AgPro, −1.46) and hydroxyproline (AgHyp, −1.16) exhibit higher log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values. The rigid pyrrolidine ring in proline reduces solvation, while the hydroxyl group in hydroxyproline increases polarity and slightly offsets this effect.
P values. The rigid pyrrolidine ring in proline reduces solvation, while the hydroxyl group in hydroxyproline increases polarity and slightly offsets this effect.
Phenylalanine (AgPhe, −0.96), containing an aromatic ring, yields the highest log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P value due to its hydrophobic π-conjugated system, which favours partitioning into the organic phase.
P value due to its hydrophobic π-conjugated system, which favours partitioning into the organic phase.
The newly studied complexes align with this trend. AgSar (−2.11) is the most hydrophilic. AgBet (−1.23) contains a quaternary ammonium group and is moderately lipophilic. AgDmg (−1.18), a dimethylated derivative, exhibits enhanced lipophilicity due to the presence of two additional methyl groups.
Based on the DFT calculated free energy values for the same structure types of the discussed silver(I) complexes [AgL] in water and in pure octan-1-ol (4), the corresponding equilibrium constants (partition coefficients) were estimated. The resulting values are listed in Table 11 and are valid only under the assumption that the structure type (OAgO or OAgN) remains the same in both solvents. The trend in values suggests generally larger hydrophobicity for the variants OAgO proposed to exist in the solution than the other hypothetical OAgN types, which should be more thermodynamically stable. Although the experimental log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values exhibit larger inter-individual differences, at least the lipophilicity range for the neutral complexes (AgGly < AgSar < AgDmg) is predicted correctly. The complex with betaine is a cation (AgBet+) unlike the previously mentioned complexes AgL (L = Gly, Sar, Dmg). The only coordination mode considered is OAgO, as the nitrogen atom in the ammonium moiety has no electron donating capabilities any more. Because of the ionic character, the calculated partition coefficient for AgBet+ is expected to be shifted towards hydrophilic values. However, the experimental log
P values exhibit larger inter-individual differences, at least the lipophilicity range for the neutral complexes (AgGly < AgSar < AgDmg) is predicted correctly. The complex with betaine is a cation (AgBet+) unlike the previously mentioned complexes AgL (L = Gly, Sar, Dmg). The only coordination mode considered is OAgO, as the nitrogen atom in the ammonium moiety has no electron donating capabilities any more. Because of the ionic character, the calculated partition coefficient for AgBet+ is expected to be shifted towards hydrophilic values. However, the experimental log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P is significantly less negative, indicating much more lipophilic behaviour. With regard to the assumption that the charge balance of all the species transferred across the liquid–liquid interface must be zero (in order to keep both immiscible phases electroneutral), the next attempt at modelling the betaine complex included also the counter-ion (nitrate) supposed to be available under the experimental conditions to form a ternary complex AgBetNO3 (or an ionic pair with the AgBet+ cation). Its calculated log
P is significantly less negative, indicating much more lipophilic behaviour. With regard to the assumption that the charge balance of all the species transferred across the liquid–liquid interface must be zero (in order to keep both immiscible phases electroneutral), the next attempt at modelling the betaine complex included also the counter-ion (nitrate) supposed to be available under the experimental conditions to form a ternary complex AgBetNO3 (or an ionic pair with the AgBet+ cation). Its calculated log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P value is according to expectations less hydrophilic than that of the bare AgBet+ cation, yet not in good agreement with the experimental one. This discrepancy between the theory and experiment suggests that there is a different structure modelled (geometry, composition) and estimated, at least in one of the solvents:
P value is according to expectations less hydrophilic than that of the bare AgBet+ cation, yet not in good agreement with the experimental one. This discrepancy between the theory and experiment suggests that there is a different structure modelled (geometry, composition) and estimated, at least in one of the solvents:
| [AgL](water) = [AgL](octan-1-ol) | (4) |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P
P
| Complex | Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P P |
||
|---|---|---|---|
| OAgN | OAgO | Exp. | |
| [AgGly] | −2.9 | −2.1 | −2.74 |
| [AgGlyGly] | −2.5 | −2.6 | — |
| [AgGlyGlyGly] | −2.6 | −2.6 | — |
| [AgSar] | −2.8 | −2.0 | −2.11 |
| [AgDmg] | −2.7 | −1.9 | −1.18 |
| [AgBet]+ | — | −4.4 | −1.23 |
| [AgBetNO3] | — | −2.7 | — |
Although log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P is formally defined for the neutral species and is considered pH-independent, many amino acid-based ligands contain ionizable groups. In aqueous environments, the protonation state of these groups can vary with pH, potentially altering the speciation of the complex. Therefore, the apparent lipophilicity under physiological or experimental conditions may be better described by the distribution coefficient (log
P is formally defined for the neutral species and is considered pH-independent, many amino acid-based ligands contain ionizable groups. In aqueous environments, the protonation state of these groups can vary with pH, potentially altering the speciation of the complex. Therefore, the apparent lipophilicity under physiological or experimental conditions may be better described by the distribution coefficient (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D), which accounts for both ionised and unionised forms at a given pH. As such, the measured log
D), which accounts for both ionised and unionised forms at a given pH. As such, the measured log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values should be interpreted with this limitation in mind, particularly when comparing structurally diverse complexes with different acid–base properties.78,79 A summary of the Lipinski's parameter data (Table 12) shows that only in the case of AgGly and AgBet complexes are the rules closely related to the composition of the complexes violated. Moreover, log
P values should be interpreted with this limitation in mind, particularly when comparing structurally diverse complexes with different acid–base properties.78,79 A summary of the Lipinski's parameter data (Table 12) shows that only in the case of AgGly and AgBet complexes are the rules closely related to the composition of the complexes violated. Moreover, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values confirm that the lipophilicity of silver(I) complexes is highly dependent on ligand structure, particularly on side-chain polarity, molecular size, and the presence of hydrophobic or charged functional groups. Structural modifications of the amino acid ligands provide a viable strategy for tuning the physicochemical properties of silver(I) complexes, which may be relevant to their solubility, transport behaviour, and potential biological activity. In addition, biological activity values (pMIC and pIC50, Table 11) correlated with log
P values confirm that the lipophilicity of silver(I) complexes is highly dependent on ligand structure, particularly on side-chain polarity, molecular size, and the presence of hydrophobic or charged functional groups. Structural modifications of the amino acid ligands provide a viable strategy for tuning the physicochemical properties of silver(I) complexes, which may be relevant to their solubility, transport behaviour, and potential biological activity. In addition, biological activity values (pMIC and pIC50, Table 11) correlated with log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P indicate a more pronounced activity of more hydrophilic complexes, as well as a stronger interaction with HAS.
P indicate a more pronounced activity of more hydrophilic complexes, as well as a stronger interaction with HAS.
| Complex | MW (g mol−1) | Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P (meas.) P (meas.) |
Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P (calc.) P (calc.) |
HA | HD | n | pMIC (S. aureus) | pIC50 (MBA-MB-231) | Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kb Kb |
|---|---|---|---|---|---|---|---|---|---|
a Ref. 21.
b Ref. 17, molecular weight, MW < 500; number of hydrogen bond acceptors, HA < 10; number of hydrogen bond donors, HD < 5; n-octanol–water partition coefficient, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P < 5, pMIC = −log P < 5, pMIC = −log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MIC, pIC50 = −log MIC, pIC50 = −log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) IC50, n = number of rule violations. IC50, n = number of rule violations.
|
|||||||||
| {[Ag2(HGly)2]}n(NO3)2na | 489.90 | −2.74 | −2.1 | 14 | 6 | 2 | 5.3 | — | — |
| {[Ag(HGlyGly)]}n(NO3)nb | 301.99 | — | −2.6 | 9 | 3 | 0 | 4.7 | 4.9 | — |
| [Ag(HGlyGlyGly)(NO3)] | 359.04 | — | −2.6 | — | — | — | 5.6 | 4.7 | 4.51 |
| {[Ag(HSar)(NO3)]}n | 258.98 | −2.11 | −2.0 | 7 | 3 | 0 | 5.3 | 5.11 | 4.41 |
| {[Ag(HDmg)(NO3)]}n | 273.00 | −1.18 | −1.9 | 7 | 3 | 0 | 5.3 | 5.07 | 4.77 |
| {[Ag3(HBet)2(NO3)3]}n | 743.93 | −1.23 | −4.4(−2.7) | 18 | 2 | 2 | 5.6 | 4.93 | 4.89 |
Potential action mechanism
| % of enzyme activity | ||||||
|---|---|---|---|---|---|---|
| c (µM) | AgDmg | AgSar | AgBet | AgGlyGlyGly | AgGlyGly | AgGly |
| 10 | 54.3 | 51.2 | 42.1 | 31.9 | 56.5 | 42.6 |
| 5 | 85.9 | 70.0 | 52.1 | 73.7 | 53.9 | 59.7 |
| 2.5 | 100 | 100 | 100 | 98.5 | 94.5 | 73.0 |
| 1 | 100 | 100 | 100 | 100 | 100 | 100 |
The combined results of this study indicate that the antibacterial activity of the tested silver(I) complexes is mediated by multiple converging mechanisms, including oxidative stress and direct interference with essential enzymatic functions. All complexes induced a measurable increase in intracellular ROS levels, with the glycine-derived complexes showing the strongest effect. Elevated ROS is known to damage cellular macromolecules, destabilize membranes and disrupt protein function, and therefore represents one of the early and important components of silver-induced toxicity.
In parallel with ROS production, the complexes also acutely inhibited two unrelated bacterial enzymes—β-galactosidase and Taq DNA polymerase. Inhibition of β-galactosidase was observed already within hours of exposure, under conditions where transcriptional downregulation could not occur, indicating a direct or microenvironment-mediated loss of enzymatic activity. All complexes caused a concentration-dependent decrease in ONPG-hydrolysing activity, consistent with rapid inactivation of pre-existing membrane-associated proteins.
A similar pattern was observed in the PCR inhibition assay. The complete or partial suppression of 16S rDNA amplification in the presence of silver(I) complexes—most evident at 10–25 μmol L−1—strongly suggests that Ag complexes interfere with core catalytic residues of DNA polymerases. The high sensitivity of polymerases to silver(I) ions agrees with the known reactivity of Ag+ toward thiol groups, histidine and other amino acid residues essential for polymerase structure and processivity.
Together, these findings support a model in which silver(I) complexes exert their antimicrobial activity through a multitarget mode of action. ROS overproduction creates a pro-oxidative intracellular environment, while direct binding of Ag+ or Ag-ligand species to enzymes leads to functional inactivation of key bacterial proteins. The simultaneous impairment of membrane-associated enzymes and DNA replication machinery may explain the strong bactericidal potency of the complexes and reduce the likelihood of resistance development.
In addition to enzyme inhibition and ROS-mediated stress, our data show that the antimicrobial effect of the silver(I) complexes is equally effective against both the antibiotic-sensitive S. aureus strain and the methicillin-resistant MRSA strain. The comparable susceptibility of MRSA demonstrates that the activity of the complexes is independent of classical resistance mechanisms, such as altered penicillin-binding proteins or β-lactam target modification. This supports the notion that silver(I) complexes exert a multitarget mechanism of action involving oxidative stress, protein inactivation, and broader cellular damage, which MRSA cannot readily evade. Such activity against resistant strains highlights the therapeutic potential of silver(I) complexes in overcoming antibiotic resistance.
Analysis of cell-cycle distribution (Table 14 and Fig. S44) revealed that AgSar and AgDmg induced accumulation of cells in the G1 phase after 6 h of treatment, indicating early cell-cycle arrest. Similar ligand-dependent G1 arrest has been reported for silver(I) complexes bearing biologically relevant ligands.81 In contrast, AgBet and AgGlyGlyGly reduced the G1 population at this early time point, consistent with rapid cytotoxicity and initiation of apoptotic processes. After 24 h of exposure, only AgSar maintained partial G1 arrest, whereas AgDmg, AgBet, and AgGlyGlyGly induced a pronounced increase in the sub-G0/G1 population, indicative of DNA fragmentation and apoptotic cell death, as commonly observed for apoptosis-inducing Ag(I) complexes.82
| subG0/G1 | G1 | S | G2/M | |
|---|---|---|---|---|
| Results are expressed as mean ± standard deviation of independent experiments (*p < 0.05 compared to untreated control, based on ordinary one-way ANOVA with Dunnett's post hoc test). | ||||
| ctrl_6h | 1.4 ± 0.6 | 42.9 ± 3.6 | 29.3 ± 1.4 | 26.5 ± 3.2 |
| AgSar | 1.7 ± 0.4 | 51.6 ± 0.9* | 29.1 ± 1.7 | 17.7 ± 0.3* |
| AgDmg | 2.8 ± 1.4 | 49.5 ± 1.0* | 29.5 ± 0.2 | 18.2 ± 0.2* |
| AgBet | 40.3 ± 5.6* | 23.8 ± 5.5* | 25.0 ± 2.9 | 10.9 ± 1.4* |
| AgGlyGlyGly | 30.9 ± 4.8* | 30.3 ± 1.0* | 25.5 ± 2.0 | 13.4 ± 3.0* |
| ctrl_24h | 1.1 ± 0.4 | 47.6 ± 1.5 | 29.7 ± 1.6 | 21.7 ± 2.7 |
| AgSar | 4.0 ± 2.3 | 52.2 ± 1.9 | 27.5 ± 2.3 | 16.3 ± 1.6* |
| AgDmg | 28.2 ± 7.6* | 36.9 ± 1.9* | 20.1 ± 4.9* | 14.9 ± 0.8* |
| AgBet | 51.6 ± 8.0* | 23.2 ± 2.8* | 15.9 ± 3.1* | 9.4 ± 0.7* |
| AgGlyGlyGly | 41.1 ± 9.1* | 28.2 ± 6.3* | 20.0 ± 3.6* | 10.6 ± 1.6* |
Phosphatidylserine externalization, a hallmark of apoptosis, was assessed by Annexin V/propidium iodide double staining (Fig. S45). As shown in Table 15, AgBet—followed by AgGlyGlyGly—was the most potent apoptosis inducer, rapidly driving cells into late apoptosis within 6 h and maintaining high apoptotic levels at 24 h. Comparable rapid induction of late apoptosis by potent silver(I) complexes has been previously reported.83 AgDmg exhibited a marked time-dependent increase in both early and late apoptotic populations, while AgSar triggered apoptosis more slowly and to a lesser extent, reflecting ligand-dependent differences in apoptotic potency that are well documented for Ag(I) coordination compounds.30,80 Notably, at 24 h, AgBet and AgGlyGlyGly produced the highest proportion of early apoptotic cells, suggesting strong activation of initial apoptotic signaling pathways, potentially involving mitochondrial dysfunction.83
| Live | Early Apo | Late Apo | Necrotic | |
|---|---|---|---|---|
| Results are expressed as mean ± standard deviation of independent experiments (*p < 0.05 compared to untreated control, based on ordinary one-way ANOVA with Dunnett's post hoc test). | ||||
| ctrl_6h | 90.2 ± 2.2 | 4.9 ± 0.1 | 3.8 ± 1.8 | 1.1 ± 0.5 |
| AgSar | 75.5 ± 3.9* | 15.5 ± 3.1* | 7.2 ± 2.9 | 1.8 ± 0.4 |
| AgDmg | 65.1 ± 9.2* | 20.4 ± 4.5* | 12.4 ± 4.2* | 2.2 ± 0.9 |
| AgBet | 2.2 ± 0.4* | 22.6 ± 2.8* | 74.7 ± 3.2* | 0.4 ± 0.1 |
| AgGlyGlyGly | 47.3 ± 5.6* | 15.3 ± 1.9* | 35.7 ± 2.9* | 1.8 ± 0.4 |
| ctrl_24h | 89.6 ± 3.4 | 4.6 ± 0.4 | 4.3 ± 2.3 | 1.5 ± 0.8 |
| AgSar | 51.7 ± 3.4* | 25.9 ± 5.5* | 20.1 ± 6.6* | 2.3 ± 0.4 |
| AgDmg | 26.6 ± 2.4* | 37.6 ± 5.4* | 33.8 ± 0.3* | 2.0 ± 0.2 |
| AgBet | 2.1 ± 0.1* | 54.8 ± 6.4* | 42.7 ± 6.4* | 0.3 ± 0.2 |
| AgGlyGlyGly | 13.9 ± 4.7* | 53.5 ± 5.6* | 31.0 ± 5.1* | 1.5 ± 0.2 |
Based on these findings, mitochondrial membrane potential (MMP) alterations were further examined. As shown in Fig. 15, AgBet induced the most pronounced decrease in MMP, in agreement with its strong apoptotic activity observed in cell-cycle and Annexin V analyses. Loss of MMP is a well-established mechanism underlying the intrinsic apoptotic pathway activated by silver(I) complexes.83 Overall, all tested Ag(I) complexes caused time-dependent disruption of mitochondrial membrane potential, supporting the involvement of mitochondria-mediated apoptosis in their cytotoxic effects.
One of the established mechanisms of cytotoxic drug action involves interaction with DNA. To assess whether this mechanism may contribute to the biological activity of the investigated silver(I) complexes, their interaction with ctDNA was studied by direct spectrofluorimetric titration. The fluorescence emission spectra obtained during the titration experiments are shown in Fig. S46 and the corresponding binding constants (Kb), calculated from these data, are summarized in Table 16.
In the literature, in addition to the commonly used competitive fluorimetric titrations employing ethidium bromide (EB) or Hoechst dyes, DNA-binding studies of metal complexes are also often based on their intrinsic fluorescence properties.84–86 Similarly, in the present study, the silver(I) complexes exhibited weak but sufficient intrinsic fluorescence under the applied experimental conditions, which can be attributed to Ag(I)–Ag(I)-ligand attractive interactions.87 This enabled their interaction with ctDNA to be investigated by spectrofluorimetric titration. The corresponding fluorescence emission spectra are provided in the SI (Fig. S46).
The binding constants (Kb = 2.1 × 104–3.57 × 105 M−1) determined for the silver(I) complexes are comparable to those reported for zinc–naproxen complex–ctDNA interactions or silver-diazafluorene complex–ctDNA determined by non-competitive fluorescence titrations (Kb = 0.2 × 105 or 2.4 × 105 M−1).84,86 This similarity supports the reliability of the spectrofluorimetric approach and highlights the relevance of metal complexes as DNA-binding agents. To assess the binding mode of the investigated silver(I) complexes, comparisons were made with classical DNA-binding molecules. Intercalators such as ethidium bromide exhibit high binding affinities (Kb ≈ 105–106 M−1), whereas minor groove binders, including Hoechst 33258, show lower binding constants (Kb ≈ 104–105 M−1) in non-competitive assays.88–90 Therefore, the obtained results place the silver(I) complexes within the binding affinity range characteristic of minor groove binders rather than strong intercalators. Collectively, these findings suggest that the studied silver(I) complexes most likely interact with DNA via a minor groove binding mode.
Experimental
Materials for synthesis and chemical studies
All chemicals were analytically pure and used without purification. Glycine, N-methylglycine (Sarcosine), N,N-dimethylglycine, N,N,N-trimethylglycine (betaine), glycylglycine, glycylglycylglycine and dimethyl sulfoxide (DMSO) were received from Sigma-Aldrich Chemicals and silver(I) nitrate from Lachema. Moreover, for potentiometric measurements, nitric acid, potassium nitrate, potassium hydroxide, sodium chloride, sodium carbonate, and potassium hydrogen phthalate were purchased from Mikrochem. For voltammetry measurements the stock electrolyte KNO3 was received from Lachema.Materials for biological studies
(Firmicutes) Staphylococcus aureus (S. aureus) CCM 3953 (Czech Collection of Microorganisms), G− bacteria (γ-proteobacteria) Escherichia coli (E. coli) CCM 3988, (Czech Collection of Microorganisms), Rhizopus oryzae (R. oryzae) CCM F-8284, Alternaria alternata (A. alternata) CCM F-128 and Microsporum gypseum (M. gypseum) CCM F-8342 (Czech Collection of Microorganisms) were obtained from Masaryk University, Brno and the yeast Candida parapsilosis (C. parapsilosis) ATCC 22019 (American type culture collection) from ATCC, Manassas, VA, USA. Mueller Hinton Broth (MHB, SigmaAldrich) and Sabouraud growth medium, (SB, SigmaAldrich) were obtained from Sigma-Aldrich Chemicals, Cell lines HCT116 (human colorectal carcinoma), MDA-MB-231 (human mammary gland adenocarcinoma), A549 (human alveolar adenocarcinoma), A2058 (human metastatic melanoma), PaTu 8902 (human pancreatic adenocarcinoma), HepG2 (human hepatocellular carcinoma), Jurkat (human leukaemic T-cell lymphoma), and human dermal fibroblasts (BJ-5ta) were obtained from ATCC, Manassas, VA, USA, growth medium RPMI 1640, and Dulbecco's Modified Eagle Medium (DMEM) + sodium pyruvate from Biosera, Kansas City, MO, USA, 10% fetal bovine serum (FBS) from Invitrogen, Carlsbad, CA, USA, and Antibiotic/Antimycotic Solution and Hygromycin B from Merck, Darmstadt, Germany, (Merck). MTS (3-(4,5-dimethylthiazol-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H tetrazolium) assay was received from Promega, Madison, WI, USA. Fertilized chicken embryos came from a certified hatchery, Párovské Háje (Nitra, Slovakia) and human serum albumin (HSA) from Sigma-Aldrich Chemicals.Instruments used
Potentiometric measurements were obtained on a TitroLine 7000 dosage system equipped with a half-cell Ross Type glass electrode (model 8101 from Thermo Fisher Scientific, Waltham, MA, USA) and a double-junction reference electrode (type 900, 200, from Thermo Fisher Scientific, Waltham, MA, USA). 1H NMR spectra were recorded on a Varian VNMRS 600 MHz spectrometer (operating at 599.87 MHz for 1H) using MestReNova 15.1.0 (2024) software. Single-crystal diffraction data were collected on a Bruker D8 VENTURE Kappa Duo PHOTON 100 diffractometer. Infrared spectra were recorded on an Avatar FT-IR 6700 (Fourier transform infrared spectroscopy) spectrometer from 4000 to 400 cm−1 using the ATR (attenuated total reflectance) technique. Elemental analysis for carbon, hydrogen and nitrogen was performed with a CHNOS Elemental Analyzer Vario MICRO from Elementar Analysensysteme GmbH. Thermogravimetric analysis (TG) was studied by using a Setaram Setsys Evolution analyser-1750. The voltametric measurements were performed using an Autolab PGSTAT101 potentiostat (Metrohm, Switzerland), operated via Nova 1.11 software. UV-VIS spectra of tested complexes in growth media were performed with a Jasco V-730 spectrophotometer. The fluorescence experiments were carried out on a Varian Cary Eclipse fluorimeter. Irritation potential by HET-CAM test was carried out by using an Olympus SZ61 stereomicroscope (Tokyo, Japan) equipped with a PROMICRA 3.2 digital camera (Prague, Czech Republic). Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P determinations were determined on a Varian VNMRS 600 MHz spectrometer (operating at 599.87 MHz for 1H).
P determinations were determined on a Varian VNMRS 600 MHz spectrometer (operating at 599.87 MHz for 1H).
Complexes formation in aqueous solution
Potentiometric measurements were carried out in a glass vessel (25 mL) thermostated at 25 ± 0.1 °C using a TitroLine 7000 dosage system equipped with a half-cell Ross Type glass electrode (model 8101 from Thermo Fisher Scientific, Waltham, MA, USA) and a double-junction reference electrode (type 900, 200, from Thermo Fisher Scientific, Waltham, MA, USA). Its liquid junction (the external filling solution) consisted of KNO3(aq) at I = 0.1 M. An inert atmosphere was ensured by a constant flow of a nitrogen gas. Precise calibration of the electrode was carried out by the titration of 0.1 M HNO3 with 0.15 M KOH in the range −log[H+] = 2.0–12.0, with the pH-meter yielding E values. The relation between E and −log[H+] is expressed by (eqn 5) where the term E° contains the standard potentials of the electrode and the contribution of inert ions to the liquid-junction potential. The value S corresponds to the Nernstian slope, the value of which should be close to the theoretical value, and j1[H+] and j2[OH−] = j2Kw/[H+] are contributions of the H+ and OH− ions, respectively, to the liquid-junction potential. The pKw value is equal to 13.68 in eqn (5).
| E = E° − S(−log[H+] + j1[H+] + j2Kw/[H+] | (5) |
The parameters j1 and j2 cause deviation from linear dependence between E and −log[H+] only in strongly acidic and strongly alkaline solutions.
The titrand solutions consisted of appropriate ligand (Sar/Dmg/GlyGly/GlyGlyGly) (cL = 0.003–0.006 M), HNO3 (cH = 0.0061 M), the supporting electrolyte (KNO3 previously dried in an oven at t = 120 °C for at least 2 h) to reach the desired ionic strength value of I = 0.1 M, and silver(I) nitrate (cM = 0–0.005 M; cM = 0 during measurements for the determination of ligand's acid–base properties).
The protonation constants β110, β210 are the concentration constants and are defined by β110 = [HL1,2,3,4]/[H][L1,2,3,4], β210 = [H2L1,2,3,4]/[H]2[L1,2,3,4] (pK2 = log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β110, pK1 = log
β110, pK1 = log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β210 − log
β210 − log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β110), L1 = Sar, L2 = Dmg, L3 = GlyGly, L4 = GlyGlyGly; the stability constants are defined by βrqp = [HrLqMp]/[H]r[L]q[M]p, M = Ag(I). The equilibrium constants were obtained by fitting the titration data with OPIUM.27 The following hydrolysis constants were used for the calculations: βpr = [Mp(OH)r3−r][H]r/[M]p; log
β110), L1 = Sar, L2 = Dmg, L3 = GlyGly, L4 = GlyGlyGly; the stability constants are defined by βrqp = [HrLqMp]/[H]r[L]q[M]p, M = Ag(I). The equilibrium constants were obtained by fitting the titration data with OPIUM.27 The following hydrolysis constants were used for the calculations: βpr = [Mp(OH)r3−r][H]r/[M]p; log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β11 = −11.75; log
β11 = −11.75; log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β12 = − 24.34.91
β12 = − 24.34.91
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ligand systems. Solution pH values were adjusted by HNO3 solution (0.1 M) and KOH solution (0.1 M).
ligand systems. Solution pH values were adjusted by HNO3 solution (0.1 M) and KOH solution (0.1 M).
Syntheses of complexes AgSar, AgDmg, AgBet and AgGlyGlyGly. {[Ag(HSar)(NO 3 )]} n (AgSar): N-Methylglycine (Sarcosine, HSar = means protonation on the nitrogen atom of the amino group) (52.4 mg, 0.589 mmol) was dissolved in EtOH
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O solvent mixture in 1
H2O solvent mixture in 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 molar ratio (5 mL) and was added into the silver(I) nitrate solution (100 mg; 0.589 mmol) prepared from the same solvent mixture (5 mL). The reaction mixture was stirred for 10 min and then left for free crystallization in the absence of light. Crystals formed by slow evaporation (2 weeks) were filtered off, dried on filter paper in a Petri dish and used for further characterization. Light-gray crystals; elemental analysis found C: 13.91; H: 2.73; N: 10.82; calculated for C3H7AgN2O5: C: 13.58; H: 2.68; N: 10.57. Yield (based on AgNO3): 64.5%. IR (cm−1): 3029 (br), 2903 (br), 1568 (s), 1472 (m), 1456 (m), 1414 (m), 1392 (m), 1315 (br), 1063 (m), 865 (s), 680 (m), 452 (s). 1H NMR (ppm) in 1% DMSO-d6: 3.62 (s, H3 [CH2–]), 2.74 (s, H1 [CH3–]) of Sar.
2 molar ratio (5 mL) and was added into the silver(I) nitrate solution (100 mg; 0.589 mmol) prepared from the same solvent mixture (5 mL). The reaction mixture was stirred for 10 min and then left for free crystallization in the absence of light. Crystals formed by slow evaporation (2 weeks) were filtered off, dried on filter paper in a Petri dish and used for further characterization. Light-gray crystals; elemental analysis found C: 13.91; H: 2.73; N: 10.82; calculated for C3H7AgN2O5: C: 13.58; H: 2.68; N: 10.57. Yield (based on AgNO3): 64.5%. IR (cm−1): 3029 (br), 2903 (br), 1568 (s), 1472 (m), 1456 (m), 1414 (m), 1392 (m), 1315 (br), 1063 (m), 865 (s), 680 (m), 452 (s). 1H NMR (ppm) in 1% DMSO-d6: 3.62 (s, H3 [CH2–]), 2.74 (s, H1 [CH3–]) of Sar.
{[Ag(HDmg)(NO 3 )]} n (AgDmg): Complex AgDmg was prepared by a similar process as AgSar (amount of N,N-dimethylglycine, HDmg = means protonation on the nitrogen atom of the amino group): (60.8 mg, 0.589 mmol). Crystals were filtered off after 2 weeks and dried on filter paper. Light-gray crystals; elemental analysis found C: 17.59; H: 3.32; N: 10.26; calculated for C4H9AgN2O5: C: 17.85; H: 3.80; N: 10.29. Yield (based on AgNO3): 60.2%. IR (cm−1): 3044 (br), 2948 (br), 1585 (s), 1479 (m), 1417 (m), 1398 (m), 1317 (br), 1174 (m), 1033 (m), 861 (m), 460 (m). 1H NMR (ppm) in 1% DMSO-d6: 3.73 (s, H3 [CH2–]), 2.93 (s, H1, 7 [CH3–]) of Dmg.
{[Ag3(HBet)2(NO3)3]}n (AgBet): Silver(I) nitrate (200 mg, 1.177 mmol) was dissolved in 5 mL deionized water and was added dropwise to the 5 mL aqueous solution of N,N,N-trimethylglycine (betaine) (68.96 mg, 0.589 mmol). The reaction mixture was stirred for 15 min and then left for crystallization in the absence of light. Crystals were filtered off after three weeks and dried in a stream of air. Colourless crystals; elemental analysis found C: 15.90; H: 2.34; N: 8.93; calculated for C10H22Ag3N5O13: C: 16.14; H: 2.98; N: 9.41. Yield (based on AgNO3): 65.9%. IR (cm−1): 3034 (w), 2984 (w), 1581 (s), 1485 (m), 1446 (m), 1381 (m), 1310 (br), 1135 (m), 1011 (m), 890 (s), 707 (m), 442 (w). 1H NMR (ppm) in 1% DMSO-d6: 3.91 (s, H3 [CH2–]), 3.27 (s, H1,7,8 [CH3–]) of Bet.
[Ag(HGlyGlyGly)(NO 3 )] (AgGlyGlyGly): Glycylglycylglycine (111.36 mg, 0.589 mmol) dissolved in 5 mL of deionized water was added dropwise to the 5 mL silver(I) nitrate solution (100 mg, 0.589 mmol). The reaction mixture was stirred for 10 min and then allowed for crystallization in the absence of light. The final compound in non-crystallic form was isolated from the beaker after 2 weeks and dried on filter paper in a Petri dish. Light-brown powder; elemental analysis found C: 20.07; H: 3.09; N: 15.60; calculated for C6H11AgN4O7: C: 20.43; H: 3.19; N: 15.73. Yield (based on AgNO3): 42.5%. IR (cm−1): 3271 (br), 3086 (br), 2967 (br), 1634 (m), 1538 (s), 1456 (m), 1416 (m), 1293 (br), 1121 (m), 1040 (m), 814 (w), 675 (s), 467 (w). 1H NMR (ppm) in 1% DMSO-d6: 4.05 (s, H2 [CH2–]), 3.91 (s, H5 [CH2–]), 3.80 (s, H8 [CH2–]) of GlyGlyGly.
X-ray structure determination. Single crystals of AgSar (0.530 × 0.213 × 0.084), AgDmg (0.171 × 0.162 × 0.058) and AgBet (0.270 × 0.178 × 0.116) were collected on a Bruker D8 VENTURE Kappa Duo PHOTON 100 diffractometer using Mo Kα radiation for (λ = 0.71073 Å). The structures were solved by intrinsic phasing using SHELXT105 and refined on F2 using SHELXL.106 All non-hydrogen atoms were refined anisotropically and aromatic hydrogens were included in the calculated positions. The intermolecular interactions were analysed using PLATON.107 Detailed crystal data for complexes are listed in Table S1. The structure figures were drawn using the DIAMOND software.108
AgSar: C3H7AgN2O5; MW = 258.98 g mol−1; monoclinic lattice; space group P21/n; a = 5.9666(3) Å, b = 8.4114(4) Å, c = 13.8333(6) Å; β = 101.712(2)°; V = 679.80(6) Å3; Z = 4; T = 120(2) K; radiation Mo Kα; ρ(calcd) = 2.530 g cm−3; μ = 2.942 mm−1; R1 = 0.0150, wR2 = 0.0348; reflections collected/unique 12879/1685; goodness-of fit on F2 1.115; (Δρ)max/(Δρ)min = 0.473/−0.492 e Å−3; R1/wR2 (for all data), 0.0164/0.0353.
AgDmg: C4H9AgN2O5; MW = 273.00 g mol−1; monoclinic lattice; space group P21/c; a = 5.5261(4) Å, b = 11.2028(8) Å, c = 14.0565(12) Å; β = 97.992(3)°; V = 862.17(11) Å3; Z = 2; T = 120(2) K; radiation Mo Kα; ρ(calcd) = 2.319 g cm−3; μ = 2.346 mm−1; R1 = 0.0351, wR2 = 0.1062; reflections collected/unique 31594/3526; goodness-of fit on F2 1.108; (Δρ)max/(Δρ)min = 2.396/−1.008 e Å−3; R1/wR2 (for all data), 0.0358/0.1071.
AgBet: C10H22Ag3N5O13; MW = 743.93 g mol−1; orthorhombic lattice; space group P21212; a = 15.7832(8) Å, b = 23.2072(12) Å, c = 5.6320(2) Å; V = 2062.91(17) Å3; Z = 4; T = 120(2) K; radiation Mo Kα; ρ(calcd) = 2.395 g cm−3; μ = 2.895 mm−1; R1 = 0.0154, wR2 = 0.0374; reflections collected/unique 47613/5135; goodness-of fit on F2 1.068; (Δρ)max/(Δρ)min = 0.638/−0.513 e Å−3; R1/wR2 (for all data), 0.0157/0.0375.
Crystallographic data (excluding structure factors) for the structures reported in this paper have been deposited with the Cambridge Crystallographic Data Centre as supporting publication no. 2473205–2473207.
Thermal analysis. The thermal behaviour of compounds AgSar, AgDmg, AgBet and AgGlyGlyGly was studied by thermogravimetry (TG) using a Setaram Setsys Evolution analyser-1750 under an atmosphere of air. The samples were heated with a heating rate of 10 °C min−1 in the temperature range from 20 to 600 °C and with an air flow rate of 60 mL min−1. Before the thermal measurements, gentle grinding of the samples and careful packing into the corundum crucibles were performed. The mass of samples used in the analyses was within 6–10 mg. The obtained thermoanalytical curves were analysed using the Origin computational program (version 6.1052, Origin Lab Northampton, MA, USA).
Complexes’ stability in testing and growth media
Complexes’ bioassays
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O = 1
H2O = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2. The final concentration of the solvent never exceeded 1.0 vol%, neither in control nor treated samples. The concentration of metal complexes used in the experimental work for the evaluation of antimicrobial activity was in the range of 0.5–100 µM, and for the free ligands in the range of 0.05–1.0 mM in all experiments. The antimicrobial activity of tested compounds was characterized by MIC80 resp. MIC90 values (minimal inhibitory concentration of a compound that inhibits the growth of model microorganism on corresponding value (%) indicated by the MIC index when compared to the untreated control). Since the bacterial cells were most sensitive, except for the addition of the complexes to the lag phase, complexes were added to the growing bacterial cells in the exponential growth phase at the mentioned rage of concentration above. The growth of the culture was measured, and the growth curve was visualised graphically. All the obtained results of antimicrobial activity were compared to the activity of AgNO3 and AgSD.
2. The final concentration of the solvent never exceeded 1.0 vol%, neither in control nor treated samples. The concentration of metal complexes used in the experimental work for the evaluation of antimicrobial activity was in the range of 0.5–100 µM, and for the free ligands in the range of 0.05–1.0 mM in all experiments. The antimicrobial activity of tested compounds was characterized by MIC80 resp. MIC90 values (minimal inhibitory concentration of a compound that inhibits the growth of model microorganism on corresponding value (%) indicated by the MIC index when compared to the untreated control). Since the bacterial cells were most sensitive, except for the addition of the complexes to the lag phase, complexes were added to the growing bacterial cells in the exponential growth phase at the mentioned rage of concentration above. The growth of the culture was measured, and the growth curve was visualised graphically. All the obtained results of antimicrobial activity were compared to the activity of AgNO3 and AgSD.
Cell culture. The cell lines HCT116 (human colorectal carcinoma), MDA-MB-231 (human mammary gland adenocarcinoma), A549 (human alveolar adenocarcinoma), A2058 (human metastatic melanoma), PaTu 8902 (human pancreatic adenocarcinoma), HepG2 (human hepatocellular carcinoma), Jurkat (human leukaemic T cell lymphoma) and human dermal fibroblasts (BJ-5ta) were obtained from the American type culture collection (ATCC, Manassas, VA, USA). HCT116, HepG2, MDA-MB-231 and Jurkat cells were cultured in growth medium RPMI 1640 (Biosera, Kansas City, MO, USA) supplemented with a 10% fetal bovine serum (FBS) (Invitrogen, Carlsbad, CA, USA) and Antibiotic/Antimycotic solution (Merck, Darmstadt, Germany). A549, A2058, PaTu 8902 cells were cultured in growth medium consisting of high-glucose Dulbeccós Modified Eagle Medium (DMEM) + sodium pyruvate (Biosera) supplemented with a 10% FBS, Antibiotic/Antimycotic solution and 25 mM HEPES (only PaTu 8902). BJ-5ta were maintained in growth medium consisting of high-glucose DMEM
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) M199 4
M199 4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 medium mixture and supplemented with a 10% fetal bovine serum (FBS) + hygromycin B (0.01 mg mL−1); Merck. Cells were maintained in standard conditions under an atmosphere containing 5% CO2 at 37 °C. Prior to each experiment, cell viability was greater than 95%.
1 medium mixture and supplemented with a 10% fetal bovine serum (FBS) + hygromycin B (0.01 mg mL−1); Merck. Cells were maintained in standard conditions under an atmosphere containing 5% CO2 at 37 °C. Prior to each experiment, cell viability was greater than 95%.
MTS assay. The antiproliferative activity of metal complexes AgSar, AgDmg, AgBet, AgGlyGlyGly, free ligands Sar, Dmg, Bet, GlyGlyGly, AgNO3 and cisPt was evaluated by colorimetric microculture MTS (3-(4,5-dimethylthiazol-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H tetrazolium) assay (Promega, Madison, WI, USA). Cancer and non-cancer cells were seeded at density 5 × 103 cells/well in 96-well polystyrene microplates. Twenty-four hours after cell seeding, different concentrations (1.0–100.0 µM) of the compounds were tested. After 72 hours of incubation, 10 µL of MTS was added to each well. After an additional 2 hours, cell proliferation was evaluated by measuring of absorbance at wavelength 490 nm using the automated Cytation™ 3 Cell Imaging Multi-Mode Reader (Biotek, Winooski, VT, USA). The absorbance of control wells was taken as 100%, and the results were expressed as a percent of untreated control. IC50 values were calculated from MTS analyses by GraphPad Prism version 9.0.2 (GraphPad Software, La Jolla, CA, USA).
In ovo methods. Fertilized chicken embryos (150) from a certified hatchery, Párovské Háje (Nitra, Slovakia), were used to evaluate irritation potential and effects on angiogenesis. The chicken eggs were incubated in a horizontal position in an incubator with a temperature of 37.5 °C and humidity around 60% in a River System incubator (River, Italy). After 24 hours of incubation, 3 mL of albumin was removed using a needle and syringe, and the hole was sealed with liquid paraffin. On embryonic day (ED) 8, the eggshell was opened, and a sterilized silicon ring was applied to the chorioallantoic membrane (CAM). Approval for the animal protocol was not necessary, as it is exempt under the legislation regarding the protection of animals used for scientific purposes (2010/63/EU), as well as applicable laws in the United States.
Irritation potential by HET-CAM test. The safety of application of the tested substances was evaluated using the Hen's Egg Test on the CAM (HET-CAM) according to the ICCVAM-recommended protocol (NIH Publication No. 10-7553, 2010). 30 µL of the tested substances, dissolved in phosphate buffer (30 µM), was applied to the chorioallantoic membrane (CAM) within a silicone ring. Phosphate buffer served as the control, and 1% SDS was used as the negative control. The designated zone was photographed before application and at 30, 120, and 300 seconds after application using an Olympus SZ61 stereomicroscope (Tokyo, Japan) equipped with a PROMICRA 3.2 digital camera (Prague, Czech Republic). The captured images were subsequently processed using QuickPHOTO MICRO software version 3.2 (Promicra, Prague, Czech Republic). Hemorrhage, vascular lysis, and coagulation were evaluated on each photograph by three independent observers to eliminate subjective errors. The values were assessed using the Luepke score system.43
Angiogenesis by CAM method. The tested substances, diluted as described in the previous section, were applied (30 µL) into the sterile silicone ring and photographed using the camera mentioned in the previous paragraph. The hole in the eggshells was then sealed with tape, and the eggs were returned to the incubator and incubated under the same conditions (37.5 °C, 60% humidity). After 72 hours, the zones determined by rings were photographed, and the chicken embryos were sacrificed. The photographs were evaluated using software enhanced with artificial intelligence, IKOSA® software (KOLAIDO GmbH, Altenrhein, Switzerland).
Structure–activity relationships
| F0/F = 1 + KSV[Q] = 1 + Kqτ0[Q] | (6) |
log[(F0 − F/F)] = log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kb + n Kb + n![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log[Q] log[Q] | (7) |
The displacement experiments were carried out by using HSA site probes, warfarin (binding to site IIA) and ibuprofen (binding to site IIIA). The concentration of HSA and warfarin/ibuprofen was the same, i.e. 2.0 × 10−6 M. Increasing concentrations of silver(I) complexes (0–4.9 × 10−5 M) were added to the HSA-WAR/IBU system. Every emission spectrum was recorded upon the excitation wavelength at 280 nm, at 25 °C in 10 mM PBS buffer (pH 7.4), with a slit width of 10 nm for the excitation and 10 nm for the emission beam in the range of 300–450 nm.114
Log
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/i_char_2009.gif) P determination.
The octanol–water partition coefficients Kow for three silver(I) complexes (AgBet, AgDmg AgSar) were determined experimentally using the NMR-based method described by Rücker.77 This method is a simple miniature shake-flask technique conducted in an NMR tube, where only the aqueous phase is analysed by quantitative 1H NMR spectroscopy. In this approach, the aqueous phase is measured both before and after equilibration with n-octanol. When equal volumes of aqueous and octanol phases are used, the equilibrium concentration of the analyte in the octanol phase can be determined indirectly as the difference between its initial concentration in the aqueous phase and its concentration after equilibration. The partition coefficient is then calculated as:
P determination.
The octanol–water partition coefficients Kow for three silver(I) complexes (AgBet, AgDmg AgSar) were determined experimentally using the NMR-based method described by Rücker.77 This method is a simple miniature shake-flask technique conducted in an NMR tube, where only the aqueous phase is analysed by quantitative 1H NMR spectroscopy. In this approach, the aqueous phase is measured both before and after equilibration with n-octanol. When equal volumes of aqueous and octanol phases are used, the equilibrium concentration of the analyte in the octanol phase can be determined indirectly as the difference between its initial concentration in the aqueous phase and its concentration after equilibration. The partition coefficient is then calculated as:| Kow = (cinitw − cequilw)/cequilw | (8) |
Because the integrated area (I) of a 1H NMR signal is directly proportional to the concentration of the corresponding protons, and hence to the concentration of the analyte, it is possible to replace concentration values with NMR integrals. Although small variations in integral values before and after equilibrium may occur due to the magnetic field drift, this drift affects both the analyte signal and the water signal equally. Therefore, relative integrals (RI), defined as RI = I/Iwater, can be used to correct for such effects. The eqn (8) then becomes (9):
| Kow = (RIinitw−RIequilw)/RIequilw | (9) |
A microtube containing 700 µL of a H2O/D2O mixture (9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) was used to dissolve the Ag(I) complexes (AgBet: 3 mg, 4.03 µmol; AgDmg: 3 mg, 11.00 µmol; AgSar: 3 mg, 11.58 µmol). The mixture was placed in an ultrasonic bath at 30 °C for 10 minutes to ensure complete dissolution of the analytes. The solution was then quantitatively transferred to an NMR tube, and quantitative 1H NMR spectra were acquired. Subsequently, 700 µL of n-octanol was added, and the NMR tube was inverted at least 40 times to achieve equilibration. After allowing the phases to separate, a second 1H NMR spectrum was recorded 24 hours later. This extended equilibration period was necessary not only to ensure more accurate partition coefficient measurements but also to allow for a more complete separation of the individual phases before analysis.78
1) was used to dissolve the Ag(I) complexes (AgBet: 3 mg, 4.03 µmol; AgDmg: 3 mg, 11.00 µmol; AgSar: 3 mg, 11.58 µmol). The mixture was placed in an ultrasonic bath at 30 °C for 10 minutes to ensure complete dissolution of the analytes. The solution was then quantitatively transferred to an NMR tube, and quantitative 1H NMR spectra were acquired. Subsequently, 700 µL of n-octanol was added, and the NMR tube was inverted at least 40 times to achieve equilibration. After allowing the phases to separate, a second 1H NMR spectrum was recorded 24 hours later. This extended equilibration period was necessary not only to ensure more accurate partition coefficient measurements but also to allow for a more complete separation of the individual phases before analysis.78
Log
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/i_char_2009.gif) P theoretical estimation.
The DFT-optimized structures of the amino acid silver(I) complexes in water were subsequently re-optimized in octan-1-ol under the same conditions to obtain the corresponding free energy values. The octanol–water partition coefficient was then evaluated as the equilibrium constant between the species in water and in octan-1-ol without other corrections (e.g. for the partial miscibility of both solvents resulting in shifts of their permittivity).
P theoretical estimation.
The DFT-optimized structures of the amino acid silver(I) complexes in water were subsequently re-optimized in octan-1-ol under the same conditions to obtain the corresponding free energy values. The octanol–water partition coefficient was then evaluated as the equilibrium constant between the species in water and in octan-1-ol without other corrections (e.g. for the partial miscibility of both solvents resulting in shifts of their permittivity).
Molecular docking. The receptor (HSA) geometry was adopted from the crystal structure,115 and prepared (water molecules removed) using AutoDockTools 1.5.7 (part of MGLTools).116,117 The ligand structures (Ag amino acid complexes and drugs – ibuprofen, warfarin isomers) were optimized at the same level as for the other calculations in this study (M06-2X/6-311+G(d,p)/LANL2TZ(f) in CPCM for water, see the Theoretical estimation paragraph) to obtain the input geometry for the subsequent molecular docking. The VINA program74,75 was modified in order to enable calculations on silver-containing species (Fe atom was replaced by Ag atom in the source file atom_constants.h) using the parameters from the updated AutoDock 4 database.116 The docking calculations were run over the whole receptor structure (size 126 Å in each direction from the centre of the molecule) to generate the 8 best configurations. The PyMol program118 was used for analysis and depictions of the predicted ligand topology with respect to the receptor.
Inhibition of β-galactosidase activity of E. coli by silver(I) complexes. The ability of silver(I) complexes to influence enzyme activity was assessed based on the inhibition of β-galactosidase activity. E. coli cells were harvested at mid-logarithmic phase (OD590 = 0.6) by centrifugation at 3000g for 10 min. The pellet was washed three times with phosphate-buffered saline (PBS) and adjusted to OD420 = 1.0 in the same buffer. In a 96-well microplate, 60 µL of o-nitrophenyl-β-D-galactopyranoside (ONPG; 3 M) was combined with 2 µL of complexes (at various concentrations) and 138 µL of the washed E. coli suspension. Absorbance at 420 nm was recorded every 5 min for 60 min using a microplate reader. The enzyme activity in control conditions without silver(I) complexes containing just the solvent 100× diluted ethanol
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) water (1
water (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2) served as the control. The enzyme activity was evaluated as the % of enzyme activity in control conditions.
2) served as the control. The enzyme activity was evaluated as the % of enzyme activity in control conditions.
Evaluation of the inhibitory effect of silver(I) complexes on PCR reaction – inhibition of DNA polymerase activity. To assess whether the silver(I) complexes interfere with PCR amplification and thus inhibit DNA polymerase activity, we performed a standard PCR targeting the bacterial 16S rDNA in the presence and absence of the tested complexes. Exponentially growing bacterial cells were used as the source of template DNA. A ∼1400 bp fragment of the 16S rRNA gene was amplified using the universal primers 27F (forward) and 1495R (reverse). PCR reactions were prepared as a standard master mix according to the manufacturer's instructions, and silver(I) complexes were added to the reaction mixture at defined concentrations immediately before amplification. Control reactions without silver(I) complexes were run in parallel.
The PCR program consisted of an initial activation step at 95 °C for 2 min to activate the DNA polymerase, followed by 30 amplification cycles. Each cycle comprised denaturation of the double-stranded DNA at 95 °C for 30 s, annealing of the primers at 65 °C for 30 s, and extension at 72 °C for 1 min. After completion of the 30 cycles, a final extension step was carried out at 72 °C for 5 min to ensure complete synthesis of all amplification products.
The impact of the silver(I) complexes on PCR efficiency, and thus on DNA polymerase activity, was evaluated by comparing the presence and intensity of the 1400 bp amplification product in reactions containing silver(I) complexes with the control reactions without silver(I) complexes and with the reaction where just the solvent (ethanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) water = 1
water = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2) of the complexes was added.
2) of the complexes was added.
Measurement of intracellular reactive oxygen species (ROS) production. Intracellular ROS production in E. coli was quantified using the dichloro–dihydro–fluorescein diacetate (DCFH-DA) assay. Cells were grown to mid-log phase, collected by centrifugation at 3.000g for 10 min, washed twice with PBS, and adjusted to OD590 = 0.5. The bacterial suspension was mixed with 10 μM concentrations of silver(I) complexes in a 96-well microplate. Subsequently, DCFH-DA (0.1 mM) was added immediately to each well, and the plate was incubated at 37 °C for 1 h. After that fluorescence was measured at excitation 488 nm and emission 525 nm using a Synergy H1 microplate reader (BioTek, USA) each 15 min during 3 h. A bacterial suspension containing 100× diluted ethanol
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) water (1
water (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2) served as the control, whereas 1.5% H2O2 was used as the positive control for ROS induction. Statistical differences in ROS levels among treatments were evaluated using one-way ANOVA followed by Tukey's HSD post-hoc test.
2) served as the control, whereas 1.5% H2O2 was used as the positive control for ROS induction. Statistical differences in ROS levels among treatments were evaluated using one-way ANOVA followed by Tukey's HSD post-hoc test.
Flow cytometry analyses. For the purposes of flow cytometry analyses, MDA-MB-231 breast cancer cells were seeded into medium-sized Petri dishes (25 × 104 cells per Petri dish) and cultured for 24 hours at 37 °C and 5% CO2. Subsequently, the tested compounds AgSar, AgBet, AgDmg, and AgGlyGlyGly were applied to the cells at their respective IC50 concentrations. After 6 and 24 hours of exposure, the cells were collected, trypsinized, and centrifuged. The resulting cell pellets were resuspended in PBS and divided into three analyses: MMP, Annexin, and cell cycle. Fluorescence was measured using a FACSCalibur flow cytometer (Becton Dickinson, San Jose, CA, USA). Flow cytometry output data were evaluated using FlowJo 10.0 software (Tree Star, Inc., Ashland, OR, USA, RRID:SCR_008520).
Cell cycle analysis. The redistribution of cells among the different phases of the cell cycle was analysed using flow cytometry. Cells were seeded and cultured according to the method described above. After each time interval (6 and 24 hours), the cells were trypsinized, centrifuged, and resuspended in PBS. For the purpose of cell cycle analysis, the cell suspension was fixed with ice-cold 70% ethanol and stored at −20 °C until analysis. On the day of analysis, the samples were centrifuged and incubated for 30 minutes at room temperature with a staining solution consisting of 0.5 mg ml−1 ribonuclease A, 0.2% final concentration of Triton X-100, and 0.025 mg ml−1 propidium iodide in 500 µl PBS (Sigma Aldrich, St. Louis, MO, USA). After incubation, flow cytometric analysis of cell cycle distribution was performed. Fluorescence was measured using a FACSCalibur flow cytometer (Becton Dickinson, San Jose, CA, USA). Cell cycle distribution was determined by deconvolution of DNA content histograms, after discrimination of doublets (FL-2-W vs. FL-2-A) and other cellular aggregates. Experimental data were analyzed using FlowJo 10.0 software (Tree Star, Inc., Ashland, OR, USA, RRID:SCR_008520) with the Dean–Jett–Fox model.
Mitochondrial membrane potential analysis. For the purpose of MMP analysis, cells were seeded and cultured according to the method described above. To monitor changes in mitochondrial membrane potential, the fluorescent probe tetramethylrhodamine ethyl ester (TMRE, 1
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100) was added to the cells resuspended in PBS. This probe functions by incorporating into intact mitochondria with a high membrane potential. After a 30 minute incubation at room temperature, flow cytometry was performed and the signal was detected in the FL-2 channel (585/42) and displayed as a single-parameter density plot. Subsequently, the percentage of cells with decreased MMP was determined.
100) was added to the cells resuspended in PBS. This probe functions by incorporating into intact mitochondria with a high membrane potential. After a 30 minute incubation at room temperature, flow cytometry was performed and the signal was detected in the FL-2 channel (585/42) and displayed as a single-parameter density plot. Subsequently, the percentage of cells with decreased MMP was determined.
Annexin V/PI staining. Cells were treated, collected, and centrifuged, and the resulting cell pellets were resuspended in PBS. The samples were then incubated with Annexin V–Alexa Fluor® 647 (1
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100, Thermo Scientific, Rockford, IL, USA) for 30 minutes at room temperature. After incubation, PI (5 mg ml−1, 1
100, Thermo Scientific, Rockford, IL, USA) for 30 minutes at room temperature. After incubation, PI (5 mg ml−1, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500, Sigma-Aldrich, Steinheim, Germany) was added to the cells to distinguish intact, apoptotic, and necrotic cells. Flow cytometric analysis was subsequently performed based on FL-2 vs. FL-4 channel signals.
500, Sigma-Aldrich, Steinheim, Germany) was added to the cells to distinguish intact, apoptotic, and necrotic cells. Flow cytometric analysis was subsequently performed based on FL-2 vs. FL-4 channel signals.
Statistical analyses. Results are expressed as mean ± standard deviation (SD). Statistical analyses were performed using one-way analysis of variance (ANOVA) with Dunnett's post hoc test. All experimental data were analysed using GraphPad Prism (version 9.0.2., La Jolla, California, USA; RRID:SCR_002798). P-Values < 0.05 (vs. untreated cells) were considered statistically significant. All experiments were performed in independent replicates.
ctDNA interactions. The fluorescence emission spectra of the silver(I) complexes were recorded since the complexes exhibited intrinsic fluorescence under the experimental conditions, allowing their interaction with ctDNA to be investigated by spectrofluorimetric titration.
Working solutions of silver(I) complexes (AgSar, AgDmg, AgBet, AgGlyGlyGly) were prepared by dissolving their powdered forms in distilled water to obtain a final concentration of 10 mM.
A ctDNA (calf thymus deoxyribonucleic acid) stock solution was prepared by dissolving the sodium salt of ctDNA in Tris-HCl buffer (pH = 7.4) at a ratio of 3 mg of ctDNA per 1 mL of buffer. The solution was stirred continuously at 4 °C for 24 h to ensure complete dissolution. The concentration of ctDNA was determined spectrophotometrically using a UV-Vis spectrophotometer (Specord S300) and was found to be 2.8901 mM.
Fluorometric titration of silver(I) complexes with ctDNA. Fluorescence emission spectra were recorded using a Varian Cary Eclipse fluorescence spectrophotometer with a 1 × 1 cm quartz cuvette at ambient temperature. The excitation wavelengths, λex, were set at 275 nm for AgSar, 267 nm for AgDmg, 267 nm for AgBet, and 255 nm for AgGlyGlyGly. Emission spectra were collected over the wavelength range of 283–340 nm.
For each experiment, the cuvette contained Tris-HCl buffer (pH = 7.4) and the respective silver(I) complex (AgSar or AgDmg or AgBet or AgGlyGlyGly) at an initial concentration of 49.75 μM. The solution was then titrated by successive additions of ctDNA over a concentration range of 0–21.41 μM, and the changes in fluorescence emission intensity were recorded after each addition.
The obtained data were processed graphically using GraphPad Prism 6 software, and numerical calculations were performed using Microsoft Office Excel 365. Binding constants (Kb) and the number of binding sites (n) was determined using the logarithmic form of the Stern–Volmer eqn (10):64
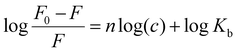 | (10) |
Conclusion
Naturally occurring N-methyl and N-glycyl derivatives of glycine, in combination with antimicrobial and anticancer silver(I) ions, yielded potent, water-soluble drug candidates. Titration data and density functional theory (DFT) calculations confirmed a pronounced preference for carboxylate coordination of the ligand to the central silver(I) ion, particularly in the presence of ligand excess in solution, across all Ag(I)-N-methyl and Ag(I)-N-glycyl glycine derivative systems. The dominance of carboxylate coordination was further supported by single-crystal structures of the AgSar, AgDmg, and AgBet complexes, obtained via a straightforward, one-step synthesis using biologically and environmentally acceptable solvents. Structural characterization revealed recurring mononuclear motifs in AgSar and AgDmg, while AgBet displayed a polynuclear architecture with crystallographically independent silver(I) ions forming extended polymeric structures. All complexes exhibited argentophilic interactions as a common structural feature.Stability assessments of the complexes in 1% DMSO and in a 0.1 M KNO3 stock electrolyte revealed only minor, time-invariant changes in 1H NMR chemical shifts and reduction potentials upon carboxylate coordination to the silver(I) center, suggesting moderate kinetic lability. In contrast, the complexes demonstrated high stability in biological growth media (RPMI, SB, and MHB), showing no evidence of precipitation and distinct UV-Vis spectral responses compared to the individual components (AgNO3 and amino acids/triglycine), indicating their persistence in complex biological environments.
The antimicrobial and antifungal efficacy of the complexes was significantly enhanced compared to AgNO3. Against Staphylococcus aureus, AgSar and AgDmg showed a 5-fold, and AgBet and AgGlyGlyGly a 10-fold, increase in activity. Against Escherichia coli, AgSar exhibited a 5-fold, and AgDmg, AgBet, and AgGlyGlyGly a 10-fold, enhancement. In the case of Candida parapsilosis, AgSar and AgGlyGlyGly showed a 2-fold, AgDmg a 4-fold, and AgBet a 10-fold increase in activity.
Remarkable antifungal activity was observed, surpassing that of silver(I) sulfadiazine (AgSD): activity against Rhizopus oryzae was more than 6-fold higher for AgSar and AgDmg, and over 10-fold for AgBet and AgGlyGlyGly. Activity against Alternaria alternata was increased 4-fold (AgSar and AgDmg) and 2-fold (AgBet), while against Microsporum gypseum, activity was 20-fold (AgSar and AgDmg) and 10-fold (AgBet) higher than AgNO3.
In anticancer assays, the MDA-MB-231 human mammary adenocarcinoma cell line showed the highest sensitivity to all tested complexes. Cytotoxicity was over 3-fold higher for AgSar and AgDmg, more than 2-fold for AgBet, and comparable (1-fold) for AgGlyGlyGly relative to cisPt. Additionally, AgSar and AgDmg exhibited significantly greater selectivity indices (SI = 4.3 and 4.0, respectively) compared to cisPt (SI = 1.2), highlighting their therapeutic potential.
Safety profiles assessed using the in ovo CAM assay confirmed the non-toxic nature of the complexes upon topical application to skin and mucosal surfaces. Furthermore, the complexes demonstrated potential to reduce neovascularization, suggesting possible utility in anticancer therapies.
Structure–activity relationship analyses and bioavailability assessments, based on Lipinski's parameters and human serum albumin (HSA) binding studies, indicated that ligand structure – particularly side chain polarity, molecular size, and the presence of hydrophobic or charged groups – plays a key role in determining biological performance. Considering the potential mechanism of action, the results indicate that the antimicrobial and anticancer activity of the complexes is mediated through a multi-targeted mode of action. Excessive production of reactive oxygen species (ROS) induces a pro-oxidant intracellular environment, while direct binding of Ag+ ions or Ag–ligand species to enzymes results in functional inactivation of key bacterial proteins. Concurrent damage to membrane-associated enzymes and the DNA replication machinery provides a plausible explanation for the pronounced bactericidal efficacy of the complexes, as well as for the reduced likelihood of resistance development. Moreover, analysis of the effects of the silver(I) complexes on cell cycle progression in breast cancer cells, in which the complexes demonstrated the highest efficacy, indicated that all tested complexes induced a time-dependent disruption of the mitochondrial membrane potential, supporting the potential involvement of mitochondria-mediated apoptosis in their cytotoxic effects. In addition, interaction with DNA may represent another contributing mechanism of action.
In conclusion, water-soluble silver(I) complexes with N-methyl and N-glycyl derivatives of glycine represent promising, bioavailable candidates for pharmaceutical applications. These non-toxic ligands serve as effective carriers for antimicrobial and antiproliferative silver(I) ions and likely facilitate their transport into pathogenic microorganisms or cancer cells. However, due to their relatively low thermodynamic stability and kinetic lability, it is essential to employ a controlled excess of ligand to ensure effective delivery at the target site.
Author contributions
Gabriela Kuzderová performed the potentiometric titrations and synthesis, and she investigated the complexes and contributed to the writing and finalization of the original draft manuscript (investigation, writing – original draft, visualization). Róbert Gyepes measured and solved the crystal structures of the complexes (investigation). Alan Liška investigated, designed and performed computational methods and contributed to the writing of the original draft manuscript (investigation, methodology, software, writing – original draft, visualization). Jana Havlíčková performed the thermal analysis measurements (investigation). Mária Vilková investigated and validated the NMR spectra and contributed to the writing of the original draft manuscript (investigation, writing – original draft, visualization). Simona Žiláková measured anticancer activity (investigation). Martin Kello designed and supervised the anticancer experiments, analysed and interpreted the data, and performed statistical analyses (investigation, validation). Tomáš Pagáč measured antimicrobial activity (investigation). Petra Olejníková designed and supervised the antimicrobial experiments and contributed to the writing of the original draft manuscript (investigation, writing – original draft, visualization). Eva Petrovová performed in ovo measurements (investigation). Ľudmila Balážová designed and investigated the in ovo experiments and contributed to the writing of original draft manuscript (investigation, validation, writing – original draft, visualization). Henrieta Matajová investigated DNA binding properties (investigation). Mária Kožurková designed and supervised the spectral experiments, analysed, interpreted, and validated data. Danica Sabolová designed and supervised the spectral experiments, analysed, interpreted, and validated the data, and contributed to the original draft manuscript (supervision, validation, writing – original draft, visualization). Michaela Rendošová contributed to the writing of the original draft manuscript (investigation, writing – original draft, visualization). Zuzana Vargová supervised the students during the complexes’ preparation and investigation and is responsible for the idea and concept of the article (conceptualization, methodology, validation, supervision, visualization, writing – original draft, writing – review & editing). All authors have read and agreed to the published version of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
Data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5dt03032c.CCDC 2473205–2473207 contain the supplementary crystallographic data for this paper.119a–c
Acknowledgements
This work was financially supported by Slovak grant agencies VEGA, KEGA and APVV (Scientific Grant Agency, Cultural and Educational Grant Agency of Ministry of Education, Science, Research and Sport of the Slovak Republic and Slovak Research and Development Agency): VEGA 1/0268/24, 1/0498/23, KEGA 007UPJŠ-4/2024, APVV-23-0372 and VVGS vvgs-2025-3528. Computational resources were provided by the e-INFRA CZ project (ID: 90254), supported by the Ministry of Education, Youth and Sports of the Czech Republic.References
- L. Ronga, M. Varcamonti and D. Tesauro, Structure–Activity Relationships in NHC–Silver Complexes as Antimicrobial Agents, Molecules, 2023, 28(11), 4435, DOI:10.3390/molecules28114435.
- A. Salleh, R. Naomi, E. Mahmoudi, N. D. Utami, A. W. Mohammad, N. Mustafa and M. B. Fauzi, The Potential of Silver Nanoparticles for Antiviral and Antibacterial Applications: A Mechanism of Action, Nanomaterials, 2020, 10(8), 1566, DOI:10.3390/nano10081566.
- F. C. Carolin and T. Kamalesh, Advances in Stabilization of Metallic Nanoparticles with Biosurfactants—A Review on Current Trends, Heliyon, 2024, 10, e29773 CrossRef PubMed.
- Y.-F. Zhang, Y.-K. Yin, H. Zhang and Y.-F. Han, Metal N-Heterocyclic Carbene Complexes as Potential Metallodrugs in Antitumor Therapy, Coord. Chem. Rev., 2024, 514, 215941, DOI:10.1016/j.ccr.2024.215941.
- J. S. Möhler, W. Sim, M. A. Blaskovich, M. A. Cooper and Z. M. Ziora, Silver Bullets: A New Lustre on an Old Antimicrobial Agent, Biotechnol. Adv., 2018, 36(5), 1391–1411, DOI:10.1016/j.biotechadv.2018.05.004.
- R. H. Allen, S. P. Stabler and J. Lindenbaum, Serum Betaine, N,N-Dimethylglycine and N-Methylglycine Levels in Patients with Cobalamin and Folate Deficiency and Related Inborn Errors of Metabolism, Metabolism, 1993, 42, 1448–1460, DOI:10.1016/0026-0495(93)90198-W.
- M. Pietzke, J. Meiser and A. Vazquez, Formate Metabolism in Health and Disease, Mol. Metab., 2020, 33, 23–37, DOI:10.1016/j.molmet.2019.05.012.
- J.-C. Lin, M.-H. Chan, M.-Y. Lee, Y.-C. Chen and H.-H. Chen, N,N-Dimethylglycine Differentially Modulates Psychotomimetic and Antidepressant-Like Effects of Ketamine in Mice, Prog. Neuropsychopharmacol. Biol. Psychiatry, 2016, 71, 7–13, DOI:10.1016/j.pnpbp.2016.06.002.
- G. Zhao, F. He, C. Wu, P. Li, N. Li, J. Deng, G. Zhu, W. Ren and Y. Peng, Betaine in Inflammation: Mechanistic Aspects and Applications, Front. Immunol., 2018, 9, 1070, DOI:10.3389/fimmu.2018.01070.
- N. C. Kasuga, R. Yamamoto, A. Hara, A. Amano and K. Nomiya, Molecular Design, Crystal Structure, Antimicrobial Activity and Reactivity of Light-Stable and Water-Soluble Ag–O Bonding Silver(I) Complexes, Dinuclear Silver(I) N-Acetylglycinate, Inorg. Chim. Acta, 2006, 359(13), 4412–4416, DOI:10.1016/j.ica.2006.04.007.
- K. Nomiya and H. Yokoyama, Syntheses, Crystal Structures and Antimicrobial Activities of Polymeric Silver(I) Complexes with Three Amino Acids [Aspartic Acid (H2asp), Glycine (Hgly) and Asparagine (Hasn)], J. Chem. Soc., Dalton Trans., 2002,(12), 2483–2490, 10.1039/B200684G.
- N. C. Kasuga, R. Yoshikawa, Y. Sakai and K. Nomiya, Syntheses, Structures, and Antimicrobial Activities of Remarkably Light-Stable and Water-Soluble Silver Complexes with Amino Acid Derivatives, Silver(I) N-Acetylmethioninates, Inorg. Chem., 2012, 51(3), 1640–1647, DOI:10.1021/ic202358e.
- K. Nomiya, S. Takahashi, R. Noguchi, S. Nemoto, T. Takayama and M. Oda, Synthesis and Characterization of Water-Soluble Silver(I) Complexes with L-Histidine (H2his) and (S)-(−)-2-Pyrrolidone-5-Carboxylic Acid (H2pyrrld) Showing a Wide Spectrum of Effective Antibacterial and Antifungal Activities. Crystal Structures of Chiral Helical Polymers [Ag(Hhis)]n and {[Ag(Hpyrrld)]2}n in the Solid State, Inorg. Chem., 2000, 39(15), 3301–3311, DOI:10.1021/ic990526o.
- K. Nomiya, K. Tsuda, T. Sudoh and M. Oda, Ag(I)–N Bond-Containing Compound Showing Wide Spectra in Effective Antimicrobial Activities: Polymeric Silver(I), Imidazolate, J. Inorg. Biochem., 1997, 68(1), 39–44, DOI:10.1016/S0162-0134(97)00006-8.
- G. Kuzderová, M. Rendošová, R. Gyepes, M. Almáši, D. Sabolová, M. Vilková, P. Olejníková, D. Hudecová, M. Kello and Z. Vargová, In Vitro Biological Evaluation and Consideration about Structure–Activity Relationship of Silver(I) Aminoacidate Complexes, J. Inorg. Biochem., 2020, 210, 111170, DOI:10.1016/j.jinorgbio.2020.111170.
- G. Kuzderová, S. Sovová, M. Rendošová, R. Gyepes, D. Sabolová, I. Kožárová, Ľ. Balážová, M. Vilková, M. Kello, A. Liška and Z. Vargová, Influence of Proline and Hydroxyproline as Antimicrobial and Anticancer Peptide Components on the Silver(I) Ion Activity: Structural and Biological Evaluation with a New Theoretical and Experimental SAR Approach, Dalton Trans., 2024, 53(26), 10834–10850, 10.1039/D4DT00389F.
- G. Kuzderová, M. Rendošová, R. Gyepes, S. Sovová, D. Sabolová, M. Vilková, P. Olejníková, I. Bačová, S. Stokič and M. Kello, et al., Antimicrobial and Anticancer Application of Silver(I), Dipeptide Complexes, Molecules, 2021, 26, 6335, DOI:10.3390/molecules26216335.
- M. Rendošová, R. Gyepes, I. C. Maruščáková, D. Mudroňová, D. Sabolová, M. Kello, M. Vilková, M. Almáši, V. Huntošová, O. Zemek and Z. Vargová, An In Vitro Selective Inhibitory Effect of Silver(I) Aminoacidates against Bacteria and Intestinal Cell Lines and Elucidation of the Mechanism of Action by Means of DNA Binding Properties, DNA Cleavage and Cell Cycle Arrest, Dalton Trans., 2021, 50(3), 936–953, 10.1039/D0DT03332D.
- R. Groom, I. J. Bruno, M. P. Lightfoot and S. C. Ward, The Cambridge Structural Database, Acta Crystallogr., Sect. B: Struct. Sci., 2016, 72(2), 171–179, DOI:10.1107/S2052520616003954.
- E. Martell, R. M. Smith and R. J. Motekaitis, NIST Standard Reference Database 46: Critically Selected Stability Constants of Metal Complexes, Version 7.0; NIST Standard Reference Data, Gaithersburg, MD, USA, 2003 Search PubMed.
- M. Rendošová, Z. Vargová, J. Kuchár, D. Sabolová, Š. Levoča, J. Kudláčová, D. Paulíková, D. Hudecová, V. Helebrandtová, M. Almáši, M. Vilková, M. Dušek and D. Bobáľová, New Silver Complexes with Bioactive Glycine and Nicotinamide Molecules—Characterization, DNA Binding, Antimicrobial and Anticancer Evaluation, J. Inorg. Biochem., 2017, 168, 1–12, DOI:10.1016/j.jinorgbio.2016.11.010.
- E. Martell and R. M. Smith, Aminocarboxylic Acids, in Critical Stability Constants, Springer, Boston, MA, 1982, pp. 1–58. DOI:10.1007/978-1-4615-6761-5_1.
- J. Israéli and R. Volpé, Mixed Complexes of Bivalent Cations Containing Methyliminodiacetate and Glycinates, Inorg. Chim. Acta, 1972, 6, 5–7, DOI:10.1016/S0020-1693(00)91748-1.
- E. Martell and R. M. Smith, Peptides, in Critical Stability Constants, Springer, Boston, MA, 1982, pp. 91–120. DOI:10.1007/978-1-4615-6761-5_3.
- Rostášová, M. Vilková, Z. Vargová, M. Walko, M. Almáši, J. Imrich and I. Lukeš, Dipeptide interactions with Zn(II)–cyclen artificial model for molecular recognition, J. Mol. Recognit., 2015, 28(4), 211–219, DOI:10.1002/jmr.2416.
- Y. Rajhi, Y.-H. Ju, A. E. Angkawijaya and A. E. Fazary, Complex Formation Equilibria and Molecular Structure of Divalent Metal Ions–Vitamin B3−Glycine Oligopeptides Systems, J. Solution Chem., 2013, 42, 2409–2442, DOI:10.1007/s10953-013-0116-5.
- M. Kývala and I. Lukeš, International Conference, Chemometrics ‘95, Pardubice, Czech Republic, p. 63, full version of “OPIUM” available (free of charge) on https://www.natur.cuni.cz/~kyvala/opium.html, 1995.
- Y. Podolyan and J. Leszczynski, MaSK: A Visualization Tool for Teaching and Research in Computational Chemistry, Int. J. Quantum Chem., 2009, 109(1), 8–16, DOI:10.1002/qua.21662.
- H. Schmidbaur and A. Schier, Argentophilic Interactions, Angew. Chem., Int. Ed., 2015, 54(3), 746–784, DOI:10.1002/anie.201405936.
- N. Banti, C. P. Raptopoulou, V. Psycharis and S. K. Hadjikakou, Novel Silver Glycinate Conjugate with 3D Polymeric Intermolecular Self-Assembly Architecture: An Antiproliferative Agent Which Induces Apoptosis on Human Breast Cancer Cells, J. Inorg. Biochem., 2021, 216, 111351, DOI:10.1016/j.jinorgbio.2020.111351.
- Z. Rzączyńska, R. Mrozek and M. Sikorska-Iwan, Crystal Structure, Spectroscopy and Thermal Study of Manganese(II) Complex with Sarcosine, Pol. J. Chem., 2002, 76, 29–35 Search PubMed.
- M. Fleck, V. V. Ghazaryan and A. M. Petrosyan, Crystal Structure at 296 and 150 K, Vibrational Spectra and Thermal Behaviour of Sarcosine Sarcosinium Nitrate, Z. Kristallogr. - Cryst. Mater., 2012, 227(12), 819–824, DOI:10.1524/zkri.2012.1536.
- R. V. Krishnakumar, M. P. Rameela and S. Natarajan, Crystal Structure of Sarcosine Cadmium Chloride, Cryst. Res. Technol., 1996, 31(2), 203–207, DOI:10.1002/crat.2170310214.
- L. Yang, D. R. Powell and R. P. Houser, Structural Variation in Copper(I) Complexes with Pyridylmethylamide Ligands: Structural Analysis with a New Four-Coordinate Geometry Index, τ4, Dalton Trans., 2007,(9), 955–964, 10.1039/B617136B.
- M. R. Silva, A. M. Beja, J. A. Paixão and J. S. Martín-Gil, Synthesis, Crystal Structure and Magnetic Properties of an Alternating Manganese Chain, J. Solid State Chem., 2006, 179(7), 2054–2058, DOI:10.1016/j.jssc.2006.03.024.
- M. R. Silva, A. M. Beja, J. A. Paixão and J. S. Martín-Gil, Di-μ-chloro-1:2κ2Cl;3:4κ2Cl-Hexachloro-1κ3Cl,4κ3Cl-Tetra-μ-dimethylglycine-2:3κ8O:O′-Tetracopper(II), Acta Crystallogr., Sect. C: Cryst. Struct. Commun., 2005, 61(8), m380–m382, DOI:10.1107/S0108270105021591.
- X.-M. Chen and T. C. W. Mak, Metal–Betaine Interactions. Part 3. Crystal Structures of Polymeric Diaquabis(betaine)disilver(I) Dinitrate and Bis(pyridine Betaine)disilver(I) Diperchlorate, J. Chem. Soc., Dalton Trans., 1991,(5), 1219–1222, 10.1039/dt9910001219.
- Z. Vargová, P. Olejníková, G. Kuzderová, M. Rendošová, J. Havlíčková, R. Gyepes and M. Vilková, Silver(I) Complexes with Amino Acid and Dipeptide Ligands—Chemical and Antimicrobial Relevant Comparison (Mini Review), Bioorg. Chem., 2023, 141, 106907, DOI:10.1016/j.bioorg.2023.106907.
- J. Yamada, H. Hashimoto, Y. Inomata and T. Takeuchi, Preparation and Characterization of Cadmium(II) Halide Complexes with N-Substituted Glycines, and the Crystal Structures of Dichloro(N-methylglycine)cadmium(II) and Diaquadichloro(N,N-dimethylglycine)cadmium(II), Bull. Chem. Soc. Jpn., 1994, 67(12), 3224–3230, DOI:10.1246/bcsj.67.3224.
- N. D. Savić, B. B. Petković, S. Vojnovic, M. Mojicevic, H. Wadepohl, K. Olaifa, E. Marsili, J. Nikodinovic-Runic, M. I. Djuran and B.Đ Glišić, Dinuclear Silver(I) Complexes with a Pyridine-Based Macrocyclic Type of Ligand as Antimicrobial Agents against Clinically Relevant Species: The Influence of the Counteranion on the Structure Diversification of the Complexes, Dalton Trans., 2020, 49(31), 10880–10894, 10.1039/D0DT01272F.
- J. M. S. Cardoso, A. M. Galvão, S. I. Guerreiro, J. H. Leitão, A. C. Suarez and M. F. N. N. Carvalho, Antibacterial Activity of Silver Camphorimine Coordination Polymers, Dalton Trans., 2016, 45(16), 7114–7123, 10.1039/C6DT00099A.
- P. F. Wen, X. X. Zhuo and L. Wang, Skin Barrier Dysfunction in Cutaneous T-Cell Lymphoma: From Pathogenic Mechanism of Barrier Damage to Treatment, Crit. Rev. Oncol. Hematol., 2025, 205, 104559, DOI:10.1016/j.critrevonc.2024.104559.
- N. P. Luepke, Hen's Egg Chorioallantoic Membrane Test for Irritation Potential, Food Chem. Toxicol., 1985, 23(2), 287–291, DOI:10.1016/0278-6915(85)90030-4.
- Ľ. Balážová, Z. Bedlovičová, N. Daneu, M. Baláž, Z. Lukáčová Bujňáková, Ľ. Tkáčiková, M. Takáčová and A. Jacková, Silver Nanoparticles Produced In Vitro by Berberis vulgaris Fruit and Their Antioxidant, Antimicrobial and Ex Ovo Irritation Potential Study, BioNanoScience, 2024, 14(2), 867–879, DOI:10.1007/s12668-024-01400-5.
- D. Krchnák, L. Balazova, M. Hanko, D. Zigrayová and M. Spaglová, In Situ Gelling Dexamethasone Oromucosal Formulation: Physical Characteristics Influencing Drug Delivery, Gels, 2025, 11(1), 26, DOI:10.3390/gels11010026.
- M. H. Ali, O. Alam, A. Ali, M. U. Ali, S. Parvez, E. Aldosari, S. Baboota and J. Ali, Donepezil and Embelin Loaded Nanostructured Lipid Carriers for Direct Brain Delivery as An Intervention for Alzheimer's Disease: Formulation Design, Optimization and Evaluation, J. Cluster Sci., 2024, 35(4), 1021–1044, DOI:10.1007/s10876-023-02531-7.
- R. H. F. Costa, A. P. Krawczyk-Santos, J. F. M. Andrade, G. N. Barbalho, R. M. Almeida, Y. K. M. Nóbrega, M. Cunha, G. M. Gelfuso, S. F. Taveira and T. Gratieri, α-Cyclodextrin-Based Poly(pseudo)rotaxane for Antifungal Drug Delivery to the Vaginal Mucosa, Carbohydr. Polym., 2023, 302, 120420, DOI:10.1016/j.carbpol.2022.120420.
- J. H. Draize, G. Woodard and H. O. Calvery, Methods for the Study of Irritation and Toxicity of Substances Applied Topically to the Skin and Mucous Membranes, J. Pharmacol. Exp. Ther., 1944, 82(3), 377–390 CrossRef CAS.
- P. Blasi, A. Schoubben, G. Traina, G. Manfroni, L. Barberini, P. F. Alberti, C. Cirotto and M. Ricci, Lipid Nanoparticles for Brain Targeting III. Long-Term Stability and In Vivo Toxicity, Int. J. Pharm., 2013, 454(1), 316–323, DOI:10.1016/j.ijpharm.2013.06.037.
- A. Kessler, R. S. Langer, N. A. Pless and J. Folkman, Mast-Cells and Tumor Angiogenesis, Int. J. Cancer, 1976, 18(5), 703–709, DOI:10.1002/ijc.2910180520.
- P. Carmeliet, Mechanisms of Angiogenesis and Arteriogenesis, Nat. Med., 2000, 6(4), 389–395, DOI:10.1038/74651.
- J. Xiao and Z. P. Xu, Roles of Noncoding RNAs in Diabetic Retinopathy: Mechanisms and Therapeutic Implications, Life Sci., 2024, 357, 123092, DOI:10.1016/j.lfs.2024.123092.
- F. Zhao, Y. N. He, Z. H. Zhao, J. R. He, H. Huang, K. L. Ai, L. Liu and X. Cai, The Notch Signaling-Regulated Angiogenesis in Rheumatoid Arthritis: Pathogenic Mechanisms and Therapeutic Potentials, Front. Immunol., 2023, 14, 1272133, DOI:10.3389/fimmu.2023.1272133.
- L. J. Zhang, F. M. Xie, F. M. Zhang and B. Y. Lu, The Potential Roles of Exosomes in Pathological Cardiomyocyte Hypertrophy Mechanisms and Therapy, A Review, Medicine, 2024, 103(17), e37994, DOI:10.1097/MD.0000000000037994.
- J. Folkman, M. Bach, J. W. Rowe, F. Davidoff, P. Lambert, C. Hirsch, A. Goldberg, H. H. Hiatt, J. Glass and E. Henshaw, Tumor Angiogenesis—Therapeutic Implications, N. Engl. J. Med., 1971, 285(21), 1182 CrossRef CAS PubMed.
- P. Nowak-Sliwinska, K. Alitalo, E. Allen, A. Anisimov, A. C. Aplin, R. Auerbach, H. G. Augustin, D. O. Bates, J. R. van Beijnum, R. H. F. Bender, G. Bergers, A. Bikfalvi, J. Bischoff, B. C. Böck, P. C. Brooks, F. Bussolino, B. Cakir, P. Carmeliet, D. Castranova, A. M. Cimpean, O. Cleaver, G. Coukos, G. E. Davis, M. De Palma, A. Dimberg, R. P. M. Dings, V. Djonov, A. C. Dudley, N. P. Dufton, S. M. Fendt, N. Ferrara, M. Fruttiger, D. Fukumura, B. Ghesquière, Y. Gong, R. J. Griffin, A. L. Harris, C. C. W. Hughes, N. W. Hultgren, M. L. Iruela-Arispe, M. Irving, R. K. Jain, R. Kalluri, J. Kalucka, R. S. Kerbel, J. Kitajewski, I. Klaassen, H. K. Kleinmann, P. Koolwijk, E. Kuczynski, B. R. Kwak, K. Marien, J. M. Melero-Martin, L. Munn, R. F. Nicosia, A. Noel, J. Nurro, A. K. Olsson, T. V. Petrova, K. Pietras, R. Pili, J. W. Pollard, M. J. Post, P. H. A. Quax, G. A. Rabinovich, M. Raica, A. M. Randi, D. Ribatti, C. Ruegg, R. O. Schlingemann, S. Schulte-Merker, L. E. H. Smith, J. W. Song, S. A. Stacker, J. Stalin, A. N. Stratman, M. Van de Velde, V. W. M. van Hinsbergh, P. B. Vermeulen, J. Waltenberger, B. M. Weinstein, H. Xin, B. Yetkin-Arik, S. Yla-Herttuala, M. C. Yoder and A. W. Griffioen, Consensus Guidelines for the Use and Interpretation of Angiogenesis Assays, Angiogenesis, 2018, 21(3), 425–532, DOI:10.1007/s10456-018-9613-x.
- L. S. Gambino, N. G. Wreford, J. F. Bertram, P. Dockery, F. Lederman and P. A. W. Rogers, Angiogenesis Occurs by Vessel Elongation in Proliferative Phase Human Endometrium, Hum. Reprod., 2002, 17(5), 1199–1206, DOI:10.1093/humrep/17.5.1199.
- L. Faihs, B. Firouz, P. Slezak, C. Slezak, M. Weissensteiner, T. Ebner, N. G. Tabrizi-Wizsy, K. Schicho and P. Dungel, A Novel Artificial Intelligence-Based Approach for Quantitative Assessment of Angiogenesis in the Ex Ovo CAM Model, Cancers, 2022, 14(17), 4273, DOI:10.3390/cancers14174273.
- P. Takáč, R. Michalková, M. Čižmáriková, Z. Bedlovičová, Ľ. Balážová and G. Takáčová, The Role of Silver Nanoparticles in the Diagnosis and Treatment of Cancer: Are There Any Perspectives for the Future?, Life, 2023, 13(2), 466, DOI:10.3390/life13020466.
- F. A. di Masi, V. Trezza, M. Marino, M. Fasano and P. Ascenzi, Human Serum Albumin: From Bench to Bedside, Mol. Aspects Med., 2012, 33, 209–290, DOI:10.1016/j.mam.2011.12.002.
- M. Fasano, S. Curry, E. Terreno, M. Galliano, G. Fanali, P. Narciso and P. Ascenzi, The Extraordinary Ligand Binding Properties of Human Serum Albumin, IUBMB Life, 2005, 57, 787–796, DOI:10.1080/15216540500404093.
- J. Ghuman, P. A. Zunszain, I. Petitpas, A. A. Bhattacharya, M. Otagiri and S. Curry, Structural Basis of the Drug-Binding Specificity of Human Serum Albumin, J. Mol. Biol., 2005, 353, 38–52, DOI:10.1016/j.jmb.2005.07.075.
- Y. Sun, T. Hou, X. He, V. H. Man and J. Wang, Development and Test of Highly Accurate Endpoint Free Energy Methods. 2: Prediction of Logarithm of n-Octanol–Water Partition Coefficient (log P) for Druglike Molecules Using MM-PBSA Method, J. Comput. Chem., 2023, 44(13), 1300–1311, DOI:10.1002/jcc.27086.
- J. R. Lakowicz, Principles of fluorescence spectroscopy, Springer Publishing, New York, 3rd Edition, 2006, ISBN 03-873-1278-1 Search PubMed.
- S. Hashempour, N. Shahabadi, A. Adewoye, B. Murphy, C. Rouse, B. A. Salvatore, C. Stratton and E. Mahdavian, Binding Studies of AICAR and Human Serum Albumin by Spectroscopic, Theoretical, and Computational Methodologies, Molecules, 2020, 25(22), 5410, DOI:10.3390/molecules25225410.
- S.Ž Đurić, S. Vojnovic, T. P. Andrejević, N. L. Stevanović, N. D. Savić, J. Nikodinovic-Runic, B.Đ Glišić and M. I. Djuran, Antimicrobial Activity and DNA/BSA Binding Affinity of Polynuclear Silver(I) Complexes with 1,2-Bis(4-pyridyl)ethane/Ethene as Bridging Ligands, Bioinorg. Chem. Appl., 2020, 2020, 3812050, DOI:10.1155/2020/3812050.
- M. Rendošová, G. Kuzderová, R. Gyepes, M. Kello, P. Olejníková, M. Vilková, S. Gama, H. Matajová, M. Kožurková, A. Liška, S. Žiláková and Z. Vargová, Silver(I), Zinc(II) and Gallium(III) thiophene-2-carboxylates: Synthesis, solution and solid state characterization and bioevaluation, J. Inorg. Biochem., 2025, 270, 112936, DOI:10.1016/j.jinorgbio.2025.112936.
- F. Ayimbila, T. Tantimongcolwat, W. Ruankham, R. Pingaew, V. Prachayasittikul, V. Prachayasittikul, S. Prachayasittikul and K. Phopin, Exploring the binding interaction of 1,4-naphthoquinone derivative–human serum albumin complex by biophysics and molecular simulation, Sci. Rep., 2025, 15, 19249, DOI:10.1038/s41598-025-02787-1.
- G. D. J. B. Sudlow, D. J. Birkett and D. N. Wade, Further characterization of specific drug binding sites on human serum albumin, Mol. Pharmacol., 1976, 12, 1052–1061 CrossRef CAS PubMed.
- Y. Y. Fang, X. M. Yang, Y. Y. Li and C. L. Feng, Spectroscopic studies on the interaction of bovine serum albumin with Ginkgol C15:1 from Ginkgo biloba L., J. Lumin., 2015, 162, 203–211, DOI:10.1016/j.jlumin.2015.02.050.
- A. Thorarensen, R. W. Sarver, F. Tian, A. Ho, D. L. Romero and K. R. Marotti, Human serum albumin binding assay based on displacement of a non-selective fluorescent inhibitor, Bioorg. Med. Chem. Lett., 2007, 17, 4646–4649, DOI:10.1016/j.bmcl.2007.05.077.
- Q. Li, W.-Y. Yang, L.-L. Qu, H.-Y. Qi, Y. Huang and Z. Zhang, Interaction of Warfarin with Human Serum Albumin and Effect of Ferulic Acid on the Binding, J. Spectroscopy, 2014, 7, 834501, DOI:10.1155/2014/834501.
- S. Evoli, D. L. Mobley, R. Guzzi and B. Rizzuti, Multiple binding modes of ibuprofen in human serum albumin identified by absolute binding free energy calculations, Phys. Chem. Chem. Phys., 2016, 18, 32358–32368, 10.1039/C6CP05680F.
- J. Eberhardt, D. Santos-Martins, A. F. Tillack and S. Forli, AutoDock Vina 1.2.0: new docking methods, expanded force field, and python bindings, J. Chem. Inf. Model., 2021, 61, 3891–3898, DOI:10.1021/acs.jcim.1c00203.
- O. Trott and A. J. Olson, AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading, J. Comput. Chem., 2010, 31, 455–461, DOI:10.1002/jcc.21334.
- D. Kempińska, T. Chmiel, A. Kot-Wasik, A. Mróz, Z. Mazerska and J. Namieśnik, State of the Art and Prospects of Methods for Determination of Lipophilicity of Chemical Compounds, TrAC, Trends Anal. Chem., 2019, 113, 54–73, DOI:10.1016/j.trac.2019.01.011.
- C. Cumming and C. Rücker, Octanol–Water Partition Coefficient Measurement by a Simple 1H NMR Method, ACS Omega, 2017, 2(9), 6244–6249, DOI:10.1021/acsomega.7b01102.
- D. Soulsby and J. A. Chica, Determination of Partition Coefficients Using 1H NMR Spectroscopy and Time Domain Complete Reduction to Amplitude–Frequency Table (CRAFT) Analysis, Magn. Reson. Chem., 2017, 55(8), 724–729, DOI:10.1002/mrc.4582.
- S. K. Bhal, K. Kassam, I. G. Peirson and G. M. Pearl, The Rule of Five Revisited: Applying Log D in Place of Log P in Drug-Likeness Filters, Mol. Pharmaceutics, 2007, 4(4), 556–560, DOI:10.1021/mp0700209.
- C. N. Banti and S. K. Hadjikakou, Anti-proliferative and anti-tumor activity of silver(I) compounds, Metallomics, 2013, 5, 569–596, 10.1039/c3mt00046j.
- Z.-L. Niu, T.-T. Wu, Y.-Y. Wu, S.-H. Zhou, Z. Li, J.-C. Guo, S.-M. Zhou, S.-H. Deng, J.-Y. Xu and M.-J. Xie, Two novel silver(I) phenanthroline derivative complexes induce G2/M phase cycle arrest and apoptosis of MDA-MB-231 cancer cells by multiple mechanisms, Appl. Organomet. Chem., 2023, 37, e7123, DOI:10.1002/aoc.7123.
- J. Marinković, M. Jurišević, M. Jovanović, M. Milosavljević, I. Jovanović, S. J. Stević, M. Vesović, M. Nikolić, N. Nedeljković, A. Živanović, D. Tomović, A. Bukonjić, G. Radić and N. Gajović, Silver(I) complexes bearing S-alkyl thiosalicylic acid derivatives: DNA/BSA binding and antitumor activity in vitro and in vivo, Pharmaceutics, 2025, 17, 1340, DOI:10.3390/pharmaceutics17101340.
- K. E. Roberts, Z. Engelbrecht, K. Potgieter, R. Meijboom and M. J. Cronjé, Silver(I) bromide phosphines induce mitochondrial-mediated apoptosis in malignant human colorectal cells, Biomedicines, 2023, 11, 2794, DOI:10.3390/biomedicines11102794.
- A. A. Khandar, Z. Mirzaei-Kalar, N. Shahabadi, S. Hadidi, H. Abolhasani, S. A. Hosseini-Yazdi and A. Jouyban, Antimicrobial, cytotoxicity, molecular modeling and DNA cleavage/binding studies of zinc-naproxen complex: switching DNA binding mode of naproxen by coordination to zinc ion, J. Biomol. Struct. Dyn., 2022, 40(9), 4224–4236, DOI:10.1080/07391102.2020.1854858.
- N. Arshad, N. Abbas, M. H. Bhatti, N. Rashid, M. N. Tahir, S. Saleem and B. Mirza, Synthesis, crystal structure, DNA binding and in vitro biological studies of Ni(II), Cu(II) and Zn(II) complexes of N-phthaloylglycine, J. Photochem. Photobiol. B, Biol., 2012, 117, 228–239, DOI:10.1016/j.jphotobiol.2012.10.003.
- E. Movahedi and A. R. Rezvani, New silver(I) complex with diazafluorene based ligand: Synthesis, characterization, investigation of in vitro DNA binding and antimicrobial studies, J. Mol. Struct., 2017, 1139, 407–417, DOI:10.1016/j.molstruc.2017.03.042.
- C.-M. Che, M.-C. Tse, M. C. W. Chan, K.-K. Cheung, D. L. Phillips and K.-H. Leung, Spectroscopic Evidence for Argentophilicity in Structurally Characterized Luminescent Binuclear Silver(I) Complexes, J. Am. Chem. Soc., 2000, 122, 2464–2468 CrossRef CAS.
- A. A. Almaqwashi, T. Paramanathan, I. Rouzina and M. C. Williams, Mechanisms of small molecule–DNA interactions probed by single-molecule force spectroscopy, Nucleic Acids Res., 2016, 44(9), 3971–3988, DOI:10.1093/nar/gkw237.
- BenchChem, [Core Concepts of Hoechst 33258-DNA Binding], BenchChem, 2025 Search PubMed . [Online PDF]. Available at: https://www.benchchem.com/product/b15609105#hoechst-33258-binding-to-dna-minorgroove.
- BenchChem, [The Intricate Dance of Intercalation: A Technical Guide to Ethidium Bromide's DNA Binding Affinity and Specificity], BenchChem, 2025 Search PubMed . [Online PDF]. Available at: https://www.benchchem.com/product/b12369871#dna-intercalator-1-binding-affinity-and-specificity.
- P. L. Brown and C. Ekberg, Hydrolysis of Metal Ions, Wiley, 2016, pp. 725–730 Search PubMed.
- P. Pracht, F. Bohle and S. Grimme, Automated Exploration of the Low-Energy Chemical Space with Fast Quantum Chemical Methods, Phys. Chem. Chem. Phys., 2020, 22(14), 7169–7192, 10.1039/D0CP00252J.
- S. Grimme, Exploration of Chemical Compound, Conformer, and Reaction Space with Meta-Dynamics Simulations Based on Tight-Binding Quantum Chemical Calculations, J. Chem. Theory Comput., 2019, 15(5), 2847–2862, DOI:10.1021/acs.jctc.9b00143.
- P. Pracht, S. Grimme, C. Bannwarth, F. Bohle, S. Ehlert, G. Feldmann, J. Gorges, M. Müller, T. Neudecker, C. Plett, S. Spicher, P. Steinbach, P. Wesołowski and F. Zeller, J. Chem. Phys., 2024, 160(11), 114110, DOI:10.1063/5.0191877.
- C. Bannwarth, S. Ehlert and S. Grimme, GFN2-xTB—An Accurate and Broadly Parametrized Self-Consistent Tight-Binding Quantum Chemical Method with Multipole Electrostatics and Density-Dependent Dispersion Contributions, J. Chem. Theory Comput., 2019, 15(3), 1652–1671, DOI:10.1021/acs.jctc.8b01176.
- S. Ehlert, M. Stahn, S. Spicher and S. Grimme, Robust and Efficient Implicit Solvation Model for Fast Semiempirical Methods, J. Chem. Theory Comput., 2021, 17(7), 4250–4261, DOI:10.1021/acs.jctc.1c00471.
- Y. Zhao and D. G. Truhlar, The M06 Suite of Density Functionals for Main Group Thermochemistry, Thermochemical Kinetics, Noncovalent Interactions, Excited States, and Transition Elements: Two New Functionals and Systematic Testing of Four M06-Class Functionals and 12 Other Functionals, Theor. Chem. Acc., 2008, 120(1–3), 215–241, DOI:10.1007/s00214-007-0310-x.
- P. Pritchard, D. Altarawy, B. Didier, T. D. Gibson and T. L. Windus, A New Basis Set Exchange: An Open, Up-to-Date Resource for the Molecular Sciences Community, J. Chem. Inf. Model., 2019, 59(11), 4814–4820, DOI:10.1021/acs.jcim.9b00725.
- A. Klamt and G. Schürmann, COSMO: A New Approach to Dielectric Screening in Solvents with Explicit Expressions for the Screening Energy and Its Gradient, J. Chem. Soc., Perkin Trans., 1993, 2, 799–805, 10.1039/P29930000799.
- C. Amovilli, V. Barone, R. Cammi, E. Cancès, M. Cossi, B. Mennucci, C. S. Pomelli and J. Tomasi, Recent Advances in the Description of Solvent Effects with the Polarizable Continuum Model, Advances in Quantum Chemistry, Elsevier, 1998, 32, 227–261, DOI:10.1016/S0065-3276(08)60416-5.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, G. A. Petersson, H. Nakatsuji, X. Li, M. Caricato, A. V. Marenich, J. Bloino, B. G. Janesko, R. Gomperts, B. Mennucci, H. P. Hratchian, J. V. Ortiz, A. F. Izmaylov, J. L. Sonnenberg, D. W. Williams-Young, F. Ding, F. Lipparini, F. Egidi, J. Goings, B. Peng, A. Petrone, T. Henderson, V. G. Zakrzewski, J. Gao, N. Rega, G. Zheng, W. Liang, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, K. Throssell, J. A. Montgomery Jr., J. E. Peralta, F. Ogliaro, M. J. Bearpark, J. J. Heyd, E. N. Brothers, K. N. Kudin, V. N. Staroverov, T. A. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. P. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, J. B. Foresman and D. J. Fox, Gaussian 16, Revision C.01, Gaussian, Inc., Wallingford, CT, 2019 Search PubMed.
- M. Rendošová, R. Gyepes, A. Gucký, M. Kožurková, M. Vilková, P. Olejníková, M. Kello, A. Liška, I. Kléri, J. Havlíčková, A. Tamáš and Z. Vargová, In(III) pyridinecarboxylate complexes: Composition, solution equilibria estimation, bioevaluation and interactions with HSA, J. Inorg. Biochem., 2025, 262, 112738, DOI:10.1016/j.jinorgbio.2024.112738.
- V. S. Bryantsev, M. S. Diallo and W. A. Goddard III, Prediction of Solvation Free Energies of Charged Solutes Using Mixed Cluster/Continuum Models, J. Phys. Chem. B, 2008, 112(30), 9709–9719, DOI:10.1021/jp802665d.
- J. H. Jensen, Predicting Accurate Absolute Binding Energies in Aqueous Solution: Thermodynamic Considerations for Electronic Structure Methods, Phys. Chem. Chem. Phys., 2015, 17(19), 12441–12451, 10.1039/C5CP00628G.
- M. Sheldrick, SHELXT—Integrated Space-Group and Crystal-Structure Determination, Acta Crystallogr., Sect. A: Found. Adv., 2015, 71(1), 3–8, DOI:10.1107/S2053273314026370.
- G. M. Sheldrick, Crystal Structure Refinement with SHELXL, Acta Crystallogr., Sect. C: Struct. Chem., 2015, 71(1), 3–8, DOI:10.1107/S2053229614024218.
- L. Spek, Structure Validation in Chemical Crystallography, Acta Crystallogr., Sect. D: Biol. Crystallogr., 2009, 65(2), 148–155, DOI:10.1107/S090744490804362X.
- H. Putz and K. Brandenburg, DIAMOND—Crystal and Molecular Structure Visualization, Version 4.6, Crystal Impact GbR, Bonn, Germany, 2020 Search PubMed.
- S. Jantová, D. Hudecová, Š. Stankovský, K. Špirková and L. Ružeková, Antibacterial Effect of Substituted 4-Quinazolylhydrazines and Their Arylhydrazones Determined by a Modified Microdilution Method, Folia Microbiol., 1995, 40, 611–614, DOI:10.1007/BF02818517.
- V. Betina and D. Mičeková, Antimicrobial Properties of Fungal Macrolide Antibiotics, Z. Allg. Mikrobiol., 1972, 12, 355–364, DOI:10.1002/jobm.19720120502.
- D. Hudecová, S. Jantová, M. Melník and M. Uher, New Azidometalkojates and Their Biological Activity, Folia Microbiol., 1996, 41, 473–476, DOI:10.1007/BF02814660.
- B. Dudová, D. Hudecová, R. Pokorný, M. Mičková, M. Palicová, P. Segl’a and M. Melník, Copper Complexes with Bioactive Ligands, Folia Microbiol., 2002, 47, 225–229, DOI:10.1007/BF02817642.
- K. Paliwal, P. Haldar, P. S. Antharjanam and M. Kumar, Synthesis, Characterization, DNA/HSA Interaction, and Cytotoxic Activity of a Copper(II) Thiolate Schiff Base Complex and Its Corresponding Water-Soluble Stable Sulfinato–O Complex Containing Imidazole as a Co-Ligand, ACS Omega, 2023, 8(24), 21948–21968, DOI:10.1021/acsomega.3c01853.
- M. A. Qureshi and S. Javed, Aflatoxin B1 induced structural and conformational changes in bovine serum albumin: a multispectroscopic and circular dichroism-based study, ACS Omega, 2021, 6, 18054–18064, DOI:10.1021/acsomega.1c01799.
- S. Sugio, A. Kashima, S. Mochizuki, M. Noda and K. Kobayashi, Crystal structure of human serum albumin at 2.5 Å resolution, Protein Eng., 1999, 12, 439–446, DOI:10.1093/protein/12.6.439.
- S. J. R. S. Dallakyan, MGLTools, Reference Source., 2010 Search PubMed.
- G. M. Morris, R. Huey, W. Lindstrom, M. F. Sanner, R. K. Belew, D. S. Goodsell and A. J. Olson, AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility, J. Comput. Chem., 2009, 30, 2785–2791, DOI:10.1002/jcc.21256.
- M. A. Lill and M. L. Danielson, Computer-aided drug design platform using PyMOL, J. Comput.-Aided Mol. Des., 2011, 25(1), 13–19, DOI:10.1007/s10822-010-9395-8.
- (a) CCDC 2473205: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2p0kt3; (b) CCDC 2473206: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2p0kv4; (c) CCDC 2473207: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2p0kw5.
| This journal is © The Royal Society of Chemistry 2026 |




