Solvent- and metal-gated coordination diversity in a dynamic aminal ligand
Jen-Wen
Juan
a,
Ji-Ming
Ciou
b,
Yi-Rong
Hsiang†
ac and
Ya-Fan
Lin
 *a
*a
aDepartment of Chemistry, National Dong Hwa University, Hualien 974301, Taiwan. E-mail: yafan@gms.ndhu.edu.tw
bPhD Program in Life Science, College of Life Science, Kaohsiung Medical University, Kaohsiung 807378, Taiwan
cDepartment of Medicinal and Applied Chemistry, Kaohsiung Medical University, Kaohsiung 807378, Taiwan
First published on 2nd December 2025
Abstract
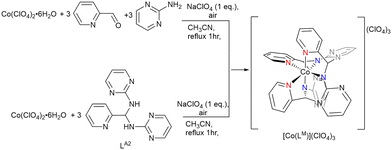
Within the broad landscape of dynamic covalent chemistry (DCC), the coordination chemistry of primary-amine-based aminals has received comparatively little systematic investigation. Here we demonstrate that a primary-amine-based dynamic covalent aminal (LA2) can serve as an adaptive scaffold for coordination chemistry diversity, where solvent and metal ion choice govern distinct outcomes. Coordination stabilizes otherwise transient ligand forms and directs divergent pathways: Co(III) stabilizes hemiaminalates in methanol, with [Co(LO2)2](ClO4) exhibiting fluxional behavior, and templates macrocyclization in acetonitrile to yield [Co(LM)](ClO4)3; Cu(II) affords hemiaminal and hemiaminal ether complexes, and Fe(III) promotes pyridinium dimerization to give cis-[(LI2)2](ClO4). These findings establish that dynamic ligands can actively generate structural diversity in metal complexes, opening new directions for adaptive ligand design and functional coordination architectures.
Introduction
Primary-amine-based aminals can be considered as precursors to imines,1–4 and many systems exist in reversible equilibrium between imine and aminal forms, displaying dynamic covalent properties.5–8 Imines are versatile scaffolds in organic synthesis, sensing, and materials chemistry,9–17 and their reversibility has been widely exploited in dynamic covalent chemistry (DCC) to construct adaptive and responsive molecular architectures.18–25 While DCC has often been applied to macromolecular self-assemblies and higher-order structures,26–33 its potential in stabilizing transient small-molecule species through metal coordination remains comparatively underexplored.34–38Dynamic aminals derived from primary amines readily interconvert with imines, which in turn serve as competent ligands for metal coordination. Although imines possess lower electrophilicity than both iminium ions and aldehydes/ketones, they can still undergo nucleophilic addition, thereby providing access to otherwise elusive species such as hemiaminals and hemiaminal ethers. Notably, most metal-stabilized hemiaminal or hemiaminal-ether systems reported to date arise from secondary amines or cyclic aminals.39–45 In contrast, primary-amine-based aminals enable a broader set of interconnected equilibria involving imines, hemiaminals, and hemiaminal ethers, yet their coordination behaviour has been far less systematically explored. In this context, Anslyn and co-workers established a highly adaptive Zn(II)-templated dynamic system that provides a key precedent. In their framework, 2-pyridinecarboxaldehyde (2-PA) and 4-methylaniline first generate an imine, and subsequent addition of the secondary amine di(2-picolyl)amine (DPA) leads to dynamic metal-templated assemblies.38 They further demonstrated aniline-exchange behaviour using para-substituted anilines, with more electron-donating anilines displaying larger equilibrium constants. In a related Zn(II)/2-PA/DPA system, they also showed reversible formation of hemiaminals and hemiaminal ethers upon alcohol addition,37,46 as well as aminal-type complexes upon introducing secondary amines.47
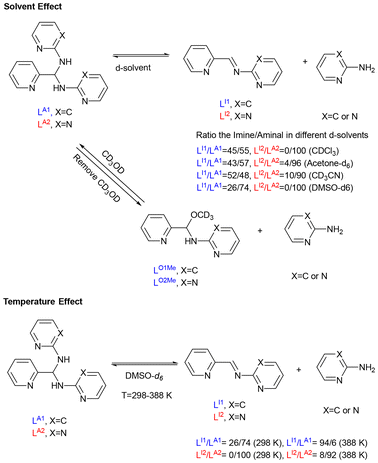
In our prior work (Scheme 1), we developed primary-amine-based heterocyclic aminals LA1 and LA2, both of which display solvent- and condition-dependent equilibria, generating imines, hemiaminals, and hemiaminal ethers that are detectable in solution but difficult to isolate.7 Previous studies of LA1-related complexes largely emphasized structural characterization, without probing their underlying equilibria or adaptive properties.48–51 In contrast, the present study systematically investigated LA2 as a dynamic covalent ligand in coordination chemistry. Inspired by the adaptivity observed in Anslyn's system, we explored whether coordination to different metal ions in different solvent systems could stabilize these inherently dynamic forms, enabling structural characterization and directing their reactivity along divergent pathways. A critical question is whether such dynamic ligands can be deliberately harnessed to expand structural diversity in metal complexes.
This study demonstrates that dynamic covalent ligand LA2 can act as an adaptive scaffold for coordination chemistry diversity. Coordination stabilizes otherwise transient ligand forms and directs their reactivity along divergent pathways: Co(III) stabilizes hemiaminalates in methanol and templates macrocyclization in acetonitrile, Cu(II) stabilizes hemiaminals or hemiaminal ethers, and Fe(III) induces pyridinium dimerization.52 These findings establish a conceptual framework in which dynamic covalent ligands serve not merely as passive participants but as active drivers of coordination diversity, opening new directions for adaptive ligand design and functional coordination architectures.
Results and discussion
Influence of solvent
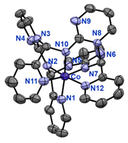
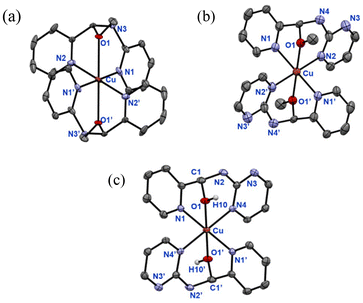
Given the ability of the dynamic aminal systems of LA1 and LA2 to reversibly transform into hemiaminal ether and imine through solvent tuning, we initiated an investigation into the coordination chemistry of these dynamic ligands by examining their behaviour in various solvents in the presence of metal ions.[Co(LO2Me)2](ClO4)2 is unstable in air, undergoing gradual oxidation to give a pink powder. After purification and characterization, the product was identified as the Co(III) complex, [Co(LO2)2](ClO4), which crystallizes in an non-centrosymmetric structure within the monoclinic space group P21/c (Fig. 1(b)). The two Co–O bond lengths (1.894(2), 1.892(2) Å) are significantly shorter than those in [Co(LO2Me)2](ClO4)2 (2.1362 (15) Å), and the four Co–N bond lengths (average ∼1.95 Å) are also shorter than in [Co(LO2Me)2](ClO4)2 (average ∼2.13 Å). These shortened donor-Co distances enforce larger bite angles in [Co(LO2)2](ClO4), reflecting the more compact geometry of a Co(III) complex.
The analogue [Co(LO1)2](ClO4) has previously been reported, though only its molecular structure was described. Attempts to prepare the corresponding hemiaminal ether complex [Co(LO1Me)2](ClO4)2 under aerobic conditions yielded only [Co(LO1)2](ClO4). UV-Vis absorption spectra revealed that [Co(LO1)2](ClO4) and [Co(LO2)2](ClO4) both display d–d transition bands at λmax = 510 nm (ε = 64 and 69 M−1 cm−1, respectively) in methanol, indicating similar coordination geometries and comparable ligand field environments.
To improve the yield, the synthesis was optimized using a one-pot reaction of Co(ClO4)2·6H2O, 2-pyridinecarboxaldehyde, 2-aminopyrimidine in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
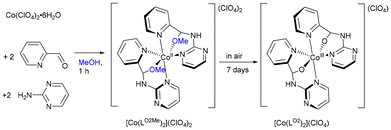
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 ratio in methanol, followed by addition of two equivalents of TBHP as oxidant and overnight stirring. Evaporation of the solvent and the recrystallization from acetone afforded [Co(LO2)2](ClO4) in good yield. Interestingly, the same product can also be obtained by reacting Co(ClO4)2·6H2O with two equivalents of the aminal ligand LA2 in methanol under aerobic conditions, without isolating the Co(II) intermediate [Co(LO2Me)2](ClO4)2 (Scheme 3).
2 ratio in methanol, followed by addition of two equivalents of TBHP as oxidant and overnight stirring. Evaporation of the solvent and the recrystallization from acetone afforded [Co(LO2)2](ClO4) in good yield. Interestingly, the same product can also be obtained by reacting Co(ClO4)2·6H2O with two equivalents of the aminal ligand LA2 in methanol under aerobic conditions, without isolating the Co(II) intermediate [Co(LO2Me)2](ClO4)2 (Scheme 3).
The 1H NMR spectra in DMSO-d6 showed the methine proton resonance of the hemiaminalate moieties at 5.54 ppm for [Co(LO1)2](ClO4) and 5.65 ppm for [Co(LO2)2](ClO4), with corresponding 13C signals at 84.2 ppm and 84.6 ppm. Both complexes displayed minor isomeric species, with major /minor ratios of approximately 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 for [Co(LO1)2](ClO4) and 12
1 for [Co(LO1)2](ClO4) and 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 for [Co(LO2)2](ClO4) (Fig. S10 and S14). For [Co(LO1)2](ClO4), the ratio was solvent-independent (Fig. S11 and S12), whereas for [Co(LO2)2](ClO4) it varied (8
1 for [Co(LO2)2](ClO4) (Fig. S10 and S14). For [Co(LO1)2](ClO4), the ratio was solvent-independent (Fig. S11 and S12), whereas for [Co(LO2)2](ClO4) it varied (8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 in CD3OD and 6
1 in CD3OD and 6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 in CD3CN), underscoring the structural inertness for [Co(LO1)2](ClO4) versus the fluxionality of [Co(LO2)2](ClO4) (Fig. S15 and S16). Dissolving crystalline [Co(LO2)2](ClO4) in DMSO-d6 reproduced the same spectral features, confirming the intrinsic lability of its hemiaminalate ligands.
1 in CD3CN), underscoring the structural inertness for [Co(LO1)2](ClO4) versus the fluxionality of [Co(LO2)2](ClO4) (Fig. S15 and S16). Dissolving crystalline [Co(LO2)2](ClO4) in DMSO-d6 reproduced the same spectral features, confirming the intrinsic lability of its hemiaminalate ligands.
Density functional theory (DFT) calculations at the B3LYP-D3 level with the LanL2DZ basis set for Co and 6-31g* for H, C, and N were carried out using Gaussian 16 program.53–55 Only isomers with equivalent hemiaminalate environments were considered, consistent with the 1H NMR spectra, which showed a single set of ligand signals. As shown in Fig. 2, the crystal form OC-6-13-[Co(LO2)2]+ was the most stable and used as the reference. The Ci-symmetric OC-6-12-[Co(LO2)2]+ was only 2.0 kcal mol−1 higher in energy, whereas OC-6-22- and OC-6-33-[Co(LO2)2]+ were 11.0 and 12.7 kcal mol−1 less stable, suggesting they are minor species. Accordingly, the major and minor isomers observed in solution were assigned to OC-6-13-[Co(LO2)2]+ and OC-6-12-[Co(LO2)2]+, respectively. The increased OC-6-13-[Co(LO2)2]+/OC-6-12-[Co(LO2)2]+ ratio in polar solvents correlates with their calculated dipole moments (41.5 D vs. 0 D), indicating the solvent polarity governs the isomer distribution.
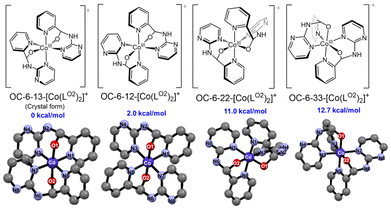 | ||
| Fig. 2 Schematic representations and optimized structures of the possible isomers, along with their relative potential energies. The isomer names follow the configuration index system recommended by IUPAC.56 | ||
Interestingly, [Co(LO2)2](NO3) can be obtained by replacing Co(ClO4)2·6H2O with Co(NO3)2·6H2O. Its NMR spectra resembled those of [Co(LO2)2](ClO4), showing two sets of 1H NMR signals but with smaller major/minor ratios of 1.6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 2.5
1, 2.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 and 3.5
1 and 3.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 in DMSO-d6, CD3OD, and CD3CN, respectively (Fig. S18–S20). This facilitated the isolation of two distinct crystals from CH3CN: one non-centrosymmetric, analogous to [Co(LO2)2](ClO4), and one centrosymmetric, corresponding to the predicted iso1-[Co(LO2)2]+ (Fig. S1(a) and (b)). Occasionally, we also obtained neutral [Co(LO2)2] in a centrosymmetric configuration (Fig. S1(c)), although this form was unstable and decomposed to an unidentified diamagnetic Co(III) species.
1 in DMSO-d6, CD3OD, and CD3CN, respectively (Fig. S18–S20). This facilitated the isolation of two distinct crystals from CH3CN: one non-centrosymmetric, analogous to [Co(LO2)2](ClO4), and one centrosymmetric, corresponding to the predicted iso1-[Co(LO2)2]+ (Fig. S1(a) and (b)). Occasionally, we also obtained neutral [Co(LO2)2] in a centrosymmetric configuration (Fig. S1(c)), although this form was unstable and decomposed to an unidentified diamagnetic Co(III) species.
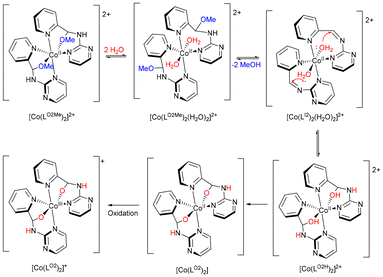
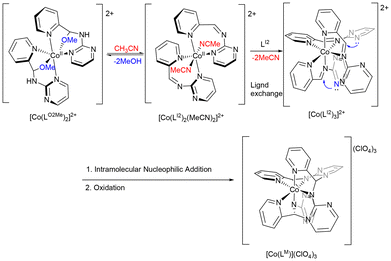
We proposed the mechanism for the conversion of [Co(LO2)2]+ from [Co(LO2Me)2]2+ (Scheme 4). In methanol, hemiaminal ether LO2Me may remain metastable upon Co(II) coordination, but the –OMe moiety is labile and competes with aqua ligands from the metal ion hydration sphere or solvent. Equilibrium with [Co(LO2Me)2(H2O)2]2+ in solution enables elimination of methanol, giving [Co(LI2)2(H2O)2]2+, which undergoes intramolecular nucleophilic addition to form the hemiaminal intermediate [Co(LO2H)2]2+. Subsequent deprotonation by environmental species (e.g. MeOH or H2O) furnishes [Co(LO2)2], which is then oxidized by air to [Co(LO2)2]+. When the reaction was carried out by directly reacting the pre-synthesized ligand LA2 with Co(ClO4)2·6H2O in methanol, 2-aminopyrimidine was released during LO2Me formation in situ. Acting as a weak base, the liberated 2-aminopyrimidine facilitates rapid deprotonation of [Co(LO2H)2]2+, accelerating its conversion to [Co(LO2)2] followed by oxidation. This explains why [Co(LO2Me)2](ClO4)2 could not be isolated under these conditions.
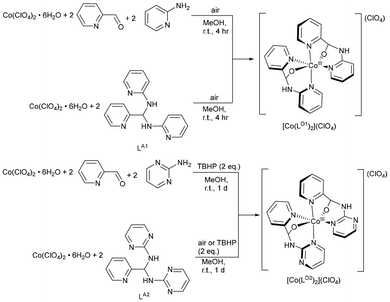
[Co(LM)](ClO4)3 was also obtained by reacting Co(ClO4)2·6H2O with three equivalents of LA2 in CH3CN, or via one-pot reaction of Co(ClO4)2·6H2O, 2-PA and 2-aminopyrimidine (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3
3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3) under aerobic conditions. Addition of one equivalent of NaClO4 to balance the charge of Co(III) and refluxing improved the yield to 60% (Scheme 6). The 1H NMR spectrum in DMSO-d6 showed a methine resonance at 7.97 ppm with a corresponding 13C signal at 80.3 ppm. The UV-Vis absorption spectrum in CH3CN showed a d–d transition band at λmax = 480 nm (ε = 315 M−1 cm−1). Unlike [Co(LO2)2](ClO4), [Co(LM)](ClO4)3 does not exhibit fluxional behaviour and is insoluble in methanol, but remains soluble and stable in CH3CN, D2O and DMSO-d6. Variable–temperature NMR (VT-NMR) analysis conducted in DMSO-d6 confirmed stability up to 403 K, above which decomposition yielded signals corresponding to 2-pyridinecarboxaldehyde and 2-aminopyrimidine (Fig. S21).
3) under aerobic conditions. Addition of one equivalent of NaClO4 to balance the charge of Co(III) and refluxing improved the yield to 60% (Scheme 6). The 1H NMR spectrum in DMSO-d6 showed a methine resonance at 7.97 ppm with a corresponding 13C signal at 80.3 ppm. The UV-Vis absorption spectrum in CH3CN showed a d–d transition band at λmax = 480 nm (ε = 315 M−1 cm−1). Unlike [Co(LO2)2](ClO4), [Co(LM)](ClO4)3 does not exhibit fluxional behaviour and is insoluble in methanol, but remains soluble and stable in CH3CN, D2O and DMSO-d6. Variable–temperature NMR (VT-NMR) analysis conducted in DMSO-d6 confirmed stability up to 403 K, above which decomposition yielded signals corresponding to 2-pyridinecarboxaldehyde and 2-aminopyrimidine (Fig. S21).
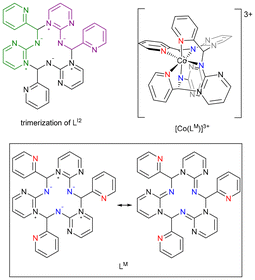
As illustrated in Scheme 7, the macrocyclic ligand LM likely form through trimerization of the imino fragment. In our previous study, approximately 10% of LA2 is converted to LI2 in CD3CN.7 Accordingly, LO2Me may undergo equilibration with LI2/LA2, releasing a methanol molecule in CH3CN. Based on these observations, [Co(LM)](ClO4)3 formation begins with [Co(LO2Me)2] undergoing –OMe substitution by CH3CN, followed by methanol elimination to form [Co(LI2)2(MeCN)2]2+. Ligand exchange then furnishes [Co(LI2)3]2+, which undergoes intramolecular nucleophilic attack of the pyrimidyl nitrogen on the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N group, driving cyclization to yield [Co(LM)]3+. Alternatively, [Co(LI2)3]2+ can also be generated directly through Co(II) templating of three LI2 units, which then undergo nucleophilic addition to produce [Co(LM)]3+ (Scheme 7). Consistently, when acetone was used as solvent, [Co(LM)](ClO4)3 was also obtained, albeit in slightly lower yield (45%), possibly due to reduced LI2 formation under these conditions.
N group, driving cyclization to yield [Co(LM)]3+. Alternatively, [Co(LI2)3]2+ can also be generated directly through Co(II) templating of three LI2 units, which then undergo nucleophilic addition to produce [Co(LM)]3+ (Scheme 7). Consistently, when acetone was used as solvent, [Co(LM)](ClO4)3 was also obtained, albeit in slightly lower yield (45%), possibly due to reduced LI2 formation under these conditions.
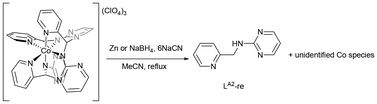
To obtain the free macrocyclic ligand LM, demetalation of [Co(LM)](ClO4)3 was attempted by adding either HCl or NaCN (six equivalents) in MeCN under reflux conditions.57 However, HCl left the complex unchanged, while NaCN led to decomposition reactions. Considering that Co(II) is more labile than Co(III), potentially facilitating demetalation, a reductant, such as NaBH4 or Zn dust was introduced to the reaction mixture to enhance the likelihood of successful demetalation.58,59 Unfortunately, this approach only yielded the amine, LA2-re, the reduction product of LA2 (Scheme 8). Analysis of the redox properties of [Co(LM)](ClO4)3 provides insight into this outcome. Cyclic voltammetry of a 10 mM solution of [Co(LM)](ClO4)3 in DMSO, using 0.1 M [NBu4][ClO4] as a supporting electrolyte and a 50 mV s−1 scan rate, was performed in the potential range of 1.0 to −2.0 V. During the anodic scan, no corresponding anodic peak was observed, whereas the cathodic scan exhibits peaks at −0.370, −0.646, and −1.224 V vs. Fc/Fc+ (Fig. S22), corresponding to the reduction of the ligand, the Co(III)/Co(II) transition and Co(II)/Co(I) transition, respectively.60–63 The experimental results suggest that the macrocyclic ligand LM undergoes reduction when treated with Zn dust, a mild reductant, with a reduction potential of −0.76 V.64 The observed formation of [Co(LM)](ClO4)3 is likely driven by its thermodynamic stability through self-assembly. Attempts to isolate the free ligand LM using acid, base or alternative demetalation strategies were unsuccessful.
Influence of metal ion
We next explored the coordination behaviour of Cu(ClO4)2·6H2O toward the picolyl heterocyclic amino aminals LA1 and LA2. In methanol, the reactions under aerobic conditions afforded [Cu(LO1Me)2](ClO4)2 and [Cu(LO2Me)2](ClO4)2 as green and blue powders, respectively (Scheme 9). Slow evaporation of methanol yielded single crystals suitable for X-ray diffraction. As shown in Fig. 4(a) and (b), both complexes crystallize in centrosymmetric structures with the space groups P21/c. The Cu–O bond lengths (∼2.4 Å) fall within the range typically observed for Cu(II)-neutral alcohol interaction,65–67 suggesting relatively weak coordination. The [Cu(LO1Me)2](ClO4)2 crystals were unstable in air, transforming into purple [Cu(pic)2] (pic = picolinate).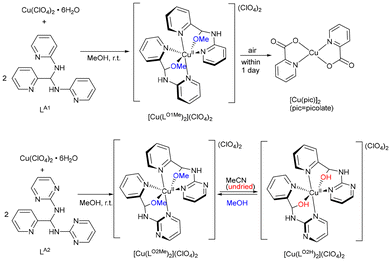 | ||
| Scheme 9 Reactions of LA1 and LA2 with Cu(ClO4)2·6H2O to generate [Cu(LO1Me)2](ClO4)2, [Cu(LO2Me)2](ClO4)2 and [Cu(LO2H)2](ClO4)2. | ||
In contrast, recrystallization of [Cu(LO2Me)2](ClO4)2 from CH2Cl2/CH3CN yielded [Cu(LO2H)2](ClO4)2, instead of a macrocyclic Cu(II) complex. Blue block-shape crystals of [Cu(LO2H)2](ClO4)2 were obtained by diffusing CH2Cl2 into an CH3CN solution. X-ray diffraction analysis (Fig. 4(c)) revealed a Cu(II) centre coordinated by two hemiaminal ligands, with Cu–O bond lengths of 2.2819(17), shorter than those observed in [Cu(LO2Me)2](ClO4)2. Thus, [Cu(LO2H)2](ClO4)2 can be prepared directly from Cu(ClO4)2·6H2O and two equivalents of LA2 in CH3CN.
Interestingly, redissolving [Cu(LO2H)2](ClO4)2 in methanol and recrystallized by slow solvent evaporation regenerated [Cu(LO2Me)2](ClO4)2. Likewise, HR-ESI-MS analysis revealed that dissolving [Cu(LO2H)2](ClO4)2 in ethanol led to the formation of [Cu(LO2Et)2](ClO4)2 (Fig. S31). These observations demonstrate that solvent choice (alcohol vs. non-alcohol) directs whether the Cu(II) centre coordinates to the hemiaminal ether or the hemiaminal ligands (Scheme 9). Consistent with its dynamic behaviour in solution, [Cu(LO2H)2](ClO4)2 exhibits distinct d–d transition bands: λmax = 656 nm (ε = 144 M−1 cm−1) in methanol and λmax = 689 nm (ε = 89 M−1 cm−1) in CH3CN.
DFT study of the formation of macrocyclic complex
We are intrigued by the novel macrocyclic LM structure and aim to address the following questions through calculations: (1) Under what conditions can the macrocyclic structure form? (2) Why does the Cu(II) system fail to yield the macrocyclic structure? Based on our hypothesis shown in Scheme 7, we performed simulations of [Co(LI2)3]3+ and [Co(LM)]3+ to explore the structural and energetic factors that govern the formation of the macrocyclic structure. The calculations were performed at the same level of theory previously applied to the isomers for [Co(LO2)]+.The optimized structure of [Co(LI2)3]3+ shows that the three LI2 ligands coordinate with Co(III) in a bidentate manner through the pyridyl N and imine N atoms. Additionally, we identified an n → π* interaction between the pyrimidyl N atom and the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N bond of the adjacent LI2 ligand (depicted by green dashed line in Fig. 5),68–72 with an energy of 0.58 kcal mol−1. The Bürgi-Dunitz (BD) trajectory parameters, including BD angle (α = 86.4°, the N–C–N angle in Fig. 5),73 were calculated, and the N–C distance (d) was found to be 2.77 Å,74–76 which is smaller than the sum of the van der Waals radii of N and C (3.25 Å). These findings suggest a significant intramolecular n → π* interaction between the pyrimidyl N and the adjacent C
N bond of the adjacent LI2 ligand (depicted by green dashed line in Fig. 5),68–72 with an energy of 0.58 kcal mol−1. The Bürgi-Dunitz (BD) trajectory parameters, including BD angle (α = 86.4°, the N–C–N angle in Fig. 5),73 were calculated, and the N–C distance (d) was found to be 2.77 Å,74–76 which is smaller than the sum of the van der Waals radii of N and C (3.25 Å). These findings suggest a significant intramolecular n → π* interaction between the pyrimidyl N and the adjacent C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N bond. When comparing the potential energies of [Co(LI2)3]3+ and [Co(LM)]3+, we found that [Co(LM)]3+ is 24.5 kcal mol−1 more stable, which explains why the reaction favours the formation of the macrocyclic [Co(LM)](ClO4)3, driven by its thermodynamic stability.
N bond. When comparing the potential energies of [Co(LI2)3]3+ and [Co(LM)]3+, we found that [Co(LM)]3+ is 24.5 kcal mol−1 more stable, which explains why the reaction favours the formation of the macrocyclic [Co(LM)](ClO4)3, driven by its thermodynamic stability.
 | ||
| Fig. 5 Comparison of the energy difference between [Co(LM)]3+ (right) and [Co(LI2)3]3+ (left), calculation of En→π* and the Bürgi–Dunitz parameters (α (°), d (Å)) for [Co(LI2)3]3+. | ||
Conversely, when Cu(II) is used as the central metal, the simulated [Cu(LI2)]2+ shows an intramolecular n → π* interaction with an energy of 0.25 kcal mol−1. The N–C distance in this case is 3.26 Å, which is nearly equal to the sum of the van der Waals radii of N and C, indicating a much weaker n → π* interaction in [Cu(LI2)3]2+ compared to [Co(LI2)3]3+. Furthermore, the potential energy of [Cu(LI2)]2+ is 7.24 kcal mol−1 lower than [Cu(LM)]3+, suggesting that the formation of the macrocyclic LM complex is thermodynamically unfavourable in the Cu(II) system. The electron-deficient nature of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N bond after Cu(II) coordination makes it susceptible to attack by water molecules, leading to the formation of [Cu(LO2H)2]2+.
N bond after Cu(II) coordination makes it susceptible to attack by water molecules, leading to the formation of [Cu(LO2H)2]2+.
To further explore the formation of the metal complexes with the macrocyclic LM ligand beyond [Co(LM)]3+, we performed theoretical calculations on 12 metal ions (Sc3+, Ti2+, Ti4+, V2+, Cr2+, Mn2+, Fe2+, Fe3+, Co2+, Ni2+, Zn2+, and Al3+). We compared the potential energies of [M(LI2)3]n+ and [M(LM)]n+ structures (Fig. 6) and examined the presence of interactions between the pyrimidyl N and the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N bond of the adjacent LI2 ligand in [M(LI2)3]n+. The results are summarized in Table 1. From the calculations, Ti2+ and Mn2+ exhibit relatively small En→π* values, while the remaining metal ions show values ranging from 0.25 to 1.44 kcal mol−1. Notably, six metal ions—Sc3+, Ti4+, Fe2+, Co2+, Ni2+, and Zn2+—exhibit En→π* values for [M(LI2)3]n+ that are comparable to or stronger than that of Co3+. Among these, Co2+ and Ni2+ show the highest values, both exceeding 1.4 kcal mol−1. However, for Sc3+, Ti2+, and Zn2+, the distance between the pyrimidyl N and the C
N bond of the adjacent LI2 ligand in [M(LI2)3]n+. The results are summarized in Table 1. From the calculations, Ti2+ and Mn2+ exhibit relatively small En→π* values, while the remaining metal ions show values ranging from 0.25 to 1.44 kcal mol−1. Notably, six metal ions—Sc3+, Ti4+, Fe2+, Co2+, Ni2+, and Zn2+—exhibit En→π* values for [M(LI2)3]n+ that are comparable to or stronger than that of Co3+. Among these, Co2+ and Ni2+ show the highest values, both exceeding 1.4 kcal mol−1. However, for Sc3+, Ti2+, and Zn2+, the distance between the pyrimidyl N and the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N group on the adjacent LI2 exceeds the sum of the van der Waals radii of C and N, suggesting weaker interactions. In terms of stability, only [Ti(LM)]4+, [Fe(LM)]3+, and [Al(LM)]3+exhibit comparable stability to [Co(LM)]3+ when using [M(LI2)3]n+ as the reference. Although [V(LM)]2+ and [Fe(LM)]2+ are also more stable than their corresponding [M(LI2)3]n+, the differences are within 10 kcal mol−1. For the remaining metal ions, [M(LI2)3]n+ is the more stable species. Interestingly, [Co(LM)]2+ is slightly less stable than [Co(LI2)3]2+ by 0.65 kcal mol−1. Given that the starting material for [Co(LM)]3+ is Co(II), the formation of the highly stable [Co(LM)]3+ in CH3CN likely drives the reaction forward under aerobic conditions.
N group on the adjacent LI2 exceeds the sum of the van der Waals radii of C and N, suggesting weaker interactions. In terms of stability, only [Ti(LM)]4+, [Fe(LM)]3+, and [Al(LM)]3+exhibit comparable stability to [Co(LM)]3+ when using [M(LI2)3]n+ as the reference. Although [V(LM)]2+ and [Fe(LM)]2+ are also more stable than their corresponding [M(LI2)3]n+, the differences are within 10 kcal mol−1. For the remaining metal ions, [M(LI2)3]n+ is the more stable species. Interestingly, [Co(LM)]2+ is slightly less stable than [Co(LI2)3]2+ by 0.65 kcal mol−1. Given that the starting material for [Co(LM)]3+ is Co(II), the formation of the highly stable [Co(LM)]3+ in CH3CN likely drives the reaction forward under aerobic conditions.
 | ||
| Fig. 6 Comparison of the energy difference between [M(LM)]n+ (right) and [M(LI2)3]n+ (left), calculation of En→π* and the Bürgi–Dunitz parameters (α (°), d (Å)) for [M(LI2)3]n+. | ||
| ΔEa |
E
n→π*![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) b b |
α![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) c c |
d![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) d d |
|
|---|---|---|---|---|
| a Comparison of the energy (kcal mol−1) difference between [MLM]n+ and [M(LI2)3]n+. b Calculation of En→π* (kcal mol−1). c The Bürgi-Dunitz angles α (°) in [M(LI2)3]n+. d The Npym⋯Cimined (Å) in [M(LI2)3]n+. | ||||
| Sc3+ | 5.73 | 0.76 | 110.7 | 3.48 |
| Ti2+ | 15.87 | 0.08 | 100.2 | 3.54 |
| Ti4+ | −22.99 | 0.89 | 100.3 | 3.24 |
| V2+ | 8.37 | 0.24 | 89.3 | 3.25 |
| Cr2+ | 7.95 | 0.31 | 76.9 | 3.00 |
| Mn2+ | 0.85 | 0.14 | 77.6 | 3.01 |
| Fe2+ | −3.64 | 1.01 | 74.3 | 2.94 |
| Fe3+ | −24.05 | 0.34 | 70.0 | 2.80 |
| Co2+ | 0.65 | 1.43 | 68.5 | 3.00 |
| Co3+ | −24.49 | 0.58 | 86.4 | 2.77 |
| Ni2+ | 2.93 | 1.44 | 79.6 | 3.11 |
| Cu2+ | 7.24 | 0.25 | 77.5 | 3.26 |
| Zn2+ | 10.79 | 0.89 | 97.8 | 3.48 |
| Al3+ | −16.58 | 0.32 | 83.4 | 3.08 |
Iron(III) as the template
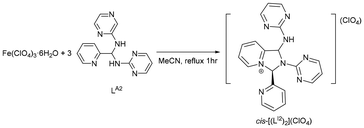
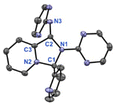
Based on DFT calculations, Fe(III) was chosen as a template to explore whether a macrocyclic complex analogue to [Co(LM)]3+ could be formed. However, unexpectedly, when the reaction was conducted under conditions similar to those used for synthesizing [Co(LM)]3+, but with Fe(ClO4)3·6H2O as the metal source, a pyridinium salt, cis-[(LI2)2](ClO4), was isolated along with unidentified Fe(III)-containing products (Scheme 10). The crystal structure of cis-[(LI2)2](ClO4), shown in Fig. 7, suggests that the presence of the Fe(ClO4)3·6H2O may promote the dimerization of LI2 in a cis configuration. Based on these observations, it is possible that the coordination environment and ligand orientation prior to dissociation may influence whether the system evolves toward macrocyclization or dimerization; however, this aspect remains unclear and is currently being investigated. The detailed mechanism and the stereoselectivity of this reaction are currently under investigation.Conclusions
In this study, we demonstrated that solvent and metal ion choice direct picolyl heterocyclic aminals into divergent coordination outcomes, including fluxional hemiaminalates, hemiaminal ethers, a Co(III)-templated polyazamacrocyclic complex, and an Fe(III)-induced pyridinium salt. A central finding is that coordination can stabilize inherently dynamic ligand forms, such as hemiaminals and hemiaminal ethers, enabling their isolation and structural characterization. This principle highlights how dynamic covalent scaffolds, often considered too labile for study, can become accessible through metal coordination.By integrating reversibility with coordination, these systems reveal an expanded dimension of structural diversity in small-molecule chemistry. Unlike classical ligands, which deliver predictable geometries, dynamic aminals adapt their form in response to metal ions and solvents, giving rise to orthogonal reactivity pathways and unanticipated structures. This ability of coordination to both stabilize and redirect dynamic species enriches the conceptual framework of coordination chemistry diversity.
Beyond fundamental interest, such adaptive scaffolds provide a versatile platform for designing responsive functional complexes. Future work will extend this strategy to other metal ions and evaluate their potential in catalysis, molecular recognition, and biomedical applications.
Author contributions
J. W. J.: writing – original draft, writing – review & editing, validation, formal analysis, investigation, data curation, visualization. J. M. C.: writing – original draft, formal analysis, investigation, data curation, visualization. Y. R. H.: writing – review & editing, software, formal analysis, data curation, visualization. Y. F. L.: writing – original draft, writing – review & editing, conceptualization, methodology, visualization, resources, supervision, project administration, funding acquisition.Conflicts of interest
There are no conflicts to declare.Data availability
All remaining data are provided in the article and supplementary information (SI). Supplementary information is available. The SI contains synthetic procedures, additional experimental details, full characterization data (NMR, IR, UV–Vis, HR-ESI-MS), variable-temperature NMR studies, crystallographic information files (CIFs), supplementary figures and tables, and computational details. See DOI: https://doi.org/10.1039/d5dt02405f.CCDC 2423656–2423658, 2493551–2493557, 2493743 contain the supplementary crystallographic data for this paper.77a–k
Acknowledgements
We sincerely appreciate the financial support from the National Science and Technology Council (NSTC), Taiwan, through research grants NSTC 111-2621-M-A49-004-MY3, NSTC 112-2113-M-259-007, NSTC 113-2113-M-259-005 and NSTC 114-2113-M-259-002. We also extend our gratitude to the National Center for High-performance Computing (NCHC), Taiwan, for providing computational and storage resources. Special thanks go to Dr Chien-Hung Liao and Mr Wei-Rong Yen for their assistance with single-crystal X-ray diffraction measurements. We thank the Advanced Instrumentation Center of the Department of Medical Research, Hualien Tzu Chi Hospital, Buddhist Tzu Chi Medical Foundation, and the Bioinnovation Center, Buddhist Tzu Chi Medical Foundation, Hualien, Taiwan, for access to mass spectrometry facilities, as well as Dr. Hsueh-Hui Yang for support with high-resolution ESI-MS analyses. We are deeply grateful to Department of Chemistry at National Taiwan University and Institute of Chemistry, Academia Sinica, for generously providing laboratory space, instrumentation, and research facilities. Their support made it possible for us to continue our research following the 2024 post-earthquake fire, which tragically destroyed the entire Chemistry Department building at NDHU.References
- T. Kano, T. Yurino, D. Asakawa and K. Maruoka, Angew. Chem., Int. Ed., 2013, 52, 5532–5534 CrossRef CAS PubMed.
- Y.-Y. Huang, C. Cai, X. Yang, Z.-C. Lv and U. Schneider, ACS Catal., 2016, 6, 5747–5763 CrossRef CAS.
- T. Yurino, Y. Aota, D. Asakawa, T. Kano and K. Maruoka, Tetrahedron, 2016, 72, 3687–3700 CrossRef CAS.
- J. F. Bai, H. Sasagawa, T. Yurino, T. Kano and K. Maruoka, Chem. Commun., 2017, 53, 8203–8206 RSC.
- M. E. Belowich and J. F. Stoddart, Chem. Soc. Rev., 2012, 41, 2003–2024 RSC.
- Y. Zhou, L. Li, H. Ye, L. Zhang and L. You, J. Am. Chem. Soc., 2016, 138, 381–389 CrossRef CAS PubMed.
- J.-M. Ciou, H.-F. Zhu, C.-W. Chang, J.-Y. Chen and Y.-F. Lin, RSC Adv., 2020, 10, 40421–40427 RSC.
- Z. Feng, S. Jia, H. Chen and L. You, Tetrahedron, 2020, 76, 131128 CrossRef CAS.
- C. Palomo, J. M. Aizpurua, I. Ganboa and M. Oiarbide, Eur. J. Org. Chem., 1999, 3223–3235 CrossRef CAS.
- I. Banik, F. F. Becker and B. K. Banik, J. Med. Chem., 2003, 46, 12–15 CrossRef CAS PubMed.
- S. F. Martin, Pure Appl. Chem., 2009, 81, 195–204 CrossRef CAS PubMed.
- L. A. Joyce, E. C. Sherer and C. J. Welch, Chem. Sci., 2014, 5, 2855–2861 RSC.
- A. L. Berhanu, Gaurav, I. Mohiuddin, A. K. Malik, J. S. Aulakh, V. Kumar and K.-H. Kim, TrAC Trends Anal. Chem., 2019, 116, 74–91 CrossRef CAS.
- S. L. Pilicer, M. Mancinelli, A. Mazzanti and C. Wolf, Org. Biomol. Chem., 2019, 17, 6699–6705 RSC.
- R. Lopez and C. Palomo, Chemistry, 2021, 27, 20–29 CrossRef CAS PubMed.
- U. Krishnan and S. K. Iyer, Photochem. Photobiol., 2022, 98, 843–855 CrossRef CAS PubMed.
- X. Qu and Z. Yang, Chem. – Asian J., 2016, 11, 2633–2641 CrossRef CAS PubMed.
- S. J. Rowan, S. J. Cantrill, G. R. Cousins, J. K. Sanders and J. F. Stoddart, Angew. Chem., Int. Ed., 2002, 41, 898–952 CrossRef PubMed.
- S. Otto, J. Mater. Chem., 2005, 15, 3357–3361 RSC.
- L. You, Chem. Commun., 2023, 59, 12943–12958 RSC.
- N. Boehnke, C. Cam, E. Bat, T. Segura and H. D. Maynard, Biomacromolecules, 2015, 16, 2101–2108 CrossRef CAS PubMed.
- W. Dai, F. Shao, J. Szczerbiński, R. McCaffrey, R. Zenobi, Y. Jin, A. D. Schlüter and W. Zhang, Angew. Chem., Int. Ed., 2016, 55, 213–217 CrossRef CAS PubMed.
- H. Zhang, Z. Su and X. Wang, ACS Sustainable Chem. Eng., 2022, 10, 8650–8657 CrossRef CAS.
- P. Shen, B. Pollard, M. Ueland, D. Lawes and L. A. Connal, Macromol. Rapid Commun., 2025, 46, e2400683 CrossRef PubMed.
- C. A. Zentner, F. Anson, S. Thayumanavan and T. M. Swager, J. Am. Chem. Soc., 2019, 141, 18048–18055 CrossRef CAS PubMed.
- F. L. C. Morgan, I. A. O. Beeren, J. Bauer, L. Moroni and M. B. Baker, J. Am. Chem. Soc., 2024, 146, 27499–27516 CrossRef CAS PubMed.
- A. Chao, I. Negulescu and D. Zhang, Macromolecules, 2016, 49, 6277–6284 CrossRef CAS.
- F. García, J. Pelss, H. Zuilhof and M. M. J. Smulders, Chem. Commun., 2016, 52, 9059–9062 RSC.
- F. Haase, E. Troschke, G. Savasci, T. Banerjee, V. Duppel, S. Dörfler, M. M. J. Grundei, A. M. Burow, C. Ochsenfeld, S. Kaskel and B. V. Lotsch, Nat. Commun., 2018, 9, 2600 CrossRef PubMed.
- H. Yang, S. Ghiassinejad, E. van Ruymbeke and C.-A. Fustin, Macromolecules, 2020, 53, 6956–6967 CrossRef CAS.
- F. Song, Z. Li, P. Jia, M. Zhang, C. Bo, G. Feng, L. Hu and Y. Zhou, J. Mater. Chem. A, 2019, 7, 13400–13410 RSC.
- J. Sun, H. He, K. Zhao, W. Cheng, Y. Li, P. Zhang, S. Wan, Y. Liu, M. Wang, M. Li, Z. Wei, B. Li, Y. Zhang, C. Li, Y. Sun, J. Shen, J. Li, F. Wang, C. Ma, Y. Tian, J. Su, D. Chen, C. Fan, H. Zhang and K. Liu, Nat. Commun., 2023, 14, 5348 CrossRef CAS PubMed.
- R. J. Kearsey, A. Tarzia, M. A. Little, M. C. Brand, R. Clowes, K. E. Jelfs, A. I. Cooper and R. L. Greenaway, Chem. Commun., 2023, 59, 3731–3734 RSC.
- M. He and J.-M. Lehn, Chem. – Eur. J., 2021, 27, 7516–7524 CrossRef CAS PubMed.
- A. van Dam, R. van Schendel, S. Gangarapu, H. Zuilhof and M. M. J. Smulders, Chem. – Eur. J., 2023, 29, e202301795 CrossRef CAS PubMed.
- S. Ghosh, R. J. Butcher and R. Angamuthu, Inorg. Chim. Acta, 2024, 570, 122167 CrossRef CAS.
- H. H. Jo, R. Edupuganti, L. You, K. N. Dalby and E. V. Anslyn, Chem. Sci., 2015, 6, 158–164 RSC.
- Y. Zhou, Y. Yuan, L. You and E. V. Anslyn, Chem. – Eur. J., 2015, 21, 8207–8213 CrossRef CAS PubMed.
- C. Kachi-Terajima, T. Shimoyama, T. Ishigami, M. Ikeda and Y. Habata, Dalton Trans., 2018, 47, 2638–2645 RSC.
- S. Raje, N. Mondivagu, M. Chahal, R. J. Butcher and R. Angamuthu, Chem. – Asian J., 2018, 13, 1458–1466 CrossRef CAS PubMed.
- S. Purkait, P. Chakraborty, A. Frontera, A. Bauzá, E. Zangrando and D. Das, New J. Chem., 2018, 42, 12998–13009 RSC.
- J. P. Saucedo-Vázquez, V. M. Ugalde-Saldívar, A. R. Toscano, P. M. H. Kroneck and M. E. Sosa-Torres, Inorg. Chem., 2009, 48, 1214–1222 CrossRef PubMed.
- S. Raje, A. Koner, S. Ghosh, R. J. Butcher, L. Mathivathanan, M. Sundararajan and R. Angamuthu, Chem. – Asian J., 2023, 18, e202300706 CrossRef CAS PubMed.
- L. You, S. R. Long, V. M. Lynch and E. V. Anslyn, Chemistry, 2011, 17, 11017–11023 CrossRef CAS PubMed.
- M. Chahal, K. Mani, C. S. Lodhi, R. J. Butcher, S. Raje and R. Angamuthu, ChemistrySelect, 2018, 3, 9960–9964 CrossRef CAS.
- L. You, S. R. Long, V. M. Lynch and E. V. Anslyn, Chemistry, 2011, 17, 11017–11023 CrossRef CAS PubMed.
- Y. Zhou, Y. Ren, L. Zhang, L. You, Y. Yuan and E. V. Anslyn, Tetrahedron, 2015, 71, 3515–3521 CrossRef CAS.
- N. Arulsamy and D. J. Hodgson, Inorg. Chem., 1994, 33, 4531–4536 CrossRef CAS.
- H. Adams, M. S. Shongwe, I. Al-Bahri, E. Al-Busaidi and M. J. Morris, Acta Crystallogr., Sect. C:Cryst. Struct. Commun., 2005, 61, m497–m500 CrossRef PubMed.
- S. Ding, C. Xia, Y. Ji, Z. Liu and Y. Ding, Acta Crystallogr., Sect. C:Cryst. Struct. Commun., 2011, 67, m622–m623 CAS.
- S. K. Padhi, R. Sahu and V. Manivannan, Inorg. Chim. Acta, 2011, 367, 57–63 CrossRef CAS.
- S. Albano, G. Olivo, L. Mandolini, C. Massera, F. Ugozzoli and S. Di Stefano, J. Org. Chem., 2017, 82, 3820–3825 CrossRef CAS PubMed.
- C. E. Check, T. O. Faust, J. M. Bailey, B. J. Wright, T. M. Gilbert and L. S. Sunderlin, J. Phys. Chem. A, 2001, 105, 8111–8116 CrossRef CAS.
- S. Grimme, J. Antony, S. Ehrlich and H. Krieg, J. Chem. Phys., 2010, 132, 154104 CrossRef PubMed.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, G. A. Petersson, H. Nakatsuji, X. Li, M. Caricato, A. V. Marenich, J. Bloino, B. G. Janesko, R. Gomperts, B. Mennucci, H. P. Hratchian, J. V. Ortiz, A. F. Izmaylov, J. L. Sonnenberg, D. Williams-Young, F. Ding, F. Lipparini, F. Egidi, J. Goings, B. Peng, A. Petrone, T. Henderson, D. Ranasinghe, V. G. Zakrzewski, J. Gao, N. Rega, G. Zheng, W. Liang, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, K. Throssell, J. A. Montgomery Jr., J. E. Peralta, F. Ogliaro, M. J. Bearpark, J. J. Heyd, E. N. Brothers, K. N. Kudin, V. N. Staroverov, T. A. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. P. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, J. M. Millam, M. Klene, C. Adamo, R. Cammi, J. W. Ochterski, R. L. Martin, K. Morokuma, O. Farkas, J. B. Foresman and D. J. Fox, Gaussian 16 Rev. C.01, in Wallingford, CT, 2016 Search PubMed.
- N. G. Connelly, T. Damhus, R. M. Hartshorn and A. T. Hutton, Nomenclature of Inorganic Chemistry: IUPAC Recommendations 2005, IUPAC and the Royal Society of Chemistry, Cambridge, 2005 Search PubMed.
- M. Bröring, S. Prikhodovski, C. D. Brandt, E. Cónsul Tejero, S. Köhler and C. Pietzonka, Inorg. Chem., 2007, 46, 917–925 CrossRef PubMed.
- G. Bottomley, I. Clark, I. Creaser, L. Engelhardt, R. Geue, K. Hagen, J. Harrowfield, G. Lawrance, P. Lay, A. Sargeson, A. See, B. Skelton, A. White and F. Wilner, Aust. J. Chem., 1994, 47, 143–179 CrossRef CAS.
- P. Osvath, A. M. Sargeson, A. McAuley, R. E. Mendelez, S. Subramanian, M. J. Zaworotko and L. Broge, Inorg. Chem., 1999, 38, 3634–3643 CrossRef CAS PubMed.
- H. Ferreira, M. M. Conradie and J. Conradie, Electrochim. Acta, 2018, 292, 489–501 CrossRef CAS.
- J. N. McPherson, R. W. Hogue, F. S. Akogun, L. Bondi, E. T. Luis, J. R. Price, A. L. Garden, S. Brooker and S. B. Colbran, Inorg. Chem., 2019, 58, 2218–2228 CrossRef CAS PubMed.
- J. P. F. Rebolledo-Chávez, G. T. Toral, V. Ramírez-Delgado, Y. Reyes-Vidal, M. L. Jiménez-González, M. Cruz-Ramírez, A. Mendoza and L. Ortiz-Frade, Catalysts, 2021, 11, 948 CrossRef.
- J. Yang, M. Ansari, D. A. Pantazis, C. H. M. Zheng, R. McDonald, M. J. Ferguson and L. Rosenberg, Inorg. Chem., 2024, 63, 22409–22421 CrossRef CAS PubMed.
- L. Rocchetti, A. Amato and F. Beolchini, J. Clean. Prod., 2016, 116, 299–305 CrossRef CAS.
- J. P. Reddy, Acta Crystallogr. E, 2020, 76, 1771–1774 CAS.
- G.-B. Li, B.-Q. Song, S.-Q. Wang, L.-M. Pei, S.-G. Liu, J.-L. Song and Q.-Y. Yang, ACS Omega, 2019, 4, 1995–2000 CrossRef CAS PubMed.
- N. Podjed, J. Črepinšek and B. Modec, Molbank, 2021, 2021, M1216 CrossRef.
- P. C. Bell and J. D. Wallis, Chem. Commun., 1999, 257–258 RSC.
- A. Lari, M. B. Pitak, S. J. Coles, G. J. Rees, S. P. Day, M. E. Smith, J. V. Hanna and J. D. Wallis, Org. Biomol. Chem., 2012, 10, 7763–7779 RSC.
- J. O'Leary and J. D. Wallis, Org. Biomol. Chem., 2009, 7, 225–228 RSC.
- R. W. Newberry and R. T. Raines, Acc. Chem. Res., 2017, 50, 1838–1846 CrossRef CAS PubMed.
- H. B. B:urgi, J. D. Dunitz, J. M. Lehn and G. Wipff, Tetrahedron, 1974, 30, 1563–1572 CrossRef.
- H. B. Burgi, J. D. Dunitz and E. Shefter, J. Am. Chem. Soc., 1973, 95, 5065–5067 CrossRef CAS.
- H. Zheng, H. Ye, X. Yu and L. You, J. Am. Chem. Soc., 2019, 141, 8825–8833 CrossRef CAS PubMed.
- H. R. Kilgore, C. R. Olsson, K. A. D'Angelo, M. Movassaghi and R. T. Raines, J. Am. Chem. Soc., 2020, 142, 15107–15115 CrossRef CAS PubMed.
- K. B. Muchowska, D. J. Pascoe, S. Borsley, I. V. Smolyar, I. K. Mati, C. Adam, G. S. Nichol, K. B. Ling and S. L. Cockroft, Angew. Chem., Int. Ed., 2020, 59, 14602–14608 CrossRef CAS PubMed.
- (a) CCDC 2423656: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2mc0gh; (b) CCDC 2423657: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2mc0hj; (c) CCDC 2423658: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2mc0jk; (d) CCDC 2493551: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2ppr49; (e) CCDC 2493552: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2ppr5b; (f) CCDC 2493553: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2ppr6c; (g) CCDC 2493554: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2ppr7d; (h) CCDC 2493555: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2ppr8f; (i) CCDC 2493556: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2ppr9g; (j) CCDC 2493557: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2pprbh; (k) CCDC 2493743: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2ppybp.
Footnote |
| † Current affiliation: Department of Chemistry, National Taiwan Normal University, Taipei 11677, Taiwan. |
| This journal is © The Royal Society of Chemistry 2026 |