Phenoxido-bridged open-cubane [Cu4(μ3-OH)2] and stepped-cubane [Cu4(μ3-OMe)2] complexes from a semicarbazone Schiff base: synthesis, anion-coordination-tunable structures, biomimicking functions, DNA binding, magnetic behavior and theoretical support
Anupama
Manna
a,
Mohammad Fawad
Ansari
b,
Farukh
Arjmand
 b,
Zvonko
Jagličić
b,
Zvonko
Jagličić
 c,
Radovan
Herchel
c,
Radovan
Herchel
 d and
Debashis
Ray
d and
Debashis
Ray
 *a
*a
aDepartment of Chemistry, Indian Institute of Technology Kharagpur, Kharagpur 721302, India. E-mail: dray@chem.iitkgp.ac.in
bDepartment of Chemistry, Aligarh Muslim University, Aligarh 202002, India
cInstitute of Mathematics, Physics and Mechanics & Faculty of Civil and Geodetic Engineering, University of Ljubljana, Ljubljana 1000, Slovenia
dDepartment of Inorganic Chemistry, Faculty of Science, Palacký University, 17. listopadu 12, 77146 Olomouc, Czech Republic
First published on 28th January 2026
Abstract
The bridging nature of in situ solvent-generated hydroxido (HO−) and methoxido (MeO−) groups has been utilized to obtain tetranuclear open-cubane and stepped-cubane [Cu4] complexes: [Cu4(L)2(H2O)2(μ3-OH)2](ClO4)4·3H2O (1) and [Cu4(L)2(μ-NO3)2(μ3-OMe)2](NO3)2·H2O (2) (HL = 2,6-bis-{(semicarbazidoimino)}-4-methylphenol). The ligand HL, possessing two metal ion-capturing bay regions, immediately seizes two copper(II) ions in solution to form Cu2L species, with their remaining coordination sites loosely occupied by H2O or NO3− groups, ultimately leading to the formation of complexes 1 and 2. The treatment of HL with Cu(ClO4)2·6H2O and Cu(NO3)2·3H2O metal ion salts is responsible for the in situ generation of the supporting HO− and MeO− linkers to sustain two different topologies from the same MeOH–H2O reaction medium. In 2, the bridging coordination of the nitrate anion to two copper(II) centers guides the approach of individual Cu2L units for Cu4 cluster formation, whereas for 1, the approach of the Cu2L units is different due to the coordination of labile H2O molecules to the copper(II) centers. These complexes have been characterized by X-ray crystallography, and their magnetic properties have been studied. Both the complexes exhibit interesting catalytic oxidation behavior for mimicking the enzyme behaviours of catecholase oxidase and phenoxazinone synthase, using model substrate 3,5-DTBCH2 and AP with the Kcat values 12.08, 1.845 and 0.101, 1.522 h−1, respectively. In vitro ct-DNA interaction studies of complexes 1 and 2 revealed electrostatic binding in the groove region of DNA. Variable-temperature magnetic studies provided a J value of approximately −180 cm−1 for 1 and −190 cm−1 for 2, which were supported by DFT-based calculations with various functionals (PBE0, B3LYP, CAM-B3LYP, and ωr2SCAN).
Introduction
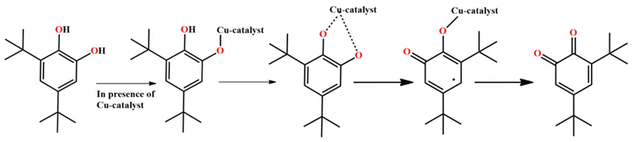
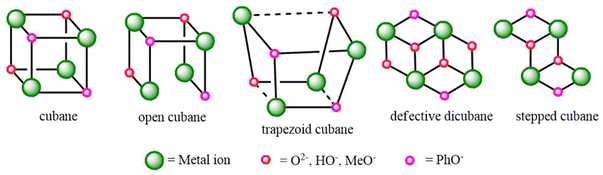
Binuclear complexes that hold two adjacent 3d ions in close proximity have gained significant attention due to their potential to empower several molecular transformations, which are otherwise infeasible, through cooperative metal ion–metal ion interactions.1,2 Many such studies rely on binucleating ligands with a single μ-phenolate group to grab the metal ions.3,4 More recent investigations from a synthetic viewpoint have focused on bimetallic systems based on the design of a new family of binucleating ligands having supporting side arms around the central μ-phenolate group. Synthetic reactivity explorations of such a ligand with copper(II) as a representative 3d ion indicate that [Cu2] fragments can be spontaneously and immediately obtained, which, on further reactions in the presence of available ancillary bridges, can finally result in the Cu4-based products following the self-assembly paths.5 Thus, designing the ligand support and fine-tuning the reaction conditions during reactivity exploration are crucial for understanding and controlling the aggregation processes.Coordination-driven self-assembly applies to a solution progression where several bio-molecular building units, characteristically integrating metal ions as coordination centers, impulsively establish and configure themselves into larger, ordered structures or assemblies via the formation of new, specific and directional coordination bonds. In this process, other weak interactions such as hydrogen bonding, π–π stacking, van der Waals forces, and electrostatic attractions drive the self-assembly. Self-assembly propelled by metal ion coordination enables particular and precise arrangement of biomolecules into functional architectures. The exact coordination between the metal ions and available functional groups, such as amino acids within proteins or bases within nucleic acids, guides the assembly of these molecules into distinctly defined configurations.6 The self-assembled final products based on the initially formed Cu2L fragments can have varying structural topologies.7,8 From an application point of view, these higher-order assemblies could show interesting structural properties, catalytic performances,9,10 biological applications,11 and magnetic properties.12,13 The coupling of two such Cu2L fragments can lead to different types of [Cu4] aggregates, like cubane, open-cubane, defective dicubane, stepped cubane, and trapezoid cubane (Scheme 3). The fine-tuning of the self-assembly path depends upon the reaction conditions, nature and number of in situ generated ancillary bridges. Herein, the implemented solvents and reaction conditions facilitated the coupling of two Cu2L fragments to obtain one open-cubane [Cu4] structure and another stepped-cubane [Cu4] structure. Synthetic [Cu4] complexes with changeable structures are of interest in bioinorganic modeling, multielectron transfer, copper(II)-based industrial and biological catalysis, and magnetostructural research. Water and water-derived linkers, such as HO− and O2− groups, are the most appropriate linkers in the stabilization of [Cu2] moieties, as well as their self-assembled growth, toward higher order congeners.14–16 The aggregate μ4-oxidotetracopper(II) cores, [Cu4(μ4-O)], in copper ion complexes are important in industrial and biological catalysis for transforming substituted phenols to quinones17 and in the environmental nitrogen cycle for the N2O reductase during N2O reduction to N2.18 Within the second domain of N2O reductase, the active site contains a catalytic site known as the CuZ site, which has a [Cu4S] aggregate that binds to N2O for its reduction to N2 in the nitrogen cycle.19,20 During the oxidation of 2,3,6-trimethylphenol (TMP) using O2 to trimethyl quinone (TMQ), which is a key intermediate of vitamin E, in the presence of a catalytic amount of CuCl2 in ionic liquids, a hexa-μ-chlorido-tetrachlorido-μ4-oxidocuprate (4−) species, [Cu4(μ4-O)Cl10]4−, was identified as an active species.21–23
The main biological role of DNA is to store genetic information, and its chemical structure is perfect for metal ion coordination via both the anionic phosphate backbone and neutral nucleobases.24 Newer synthesis techniques and the generation of 3d ion-based complexes capable of cleaving the phosphate ester backbone of DNA under physiological conditions have attracted significant attention due to their potential applications as metallo-drugs, molecular biology tools, gene expression regulators, and also having the ability to cause oxidative damage in nucleobases like 2-deoxyribose.25,26 Copper ion-bearing complexes of organic ligands have revealed improved prospects as anticancer drugs, exhibiting good DNA scission propensity via the oxidative pathway.27,28 Copper ion complexes can scavenge excess intracellular O2˙− to H2O2 and O2 by increasing the H2O2 concentration in the malignant cells and bringing cell death by apoptosis.29 Polyphenol oxidases are a family of Cu2 metalloenzymes that include tyrosinase (TYR) and catechol oxidase (CO).30 CO is a type 3 copper ion-bearing metalloenzyme that catalyzes the aerobic oxidation of o-diphenols to the corresponding o-quinones31,32 (Scheme 1).
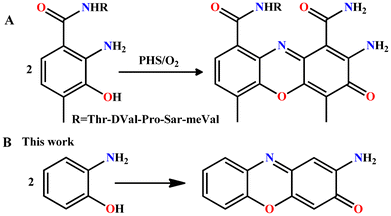
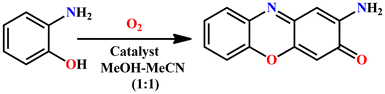
Actinomycin D, a chemotherapy medication, was first isolated from the fermentation of Streptomyces in 1940 and used to treat a number of forms of cancer. Actinomycin D is composed of a central phenoxazinone chromophore fastened to two identical cyclic peptides.33 The formation of phenoxazinone chromophore is dependent on the oligomeric metalloenzyme phenoxazinone synthase (PHS), where each subunit contains a multinuclear copper catalytic center. It catalyzes the oxidative coupling of two o-aminophenol molecules to produce one phenoxazinone chromophore using O2 as an oxidant.34 PHS is responsible for the final step in the biosynthesis of actinomycin D (Scheme 2). The phenoxazinone chromophore is a possible antioxidant and thus has potential applications in drug design.30
This work stems from our ongoing interest in the utilization of simple and semirigid phenol-centred two-metal-ion snatching ligands for the construction of tetranuclear molecular complexes of 3d ions.35 The used ligands with two side arms around the p-cresol backbone facilitate the formation of tetranuclear molecular structures with a wide variety of nuclearities and topologies, using the spontaneous metal ion coordination-induced self-assembly synthetic approaches. Though this synthetic approach lacks predictability, it allows the synthesis of large cores, which could be difficult (or impossible) to achieve using other approaches.
Cu4O4 cubane cores are of three different types based on the degree of structural distortions on six Cu2O2 faces and Cu⋯Cu separations: (i) regular cubane (6 + 0), where all six Cu⋯Cu distances are comparable, (ii) open-cubane type (2 + 4) with two short and four long Cu⋯Cu distances, and (iii) trapezoid cubane type (4 + 2) having four short and two long Cu⋯Cu distances.16,36 In the case of the Cu4O6 type defective dicubane core, there are five Cu2O2 faces, as one of them is shared. These types can therefore be classified into two categories: (i) dicubane core (5 + 0), where all five Cu⋯Cu distances are similar, and (ii) stepped-cubane core (3 + 2), featuring three short and two long Cu⋯Cu distances. Herein, we report one Cu4O4 open cubane core (2 + 4) and one Cu4O6 stepped-cubane core (3 + 2).
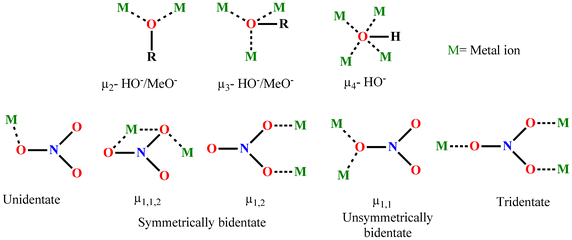
Secondary bridges, like in situ solvent-derived HO− and MeO− ions, along with available NO3− groups, can be employed for the self-assembly follow-up due to their versatile bridging capabilities (Scheme 4). The HO− and MeO− groups can connect two to three copper(II) ions in μ2 and μ3 modes, depending on the available basal or apical positions around copper(II) ions participating in the aggregation process.
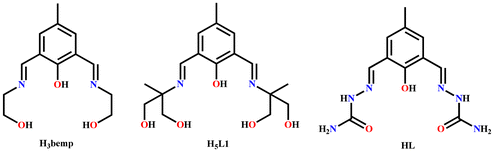
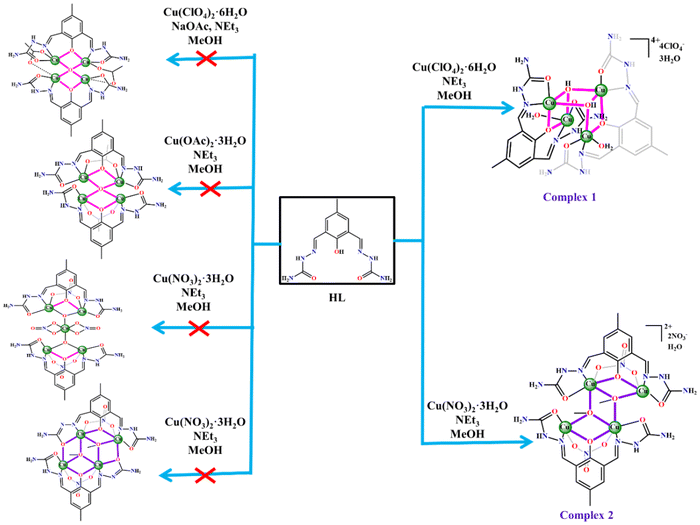
In our efforts to synthesize and investigate the new [Cu4] aggregates through the coordination self-assembly from anionic ligand bound {Cu2} building units, we have explored the reactivity of HL (2,6-bis-{(semicarbazidoimino)}-4-methylphenol) (Scheme 5, right) with Cu(ClO4)2·6H2O and Cu(NO3)2·3H2O in the presence of NEt3. The synergistic aggregating potential of solvent-derived HO− and MeO− groups functions differently during the formation of complexes 1 and 2. It is worth noting that the use of closely related ligands, H3bemp and H5L1 (Scheme 5, left and middle), led to the formation of the systems consisting of [Na2Cu4],37 [Cu18],38 [Cu2], [Cu4] and [Cu5] complexes.2,39
With one-sided semicarbazide arm on the salicyldehyde platform gave Cu1, Cu2 and Cu4 complexes which were known for the anticancer activities.40,41 In the case of ortho vanillin in place of salicylaldehyde, the obtained Cu4 complex exhibited interesting magnetic behaviour.42 In the present reaction settings, we were unable to separate any form of the dinuclear {Cu2} precursor from the reaction medium due to the facile and spontaneous in situ self-aggregation reactions.
In the present work, the principal coordinating capacity of HL, together with the bridging potential of secondary HO−, MeO− and NO3− have been utilized to obtain two new dimer-derived coordination aggregates [Cu4(L)2(H2O)2(μ3-OH)2](ClO4)4·3H2O (1) and [Cu4(L)2(μ-NO3)2(μ3-OMe)2](NO3)2·H2O (2). The crystal structures, magnetic properties and catalytic catechol and 2-aminophenol oxidation behaviors of these complexes are described and discussed with the theoretical supports. Both the synthesized complexes have been examined for their potential as biomimetic models of CO and PHS for their activity, and tentative mechanistic pathways involved during the catalysis have also been presented. From a catalytic point of view, the dinuclear and tetranuclear copper(II) complexes mostly depend on factors like ligand geometry for metal ion capture, steric properties and the basicity of the involved donors, which affect their formation and stabilization and further influence catalytic activity.
Experimental section
Reagents and starting materials
2,6-Diformyl-4-methyl phenol was synthesized by following a literature procedure.43 Semicarbazide hydrochloride was purchased from SRL, India; 3,5-DTBCH2 was purchased from TCL, India; Cu(NO3)2·3H2O and triethylamine (NEt3) were bought from Merck, India. 2-Aminophenol (AP) was purchased from Alfa Aesar, India. Cu(ClO4)2·6H2O was freshly prepared by treating copper hydroxide carbonate (22.12 g, 0.1 mol) with 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 perchloric acid. The chemicals and solvents used in this work were of reagent-grade and used without further purification.
1 perchloric acid. The chemicals and solvents used in this work were of reagent-grade and used without further purification.
Synthesis of 2,6-bis-{(semicarbazidoimino)}-4-methylphenol (HL)
HL was prepared, using a modified synthetic procedure, by the condensation of 2,6-diformyl-4-methylphenol (0.328 g, 2 mmol) and semicarbazide hydrochloride (0.446 g, 4 mmol) in MeOH (25 ml) and sodium acetate (4 mmol, 0.328 g).44 The reaction mixture was then stirred for 3 h at room temperature. The white precipitate of HL thus formed was collected by filtration and dried under a vacuum over CaCl2 and P4O10in vacuo to obtain a white powder (Scheme S1). The formed HL was characterized by ATR-FTIR (1652 cm−1), UV-vis spectra and CHN analysis. Anal. calcd (%) for C11H14N6O3: C, 47.47; H, 5.07; N, 30.20; found: C, 47.48; H, 5.07; N, 30.47.Synthesis of 1 and 2
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 687), 384 (16
687), 384 (16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 920), 674 (326.2).
920), 674 (326.2).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 859), 397 (5856), 606 (344).
859), 397 (5856), 606 (344).
In both cases above, the final aggregation of Cu2L dimers was dependent on (i) the availability of the exogenous HO− or MeO− bridges generated in situ from the reaction medium, and (ii) structure-directing apical site coordination of H2O and NO3− groups.
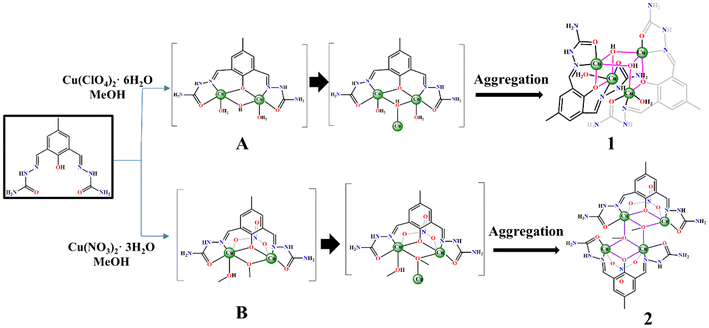
In the former case, the Cu2L(H2O)2 fragments condensed with each other following the deprotonation of one coordinated H2O to provide the Cu4L2(OH)2(H2O)2 skeleton in complex 1. After deprotonation, the Cu2L(OH)2 fragments did not condense into a hitherto unknown alternative structure based on quasi-tetrahedral Cu4L2(O). In the latter situation, the Cu2L(NO3)(MeOH)2 species underwent aggregation after the deprotonation of one coordinated MeOH, like in the previous case, to result in the Cu4L2(MeO)2(NO3)2 frame in complex 2. Here, the aggregation processes took two different paths to generate 1 and 2.
Proposed routes for the tetramer formations
In solution, the coordination reactivity of two different metal ion salts, Cu(ClO4)2·6H2O and Cu(NO3)2·3H2O, toward HL resulted in two different {Cu2} fragments, A and B. These two species are different due to the competitive coordination of H2O vs. MeOH and NO3−. Fragment A has only an intermetallic ancillary HO− bridge, whereas B has MeO− and NO3−. As a result, two types of anion coordination-dependent self-aggregation paths were followed for the final products 1 and 2. For the catalytic solution studies, it was anticipated that the Cu4 complexes 1 and 2 would generate the actual dimeric forms A and B for the interactions with DNA and enzyme mimetics for catecholase oxidase and phenoxazinone synthase (vide infra) (Scheme 7).Physical measurements
ATR-FTIR spectra were recorded using a PerkinElmer Spectrum Two spectrometer. Elemental (C, H, N, S) analysis of the complexes was conducted by using a Thermo Fisher Scientific (Model: FLASHSmart) CHNS Elemental Analyzer equipped with a MAS Plus autosampler and a fully software-controlled system. Powder X-ray diffraction patterns were recorded on a Bruker D2 PHASER X-ray diffractometer (30 kV, 10 mA) with Cu-Kα radiation (λ = 1.5418 Å) in the 2–50° (2θ) range with fixed time counting of 4 s at 25 °C. Electronic spectra were recorded with the Cary 5000 UV-vis spectrophotometer (G9825A, Agilent Technology). TGA and DTA studies were carried out with a PerkinElmer Pyris Diamond TG-DTA instrument. Variable temperature magnetic behaviors of the powder samples were investigated using a Quantum Design MPMS3 magnetometer.Crystal data collection and refinement
A Bruker SMART APEX-V CCD X-ray diffractometer with graphite-monochromated Mo-Kα (λ = 0.71073) radiation gave the diffraction data for complexes 1 and 2. The data were collected with ω scan, where the width was 0.3° per scan and 4 s per frame at 293 K. XPREP and SAINT software were utilized to determine the space group and for data processing.45 The direct method protocol in SHELXS-2014![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 46 was used to solve the crystal structure and full-matrix least squares refinement was done using the SHELXS (2014/7).47 This program package was incorporated in WINGX version 2014.1.48 The SADABS program was applied for the multiscan absorption corrections of the data.49 DIAMOND50 and POV-Ray51 were used to generate the crystallographic illustrations and figures. Crystal structure determination and other parameters are summarized in Table S1. Crystallographic data of the complexes have been deposited at the Cambridge Crystallographic Data Center as supplementary publications CCDC 2514870 and 2514871.
46 was used to solve the crystal structure and full-matrix least squares refinement was done using the SHELXS (2014/7).47 This program package was incorporated in WINGX version 2014.1.48 The SADABS program was applied for the multiscan absorption corrections of the data.49 DIAMOND50 and POV-Ray51 were used to generate the crystallographic illustrations and figures. Crystal structure determination and other parameters are summarized in Table S1. Crystallographic data of the complexes have been deposited at the Cambridge Crystallographic Data Center as supplementary publications CCDC 2514870 and 2514871.
Hirshfeld surface analysis
The intermolecular interactions of compounds 1 and 2 were quantified using Hirshfeld surface analysis. This approach is a graphical tool for the visualization and understanding of intermolecular interactions. Hirshfeld surfaces (HS) and 2D fingerprint (FT) plots were calculated by using the CIFs of 1 and 2 with Crystal Explorer (version 21.5).52 Different colour codes (red–blue–white) shown in HS after running the experiment were plotted with the normalized contact distance (dnorm). The red spots in the graphical designs quantified the shorter contacts of hydrogen bonding interactions, white for van der Waals distances, whereas the blue region specified no clear interactions or longer contacts. 2D FT plots were also generated from the Hirshfeld surfaces to show the relative contributions of the major intermolecular contacts within the complexes.53Computational method
The magnetic isotropic exchange among 3d metal ions in polynuclear complexes is usually estimated by using broken-symmetry DFT calculations by comparing the energies of the high-spin (HS) and broken-symmetry spin states (BS).54 Herein, we have utilized the freely available quantum-chemical package ORCA 6.1.0.55,56 Based on our previous works,57,58 we selected several DFT functionals for such theoretical calculations, namely, hybrid functionals PBE0![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 59 and B3LYP,60 and range-separated hybrid functionals CAM-B3LYP61 and ωr2SCAN.62 The polarized triple-ζ quality basis set def2-TZVPP was used for Cu atoms, def2-TZVP was used for N and O atoms, and the def2-SVP basis set was applied to carbon and hydrogen atoms.63 The calculations utilized the RI approximation with the decontracted auxiliary def2/J Coulomb fitting basis set64 and the chain-of-spheres (RIJCOSX) approximation to exact exchange,65 as implemented in ORCA.
59 and B3LYP,60 and range-separated hybrid functionals CAM-B3LYP61 and ωr2SCAN.62 The polarized triple-ζ quality basis set def2-TZVPP was used for Cu atoms, def2-TZVP was used for N and O atoms, and the def2-SVP basis set was applied to carbon and hydrogen atoms.63 The calculations utilized the RI approximation with the decontracted auxiliary def2/J Coulomb fitting basis set64 and the chain-of-spheres (RIJCOSX) approximation to exact exchange,65 as implemented in ORCA.
Catalytic oxidation of 3,5-DTBCH2 and AP
To evaluate the efficiency of complexes 1 and 2 as catalysts for the oxidation of DTBCH2 and AP, complexes with concentrations of 2 × 10−5 M−1 cm−1 were treated with model substrates (2 × 10−3 M−1 cm−1) in mixed MeOH–MeCN (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) solvent medium under aerobic conditions at room temperature. The entire experiment was done with the Cary 5000 UV-vis spectrophotometer (G9825A, Agilent Technology). The oxidized form of 3,5-ditertbutyl quinone (3,5-DTBQ) is stable and can be detected by its characteristic absorption band at 402 nm (ε402 nm = 1220 M−1 cm−1) in the UV-vis spectral measurement.66 The oxidized product APX in solution displays a specific absorption band at 425 nm (ε425 nm = 2.23 × 104 M−1 cm−1), facilitating effective monitoring using UV–vis absorption spectroscopy.66 The kinetic study was done using the Michaelis–Menten equation, and kinetics were generally used for the interpretation of the model enzyme-like dynamics, and also to evaluate the Kcat value.
1) solvent medium under aerobic conditions at room temperature. The entire experiment was done with the Cary 5000 UV-vis spectrophotometer (G9825A, Agilent Technology). The oxidized form of 3,5-ditertbutyl quinone (3,5-DTBQ) is stable and can be detected by its characteristic absorption band at 402 nm (ε402 nm = 1220 M−1 cm−1) in the UV-vis spectral measurement.66 The oxidized product APX in solution displays a specific absorption band at 425 nm (ε425 nm = 2.23 × 104 M−1 cm−1), facilitating effective monitoring using UV–vis absorption spectroscopy.66 The kinetic study was done using the Michaelis–Menten equation, and kinetics were generally used for the interpretation of the model enzyme-like dynamics, and also to evaluate the Kcat value.
Results and discussion
Synthetic methodology
The bis-ONO Schiff base HL was obtained from the imine condensation reaction of 2,6-diformyl-4-methyl phenol and semicarbazide hydrochloride in the presence of sodium acetate in MeOH medium. The synthesized ligand has two adjacent ONO coordination pockets to bind metal ions (Fig. 1). Earlier, HL was used for the synthesis of Co2, Ni2, and Cu2 complexes.67 | ||
| Fig. 1 (a) Available binding sites of HL and (b) coordination of the adjacent ONO donor sets to two CuII centers. | ||
In this work, the treatment of HL with Cu(ClO4)2·6H2O and NEt3 in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 molar ratio in MeOH under aerobic conditions led to the isolation of deep green block-shaped crystals of complex 1 in 56% yield (eqn (1)). However, the treatment of Cu(NO3)2·3H2O and NEt3 with HL in a 1
1 molar ratio in MeOH under aerobic conditions led to the isolation of deep green block-shaped crystals of complex 1 in 56% yield (eqn (1)). However, the treatment of Cu(NO3)2·3H2O and NEt3 with HL in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 molar ratio at room temperature in MeOH produced complex 2 in 61% yield (eqn (2)).
1 molar ratio at room temperature in MeOH produced complex 2 in 61% yield (eqn (2)).
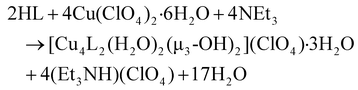 | (1) |
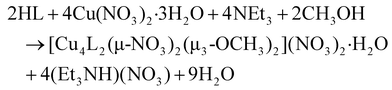 | (2) |
Spectroscopic characterization
![[small nu, Greek, macron]](https://www.rsc.org/images/entities/i_char_e0ce.gif) (C
(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), against the free ligand value at 1648 to 1655 cm−1 for complexes 1 and 2, respectively. The stretching vibration
N), against the free ligand value at 1648 to 1655 cm−1 for complexes 1 and 2, respectively. The stretching vibration ![[small nu, Greek, macron]](https://www.rsc.org/images/entities/i_char_e0ce.gif) (C
(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N appeared at 1662 cm−1 for HL (Fig. S5). The broad bands at 3346, 3298, and 3339 cm−1 were attributed to
N appeared at 1662 cm−1 for HL (Fig. S5). The broad bands at 3346, 3298, and 3339 cm−1 were attributed to ![[small nu, Greek, macron]](https://www.rsc.org/images/entities/i_char_e0ce.gif) O–H of lattice water and bridging hydroxido groups of 1 and 2 (Fig. S5). The presence of four ClO4− ions in 1 was confirmed from the signature peak at 1064 cm−1 for ν3(T2)(νClO), and at 627 cm−1 for the ν4(T2)(δdOClO), and at 476 cm−1 for the ν4 modes.68 In complex 2, the presence of lattice NO3− ions was established by the stretching frequency at 1320 cm−1 for the ν4 asymmetric mode (
O–H of lattice water and bridging hydroxido groups of 1 and 2 (Fig. S5). The presence of four ClO4− ions in 1 was confirmed from the signature peak at 1064 cm−1 for ν3(T2)(νClO), and at 627 cm−1 for the ν4(T2)(δdOClO), and at 476 cm−1 for the ν4 modes.68 In complex 2, the presence of lattice NO3− ions was established by the stretching frequency at 1320 cm−1 for the ν4 asymmetric mode (![[small nu, Greek, macron]](https://www.rsc.org/images/entities/i_char_e0ce.gif) as(NO)), at 1033 cm−1 for the ν1 symmetric mode, at 827 cm−1 for ν2 and at 746 cm−1 for the ν4 (δNO3) bending modes.69
as(NO)), at 1033 cm−1 for the ν1 symmetric mode, at 827 cm−1 for ν2 and at 746 cm−1 for the ν4 (δNO3) bending modes.69
PXRD patterns
The phase purities of complexes 1 and 2 were checked via their powder X-ray diffraction patterns. The experimental and simulated (obtained from SC-XRD data by using Mercury software) patterns were matched (Fig. S6). A comparison of the measured diffraction pattern with one simulated from the crystal structure (solved using single crystal X-ray diffraction) showed a near-perfect match; however, only a whole pattern fit will give a measure of phase purity. At certain 2θ positions, variations in intensity arise due to the difference in the orientation of the crystallites. The overall similarity in the diffraction patterns suggests that the synthesized bulk samples are pure in nature and share the same composition as the single crystals used for structure determination. When used in conjunction with other analytical techniques, such as elemental analysis, this allows the determination of the absolute purity of the materials under study.Electronic spectra
The electronic spectra in the 200 to 800 nm range were recorded in DMF and MeOH![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MeCN (1
MeCN (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) for the ligand and the complexes, respectively. For HL, n → π* and π → π* transitions were observed. Spectra are presented in Fig. S7 and S8 for HL and complexes 1 and 2. The low intensity bands for d–d transitions appeared at 674 nm (ε = 326 L mol−1 cm−1) for 1 and 606 nm (ε = 344 L mol−1 cm−1) for complex 2. The low energy bands at 384 nm (ε = 16
1) for the ligand and the complexes, respectively. For HL, n → π* and π → π* transitions were observed. Spectra are presented in Fig. S7 and S8 for HL and complexes 1 and 2. The low intensity bands for d–d transitions appeared at 674 nm (ε = 326 L mol−1 cm−1) for 1 and 606 nm (ε = 344 L mol−1 cm−1) for complex 2. The low energy bands at 384 nm (ε = 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 920 L mol−1 cm−1) for 1 and 397 nm (ε = 5856 L mol−1 cm−1) for 2, are due to the LMCT transitions.70 The most intense absorption bands at higher energy with maxima at 298 nm (ε = 30
920 L mol−1 cm−1) for 1 and 397 nm (ε = 5856 L mol−1 cm−1) for 2, are due to the LMCT transitions.70 The most intense absorption bands at higher energy with maxima at 298 nm (ε = 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 687 L mol−1 cm−1) and 191 nm (ε = 28
687 L mol−1 cm−1) and 191 nm (ε = 28![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 859 L mol−1 cm−1) for complexes 1 and 2 are due to the metal ion coordinated ligand-based π → π* transitions.
859 L mol−1 cm−1) for complexes 1 and 2 are due to the metal ion coordinated ligand-based π → π* transitions.
Thermal analysis
TGA and combined TGA–DTA plots of the complexes are presented in Fig. S9 and S10. The analysis clearly indicated the dehydration and decomposition phenomena of the complexes in different temperature ranges. The TGA plot of 1 showed a steady weight loss within 110–233 °C for the removal of lattice water molecules and perchlorate anions. For 2, the same occurred within 87–109 °C and 223–243 °C for the loss of lattice water molecules and nitrate anions. Other weight losses at higher temperatures may be attributed to the degradation of organic ligand scaffolds. DTA thermograms showed endothermic peaks at 237 and 255 °C for complexes 1 and 2 due to melting or phase transition.71,72Descriptions of the crystal structures
Due to the strong affinity of CuII centers for apical H2O coordination in two opposite directions, the attachment of two Cu2L units disqualifies the formation of a regular Cu4O4 cube structure. All four CuII centers remained in the square pyramidal geometry as indicated by the Addison parameter (τ) values ranging from 0.0016 to 0.11, where τ5 = (α − β)/γ.73 Values for open faces, i.e., the face bearing Cu1 and Cu4, bound to an apical water molecule, recorded a higher magnitude of τ5 values 0.014 and 0.119, for Jahn–Teller elongation. From the SHAPE measurements,74,75 we found that all the CuII centers remained in the square pyramidal (SPY 5, 1.054 for Cu1; 1.382 for Cu2; 1.916 for Cu3; and 1.117 for Cu4) coordination geometry (Fig. 2c).
![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) space group with Z = 2. The molecular structure of 2 is shown in Fig. 3a and the selected matrix parameters are summarized in Tables S1, S4 and S5. Herein, two solvent MeOH-derived MeO− bridges were responsible for the attachment of two Cu2L units for the Cu4O4 stepped-cubane structure. Within the crystal lattice, the dicationic part [Cu4L2(μ-NO3)2(μ3-OCH3)2]2+ is stabilized by two NO3− counterions and one lattice water molecule. For the growth of the Cu4O4 stepped cubane aggregate, ten Cu–O connections are needed from two L− and supporting ancillaries. The C
space group with Z = 2. The molecular structure of 2 is shown in Fig. 3a and the selected matrix parameters are summarized in Tables S1, S4 and S5. Herein, two solvent MeOH-derived MeO− bridges were responsible for the attachment of two Cu2L units for the Cu4O4 stepped-cubane structure. Within the crystal lattice, the dicationic part [Cu4L2(μ-NO3)2(μ3-OCH3)2]2+ is stabilized by two NO3− counterions and one lattice water molecule. For the growth of the Cu4O4 stepped cubane aggregate, ten Cu–O connections are needed from two L− and supporting ancillaries. The C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O functionalities in the semicarbazone side arms did not tautomerize to bring more charge on the ligand anion as L2− and through this, a chance for bridging two copper(II) centers. Instead, two NO3− groups function as ancillary bridges to adjacent copper ions. Only two solvent-derived MeO− groups were utilized to grow the Cu4O4-stepped cubane structure, possessing dissimilar coordination geometries around the L− bound adjacent CuII centers. One CuII center remained in a square pyramidal NO4 environment (SPY 5, 1.181), while the other fulfilled the octahedral NO5 environment (OC 6, 2.314) from SHAPE analysis74,76 (Fig. 3c). The Cu⋯Cu separation within one Cu2L part is shorter at 2.884 Å compared to the value of 3.255 Å for the Cu2(μ3-OMe)2 core between the two Cu2L parts. Four categories of Cu–O bonds showed a variance within the 1.930 to 2.594 Å range. This clearly establishes the Cu–O bond length order of Cu–OPh > Cu–O(carbonyl) > Cu–OMe > Cu–O(NO3). For Cu–N(imine) bonds, the range is from 1.926 to 1.930 Å. The Addison parameter (τ5) for the pentacoordinated CuII centers is very small (τ5 = 0.001), indicating the perfect square pyramidal geometry. The τ6 value for the CuII center is 0.131, indicating only ∼13% distortion toward trigonal prismatic geometry. This τ6 value was obtained from the use of eqn (3), wherein the angles α, β and γ are three extreme valence angles of the coordination centre.77
O functionalities in the semicarbazone side arms did not tautomerize to bring more charge on the ligand anion as L2− and through this, a chance for bridging two copper(II) centers. Instead, two NO3− groups function as ancillary bridges to adjacent copper ions. Only two solvent-derived MeO− groups were utilized to grow the Cu4O4-stepped cubane structure, possessing dissimilar coordination geometries around the L− bound adjacent CuII centers. One CuII center remained in a square pyramidal NO4 environment (SPY 5, 1.181), while the other fulfilled the octahedral NO5 environment (OC 6, 2.314) from SHAPE analysis74,76 (Fig. 3c). The Cu⋯Cu separation within one Cu2L part is shorter at 2.884 Å compared to the value of 3.255 Å for the Cu2(μ3-OMe)2 core between the two Cu2L parts. Four categories of Cu–O bonds showed a variance within the 1.930 to 2.594 Å range. This clearly establishes the Cu–O bond length order of Cu–OPh > Cu–O(carbonyl) > Cu–OMe > Cu–O(NO3). For Cu–N(imine) bonds, the range is from 1.926 to 1.930 Å. The Addison parameter (τ5) for the pentacoordinated CuII centers is very small (τ5 = 0.001), indicating the perfect square pyramidal geometry. The τ6 value for the CuII center is 0.131, indicating only ∼13% distortion toward trigonal prismatic geometry. This τ6 value was obtained from the use of eqn (3), wherein the angles α, β and γ are three extreme valence angles of the coordination centre.77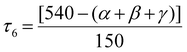 | (3) |
The use of SHAPE analysis, on the other hand, indicated one CuII center type within a square pyramid (SPY 5, 1.181 for Cu1) environment and the second one in an octahedral (OC 6, 2.314 for Cu2) geometry (Fig. 3c).
Hirshfeld surface analysis
The Hirshfeld surface analysis of complexes 1 and 2 are depicted in Fig. S13(a), with dnorm mapped in the range of 0.5 to 1.5 Å. The complexes show different colors for different types of interactions. The bright red area represents the intermolecular H-bonding overall; i.e., the complexes that dominate the structures. The intense red region around the oxygen is due to the strong H-bonding character. The white zone indicates the C⋯H/H⋯H interactions and the blue space represents no noticeable interactions. A noticeable green area can be seen on the curvedness surface maps of the complex, which signifies the flat region above the aromatic ring and the chelate rings that Cu and its ligand (HL) produce, as shown in Fig. S13(b).78 The 2D fingerprint plot region is generated from the Hirshfeld surface, which describes the percentage of various interactions, including O⋯H, N⋯H. The two spikes in the 2D fingerprint plots are for the O⋯H/H⋯O interactions of 48.5% (1) and 36.9% (2) with the overall region of di + de. The interactions of N⋯H/N⋯H are 1% (1) and 4.8% (2) shown in Fig. S13(d).Structural comparison of 1 with 2
In both complexes, each CuII center is bound by the imine N atom of the ligand and bridged by a central phenoxido functionality. The average Cu–N bond lengths in these compounds are in the 1.923–1.931 Å range. Metal ion salt-dependent aggregation reactions were achieved due to the contrasting coordination behaviour of available perchlorate and nitrate ions toward the copper(II) centers. A range of Cu–O bonds were seen, which differed from each other with respect to the coordination geometry and donor-atom environment, e.g., square pyramidal O4N, and octahedral O5N. Six different Cu–O bonds were formed from the coordination of water, semicarbazone oxygen, bridging nitrate, bridging phenoxido, hydroxido and methoxido groups. The longest one in this Cu–O bond group is from the nitrato oxygen atoms and the CuII centers of 2 at 2.594 Å. The oxygen of the water molecule coordinated to the copper(II) centers in 1 register a large Cu–O separation (2.311 and 2.392 Å). The Cu–O bonds from the bridging phenoxido groups of ligand anion are shorter at 1.936 and 1.945 Å compared to the in-plane Cu–O(H) bonds (2.418 Å), which take part in the association of two {Cu2(μ-L)(μ-OH)} fragments. Two μ3-OH connections in 1, connecting three CuII centers, clearly showed two short (1.927 and 1.946 Å) and one long (2.303 Å) Cu–O bonds. The Cu4 entity in complex 1 solely rests on these two connections. The phenolate O atoms only bridge two copper(II) centers at a 1.936 Å distance. The Cu–O–Cu angles within the {Cu2(μ-L)(μ-OH)} fragments in both 1 and 2 are in the 97.41–98.93° range for phenoxido bridges as against 96.27° for hydroxido bridges and methoxido bridges. In 1, the Cu–O(H)–Cu angle is further shortened due to the involvement of the μ3-OH connector required for the growth of the [Cu4] aggregate. In all the complexes, due to the involvement in hydrogen bonding interactions, the H atom of the bridging hydroxido group remains out of plane with respect to the planar Cu2O2 diamond core.Magnetic measurements
Magnetic susceptibility measurements of the powder samples were performed in the temperature range from 300 K to 2 K under an applied magnetic field of 1 kOe. Additionally, at 2 K, the magnetization curves were measured between −50 kOe and +50 kOe. The data presented in the manuscript have been corrected for the temperature-independent contribution of inner shell electrons using Pascal's constants,79 as well as for the magnetic response of the sample holder.Temperature-dependent magnetic susceptibility was measured under two conditions: after cooling the sample in a zero magnetic field (ZFC susceptibility) and after cooling in the presence of a 1 kOe magnetic field (FC susceptibility). In both cases, the measurements were conducted in a magnetic field of 1 kOe (Fig. S11). No significant difference was observed between the zfc and fc susceptibilities, indicating the absence of long-range magnetic ordering in the compounds.
The inset in Fig. S11 displays the magnetic susceptibility on an expanded y-axis scale. Above 150 K, the susceptibilities of both compounds are quite similar. Around 150 K, compound 2 exhibits a local maximum, whereas the susceptibility of compound 1 increases monotonically as the temperature decreases. This local maximum in compound 2 is a clear indication of dominant antiferromagnetic intramolecular interactions. The upturn in susceptibility observed in compound 2 below 50 K is likely due to uncoupled Cu(II) ions or structural defects—such as missing Cu(II) ions or broken bonds between Cu(II) ions in dimers—in the basic magnetic unit, which consists of four Cu(II) ions, as illustrated in Fig. 3b.
To gain deeper insight into the magnetic behavior, Fig. 4 presents a plot of the product χ·T as a function of temperature. At room temperature, the χ·T values are 1.70 emu K mol−1 for compound 1 and 1.54 emu K mol−1 for compound 2. These values are slightly lower than the expected value of 1.81 emu K mol−1 for four Cu(II) ions per molecule, assuming an effective magnetic moment of 1.9μB per Cu(II) ion.79 Nevertheless, the χ·T product still increases with temperature.
 | ||
| Fig. 4 Temperature dependence of the χ·T product of 1 and 2 measured in a magnetic field of 1 kOe. The full green lines are the results of the fit as described in the main text. | ||
As the temperature decreases, the χ·T product of both compounds 1 and 2 decreases, indicating dominant antiferromagnetic interactions not only in compound 2 but also in compound 1. Below approximately 50 K, the χ·T product approaches a nearly constant value. At the lowest measured temperature of 2 K, the χ·T product is approximately 0.6 emu K mol−1 for compound 1 and about 0.1 emu K mol−1 for compound 2. The significantly higher value for compound 1 suggests a greater concentration of structural defects, which contribute to the paramagnetic signal from Cu(II) ions that are not fully incorporated into the antiferromagnetically coupled four-ion magnetic unit. This conclusion is further supported by the M(H) data shown in Fig. S12, where the magnetization of compound 1 at 2 K is roughly seven times greater than that of compound 2.
The temperature-dependent product χ·T(T) was fitted using the freely available software PHI.80 To avoid overparameterization and to remain consistent with the DFT results, the fits were performed using the simplest possible interaction Hamiltonians. For compound 1, we used the Hamiltonian H = −J12(S1·S2) − J34(S3·S4), yielding the parameters J12 = −286 cm−1, J34 = −278 cm−1, g = 2.1. Additionally, 32% of uncoupled Cu(II) centers were required to account for the paramagnetic plateau at 0.6 emu K mol−1 observed in the χ·T(T) plot.
For compound 2, the Hamiltonian H = −J12(S1·S2 + S′1·S′2) − J11′(S1·S′1) was used, resulting in J12 = −188 cm−1, J11′ = 23 cm−1, g = 2.0. In this case, 6% of uncoupled Cu(II) centers were needed to reproduce the paramagnetic plateau at 0.1 emu K mol−1 in the χ·T(T) plot. The results of the fits are shown as green lines in Fig. 4.
DFT calculations
Increased integration grids (defgrid3 in ORCA convention) and tight SCF convergence criteria were used in all calculations.In the case of 1, the following spin Hamiltonian was postulated
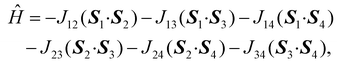 | (4) |
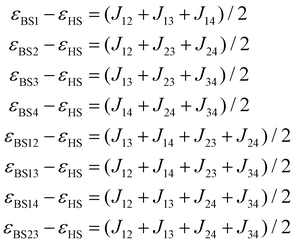 | (5) |
The respective energy differences and relative errors are presented in Table S9. The best agreement was achieved for the parameters listed in Table 1.
| J | PBE0 | CAM-B3LYP | ωr2SCAN | B3LYP |
|---|---|---|---|---|
| J 12 | −319 | −281 | −374 | −455 |
| J 13 | 4.71 | 4.07 | 4.53 | 10.1 |
| J 14 | 2.11 | −3.81 | −1.81 | −10.3 |
| J 23 | −2.20 | −0.672 | −0.756 | −1.86 |
| J 24 | 1.51 | 6.35 | 7.60 | 16.9 |
| J 34 | −162 | −130 | −183 | −245 |
All four PBE0, B3LYP, CAM-B3LYP, and ωr2SCAN functionals predict strong antiferromagnetic exchange between Cu1–Cu2 and Cu3–Cu4 pairs, and weak antiferromagnetic or ferromagnetic exchange between the rest of the Cu(II) ions. These results are in accordance with the experimental magnetic data. The spin density for selected BS-states and DFT functional is depicted in Fig. S14. As expected, the spin density distribution corresponds to 3dx2−y2 magnetic orbitals. The largest overlap between magnetic orbitals is possible only in the planes defined by Cu1–(O)2–Cu2 or Cu3–(O)2–Cu4 atoms, which explains strong AF exchange for these couples.
For the next Cu4-complex 2, the spin Hamiltonian is as follows:
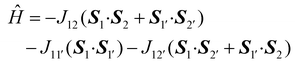 | (6) |
Similarly, several BS-DFT states were computed, and the respective energy differences were related to J-parameters:
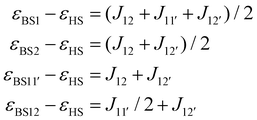 | (7) |
The respective energy differences and relative errors are tabulated in Table S10. The best agreement was achieved for the parameters listed in Table 2.
| J | PBE0 | CAM-B3LYP | ωr2SCAN | B3LYP |
|---|---|---|---|---|
| J 12 | −123 | −94.5 | −139 | −198 |
| J 11′ | 5.52 | −1.23 | 2.19 | 3.33 |
| J 12′ | 6.19 | 5.93 | 8.49 | 11.8 |
All four PBE0, B3LYP, CAM-B3LYP, and ωr2SCAN functionals predict strong antiferromagnetic exchange between Cu1–Cu2 and Cu1′–Cu2′ pairs, and other interactions are either weakly antiferromagnetic or weakly ferromagnetic, which seem to be in agreement with the results of the DC magnetic data. The spin density for selected BS-states and DFT functional is depicted in Fig. S15. As expected, the spin density distribution corresponds to 3dx2−y2 magnetic orbitals. The largest overlap between magnetic orbitals is possible only in the planes defined by Cu1–(O)2–Cu2 or Cu1′–(O)2–Cu2′ atoms, which explains strong AF exchange for these couples.
Further discussion is devoted to searching for the correlation between the molecular structures of 1 and 2, and the strongest AF magnetic exchange parameters, J12 and J34 for 1, J12 for 2. In Table 3, the relevant geometrical parameters are summarized, namely, distances between copper atoms, Cu–O–Cu angles defined by bridging oxygen atoms from L and either OH or OMe ligands, and angles between the planes of the respective chelate rings comprising Cu–O–C–N–N atoms (where O is the terminal atom of L). It is intriguing that J12 is approximately twice as large as J34 in 1, despite having very similar Cu⋯Cu distances and ∠(Cu–O–Cu) angles. The significant difference, however, is observed for the ∠(Cu-planes) angles, with 5.87° for the Cu1–Cu2 planes and 12.29° for the Cu3–Cu4 planes. This means that the overlap between magnetic orbitals should be less efficient in the latter case, which can explain the weaker AF exchange. To support this hypothesis, the decomposition of magnetic exchange coupling82,83 was investigated with the help of ORCA software. This procedure is implemented only for dimers in ORCA, therefore, these calculations were performed for Cu2Zn2 molecular complexes of 1 and 2, where irrelevant Cu atoms were replaced with diamagnetic Zn atoms. The procedure is based on the Yamaguchi formula; thus, the J-values are a bit different from those obtained according to eqn (5) and (7). The decomposition procedure calculated three contributions to J-parameters, JΣ = J0 + ΔJKE + ΔJSP, where J0 is the direct exchange contribution, ferromagnetic in nature, ΔJKE is the antiferromagnetic kinetic exchange contribution, and ΔJSP quantifies the spin polarisation effects, which may be ferro- or antiferromagnetic. This procedure confirmed a larger ΔJKE contribution for the Cu1–Cu2 interaction of 1, as per our presumption. In the case of the Cu1–Cu2 interaction of 2, relevant geometric parameters are similar to the Cu3–Cu4 interaction of 1, and J0 and ΔJKE are quite similar. However, there is a significant increase in ferromagnetic ΔJSP contribution in 2, which may be connected to a different bridging co-ligand (OMe vs. OH) and the presence of an additional nitrato bridging ligand.
| Pair | d(Cu⋯Cu) | ∠(Cu–O–Cu) | ∠(Cu-planes) | J 0 | ΔJKE | ΔJSP | J Σ |
|---|---|---|---|---|---|---|---|
| Cu1–Cu2 (1) | 2.916 | 97.27, 98.89 | 5.87 | 217 | −498 | 20.7 | −260 |
| Cu3–Cu4 (1) | 2.910 | 97.16, 97.41 | 12.29 | 203 | −353 | 45.8 | −105 |
| Cu1–Cu2 (2) | 2.884 | 96.29, 96.28 | 10.37 | 233 | −355 | 73.0 | −48.7 |
DNA binding activity
Absorption spectral studies
To determine the binding propensity of complexes 1 and 2, absorption titration experiments with ct-DNA were performed. Titration curves of complexes with increasing ct-DNA concentrations in aqueous Tris-buffer revealed a decrease in the absorption spectrum at 315 nm in complex 1 and 319 nm in complex 2 (Fig. 5). Such spectral changes imply an intercalative binding mode of the complexes.84 To further quantify the extent of binding, the Wolfe–Shimer equation85 was used to calculate Kb values, and the values are presented in Table 4. | ||
| Fig. 5 Absorption spectra of complexes 1 and 2 in the presence of increasing concentrations of ct-DNA (0.2–1.4 × 10−4 M−1). | ||
| Complexes | K b (M−1) | K sv (M−1) |
|---|---|---|
| 1 | 2.72 × 104 | 3.89 × 105 |
| 2 | 1.43 × 104 | 3.19 × 105 |
Ethidium bromide (EB) displacement assay
Ethidium bromide or EB, a commonly used intercalating agent, exhibits the characteristic property of intercalation by forming soluble complexes with nucleic acids. This leads to the emission of highly intense fluorescence, which can be attributed to the intercalation of the planar phenanthridine ring between adjacent base pairs on the double helix structure of ct-DNA. The alterations observed in the spectra of EB upon its binding to ct-DNA are frequently employed to investigate the interactions between DNA and metal complexes.86 Due to strong interactions with DNA, EB emits fluorescence at 565 nm, which is quenched on the addition of other molecules, causing the replacement of bound EB molecules as the number of binding sites decreases.87 Upon adding complexes 1 and 2 to the EB–ct-DNA system in increasing aliquots of 200 μL, a notable reduction in fluorescence was observed. This reduction indicates that a competitive interaction between EB and the complexes for binding to ct-DNA takes place. To quantitatively assess the strength of the interaction, the quenching efficiency was determined using the Stern–Volmer equation. The calculated Stern–Volmer constants (Ksv values) for complexes 1 and 2 were evaluated (Table 4). These values followed the order of 1 > 2 in terms of quenching efficiency (Fig. 6).Circular dichroism (CD) spectroscopy is a powerful technique for investigating conformational changes in DNA induced by interactions with small molecules or drug candidates. DNA exhibits two characteristic CD signals: a positive band at approximately 275 nm, attributed to base stacking interactions, and a negative band near 245 nm, associated with the right-handed B-form helicity of the DNA double helix. Both bands are highly sensitive to structural perturbations arising from DNA–drug interactions.88 Drug molecules that engage in groove binding or electrostatic interactions typically induce minor spectral changes, while intercalative binding causes pronounced alterations in the CD spectra due to significant conformational disruption. In the present study, upon incubation of calf thymus DNA (ct-DNA) with complexes 1 and 2 at a molar ratio of [complex]/[DNA] = 1, notable reductions in the intensities of both the positive and negative CD bands were observed compared to unbound DNA (Fig. 7). This reduction indicates a perturbation of base pair stacking and a disruption of the native helical conformation of the DNA, likely arising from an interaction mode involving intercalation.
Taken together, the consistent observations from various complementary optical methods viz., UV-vis, fluorescence and, CD spectroscopy exhibited a ‘hypochromic’ effect in absorption bands, effective EB displacement with high binding and quenching constants, and pronounced CD spectral perturbations, which corroborate well, suggesting a strong intercalative binding mode of complexes 1 and 2via π–π stacking interactions that ‘wedge’ in between the nucleobases of the DNA double helix. The stronger binding affinity of complex 1 as compared to complex 2 is suggestive of preferential binding selectivity and the higher therapeutic potency of complex 1 that could be attributed to the different coordination environments in the complexes, viz., complex 1 possesses an intermetallic ancillary HO− bridge vs. the MeO skeleton in complex 2.
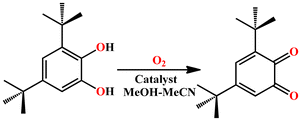
Catechol oxidation activity of complexes
To estimate the catecholase activity of complexes 1 and 2, 3,5-DTBCH2 was used as a model substrate. The moderate reaction potential and the steric hindrance of the bulky tert-butyl group of 3,5-ditertbutylcatechol (3,5-DTBCH2) help to prevent unwanted oxidation (Scheme 8). As a result, it is an excellent substrate for evaluating catechol oxidase by ligand anion-bound metal ion fragments in the catalysts.Electronic spectra of the complexes 1 and 2 in the presence of 3,5-DTBCH2 in a methanol–acetonitrile (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) mixed solvent medium show catecholase activity (Fig. 8), where 3,5-DTBCH2 is used as a model substrate for catecholase activity. For the experiment, 1 × 10–3 M complex 1 and 2 in MeOH
1) mixed solvent medium show catecholase activity (Fig. 8), where 3,5-DTBCH2 is used as a model substrate for catecholase activity. For the experiment, 1 × 10–3 M complex 1 and 2 in MeOH![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MeCN solution and 1 × 10–1 M 3,5-DTBCH2 were prepared. For the catecholase study, 20 μL 10–3 (M) complex and 20 μL 10–1 (M) 3,5-DTBCH2 were taken in 960 μL mixed solvent, and the experiment was carried out for up to 1 h at room temperature. In pure MeOH
MeCN solution and 1 × 10–1 M 3,5-DTBCH2 were prepared. For the catecholase study, 20 μL 10–3 (M) complex and 20 μL 10–1 (M) 3,5-DTBCH2 were taken in 960 μL mixed solvent, and the experiment was carried out for up to 1 h at room temperature. In pure MeOH![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MeCN (1
MeCN (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) mixed solvent medium, complexes 1 and 2 showed an absorption band at 394 nm. When the 3,5-DTBCH2 was added to complexes 1 and 2, intense peaks appeared at 404 nm and 402 nm, characteristic of 3,5-DTBQ. The spectra remain unchanged after 1 h of mixing.
1) mixed solvent medium, complexes 1 and 2 showed an absorption band at 394 nm. When the 3,5-DTBCH2 was added to complexes 1 and 2, intense peaks appeared at 404 nm and 402 nm, characteristic of 3,5-DTBQ. The spectra remain unchanged after 1 h of mixing.
2-Aminophenol oxidation activity of complexes
The oxidative capabilities of complexes 1 and 2 were subsequently assessed through their efficacy in catalyzing the transformation of 2-aminophenol (AP) into 2-amino-3H-phenoxazin-3-one (APX) (Scheme 9), which serves as a model reaction system for evaluating phenoxazinone synthase-mimicking activity.For this catalytic experiment, 2 × 10−5 M−1 cm−1 of catalyst (complexes 1 and 2) were taken with a 100 times concentration of the model substrate (AP) (2 × 10−3 M−1 cm−1) in the same 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 MeOH–MeCN mixed solvent medium at room temperature. The reaction progress was monitored over 1 h by obtaining consecutive UV–vis spectra within the range of 300–500 nm at 5 min intervals (Fig. 9). During this process, AP undergoes oxidative dimerization to yield APX, utilizing molecular oxygen from the atmosphere as the oxidant in the catalytic presence of the complexes.
1 MeOH–MeCN mixed solvent medium at room temperature. The reaction progress was monitored over 1 h by obtaining consecutive UV–vis spectra within the range of 300–500 nm at 5 min intervals (Fig. 9). During this process, AP undergoes oxidative dimerization to yield APX, utilizing molecular oxygen from the atmosphere as the oxidant in the catalytic presence of the complexes.
Kinetics study
Kinetic measurements of catecholase activity and phenoxazinone synthase activity were performed for both complexes 1 and 2, using the model substrates 3,5-DTBCH2 and AP in MeOH–MeCN mixed solvent medium at room temperature at the respective wavelengths89 (Fig. 10 and 11). This was carried out by taking a constant concentration (2 × 10−5 mol L−1) of the catalyst (complex 1 and 2) while varying the model substrate (3,5-DTBCH2 and AP) concentration from 20 to 100 times (4 × 10−4 mol L−1 to 2 × 10−3 mol L−1) and the development of the oxidized products (3,5-DTBQ and APX) was monitored using the UV-vis spectrophotometer.2 The Michaelis–Menten equation and kinetics are generally used for the interpretation of the model enzyme-like dynamics. The model helps to describe how the dinuclear CuII fragments, mimicking the enzymes, enhance the kinetic rates. The Kcat, Vmax values of the respective experiments for both complexes are summarized in Table 5. We have also calculated the TON and TOF values for the respective complexes for both catalytic activities (shown in Table 5). A comparison of the Kcat values with other reported Cu(II) complexes is shown in Table S11. | ||
Fig. 10 Concentration dependence of the reaction rates in catechol oxidation by complexes 1 and 2 in MeOH–MeCN (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) solvent medium; the Lineweaver–Burk plots are in the insets. 1) solvent medium; the Lineweaver–Burk plots are in the insets. | ||
 | ||
Fig. 11 Concentration dependence of the reaction rates in 2-aminophenol oxidation by complexes 1 and 2 in MeOH–MeCN (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) solvent medium; the Lineweaver–Burk plots are in the insets. 1) solvent medium; the Lineweaver–Burk plots are in the insets. | ||
| Model substrates | Complexes | K cat (h−1) | V max (mol L−1 s−1) | K cat/KM (M−1 s−1) | TON | TOF (min−1) |
|---|---|---|---|---|---|---|
| 3,5-DTBCH2 | 1 | 12.08 | 6.711 × 10−8 | 2.289 | 8.2 | 0.13 |
| 3,5-DTBCH2 | 2 | 1.845 | 1.025 × 10−8 | 0.35 | 5.65 | 0.09 |
| AP | 1 | 0.101 | 5.64 × 10−10 | 0.0198 | 3.28 | 0.054 |
| AP | 2 | 1.522 | 8.46 × 10−9 | 0.362 | 3.05 | 0.05 |
Conclusions
The contributions of perchlorate and nitrate salts of copper(II) have significantly influenced the coordination geometries around two entities. The dimer–dimer aggregations took place with {Cu2L} fragments, where one or more copper(II) ions showed apical coordination from H2O and MeOH molecules. Diverse types of Jahn–Teller distortions of square-pyramidal and octahedral CuII centers were accomplished through the formation of bridges from hydroxide and methoxido O donors. The ligand HL in its phenoxido form directed the formation of two types of Cu4 aggregates. Bridging coordination from the nitrate anion facilitated a different course of attachment of two {Cu2L} fragments in obtaining a stepped-cubane Cu4 structure. The linking of two such {Cu2L} units was achieved by reaction medium solvent-derived methoxido bridges for interdimeric clipping. The non-availability of such an ancillary bridge allowed solvent water coordination and the hydroxido bridge-based fixing of two {Cu2L} units. For both complexes 1 and 2, only three Cu2O2 faces were generated, involving four copper(II) ions during interdimer aggregation. The 3D Hirshfeld surface analysis, together with the 2D fingerprint plots, showed intermolecular interactions in the crystal packing, which were maintained by the distinct H⋯H, O⋯H and C⋯H contacts. This sort of interdimeric fixing provided three Cu⋯Cu magnetic interactions in these faces but with different spatial dispositions. Variable temperature magnetic studies provided a single exchange constant J with values of approximately −180 cm−1 for 1 and −190 cm−1 for 2, originating from strong antiferromagnetic interactions between the four intertwined copper(II) centers in different spatial arrangements. The utilization of biophysical methods indicated in vitro interactions with complexes 1 and 2 with ct-DNA, showing a strong intercalative mode (via π–π stacking interactions) of binding in the groove region of DNA, and it was observed that complex 1 has a stronger binding affinity than complex 2. Complexes 1 and 2 have shown enzyme (catechol oxidase)-simulated solution catechol oxidation behavior toward the chosen model substrate, 3,5-DTBCH2, in a mixed MeOH–MeCN (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) reaction medium. From the Michaelis–Menten enzyme kinetic study, it was observed that the Kcat values varied from 1.854 h−1 to 12.08 h−1 for complexes 1 and 2, respectively. Complexes 1 and 2 also showed solution catalytic activity toward AP for 2-aminophenoxazine-3-one (APX) (phenoxazinone synthase mimicking activity) conversion in mixed MeOH–MeCN (1
1) reaction medium. From the Michaelis–Menten enzyme kinetic study, it was observed that the Kcat values varied from 1.854 h−1 to 12.08 h−1 for complexes 1 and 2, respectively. Complexes 1 and 2 also showed solution catalytic activity toward AP for 2-aminophenoxazine-3-one (APX) (phenoxazinone synthase mimicking activity) conversion in mixed MeOH–MeCN (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) solvent. The Michaelis–Menten kinetic studies provided Kcat values of 0.101 h−1 and 1.522 h−1 for complexes 1 and 2, respectively. The kinetic studies indicated that complex 1 is catalytically superior to complex 2 for catechol oxidation, but the opposite was observed for 2-aminophenol oxidation.
1) solvent. The Michaelis–Menten kinetic studies provided Kcat values of 0.101 h−1 and 1.522 h−1 for complexes 1 and 2, respectively. The kinetic studies indicated that complex 1 is catalytically superior to complex 2 for catechol oxidation, but the opposite was observed for 2-aminophenol oxidation.
Conflicts of interest
The authors have no conflicts of interest.Data availability
The authors declare that the data are available on request.The data supporting this article have been included as part of the SI. Crystallographic data for the complexes 1 and 2 have been deposited in CCDC. Supplementary Information: Scheme for ligand synthesis, Summary of the crystal data and refinement of 1 and 2. Table of selected bond lengths and bond angles of 1-2, SHAPE measurement values, IR spectra, UV-vis spectra, PXRD patterns, TGA plots, Packing diagrams, additional data for catalytic oxidation, magnetic characterization and theoretical calculations. Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5dt03005f.
CCDC 2514870 and 2514871 contain the supplementary crystallographic data for this paper.90a,b
Acknowledgements
AM is thankful to IIT Kharagpur for the research facility and her research fellowship. ZJ acknowledges the financial support from the Slovenian Research Agency (research core funding No. P2-0348). RH acknowledges the financial support from the institutional sources of the Department of Inorganic Chemistry, Palacký University Olomouc, Czech Republic. FA acknowledges the Department of Chemistry, AMU for providing the necessary research facility.References
- B. Dutta, T. Guizouarn, K. Kotrle, F. Pointillart, R. Herchel and D. Ray, Cryst. Growth Des., 2025, 25, 3915–3927 CrossRef CAS.
- A. Manna, Z. Jagličić and D. Ray, Chem. – Asian J., 2026, e70497, DOI:10.1002/asia.70497.
- P. K. Hota, A. Jose, S. Panda, E. M. Dunietz, A. E. Herzog, L. Wojcik, N. Le Poul, C. Belle, E. I. Solomon and K. D. Karlin, J. Am. Chem. Soc., 2024, 146, 13066–13082 CrossRef CAS PubMed.
- Y. Kadoya, M. Hata, Y. Tanaka, A. Hirohata, Y. Hitomi and M. Kodera, Inorg. Chem., 2021, 60, 5474–5482 Search PubMed.
- M. Das, A. B. Canaj, V. Bertolasi, M. Murrie and D. Ray, Dalton Trans., 2018, 47, 17160–17176 RSC.
- Y. Su, Y. Liu, Y. Zhong, S. Ma, Q. Ouyang, X. Chen, Y. Li, E. Xiong, Z. Zou and J. Liu, Coord. Chem. Rev., 2025, 527, 216403 Search PubMed.
- J. P. Karnes, N. M. Lind, A. G. Oliver, C. S. Day, V. W. Day and J. D. Blakemore, Inorg. Chem., 2025, 64, 571–593 Search PubMed.
- T. S. Mahapatra, D. Basak, S. Chand, M. Shatruk, V. Bertolasi and D. Ray, Dalton Trans., 2016, 45, 13576–13589 RSC.
- H. Huang, X. Jing, J. Deng, C. Meng and C. Duan, J. Am. Chem. Soc., 2023, 145, 2170–2182 CrossRef CAS PubMed.
- K. C. Gupta and A. K. Sutar, Coord. Chem. Rev., 2008, 252, 1420–1450 CrossRef CAS.
- K. Tummalapalli, C. S. Vasavi, P. Munusami, M. Pathak and M. M. Balamurali, Appl. Organomet. Chem., 2017, 31, 1–10, DOI:10.1002/aoc.3680.
- K. Iman, M. K. Raza, M. Ansari, Monika, A. Ansari, M. Ahmad, M. N. Ahamad, K. M. A. Qasem, S. Hussain, M. N. Akhtar and M. Shahid, Dalton Trans., 2021, 50, 11941–11953 RSC.
- J. Das, S. Ta and D. Das, New J. Chem., 2023, 47, 18126–18129 RSC.
- M. Bera, W. T. Wong, G. Aromi, J. Ribas and D. Ray, Inorg. Chem., 2004, 43, 4787–4789 CrossRef CAS PubMed.
- K. Dhara, J. Ratha, M. Manassero, X. Y. Wang, S. Gao and P. Banerjee, J. Inorg. Biochem., 2007, 101, 95–103 CrossRef CAS PubMed.
- N. C. Jana, A. Escuer, P. Brandão and A. Panja, Dalton Trans., 2025, 54, 3000–3012 RSC.
- H. Sun, K. Harms and J. Sundermeyer, J. Am. Chem. Soc., 2004, 126, 9550–9551 Search PubMed.
- M. Prudêncio, A. S. Pereira, P. Tavares, S. Besson, I. Cabrito, K. Brown, B. Samyn, B. Devreese, J. Van Beeumen, F. Rusnak, G. Fauque, J. J. G. Moura, M. Tegoni, C. Cambillau and I. Moura, Biochemistry, 2000, 39, 3899–3907 Search PubMed.
- C. Hsu, S. C. Rathnayaka, S. M. Islam, S. N. MacMillan and N. P. Mankad, Angew. Chem., 2020, 132, 637–641 Search PubMed.
- B. J. Johnson, S. V. Lindeman and N. P. Mankad, Inorg. Chem., 2014, 53, 10611–10619 Search PubMed.
- Z. Ke, A. Abtahi, J. Hwang, K. Chen, J. Chaudhary, I. Song, K. Perera, L. You, K. N. Baustert, K. R. Graham and J. Mei, J. Am. Chem. Soc., 2023, 145, 3706–3715 Search PubMed.
- J. Dupont, B. C. Leal, P. Lozano, A. L. Monteiro, P. Migowski and J. D. Scholten, Chem. Rev., 2024, 124, 5227–5420 CrossRef CAS PubMed.
- H. Sun, K. Harms, J. Sundermeyer, F. Chemie and D. P. V. Marburg, J. Am. Chem. Soc., 2004, 126, 9550–9551 CrossRef CAS PubMed.
- W. Zhou, R. Saran and J. Liu, Chem. Rev., 2017, 117, 8272–8325 Search PubMed.
- M. Pitié, C. J. Burrows and B. Meunier, Nucleic Acids Res., 2000, 28, 4856–4864 CrossRef PubMed.
- F. Mancin, P. Scrimin, P. Tecilla and U. Tonellato, Chem. Commun., 2005, 2540–2548 RSC.
- D. Díaz-garcía, P. R. Ardiles, M. Díaz-sánchez, I. Mena-palomo, S. Prashar, A. Rodríguez-diéguez and P. L. Páez, J. Inorg. Biochem., 2020, 203, 110912 CrossRef PubMed.
- D. Díaz-García, P. R. Ardiles, S. Prashar, A. Rodríguez-Diéguez, P. L. Páez and S. Gómez-Ruiz, Pharmaceutics, 2019, 11, 30, DOI:10.3390/pharmaceutics11010030.
- R. N. Patel, K. K. Shukla, A. Singh, M. Choudhary, U. K. Chauhan and S. Dwivedi, Inorg. Chim. Acta, 2009, 362, 4891–4898 Search PubMed.
- S. K. Dey and A. Mukherjee, Coord. Chem. Rev., 2016, 310, 80–115 Search PubMed.
- S. Araji, T. A. Grammer, R. Gertzen, S. D. Anderson, M. Mikulic-Petkovsek, R. Veberic, M. L. Phu, A. Solar, C. A. Leslie, A. M. Dandekar and M. A. Escobar, Plant Physiol., 2014, 164, 1191–1203 Search PubMed.
- E. I. Solomon, U. M. Sundaram and T. E. Machonkin, Chem. Rev., 1996, 96, 2563–2605 Search PubMed.
- L. J. Sun, H. Yuan, J. K. Xu, J. Luo, J. J. Lang, G. B. Wen, X. Tan and Y. W. Lin, Biochemistry, 2023, 62, 369–377 CrossRef CAS PubMed.
- M. Le Roes-Hill, C. Goodwin and S. Burton, Trends Biotechnol., 2009, 27, 248–258 CrossRef CAS PubMed.
- M. Das, G. A. Craig, D. Escudero, M. Murrie, A. Frontera and D. Ray, New J. Chem., 2018, 42, 14349–14364 RSC.
- J. Tercero, E. Ruiz, S. Alvarez, A. Rodríguez-Fortea and P. Alemany, J. Mater. Chem., 2006, 16, 2729–2735 RSC.
- D. Mandal, M. Chauhan, F. Arjmand, G. Aromí and D. Ray, Dalton Trans., 2009, 9183–9191 Search PubMed.
- T. Shiga, K. Maruyama, L. Han and H. Oshio, Chem. Lett., 2005, 34, 1648–1649 CrossRef CAS.
- N. Ahmed, S. Tripathi, A. Sarkar, K. U. Ansari, C. Das, N. Prajesh, S. Horike, R. Boomishankar and M. Shanmugam, New J. Chem., 2020, 44, 16845–16855 RSC.
- Z. Afrasiabi, E. Sinn, S. Padhye, S. Dutta, S. Padhye, C. Newton, C. E. Anson and A. K. Powell, J. Inorg. Biochem., 2003, 95, 306–314 CrossRef CAS PubMed.
- N. M. H. Salem, A. R. Rashad, L. El Sayed, S. Foro, W. Haase and M. F. Iskander, Inorg. Chim. Acta, 2015, 432, 231–242 CrossRef CAS.
- M. Das, B. K. Shaw, B. N. Ghosh, K. Rissanen, S. K. Saha and S. Chattopadhyay, RSC Adv., 2015, 5, 46869–46872 RSC.
- R. R. Gagné, C. L. Spiro, C. A. Hamann, T. J. Smith, W. R. Thies and A. K. Shiemke, J. Am. Chem. Soc., 1981, 103, 4073–4081 CrossRef.
- S. M. Annigeri, M. P. Sathisha and V. K. Revankar, J. Coord. Chem., 2008, 61, 4011–4024 CrossRef CAS.
- SAINT, SMART and XPREP, Siemens Analytical X-ray Instruments Inc., Madison, WI, 1995 Search PubMed.
- G. M. Sheldrick, SHELXS-2014, Program for Crystal Structure Solution, University of Göttingen, Göttingen, Germany, 2014 Search PubMed.
- G. M. Sheldrick, Acta Crystallogr., Sect. A: Found. Crystallogr., 2008, 64, 112–122 CrossRef CAS PubMed.
- L. J. Farrugia, J. Appl. Crystallogr., 2012, 45, 849–854 CrossRef CAS.
- G. M. Sheldrick, SADABS: Software for Empirical Absorption Correction, University of Göttingen, Institute fur Anorganische Chemieder Universitat, Göttingen, Germany, 1999–2003 Search PubMed.
- W. T. Pennington, J. Appl. Crystallogr., 1999, 32, 1028–1029 CrossRef CAS.
- L. J. Farrugia, POV-Ray - 3.5, University of Glasgow, Glasgow,U.K., 2003 Search PubMed.
- P. R. Spackman, M. J. Turner, J. J. McKinnon, S. K. Wolff, D. J. Grimwood, D. Jayatilaka and M. A. Spackman, J. Appl. Crystallogr., 2021, 54, 1006–1011 CrossRef CAS PubMed.
- M. A. Spackman and D. Jayatilaka, CrystEngComm, 2009, 11, 19–32 RSC.
- F. Neese, Coord. Chem. Rev., 2009, 253, 526–563 CrossRef CAS.
- F. Neese, Faraday Discuss., 2024, 254, 295–314 RSC.
- F. Neese, Software Update: The ORCA Program System—Version 6.0, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2025, 15, e70019, DOI:10.1002/wcms.70019.
- E. M. Manohar, S. Roy, X. Li, S. Tothadi, J. Mok, J. Tang, R. Herchel, J. Lee, A. Dey and S. Das, Dalton Trans., 2024, 53, 10499–10510 RSC.
- B. Rybníčková, J. Kuchár, P. Antal and R. Herchel, Inorg. Chim. Acta, 2020, 509, 119689, DOI:10.1016/j.ica.2020.119689.
- C. Adamo and V. Barone, J. Chem. Phys., 1999, 110, 6158–6170 CrossRef CAS.
- P. J. Stephens, F. J. Devlin, C. F. Chabalowski and M. J. Frisch, J. Phys. Chem., 1994, 98, 11623–11627 CrossRef CAS.
- T. Yanai, D. P. Tew and N. C. Handy, Chem. Phys. Lett., 2004, 393, 51–57 CrossRef CAS.
- L. Wittmann, H. Neugebauer, S. Grimme and M. Bursch, J. Chem. Phys., 2023, 159, 224103, DOI:10.1063/5.0174988.
- F. Weigend, R. Ahlrichs and F. K. Gmbh, Phys. Chem. Chem. Phys., 2005, 7, 3297–3305 RSC.
- F. Weigend, Phys. Chem. Chem. Phys., 2006, 8, 1057–1065, 10.1039/B515623H.
- F. Neese, F. Wennmohs, A. Hansen and U. Becker, Chem. Phys., 2009, 356, 98–109 CrossRef CAS.
- P. Mahapatra, S. Ghosh, S. Giri, V. Rane, R. Kadam, M. G. B. Drew and A. Ghosh, Inorg. Chem., 2017, 56, 5105–5121 CrossRef CAS PubMed.
- M. H. Inoue, R. R. Ribeiro, J. R. Sabino and F. S. Nunes, Spectrochim. Acta, Part A, 2016, 164, 76–83 CrossRef CAS PubMed.
- R. Gil-García, G. Madariaga, A. Jiménez-Pérez, I. Herrán-Torres, A. Gago-González, M. Ugalde, V. Januskaitis, J. Barrera-García, M. Insausti, M. S. Galletero, J. Borrás, J. V. Cuevas, R. Pedrido, P. Gómez-Saiz, L. Lezama and J. García-Tojal, CrystEngComm, 2023, 25, 2213–2226 RSC.
- M. Y. Mihaylov, V. R. Zdravkova, E. Z. Ivanova, H. A. Aleksandrov, P. S. Petkov, G. N. Vayssilov and K. I. Hadjiivanov, J. Catal., 2021, 394, 245–258 CrossRef CAS.
- M. Das, S. R. Chaudhuri, D. Basak, S. Dasgupta and D. Ray, Inorg. Chim. Acta, 2019, 485, 140–154 CrossRef CAS.
- J. Guo, J. Zhang, P. Shu, L. Kong, X. Hao, Y. Xue, Z. Luo, Y. Li, G. Li, G. Yao and Y. Zhang, Molecules, 2012, 17, 6424–6433 CrossRef CAS PubMed.
- U. Ibotomba Singh, R. K. Bhubon Singh, W. Radhapiyari Devi and C. Brajakisor Singh, J. Chem. Pharm. Res., 2012, 4, 1130–1135 CAS.
- A. G. Blackman, E. B. Schenk, R. E. Jelley, E. H. Krenske and L. R. Gahan, Dalton Trans., 2020, 49, 14798–14806 RSC.
- J. Cirera, E. Ruiz and S. Alvarez, Organometallics, 2005, 24, 1556–1562 CrossRef CAS.
- I. D. Brown and K. K. Wu, Acta Crystallogr., Sect. B, 1976, 32, 1957–1959 CrossRef.
- S. Alvarez, P. Alemany, D. Casanova, J. Cirera, M. Llunell and D. Avnir, Coord. Chem. Rev., 2005, 249, 1693–1708 CrossRef CAS.
- K. Kotrle, I. Nemec, P. Antal, K. Petrželová, E. Čižmár and R. Herchel, Inorg. Chem. Front., 2023, 10, 7319–7332 RSC.
- S. Sundareswaran and S. Karuppannan, Cryst. Res. Technol., 2020, 55, 1–8 Search PubMed.
- N. W. Ashcroft and N. D. Mermin, Solid State Physics, Saunders College Publishing, USA, 1976 Search PubMed.
- N. F. Chilton, R. P. Anderson, L. D. Turner, A. Soncini and K. S. Murray, J. Comput. Chem., 2013, 34, 1164–1175 CrossRef CAS PubMed.
- A. Bencini and F. Totti, J. Chem. Theory Comput., 2009, 5, 144–154 CrossRef CAS PubMed.
- E. Coulaud, J.-P. Malrieu, N. Guihéry and N. Ferré, J. Chem. Theory Comput., 2013, 9, 3429–3436 CrossRef CAS PubMed.
- G. David, G. Trinquier and J. P. Malrieu, J. Chem. Phys., 2020, 153, 194107 CrossRef CAS PubMed.
- A. Erxleben, Coord. Chem. Rev., 2018, 360, 92–121 Search PubMed.
- H. Y. Khan, M. F. Ansari, S. Tabassum and F. Arjmand, Drug Discovery Today, 2024, 29, 104055 CrossRef CAS PubMed.
- S. Fountoulaki, F. Perdih, I. Turel, D. P. Kessissoglou and G. Psomas, J. Inorg. Biochem., 2011, 105, 1645–1655 CrossRef CAS PubMed.
- J. B. LePecq and C. Paoletti, J. Mol. Biol., 1967, 27(1), 87–106 Search PubMed.
- F. Bisceglie, S. Pinelli, R. Alinovi, M. Goldoni, A. Mutti, A. Camerini, L. Piola, P. Tarasconi and G. Pelosi, J. Inorg. Biochem., 2014, 140, 111–125 Search PubMed.
- K. Chattopadhyay, B. Dutta, S. Dutta, A. Manna, G. A. Craig, M. Mandal, M. Murrie and D. Ray, Inorg. Chim. Acta, 2024, 572, 122270 CrossRef CAS.
- (a) CCDC 2514870: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2qdxvx; (b) CCDC 2514871: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2qdxwy.
| This journal is © The Royal Society of Chemistry 2026 |