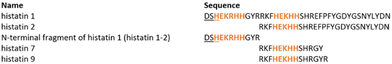From speciation to action: Cu(II) and Zn(II) tune histatins, but pH and enamel drive efficacy
Emilia
Dzień
a,
Aleksandra
Mikołajczyk-Tarnawa
b,
Agnieszka
Matera-Witkiewicz
*b,
Krzysztof
Szewczyk
c,
Miquel
Barceló-Oliver
 d,
Lilla
Pawlik-Sobecka
e,
Joanna
Wątły
d,
Lilla
Pawlik-Sobecka
e,
Joanna
Wątły
 a and
Magdalena
Rowińska-Żyrek
a and
Magdalena
Rowińska-Żyrek
 *a
*a
aFaculty of Chemistry, University of Wrocław, F. Joliot-Curie 14, 50-383 Wrocław, Poland. E-mail: magdalena.rowinska-zyrek@uwr.edu.pl
bScreening of Biological Activity Assays and Collection of Biological Material Laboratory, Faculty of Pharmacy, Wroclaw Medical University, Borowska 211A, 50-556 Wrocław, Poland. E-mail: agnieszka.matera-witkiewicz@umw.edu.pl
cDepartment of Oncology, Wrocław Medical University, pl. L. Hirszfelda 12, 53-413 Wrocław, Poland
dDepartment of Chemistry, University of Balearic Islands, Cra. de Valldemossa, km 7.5., 07122 Palma de Mallorca, Spain
eDepartment of Basic Medical Sciences, Faculty of Farmacy, Wroclaw Medical University, Borowska 211, Wroclaw, Poland
First published on 18th November 2025
Abstract
Histatins are histidine-rich salivary peptides whose antimicrobial activity emerges from a delicate interplay between proteolytic cleavage and metal coordination. We quantified Cu(II) and Zn(II) binding to histatin 1 and its hydrolytic products (histatin 1-2 and histatin 2), as well as to histatin 7 and histatin 9, and related thermodynamic and spectroscopic properties to in vitro activity. Histatins form stable metal complexes, with Cu(II) binding occurring primarily via the ATCUN motif in histatin 1 and histatin 1-2, and with Zn(II) coordination following the –HEXXH– motif. In contrast to simple electrostatic expectations, adding terminal Arg residues neither measurably stabilizes Zn(II) complexes nor enhances bactericidal activity. Across ATCC pathogens tested, activities remain modest and largely decoupled from complex stability, with only isolated effects upon metallation. Overall, two main conclusions may be drawn: (i) proteolysis mainly reshapes peptide topology and surface contacts rather than activating a metal-dependent mechanism and (ii) environmental pH together with anchoring to hydroxyapatite are likely the main drivers of efficacy in situ. We propose a working model in which site-selective hydrolysis positions histatins at the enamel–biofilm interface, while Cu(II)/Zn(II) binding acts as a structural governor rather than a direct antimicrobial switch. This reframes design rules for histatin-like therapeutics: optimize localization and pH-gated charge distribution first and then treat metallation as a context-dependent modulator.
Introduction
Saliva is a key element of the innate immune system in humans and animals, as it contains antimicrobial peptides (AMPs). Among salivary AMPs, the following can be distinguished: lysozyme,1 defensins (α- and β-defensins),2 cystatins,3 mucins,4 statherin,5 and histatins.6 Histatins are a family of cationic, histidine-rich peptides varying in size between 7 and 38 amino acids.6 The concentration of histatins in saliva extends from 17 to 33 μg mL−1,7 of which about 80% are histatin 1, histatin 3 and histatin 5. Castagnola et al. observed that the highest concentration of histatins in saliva occurs around 5:00 and 6:00 p.m.8 Johnson et al., in turn, indicate a decrease in histatin concentration with age.9 Histatins, as natural peptides, are sensitive to proteolytic cleavage; therefore, another factor influencing the concentration of histatins is the presence of proteolytic enzymes in saliva.10,11 This is important due to differences in the antimicrobial activity of peptides from the histatin family. Of all histatins, histatin 5, which is one of the main proteolytic cleavage products of histatin 3,12 demonstrates the greatest antifungal activity, especially against Candida albicans.13 Recent work by Franz and co-workers has shown that well-defined Cu(II)-binding sites in histatin 5 can directly couple copper coordination to antifungal activity against Candida albicans.14 The antimicrobial activity of histatins has also been demonstrated against ESKAPE pathogens.15Histatins are mainly produced and released by the serous acini of the parotid, submandibular, sublingual and von Ebner glands.16,17 Expression of histatin 1, as the only one of the histatin family, has also been found in human lacrimal glands, melanoma cell lines, metastatic lesions of melanoma patients18,19 and head and neck squamous cell carcinoma (HNSCC) tumors.20 The presence of histatin 1 in cancer tissues suggests the potential of using the peptide as a tumor marker.19 Moreover, a recent study by Jenwanichkul et al. indicates the potential use of histatin 1 as an anticancer drug. Due to its cationic nature, it binds to the cancer cell membrane, leading to its disruption. Additionally, it has been noted that histatin 1 sensitizes cancer cells to the commonly used drug cisplatin, which enhances its cytotoxicity.21 In addition to its potential anticancer activity, histatin 1 also exhibits antifungal activity against C. albicans (strain B311 and ATCC 44505), although lower than that of histatin 3 and histatin 5.22 In turn, in conjunction with statherin 1, it has an antibacterial effect by reducing the adhesion of the oral bacteria Streptococcus mutans to the hydroxyapatite surface,23 which may limit the development of caries.24,25 It is possible due to its full phosphorylation on serine 2, which allows histatin 1 to effectively bind to hydroxyapatite crystals,26 influencing the acquisition of the enamel pellicle27 and facilitating the remineralization of enamel lesions.28 Moreover, as shown by McDonald et al., the binding of histatin 1 to the hydroxyapatite surface protects it from the action of proteolytic enzymes.29 In addition to its antimicrobial activity and role in maintaining tooth enamel homeostasis described above, histatin 1 promotes wound healing in the oral mucosa, skin and epithelial cells, in vitro and in vivo, by stimulating epithelial migration.30–32
Previously, we discussed the coordination chemistry and microbiological activity of histatin 3 and 5 and their hydrolysis products, i.e. histatin 3-4, histatin 4, histatin 5-8 and histatin 8 in complexes with Cu(II) and Zn(II) ions, showing that: (i) Cu(II) binds to the albumin-like binding motif, if available (in histatin 3-4, histatin 4, histatin 5-8 and histatin 8); (ii) Zn(II) most typically coordinates to either the carboxylate side chain of glutamic acid and two histidine imidazole groups or two histidyl residues, with polymorphic binding sites, where zinc is most likely bound to two different sets of {2Nim}, with two of such forms being in equilibrium; (iii) the antimicrobial efficacy of histatins and their complexes is highly pH-dependent, with Zn(II) coordination often enhancing their activity; and (iv) histatin 3-4 and histatin 5-8 undergo structural changes, adopting a more α-helical conformation after binding Zn(II).33,34
This study demonstrates the relationship between the coordination mode, thermodynamic properties and the antimicrobial effectiveness of (i) genetically encoded histatin 16,35 and products of its autoproteolysis: its N-terminal fragment (here called histatin 1-2) and histatin 2 and (ii) cleavage products of histatin 3 – histatin 7 and histatin 9 – in this case, the influence of non-binding arginine residues on the metal complex's thermodynamic stability is discussed (Fig. 1).
Experimental
Materials
All peptides, histatin 1 (DSHEKRHHGYRRKFHEKHHSHREFPFYGDYGSNYLYDN), the N-terminal fragment of histatin 1 (here called histatin 1-2, DSHEKRHHGYR), histatin 2 (RKFHEKHHSHREFPFYGDYGSNYLYDN), histatin 7 (RKFHEKHHSHRGY) and histatin 9 (RKFHEKHHSHRGYR) were purchased from KareBay Biochem (USA) (certified purity of 98%) and were used as received. The purity was checked potentiometrically. Cu(ClO4)2·6H2O and Zn(ClO4)2·6H2O were extra pure products (Sigma-Aldrich) and the concentrations of stock solutions of these salts were determined by inductively coupled plasma optical emission spectroscopy (ICP-OES). The carbonate-free stock solution of 0.1 M NaOH was purchased from Sigma-Aldrich and then potentiometrically standardized with the primary standard potassium hydrogen phthalate (KHP, 99.9% purity). The ionic strength (I) was adjusted to 0.1 M by addition of NaClO4 (Sigma-Aldrich). Peptides were soluble in water solutions.Mass spectrometric measurements
High-resolution mass spectra were obtained on a Shimadzu LCMS-9030 qTOF (Shimadzu, Kyoto, Japan) device, equipped with a standard ESI source and a Nexera X2 system, and a Bruker Apex Ultra FT-ICR mass spectrometer (Bruker Daltonik, Bremen, Germany), equipped with an Apollo II electrospray ionization source with an ion funnel. The Shimadzu LCMS-9030 qTOF was operated in positive and negative ion modes. The instrumental parameters were as follows: scan range: m/z 100–2000, nebulizing gas: nitrogen, nebulizing gas flow: 3.0 L·min−1, drying gas flow: 10 L·min−1, heating gas flow: 10 L·min−1, interface temperature: 300 °C, desolvation line temperature: 400 °C, detector voltage: 2.02 kV, interface voltage: 4.0 kV, collision gas: argon, mobile phases: (A) H2O + 0.1% HCOOH and (B) MeCN + 0.1% HCOOH, and mobile phase total flow: 0.3 mL min−1. The Bruker Apex Ultra FT-ICR mass spectrometer was operated in positive ion mode. The instrumental parameters were as follows: scan range: m/z 100–2000, dry gas: nitrogen, temperature: 473 K, and ion energy: 5 eV. The capillary voltage was optimized to the highest S/N ratio, and it was 4200 V. The injection volume was optimized depending on the intensity of the signals observed in the mass spectrum within the range of 0.1 to 3 μL. The samples were prepared in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 methanol–water mixture at a pH value of 7.4. The sample concentration was [ligand]tot = 10−4 M and the M
1 methanol–water mixture at a pH value of 7.4. The sample concentration was [ligand]tot = 10−4 M and the M![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) L molar ratio was 1
L molar ratio was 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. The obtained data were processed using ACDLabs Spectrus Processor v2021.1.3 and Bruker Compass DataAnalysis 4.0 programs.
1. The obtained data were processed using ACDLabs Spectrus Processor v2021.1.3 and Bruker Compass DataAnalysis 4.0 programs.
Potentiometric measurements
Proton–ligand and metal–ligand stability constants were calculated from titration curves carried out over the pH range of 2–11 at 298 K under an argon atmosphere and an ionic strength of 0.1 M (NaClO4). The total volume of the solution used was 2.8 cm3. The potentiometric titrations were performed using a Metrohm Titrando 905 titrator and a Mettler Toledo InLab Micro combination pH electrode. The thermostabilized glass cell was equipped with a magnetic stirring system and a microburette delivery tube. The solutions were titrated with 0.1 M carbonate-free NaOH. The electrodes were daily calibrated for hydrogen ion concentration by titrating 4 mM HClO4 with NaOH under the same experimental conditions as mentioned above. The purities and the exact concentrations of the ligand solutions were determined by the Gran method.36 The ligand concentration was 0.5 mM and the ratio of Cu(II) and Zn(II) to the ligand was 0.8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1.
1.
The standard potential and the slope of the electrode couple were computed by means of the GLEE program.37 HYPERQUAD 2006 was used for the stability constant calculations.38 The standard deviations were computed by HYPERQUAD 2006 and refer to random errors only. The constants for the hydrolytic Zn(II) species were used in these calculations.39 The speciation and competition diagrams were computed with the HYSS program.40
Spectroscopic studies
The UV-Vis absorption spectra were recorded on a Jasco-730 spectrophotometer and circular dichroism (CD) measurements were obtained on a Chirascan VX spectropolarimeter at 25 °C. For both methods, spectra were collected over the 200–800 nm range using quartz cuvettes with 1 cm optical path length. To determine the secondary structure of the analyzed samples, measurements were made in the 180–250 nm range, using a quartz cuvette with an optical path of 0.01 cm. The concentrations of the solutions used for spectroscopic studies were similar to those employed in the potentiometric experiments. The metal![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) peptide ratio was 0.8
peptide ratio was 0.8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. All spectroscopic measurements were recorded in the pH range of 3–11. The pH of the samples was adjusted with the appropriate amounts of HClO4 and NaOH solutions. The UV-Vis and CD spectroscopy parameters were calculated from the spectra obtained at the pH values corresponding to the maximum concentration of each particular species, based on distribution diagrams. OriginPro 8 was used to process and visualize the obtained spectra.
1. All spectroscopic measurements were recorded in the pH range of 3–11. The pH of the samples was adjusted with the appropriate amounts of HClO4 and NaOH solutions. The UV-Vis and CD spectroscopy parameters were calculated from the spectra obtained at the pH values corresponding to the maximum concentration of each particular species, based on distribution diagrams. OriginPro 8 was used to process and visualize the obtained spectra.
Antimicrobial activity assay
Seven reference strains from ATCC (Enterococcus faecalis ATCC 29212, MRSA Staphylococcus aureus ATCC 43300, Klebsiella pneumoniae ATCC 700603, Acinetobacter baumannii ATCC 19606, Pseudomonas aeruginosa ATCC 27853, Escherichia coli ATCC 25922 and Candida albicans ATCC 10231) were used for antimicrobial activity assays. The MIC50, as the lowest concentration of the tested compound at which 50% growth of the isolates was inhibited, was determined by a microdilution method with spectrophotometric measurements according to ISO 20776-1:201941 and ISO 16256:2012,42 whereas a modified Richard's method using the redox indicator (2,3,5-triphenyltetrazolium chloride, TTC) was used to determine the minimal bactericidal/fungicidal concentration (MBC/MFC).43–45 Serial dilutions of the peptides in Tryptic Soy Broth (TSB), with or without an equimolar concentration of Cu(II) or Zn(II), were made on 96-well microplates in the range between 0.5 and 512 µg mL−1. Tryptone Soya Agar (TSA) plates were inoculated with microbial strains from preformed stocks. After 24 h/37 °C incubation (for bacteria) or 24 h/25 °C (for fungi), a proper density of bacterial and fungal suspension was prepared using a densitometer (final inoculum 5 × 105 CFU mL−1) in TSB. Positive (TSB + strain) and negative controls (only TSB) were also included in the test. Spectrophotometric solubility control of each peptide and the peptide–metal ion system was performed. To avoid using strains that had developed additional resistance, the following antibacterial/antifungal agents were used as positive controls, according to breakpoint values established by the EUCAST:46,47E. faecalis: 4 µg mL−1 levofloxacin, S. aureus: 1 µg mL−1 levofloxacin, K. pneumoniae: 4 µg mL−1 gentamicin, A. baumannii: 0.5 µg mL−1 levofloxacin, P. aeruginosa: 1 µg mL−1 levofloxacin, E. coli: 4 µg mL−1 gentamicin and C. albicans: 1 µg mL−1 amphotericin B. MIC was determined from a series of at least three experiments as the lowest concentration of an antimicrobial agent that decreased the measured microbial growth to 50% relative to the positive control.Microplates were incubated at 37 ± 1 °C or 25 ± 1 °C for 24 h on a shaker (500 rpm). After this, the spectrophotometric measurement was performed at 580 nm and then 50 μL aliquots of 1% (m/v) 2,3,5-triphenyltetrazolium chloride (TTC) solution were added to each well. TTC is a commonly used indicator of cellular respiration. In oxidized form, TTC is colorless, while it turns pink after reduction due to reactions in the respiratory chain. MBC/MFC (Minimum Bactericidal/Fungicidal Concentration) can be defined as the lowest concentration required to kill a particular microbial strain, determined by visual analysis after 24 h incubation with TTC. The pink color indicates the presence of living microorganisms, while the lack of color indicates that the colonies do not survive. Thanks to both methods, MIC and MBC or MFC can be determined.
Neutral red (NR) uptake assay
The experiment was conducted according to the established protocol.48 The cells were seeded into 96-well plates at a concentration of 1 × 104 cells per well. After reaching 80% confluency (usually after 24 h), the cells were treated with dedicated compounds with/without metal ions (concentrations to assess cytotoxicity for normal cells (L929)). Additionally, the following concentrations were used: 150, 100, 50, and 10 µM. Negative controls with medium only and positive controls with 1 µM staurosporine were also included. The cells were incubated for 24, 48 and 72 hours prior to the experiment. After the exposure, the medium was replaced with 40 µg mL−1 NR in the appropriate culture medium and the cells were further incubated for up to 3 hours. The cells were subsequently washed with phosphate-buffered saline (PBS) and the dye was extracted with NR destain solution (1% acetic acid, 50% ethanol, 49% deionized water). To form a homogeneous solution, plates with cells were shaken for 30 minutes before absorbance reading at 540 nm. Untreated cells (negative control) represented 100% potential cellular growth.Results and discussion
To investigate the precise stoichiometry and structural and thermodynamic properties of the complexes of histatin 1, histatin 1-2, histatin 2, histatin 7 and histatin 9 with Cu(II)/Zn(II) ions, a set of experimental methods were used: mass spectrometry (ESI-MS), potentiometric titrations, and UV-Vis and CD spectroscopy.The mass spectra confirmed that the tested peptides, under the experimental conditions here employed, are capable of forming Cu(II) and Zn(II) complexes in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (metal
1 (metal![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ligand ratio) stoichiometry. Signals and isotopic distributions in the experimental and simulated spectra for the chosen signals are consistent and confirm the correct interpretation (Fig. S1A–J).
ligand ratio) stoichiometry. Signals and isotopic distributions in the experimental and simulated spectra for the chosen signals are consistent and confirm the correct interpretation (Fig. S1A–J).
Protonation constants
Based on a series of potentiometric titrations, nineteen deprotonating constants (pKa) were established for histatin 1, eight deprotonating constants (pKa) for the N-terminal fragment of histatin 1 (histatin 1-2), thirteen deprotonating constants (pKa) for histatin 2 and nine deprotonating constants (pKa) for histatin 7 and histatin 9, respectively (Table S1). The determined values for the deprotonation of histidine residues (with pKa values in the range of 5.32–8.00 for histatin 1, 5.76–7.33 for histatin 1-2, 5.64–7.28 for histatin 2, 5.56–7.26 for histatin 7 and 5.57–7.22 for histatin 9) are in agreement with those found in the literature for similar poly-His systems.49–52Cu(II)complexes
Histatin 2, just as histatin 4 (RKFHEKHHSHRGYRSNYLYDN), lacks the ATCUN motif. Its first 11 N-terminal amino acid residues are in the same sequence as histatin 4 (NH2–RKFHEKHHSHR11–). Based on the NMR spectrum of the Cu(II)-histatin 4 complex, which confirmed that the coordination {Nim, 3N−} mode involved His4 in the coordination sphere,34 we may conclude that the coordination mode is analogous in this case.
Stabilities and spectroscopic data for Cu(II)-histatin 9 are analogous to those of Cu(II)-histatin 7, which is discussed above as a representative example.
Zn(II) complexes
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β of 53.68, reaches its maximum at pH 6.1 (Fig. S8D) and most likely, the carboxylate side chain of Glu and two His imidazole nitrogens from the typical Zn(II)-binding motif, –HEXXH–, are involved in Zn(II) coordination. The next two forms, ZnH4L and ZnH3L, with pKa values of 6.50 and 7.63, respectively, probably come from the deprotonation of two non-binding His imidazoles. Subsequently, the ZnH2L (pKa = 8.70) form is the most abundant at around pH 9.2. In this species, the deprotonation of a bound water molecule is observed. The last two forms, ZnHL and ZnL, with pKa values of 9.78 and 10.54, respectively, arise from the deprotonation of the non-bonding N-terminal amine group and Tyr residue. The Zn(II)-histatin 7 complex results in a {2Nim, O−} binding mode.
β of 53.68, reaches its maximum at pH 6.1 (Fig. S8D) and most likely, the carboxylate side chain of Glu and two His imidazole nitrogens from the typical Zn(II)-binding motif, –HEXXH–, are involved in Zn(II) coordination. The next two forms, ZnH4L and ZnH3L, with pKa values of 6.50 and 7.63, respectively, probably come from the deprotonation of two non-binding His imidazoles. Subsequently, the ZnH2L (pKa = 8.70) form is the most abundant at around pH 9.2. In this species, the deprotonation of a bound water molecule is observed. The last two forms, ZnHL and ZnL, with pKa values of 9.78 and 10.54, respectively, arise from the deprotonation of the non-bonding N-terminal amine group and Tyr residue. The Zn(II)-histatin 7 complex results in a {2Nim, O−} binding mode.
In the case of the Zn(II)-histatin 9 complex, overall stabilities are very similar (Table S2 and Fig. S8E), so the coordination mode is analogous to the Zn(II)-histatin 7 one, with one extra deprotonation of a Lys residue (pKa = 10.81).
Antimicrobial activity and cytotoxicity
Antimicrobial susceptibility testing was performed using an expanded panel of microorganisms: two Gram-positive (E. faecalis ATCC 29212 and MRSA S. aureus ATCC 43300), four Gram-negative (K. pneumoniae ATCC 700603, A. baumannii ATCC 19606, P. aeruginosa ATCC 27853, and E. coli ATCC 25922) and one fungal strain (C. albicans ATCC 10231). The results showed no MBC/MFC activity. MIC50 values were determined for all active compounds and their metal complexes (Table 1). Moreover, for all ligands, ligand–metal ion systems, and regular L929 cells, viability (NR assay) was over 90%. Thus it can be clearly assumed that the investigated histatin and histatin–metal ion connections are safe in the analysed concentration range.| Strain | S. aureus (+) ATCC 43300 | E. faecalis (+) ATCC 29212 | P. aeruginosa (−) ATCC 27853 | K. pneumoniae (−) ATCC 700603 | C. albicans ATCC 10231 |
|---|---|---|---|---|---|
| MIC (µg mL−1) | MIC (µg mL−1) | MIC (µg mL−1) | MIC (µg mL−1) | MIC (µg mL−1) | |
| Histatin 1 | n/d | n/d | 128 | 128 | 256 |
| Cu(II)-histatin 1 | n/d | n/d | n/d | 128 | 256 |
| Zn(II)-histatin 1 | 256 | n/d | n/d | 256 | 256 |
| Histatin 1-2 | n/d | n/d | n/d | n/d | 256 |
| Cu(II)-histatin 1-2 | n/d | n/d | n/d | n/d | n/d |
| Zn(II)-histatin 1-2 | n/d | n/d | n/d | n/d | 256 |
| Histatin 2 | n/d | n/d | n/d | n/d | n/d |
| Cu(II)-histatin 2 | n/d | n/d | n/d | n/d | 256 |
| Zn(II)-histatin 2 | n/d | n/d | n/d | n/d | n/d |
| Histatin 7 | 256 | 256 | n/d | n/d | n/d |
| Cu(II)-histatin 7 | n/d | 256 | n/d | n/d | n/d |
| Zn(II)-histatin 7 | n/d | 256 | n/d | n/d | n/d |
| histatin 9 | n/d | n/d | n/d | n/d | n/d |
| Cu(II)-histatin 9 | n/d | n/d | n/d | n/d | n/d |
| Zn(II)-histatin 9 | n/d | n/d | n/d | n/d | n/d |
The only two active compounds against the Gram-positive bacterium S. aureus were the complex of histatin 1 with Zn(II) ions (MIC = 256 µg mL−1) and histatin 7 (MIC = 256 µg mL−1). Native histatin 7 is also active against E. faecalis (MIC = 256 µg mL−1). After binding both Cu(II) and Zn(II) ions, the biological effect against E. faecalis did not change compared to the native peptide (MIC = 256 µg mL−1). Unlike the previously analysed histatins, at physiological pH, histatin 1 is moderately active against Gram-negative bacteria P. aeruginosa (MIC = 128 µg mL−1) and K. pneumoniae (MIC = 128 µg mL−1). In the case of K. pneumoniae, a microbiological effect is also observed for the copper(II) and zinc(II) complexes of histatin 1 (MIC = 128 µg mL−1 and 256 µg mL−1, respectively), but the binding of the metal ion by this peptide does not enhance its antimicrobial properties.
Under the studied conditions, MICs for histatin 1 (128–256 µg mL−1) exceed the typical whole-saliva histatin levels (with a median for histatin 1 ≈ 17 µg mL−1; for total histatins ≈ 33 ± 17 µg mL−1);7 however, local histatin 1 levels at the enamel–biofilm interface are expected to be considerably higher due to specific binding to hydroxyapatite and protection from proteolysis, which renders our activities compatible with a biologically relevant, interfacial mechanism of action rather than bulk-phase killing.
Histatin activity does not depend simply on Cu(II)/Zn(II) affinity or complex stoichiometry. Proteolytic trimming changes peptide topology and redistributes positive charge, which modulates binding to hydroxyapatite and the pellicle and the electrostatic interaction with bacterial surfaces. Coordination of Cu(II) at the ATCUN site and of Zn(II) at the –HEXXH– motif mainly restricts backbone flexibility and does not uniformly increase activity; the secondary structure remains largely unchanged after metal binding and MIC values do not consistently decrease. In vivo, three factors appear critical: (i) localization at the enamel or pellicle, which concentrates the peptide and limits proteolysis; (ii) pH-dependent protonation, which enhances surface attraction; and (iii) metal availability, which is lowered by competing salivary ligands. Within this framework, metal coordination fine-tunes conformation and speciation but is not the principal determinant of the antimicrobial effect. This is consistent with the lack of benefit from adding terminal arginine residues and with the higher activity of native histatin 7 over its Arg-extended variant against selected Gram-positive strains.
A comparison of metal binding abilities
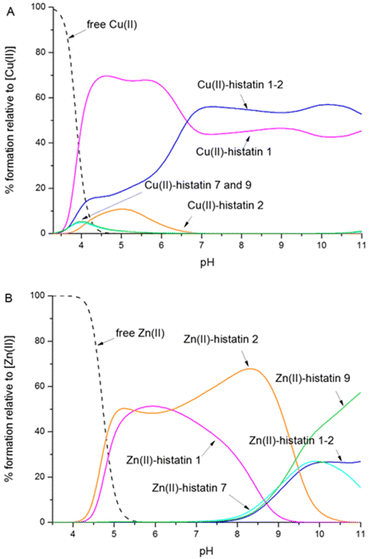
Histatins form thermodynamically stable complexes with Cu(II) and Zn(II) (Table S2). As expected, histatin 1 and its N-terminal fragment (histatin 1-2), in which the ATCUN motif is present, bind Cu(II) more effectively compared to histatins 2, 7 and 9, in which the copper ions are coordinated by the histidine imidazole and three amide nitrogens (Fig. 2A). Histatin 1 is a more effective Cu(II) binder, most likely because of the longer C-terminal part (as compared to histatin 2), which is able to sterically protect the metal binding site by (i) partial wrapping of the ATCUN site, (ii) providing additional electrostatic and hydrogen-bonding interactions, and (iii) favouring a conformation that shields the Cu(II) site from solvent.Across the Zn(II) series, coordination involves two His imidazoles and a Glu carboxylate within the –HEXXH motif; sequence context matters, yet the longest peptides do not uniformly outperform shorter ones – histatins 1 and 2 show the greatest affinity towards Zn(II) in almost the entire tested pH range. This is probably related to the presence of the –FPF– sequence near the metal ion binding site. Proline, due to its unique cyclic structure that restricts rotation around the peptide bond, limits the flexibility of the peptide, resulting in local rigidity of the peptide chain.53 In turn, the proximity of two phenylalanine residues may influence the formation of hydrophobic interactions or π–π stacking interactions, stabilizing the local structure of the peptide.54 It is worth noting that the HEKHH motif, present in histatins 1 and 2, is a much more tempting site for Zn(II) than the HEKRHH site available in histatin 1-2. This finding highlights the importance of the non-binding residues in the typical –HEXXH– Zn(II) binding motif.
Due to the similarity of the amino acid sequence of histatin 7, histatin 8 and histatin 9, it was intriguing to compare the stability of the complexes of these peptides with Cu(II) and Zn(II). The case of Cu(II) complexes is what may have been expected – the Cu(II)-histatin 8 complex, in which the metal ion is coordinated by the N-terminus, the imidazole nitrogen of histidine in position 3 and two amide nitrogens in between (the ATCUN motif), is the most stable complex in the entire pH range. In histatins 7 and 9, the imidazole nitrogen of His 4 is the anchoring site for Cu(II), showing comparable stability (Fig. 3A).
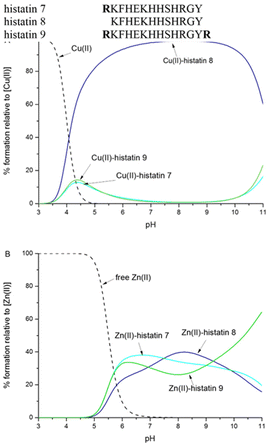 | ||
| Fig. 3 Competition plots between histatin 7, histatin 8 and histatin 9 binding (A) Cu(II) and (B) Zn(II). The plots describe the complex formation at different pH values in a hypothetical situation, in which equimolar amounts of reagents are mixed. Calculations are based on binding constants from Table S2 and ref. 33. Conditions: T = 298 K, [Cu(II)/Zn(II)] = [histatin 7] = [histatin 8] = [histatin 9] = 0.001 M. | ||
Quite a surprising issue is observed for zinc(II) complexes (Fig. 3B). Our aim was to observe and explain differences in complex stabilities due to the presence or absence of C-/N-terminal Arg residues – arginines carry a significant positive charge and could theoretically have a stabilizing effect on the complexes by forming hydrogen bonds between the positively charged NH2 group of the Arg side chain and the backbone carbonyl oxygens55 or other electrostatic effects and less strong interactions with the solvent molecules that are coordinated.56
The results are quite far from what was expected – there are barely any differences that can be commented on. Up to pH 9.3, the almost same thermodynamic stability is related to the same coordination mode {2Nim, O−}. Above pH 10, the Zn(II)-histatin 9 complex becomes somewhat more stable, with the difference not being significantly higher than the measurement error. The stabilities of Zn(II) complexes of histatin 7, in which the Arg residue is present only at the N-terminus, and histatin 9, without Arg residues at the N- and C-termini, are almost identical. We may conclude that surprisingly, the increase of complex stability by a simple electrostatic interaction is not the case in the studied histatin ligands.
None of the histatins studied in this work showed a tendency to change its secondary structure after binding Cu(II) or Zn(II) ions (as confirmed by near-ultraviolet circular dichroism spectroscopy (data not shown)). Thus, the secondary structure does not affect the antimicrobial activity, which is consistent with the results obtained for the other peptides from the histatin family, especially for the complexes of the N-terminal fragment of histatin 3 (histatin 3-4) and the N-terminal fragment of histatin 5 (histatin 5-8), which, although after binding Zn(II), tend to adopt an α-helical structure, and yet the activity of the complexes is not higher than that of the native peptides.33,34 Also, Cu(II)/Zn(II) binding does not significantly enhance the antimicrobial activity of the histatins studied in this work and no relationship was observed between the stability of the complexes and their antimicrobial activity – the mechanism of their action is not based on the process of nutritional immunity (AMPs bind metal ions so that microbes cannot get enough metals essential for their life and virulence).
What then influences the antimicrobial activity of histatins and their complexes and the mechanism of their action? Undoubtedly, the pH of the environment has an impact on biological activity. This is related to the cationic nature of histatins – a positively charged peptide interacts more easily with the negatively charged membrane of microorganisms. Previous studies have shown that the anticandidal activity of histatins increases at mildly acidic pH,22,33 similar to that found in cariogenic plaque, which can reach pH values as low as 5.5. In this sense, the reported MIC values at pH 7.40 should be regarded as conservative estimates of activity under more acidic, clinically relevant conditions.
The effective binding of histatin 1 to the surface of hydroxyapatite is another significant factor that protects it from proteolytic enzyme activity. In addition, specific protease inhibitors present in saliva inhibit protease activity, thereby prolonging the presence and activity of histatin 1 in the oral cavity and indirectly maintaining its functional concentration at the site of action.57
The hydrolysis of the genetically encoded histatin 1 and histatin 3 into smaller, often more active fragments, suggests a functional adaptation rather than a simple degradation process. Histatin 1, due to its high affinity for tooth enamel, plays a key role in the formation of the acquired enamel pellicle, where it undergoes controlled proteolysis.58 This localized hydrolysis may serve as a regulatory mechanism, ensuring that the antimicrobial activity of histatins is maximized precisely where it is needed – at the interface between the enamel and the oral microbiome. The enhanced activity of hydrolysis products highlights the importance of peptide fragmentation as a factor modulating their biological function, which should be considered when designing synthetic analogues or peptide-based therapeutics.
Conclusions
Histatins form thermodynamically stable complexes with Cu(II) and Zn(II). Cu(II) engages ATCUN sites when available, whereas Zn(II) binds via –HEXXH– coordination across sequences. Despite robust binding, metallation does not yield a systematic gain in antimicrobial potency: MIC values are modest and largely decoupled from complex stability, indicating that metal coordination is a context-dependent modulator rather than a universal activity switch.Across the Zn(II) series, coordination involves two His imidazoles and a Glu carboxylate within the –HEXXH– motif, but sequence context governs performance. Histatin 8 shows the expected Cu(II) robustness due to ATCUN chemistry, whereas the Zn(II) complexes of histatin 7 and 9 display near-overlapping thermodynamics over most of the pH range. The strong polarizing character of Zn(II), particularly when an aqua ligand completes a three-nitrogen coordination sphere, likely contributes to these subtle speciation differences without directly boosting antimicrobial potency. Terminal arginine additions do not measurably stabilize Zn(II) binding or enhance killing, underscoring the limits of simple electrostatics.
Metallation does not measurably alter the secondary structure under our conditions, and no clear link emerges between conformation and activity. Together with the weak MIC–stability relationship, this argues against nutritional immunity as the dominant mechanism for these peptides.
Overall, three environmental factors dominate efficacy: localization on the enamel/pellicle (concentrates peptides and limits proteolysis), pH-dependent protonation (helps to interact with pathogen membranes and strengthens surface binding), and metal availability. In this context, proteolysis of histatin 1 or 3 redistributes charge and places active fragments at the enamel–biofilm interface.
Design implications follow directly from this context: aim for pellicle targeting, tune net cationic charge via pH or the protonation state rather than only by residue additions, adjust Zn(II)/Cu(II) binding to modulate speciation and conformation as needed but not as a sole lever of potency, and favour compact, optimized fragments that retain enamel affinity.
Author contributions
E. D., A. M-T., L. P-S., K. S., M. B-O., J. W., A. M.-W., and M. R-Ż. conceived and planned the experiments. E. D., A. M.-T. and L. P-S. conducted the experiments. E. D., A. M-T., L. P-S., K. S., M. B-O., A. M.-W., J. W and M. R.-Ż. analysed the results. All authors reviewed the manuscript.Data availability
The data supporting the article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5dt02485d.If any additional questions regarding experimental details may arise, the corresponding authors remain at the readers’ disposal.
Conflicts of interest
There are no conflicts to declare.Acknowledgements
Support from the National Science Centre (UMO-2022/47/O/ST4/01865) is gratefully acknowledged.References
- E. C. Veerman, P. A. van den Keybus, A. Vissink and A. V. Nieuw Amerongen, Eur. J. Oral Sci., 1996, 104, 346–352 CrossRef CAS PubMed.
- K. Yamasaki and R. L. Gallo, Eur. J. Dermatol., 2008, 18, 11–21 CAS.
- S.-U. Gorr, Periodontology, 2009, 51, 152–180 CrossRef.
- W.-I. Chang, J.-Y. Chang, Y.-Y. Kim, G. Lee and H. S. Kho, Arch. Oral Biol., 2011, 56, 885–890 CrossRef CAS PubMed.
- S. S. Schwartz, D. I. Hay and S. K. Schluckebier, Calcif. Tissue Int., 1992, 50, 511–517 CrossRef CAS PubMed.
- F. G. Oppenheim, T. Xu, F. M. McMillian, S. M. Levitz, R. D. Diamond, G. D. Offner and R. F. Troxler, J. Biol. Chem., 1988, 263, 7472–7477 CrossRef CAS.
- M. Campese, X. Sun, J. A. Bosch, F. G. Oppenheim and E. J. Helmerhorst, Arch. Oral Biol., 2009, 54(4), 345–353 CrossRef CAS.
- M. Castagnola, T. Cabras, G. Denotti, M. B. Fadda, G. Gambarini, A. Lupi, I. Manca, G. Onnis, V. Piras, V. Soro, S. Tambaro and I. Messana, Biol. Rhythm Res., 2002, 33, 213–222 CrossRef CAS.
- D. A. Johnson, C. K. Yeh and M. W. Dodds, Arch. Oral Biol., 2000, 45, 731–740 CrossRef CAS.
- M. Campese, X. Sun, J. A. Bosch, F. G. Oppenheim and E. J. Helmerhorst, Arch. Oral Biol., 2009, 54, 345–353 CrossRef CAS PubMed.
- X. Sun, E. Salih, F. G. Oppenheim and E. J. Helmerhorst, FASEB J., 2009, 23, 2691–2701 CrossRef CAS.
- I. Messana, T. Cabras, E. Pisano, M. T. Sanna, A. Olianas, B. Manconi and M. Castagnola, Mol. Cell. Proteomics, 2008, 7, 911–926 CrossRef CAS.
- S. Puri and M. Edgerton, Eukaryotic Cell, 2014, 13(8), 958–964 CrossRef.
- S. E. Conklin, E. C. Bridgman, Q. Su, P. Riggs-Gelasco, K. L. Haas and K. J. Franz, Biochemistry, 2017, 56(32), 4244–4255 CrossRef CAS PubMed.
- H. Du, S. Puri, A. McCall, H. L. Norris, T. Russo and M. Edgerton, Front. Cell. Infect. Microbiol., 2017, 7, 41 Search PubMed.
- K. Takano, D. Malamud, A. Bennick, F. G. Oppenheim and A. R. Hand, Crit. Rev. Oral Biol. Med., 1993, 4, 399–405 CrossRef CAS PubMed.
- M. Ahmad, M. Piludu, F. G. Oppenheim, E. J. Helmerhorst and A. R. Hand, J. Histochem. Cytochem., 2004, 52, 361–370 CrossRef CAS.
- D. Shah, M. Ali, Z. Pasha, A. J. Jaboori, S. H. Jassim, S. Jain and V. K. Aakalu, PLoS One, 2016, 11, e0148018 CrossRef.
- T. Yaguchi and Y. Kawakami, PATENT WO/2012/023588A1, 2012.
- W. Wongpanuwich, S. Yodsanga, R. Chaisuparat and P. Amornphimoltham, Anticancer Res., 2022, 42, 2689–2699 CrossRef CAS.
- P. Jenwanichkul and P. Amornphimoltham, Exp. Oncol., 2024, 46(2), 101–109 CrossRef CAS.
- T. Xu, S. M. Levitz, R. D. Diamond and F. G. Oppenheim, Infect. Immun., 1991, 59(8), 2549–2554 CrossRef CAS PubMed.
- A. Shimotoyodome, H. Kobayashi, I. Tokimitsu, T. Matsukubo and Y. Takaesu, Caries Res., 2006, 40, 403–411 CrossRef CAS.
- R. Vitorino, M. J. Lobo, J. R. Duarte, A. J. Ferrer-Correia, P. M. Domingues and F. M. Amado, Biomed. Chromatogr., 2005, 19, 214–222 CrossRef CAS.
- X. Sun, X. Huang, X. Tan, Y. Si, X. Wang, F. Chen and S. Zheng, J. Transl. Med., 2016, 14, 240 CrossRef.
- J. Driscoll, Y. Zuo, T. Xu, J. R. Choi, R. F. Troxler and F. G. Oppenheim, J. Dent. Res., 1995, 74, 1837–1844 CrossRef CAS.
- W. L. Siqueira, H. C. Margolis, E. J. Helmerhorst, F. M. Mendes and F. G. Oppenheim, J. Dent. Res., 2010, 89, 626–630 CrossRef CAS.
- H. C. Margolis, S. Y. Kwak and H. Yamazaki, Front. Physiol., 2024, 5, 339 Search PubMed.
- E. E. McDonald, H. A. Goldberg, N. Tabbara, F. M. Mendes and W. L. Siqueira, J. Dent. Res., 2011, 90, 268–272 CrossRef CAS.
- M. J. Oudhoff, J. G. Bolscher, K. Nazmi, H. Kalay, W. Van't Hof, A. V. Amerongen and E. C. Veerman, FASEB J., 2008, 22, 3805–3812 CrossRef CAS PubMed.
- P. Torres, M. Castro, M. Reyes and V. A. Torres, Oral Dis., 2018, 24, 1150–1160 CrossRef CAS PubMed.
- I. A. Van Dijk, E. C. I. Veerman, E. A. J. Reits, J. G. M. Bolscher and J. Stap, Biol. Chem., 2018, 399, 1409–1419 CrossRef CAS.
- E. Dzień, J. Wątły, A. Kola, A. Mikołajczyk, A. Miller, A. Matera-Witkiewicz, D. Valensin and M. Rowińska-Żyrek, Dalton Trans., 2024, 53, 7561–7570 RSC.
- E. Dzień, J. Wątły, A. Hecel, A. Mikołajczyk, A. Matera-Witkiewicz, M. Adrover, M. Barceló-Oliver, A. Domínguez-Martín and M. Rowińska-Żyrek, Dalton Trans., 2024, 53, 19202–19213 RSC.
- L. Sabatini and E. Azen, Biochem. Biophys. Res. Commun., 1989, 160, 495–502 CrossRef CAS.
- G. Gran, Acta Chem. Scand., 1950, 4, 559–577 CrossRef CAS.
- P. Gans and B. O'Sullivan, Talanta, 2000, 51, 33–37 CrossRef CAS.
- P. Gans, A. Sabatini and A. Vacca, Talanta, 1996, 43, 1739–1753 CrossRef CAS PubMed.
- L. Petitt, IUPAC Stability Constants Database, Vol. 23, Chemistry International – Newsmagazine for IUPAC, 2001.
- L. Alderighi, P. Gans, A. Ienco, D. Peters, A. Sabatini and A. Vacca, Coord. Chem. Rev., 1999, 184, 311–318 CrossRef CAS.
- International Organization for Standardization, Susceptibility testing of infectious agents and evaluation of performance of antimicrobial susceptibility test devices–Part 1: Broth micro-dilution reference method for testing the in vitro activity of antimicrobial agents against rapidly growing aerobic bacteria involved in infectious diseases, 2019.
- International Organization for Standardization, Clinical laboratory testing and in vitro diagnostic test systems–Reference method for testing the in vitro activity of antimicrobial agents against yeast fungi involved in infectious diseases, 2012.
- J. Gabrielson, M. Hart, A. Jarelov, I. Kuhn, D. McKenzie and R. Mollby, J. Microbiol. Methods, 2002, 50, 63–73 CrossRef CAS.
- F. L. Francisco, A. M. Saviano, T. D. J. A. Pinto and F. R. Lourenço, J. Microbiol. Methods, 2014, 103, 104–111 CrossRef CAS PubMed.
- P. Sabaeifard, A. Abdi-Ali, M. R. Soudi and R. Dinarvand, J. Microbiol. Methods, 2014, 105, 134–140 CrossRef CAS PubMed.
- The European Committee on Antimicrobial Susceptibility Testing, Breakpoint tables for interpretation of MICs and zone diameters, Version 14.0, 2024, https://www.eucast.org (accessed on 31 March 2025).
- The European Committee on Antimicrobial Susceptibility Testing, Breakpoint tables for interpretation of MICs for antifungal agents, version 10.0, 2020. https://www.eucast.org/astoffungi/clinicalbreakpointsforantifungals/ (accessed on 31 March 2025).
- G. Repetto, A. Del Peso and J. L. Zurita, Nat. Protoc., 2008, 3, 1125–1131 CrossRef CAS.
- A. Miller, A. Matera-Witkiewicz, A. Mikołajczyk-Tarnawa, A. Kola, M. Wiloch, M. Jonsson-Niedziolka, R. Wieczorek, J. Wątły, D. Valensin and M. Rowińska-Żyrek, Chem. Sci., 2025, 16, 3447–3458 RSC.
- J. Wątły, K. Szarszoń, A. Mikołajczyk, M. Grelich-Mucha, A. Matera-Witkiewicz, J. Olesiak-Bańska and M. Rowińska-Żyrek, Inorg. Chem., 2023, 62, 19786–19794 CrossRef.
- A. Hecel, A. Kola, D. Valensin, H. Kozłowski and M. Rowińska-Żyrek, Inorg. Chem., 2020, 59, 1947–1958 CrossRef CAS PubMed , Express.
- A. Miller, A. Matera-Witkiewicz, A. Mikołajczyk, R. Wieczorek and M. Rowińska-Żyrek, Inorg. Chem., 2021, 60, 12730–12734 CrossRef CAS PubMed.
- J. Huang, B. J. Jones and R. J. Kazlauskas, Biochemistry, 2015, 54(28), 4330–4341 CrossRef CAS.
- L. Adibi, P. Yaghmaei, P. Maghami and A. Ebrahim-Habibi, AMB, 2024, 14(1), 69 CAS.
- C. L. Jr Borders, J. A. Broadwater, P. A. Bekeny, J. E. Salmon, A. S. Lee, A. M. Eldridge and V. B. Pett, Protein Sci., 1994, 3(4), 541–548 CrossRef.
- S. Leveraro, M. D'Accolti, E. Marzola, E. Caselli, R. Guerrini, M. Rowińska-Żyrek, M. Remelli and D. Bellotti, J. Inorg. Biochem., 2025, 262, 112761, DOI:10.1016/j.jinorgbio.2024.112761.
- T. Baumann, S. H. Niemeyer, M. A. R. Buzalaf and T. S. Carvalho, Sci. Rep., 2023, 13, 8618 CrossRef CAS.
- C. F. Richardson, M. Johnsson, P. A. Raj, M. J. Levine and G. H. Nancollas, Arch Oral Biol., 1993, 38(11), 997–1002 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |