Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
CReM-dock: de novo design of synthetically feasible structures guided by molecular docking
Guzel
Minibaeva
 a,
Haolin
Du
a,
Haolin
Du
 b,
Finlay
Clark
b,
Finlay
Clark
 c,
Julien
Michel
c,
Julien
Michel
 b and
Pavel
Polishchuk
b and
Pavel
Polishchuk
 *a
*a
aInstitute of Molecular and Translational Medicine, Faculty of Medicine and Dentistry, Palacky University, Hněvotínská 1333/5, 779 00 Olomouc, Czech Republic. E-mail: pavlo.polishchuk@upol.cz
bEaStCHEM School of Chemistry, University of Edinburgh, Edinburgh, EH9 3FJ, UK
cSchool of Natural and Environmental Sciences, Newcastle University, Newcastle upon Tyne, NE1 7RU, UK
First published on 28th April 2026
Abstract
The de novo generation of chemical compounds represents a compelling strategy for the exploration of a significantly broader chemical space compared to traditional virtual screening methods. However, fragment-based approaches often encounter challenges related to the low synthetic accessibility of generated structures. In this study, we integrated the previously established fragment-based structure generator, CReM, with molecular docking techniques implemented in EasyDock to facilitate the exploration of chemical space. This novel methodology enables indirect control over the synthetic accessibility and diversity of the generated structures, enhances an objective function to yield compounds with desirable physicochemical properties, and maintains key protein–ligand interactions and ligand conformations. The structures generated through this approach exhibited a high degree of novelty and demonstrated competitiveness with those produced by leading methodologies in the field, as demonstrated by Absolute Binding Free Energy calculations. We illustrated the versatility of the developed approach through several case studies, showcasing its applicability to both de novo generation and fragment expansion tasks. The tool developed in this study is open-source and can be accessed at https://github.com/ci-lab-cz/crem-dock.
Introduction
The vastness of drug-like chemical space, estimated to encompass approximately 1036 compounds, makes it impractical to fully enumerate or even to sample a representative subset for the purpose of virtual screening.1De novo design offers a promising approach to identify novel chemical entities within this extensive chemical space by exploring only a small fraction of it. This method integrates a structure generation tool with an objective function that is optimized during the exploration of the chemical space, directing the generator towards the most promising regions.2,3 Currently, there exist various methodologies for the generation of molecular structures from first principles.4,5 However, a significant limitation of many structure-generation techniques is the limited synthetic accessibility of the molecules generated.6 Additionally, the choice of objective function employed in the search for promising molecular structures can impose further constraints. The use of machine learning models to guide chemical space exploration introduces challenges associated with their limited applicability domain. In particular, such models often perform poorly when predicting properties of molecules that differ substantially from those represented in their training data,7 thereby constraining the practically accessible search space for de novo design.8 In contrast, molecular docking offers broader applicability and is less biased towards specific chemotypes or structural motifs,9–11 making docking scores a potentially more suitable objective function for the identification of novel promising compounds. However, docking is still a proxy for binding affinity and has its own set of challenges. The main issues arise from a lack of consideration for protein flexibility (known as induced fit due to ligand binding), the neglect of explicit water molecules that may influence protein–ligand interactions, the difficulty in assessing ligand flexibility, which can lead to inaccurate binding entropy estimates, and challenges in accurately estimating solvation/desolvation factors.9,10,12,13 Such effects can be accounted for using free energy methods such as alchemical absolute binding free energy calculations,14,15 but their high associated computational cost restricts their use to a late-stage filter in de novo design pipelines. Therefore, in spite of these constraints, docking has proven valuable in identifying promising candidates when exploring extremely large chemical libraries.16–18Numerous approaches combine de novo structure generation with molecular docking to identify novel bioactive compounds. Reaction-based methodologies enumerate molecules using predefined reaction rules to improve synthetic plausibility. For example, AutoCouple19 generated ∼70![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ligands for the CBP bromodomain from three reactions, leading to 53 synthesized compounds and a highly potent and selective inhibitor (IC50 = 35 nM). Similarly, Chevillard et al.20 expanded fragment hits for the β2-adrenergic receptor via reaction-based linking, yielding compounds with up to 40-fold affinity improvements. Other tools such as NAOMInext21 and AutoGrow4
000 ligands for the CBP bromodomain from three reactions, leading to 53 synthesized compounds and a highly potent and selective inhibitor (IC50 = 35 nM). Similarly, Chevillard et al.20 expanded fragment hits for the β2-adrenergic receptor via reaction-based linking, yielding compounds with up to 40-fold affinity improvements. Other tools such as NAOMInext21 and AutoGrow4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 22 iteratively grow or recombine molecules using reaction rules and docking-based evaluation, with AutoGrow4 demonstrating docking scores exceeding those of known ligands. However, while reaction-based approaches are intended to enhance the likelihood of synthetic accessibility of the designed structures, they do not guarantee it. For example, the top-scoring structures generated by AutoGrow4 have been found to lack synthetic feasibility.22 Another limitation pertains to the coverage of chemical space, which depends on the selected reaction rules and the libraries of building blocks.
22 iteratively grow or recombine molecules using reaction rules and docking-based evaluation, with AutoGrow4 demonstrating docking scores exceeding those of known ligands. However, while reaction-based approaches are intended to enhance the likelihood of synthetic accessibility of the designed structures, they do not guarantee it. For example, the top-scoring structures generated by AutoGrow4 have been found to lack synthetic feasibility.22 Another limitation pertains to the coverage of chemical space, which depends on the selected reaction rules and the libraries of building blocks.
Fragment-based methods offer broader chemical space exploration but often at the cost of reduced synthetic accessibility. Strategies to mitigate this include post hoc filtering or explicit biasing during optimization. LigBuilder23 applies retrosynthetic filtering after fragment growth, whereas Graph-GA24 integrates a synthetic accessibility (SA)25 score directly into the objective function, markedly increasing the proportion of synthetically feasible top-ranked compounds. Other tools, such as OpenGrowth26 and FragExplorer,27 introduce implicit synthetic bias by restricting allowable fragment connections based on observed chemical patterns. While these constraints improve chemical plausibility, they provide limited control and do not consistently yield drug-like or synthetically accessible molecules.
Machine learning-based generative models represent a third major category. OptiMol28 and SampleDock29 employ variational autoencoders guided by iterative docking, while DockStream30 provides a docking wrapper that integrates with recurrent neural network-based reinforcement learning frameworks such as REINVENT SBMolGen31 combines recurrent neural networks with Monte Carlo tree search and docking. These approaches can generate molecules with docking scores comparable to or exceeding those of known actives and often maintain reasonable drug-likeness and SA scores. Nevertheless, issues such as declining novelty, limited explicit control over synthetic feasibility, and high computational cost remain. Reaction-aware neural models, such as SynFlowNet32 and RGFN,33 further integrate docking and retrosynthesis signals but are computationally demanding and tend to generate highly aromatic structures with suboptimal drug-likeness and limited capacity for further optimization.
Overall, fragment-based and machine learning-driven approaches provide superior chemical space coverage relative to reaction-based methods, but synthetic feasibility remains a central challenge. Synthetic biasing can be introduced explicitly through objective functions or implicitly through structural constraints; however, explicit methods require careful scoring and weighting, while implicit strategies are often overly simplistic and lack fine-grained control.
Previously, we introduced CReM,34 a fragment-based framework that indirectly biases generation toward synthetically accessible molecules by restricting bond formation to chemical contexts observed in synthesized compounds. CReM supports fragment growth, mutation, and linking, and its synthetic relevance has been validated using SA scores and retrosynthetic analysis.35 Building on CReM, Moldrug36 was developed as a structure-based optimization tool for hit-to-lead design. Moldrug employs a genetic algorithm to optimize docking, drug-likeness, and synthetic accessibility. While Moldrug supports multi-objective optimization and interactive analysis, it is currently limited to AutoDock Vina and does not incorporate protein–ligand interaction fingerprints into its objective function; however, it may be extended with additional objectives.
In this study, we developed and implemented a de novo generation pipeline, named CReM-dock, which employs a fragment growing strategy informed by the CReM generative approach34 and is guided by molecular docking techniques integrated within the EasyDock module.37 The development of EasyDock was largely driven by the lack of accessible, fully automated open-source docking pipelines capable of seamless integration with third-party software. Consequently, its development proceeded concurrently with that of CReM-dock. The modular architecture of CReM-dock facilitates the modification and enhancement of individual components. While CReM offers fine-tuning control over the synthetic accessibility of the generated molecules,35 the integration of EasyDock provides a unified interface for a fully automated docking pipeline, encompassing ligand preparation and various molecular docking methodologies. This integration significantly streamlines the execution, processing, and analysis of docking results. Currently, EasyDock is compatible with AutoDock Vina,38 Smina,39 Gnina40 programs and other derivatives of Vina: QVina2,41 QVina-W42 and their GPU implementations.43 Additional program support can be incorporated into EasyDock, thereby making them readily accessible to CReM-dock.
Current de novo design pipelines guided by molecular docking often incorporate proprietary components, limiting their accessibility to the broader research community. Among these, protonation tools are the most commonly used proprietary modules within existing docking workflows. While some pipelines neglect ligand protonation altogether, others employ OpenBabel, which utilizes a simple rule-based approach. To address these limitations, we have integrated multiple open-source protonation tools into EasyDock and validated them. Notably, the protonation tool based on the Uni-pKa model44 exhibited the highest agreement with the proprietary software from Chemaxon. The incorporation of open-source protonation tools has made the entire CReM-dock pipeline fully open-source and freely accessible.
CReM-dock is capable of addressing two distinct scenarios: (i) fragment expansion or scaffold decoration, and (ii) de novo design. In the first scenario, the generation process commences from a defined pose of a seed molecule or fragment, which may be derived from the X-ray structure of a protein–ligand complex or obtained through alternative computational modelling techniques, such as molecular dynamics simulations. In the second scenario, the initial fragments can be sourced from any existing library of fragments or building blocks. These fragments undergo sequential expansion, with the most promising candidates advancing to subsequent iterations. A notable feature of CReM-dock is its ability to specify important protein–ligand interactions that must be preserved throughout the generation process, as well as to maintain the coordinates of the seed fragment in the resulting structures, which is particularly pertinent in the context of the fragment expansion scenario.
Methods
In alignment with established methodologies, our approach utilizes an iterative strategy for exploring chemical space through fragment growth, which encompasses several stages (Fig. 1): (i) the docking of initial fragments or molecules, (ii) the selection of compounds exhibiting promising scores and meeting a user-defined protein–ligand interaction fingerprint (the latter being optional), (iii) the growth of the selected molecules, and (iv) the filtering of molecules based on their physicochemical properties. This process is reiterated until the physicochemical properties of the compounds exceed predetermined user-defined thresholds.There are two distinct modes of structure generation: de novo design and fragment/molecule expansion. The de novo design mode requires a set of initial fragments provided in the form of SMILES or 2D structures. The fragment/molecule expansion mode requires 3D structures as input, corresponding to either empirically observed or predicted binding conformations. The primary distinction between these modes lies in the fact that 3D input molecules are directly subjected to the growth phase, thereby bypassing the docking and selection stages.
The implemented generative pipeline consists exclusively of open-source modules and can therefore be used without additional restrictions. This eliminates limitations commonly associated with the ligand-preparation step, for which proprietary software is often employed for tasks such as protonation-state assignment.
Molecular docking
Molecular docking is conducted utilizing EasyDock,37 which presently accommodates AutoDock Vina, Gnina, and multiple tools from the Vina family: QVina2, QVina-W, and their corresponding GPU implementations. EasyDock offers a unified interface for the preparation and docking of molecules, as well as for the integration of docking processes within external applications. The input for this software includes a protein structure and a configuration file that defines all necessary parameters, such as grid box coordinates and dimensions, the level of search exhaustiveness. We employed the same database architecture as EasyDock to store all outputs from the design and docking phases, along with the characteristics of the generated structures.A major advance in the development of CReM-dock was the incorporation of several open-source tools for ligand protonation, thereby rendering the entire design pipeline fully open-source. Specifically, we integrated and evaluated the MolGpKa45 and Uni-pKa44 models, comparing their predicted protonation states at pH 7.4 against those generated by the Chemaxon cxcalc utility. This comparison was conducted using molecular structures from the ChEMBL database, restricted to compounds with molecular weights below 500 Da. The concordance between Chemaxon predictions and those of MolGpKa, Uni-pKa, and OpenBabel was 78%, 82%, and 70%, respectively. Based on these results, the Uni-pKa protonation tool was selected for all subsequent experiments in this study.
Implemented objective functions
Structures are ranked by their docking scores by default. However, to enhance the balance and alignment of the generated structures with the desired property space, the docking scores can be supplemented with additional properties. Specifically, we incorporated the quantitative estimate of drug-likeness (QED)46 into the docking scores. Initially, docking scores are normalized to a range of 0 to 1 using the formula s = (x − xmin)/(xmax − xmin), where x represents the docking score of a molecule, and xmin and xmax denote the minimum and maximum docking scores among the compounds generated during a given iteration that are eligible for selection. Subsequently, the scaled docking scores are multiplied by the corresponding QED values to yield a final score.Additionally, we implemented an augmentation involving the fraction of sp3 carbon atoms within Bemis–Murcko scaffolds (Csp3BM). The Csp3BM values, which range from 0 to 0.3, were linearly scaled to a range of 0 to 1, while values above 0.3 were capped at 1. The scaled Csp3BM values were squared to increase the selection pressure associated with this parameter and subsequently multiplied by the previously normalized docking score. The threshold of 0.3 was chosen based on recommendations from the authors of the initial CACHE challenge,47 in which our participation resulted in one of top-ranking positions.48 Other objective functions were also considered, such as those based on docking efficiency (calculated as the docking score divided by the number of heavy atoms); however, these were not utilized in the studies presented herein.
Structure selection strategies
We employed three distinct strategies for molecule selection during each iteration: greedy, Pareto, and clustering-based selection. The first strategy consists of selecting the top N molecules based solely on their objective scores. Although this approach may yield compounds with high scores, it may also reduce the diversity of the selected molecules. To address this limitation, we introduced two additional protocols. The first alternative utilizes Pareto ranking to identify molecules on the Pareto front that exhibit low molecular mass and high objective scores, thereby fostering the development of promising low molecular mass candidates. The choice of molecular weight as a selection criterion was motivated by the objective of prioritizing smaller molecular structures that possess the potential for further expansion in subsequent iterations. The second alternative employs a K-means clustering method to categorize structures into a user-defined number of clusters based on 2048-bit Morgan fingerprints with a radius of 2. For each cluster, a specified number of top-scoring structures are selected. In clusters where top-scoring structures are unsuitable for further development (e.g., due to the absence of hydrogen atoms for growth), the next highest-scoring structure is chosen until the desired number of molecules is obtained from that cluster. The clustering protocol offers a more predictable number of selected structures and runtime, whereas the Pareto approach yields a more variable number of generated structures, resulting in greater diversity in the final solutions.Additional selection protocols were also implemented, but they were not used in the present study because they did not provide a clear advantage. One such approach is based on Pareto ranking with respect to docking and SA scores; however, in test runs this strategy led to a substantial decrease in docking scores while it did not improve SA scores sufficiently to outperform other selection strategies. Another approach clusters structures according to their scaffolds, rather than whole structures. Although this was expected to increase scaffold diversity in the output set, only a minor effect was observed experimentally.
Users have the option to specify additional criteria prior to the actual selection process. One such criterion is the root-mean-square deviation (RMSD) threshold, which serves to maintain the pose of a parent structure in its derivative structures. If the docking poses of the generated compounds differ from their parent molecule by more than the established RMSD threshold, those compounds will be excluded from consideration. The RMSD value is computed based on the heavy atoms of the maximum common substructure between the parent and successor molecules. This approach is particularly advantageous in fragment expansion studies, where the conformation of the initial fragment is determined experimentally, and it is anticipated that the successor structures will retain this conformation.
Additionally, users may provide a list of important protein–ligand interactions and establish a minimum similarity threshold based on protein–ligand interaction fingerprints. These fingerprints, calculated using ProLIF,49 encompass hydrogen bond donors and acceptors, hydrophobic interactions, aromatic contacts, and charged interactions, as well as metal coordination sites. Compounds that do not meet the specified similarity threshold are excluded from the selection process. For instance, if three contacts are specified and a threshold of 0.5 is set, this implies that at least two of these contacts must be present in the candidate structures (resulting in a similarity score of 0.66); otherwise, they will not qualify for selection. This methodology is particularly beneficial when important contacts are known in advance, ensuring that the designed molecules retain these essential interactions.
Molecule growing
The selected molecules undergo a growth process facilitated by the CReM approach.34 In this process, hydrogen atoms are substituted with larger molecular fragments sourced from a database of interchangeable fragments, referred to as CReM databases. These fragments are derived from existing molecules through a comprehensive fragmentation process that involves cleaving up to four single bonds. For each fragment, a chemical context is established, defined as a substructure that includes atoms within a specified number of bonds (context radius) from the attachment points of the fragment. Consequently, fragments that occur within the same chemical context are deemed interchangeable, and their mutual substitution is expected to yield synthetically accessible molecules. Prior research has indicated that synthetic feasibility can be indirectly controlled by selecting a larger context radius and utilizing CReM databases comprising fragments from more synthetically accessible molecules.35During the growth process, only hydrogen atoms that are situated at least 2 Å away from any heavy atom of the protein are replaced. This precaution is taken to prevent growth in spatial directions that lack sufficient volume to accommodate larger fragments. To regulate the number of generated compounds and enhance the predictability of runtime, a maximum number of randomly selected replacements can be specified. In the absence of such a specification, all potential replacements will be executed, which may result in a substantial number of structures, particularly for smaller context radii. Our findings suggest that employing 1000 random replacements yields satisfactory results, and this value has been consistently utilized in all reported studies. Furthermore, the size of the steps within chemical space can be adjusted; by default, fragments containing up to 10 heavy atoms are attached. Given that CReM cannot create new ring systems, it is essential to select a sufficiently large fragment size to allow for the incorporation of rings into the molecules. The ten-atom limit permits the addition of bicyclic fragments that include five- or six-membered rings.
By default, the fragment selection process in CReM utilizes a uniform distribution. However, to impose selective pressure and prioritize fragments with more favourable properties, we incorporated customizable selection functions within CReM, enabling fragment selection based on user-defined criteria. In one of the studies presented in this work, fragment selection was weighted in proportion to the square of the fraction of sp3 carbon atoms within the fragments. This approach is intended to preferentially select fragments with a higher fraction of saturated carbon atoms, thereby enriching the generated structures in sp3 carbon atoms in their frameworks.
We have implemented controls over important physicochemical properties that influence drug-likeness, including molecular weight (MW), topological polar surface area (TPSA), lipophilicity (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P), and the number of rotatable bonds (RTB), to ensure that the generation process predominantly yields drug-like molecules. All these parameters, except for lipophilicity, either increase or remain constant as the number of atoms in a molecule increases and are primarily additive in nature. If any of these parameters reaches or exceed a predetermined threshold, the corresponding molecule is excluded from further consideration. Due to the predominantly additive nature of these properties, we can pre-filter CReM fragments in real time, selecting those that are unlikely to produce structures that exceed the predefined thresholds for physicochemical properties. This strategy effectively circumvents the need to enumerate and dock of structures with undesirable properties.
P), and the number of rotatable bonds (RTB), to ensure that the generation process predominantly yields drug-like molecules. All these parameters, except for lipophilicity, either increase or remain constant as the number of atoms in a molecule increases and are primarily additive in nature. If any of these parameters reaches or exceed a predetermined threshold, the corresponding molecule is excluded from further consideration. Due to the predominantly additive nature of these properties, we can pre-filter CReM fragments in real time, selecting those that are unlikely to produce structures that exceed the predefined thresholds for physicochemical properties. This strategy effectively circumvents the need to enumerate and dock of structures with undesirable properties.
In all studies described below, specific constraints were imposed on the physicochemical properties of the generated structures. These constraints included a molecular mass not exceeding 450 Da, a maximum of five rotatable bonds, a lipophilicity value of no more than 4, and a topological polar surface area limited to 120 Å2. These parameters are consistent with Lipinski's rule and allow for potential further optimization of the generated structures. The deliberate restriction on the number of rotatable bonds is intended to reduce the flexibility of the designed structures, thereby improving the accuracy of docking simulations.50,51
Protein preparation protocol
Receptor structures were prepared utilizing the DockPrep protocol available in Chimera.52 This preparation process included the remodeling of absent side chains and sequences through the application of the Dunbrack rotamer library53 and MODELLER,54 respectively. Hydrogen atoms were added, taking into account a pH of 7.4, while solvent molecules were removed. Subsequently, the structures were converted to the PDBQT format using the prepare_receptor4.py utility from AutoDock Tools. The dimensions of the grid boxes for docking were established based on the coordinates of the co-crystallized ligands. Specifically, the center of each grid box was determined as the geometric center of a ligand, with the box size defined by extending 7 Å beyond the minimum and maximum coordinates of the ligand's heavy atoms. All prepared structures and grid box parameters were deposited in the repository.55Novelty assessment
To evaluate the novelty of the generated structures, we calculated Tanimoto similarity to the nearest compound within a dataset of known compounds sourced from ChEMBL (version 33). The similarity assessment was conducted using the chemfp tool56 and employed 2048-bit Morgan fingerprints with a radius of 2. Lower Tanimoto similarity values indicate a higher degree of novelty for the generated structures. To establish a baseline for similarity, we utilized randomly selected 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 compounds from ChEMBL with a molecular weight of 500 Da or less and computed pairwise similarities among them. The results indicated a mean similarity of 0.105, with the 95th percentile at 0.178 and the 99th percentile at 0.230.
000 compounds from ChEMBL with a molecular weight of 500 Da or less and computed pairwise similarities among them. The results indicated a mean similarity of 0.105, with the 95th percentile at 0.178 and the 99th percentile at 0.230.
UMAP and reference space
To examine the distribution of generated structures in relation to reference compounds, we selected a baseline reference set comprising 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 random compounds with a molecular weight of 500 Da or less from ChEMBL33. Additionally, we compiled distinct sets of active compounds for each individual target. These active compounds were selected from those evaluated in a single protein assay format and exhibited pIC50, pKi, or pKd values of 6 or higher. For the purpose of visualizing the chemical space, we employed UMAP57 (utilizing the umap-learn Python package) alongside 2048-bit Morgan fingerprints with a radius of 3. All parameters were maintained at their default settings, except for the number of neighbours, which was set to 10, and the metric, which was specified as “jaccard”.
000 random compounds with a molecular weight of 500 Da or less from ChEMBL33. Additionally, we compiled distinct sets of active compounds for each individual target. These active compounds were selected from those evaluated in a single protein assay format and exhibited pIC50, pKi, or pKd values of 6 or higher. For the purpose of visualizing the chemical space, we employed UMAP57 (utilizing the umap-learn Python package) alongside 2048-bit Morgan fingerprints with a radius of 3. All parameters were maintained at their default settings, except for the number of neighbours, which was set to 10, and the metric, which was specified as “jaccard”.
Absolute binding free energy calculations (ABFE)
To assess the likelihood that the designed compounds would be potent binders we use alchemical Absolute Binding Free Energy calculations to estimate their free energy of binding using the software A3FE.58,59Subsequent solvation, energy minimization, and pre-equilibration steps were performed automatically within A3FE. Specifically, the bound complexes and free systems were solvated with TIP3P water molecules in a 0.15 M NaCl solution in rhombic dodecahedral boxes, with a minimum distance of 15 Å maintained between the solute(s) and the box edge. All simulations were conducted using GROMACS 2024.4-foss-2023b-CUDA-12.4.0 via BioSimSpace (version 2024.4.0).62 Energy minimization was performed for 1000 steps, followed by several stages of NVT equilibration: 5 ps with all nonsolvent atoms positionally restrained while heating from 0 to 298 K, 50 ps with position restraints applied to all backbone atoms for complexes only, and finally a further 50 ps with no restraints. Subsequently, NPT equilibration was carried out at 1 atm and 298 K for a duration of 50 ps with positional restraints on non-solvent heavy atoms, then followed by a 50 ps simulation without positional restraints. For each system, independent 100 ps NPT equilibration runs were performed for every replicate to generate diverse starting conformations. A total of 5 replicates per system were generated. All positional restraints employed a force constant of 10 kcal mol−1 Å−2. The V-rescale and C-rescale algorithms were used for temperature and pressure coupling, with coupling constants of 1 ps and 4 ps, respectively.63,64 All equilibration molecular dynamics simulations were performed with a 2 fs time step and an 10 Å cutoff for short-range interactions, with long-range electrostatics calculated using the PME algorithm.65,66 After equilibration, the trajectory of the first replicate was analysed with BioSimSpace to determine the optimal Boresch restraints. The BioSimSpace restraints generator algorithm evaluates fluctuations for selecting the stable anchor points (three atoms in receptor and three atoms in ligand) for restraining the configurational volume.67 This approach emulated the effect of native interactions by ensuring the ligand remaining localized within the binding site upon decoupling. To ensure setup consistency across replicates, the same restraints were applied across all replicate simulations. Different starting conformations for each leg of every repeat were obtained from the last frame of 100 ps equilibration.
Simulations were carried out with five replicates, each initiated with independently assigned starting velocities and starting coordinates from the equilibration stage. MBAR was used to estimate free energy changes.71,72 Free energies were reported as the mean of the five replicates. Statistical uncertainties were obtained from the 95% confidence interval, assuming Gaussian distribution and employing values to four degrees of freedom. Statistical significance was evaluated by using Student's t-test. The total sampling time allocated to each ligand was 260 ns.
Results
The results section is organized into several subsections. The initial subsection details the development of CReM fragment databases, which serve as the foundational fragments for de novo design tasks, along with a theoretical assessment of the number of structures that can be generated by applying the CReM grow operator to these fragment sets.The subsequent subsection presents a de novo investigation utilizing CDK2 as the target protein. This analysis examines the influence of various parameters on the docking and SA scores of the generated structures, as well as their novelty and the reproducibility of results across independent runs. From these investigations, we identified optimal settings that were subsequently employed in all further experiments. Additional experiments focused on: (i) the impact of different selection strategies on the diversity of the generated structures; (ii) methods that may skew generation towards structures enriched with sp3 carbon atoms. Ultimately, we compared the results of the CReM-dock pipeline with those of traditional virtual screening methods.
In the third subsection, we demonstrate the consistency in the quality of the generated structures across various proteins from different families, utilizing the previously established settings, and compare of these results with the outputs of state-of-the-art approaches. A separate section is devoted to the analysis of calculated absolute binding free energies for the top-scoring structures obtained using different approaches, in order to demonstrate the utility of the developed pipeline in comparison with state-of-the-art approaches. The final subsection presents a fragment expansion study aimed at retrospectively assessing the capability to generate structures that are identical or highly similar to those previously discovered, starting from a small fragment co-crystallized with a protein.
Preparation of fragment databases and estimation of the size of accessible chemical space
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 554
554![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 260 structures. Subsequently, we refined this subset by excluding molecules that matched to at least one structural alert from the BMS, Dundee, Glaxo, Inpharmatica, and PAINS filters, as implemented by Pat Walters.75 Our previous research has demonstrated that the exclusion of such molecules prior to the creation of the fragment database ensures the generation of molecules devoid of these patterns, provided that the size of the alert patterns does not exceed the designated context radius.34 This process reduced the dataset to 818
260 structures. Subsequently, we refined this subset by excluding molecules that matched to at least one structural alert from the BMS, Dundee, Glaxo, Inpharmatica, and PAINS filters, as implemented by Pat Walters.75 Our previous research has demonstrated that the exclusion of such molecules prior to the creation of the fragment database ensures the generation of molecules devoid of these patterns, provided that the size of the alert patterns does not exceed the designated context radius.34 This process reduced the dataset to 818![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 174 molecules. Furthermore, we identified subsets of molecules characterized by restricted synthetic accessibility (SA) values, as predicted by the methodology of Ertl and Schuffenhauer.25 We established thresholds of 2 and 2.5, which are below the average (3.0) and the median SA score (2.73) for all compounds in ChEMBL22. This resulted in subsets containing 67
174 molecules. Furthermore, we identified subsets of molecules characterized by restricted synthetic accessibility (SA) values, as predicted by the methodology of Ertl and Schuffenhauer.25 We established thresholds of 2 and 2.5, which are below the average (3.0) and the median SA score (2.73) for all compounds in ChEMBL22. This resulted in subsets containing 67![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 970 and 338
970 and 338![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 422 molecules, respectively. These sets were then subjected to exhaustive fragmentation and subsequently converted into CReM fragment databases (Table 1).
422 molecules, respectively. These sets were then subjected to exhaustive fragmentation and subsequently converted into CReM fragment databases (Table 1).
| CReM DB | n (fragmented molecules) | n (distinct fragments) | Number of distinct fragment/context pairs for each radius | ||||
|---|---|---|---|---|---|---|---|
| Radius 1 | Radius 2 | Radius 3 | Radius 4 | Radius 5 | |||
| ChEMBL | 818![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 174 174 |
988![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 585 585 |
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 263 263![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 436 436 |
4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 051 051![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 790 790 |
7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 133 133![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 534 534 |
11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 007 247 007 247 |
15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 271 543 271 543 |
| ChEMBL SA2.5 (SA ≤ 2.5) | 338![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 422 422 |
272![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 988 988 |
671![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 140 140 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 263 263![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 268 268 |
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 319 319![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 377 377 |
3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 752 752![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 375 375 |
5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 419 419![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 544 544 |
| ChEMBL SA2 (SA ≤ 2) | 67![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 970 970 |
55![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 498 498 |
143![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 434 434 |
267![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 156 156 |
472![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 126 126 |
754![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 905 905 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 087 087![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 492 492 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 970 ChEMBL molecules that exhibited a SA score of less than 2. All attachment points were capped with hydrogen atoms, and the resulting molecules were subsequently converted into canonical SMILES format, with duplicates removed in the process. Following this, we applied a filtering process based on the following physicochemical properties of the resulting 200
970 ChEMBL molecules that exhibited a SA score of less than 2. All attachment points were capped with hydrogen atoms, and the resulting molecules were subsequently converted into canonical SMILES format, with duplicates removed in the process. Following this, we applied a filtering process based on the following physicochemical properties of the resulting 200![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 fragments, which yielded a final set of 20
000 fragments, which yielded a final set of 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 164 structures:
164 structures:
• The number of heavy atoms was within the range 8–15.
• The number of distinct H-bond donors and acceptors was within the range 1–5. If an atom was labeled as an H-bond donor and an acceptor it was counted only once. This gives an estimate on the number of specific contacts.
• The number of rings was 1–3.
• The number of fused ring systems was 0–2.
• The number of rotatable bonds was 0–2.
• Lipophilicity was less than 2.
• Topological polar surface area (TPSA) was greater than 25 Å2
• The total number of halogen atoms (Cl, Br, and I) was 0–1.
• The maximum size of rings was 7.
For this subset, we enumerated all stereoisomers utilizing an RDKit script sourced from the repository,76 as well as tautomers using the cxcalc utility from Chemaxon.77 A check for duplicates was conducted, yielding a final count of 23![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 840 structures, which were designated as starting fragments for de novo generation.
840 structures, which were designated as starting fragments for de novo generation.
The chemical environment for each non-equivalent hydrogen atom in the initial molecular fragment was identified utilizing CReM. We analyzed all possible combinations of four hydrogen atoms. However, by restricting the analysis to non-equivalent hydrogens, we underestimated the potential number of derivatives, as this approach excludes scenarios where substituents are attached to the same methyl group, for example. This simplification is a consequence of the constraints inherent in the current implementation of CReM.
For each combination of four hydrogens, the total number of potentially enumerated compounds was calculated as the product of the number of substituents available at each hydrogen position, adhering to the constraint of a maximum total number of heavy atoms in the generated structures. The overall number of molecules was determined by aggregating the number of enumerated structures across all combinations of four substituents. To improve computational efficiency, calculations were conducted on a subset of 1000 randomly selected molecules, and the final estimate was extrapolated by applying a scaling factor of 23.84 (with a total of 23![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 840 starting fragments). This estimate was predicated on the assumption that substituents would only be attached to the initial starting fragment.
840 starting fragments). This estimate was predicated on the assumption that substituents would only be attached to the initial starting fragment.
However, it is important to note that substituents can also be attached to previously introduced substituents. To account for this, the molecular structure was considered as a tree, where the five fragments (the starting fragment and four substituents) represent nodes. The possible number of connection combinations between these nodes was estimated using Cayley's formula, nn−2, which indicates that five nodes can be interconnected in 125 distinct configurations. This methodology introduces an additional simplification, resulting in an overestimation of the number of derivatives, as not all linkage combinations are chemically feasible due to chemical context constraints. These approximations may offset one another, yielding a rough estimate of the order of magnitude for the size of the chemical space covered.
A radius of 3 was selected as it represents a sufficiently large default value to produce synthetically feasible structures. For the largest CReM fragment database, the estimated coverage of chemical space was 1017 (Table 2). This coverage diminished when utilizing more restricted fragment databases: 1016 structures for ChEMBL SA2.5 and 1015 for ChEMBL SA2. For the smallest fragment database (ChEMBL SA2), the chemical space size was also assessed for radii of 2 and 4, resulting in predictable variations. The coverage increased to 1016 structures for a radius of 2 and decreased to 1013 for a radius of 4 (Table 2). These results suggest that even under highly constrained conditions, the covered chemical space remains considerable.
| CReM DB | Radius | Estimated size of covered chemical space |
|---|---|---|
| ChEMBL | 3 | 2.8 × 1017 |
| ChEMBL SA2.5 | 3 | 4.2 × 1016 |
| ChEMBL SA2 | 3 | 1.8 × 1015 |
| ChEMBL SA2 | 2 | 8.4 × 1016 |
| ChEMBL SA2 | 4 | 2.7 × 1013 |
Study of settings and features of CReM-dock in the case of de novo design of CDK2 inhibitors
In order to examine the impact of various settings on generation output, we selected CDK2 kinase as our target due to its clinical significance, the availability of multiple X-ray protein–ligand complexes, and its frequent utilization in the validation of modelling methodologies. We utilized the CDK2 structure (PDB 2BTR) in complex with the inhibitor PNU-198873 (Ki = 95 nM). This ligand establishes both hydrogen bond donor and acceptor interactions with the Leu83 residue located in the hinge region. Given the importance of hinge region interactions for competitive kinase inhibitors, we considered these two contacts (protein–ligand interaction pattern, PLIP) as essential for all designed molecules. Clustering was employed as the search algorithm: structures were divided into 25 clusters, from which the top two molecules were selected per cluster, resulting in the selection of up to 50 molecules for further growth in each iteration. Each compound was expanded to yield up to 1000 new molecules. In instances where the number of potential expansions exceeded 1000, a random selection of 1000 expansions was drawn from the CReM database. Simulations were conducted across all three CReM databases and five context radii (1–5). Each simulation was executed three times to assess the robustness of the search, as stochastic variability in the selection of growing fragments could influence the outcomes. For docking procedures, a consistent seed was employed, ensuring deterministic results. The overall number of generated structures demonstrated high reproducibility across different runs and exhibited a predictable decline when more restrictive fragment databases and larger radii were utilized (Fig. S1).Furthermore, we assessed the diversity of the top 100 generated structures based on Murcko scaffolds. The average number of distinct Murcko scaffolds was relatively low (ranging from 16 to 24) for smaller context radii of 1 to 3, irrespective of the fragment database (Fig. S2b). This observation suggests that under these conditions, certain scaffolds may yield multiple successful derivatives that outperform others. Conversely, for larger context radii of 4 and 5, the average number of distinct scaffolds increased significantly, ranging from 35 to 57 out of a maximum of 100. The impact of larger radii was particularly pronounced when utilizing more restricted fragment databases. This phenomenon can be attributed to the limited number of expansions available for each structure during each iteration, resulting in a reduced number of successor molecules generated per scaffold, thereby decreasing the likelihood of any single scaffold being overrepresented and outperforming others. Despite the increased diversity of scaffolds observed for context radii of 4 and 5, these scaffolds were often reproducible across different fragment databases (Fig. S3).
Two near-optimal configurations were identified: the use of the SA2 fragment database with a radius of 2 and the SA2.5 fragment database with a radius of 3. Both settings yielded high docking scores, ranging from −12.5 to −12.1, together with favorable SA scores, ranging from 2.8 to 2.95. These configurations also demonstrated comparable diversity in terms of distinct structures and scaffolds (Fig. S2), as well as novelty of the generated compounds (Fig. 3). Given the minimal differences between these conditions, the former configuration – the SA2 fragment database with a radius of 2 – was arbitrarily selected for all subsequent experiments.
As anticipated, the application of the greedy selection strategy resulted in partially reproducible molecular structures across independent runs, characterized by low scaffold diversity (Fig. S4b). The clustering selection exhibited moderate scaffold diversity, while the Pareto selection demonstrated the highest scaffold diversity. Similarly to previous strategies, there was partial reproducibility of generated scaffolds within top 100 structures (Fig. S4b).
The percentage of generated structures that bind to the hinge region was highest for the Pareto strategy (19–25%), followed by the greedy approach (19–20%) and clustering (15–17%). The number of generated and docked structures varied significantly across the different strategies. The clustering strategy consistently produced the lowest number of structures (87![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000–89 000). In contrast, the greedy strategy generated between 90
000–89 000). In contrast, the greedy strategy generated between 90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 and 92
000 and 92![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 molecules, while the Pareto strategy yielded substantially more structures, ranging from 108
000 molecules, while the Pareto strategy yielded substantially more structures, ranging from 108![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 to 148
000 to 148![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 (Table S1).
000 (Table S1).
While the Pareto strategy offers the highest diversity of scaffolds among the generated compounds, its runtime is less predictable compared to the other strategies due to the variable number of compounds selected in each iteration and the potential for a greater number of iterations. The greedy search, on the other hand, results in partially reproducible outputs with compounds exhibiting high docking scores, thus negating the necessity for multiple runs. However, the diversity of the generated structures remains relatively low. The clustering strategy presents a balanced approach, offering predictable runtime and reasonable diversity.
All simulations were conducted in a single replicate. The initial strategy involved augmentation of the docking score by the fraction of sp3 carbon atoms in Bemis–Murcko scaffolds (Csp3BM). This method resulted in only a marginal increase in the proportion of generated molecules that met the specified criteria (Fig. 4a). A more effective strategy involved using a custom sampling function that selected fragments from the CReM database in proportion to the squared fraction of sp3 carbon atoms. This increased the proportion of compounds with the Csp3BM fraction ≥ 0.3 on 15–20%.
The most significant improvement was achieved by pre-filtering the starting fragments to retain only those with Csp3BM values of 0.3 or higher. This filtering reduced the number of starting fragments from 23![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 840 (SA2 fragments) to 2851 (SA2 Csp3-rich fragments), while markedly increasing the proportion of generated molecules with Csp3BM ≥ 0.3, rising from ∼15% to ∼57% (Fig. 4a). Additionally, this filtering led to increased synthetic accessibility scores for the top-scoring compounds that satisfied PLIP and had Csp3BM ≥ 0.3, while maintaining docking scores comparable to or better than those obtained using the complete fragment set (Fig. 4b).
840 (SA2 fragments) to 2851 (SA2 Csp3-rich fragments), while markedly increasing the proportion of generated molecules with Csp3BM ≥ 0.3, rising from ∼15% to ∼57% (Fig. 4a). Additionally, this filtering led to increased synthetic accessibility scores for the top-scoring compounds that satisfied PLIP and had Csp3BM ≥ 0.3, while maintaining docking scores comparable to or better than those obtained using the complete fragment set (Fig. 4b).
To further explore this phenomenon, an alternative set of starting fragments was prepared following the same protocol described in the Methods section, utilizing the CReM SA2.5 database. This approach yielded 27![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 802 starting fragments enriched in sp3 carbon atoms (SA2.5 Csp3-rich fragments). Utilizing this extended initial set, a similarly high proportion of compounds meeting the desired criteria (55–77%) was achieved. The top-scoring molecules from this dataset exhibited slightly improved docking scores and increased SA scores. Consequently, the proportion of structures containing a substantial fraction of sp3 carbon atoms appears to be largely independent of the initial fragment set size, but is instead influenced by its composition. Examples of these top-ranked molecules are presented in Table S2.
802 starting fragments enriched in sp3 carbon atoms (SA2.5 Csp3-rich fragments). Utilizing this extended initial set, a similarly high proportion of compounds meeting the desired criteria (55–77%) was achieved. The top-scoring molecules from this dataset exhibited slightly improved docking scores and increased SA scores. Consequently, the proportion of structures containing a substantial fraction of sp3 carbon atoms appears to be largely independent of the initial fragment set size, but is instead influenced by its composition. Examples of these top-ranked molecules are presented in Table S2.
In summary, the commonly employed strategy of augmenting docking scores with additional parameters to guide molecular generation towards a specific region of chemical space proved to be ineffective in this context, likely due to the necessity for fine-tuning the augmented objective function. A more effective approach involved direct control over the composition of starting fragments and the fragments utilized during the molecular growth process.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000) was somewhat greater than the average number of compounds docked during the de novo generation process utilizing the ChEMBL SA2 fragment database with a radius of 2. This selection was undertaken to evaluate the efficacy of virtual screening relative to our de novo design pipeline, employing comparable computational resources for both methodologies. For comparison, we additionally conducted de novo generation using the previously established settings, but without PLIP guidance. This approach aimed to provide a clearer understanding of the relationship between virtual screening and de novo generation.
000) was somewhat greater than the average number of compounds docked during the de novo generation process utilizing the ChEMBL SA2 fragment database with a radius of 2. This selection was undertaken to evaluate the efficacy of virtual screening relative to our de novo design pipeline, employing comparable computational resources for both methodologies. For comparison, we additionally conducted de novo generation using the previously established settings, but without PLIP guidance. This approach aimed to provide a clearer understanding of the relationship between virtual screening and de novo generation.
Notably, only 617 ZINC compounds (0.51%) were able to establish the required interactions with the hinge region, a proportion lower than that observed among de novo generated compounds, for which 16.8% and 5.4% of structures were predicted to bind the hinge region in the PLIP-guided and non-guided runs, respectively. Furthermore, the docking scores of the highest-ranked compounds from the ZINC database were inferior to those of the de novo generated molecules, regardless of the compounds' ability to bind to the hinge region (Fig. 5). This indicates that de novo generation techniques can yield superior docking scores compared to traditional virtual screening methods when comparable computational resources are employed. Additionally, de novo generation, regardless of biasing towards favorable interactions, demonstrated a greater success rate in identifying structures that conform to a specified protein–ligand interaction pattern compared to the conventional screening approach.
De novo design of ligands for different protein families and comparison with state-of-the-art approaches
REINVENT80 and ChemTS (version 2)81 were chosen as reference methods because both are established open-source platforms for de novo molecular design, while representing algorithmic paradigms distinct from the fragment-based CReM framework. REINVENT is a recurrent neural network model trained on SMILES representations of ChEMBL structures. ChemTS is based on a recurrent neural network trained on ChEMBL as a molecule generator and Monte Carlo Tree Search as an exploration approach. Both methods can be coupled to structure-based scoring workflows, making them suitable comparators in a docking-guided setting. Both tools are mature and can be easily customized to align with the settings used by CReM-dock.To adjust REINVENT and ChemTS to the settings used by CReM-dock we applied several modifications to their pipelines. We fine-tuned the REINVENT model pre-trained on ChEMBL data to generate structures satisfying physicochemical criteria (MW ≤ 450, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P ≤ 4, RTB ≤ 5, TPSA ≤ 120) and containing no structural alerts from the BMS, Dundee, Glaxo, Inpharmatica, and PAINS filter lists. In the second stage, structures were generated to satisfy the criteria above and were additionally biased by docking scores from AutoDock Vina38 and SA scores (the upper bound of a sigmoid transformation function was set to 3.5). We activated the Murcko scaffold diversity filter recommended by the authors of REINVENT, which is intended to explicitly manage diversity of generated structures and maintain it at a high level. To adjust ChemTS, we applied strict filters by physicochemical properties (MW ≤ 450, log
P ≤ 4, RTB ≤ 5, TPSA ≤ 120) and containing no structural alerts from the BMS, Dundee, Glaxo, Inpharmatica, and PAINS filter lists. In the second stage, structures were generated to satisfy the criteria above and were additionally biased by docking scores from AutoDock Vina38 and SA scores (the upper bound of a sigmoid transformation function was set to 3.5). We activated the Murcko scaffold diversity filter recommended by the authors of REINVENT, which is intended to explicitly manage diversity of generated structures and maintain it at a high level. To adjust ChemTS, we applied strict filters by physicochemical properties (MW ≤ 450, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P ≤ 4, RTB ≤ 5, TPSA ≤ 120), structural alerts, and SA score (≤3.5), and generation was guided by AutoDock Vina docking score. The default docking protocols of REINVENT and ChemTS differed from that of CReM-dock. REINVENT required proprietary protonation modules, while ChemTS pipeline did not include a protonation step. Therefore, we incorporated protonation with the Uni-pKa model into REINVENT and updated Vina to version 1.2.7 used in CReM-dock. In ChemTS, we integrated the EasyDock pipeline including protonation with Uni-pKa and docking with Vina 1.2.7. Exhaustiveness was set to 8 for all programs.
P ≤ 4, RTB ≤ 5, TPSA ≤ 120), structural alerts, and SA score (≤3.5), and generation was guided by AutoDock Vina docking score. The default docking protocols of REINVENT and ChemTS differed from that of CReM-dock. REINVENT required proprietary protonation modules, while ChemTS pipeline did not include a protonation step. Therefore, we incorporated protonation with the Uni-pKa model into REINVENT and updated Vina to version 1.2.7 used in CReM-dock. In ChemTS, we integrated the EasyDock pipeline including protonation with Uni-pKa and docking with Vina 1.2.7. Exhaustiveness was set to 8 for all programs.
The targets selected for this study belonged to different protein families: CDK2 (PDB 2BTR, kinase), BACE1 (PDB 6UWP, protease), DRD2 (PDB 6CM4, GPCR), ESR1 (PDB 8DV7, nuclear receptor), HDAC2 (PDB 7ZZT, epigenetic regulator), and PARP1 (PDB 7ONT, transferase) (Table S3). For CReM-dock, we used the same settings as in the previous experiments (SA2 fragment database and context radius of 2). Since both REINVENT and ChemTS do not incorporate biasing of structure generation by protein–ligand interactions, we ran CReM-dock without PLIP consideration. However, for the analysis we included results obtained previously, where PLIP was considered. For REINVENT and ChemTS, we ran a sufficient number of iterations to obtain approximately the same number of docking events as for CReM-dock.
The proportion of generated structures satisfying physicochemical criteria was high for both programs, REINVENT (∼95%) and ChemTS (∼85%). The number of structures for which important protein–ligand interactions were observed was low (Table 3). In most cases, fewer than 1% of structures exhibited such interactions. Only for HDAC2, where only a single interaction was identified as essential, the proportion of such structures was high. Conversely, CReM-dock runs, which were not guided by PLIP, demonstrated a much higher fraction of structures exhibiting important protein–ligand interactions, up to 9.5%, apart from HDAC2 target, for which 45% of ligands exhibited such interactions. We can only hypothesize that such a large number of structures with the expected PLIP were generated by CReM-dock owing to the iterative growing approach. While structures are smaller in earlier iterations, they have greater chances to establishing particular contacts within the binding site. This is supported by the observation that in early iterations, the fraction of generated structures satisfying given PLIPs is greater (Table S4). Therefore, even without direct guidance and explicit selection of structures having desired interactions, CReM-dock generated a higher number of structures satisfying the desired PLIPs than approaches generating complete structures in a single step, such as REINVENT or ChemTS. CReM-dock guided explicitly by PLIP, as expected, resulted in the highest number of compounds satisfying the desired interaction pattern among all tested approaches (Table 3).
| Protein | Program | n | n (physchem) | % | n (PLIP ≥ 0.6) | % | n (physchem & PLIP ≥ 0.6) | % |
|---|---|---|---|---|---|---|---|---|
| BACE1 | CReM-dock (with PLIP) | 80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 077 077 |
80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 077 077 |
100 | 7155 | 8.94 | 7155 | 8.94 |
| BACE1 | CReM-dock (w/o PLIP) | 94![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 134 134 |
94![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 134 134 |
100 | 418 | 0.44 | 418 | 0.44 |
| BACE1 | REINVENT | 95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 022 022 |
89![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 612 612 |
94.31 | 17 | 0.02 | 16 | 0.02 |
| BACE1 | CHEMTS | 95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 012 012 |
80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 925 925 |
85.17 | 115 | 0.12 | 82 | 0.09 |
| CDK2 | CReM-dock (with PLIP) | 89![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 530 530 |
89![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 530 530 |
100 | 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 002 002 |
16.76 | 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 002 002 |
16.76 |
| CDK2 | CReM-dock (w/o PLIP) | 91![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 866 866 |
91![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 866 866 |
100 | 4911 | 5.35 | 4911 | 5.35 |
| CDK2 | REINVENT | 94![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 726 726 |
89![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 387 387 |
94.36 | 315 | 0.33 | 272 | 0.29 |
| CDK2 | CHEMTS | 91![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 779 779 |
85![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 259 259 |
92.9 | 967 | 1.05 | 928 | 1.01 |
| DRD2 | CReM-dock (with PLIP) | 99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 433 433 |
99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 433 433 |
100 | 24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 743 743 |
24.88 | 24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 743 743 |
24.88 |
| DRD2 | CReM-dock (w/o PLIP) | 97![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 342 342 |
97![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 342 342 |
100 | 4616 | 4.74 | 4616 | 4.74 |
| DRD2 | REINVENT | 98![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 429 429 |
92![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 939 939 |
94.42 | 0 | 0 | 0 | 0 |
| DRD2 | CHEMTS | 93![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 617 617 |
80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 472 472 |
85.96 | 0 | 0 | 0 | 0 |
| ESR1 | CReM-dock (with PLIP) | 96![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 858 858 |
96![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 858 858 |
100 | 1824 | 1.88 | 1824 | 1.88 |
| ESR1 | CReM-dock (w/o PLIP) | 96![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 388 388 |
96![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 388 388 |
100 | 186 | 0.19 | 186 | 0.19 |
| ESR1 | REINVENT | 97![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 011 011 |
93![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 310 310 |
96.18 | 12 | 0.01 | 10 | 0.01 |
| ESR1 | CHEMTS | 94![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 607 607 |
81![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 962 962 |
86.63 | 110 | 0.12 | 104 | 0.11 |
| HDAC2 | CReM-dock (with PLIP) | 95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 109 109 |
95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 109 109 |
100 | 45![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 853 853 |
48.21 | 45![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 853 853 |
48.21 |
| HDAC2 | CReM-dock (w/o PLIP) | 95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 458 458 |
95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 458 458 |
100 | 43![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 478 478 |
45.55 | 43![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 478 478 |
45.55 |
| HDAC2 | REINVENT | 94![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 848 848 |
89![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 791 791 |
94.67 | 57![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 910 910 |
61.06 | 54![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 396 396 |
57.35 |
| HDAC2 | CHEMTS | 92![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 462 462 |
77![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 842 842 |
84.19 | 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 471 471 |
27.55 | 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 472 472 |
22.14 |
| PARP1 | CReM-dock (with PLIP) | 90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 488 488 |
90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 488 488 |
100 | 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 842 842 |
20.82 | 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 842 842 |
20.82 |
| PARP1 | CReM-dock (w/o PLIP) | 91![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 429 429 |
91![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 429 429 |
100 | 8652 | 9.46 | 8652 | 9.46 |
| PARP1 | REINVENT | 92![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 003 003 |
86![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 199 199 |
93.69 | 408 | 0.44 | 389 | 0.42 |
| PARP1 | CHEMTS | 94![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 681 681 |
82![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 262 262 |
86.88 | 848 | 0.9 | 768 | 0.81 |
For the analysis, we selected the top 100 structures by the highest docking score for each program. Additionally, we selected top 100 structures from corresponding subsets of compounds that satisfied physicochemical criteria only and both physicochemical and PLIP simultaneously (Fig. 6). Docking scores for ligands generated by CReM-dock were comparable to or better than those of REINVENT. Docking scores for ligands generated by ChemTS were systematically worse relative to both CReM-dock and REINVENT. When the PLIP criterion was applied for compound selection, docking scores of the top 100 compounds dropped for all programs, except CReM-dock generations explicitly guided by PLIP. In many cases, the generated structures outperformed known actives from ChEMBL in terms of docking scores.
SA scores of generated structures for all programs were at the acceptable level for different scenarios of compound selection, mainly below 3 (Fig. 6). Explicit biasing in REINVENT and ChemTS as well as implicit biasing in CReM-dock worked effectively. The only exception was DRD2 ligands generated by CReM-dock with the PLIP criterion, where the median SA score of the top 100 compounds was 3.6. However, these compounds had high docking scores, while REINVENT and ChemTS were unable to generate ligands capable of interacting with the aspartic acid residue, which is considered a canonical interaction for the dopamine D2 receptor and many other GPCRs.82,83
Structures generated by REINVENT and ChemTS exhibited greater similarity to compounds from ChEMBL, resulting in lower novelty compared to those generated by CReM-dock (Fig. S5). Although CReM-dock structures were composed of fragments from ChEMBL molecules, they demonstrated reduced overlap with the reference ChEMBL chemical space (Fig. S6). Interestingly, the top 100 compounds selected from REINVENT runs frequently clustered together on UMAPs (Fig. S6). Pairwise similarity calculated for the top 100 compounds showed that structures generated by ChemTS had the highest diversity, while REINVENT, explicitly biased by scaffold diversity, was comparable to or worse than CReM-dock (Fig. 7). The observed similarity of 1 in Fig. 7 is due to the stereoisomers. Examples of top scored structures are given in Table S5.
 | ||
| Fig. 7 Pairwise Tanimoto similarity computed using 2048-bit Morgan fingerprints with radius 3 for top 100 compounds selected by docking score. | ||
Evaluation of designed compounds by calculation of binding free energies
Docking scores alone do not provide a sufficiently reliable metric for assessing the promise of designed compounds. In a complete design pipeline, the generated structures should therefore undergo additional post-processing to identify the most promising candidates. In the present study, this was achieved by calculating binding free energies using the A3FE approach.58 For this evaluation, we selected structures generated for CDK2 by CReM-dock and by the reference methods. Three CReM-dock sets were considered: (i) structures generated without PLIP guidance and without selection for hinge-region interactions; (ii) structures generated with explicit PLIP guidance and selected to preserve interactions with the hinge region; and (iii) structures generated with explicit PLIP guidance, initiated from SA2 Csp3-rich fragments, sampled using a Csp3-enriched fragment selection strategy, and subsequently selected for maintaining hinge-region interactions. This design enabled us to test two hypotheses: first, whether preservation of hinge-region interactions is important for obtaining more favorable calculated free energies, and second, whether enrichment in Csp3 atoms influences the calculated free-energy values. For comparison, we also selected structures generated by REINVENT and ChemTS. From each set, the top 20 compounds ranked by docking score and possessing distinct Murcko scaffolds were chosen.To achieve a reasonable trade-off between throughput and accuracy the MD sampling time allocated to each window of the ABFE protocol was 1 ns. ABFE calculations executed with such short sampling time are known to tend to systematically overestimate experimental binding free energies.58,70,84 To account for the bias introduced with this ABFE protocol, two reference sets of compounds were included to facilitate discrimination between active and inactive molecules. As inactive compounds, we selected 20 ChEMBL molecules with pIC50 < 5. These compounds were docked using Vina, and the top-ranked poses were used for ΔG calculations. None of these compounds formed interactions with the hinge region. As active compounds, we selected 12 ligands from PDB complexes (1FVV, 1OIR, 2B52, 2B55, 2UZL, 2VV9, 2XMY, 3EZV, 4BCP, 4BCQ, 5K4J, and 5NEV) with pIC50 ≥ 6. For these ligands, the native poses observed in the corresponding X-ray structures were used. In all cases except 2UZL, the ligands formed contacts with the hinge region in their crystallographic binding modes.
The calculated binding free energies clearly discriminated the active PDB ligands from the inactive ChEMBL compounds (Fig. 8). Compounds generated by CReM-dock without guidance and without selection for hinge-region interactions were not statistically distinguishable from the inactive reference set, whereas structures generated with PLIP guidance and selected for hinge-region interactions differed significantly from the true inactives. Csp3-enriched structures designed under PLIP guidance showed ΔG values comparable to those of structures generated without explicit Csp3 enrichment (Fig. 8). These findings suggest that, for CReM-dock, guidance by a known interaction pattern may be important for obtaining favorable binding free energies, whereas enrichment in Csp3 atoms does not substantially affect this metric.
Structures generated by ChemTS were likewise not statistically distinguishable from the inactive set, although one compound exhibited a particularly favorable calculated free energy and may therefore represent a promising candidate. Structures generated by REINVENT differed significantly from the ChEMBL inactive compounds and displayed the narrowest distribution of calculated ΔG values among all methods. Notably, REINVENT-generated structures generally did not interact with the hinge region, with only five compounds forming a single hydrogen bond out of the three possible hinge-region interactions. Thus, unlike CReM-dock, REINVENT did not appear to require explicit PLIP guidance to achieve favorable ΔG values. Overall, the results obtained for REINVENT and for PLIP-guided CReM-dock were statistically indistinguishable. Nevertheless, CReM-dock produced several compounds with particularly promising calculated binding free energies. Representative top-ranked structures are shown in Fig. 9.
 | ||
| Fig. 9 Structures designed by CReM-dock, REINVENT and ChemTS having the lowest calculated binding free energy to CDK2. | ||
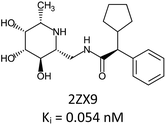
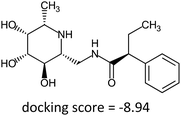
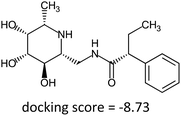
Fragment expansion study
The CReM-dock methodology, which is based on the principle of fragment growth, is particularly well-suited for scenarios in which a smaller ligand is to be expanded within a binding site to yield a larger molecule that more effectively occupies the cavity. To assess the efficacy of the CReM-dock approach, we selected pairs of compounds from the prior research by Malhotra and Karanicolas,85 who compiled an extensive collection of smaller and larger ligands that were co-crystallized with the same protein and are accessible in the Protein Data Bank (PDB). The criteria for selecting ligand pairs included: (i) the larger ligand must contain fewer than 36 heavy atoms to approximately satisfy the molecular weight constraint of ≤500, (ii) both the smaller and the larger ligands should exhibit a high degree of volume overlap, indicating minimal pose alteration upon expansion, (iii) the root mean square deviation (RMSD) between the binding pockets of the corresponding protein conformations should be low, signifying that there are no significant positional changes in the side-chain residues, and (iv) the difference in biological activity between the smaller and larger ligands must exceed two orders of magnitude. Ultimately, three ligand pairs meeting these criteria were selected (Table 4).In conducting the simulations, we modified certain parameters to avoid excessive searching and to conserve computational resources. Given that molecular growth results in an increase in molecular weight, it was deemed impractical to continue the generation process after achieving the desired molecular mass of the larger ligand. The threshold values for physicochemical parameters were established at the corresponding values of the larger ligand plus 15% (for molecular weight, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, and topological polar surface area), while the rotatable bond count (RTB) was set to the corresponding value of the larger ligand plus 1. To further constrain the generation process, we established an RMSD threshold of 1 Å and a minimum PLIP similarity of 0.6 for the 2ZWZ and 3S1G pairs, and 0.5 for the 2HB1 pair (Table S6). These parameters were intended to direct the generation of compounds that would maintain the binding pose of the initial fragment and its interactions with the corresponding protein. For the generation process, we utilized the ChEMBL SA2 fragment library with a radius of 2. Our selection strategy involved clustering into 100 groups and selecting the top-scoring structure from each cluster, thereby enhancing the comprehensiveness of the search and expanding the chemical space explored. This approach did not significantly increase computational costs, as we commenced with relatively large fragments and anticipated that the target molecules would be generated with a limited number of steps. Overall, between 8200 and 24
P, and topological polar surface area), while the rotatable bond count (RTB) was set to the corresponding value of the larger ligand plus 1. To further constrain the generation process, we established an RMSD threshold of 1 Å and a minimum PLIP similarity of 0.6 for the 2ZWZ and 3S1G pairs, and 0.5 for the 2HB1 pair (Table S6). These parameters were intended to direct the generation of compounds that would maintain the binding pose of the initial fragment and its interactions with the corresponding protein. For the generation process, we utilized the ChEMBL SA2 fragment library with a radius of 2. Our selection strategy involved clustering into 100 groups and selecting the top-scoring structure from each cluster, thereby enhancing the comprehensiveness of the search and expanding the chemical space explored. This approach did not significantly increase computational costs, as we commenced with relatively large fragments and anticipated that the target molecules would be generated with a limited number of steps. Overall, between 8200 and 24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 molecules were generated for each of the three targets.
500 molecules were generated for each of the three targets.
From the pool of generated structures, we identified those most similar to the target compounds using Tanimoto similarity based on 2048-bit Morgan fingerprints with a radius of 2. In all instances, the most similar generated ligands exhibited the same binding mode and preserved key protein–ligand interactions (Table 4, Fig. 10). For the 2HB1–2QBS pair, the pipeline successfully identified a structure identical to the larger ligand, 2QBS. Additionally, another structure with a similarity score of 1 was found, which contained a cycloheptyl residue in place of cyclohexyl; both compounds demonstrated highly similar binding modes relative to the target ligand. In the case of the 3S1G–3GC4 pair, the most similar generated ligand lacked a positively charged secondary amine center and, consequently, could not establish contact with Asp280 as the target ligand did. However, the generated compound included an amino group that enhanced binding to Asp156 and featured a hydrophobic moiety that filled the pocket in a manner analogous to the target compound. For the 2ZWZ–2ZX9 pair, the most similar generated ligand effectively occupied the hydrophobic pocket in a manner closely resembling the target compound, although it contained an ethyl group instead of a cyclopentyl residue. The most similar structure was represented by two enantiomers; both could adapt to the binding site and maintain the necessary contacts.
 | ||
| Fig. 10 Binding poses of smaller (green) and larger (magenta) ligands as well as generated compounds (yellow/blue) the most similar to the target (larger) ligand. | ||
In all cases, the generated structures were able to approach the chemical space represented by the target molecules and fit within the binding site while preserving the pose and interactions (Fig. 10). This demonstrates the capability of the CReM-dock approach to explore not only novel chemical spaces but also relevant spaces that have been previously validated through experimental means.
Discussion
Several pertinent questions warrant discussion in the context of fragment-based approaches to drug discovery. A primary question is whether the extensive chemical space explored by these methodologies genuinely correlates with the identification of superior drug-like candidates. Although systematic investigations on this matter are lacking, we can engage in some speculation. The exploration of broader chemical spaces is anticipated to yield advantages, such as the identification of novel chemotypes and more potent hits. This assertion is supported by instances where the docking of millions of compounds has led to the discovery of structurally novel and more effective hits.16–18 These cases illustrate the benefits of exploring larger chemical spaces.A critical consideration, however, is the potential generation of numerous synthetically unfeasible structures, as it is conceivable that an approach may predominantly explore regions of chemical space that are not synthetically accessible. Nonetheless, this does not inherently imply that an approach will fail to identify promising hits; rather, it suggests that such an approach may be computationally inefficient, necessitating additional time for output analysis. Previous research demonstrated that AiZynthFinder can generate retrosynthetic pathways for structures with SA scores below 2.5–3 in 50% of cases or more.35 Consequently, we hypothesize that structures generated with SA scores below 3 or 2.5 are likely to be synthetically accessible. The CReM-dock tool primarily produces structures with low SA scores, as the complexity of the structures is incrementally increased across iterations (Fig. S7).
We conducted an analysis of the Retrosynthetic Accessibility (RA) scores86 for the top 100 generated structures and observed that the RA scores were predominantly high across nearly all protein targets, with a median exceeding 0.95. This suggests that at least half of these structures may have feasible retrosynthetic pathways (Fig. 11). It is important to acknowledge that the RA score has inherent limitations; for instance, previously synthesized compounds, such as inhibitors of BACE1, did not achieve high RA scores. To enhance the likelihood that the majority of the top-scoring generated structures attain high RA scores, we recommend utilizing a radius of 4 or greater, along with a fragment database derived from more synthetically accessible compounds (Fig. S8). However, this approach may result in a reduction of the docking scores for those structures (Fig. 2). Consequently, we do not anticipate significant challenges for CReM-dock concerning the synthetic accessibility of the designed structures, as it allows for flexible control over this aspect. Nonetheless, experimental validation remains crucial to confirm the practical applicability of the proposed pipeline.
Another aspect to consider in addressing the primary question is the lack of evidence that a single generative tool can efficiently explore the entirety of the synthetically accessible chemical space and outperform other tools. It is more plausible that each tool covers distinct regions of chemical space, which may overlap but are not entirely encompassed by other tools. Thus, employing multiple tools in conjunction may yield complementary insights and facilitate the selection of the most promising structures for further investigation.
While there is currently no experimental validation of the specific pipeline described herein, nor evidence that the designed structures are synthetically accessible, the developed tool has been integrated into a more comprehensive design and virtual screening pipeline, demonstrating its applicability. In the CACHE challenge #1, where our group achieved third place,48 participants were tasked with identifying binders for the WDR40 domain of the LRRK2 kinase, implicated in the pathogenesis of Parkinson's disease. CReM-dock was utilized as one of the tools to generate promising hits, which were subsequently employed as queries for similarity searches in ultra-large chemical databases. The suggested hits were confirmed through experimental validation. This underscores the utility of the developed tool.
Practical recommendations:
• It is advisable to utilize larger context radii and/or fragments derived from more synthetically accessible molecules to facilitate the generation of structures that are themselves synthetically accessible.
• In scenarios involving larger radii and smaller fragment databases, repeated runs are unnecessary, as the results exhibit high reproducibility.
• To achieve greater diversity of solutions, Pareto selection is recommended; conversely, for more stable runtime, selection based on clustering should be employed.
• Whenever feasible, incorporating multiple protein conformations can enhance the diversity of the generated structures.
• It is beneficial to impose restrictions on protein–ligand interactions, if possible.
• The augmentation of an objective function has been shown to be effective in enhancing drug-likeness.
• When generating structures enriched with sp3 carbon atoms, it is advisable to utilize fragments that are themselves enriched in sp3 carbon atoms, as the augmentation of the objective function tends to perform poorly in this context.
• The number of iterations may be constrained to five or six, as subsequent iterations often lead to increased molecular complexity while resulting in diminished docking scores (Fig. S7).
Upon analyzing the outputs of CReM-dock, it was observed that many of the top-scoring structures contained numerous methyl groups or halogen atoms. These substituents are typically introduced during the later iterations and are unlikely to be critically important for ligand-protein recognition and binding. However, their incorporation can increase the complexity of the structures and reduce their synthetic feasibility. To mitigate this issue, it is recommended to specify a minimum size for the attached fragment that exceeds the default value of one.
An alternative approach involves the application of a fragment expansion strategy. Specifically, an anchor fragment that demonstrates binding affinity to a protein can be extracted from a co-crystallized ligand. This fragment can then be expanded using CReM-dock while maintaining its binding pose and interactions. This method presents potential advantages over de novo design, as the binding pose of at least one fragment is already established, thereby increasing the probability of generating active molecules.
Conclusions
The proposed fragment-based methodology for structure generation, informed by molecular docking, has yielded encouraging results. The approach partially addresses the challenge of synthetic accessibility by providing indirect yet effective control over this property. In particular, the use of a larger context radius and fragment databases derived from more synthetically accessible molecules improved the synthetic accessibility of the generated compounds. In addition, the framework allows docking scores to be combined with physicochemical parameters, thereby enabling the generation of structures with more favourable overall profiles. Although this strategy was effective in promoting drug-like properties, it was less suitable for increasing the fraction of sp3-hybridized carbon atoms; in such cases, explicit biasing through the selection of Csp3-enriched starting fragments or growth fragments proved more effective. The methodology also supports multiple selection strategies, thereby providing control over the diversity of the generated molecules.An important advantage of the approach is its ability to preserve predefined protein–ligand interactions during generation, particularly when such interactions are known to be essential. This substantially increases the number of structures exhibiting the desired interaction patterns. An additional level of control can be introduced by imposing an RMSD threshold relative to the pose of the initial fragment, thereby helping to preserve the binding conformation of the generated molecules. In three retrospective case studies, this strategy enabled the generation of compounds that were either identical or highly similar to known target molecules previously developed by medicinal chemists during fragment optimization.
The developed tool was found to be competitive with the state-of-the-art REINVENT4 and ChemTS approaches. Molecules generated by CReM-dock frequently displayed greater novelty while maintaining comparable docking scores and synthetic accessibility. Furthermore, in a separate study, structures generated by CReM-dock under guidance from protein–ligand interaction patterns exhibited favourable calculated binding free energies for CDK2 and achieved ΔG values comparable to those of known inhibitors, whereas structures generated without consideration of PLIP-derived interactions showed poorer calculated binding energies. Overall, the method provides substantial flexibility and can be adapted to a range of tasks, including de novo molecular generation, fragment expansion, and scaffold decoration.
Author contributions
G. M.: methodology, software, validation, investigation, visualization, writing – original draft. H. D.: software, investigation, writing – review & editing. F. C.: software; J. M.: writing – review & editing, supervision. P. P.: conceptualization, methodology, software, visualization, writing – original draft, review & editing, supervision, funding acquisition.Conflicts of interest
JM is member of the Scientific Advisory Board of Cresset. The other author declares no competing interests.Abbreviations
| Csp3 | The fraction of sp3 carbon atoms in a molecule |
| Csp3BM | The fraction of sp3 carbon atoms in a Bemis–Murcko scaffold of molecule |
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P P | Lipophilicity |
| MW | Molecular weight |
| PLIP | Protein–ligand interaction pattern (a set of protein–ligand contacts which should be at least partially observed in generated structures docked to a corresponding protein) |
| QED | Quantitative estimate of drug-likeness |
| RTB | The number of rotatable bonds |
| SA | Synthetic accessibility score |
| TPSA | Topological surface area |
Data availability
Source code of CReM-dock, documentation and examples are available at the repository https://github.com/ci-lab-cz/crem-dock. ChEMBL22 fragment databases used in the study are available at https://doi.org/10.5281/zenodo.16909328. Structures of all molecules generated in the course of this study are accessible by this link – https://doi.org/10.5281/zenodo.19136389.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d6dd00131a.
Acknowledgements
The work was supported by the Ministry of Education, Youth and Sports of the Czech Republic through INTER_EXCELLENCE II grant LUAUS23262 and the e-INFRA CZ (ID:90254), and partially by ELIXIR-CZ (LM2023055) and CZ-OPENSCREEN (LM2023052). The authors express their gratitude to Guzel Mindubaeva for her contributions during the initial phases of the project, to Aleksandra Ivanova for her assistance in the implementation of certain features, and to Veincent Yap for his involvement in the EasyDock project, which enhanced the sampling of ring conformations for improved docking accuracy. Additionally, the authors acknowledge Andrew Dalke for granting the license for the chemfp tool.References
- P. G. Polishchuk, T. I. Madzhidov and A. Varnek, J. Comput.-Aided Mol. Des., 2013, 27, 675–679 CrossRef CAS PubMed.
- G. Schneider and U. Fechner, Nat. Rev. Drug Discovery, 2005, 4, 649–663 CrossRef CAS PubMed.
- M. Hartenfeller and G. Schneider, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2011, 1, 742–759 CAS.
- J. Meyers, B. Fabian and N. Brown, Drug Discovery Today, 2021, 26, 2707–2715 CrossRef CAS PubMed.
- C. Bilodeau, W. Jin, T. Jaakkola, R. Barzilay and K. F. Jensen, Wiley Interdiscip. Rev.:Comput. Mol. Sci., 2022, e1608 Search PubMed.
- W. Gao and C. W. Coley, J. Chem. Inf. Model., 2020, 60, 5714–5723 CrossRef CAS PubMed.
- G. Domenico, M. Giuseppe Felice, C. Marco, C. Angelo and N. Orazio, Int. J. Quant. Struct.-Prop. Relat., 2016, 1, 45–63 Search PubMed.
- S. Ochi, T. Miyao and K. Funatsu, Mol. Inf., 2017, 36, 1700076 CrossRef PubMed.
- J. Li, A. Fu and L. Zhang, Interdiscip. Sci.: Comput. Life Sci., 2019, 11, 320–328 CrossRef CAS PubMed.
- L. Pinzi and G. Rastelli, Int. J. Mol. Sci., 2019, 20, 4331 CrossRef CAS PubMed.
- C. Shen, J. Ding, Z. Wang, D. Cao, X. Ding and T. Hou, Wiley Interdiscip. Rev.:Comput. Mol. Sci., 2020, 10, e1429 CAS.
- N. S. Pagadala, K. Syed and J. Tuszynski, Biophys. Rev., 2017, 9, 91–102 CrossRef CAS PubMed.
- V. Salmaso and S. Moro, Front. Pharmacol, 2018, 9, 923 CrossRef PubMed.
- A. S. J. S. Mey, B. K. Allen, H. E. Bruce McDonald, J. D. Chodera, D. F. Hahn, M. Kuhn, J. Michel, D. L. Mobley, L. N. Naden, S. Prasad, A. Rizzi, J. Scheen, M. R. Shirts, G. Tresadern and H. Xu, Living J. Comput. Mol. Sci., 2020, 2, 18378 Search PubMed.
- Z. Cournia, B. K. Allen, T. Beuming, D. A. Pearlman, B. K. Radak and W. Sherman, J. Chem. Inf. Model., 2020, 60, 4153–4169 CrossRef CAS PubMed.
- A.-T. Ton, F. Gentile, M. Hsing, F. Ban and A. Cherkasov, Mol. Inf., 2020, 39, 2000028 CrossRef CAS PubMed.
- J. Lyu, S. Wang, T. E. Balius, I. Singh, A. Levit, Y. S. Moroz, M. J. O'Meara, T. Che, E. Algaa, K. Tolmachova, A. A. Tolmachev, B. K. Shoichet, B. L. Roth and J. J. Irwin, Nature, 2019, 566, 224–229 CrossRef CAS PubMed.
- F. Liu, O. Mailhot, I. S. Glenn, S. F. Vigneron, V. Bassim, X. Xu, K. Fonseca-Valencia, M. S. Smith, D. S. Radchenko, J. S. Fraser, Y. S. Moroz, J. J. Irwin and B. K. Shoichet, Nat. Chem. Biol., 2025, 21, 1039–1045 CrossRef CAS PubMed.
- L. Batiste, A. Unzue, A. Dolbois, F. Hassler, X. Wang, N. Deerain, J. Zhu, D. Spiliotopoulos, C. Nevado and A. Caflisch, ACS Cent. Sci., 2018, 4, 180–188 CrossRef CAS PubMed.
- F. Chevillard, H. Rimmer, C. Betti, E. Pardon, S. Ballet, N. van Hilten, J. Steyaert, W. E. Diederich and P. Kolb, J. Med. Chem., 2018, 61, 1118–1129 CrossRef CAS PubMed.
- K. Sommer, F. Flachsenberg and M. Rarey, Eur. J. Med. Chem., 2019, 163, 747–762 CrossRef CAS PubMed.
- J. O. Spiegel and J. D. Durrant, J. Cheminf., 2020, 12, 25 CAS.
- Y. Yuan, J. Pei and L. Lai, Front. Chem., 2020, 8, 142 CrossRef CAS PubMed.
- C. Steinmann and J. H. Jensen, PeerJ Phys. Chem., 2021, 3, e18 CrossRef.
- P. Ertl and A. Schuffenhauer, J. Cheminf., 2009, 1, 8 Search PubMed.
- N. Chéron, N. Jasty and E. I. Shakhnovich, J. Med. Chem., 2016, 59, 4171–4188 CrossRef PubMed.
- S. Cross and G. Cruciani, J. Chem. Inf. Model., 2022, 62, 1224–1235 CrossRef CAS PubMed.
- J. Boitreaud, V. Mallet, C. Oliver and J. Waldispühl, J. Chem. Inf. Model., 2020, 60, 5658–5666 CrossRef CAS PubMed.
- Z. Xu, O. R. Wauchope and A. T. Frank, J. Chem. Inf. Model., 2021, 61, 5589–5600 CrossRef CAS PubMed.
- J. Guo, J. P. Janet, M. R. Bauer, E. Nittinger, K. A. Giblin, K. Papadopoulos, A. Voronov, A. Patronov, O. Engkvist and C. Margreitter, J. Cheminf., 2021, 13, 89 Search PubMed.
- B. Ma, K. Terayama, S. Matsumoto, Y. Isaka, Y. Sasakura, H. Iwata, M. Araki and Y. Okuno, J. Chem. Inf. Model., 2021, 61, 3304–3313 CrossRef CAS PubMed.
- M. Cretu, C. Harris, I. Igashov, A. Schneuing, M. Segler, B. Correia, J. Roy, E. Bengio and P. Lio, 2024.
- M. Koziarski, A. Rekesh, D. Shevchuk, A. M. v. d. Sloot, P. Gaiński, Y. Bengio, C.-H. Liu, M. Tyers and R. A. Batey, RGFN: synthesizable molecular generation using GFlowNets, in Proceedings of the 38th International Conference on Neural Information Processing Systems, Vancouver, BC, Canada, 2024, article 1488, pp. 46908-46955. Search PubMed.
- P. Polishchuk, J. Cheminf., 2020, 12, 28 Search PubMed.
- P. Polishchuk, J. Chem. Inf. Model., 2020, 60, 6074–6080 CrossRef CAS PubMed.
- A. Martínez León, B. Ries, J. S. Hub and A. Magarkar, J. Cheminf., 2025, 17, 85 Search PubMed.
- G. Minibaeva, A. Ivanova and P. Polishchuk, J. Cheminf., 2023, 15, 102 CAS.
- J. Eberhardt, D. Santos-Martins, A. F. Tillack and S. Forli, J. Chem. Inf. Model., 2021, 61, 3891–3898 CrossRef CAS PubMed.
- D. R. Koes, M. P. Baumgartner and C. J. Camacho, J. Chem. Inf. Model., 2013, 53, 1893–1904 CrossRef CAS PubMed.
- A. T. McNutt, P. Francoeur, R. Aggarwal, T. Masuda, R. Meli, M. Ragoza, J. Sunseri and D. R. Koes, J. Cheminf., 2021, 13, 43 Search PubMed.
- A. Alhossary, S. D. Handoko, Y. Mu and C.-K. Kwoh, Bioinformatics, 2015, 31, 2214–2216 CrossRef CAS PubMed.
- N. M. Hassan, A. A. Alhossary, Y. Mu and C.-K. Kwoh, Sci. Rep., 2017, 7, 15451 CrossRef PubMed.
- S. Tang, J. Ding, X. Zhu, Z. Wang, H. Zhao and J. Wu, IEEE/ACM Trans. Comput. Biol. Bioinf., 2024, 21, 2382–2393 CAS.
- W. Luo, G. Zhou, Z. Zhu, Y. Yuan, G. Ke, Z. Wei, Z. Gao and H. Zheng, JACS Au, 2024, 4, 3451–3465 CrossRef CAS PubMed.
- X. Pan, H. Wang, C. Li, J. Z. H. Zhang and C. Ji, J. Chem. Inf. Model., 2021, 61, 3159–3165 CrossRef CAS PubMed.
- G. R. Bickerton, G. V. Paolini, J. Besnard, S. Muresan and A. L. Hopkins, Nat. Chem., 2012, 4, 90 CrossRef CAS PubMed.
- S. Ackloo, R. Al-awar, R. E. Amaro, C. H. Arrowsmith, H. Azevedo, R. A. Batey, Y. Bengio, U. A. K. Betz, C. G. Bologa, J. D. Chodera, W. D. Cornell, I. Dunham, G. F. Ecker, K. Edfeldt, A. M. Edwards, M. K. Gilson, C. R. Gordijo, G. Hessler, A. Hillisch, A. Hogner, J. J. Irwin, J. M. Jansen, D. Kuhn, A. R. Leach, A. A. Lee, U. Lessel, M. R. Morgan, J. Moult, I. Muegge, T. I. Oprea, B. G. Perry, P. Riley, S. A. L. Rousseaux, K. S. Saikatendu, V. Santhakumar, M. Schapira, C. Scholten, M. H. Todd, M. Vedadi, A. Volkamer and T. M. Willson, Nat. Rev. Chem., 2022, 6, 287–295 CrossRef PubMed.
- F. Li, S. Ackloo, C. H. Arrowsmith, F. Ban, C. J. Barden, H. Beck, J. Beránek, F. Berenger, A. Bolotokova, G. Bret, M. Breznik, E. Carosati, I. Chau, Y. Chen, A. Cherkasov, D. D. Corte, K. Denzinger, A. Dong, S. Draga, I. Dunn, K. Edfeldt, A. Edwards, M. Eguida, P. Eisenhuth, L. Friedrich, A. Fuerll, S. S. Gardiner, F. Gentile, P. Ghiabi, E. Gibson, M. Glavatskikh, C. Gorgulla, J. Guenther, A. Gunnarsson, F. Gusev, E. Gutkin, L. Halabelian, R. J. Harding, A. Hillisch, L. Hoffer, A. Hogner, S. Houliston, J. J. Irwin, O. Isayev, A. Ivanova, C. Jacquemard, A. J. Jarrett, J. H. Jensen, D. Kireev, J. Kleber, S. B. Koby, D. Koes, A. Kumar, M. G. Kurnikova, A. Kutlushina, U. Lessel, F. Liessmann, S. Liu, W. Lu, J. Meiler, A. Mettu, G. Minibaeva, R. Moretti, C. J. Morris, C. Narangoda, T. Noonan, L. Obendorf, S. Pach, A. Pandit, S. Perveen, G. Poda, P. Polishchuk, K. Puls, V. Pütter, D. Rognan, D. Roskams-Edris, C. Schindler, F. Sindt, V. Spiwok, C. Steinmann, R. L. Stevens, V. Talagayev, D. Tingey, O. Vu, W. P. Walters, X. Wang, Z. Wang, G. Wolber, C. A. Wolf, L. Wortmann, H. Zeng, C. A. Zepeda, K. Y. J. Zhang, J. Zhang, S. Zheng and M. Schapira, J. Chem. Inf. Model., 2024, 64, 8521–8536 CrossRef CAS PubMed.
- C. Bouysset and S. Fiorucci, J. Cheminf., 2021, 13, 72 Search PubMed.
- P. T. Lang, S. R. Brozell, S. Mukherjee, E. F. Pettersen, E. C. Meng, V. Thomas, R. C. Rizzo, D. A. Case, T. L. James and I. D. Kuntz, RNA, 2009, 15, 1219–1230 CrossRef CAS PubMed.
- D. R. Houston and M. D. Walkinshaw, J. Chem. Inf. Model., 2013, 53, 384–390 CrossRef CAS PubMed.
- E. F. Pettersen, T. D. Goddard, C. C. Huang, G. S. Couch, D. M. Greenblatt, E. C. Meng and T. E. Ferrin, J. Comput. Chem., 2004, 25, 1605–1612 CrossRef CAS PubMed.
- M. Shapovalov and R. Dunbrack, Structure, 2011, 19, 844–858 CrossRef CAS PubMed.
- B. Webb and A. Sali, Curr. Protoc. Bioinf., 2016, 54, 5.6.1–5.6.37 Search PubMed .6.1–5.6.37..
- https://github.com/ci-lab-cz/docking-files .
- A. Dalke, J. Cheminf., 2019, 11, 76 Search PubMed.
- L. McInnes and J. Healy, arXiv, 2018, preprint, arXiv:1802.03426, DOI:10.48550/arXiv.1802.03426.
- F. Clark, G. R. Robb, D. J. Cole and J. Michel, J. Chem. Theory Comput., 2024, 20, 7806–7828 CAS.
- F. Clark and R. H. Du, a3fe: Automated Adaptive Absolute Alchemical Free Energy Calculator, (0.4.0), 2025, DOI:10.5281/zenodo.17298077.
- D. A. Case, T. E. Cheatham III, T. Darden, H. Gohlke, R. Luo, K. M. Merz Jr, A. Onufriev, C. Simmerling, B. Wang and R. J. Woods, J. Comput. Chem., 2005, 26, 1668–1688 CrossRef CAS PubMed.
- J. A. Maier, C. Martinez, K. Kasavajhala, L. Wickstrom, K. E. Hauser and C. Simmerling, J. Chem. Theory Comput., 2015, 11, 3696–3713 CrossRef CAS PubMed.
- M. J. Abraham, T. Murtola, R. Schulz, S. Páll, J. C. Smith, B. Hess and E. Lindahl, SoftwareX, 2015, 1–2, 19–25 CrossRef.
- G. Bussi, D. Donadio and M. Parrinello, J. Chem. Phys., 2007, 126, 014101 CrossRef PubMed.
- M. Bernetti and G. Bussi, J. Chem. Phys., 2020, 153, 114107 CrossRef CAS PubMed.
- T. Darden, D. York and L. Pedersen, J. Chem. Phys., 1993, 98, 10089–10092 CrossRef CAS.
- Y. Ge, D. F. Hahn and D. L. Mobley, J. Chem. Inf. Model., 2021, 61, 1048–1052 CrossRef CAS PubMed.
- L. O. Hedges, S. Bariami, M. Burman, F. Clark, B. P. Cossins, A. Hardie, A. M. Herz, D. Lukauskis, A. S. J. S. Mey, J. Michel, J. Scheen, M. Suruzhon, C. J. Woods and Z. Wu, Living J. Comput. Mol. Sci., 2023, 5, 2375 CrossRef.
- M. K. Gilson, J. A. Given, B. L. Bush and J. A. McCammon, Biophys. J., 1997, 72, 1047–1069 CrossRef CAS PubMed.
- M. Zacharias, T. P. Straatsma and J. A. McCammon, J. Chem. Phys., 1994, 100, 9025–9031 CrossRef CAS.
- F. Clark, G. Robb, D. J. Cole and J. Michel, J. Chem. Theory Comput., 2023, 19, 3686–3704 CrossRef CAS PubMed.
- C. H. Bennett, J. Comput. Phys., 1976, 22, 245–268 CrossRef.
- Z. X. Sun, X. H. Wang and J. Z. H. Zhang, Phys. Chem. Chem. Phys., 2017, 19, 15005–15020 RSC.
- D. Mendez, A. Gaulton, A. P. Bento, J. Chambers, M. De Veij, E. Félix, M. P. Magariños, J. F. Mosquera, P. Mutowo, M. Nowotka, M. Gordillo-Marañón, F. Hunter, L. Junco, G. Mugumbate, M. Rodriguez-Lopez, F. Atkinson, N. Bosc, C. J. Radoux, A. Segura-Cabrera, A. Hersey and A. R. Leach, Nucleic Acids Res., 2019, 47, D930–D940 CrossRef CAS PubMed.
- standardizer version 22.19.0, ChemAxon, https://www.chemaxon.com Search PubMed.
- P. Walters, https://github.com/PatWalters/rd_filters.
- P. Polishchuk, https://github.com/DrrDom/rdkit-scripts.
- cxcalc version 22.19.0, ChemAxon, https://www.chemaxon.com Search PubMed.
- F. Lovering, J. Bikker and C. Humblet, J. Med. Chem., 2009, 52, 6752–6756 CrossRef CAS PubMed.
- J. J. Irwin, K. G. Tang, J. Young, C. Dandarchuluun, B. R. Wong, M. Khurelbaatar, Y. S. Moroz, J. Mayfield and R. A. Sayle, J. Chem. Inf. Model., 2020, 60, 6065–6073 CrossRef CAS PubMed.
- H. H. Loeffler, J. He, A. Tibo, J. P. Janet, A. Voronov, L. H. Mervin and O. Engkvist, J. Cheminf., 2024, 16, 20 Search PubMed.
- S. Ishida, T. Aasawat, M. Sumita, M. Katouda, T. Yoshizawa, K. Yoshizoe, K. Tsuda and K. Terayama, Wiley Interdiscip. Rev.:Comput. Mol. Sci., 2023, 13, e1680 CAS.
- A. Mansour, F. Meng, J. H. Meador-Woodruff, L. P. Taylor, O. Civelli and H. Akil, Eur. J. Pharmacol., Mol. Pharmacol., 1992, 227, 205–214 CrossRef CAS PubMed.
- K. Jóźwiak and A. Płazińska, Molecules, 2021, 26, 851 CrossRef PubMed.
- W. Chen, D. Cui, S. V. Jerome, M. Michino, E. B. Lenselink, D. J. Huggins, A. Beautrait, J. Vendome, R. Abel, R. A. Friesner and L. Wang, J. Chem. Inf. Model., 2023, 63, 3171–3185 CrossRef CAS PubMed.
- S. Malhotra and J. Karanicolas, J. Med. Chem., 2017, 60, 128–145 CrossRef CAS PubMed.
- A. Thakkar, V. Chadimová, E. J. Bjerrum, O. Engkvist and J.-L. Reymond, Chem. Sci., 2021, 12, 3339–3349 RSC.
| This journal is © The Royal Society of Chemistry 2026 |