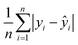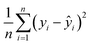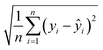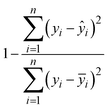Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Identification of multi-transcriptomic prognostic biomarkers to explore natural therapeutics for lung cancer integrating machine learning
Md Ahad Ali†
 *ab,
Hriddhi Sarker†
*ab,
Hriddhi Sarker† ac,
Marguba Kamrunad,
Humaira Sheikh
ac,
Marguba Kamrunad,
Humaira Sheikh ae,
Bilkis Akter Shifa
ae,
Bilkis Akter Shifa af,
Siam Ahmeda,
Tarikul Islam
af,
Siam Ahmeda,
Tarikul Islam g,
Sujoy Banikh and
Neeraj Kumari
g,
Sujoy Banikh and
Neeraj Kumari
aComputational Chemistry and Drug Design Division, Panacea Research Center, Rajshahi 6206, Bangladesh
bDepartment of Chemistry, University of Rajshahi, Rajshahi 6205, Bangladesh. E-mail: ahad.chembd@gmail.com
cDepartment of Biochemistry and Molecular Biology, University of Rajshahi, Rajshahi 6205, Bangladesh
dDepartment of Chemistry and Biochemistry, University of Oklahoma Norman, OK 73019, USA
eDepartment of Chemistry, Gopalganj Science and Technology University, Gopalganj 8100, Bangladesh
fDepartment of Biochemistry and Molecular Biology, University of Dhaka, Dhaka 1000, Bangladesh
gDepartment of Chemistry, University of Barishal, Barishal 8254, Bangladesh
hDepartment of Computer Science & Engineering, Rajshahi University of Engineering & Technology, Rajshahi, Bangladesh
iDepartment of Pharmaceutical Chemistry, Bhupal Nobles' College of Pharmacy, Udaipur 313001, Rajasthan, India
First published on 29th April 2026
Abstract
Lung cancer remains the leading cause of cancer-related mortality worldwide, underscoring the urgent need for novel therapeutic strategies. Cyclin-dependent kinase 1 (CDK1), a central cell-cycle regulator, has emerged as an oncogenic driver and potential target in lung adenocarcinoma. This study aimed to integrate transcriptomics, machine learning (ML), and advanced in silico approaches to identify natural product-derived potential inhibitors targeting CDK1. To identify robust differentially expressed genes, first we analyzed four different datasets (GSE19804, GSE10072, GSE18842, and GSE10799). Protein–protein interaction network and topological analysis highlighted CDK1 as a primary key hub gene (pKHG) enriched in cell-cycle and p53 pathways. Target validation confirmed CDK1 overexpression, prognostic significance, immune infiltration links, and mutation associations. In addition, a collected library of 9667 natural phytocompounds was reduced through ML-based bioactivity (pIC50) prediction targeting pKHG to discover potential lead molecules. Then, the selected top lead molecules were considered for further evaluation via molecular docking, molecular dynamics simulations, ADMET analysis, and binding free-energy calculations (MM-GBSA). Among the selected phytochemicals, CID_14218027 (−7.69 kcal mol−1), CID_487089 (−6.80 kcal mol−1), and CID_174880 (−6.70 kcal mol−1) showed the highest binding affinity score (GLIDE_XP score) and stable molecular interactions. Furthermore, MD simulations confirmed the conformational stability of ligand–protein complexes, supporting their potential as CDK1 inhibitors. This integrated omics-to-in silico pipeline identifies CDK1 as a robust therapeutic target and highlights natural product-derived inhibitors with favorable pharmacological and physicochemical properties. Therefore, these findings present a viable framework for accelerating precision drug discovery, with experimental validation underway. However, these findings are based solely on computational analyses and require further experimental validation to confirm CDK1 inhibitory activity, anticancer efficacy, and safety.
| Md Ahad Ali Md. Ahad Ali is a computational chemist and bioinformatics researcher affiliated with Panacea Research Center and the University of Rajshahi, where he works on transcriptomics, machine learning, and structure-based drug discovery. |
1. Introduction
Lung cancer (LC) is the deadliest cancer in the world because it causes more deaths annually than all breast, colon and prostate cancers put together.1,2 In 2022, LC was responsible for about 1.82 million deaths, representing 18.7% of all cancer fatalities, far exceeding the mortality from colorectal cancer (9.3%) and breast cancer (6.9%). Approximately 2.48 million new LC diagnoses were made that same year, along with an age-standardized mortality rate of around 16.8 per 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000, a pattern that tends to rise in countries with higher Human Development Index scores. If today's rates stay unchanged, population aging and growth could push the numbers to about 4.62 million new cases and roughly 3.55 million deaths by 2050, marking increases in total cases and deaths rather than in standardized mortality rates.2,3 LC is highly lethal due to late-stage diagnosis, inherent molecular heterogeneity, and low sensitivity to the current treatment.4,5 Though there has been a great deal of development of synthetic FDA-approved drugs, including cytotoxic chemotherapy, molecularly targeted agents, and immune checkpoint inhibitors,6–8 the overall prognosis of the patients of LC remains challenging. However, drug–drug interactions, off-target toxicity, inadequate potency, and low pharmacokinetics have limited the translation of these insights into effective therapies.9–11 Though standard synthetic drugs have played a significant role in the treatment of cancer, these drugs are usually restricted by their adverse side effects, toxicities, resistance and high cost of production.12–15 In particular, conventional chemotherapy is effective but frequently causes clinically meaningful adverse effects (notably myelosuppression and gastrointestinal toxicity), which can reduce quality of life and limit dosing intensity.16 The number of patients who initially respond eventually develop acquired resistance (e.g., resistance after EGFR-TKI/osimertinib treatment), which drives disease progression and the need for next-line options.17,18 Overall, these limitations justify exploration of alternative vulnerabilities such as cell-cycle control: cyclin-dependent kinase is a central mitotic kinase/enzyme and is reported to be overexpressed and prognostically relevant in LC. Abnormal patterns in oncogenic signaling pathways, DNA repair mechanisms, cell death, and uncontrolled cell-cycle progression collectively contribute to tumor initiation and progression.19–22 These molecular abnormalities suggest that cell-cycle regulators, mitotic proteins, and other important modulators represent promising points of therapeutic intervention. Transcriptomics and multiomics studies help to identify significant regulatory genes and signaling networks with clinical relevance. Network pharmacology-based methodologies, including protein–protein interaction (PPI) networking, hub gene identification, transcriptomics factors, and pathway enrichment analysis, are some effective ways to discover the key molecular regulators.23–26
000, a pattern that tends to rise in countries with higher Human Development Index scores. If today's rates stay unchanged, population aging and growth could push the numbers to about 4.62 million new cases and roughly 3.55 million deaths by 2050, marking increases in total cases and deaths rather than in standardized mortality rates.2,3 LC is highly lethal due to late-stage diagnosis, inherent molecular heterogeneity, and low sensitivity to the current treatment.4,5 Though there has been a great deal of development of synthetic FDA-approved drugs, including cytotoxic chemotherapy, molecularly targeted agents, and immune checkpoint inhibitors,6–8 the overall prognosis of the patients of LC remains challenging. However, drug–drug interactions, off-target toxicity, inadequate potency, and low pharmacokinetics have limited the translation of these insights into effective therapies.9–11 Though standard synthetic drugs have played a significant role in the treatment of cancer, these drugs are usually restricted by their adverse side effects, toxicities, resistance and high cost of production.12–15 In particular, conventional chemotherapy is effective but frequently causes clinically meaningful adverse effects (notably myelosuppression and gastrointestinal toxicity), which can reduce quality of life and limit dosing intensity.16 The number of patients who initially respond eventually develop acquired resistance (e.g., resistance after EGFR-TKI/osimertinib treatment), which drives disease progression and the need for next-line options.17,18 Overall, these limitations justify exploration of alternative vulnerabilities such as cell-cycle control: cyclin-dependent kinase is a central mitotic kinase/enzyme and is reported to be overexpressed and prognostically relevant in LC. Abnormal patterns in oncogenic signaling pathways, DNA repair mechanisms, cell death, and uncontrolled cell-cycle progression collectively contribute to tumor initiation and progression.19–22 These molecular abnormalities suggest that cell-cycle regulators, mitotic proteins, and other important modulators represent promising points of therapeutic intervention. Transcriptomics and multiomics studies help to identify significant regulatory genes and signaling networks with clinical relevance. Network pharmacology-based methodologies, including protein–protein interaction (PPI) networking, hub gene identification, transcriptomics factors, and pathway enrichment analysis, are some effective ways to discover the key molecular regulators.23–26
On the other hand, natural compounds—to be more specific, medicinal plant derived phytochemicals—have emerged as an attractive alternative to synthetic drug molecules, because their chemical diversity and evolved bioactivity can provide novel scaffolds and mechanisms that differ from current synthetic libraries.27 Previous studies show that more than 60 percent of the present anticancer agents are natural, including famous drugs such as paclitaxel, camptothecin, or vincristine.28–30 The IMPATT database is one of the large databases that contain information on 4010 medicinal plants and 17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 967 phytochemicals, along with their properties, which are often not well represented in other databases.31 Thus, this database is a useful source of lung cancer drugs.
967 phytochemicals, along with their properties, which are often not well represented in other databases.31 Thus, this database is a useful source of lung cancer drugs.
Nowadays, to reduce the time and cost, scientists are considering computational screening using the in silico methodology prior to evaluating the study through experimental (in vivo and in vitro) validation. This research developed a holistic computational workflow integrating transcriptomics, machine learning (ML)-based lead screening, molecular docking and a dynamic simulation study to screen a vast set of compounds in a library and develop therapeutic drug molecules with reduced toxicity and enhanced efficacy. Previous studies have used transcriptomic and machine learning (ML) approaches in LUAD primarily to identify potential biomarkers and therapeutic targets or to predict responses to approved and investigational drugs.32–36 Similarly, several studies have developed ML models using LUAD mRNA and mutation profiles to predict sensitivity to existing targeted and chemotherapeutic agents, achieving good predictive performance across dozens of drugs.37–40 In contrast to these studies, which mainly focus on biomarker/drug target identification, validation, and response prediction for existing drugs, this study integrates LUAD transcriptomics-based drug target identification with classical ML-based QSAR modelling to predict pIC50 values for plant-derived phytocompounds, followed by docking and molecular dynamics (MD) simulations to prioritize potent natural candidates.
Therefore, this study builds upon transcriptomic information from various publicly available GEO datasets to construct PPI networks, a classical ML-based regression model to predict the bioactivity (pIC50), docking validation integrating different algorithms, evaluation of the binding strength by calculating post-docking MM-GBSA, large-scale MD simulation, and pharmacokinetics analysis to create a comprehensive approach to natural product-based drug discovery for lung cancer treatment. Compared with conventional docking-centered studies, this biologically informed workflow allows both disease-relevant target prioritization and more systematic lead selection from a large natural compound library. Thus, the present work provides a comprehensive and translationally oriented computational strategy for prioritizing CDK1-targeting phytochemicals in lung adenocarcinoma and supports future experimental validation of the identified candidate compounds. A complete guideline for this work is given below in Fig. 1.
2. Materials and methods
2.1. Target identification
In this section, we present a systematic analysis of lung cancer gene expression profiles to uncover differentially expressed genes (DEGs) and highlight the primary key hub gene (pKHG), which plays a central role in understanding the underlying disease mechanisms.| GEO dataset | Number of samples | Cancer | Control | Platform | Ref. |
|---|---|---|---|---|---|
| GSE19804 | 120 | 60 | 60 | GPL570 [HG-U133_Plus_2] Affymetrix Human Genome U133 Plus 2.0 Array | 41,42 |
| GSE10072 | 107 | 58 | 49 | GPL96[ HG-U133A] Affymetrix Human Genome U133A Array | 43 |
| GSE18842 | 91 | 46 | 45 | GPL570 [HG-U133_Plus_2] Affymetrix Human Genome U133 Plus 2.0 Array | 44 |
| GSE10799 | 19 | 16 | 3 | GPL570 [HG-U133_Plus_2] Affymetrix Human Genome U133 Plus 2.0 Array | 45 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 transformation. The limma package46 was applied to detect DEGs, while the Benjamini–Hochberg false discovery rate (FDR) method was used to adjust for multiple testing. Genes with an adjusted p-value < 0.05 and log
2 transformation. The limma package46 was applied to detect DEGs, while the Benjamini–Hochberg false discovery rate (FDR) method was used to adjust for multiple testing. Genes with an adjusted p-value < 0.05 and log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) FC > +1.0 were considered significantly upregulated, whereas those with log
FC > +1.0 were considered significantly upregulated, whereas those with log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) FC < −1.0 were considered significantly downregulated.
FC < −1.0 were considered significantly downregulated.2.2. Target validation
To validate the identified prime key hub gene (pKHG) in lung cancer, we focused on the lung adenocarcinoma (LUAD) dataset from the available databases.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 FC cutoff = 1, p-value cutoff = 0.01, jitter size = 0.4, and values were log
2 FC cutoff = 1, p-value cutoff = 0.01, jitter size = 0.4, and values were log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 (TPM + 1)-transformed for visualization. The “Stage Plot” module of GEPIA2 was also utilized to determine the correlation between the expression of genes and the pathological stages (I–IV) of LUAD. In addition, two levels were applied to the UALCAN database (https://ualcan.path.uab.edu):61 (i) the study utilized TCGA RNA-seq data to validate the differences in expression of mRNA between the normal and LUAD tissues, and (ii) the study used Clinical Proteomic Tumor Analysis Consortium (CPTAC) data to test the levels of protein expression of the pKHG. This validation was consistent both at the transcriptomic and proteomic levels with this combined approach.
2 (TPM + 1)-transformed for visualization. The “Stage Plot” module of GEPIA2 was also utilized to determine the correlation between the expression of genes and the pathological stages (I–IV) of LUAD. In addition, two levels were applied to the UALCAN database (https://ualcan.path.uab.edu):61 (i) the study utilized TCGA RNA-seq data to validate the differences in expression of mRNA between the normal and LUAD tissues, and (ii) the study used Clinical Proteomic Tumor Analysis Consortium (CPTAC) data to test the levels of protein expression of the pKHG. This validation was consistent both at the transcriptomic and proteomic levels with this combined approach.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 957 samples of patients) were performed with the help of the “Query by Gene” module. The resulting “Cancer Type Summary” was a summary of mutation and copy number change types of the pKHG in various cancers. Moreover, the application of the “Mutations” module was used to produce a schematic diagram of the exact mutation sites in the gene.
957 samples of patients) were performed with the help of the “Query by Gene” module. The resulting “Cancer Type Summary” was a summary of mutation and copy number change types of the pKHG in various cancers. Moreover, the application of the “Mutations” module was used to produce a schematic diagram of the exact mutation sites in the gene.2.3. pKHG guided in silico drug discovery
To explore therapeutic opportunities, we performed structure-based drug discovery on the validated pKHG. This included retrieval and preparation of target protein's 3D crystal structure, ligand collection, virtual screening of ligands, molecular docking, ADMET, molecular dynamics (MD) simulation, MMGBSA, PCA (Principal Component Analysis) and 3D-FEL (Free Energy Landscape).2.3.4.1. Dataset curation and preparation. For dataset construction, bioactivity records and chemical structures were obtained from the ChEMBL database, a well-established platform for QSAR investigations. Compounds were then curated by retaining only those entries that reported IC50 values against pKHG. For selecting the final ChEMBL database, we mainly focused on the types of protein, organism, availability of the compound activities, including standard type (IC50) and standard unit (nm). This careful refinement produced a dataset of reliable and biologically relevant molecules, which served as the basis for regression modeling and subsequent analyses.
2.3.4.2. Molecular descriptor calculations. Molecular descriptors play a central role in QSAR modeling because they translate the structural and physicochemical features of compounds into measurable values, allowing meaningful patterns to be recognized.73 In this study, RDKit was used to calculate Morgan fingerprints and ECFP4 descriptors for each compound. These descriptors offered a detailed numerical profile of the compounds' structural characteristics, which served as essential inputs for building predictive models to estimate bioactivity in QSAR modeling.
2.3.4.3. Data splitting into train and test. The final dataset of 987 compounds was split into training and test subsets using the train_test_split function from scikit-learn with stratification on the class labels to preserve the original class distribution in both subsets. It was split into 789 molecules in the training set and 198 molecules in the test set, typical among cheminformatics and other machine learning tasks to use 20–30% of the data to test the algorithm. Random or stratified random splits are widely used as a baseline in cheminformatics modeling, although recent work has highlighted that more stringent splitting strategies (e.g., scaffold or clustering-based splits) may provide more realistic assessments of external generalization for chemical datasets.74,75
2.3.4.4. ML Model development and validation. For predictive modeling, we selected the best regression model, with a high-performance machine learning algorithm valued for its speed, accuracy, interpretability, and the coefficient of determination (R2) values of test and train datasets. To enhance the model's effectiveness, Recursive Feature Elimination (RFE) was applied to remove less informative features, reducing noise and making the model more interpretable. Here, we used optuna to tune the hyperparameter. The predictive performance of the model was assessed using widely adopted metrics, including Mean Absolute Error (MAE), Mean Squared Error (MSE), Root Mean Squared Error (RMSE), and coefficient of determination (R2). These metrics were calculated according to previously published formulae.
Mean Absolute Error (MAE):
Mean Squared Error (MSE):
Root Mean Squared Error (RMSE):
Coefficient of determination (R2):
2.3.4.5. Model implementation for bioactivity (pIC50) predictions. After model development, we applied our trained and fine-tuned model to predict the pIC50 values of the collected compounds against the pKHG. Compounds showing the highest predicted pIC50 values (>6.5), reflecting greater potential potency, were considered for further investigation. These selected compounds were then analyzed through molecular docking studies to evaluate their binding affinities and interactions with the pKHG.
| ΔGbind = ΔGcomplex − (ΔGreceptor + ΔGligand) |
Then the MM-GBSA-based binding free energy (ΔG_bind) of the selected complex was calculated with the gmx_MMPBSA package.86,87 This approach integrates van der Waals, electrostatic, and solvation energy components, along with solvent-accessible surface area (SASA) contributions, providing a thermodynamic estimate of ligand affinity. The Desmond trajectory files were converted to GROMACS-compatible formats using Schrödinger utilities, while the corresponding topology files were generated through InterMol conversion of *.cms files to *.gro and *.top formats.88 The free energy associated with binding (ΔG_bind) was determined based on the following relationship:
| ΔGbind = 〈GPL〉 − 〈GP〉 − 〈GL〉 |
In this equation, 〈GPL〉, 〈GP〉, and 〈GL〉 correspond to the average free energies of the complex, the unbound protein, and the free ligand, respectively.
Accordingly, the overall binding energy is given by:
| ΔGbind = ΔEMM + ΔGSOLV − TΔS |
In order to supplement these results, the ProTox-III server (https://tox-new.charite.de/protox_III) was employed to make predictions regarding different types of toxicity, such as hepatotoxicity, nephrotoxicity, cardiotoxicity, and neurotoxicity.90 It uses deep machine learning algorithms to train on experimental data to give confidence in estimating toxicity.
3. Results
3.1. Target identification
To identify the most influential proteins within the network, we applied eight topological methods using the CytoHubba plugin. For each method, the top 10 hub genes were determined. The results of these analyses are summarized in Fig. 3B, highlighting the proteins that consistently appear as central hubs across multiple metrics. These key hub genes (kHGs) are likely to play critical roles in lung cancer biology and represent promising targets for further investigation. Among all the identified KHGs, CDK1 exhibited the highest number of protein–protein connections, and therefore its interaction network is highlighted in Fig. 3A.
3.2. Target validation
Based on the integrated evidence from hub gene ranking within the PPI network and the enrichment analysis highlighting its strong involvement in significant pathways, CDK1 was prioritized as the prime key hub gene (pKHG) for downstream validation.3.3. Structure-based drug discovery targeting CDK1
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 667 compounds from the database with their IMPPAT IDs and SMILES. The Lipinski's Rule of Five (RO5) results reveal that 4236 compounds had zero (0) violations out of 5786 phytochemicals, indicating favorable drug-likeness profiles. Following the removal of duplicates, 2113 unique compounds were retained from all of that selected medicinal plants (Table S6). These compounds were then selected for further computational analyses.
667 compounds from the database with their IMPPAT IDs and SMILES. The Lipinski's Rule of Five (RO5) results reveal that 4236 compounds had zero (0) violations out of 5786 phytochemicals, indicating favorable drug-likeness profiles. Following the removal of duplicates, 2113 unique compounds were retained from all of that selected medicinal plants (Table S6). These compounds were then selected for further computational analyses.3.3.3.1. Model development, optimization and validation. To calculate the bioactivity of our selected compounds targeting CDK1, we choose the ChEMBL dataset (CHEMBL308) containing 3348 experimentally validated inhibitors and 4536 activities against CDK1 with their IC50 information (accessed on November 20, 2025). The aim was to develop a classical regression model for CDK1 inhibitors, so that we can calculate the IC50 and pIC50 values of our compound library based on the trained model. According to our study, we get random forest (RF), LighGBM, XGBoost, and Stacking as the best-performing models. The evaluation metrics indicated that the difference in R2 value for RF, LightGBM, XGBoost, and Stacking were 0.177, 0.117, 0.241, and 0.214, respectively (Fig. S3A). Thus, based on these findings, we observed that LightGBM showed the lowest difference in R2 values between the test and train datasets, indicating the best fit for prediction outcomes.
The LightGBM regression model among other machine-learning methods has been receiving considerable attention due to its excellent predictive accuracy, computational efficiency, and the capacity to handle large data volumes.94,95 As an ensemble of decision trees, LightGBM is well suited to capture nonlinear structure–activity relationships while maintaining fast training on large molecular descriptor and fingerprint sets. The LightGBM model was trained on the training set using optimized hyperparameters, including n_estimators, num_leaves, max_depth, learning_rate, subsample, colsample_bytree, min_child_samples, reg_alpha, and reg_lambda. Additionally, the LightGBM model tuned to improve generalization and reduce overfitting. Model performance was then evaluated on the test set using standard QSAR regression metrics: Mean Absolute Error (MAE), Mean Squared Error (MSE), Root Mean Squared Error (RMSE), and coefficient of determination (R2). These parameters offered a comprehensive assessment of how well the model's predictions aligned with the actual experimental data. The test model demonstrated impressive performance, with MAE, MSE, RMSE, and R2 values of 0.520, 0.544, 0.737, and 0.758, respectively, shown in Fig. S3A. The overall results suggest that the accuracy is comparatively higher than the others. Therefore, finally we selected the LightGBM model for predicting the bioactivity (pIC50) value of our selected natural compounds.
3.3.3.2. Application to bioactivity calculation and compound screening. Once the model's predictive performance was validated, it was used to virtually screen a curated phytocompound library of 2113 phytocompounds by predicting their pIC50 values using the optimized ML model. A total of 380 compounds showed predicted pIC50 > 6.5 (∼IC50 < 1 µM), suggesting potential inhibitory activity against the target and prioritizing them for follow-up validation (see Table S7).96 As the experimental and predicted values lie close to each other in the scatter plot (Fig. S3B), it indicates that the QSAR model is strong and has excellent predictive accuracy, as most data points closely align along the regression line. These results indicate the extent to which the model is useful in identifying potentially promising molecules that can be subjected to further investigation either with the use of molecular docking or through experimental studies to confirm their utility as a possible drug target.
| Compound CID | Compounds name | Source | Binding affinity by AutoDock | Binding affinity by GLIDE_XP |
|---|---|---|---|---|
| 115196 | Brassinolide | Camellia sinensis | −9.2 | −7.85 |
| 5317764 | Glycyrrhisoflavon | Glycyrrhiza glabra | −9.5 | −7.70 |
| 14218027 | Licoflavanone | Glycyrrhiza glabra | −9.5 | −7.69 |
| 487089 | 3-(3,4-Dihydroxyphenyl)-5,7-dihydroxy-6,8-bis(3-methylbut-2-enyl)chroman-4-one | Glycyrrhiza glabra | −9.2 | −6.80 |
| 174880 | Lactupicrin | Cichorium intybus | −9.4 | −6.70 |
| 16747683 | AZD5438 | — | −8.8 | −5.15 |
| Compound CID | Compounds name | Post-docking MMGBSA (ΔGBind) |
|---|---|---|
| 174880 | Lactupicrin | −50.44 |
| 487089 | 3-(3,4-Dihydroxyphenyl)-5,7-dihydroxy-6,8-bis(3-methylbut-2-enyl)chroman-4-one | −46.78 |
| 14218027 | Licoflavanone | −36.28 |
| 5317764 | Glycyrrhisoflavon | −36.15 |
| 115196 | Brassinolide | −27.81 |
| 16747683 | AZD5438 | −44.58 |
3.3.9.1. Root mean square deviation (RMSD). The root mean square deviation (RMSD) evaluates the structural integrity and conformational shifts in the docked complex,70 with elevated values signaling reduced stability and lower values reflecting stable system performance.97 The RMSD analysis showed that the apo protein (6GU7) presented the highest average structural deviation (3.245 Å), which is the highest conformational flexibility that is shown in the absence of a ligand. In contrast, all ligand-bound complexes exhibited markedly lower average RMSD values, indicating enhanced structural stability upon ligand binding. Among the screened compounds, CID_487089 was the compound with the lowest average RMSD (2.446 Å), which was then followed by CID_14218027 (2.470 Å), indicating that the compounds have a strong stabilizing interaction in the binding pocket. CID_174880 showed comparable stability to CID_14218027, with an average RMSD of 2.476 Å. The control ligand showed an average RMSD of 3.002 Å, comparable to the apo form, implying minimal stabilizing influence. Overall, the average RMSD results confirm that CID_487089 forms the most stable complex with 6GU7 during the MD simulation (Fig. 9A).
3.3.9.2. Root mean square fluctuation (RMSF). The residue-level dynamics of the system were examined through RMSF analysis over the equilibrated phase of the simulation. As expected, the apo protein showed the highest average fluctuation (1.571 Å), reflecting its greater structural freedom in the absence of a bound ligand. In contrast, all ligand-bound complexes exhibited noticeably lower average RMSF values, suggesting that ligand interaction contributes to a more stable and ordered residue environment. Among the screened compounds, CID_174880 exhibited the strongest stabilizing effect, as reflected by its lowest average RMSF value (1.181 Å), suggesting tight binding and effective restriction of residue mobility. The control compound showed a moderate decrease in flexibility (1.390 Å), indicating a reasonable but less pronounced stabilizing interaction. Meanwhile, CID_487089 and CID_14218027 displayed intermediate stabilization, with average RMSF values of 1.253 Å and 1.375 Å, respectively, implying balanced binding interactions that reduce flexibility without overly restricting protein dynamics (Fig. 9B).
3.3.9.3. Solvent-accessible surface area (SASA). SASA reflects the solvent-exposed surface of a protein, with lower values generally indicating greater ligand burial and potentially stronger binding.98 Analysis of the 6GU7 protein complexes revealed notable differences in solvent-accessible surface area (SASA) among the ligands. The 6GU7_487089 complex exhibited the highest average SASA value (148.17 Å2), indicating a relatively larger solvent-exposed surface and suggesting a less compact binding conformation. In contrast, the 6GU7_174880 complex showed the lowest average SASA value (126.15 Å2), implying that the ligand is more deeply buried within the binding pocket and forms a more compact and stable protein–ligand interface. The 6GU7_14218027 (130.14 Å2) and control complex (131.81 Å2) displayed intermediate SASA values, reflecting moderately compact structural arrangements with balanced solvent exposure. Overall, 6GU7_174880 demonstrated the most favorable SASA profile in terms of structural compactness, highlighting its potential as the most effective stabilizing ligand among the tested compounds (Fig. 9C).
3.3.9.4. The radius of gyration (Rg). The radius of gyration (Rg) is a key indicator of protein structural compactness and stability in molecular dynamics simulations. In this study, the control complex (6GU7_Control) exhibited a relatively high average Rg value (4.60 Å), indicating a less compact conformation. Similarly, the 6GU7_487089 complex showed the highest Rg value among the ligand-bound systems (4.69 Å), suggesting comparatively lower compactness and weaker structural stabilization. In contrast, 6GU7_14218027 (4.40 Å) and 6GU7_174880 (4.44 Å) displayed lower average Rg values, reflecting more compact and stable conformations. Among these, 6GU7_14218027 exhibited the lowest Rg value, indicating the highest degree of structural compactness and suggesting a stronger stabilizing effect on the protein. Overall, ligand binding generally enhanced protein compactness compared to the control, with 6GU7_14218027 emerging as the most effective stabilizer based on Rg analysis, followed closely by 6GU7_174880 (Fig. 9D).
3.4. Pharmacokinetics study
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values varied among the compounds, where CID_487089 (4.35) showed relatively higher lipophilicity compared to the control (2.51), while CID_174880 (1.87) and CID_14218027 (3.33) remained within an acceptable range. All selected phytochemicals showed high gastrointestinal (GI) absorption, similar to the control compound, indicating their potential suitability for oral administration. In addition, CID_487089 and CID_14218027 were predicted as non-substrates of P-glycoprotein (P-gp), similar to the control, whereas CID_174880 was identified as a P-gp substrate, which may slightly influence its cellular efflux behavior. None of the compounds violated Lipinski's rule of five, Veber's rule, Egan rule, or Ghose filter (0 violations of all compounds), and this once again indicates that they had good drug-likeness properties.99–102 None of the compounds, including the control, were predicted to be BBB permeant, which may reduce the possibility of central nervous system-related side effects.103 The bioavailability score was 0.55 for all selected compounds, which is comparable to the control and suggests favorable oral bioavailability. Regarding structural alerts, CID_14218027 demonstrated a more favorable profile with no PAINS alert and only one Brenk alert, whereas CID_487089 and CID_174880 showed relatively higher alert counts. Among the evaluated compounds, CID_14218027 exhibited broad-spectrum inhibition across all major CYP isoforms (CYP1A2, CYP2C19, CYP2C9, CYP2D6, and CYP3A4), indicating a high likelihood of metabolic interference and an increased risk of drug–drug interactions. In contrast, CID_487089 showed selective inhibition of CYP2C9 and CYP3A4, suggesting a comparatively moderate interaction risk. Notably, CID_174880 demonstrated no inhibitory activity against any of the assessed CYP enzymes, reflecting a more favorable metabolic profile with minimal risk of CYP-mediated interactions. The control compound also inhibited CYP2C9 and CYP3A4, further emphasizing that compounds with minimal or no CYP inhibition, such as CID_174880, are more desirable from a pharmacokinetic and safety perspective. The detailed physicochemical and ADME parameters of all compounds are summarized in Table S10, and their overall ADME profiles are visualized in Fig. 11.
P values varied among the compounds, where CID_487089 (4.35) showed relatively higher lipophilicity compared to the control (2.51), while CID_174880 (1.87) and CID_14218027 (3.33) remained within an acceptable range. All selected phytochemicals showed high gastrointestinal (GI) absorption, similar to the control compound, indicating their potential suitability for oral administration. In addition, CID_487089 and CID_14218027 were predicted as non-substrates of P-glycoprotein (P-gp), similar to the control, whereas CID_174880 was identified as a P-gp substrate, which may slightly influence its cellular efflux behavior. None of the compounds violated Lipinski's rule of five, Veber's rule, Egan rule, or Ghose filter (0 violations of all compounds), and this once again indicates that they had good drug-likeness properties.99–102 None of the compounds, including the control, were predicted to be BBB permeant, which may reduce the possibility of central nervous system-related side effects.103 The bioavailability score was 0.55 for all selected compounds, which is comparable to the control and suggests favorable oral bioavailability. Regarding structural alerts, CID_14218027 demonstrated a more favorable profile with no PAINS alert and only one Brenk alert, whereas CID_487089 and CID_174880 showed relatively higher alert counts. Among the evaluated compounds, CID_14218027 exhibited broad-spectrum inhibition across all major CYP isoforms (CYP1A2, CYP2C19, CYP2C9, CYP2D6, and CYP3A4), indicating a high likelihood of metabolic interference and an increased risk of drug–drug interactions. In contrast, CID_487089 showed selective inhibition of CYP2C9 and CYP3A4, suggesting a comparatively moderate interaction risk. Notably, CID_174880 demonstrated no inhibitory activity against any of the assessed CYP enzymes, reflecting a more favorable metabolic profile with minimal risk of CYP-mediated interactions. The control compound also inhibited CYP2C9 and CYP3A4, further emphasizing that compounds with minimal or no CYP inhibition, such as CID_174880, are more desirable from a pharmacokinetic and safety perspective. The detailed physicochemical and ADME parameters of all compounds are summarized in Table S10, and their overall ADME profiles are visualized in Fig. 11.
| Target | CID_487089 | CID_174880 | CID_14218027 | Control (AZD5438) |
|---|---|---|---|---|
| Hepatotoxicity | Inactive | Inactive | Inactive | Inactive |
| Neurotoxicity | Inactive | Inactive | Inactive | Inactive |
| Cardiotoxicity | Inactive | Inactive | Inactive | Inactive |
| Respiratory toxicity | Active | Active | Active | Active |
| Immunotoxicity | Active | Active | Active | Inactive |
| Carcinogenicity | Inactive | Inactive | Inactive | Active |
| Mutagenicity | Inactive | Inactive | Inactive | Inactive |
| Cytotoxicity | Inactive | Inactive | Inactive | Inactive |
| LD50 mg kg−1 | 10 | 300 | 2000 | 500 |
| Toxicity class | 2 | 3 | 4 | 4 |
4. Discussion
Lung cancer is one of the most detrimental malignancies in the entire world. It grows fast, is often detected late, and does not respond well to available treatments.2 These challenges highlight the need for new treatments guided by a clear strategy for finding targets and developing drugs. Using transcriptomic profiling, network analysis, pathway studies, and computer-based drug discovery, this research systematically identified a key molecular target in lung cancer and proposed a drug to target it. Transcriptomics is the study of all RNA molecules produced in a cell, helping us understand which genes are active and how their activity changes under different conditions.104 In cancer research, it's often used to find important targets because cancer cells tend to show unusual patterns of gene activity compared to normal cells.105 However, the results from a single dataset can be affected by technical differences, experimental platforms, or variations among populations. To address this, four independent GEO microarray datasets (GSE19804, GSE10072, GSE18842, and GSE10799) were analyzed across different patient groups (case and control). From this combined analysis, 88 genes were consistently upregulated and 294 genes were downregulated. Repeated changes in these certain genes suggest a real connection to this cancer. After identifying these common differentially expressed genes, we studied how their proteins interact. The strong enrichment of the protein–protein interaction (PPI) network indicates that these connections are not random, showing a real biological relationship between these genes in lung cancer. Then, based on this PPI network we identified the major key hub genes (kHGs) using several analytical methods, which ensured that our results were robust and not dependent on a single approach. Among these interactions, cyclin dependent kinase 1 (CDK1) showed the highest number of interactions, suggesting that it may play a central role in regulating important cancer-related processes. To clarify the regulation of kHGs, we analyzed their interactions with transcription factors (TFs) and microRNAs (miRNAs). Network analysis (Fig. 4) identified FOXC1, GATA2, YY1, E2F1, and HINFP as major TFs, while hsa-miR-192-5p, hsa-miR-92a-3p, hsa-miR-193b-3p, hsa-miR-215-5p, and hsa-miR-155-5p were identified as central miRNA regulators. The study of TFs and miRNAs revealed important hints on the regulation of kHGs at the different levels. Transcription factors are known to activate the expression of genes,106 whereas the miRNAs are known to regulate the activity of genes once they are transcribed.107 This combined method enabled us to determine vital regulatory factors which could contribute to dysregulated gene expression in lung cancer and gives a clear understanding for further functional and therapeutic studies. Moreover, we performed functional enrichment analyses (GO and KEGG) using multiple pathway databases to understand the role of these hub genes in lung cancer cells. These enrichment results indicate that lung cancer progression is driven by widespread disruption of cell cycle regulation, mitotic structure, and signaling control rather than by isolated gene alterations. The overall involvement of CDK1 in key mitotic events, strong kinase activity, and extensive interaction with other cell cycle regulators suggest that it functions as a major coordinator of uncontrolled cell division in lung cancer. Thus, this central and recurring role makes CDK1 the most promising candidate for further functional validation and therapeutic exploration.The reason why LUAD was chosen as a validation is because it is the most prevalent type of lung cancer and it accounts for a substantial percentage of the cases of lung cancer, compared to other types.108 Multiple analyses of expressions in different independent platforms indicated that CDK1 is not only transcriptionally upregulated but also overexpressed at the protein level in LUAD. In addition, the patients whose CDK1 level was higher had poorer survival and earlier recurrence, which is an obvious clinical effect. In addition to expression, the correlation of CDK1 and immune cell infiltration indicates that its activity moves beyond the expression to influence the tumor microenvironment, especially at the T-cell subsets and macrophage polarization. This implies that CDK1 can regulate tumor growth and immune response. Lastly, the somatic mutations and copy number changes observed in CDK1 contribute to the oncogenic significance of this protein in a group of LUAD patients. From a biological perspective, CDK1 is more than a statistically prioritized hub gene. As a key regulator of the G2/M transition and mitotic entry, CDK1 promotes sustained tumor cell proliferation when aberrantly activated. Prior studies in lung cancer have also linked elevated CDK1 expression with poor survival, enhanced cell-cycle and DNA-repair signaling, and altered immune-associated pathways in LUAD.109,110 These observations strengthen the view that CDK1 may act as a functionally important driver of lung cancer progression and a rational candidate for therapeutic targeting.
In this study, we used a structure-based drug discovery approach to efficiently screen a large library of phytochemicals.111 In this field, the integration of machine learning (ML), molecular docking, and molecular dynamics (MD) simulation studies has transformed the identification and optimization of novel drug candidates.112,113 We performed cheminformatics-based screening procedure to screen physicochemical and drug-likeness properties of 9577 phytocompounds. After removing duplicate entries and unwanted sub-structures, we finally obtained 2113 phytocompounds, among which 1802 showed no violations of RO5 and Veber's rule. ML is revolutionizing the computational drug discovery approach by reducing traditional experimental time and cost. ML is the best option to calculate the bioactivity of a large dataset with high accuracy within a very short time.114 To identify the most promising candidates, we applied a ML-based pIC50 prediction model, which helped us identify compounds with higher chances of biological activity. This step reduced the dataset to identify most potential 380 phytocompounds with pIC50 > 6.5 for further computational analysis, including molecular docking, molecular dynamics simulation, and pharmacokinetics analysis. The docking results and post docking MM-GBSA binding energy (ΔG) highlighted the top-ranked three phytochemicals (CID_487089, CID_174880, and CID_14218027) as candidate drug molecules.
Furthermore, molecular dynamics (MD) simulations were used to understand how the selected phytochemicals interact with CDK1 over time. While the apo protein and AZD5438-bound complex showed noticeable structural fluctuations, all phytochemical-bound systems exhibited improved stability throughout the 100 ns simulation. Notably, CID_174880 consistently outperformed the control compound by maintaining lower RMSD and RMSF values (Fig. 9), indicating a stronger and more stable binding mode. Structural compactness analyses further highlighted this difference. The CID_174880 complex showed the lowest SASA and a tightly maintained Rg, reflecting deeper ligand burial and a more compact protein structure than AZD5438.115 In contrast, the control complex remained relatively solvent-exposed and flexible, suggesting weaker stabilization of CDK1. Post-simulation MM-GBSA results further supported these findings, with CID_487089 showing the most favorable binding free energy (−40.29 kcal mol−1), followed by CID_174880 (−36.06 kcal mol−1), both outperforming the control AZD5438 (−29.53 kcal mol−1). Dynamic motion analysis using PCA showed clear differences between the control compound (AZD5438) and the phytochemicals,116 which suggests a better fit between the ligand and the binding site. Also, the FEL analysis further supported these findings by demonstrating the stability and conformational states of the ligand–protein interactions. Among the studied compounds, CID_174880 showed a dynamic and energy profile closely comparable to the control, suggesting favorable binding stability. The control compound was predicted to be carcinogenic, while none of the phytochemicals showed this risk, indicating better overall safety. However, all compounds exhibited some potential respiratory toxicity, and the phytochemicals also showed immunotoxic effects, which may limit their development. Based on LD50 values and toxicity classes, CID_14218027 was the safest (Class 4), whereas CID_487089 showed higher toxicity (Class 2). CID_174880 displayed a toxicity profile close to the control but with slightly improved safety. Overall, CID_174880 demonstrated promising inhibitory potential with comparatively better safety profiles than the control compound. Therefore, this study could be a useful resource to identify natural CDK1 inhibitors after being validated through the experimental (in vivo & in vitro) study.
5. Conclusion
This paper displays CDK1 as an important lung cancer driver, disrupting the cell cycle, p53 pathway, and immune microenvironment, such as augmented M1 macrophages, diminished M2 polarization, and unfavorable prognosis owing to overexpression and mutations, including I136N and E57V. These discoveries indicate a shared vulnerability in lung cancer, which puts mitotic dysregulation as one of the prospective treatment targets. Among the phytochemicals evaluated, lactupicrin (CID_174880), derived from Cichorium intybus, emerged as the most promising in silico prioritized candidate compared with the reference compound AZD5438, based on its favorable docking performance, structural stability, dynamic behavior, and predicted pharmacokinetic and toxicity profiles. Although minor fluctuations were observed in some metrics, all CDK1 complexes maintained overall stability and compatibility. Docking results, MD simulations, MM-GBSA calculations and ADMET analyses consistently support lactupicrin as a computationally prioritized candidate for further development. Further experimental validation, including in vitro biochemical assays, cell-based functional studies, and in vivo investigations, is required to confirm CDK1 inhibitory activity, anticancer efficacy, and safety. Overall, this research provides a clear workflow for identifying effective CDK1-targeting compounds and offers a roadmap for developing broad-spectrum, mechanism-based therapies for lung cancer, bridging computational predictions with potential real-world applications.6. Limitations of this study
Although this study applied a comprehensive multiomics and in silico approach, several limitations should be considered. First, the transcriptomic analysis was based solely on publicly available microarray datasets, which may be affected by batch effects, platform-related biases, and limited clinical information. While using four independent cohorts improves reliability, the absence of RNA-seq data may limit the ability to capture finer gene expression variability. Second, the drug discovery pipeline, including machine learning predictions, molecular docking, MM-GBSA analysis, MD simulations, and ADEMT, relies entirely on computational models. Although these methods are useful for prioritizing candidates, they cannot fully represent the complexity of biological systems, tumor heterogeneity, or real pharmacodynamic behavior. Finally, the identified drug candidates, especially lactupicrin, have not yet been validated in vitro or in vivo models. Therefore, the therapeutic potential, optimal dosage, and safety profiles of the candidate drugs still need to be confirmed through experimental validation.Author contributions
conceptualization – Md. Ahad Ali and Hriddhi Sarker; methodology –Md. Ahad Ali, Hriddhi Sarker, and Humaira Sheikh; data curation – Hriddhi Sarker, Marguba Kamrun, Bilkis Akter Shifa, and Sujoy Banik; formal analysis – Md. Ahad Ali, Hriddhi Sarker, Humaira Sheikh, and Siam Ahmed; visualization – Hriddhi Sarker, Md. Ahad Ali, Marguba Kamrun, and Tarikul Islam; validation – Md. Ahad Ali, Hriddhi Sarker, and Tarikul Islam; project administration – Md. Ahad Ali; software & resources – Neeraj Kumar, Sujoy Banik, and Bilkis Akter Shifa; supervision – Md. Ahad Ali; writing – original draft, Hriddhi Sarker, Md. Ahad Ali, and Humaira Sheikh; writing – review & editing, Md. Ahad Ali, Marguba Kamrun, and Neeraj Kumar.Conflicts of interest
All authors declare no conflict of interests.Data availability
The original data and contributions presented in this study are included in the article and its supplementary information (SI) (Tables S1–S10 and Fig. S1–S5). The training set of our selected compounds, the test set, the chEMBL datasets (containing the consensus predictions) and the related python code for running the QSAR models to predict the bioactivity of the selected compounds can be found on our GitHub repository (https://github.com/ahad004/LUAD_ML_QSAR_Modeling) or can be accessed using the following DOI: https://doi.org/10.5281/zenodo.19841200.Supplementary information is available. See DOI: https://doi.org/10.1039/d6dd00045b.
References
- Y. Ji, Y. Zhang, S. Liu, J. Li, Q. Jin, J. Wu, H. Duan, X. Liu, L. Yang and Y. Huang, The Epidemiological Landscape of Lung Cancer: Current Status, Temporal Trend and Future Projections Based on the Latest Estimates from GLOBOCAN 2022, J. Natl. Cancer Cent., 2025, 5, 278–286, DOI:10.1016/j.jncc.2025.01.003.
- F. Bray, M. Laversanne, H. Sung, J. Ferlay, R. L. Siegel, I. Soerjomataram and A. Jemal, Global Cancer Statistics 2022: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries, CA Cancer J. Clin., 2024, 74, 229–263, DOI:10.3322/caac.21834.
- J. Zhou, Y. Xu, J. Liu, L. Feng, J. Yu and D. Chen, Global Burden of Lung Cancer in 2022 and Projections to 2050: Incidence and Mortality Estimates from GLOBOCAN, J. Cancer Epidemiol., 2024, 93, 102693, DOI:10.1016/j.canep.2024.102693.
- S. Bharadwaj, J. M. Mierzwicka, L. Vaňková and P. Malý, Unraveling the Molecular-Pathological Characteristics and Cellular Complexity of the Tumor Immune Microenvironment in Metastatic Non-Small Cell Lung Cancer, Cell Commun. Signal., 2025, 23, 400, DOI:10.1186/s12964-025-02410-w.
- Z. Gu, Y. Heng, R. Fan, J. Luo and L. Ju, Single-Cell RNA Sequencing Reveals Cellular and Molecular Heterogeneity in Extensive-Stage Small Cell Lung Cancer with Different Chemotherapy Responses, Cancer Cell Int., 2025, 25, 157, DOI:10.1186/s12935-025-03785-z.
- A. Jachowski, M. Marcinkowski, J. Szydłowski, O. Grabarczyk, Z. Nogaj, Ł. Marcin, A. Pławski, P. P. Jagodziński and B. K. Słowikowski, Modern Therapies of Nonsmall Cell Lung Cancer, J. Appl. Genet., 2023, 64, 695–711, DOI:10.1007/s13353-023-00786-4.
- M. Araghi, R. Mannani, A. Heidarnejad maleki, A. Hamidi, S. Rostami, S. H. Safa, F. Faramarzi, S. Khorasani, M. Alimohammadi and S. Tahmasebi, et al., Recent Advances in Non-Small Cell Lung Cancer Targeted Therapy; an Update Review, Cancer Cell Int., 2023, 23, 162, DOI:10.1186/s12935-023-02990-y.
- P.-L. Su, N. Furuya, A. Asrar, C. Rolfo, Z. Li, D. P. Carbone and K. He, Recent Advances in Therapeutic Strategies for Non-Small Cell Lung Cancer, J. Hematol. Oncol., 2025, 18, 35, DOI:10.1186/s13045-025-01679-1.
- L. Valdez Capuccino, T. Kleitke, B. Szokol, L. Svajda, F. Martin, F. Bonechi, M. Krekó, S. Azami, A. Montinaro and Y. Wang, et al., CDK9 Inhibition as an Effective Therapy for Small Cell Lung Cancer, Cell Death Dis., 2024, 15, 345, DOI:10.1038/s41419-024-06724-4.
- A. Osoegawa, Y. Takumi, T. Hashimoto, S. Nakatsuji, M. Hori, M. Sakai, T. Karashima, M. Abe, M. Miyawaki and K. Sugio, Cyclin-Dependent Kinase (CDK) 4/6 Inhibition in Non-Small Cell Lung Cancer with Epidermal Growth Factor Receptor (EGFR) Mutations, Invest. New Drugs, 2023, 41, 183–192, DOI:10.1007/s10637-023-01337-8.
- E. Panagiotou, G. Gomatou, I. P. Trontzas, N. Syrigos and E. Kotteas, Cyclin-Dependent Kinase (CDK) Inhibitors in Solid Tumors: A Review of Clinical Trials, Clin. Transl. Oncol., 2022, 24, 161–192, DOI:10.1007/s12094-021-02688-5.
- X. Huang, Y. Yin, G. Saha, I. Francis and S. C. Saha, A Comprehensive Numerical Study on the Transport and Deposition of Nasal Sprayed Pharmaceutical Aerosols in a Nasal-To-Lung Respiratory Tract Model, Part. Part. Syst. Charact., 2025, 42, 2400004, DOI:10.1002/ppsc.202400004.
- X.-Q. Li, X.-J. Cheng, J. Wu, K.-F. Wu and T. Liu, Targeted Inhibition of the PI3K/AKT/MTOR Pathway by (+)-Anthrabenzoxocinone Induces Cell Cycle Arrest, Apoptosis, and Autophagy in Non-Small Cell Lung Cancer, Cell. Mol. Biol. Lett., 2024, 29, 58, DOI:10.1186/s11658-024-00578-6.
- H. Kitai, P. H. Choi, Y. C. Yang, J. A. Boyer, A. Whaley, P. Pancholi, C. Thant, J. Reiter, K. Chen and V. Markov, et al., Combined Inhibition of KRASG12C and MTORC1 Kinase Is Synergistic in Non-Small Cell Lung Cancer, Nat. Commun., 2024, 15, 6076, DOI:10.1038/s41467-024-50063-z.
- J.-L. Huang, L.-M. Wu, S.-Q. Wu, F.-Y. Yuan, H.-Z. Weng, D. Huang, L. Gan, S.-B. Chen, G.-H. Tang and S. Yin, A Small Molecule Targets LIC1 to Suppress Lung Tumor Growth by Inducing Autophagy, Nat. Chem. Biol., 2026, 22, 459–470, DOI:10.1038/s41589-025-02040-w.
- J. Crawford, D. Herndon, K. Gmitter and J. Weiss, The Impact of Myelosuppression on Quality of Life of Patients Treated with Chemotherapy, Future Oncol., 2024, 20, 1515–1530, DOI:10.2217/fon-2023-0513.
- C. Lazzari, V. Gregorc, N. Karachaliou, R. Rosell and M. Santarpia, Mechanisms of Resistance to Osimertinib, J. Thorac. Dis., 2020, 12, 2851–2858, DOI:10.21037/jtd.2019.08.30.
- L. Astolfi, S. Ghiselli, V. Guaran, M. Chicca, E. Simoni, E. Olivetto, G. Lelli and A. Martini, Correlation of Adverse Effects of Cisplatin Administration in Patients Affected by Solid Tumours: A Retrospective Evaluation, Oncol. Rep., 2013, 29, 1285–1292, DOI:10.3892/or.2013.2279.
- E. S. Alsatari, K. R. Smith, S. P. L. Galappaththi, E. A. Turbat-Herrera and S. Dasgupta, The Current Roadmap of Lung Cancer Biology, Genomics and Racial Disparity, Int. J. Mol. Sci., 2025, 26, 3818, DOI:10.3390/ijms26083818.
- G. Gupta, V. P. Samuel, M. M. Rekha, B. Rani, Y. Sasikumar, P. P. Nayak, P. Sudan, K. Goyal, B. G. Oliver and A. Chakraborty, et al., Caspase-Independent Cell Death in Lung Cancer: From Mechanisms to Clinical Applications, Naunyn-Schmiedebergs Arch Pharmacol, 2025, 398, 13031–13048, DOI:10.1007/s00210-025-04149-0.
- J. Zhang, X. Zeng, Q. Guo, Z. Sheng, Y. Chen, S. Wan, L. Zhang and P. Zhang, Small Cell Lung Cancer: Emerging Subtypes, Signaling Pathways, and Therapeutic Vulnerabilities, Exp. Hematol. Oncol., 2024, 13, 78, DOI:10.1186/s40164-024-00548-w.
- N. MISHRA, A. SONI, M. KUMARI, G. SINGH, S. K. SHARMA and S. K. SINGH, Targeting Cell Cycle Regulators: A New Paradigm in Cancer Therapeutics, Biocell, 2024, 48, 1639–1666, DOI:10.32604/biocell.2024.056503.
- A. Alibakhshi, A. Alagheband Bahrami, E. Mohammadi, S. Ahangarzadeh and M. Mobasheri, In-Silico Design of a New Multi-Epitope Vaccine Candidate against SARS-CoV-2, Acta Virol., 2024, 67, 12481, DOI:10.3389/av.2023.12481.
- F. S. Abdel Razek, S. D. Ibrahim, A. A. Megahed, A. S. Sadik and S. S. A. El-Masry, In Silico Molecular Docking Analysis of Green Tea Bioactive Compounds Targeting Banana Bunchy Top Virus Proteins, Discov. Appl. Sci., 2025, 7, 1232, DOI:10.1007/s42452-025-07466-4.
- Q. M. S. Jamal, S. Khan, M. Khan, A. A. Ansai, J. M. Ashraf, M. Habibullah, A. Farasani, A. M. Madkhali and M. Lohani, Smoking May Increase the Risk of COVID-19 Infection: Evidence from In Silico Analysis, J. Pharm. Res. Int., 2021, 12–21, DOI:10.9734/jpri/2021/v33i22B31394.
- Z. Sajid, T. Akhtar, K. Ahmad and M. Haroon, Molecular Docking Simulation and ADMET/Pharmacokinetic Screening of Newly Designed 2-(2-(Aryl)-4-oxo-4,5-dihydrothiazol-5-yl)Acetohydrazides as Potential Antitubercular Agents, ChemistrySelect, 2024, 9(44), e202403715, DOI:10.1002/slct.202403715.
- P. Chunarkar-Patil, M. Kaleem, R. Mishra, S. Ray, A. Ahmad, D. Verma, S. Bhayye, R. Dubey, H. N. Singh and S. Kumar, Anticancer Drug Discovery Based on Natural Products: From Computational Approaches to Clinical Studies, Biomedicines, 2024, 12(1), 201, DOI:10.3390/biomedicines12010201.
- M. Huang, J.-J. Lu and J. Ding, Natural Products in Cancer Therapy: Past, Present and Future, Nat. Prod. Bioprospect., 2021, 11, 5–13, DOI:10.1007/s13659-020-00293-7.
- S. T. Asma, U. Acaroz, K. Imre, A. Morar, S. R. A. Shah, S. Z. Hussain, D. Arslan-Acaroz, H. Demirbas, Z. Hajrulai-Musliu and F. R. Istanbullugil, et al., Natural Products/Bioactive Compounds as a Source of Anticancer Drugs, Cancers, 2022, 14, 6203, DOI:10.3390/cancers14246203.
- G. M. Cragg and D. J. Newman, Natural Products: A Continuing Source of Novel Drug Leads, Biochim. Biophys. Acta Gen. Subj., 2013, 1830, 3670–3695, DOI:10.1016/j.bbagen.2013.02.008.
- K. Mohanraj, B. S. Karthikeyan, R. P. Vivek-Ananth, R. P. B. Chand, S. R. Aparna, P. Mangalapandi and A. Samal, IMPPAT: A Curated Database of Indian Medicinal Plants, Phytochemistry and Therapeutics, Sci. Rep., 2018, 8, 4329, DOI:10.1038/s41598-018-22631-z.
- S. M. Soyer, P. Ozbek and C. Kasavi, Lung Adenocarcinoma Systems Biomarker and Drug Candidates Identified by Machine Learning, Gene Expression Data, and Integrative Bioinformatics Pipeline, OMICS J. Integr. Biol., 2024, 28, 408–420, DOI:10.1089/omi.2024.0121.
- Y. Li, Y. Cai, L. Ji, B. Wang, D. Shi and X. Li, Machine Learning and Bioinformatics Analysis of Diagnostic Biomarkers Associated with the Occurrence and Development of Lung Adenocarcinoma, PeerJ, 2024, 12, e17746, DOI:10.7717/peerj.17746.
- W. Roh, Y. Geffen, H. Cha, M. Miller, S. Anand, J. Kim, D. I. Heiman, J. F. Gainor, P. W. Laird and A. D. Cherniack, et al., High-Resolution Profiling of Lung Adenocarcinoma Identifies Expression Subtypes with Specific Biomarkers and Clinically Relevant Vulnerabilities, Cancer Res., 2022, 82, 3917–3931, DOI:10.1158/0008-5472.CAN-22-0432.
- R. X. Huang, D. Siriwanna, W. C. Cho, T. K. Wan, Y. R. Du, A. N. Bennett, Q. E. He, J. D. Liu, X. T. Huang and K. H. K. Chan, Lung Adenocarcinoma-Related Target Gene Prediction and Drug Repositioning, Front. Pharmacol, 2022, 13, 936758, DOI:10.3389/fphar.2022.936758.
- Y. Li, K. Ma, H. Wang, Z. Liu and Z. Li, Identification of Therapeutic Targets in Lung Adenocarcinoma Using Mendelian Randomization and Multi-Omics, Discov. Oncol., 2025, 16(1), 10258, DOI:10.1007/s12672-025-02835-2.
- K. Jia, Y. Wang, Q. Cao and Y. Wang, Extensive Prediction of Drug Response in Mutation-Subtype-Specific LUAD with Machine Learning Approach, Oncol. Res., 2024, 32, 409–419, DOI:10.32604/or.2023.042863.
- R. Qureshi, S. A. Basit, J. A. Shamsi, X. Fan, M. Nawaz, H. Yan and T. Alam, Machine Learning Based Personalized Drug Response Prediction for Lung Cancer Patients, Sci. Rep., 2022, 12, 18935, DOI:10.1038/s41598-022-23649-0.
- M. Sobhan and A. M. Mondal, Explainable Machine Learning to Identify Patient-Specific Biomarkers for Lung Cancer, in. Proc. – 2022 IEEE Int. Conf. Bioinforma. Biomed. BIBM, 2022, pp. 3152–3159, DOI:10.1109/BIBM55620.2022.9995516.
- T. M. Okyay, I. Yilmaz and M. Koldas, Machine Learning-Based Bioactivity Prediction of Porphyrin Derivatives: Molecular Descriptors, Clustering, and Model Evaluation, Photochem. Photobiol. Sci., 2025, 24, 923–937, DOI:10.1007/s43630-025-00733-8.
- T.-P. Lu, M.-H. Tsai, J.-M. Lee, C.-P. Hsu, P.-C. Chen, C.-W. Lin, J.-Y. Shih, P.-C. Yang, C. K. Hsiao and L.-C. Lai, et al., Identification of a Novel Biomarker, SEMA5A, for Non-Small Cell Lung Carcinoma in Nonsmoking Women, Cancer Epidemiol. Biomarkers Prev., 2010, 19, 2590–2597, DOI:10.1158/1055-9965.EPI-10-0332.
- T.-P. Lu, C. K. Hsiao, L.-C. Lai, M.-H. Tsai, C.-P. Hsu, J.-M. Lee and E. Y. Chuang, Identification of Regulatory SNPs Associated with Genetic Modifications in Lung Adenocarcinoma, BMC Res. Notes, 2015, 8, 92, DOI:10.1186/s13104-015-1053-8.
- M. T. Landi, T. Dracheva, M. Rotunno, J. D. Figueroa, H. Liu, A. Dasgupta, F. E. Mann, J. Fukuoka, M. Hames and A. W. Bergen, et al., Gene Expression Signature of Cigarette Smoking and Its Role in Lung Adenocarcinoma Development and Survival, PLoS One, 2008, 3, e1651, DOI:10.1371/journal.pone.0001651.
- A. Sanchez-Palencia, M. Gomez-Morales, J. A. Gomez-Capilla, V. Pedraza, L. Boyero, R. Rosell and M. E. Fárez-Vidal, Gene Expression Profiling Reveals Novel Biomarkers in Nonsmall Cell Lung Cancer, Int. J. Cancer, 2011, 129, 355–364, DOI:10.1002/ijc.25704.
- M. Wrage, S. Ruosaari, P. P. Eijk, J. T. Kaifi, J. Hollmén, E. F. Yekebas, J. R. Izbicki, R. H. Brakenhoff, T. Streichert and S. Riethdorf, et al., Genomic Profiles Associated with Early Micrometastasis in Lung Cancer: Relevance of 4q Deletion, Clin. Cancer Res., 2009, 15, 1566–1574, DOI:10.1158/1078-0432.CCR-08-2188.
- G. K. Smyth, Linear Models and Empirical Bayes Methods for Assessing Differential Expression in Microarray Experiments, Stat. Appl. Genet. Mol. Biol., 2004, 3, 3, DOI:10.2202/1544-6115.1027.
- G. Singh and B. Soman, Data Transformation Using Dplyr Package in R, 2019 Search PubMed.
- P. M. Valero-Mora, Ggplot2: Elegant Graphics for Data Analysis, J. Stat. Softw., 2010, 35(1), 1–3, DOI:10.18637/jss.v035.b01.
- C. H. Gao, G. Yu and P. Cai, GgVennDiagram: An Intuitive, Easy-to-Use, and Highly Customizable R Package to Generate Venn Diagram, Front. Genet., 2021, 12, 706907, DOI:10.3389/fgene.2021.706907.
- D. Szklarczyk, J. H. Morris, H. Cook, M. Kuhn, S. Wyder, M. Simonovic, A. Santos, N. T. Doncheva, A. Roth and P. Bork, et al., The STRING Database in 2017: Quality-Controlled Protein–Protein Association Networks, Made Broadly Accessible, Nucleic Acids Res., 2017, 45, D362–D368, DOI:10.1093/nar/gkw937.
- P. Shannon, A. Markiel, O. Ozier, N. S. Baliga, J. T. Wang, D. Ramage, N. Amin, B. Schwikowski and T. C. Ideker, A Software Environment for Integrated Models of Biomolecular Interaction Networks, Genome Res., 2003, 13, 2498–2504, DOI:10.1101/gr.1239303.
- C.-H. Chin, S.-H. Chen, H.-H. Wu, C.-W. Ho, M.-T. Ko and C.-Y. Lin, CytoHubba: Identifying Hub Objects and Sub-Networks from Complex Interactome, BMC Syst. Biol., 2014, 8, S11, DOI:10.1186/1752-0509-8-S4-S11.
- A. Khan, O. Fornes, A. Stigliani, M. Gheorghe, J. A. Castro-Mondragon, R. Van Der Lee, A. Bessy, J. Chèneby, S. R. Kulkarni and G. Tan, et al., JASPAR 2018: Update of the Open-Access Database of Transcription Factor Binding Profiles and Its Web Framework, Nucleic Acids Res., 2018, 46, D260–D266, DOI:10.1093/nar/gkx1126.
- S.-D. Hsu, F.-M. Lin, W.-Y. Wu, C. Liang, W.-C. Huang, W.-L. Chan, W.-T. Tsai, G.-Z. Chen, C.-J. Lee and C.-M. Chiu, et al., MiRTarBase: A Database Curates Experimentally Validated MicroRNA–Target Interactions, Nucleic Acids Res., 2011, 39, D163–D169, DOI:10.1093/nar/gkq1107.
- J. Xia, E. E. Gill and R. E. W. Hancock, NetworkAnalyst for Statistical, Visual and Network-Based Meta-Analysis of Gene Expression Data, Nat. Protoc., 2015, 10, 823–844, DOI:10.1038/nprot.2015.052.
- B. T. Sherman, M. Hao, J. Qiu, X. Jiao, M. W. Baseler, H. C. Lane, T. Imamichi and W. Chang, DAVID: A Web Server for Functional Enrichment Analysis and Functional Annotation of Gene Lists (2021 Update), Nucleic Acids Res., 2022, 50, W216–W221, DOI:10.1093/nar/gkac194.
- M. V. Kuleshov, M. R. Jones, A. D. Rouillard, N. F. Fernandez, Q. Duan, Z. Wang, S. Koplev, S. L. Jenkins, K. M. Jagodnik and A. Lachmann, et al., Enrichr: A Comprehensive Gene Set Enrichment Analysis Web Server 2016 Update, Nucleic Acids Res., 2016, 44, W90–W97, DOI:10.1093/nar/gkw377.
- M. Helmy, R. Agrawal, J. Ali, M. Soudy, T. T. Bui and K. Selvarajoo, GeneCloudOmics: A Data Analytic Cloud Platform for High-Throughput Gene Expression Analysis, Front. Bioinform., 2021, 1, 693836, DOI:10.3389/fbinf.2021.693836.
- T. Li, J. Fu, Z. Zeng, D. Cohen, J. Li, Q. Chen, B. Li and X. S. Liu, TIMER2.0 for Analysis of Tumor-Infiltrating Immune Cells, Nucleic Acids Res., 2020, 48(W1), W509–W514, DOI:10.1093/NAR/GKAA407.
- Z. Tang, B. Kang, C. Li, T. Chen and Z. Zhang, GEPIA2: An Enhanced Web Server for Large-Scale Expression Profiling and Interactive Analysis, Nucleic Acids Res., 2019, 47, W556–W560, DOI:10.1093/nar/gkz430.
- D. S. Chandrashekar, S. K. Karthikeyan, P. K. Korla, H. Patel, A. R. Shovon, M. Athar, G. J. Netto, Z. S. Qin, S. Kumar and U. Manne, et al., UALCAN: An Update to the Integrated Cancer Data Analysis Platform, Neoplasia, 2022, 25, 18–27, DOI:10.1016/j.neo.2022.01.001.
- J. Gao, B. A. Aksoy, U. Dogrusoz, G. Dresdner, B. Gross, S. O. Sumer, Y. Sun, A. Jacobsen, R. Sinha and E. Larsson, et al., Integrative Analysis of Complex Cancer Genomics and Clinical Profiles Using the CBioPortal, Sci. Signal., 2013, 6(269), pl1, DOI:10.1126/scisignal.2004088.
- S. K. Burley, C. Bhikadiya, C. Bi, S. Bittrich, L. Chen, G. V. Crichlow, C. H. Christie, K. Dalenberg, L. Di Costanzo and J. M. Duarte, et al., RCSB Protein Data Bank: Powerful New Tools for Exploring 3D Structures of Biological Macromolecules for Basic and Applied Research and Education in Fundamental Biology, Biomedicine, Biotechnology, Bioengineering and Energy Sciences, Nucleic Acids Res., 2021, 49, D437–D451, DOI:10.1093/nar/gkaa1038.
- BIOVIA, D. S., Discovery Studio Visualizer V21.1.0.20298. BIOVIA, Dassault Systèmes, 2005 Search PubMed.
- N. Guex and M. C. Peitsch, SWISS-MODEL and the Swiss-Pdb Viewer: An Environment for Comparative Protein Modeling, Electrophoresis, 1997, 18, 2714–2723, DOI:10.1002/elps.1150181505.
- A. A. Puelles, L. L. Bastos, V. M. Paixão, S. C. Araujo and R. C. de Melo Minardi In. Virtual Screening. 2024, pp. 209–236 Search PubMed.
- G. Xiong, Z. Wu, J. Yi, L. Fu, Z. Yang, C. Hsieh, M. Yin, X. Zeng, C. Wu and A. Lu, et al., ADMETlab 2.0: An Integrated Online Platform for Accurate and Comprehensive Predictions of ADMET Properties, Nucleic Acids Res., 2021, 49, W5–W14, DOI:10.1093/nar/gkab255.
- C. A. Lipinski, Lead- and Drug-like Compounds: The Rule-of-Five Revolution, Drug Discov. Today Technol., 2004, 1, 337–341, DOI:10.1016/j.ddtec.2004.11.007.
- W. Caldwell, Z. Yan, W. Lang and A. Masucci, J. The IC50 Concept Revisited, Curr. Top. Med. Chem., 2012, 12(11), 1282–1290, DOI:10.2174/156802612800672844.
- M. A. Ali, H. Sarker, T. Khan, H. Sheikh, A. Saif, F. B. Farid, S. Afrin, M. A. Khatun and N. Kumar, Multi-Omics Pan-Cancer Profiling of CDK2 and in Silico Identification of Plant-Derived Inhibitors Using Machine Learning Approaches, RSC Adv., 2025, 15, 36938–36968, 10.1039/D5RA05535K.
- A. Saif, M. T. Islam, M. O. Raihan, N. Yousefi, M. A. Rahman, H. Faridi, A. R. Hasan, M. M. Hossain, R. M. Saleem and G. M. Albadrani, et al., Pan-Cancer Analysis of CDC7 in Human Tumors: Integrative Multi-Omics Insights and Discovery of Novel Marine-Based Inhibitors through Machine Learning and Computational Approaches, Comput. Biol. Med., 2025, 190, 110044, DOI:10.1016/j.compbiomed.2025.110044.
- H. Gubler, U. Schopfer and E. Jacoby, Theoretical and Experimental Relationships between Percent Inhibition and IC50 Data Observed in High-Throughput Screening, J. Biomol. Screen, 2013, 18, 1–13, DOI:10.1177/1087057112455219.
- K. Danishuddin and A. U. Descriptors, Their Selection Methods in QSAR Analysis: Paradigm for Drug Design, Drug Discov. Today, 2016, 21, 1291–1302, DOI:10.1016/j.drudis.2016.06.013.
- Q. Guo, S. Hernandez-Hernandez and P. J. Ballester, UMAP-Based Clustering Split for Rigorous Evaluation of AI Models for Virtual Screening on Cancer Cell Lines, J. Cheminform., 2025, 17, 94, DOI:10.1186/s13321-025-01039-8.
- S. Majumdar and S. C. Basak, Beware of External Validation! - A Comparative Study of Several Validation Techniques Used in QSAR Modelling, Curr. Comput. Aided Drug Des., 2018, 14, 284–291, DOI:10.2174/1573409914666180426144304.
- X.-Y. Meng, H.-X. Zhang, M. Mezei and M. Cui, Molecular Docking: A Powerful Approach for Structure-Based Drug Discovery, Curr. Comput. Aided Drug Des., 2012, 7, 146–157, DOI:10.2174/157340911795677602.
- S. K. Kondapuram, S. Sarvagalla and M. S. Coumar, Docking-Based Virtual Screening Using PyRx Tool: Autophagy Target Vps34 as a Case Study, In Molecular Docking for Computer-Aided Drug Design, Elsevier, 2021, pp. 463–477 Search PubMed.
- A. V. Danish Ahmad, S. W. Khan, Q. Yasar, M. S. Shaikh and M. M. Khan, Computational Biology Approach to Predict Molecular Mechanism in Cancer, Oral Oncol Rep., 2024, 12, 100651, DOI:10.1016/j.oor.2024.100651.
- A. J. Owoloye, F. C. Ligali, O. A. Enejoh, A. Z. Musa, O. Aina, E. T. Idowu and K. M. Oyebola, Molecular Docking, Simulation and Binding Free Energy Analysis of Small Molecules as PfHT1 Inhibitors, PLoS One, 2022, 17, e0268269, DOI:10.1371/journal.pone.0268269.
- U. C. Ogbodo, O. A. Enejoh, C. H. Okonkwo, P. Gnanasekar, P. W. Gachanja, S. Osata, H. C. Atanda, E. A. Iwuchukwu, I. Achilonu and O. I. Awe, Computational Identification of Potential Inhibitors Targeting Cdk1 in Colorectal Cancer, Front. Chem., 2023, 11, 1264808, DOI:10.3389/fchem.2023.1264808.
- P. A. Kollman, I. Massova, C. Reyes, B. Kuhn, S. Huo, L. Chong, M. Lee, T. Lee, Y. Duan and W. Wang, et al., Calculating Structures and Free Energies of Complex Molecules: Combining Molecular Mechanics and Continuum Models, Acc. Chem. Res., 2000, 33, 889–897, DOI:10.1021/ar000033j.
- R. A. Friesner, R. B. Murphy, M. P. Repasky, L. L. Frye, J. R. Greenwood, T. A. Halgren, P. C. Sanschagrin and D. T. Mainz, Extra Precision Glide: Docking and Scoring Incorporating a Model of Hydrophobic Enclosure for Protein−Ligand Complexes, J. Med. Chem., 2006, 49, 6177–6196, DOI:10.1021/jm051256o.
- M. K. Gilson, J. A. Given, B. L. Bush and J. A. McCammon, The Statistical-Thermodynamic Basis for Computation of Binding Affinities: A Critical Review, Biophys. J., 1997, 72, 1047–1069, DOI:10.1016/S0006-3495(97)78756-3.
- H. Arya and T. K. Bhatt, Molecular Dynamics Simulations. In The Design & Development of Novel Drugs and Vaccines, Elsevier, 2021, pp. 65–81 Search PubMed.
- K. Roos, C. Wu, W. Damm, M. Reboul, J. M. Stevenson, C. Lu, M. K. Dahlgren, S. Mondal, W. Chen and L. Wang, et al., OPLS3e: Extending Force Field Coverage for Drug-Like Small Molecules, J. Chem. Theory Comput., 2019, 15, 1863–1874, DOI:10.1021/acs.jctc.8b01026.
- M. S. Valdés-Tresanco, M. E. Valdés-Tresanco, P. A. Valiente and E. Moreno, Gmx_MMPBSA: A New Tool to Perform End-State Free Energy Calculations with GROMACS, J. Chem. Theory Comput., 2021, 17, 6281–6291, DOI:10.1021/acs.jctc.1c00645.
- B. R. Miller, T. D. McGee, J. M. Swails, N. Homeyer, H. Gohlke and A. E. Roitberg, MMPBSA.Py: An Efficient Program for End-State Free Energy Calculations, J. Chem. Theory Comput., 2012, 8, 3314–3321, DOI:10.1021/ct300418h.
- M. R. Shirts, C. Klein, J. M. Swails, J. Yin, M. K. Gilson, D. L. Mobley, D. A. Case and E. D. Zhong, Lessons Learned from Comparing Molecular Dynamics Engines on the SAMPL5 Dataset, J. Comput. Aided Mol. Des., 2017, 31, 147–161, DOI:10.1007/s10822-016-9977-1.
- A. Daina, O. Michielin and V. Zoete, SwissADME: A Free Web Tool to Evaluate Pharmacokinetics, Drug-Likeness and Medicinal Chemistry Friendliness of Small Molecules, Sci. Rep., 2017, 7, 42717, DOI:10.1038/srep42717.
- P. Banerjee, E. Kemmler, M. Dunkel and R. Preissner, ProTox 3.0: A Webserver for the Prediction of Toxicity of Chemicals, Nucleic Acids Res., 2024, gkae303, DOI:10.1093/nar/gkae303.
- M. Ashtiani, A. Salehzadeh-Yazdi, Z. Razaghi-Moghadam, H. Hennig, O. Wolkenhauer, M. Mirzaie and M. Jafari, A Systematic Survey of Centrality Measures for Protein-Protein Interaction Networks, BMC Syst. Biol., 2018, 12, 80, DOI:10.1186/s12918-018-0598-2.
- M. Wang, H. Wang and H. Zheng, A Mini Review of Node Centrality Metrics in Biological Networks, Int. J. Intell. Netw., 2022, 99–110, DOI:10.53941/ijndi0101009.
- D. J. Wood, S. Korolchuk, N. J. Tatum, L.-Z. Wang, J. A. Endicott, M. E. M. Noble and M. P. Martin, Differences in the Conformational Energy Landscape of CDK1 and CDK2 Suggest a Mechanism for Achieving Selective CDK Inhibition, Cell Chem. Biol., 2019, 26, 121–130.e5, DOI:10.1016/j.chembiol.2018.10.015.
- G. Ke, Q. Meng, T. Finley, T. Wang, W. Chen, W. Ma, Q. Ye and T.-Y. Liu, LightGBM: A Highly Efficient Gradient Boosting Decision Tree, In Proceedings of the Advances in Neural Information Processing Systems, ed. Guyon, I., Luxburg, U. Von, Bengio, S., Wallach, H., Fergus, R., Vishwanathan, S. and Garnett, R., Curran Associates, Inc., 2017, vol. 30 Search PubMed.
- X. Zhao, Y. Liu and Q. Zhao, Improved LightGBM for Extremely Imbalanced Data and Application to Credit Card Fraud Detection, IEEE Access, 2024, 12, 159316–159335, DOI:10.1109/ACCESS.2024.3487212.
- A. P. Bento, A. Gaulton, A. Hersey, L. J. Bellis, J. Chambers, M. Davies, F. A. Krüger, Y. Light, L. Mak and S. McGlinchey, et al., The ChEMBL Bioactivity Database: An Update, Nucleic Acids Res., 2014, 42, D1083–D1090, DOI:10.1093/nar/gkt1031.
- L. Zhao, W. Jiang, Z. Zhu, F. Pan, X. Xing, F. Zhou and L. Zhao, Rosemarinic Acid-Induced Destabilization of Aβ Peptides: Insights from Molecular Dynamics Simulations, Foods, 2024, 13(24), 4170, DOI:10.3390/foods13244170.
- A. S. Abouzied, S. Alqarni, K. M. Younes, S. M. Alanazi, D. M. Alrsheed, R. K. Alhathal, B. Huwaimel and A. M. Elkashlan, Structural and Free Energy Landscape Analysis for the Discovery of Antiviral Compounds Targeting the Cap-Binding Domain of Influenza Polymerase PB2, Sci. Rep., 2024, 14, 25441, DOI:10.1038/s41598-024-69816-3.
- D. F. Veber, S. R. Johnson, H. Y. Cheng, B. R. Smith, K. W. Ward and K. D. Kopple, Molecular Properties That Influence the Oral Bioavailability of Drug Candidates, J. Med. Chem., 2002, 45, 2615–2623, DOI:10.1021/jm020017n.
- F. T. Ndombera, Revisiting Cheminformatics and Mechanisms of Action of Chloroquine and Hy- Droxychloroquine in Targeting Covid-19, J. Biol. Chem., 2020, 3, 1–11, DOI:10.17303/jbcg.2020.3.101.
- M. Rashid, O. Afzal and A. S. A. Altamimi, Benzimidazole Molecule Hybrid With Oxadiazole Ring As Antiproliferative Agents: In-Silico Analysis, Synthesis And Biological Evaluation, J. Chil. Chem. Soc., 2021, 66, 5164–5182, DOI:10.4067/S0717-97072021000205164.
- L. Z. Benet, C. M. Hosey, O. Ursu and T. I. Oprea, BDDCS, the Rule of 5 and Drugability, Adv. Drug Deliv. Rev., 2016, 101, 89–98, DOI:10.1016/j.addr.2016.05.007.
- Y. Sun, M. Zabihi, Q. Li, X. Li, B. J. Kim, E. E. Ubogu, S. N. Raja, U. Wesselmann and C. Zhao, Drug Permeability: From the Blood–Brain Barrier to the Peripheral Nerve Barriers, Adv. Ther., 2023, 6, 2200150, DOI:10.1002/adtp.202200150.
- M. K. Heavey, D. Durmusoglu, N. Crook and A. C. Anselmo, Discovery and Delivery Strategies for Engineered Live Biotherapeutic Products, Trends Biotechnol., 2022, 40, 354–369, DOI:10.1016/j.tibtech.2021.08.002.
- M. Sibai, S. Cervilla, D. Grases, E. Musulen, R. Lazcano, C.-K. Mo, V. Davalos, A. Fortian, A. Bernat and M. Romeo, et al., The Spatial Landscape of Cancer Hallmarks Reveals Patterns of Tumor Ecological Dynamics and Drug Sensitivity, Cell Rep., 2025, 44, 115229, DOI:10.1016/j.celrep.2024.115229.
- A. Boija, I. A. Klein, B. R. Sabari, A. Dall'Agnese, E. L. Coffey, A. V. Zamudio, C. H. Li, K. Shrinivas, J. C. Manteiga and N. M. Hannett, et al., Transcription Factors Activate Genes through the Phase-Separation Capacity of Their Activation Domains, Cell, 2018, 175, 1842–1855.e16, DOI:10.1016/j.cell.2018.10.042.
- C. Catalanotto, C. Cogoni and G. Zardo, MicroRNA in Control of Gene Expression: An Overview of Nuclear Functions, Int. J. Mol. Sci., 2016, 17(10), 1712, DOI:10.3390/ijms17101712.
- Y. Chen, L. Jin, Z. Jiang, S. Liu and W. Feng, Identifying and Validating Potential Biomarkers of Early Stage Lung Adenocarcinoma Diagnosis and Prognosis, Front. Oncol., 2021, 11, 644426, DOI:10.3389/fonc.2021.644426.
- Q. Du, W. Liu, T. Mei, J. Wang, T. Qin and D. Huang, Prognostic and Immunological Characteristics of CDK1 in Lung Adenocarcinoma: A Systematic Analysis, Front. Oncol., 2023, 13, 1128443, DOI:10.3389/fonc.2023.1128443.
- S. Li, H. Li, Y. Cao, H. Geng, F. Ren, K. Li, C. Dai and N. Li, Integrated Bioinformatics Analysis Reveals CDK1 and PLK1 as Potential Therapeutic Targets of Lung Adenocarcinoma, Medicine, 2021, 100, e26474, DOI:10.1097/MD.0000000000026474.
- M. Batool, B. Ahmad and S. Choi, A Structure-Based Drug Discovery Paradigm, Int. J. Mol. Sci., 2019, 20, 2783, DOI:10.3390/ijms20112783.
- O. A. Enejoh, C. H. Okonkwo, H. Nortey, O. A. Kemiki, A. Moses, F. N. Mbaoji, A. S. Yusuf and O. I. Awe, Machine Learning and Molecular Dynamics Simulations Predict Potential TGR5 Agonists for Type 2 Diabetes Treatment, Front. Chem., 2024, 12, 1503593, DOI:10.3389/fchem.2024.1503593.
- M. Di Stefano, S. Galati, G. Ortore, I. Caligiuri, F. Rizzolio, C. Ceni, S. Bertini, G. Bononi, C. Granchi and M. Macchia, et al., Machine Learning-Based Virtual Screening for the Identification of Cdk5 Inhibitors, Int. J. Mol. Sci., 2022, 23(18), 10653, DOI:10.3390/ijms231810653.
- R. S. Bhimanwar, K. B. Lokhande, A. Shrivastava, A. Singh, S. S. Chitlange and A. Mittal, Identification of Potential Drug Candidates as TGR5 Agonist to Combat Type II Diabetes Using in Silico Docking and Molecular Dynamics Simulation Studies, J. Biomol. Struct. Dyn., 2023, 41, 13314–13331, DOI:10.1080/07391102.2023.2173654.
- V. Hurmach, V. Karaushu, S. Prylutska, Z. Klestova, S. Vyzhva, Y. Prylutskyy, U. Ritter and V. Garamus, In Silico Analysis of C60 Fullerene Interaction with TMPRSS2: Toward Novel COVID-19 Prevention Approaches, Molecules, 2025, 30(23), 4586, DOI:10.3390/molecules30234586.
- C. C. David and D. J. Jacobs, In, Principal Component Analysis: A Method for Determining the Essential Dynamics of Proteins. 2014. pp. 193–226 Search PubMed.
Footnote |
| † These authors contributed equally as first author to this work. |
| This journal is © The Royal Society of Chemistry 2026 |