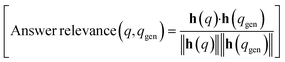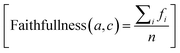Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Structured domain knowledge enables trustworthy materials science question-answering with large language models
Daegun
Lee
 ab,
Jiwoo
Choi
a,
Gyeong Hoon
Yi
d,
Seok Su
Sohn
b,
Byungju
Lee
*ac and
Donghun
Kim
ab,
Jiwoo
Choi
a,
Gyeong Hoon
Yi
d,
Seok Su
Sohn
b,
Byungju
Lee
*ac and
Donghun
Kim
 *d
*d
aComputational Science Research Center, Korea Institute of Science and Technology (KIST), Seoul 02792, Republic of Korea. E-mail: blee89@kist.re.kr
bDepartment of Materials Science and Engineering, Korea University, Seoul 02841, Republic of Korea
cNanoscience and Technology, KIST School, University of Science and Technology, Seoul, Republic of Korea
dDepartment of Materials Science and Engineering, Korea Advanced Institute of Science and Technology (KAIST), Daejeon 34141, Republic of Korea. E-mail: donghun.kim@kaist.ac.kr
First published on 29th April 2026
Abstract
Large language models (LLMs) remain unreliable for materials science question answering because correct conclusions depend on detailed experimental conditions. Here, we show that a structured, domain-specific knowledge dataset is a critical prerequisite for trustworthy LLM-assisted question answering in materials science. Using water-splitting catalysis as a proof of concept, we curate the literature into a hierarchical, machine-queryable knowledge base encoding material synthesis, composition, and performance. This structured representation improves condition-aware retrieval and reduces context mismatches that commonly arise from superficial semantic similarity. Combined with query reformulation, it achieves 85.6% accuracy on 202 DOI-identification questions versus 21.3% for an unstructured baseline, while reducing operating cost by 39%. To assess broader free-form scientific question answering beyond exact-match retrieval, we further evaluate 202 descriptive questions using the RAGAS framework, which indicates more faithful, evidence-grounded answers. Together, these results show that structured domain knowledge can substantially improve the reliability of LLM-based materials science question answering.
Introduction
Large language models (LLMs) have shown remarkable capabilities across a wide range of tasks, including summarization, question answering (Q/A), and information extraction.1–7 However, their performance often degrades in domain-specific scientific problems, where precise terminology, experimental conditions, and data structures are crucial. Two persistent issues limit their reliability: the lack of domain-specific knowledge,8 and the generation of hallucinations, i.e., plausible but incorrect statements (Fig. 1).9–11Fig. 1 illustrates these challenges. For general questions, an LLM combined with an external database (a standard retrieval-augmented generation, or RAG, approach) can well produce accurate and helpful responses (Fig. 1a).12,13 Yet for domain-specific queries, such as those in water-splitting catalysis, the same framework frequently fails because it cannot correctly interpret technical terms or experimental contexts (Fig. 1b).14–16 This highlights that while RAG enhances factual grounding in general settings, it remains insufficient for specialized scientific domains requiring fine-grained contextual understanding.14
Beyond RAG, other strategies have been proposed to adapt LLMs to specialized domains, including continued pre-training (CPT) and supervised fine-tuning (SFT).17,18 While CPT and SFT can inject domain expertise, they often suffer from catastrophic forgetting or high data labeling costs.19 In contrast, RAG augments generation with external knowledge without modifying model weights, and recent studies suggest that RAG consistently outperforms unsupervised fine-tuning for factual knowledge acquisition.20 Significant progress has been made in general domains, such as BloombergGPT in finance21 and BioGPT in biomedicine,22 demonstrating that mixing domain-specific and general corpora can preserve general capabilities while enhancing expertise.
For materials science specifically, models like MatBERT and MatSciBERT have improved performance on named entity recognition (NER) tasks.23,24 More recently, billion-parameter scale LLMs like HoneyBee and LLaMAt have been developed through instruction fine-tuning and large-scale continued pre-training, respectively, often outperforming general models like GPT-4 on materials science benchmarks.25,26 In parallel, open-source LLMs have rapidly advanced, with Meta's LLaMA 3 demonstrating performance comparable to proprietary models like GPT-4 across diverse benchmarks while enabling local deployment without API costs and full reproducibility of results.27 Beyond base models, agent systems such as ChemCrow and HoneyComb have emerged, integrating expert-designed tools and curated knowledge bases to autonomously execute complex tasks.28,29 Despite these advances, benchmarks like MaScQA reveal that even state-of-the-art models with chain-of-thought prompting still struggle with conceptual errors in specialized fields.30
The original RAG framework, or conventional RAG (C-RAG), consists of four stages: indexing, retrieval, augmentation, and generation.12 However, C-RAG often fails with complex scientific queries where semantic similarity alone is insufficient.14 To address this, query reformulation techniques have evolved. Query expansion methods such as HyDE and Query2Doc generate hypothetical or pseudo-documents to improve retrieval.31,32 Query rewriting approaches like Rewrite-Retrieve-Read and RaFe train rewriter models using reinforcement learning or ranking feedback.33,34 Query decomposition methods, such as GenDec and RQ-RAG, further break down multi-hop questions into simpler sub-queries to improve reasoning transparency.35,36 Yet, a critical gap remains: the impact of database structuring on RAG performance has not been systematically evaluated in scientific domains, and existing reformulation methods often lack the domain-specific precision required for materials science.
To address this limitation, we designed a domain-aligned RAG framework that integrates structured knowledge and query reformulation for more precise retrieval and reasoning. We selected water-splitting catalysis as a representative proof-of-concept domain due to its rich but heterogeneous literature and well-defined quantitative benchmarks. Performance in this field depends on multiple interacting variables, such as catalyst composition, synthesis route, electrolyte and its pH condition, and testing protocols,37–40 making it a stringent test for retrieval and reasoning quality. In this work, we construct a structured database of water-splitting catalysts from a large-scale scientific literature, through paragraph classification, synthesis-method classification, and named entity recognition (NER). And we develop a query reformulation RAG (QR-RAG) pipeline that couples sparse lexical retrieval with dense vector retrieval for hybrid search.
Our QR-RAG approach differs from existing methods in three key aspects: (1) it combines decomposition with query optimization to preserve critical domain-specific terms; (2) it employs an adaptive two-step process that invokes decomposition only when initial retrieval fails, reducing overhead; and (3) it integrates hybrid retrieval for exact terminology matching alongside semantic similarity. We evaluate this framework with complementary benchmarks covering distinct aspects of scientific question answering. The combination of structured domain knowledge and QR-RAG improves accuracy on 202 DOI-identification questions from 21.3% for conventional RAG on raw literature to 85.6%, while reducing operating cost by 39%. We also assess 202 descriptive questions using the RAGAS framework, showing more faithful and evidence-grounded responses beyond exact-match retrieval. Together, these results highlight the value of structured domain knowledge for reliable domain-specific scientific Q/A.
Results
System overview
From literature to Q/A We develop a comprehensive system that transforms unstructured water-splitting catalysis literature into a structured database and enables intelligent question answering (Q/A) through retrieval-augmented generation (Fig. 2). The system consists of two major components: structured database construction through a three-stage natural language processing pipeline, and a Q/A system that leverages both the structured data and original text.For structured database construction, we employ a three-stage pipeline to systematically extract and organize synthesis information from scientific literature. In stage 1, paragraph classification assigns each paragraph to one of four classes: system, performance, synthesis, or others, enabling focused extraction from relevant content. In stage 2, synthesis method classification assigns synthesis paragraphs to one of seven classes: vapor phase, solid phase, electrodeposition, hydro/solvothermal, precipitation, sol–gel, or others. In stage 3, we apply relational named entity recognition (RE-NER) to extract five critical entities: target, precursor, solvent, additive, and substrate. RE-NER selectively extracts only those entities that are relationally connected within the paragraph. The description of each class and entity was provided in advance (Fig. S1) before proceeding with the database construction process.
The Q/A system processes user queries through a multi-step pipeline leveraging the structured database constructed above. Upon receiving a user query, the system performs query reformulation that simplifies and optimizes complex questions for improved retrieval accuracy. The reformulated queries are then used to retrieve relevant source documents from the structured database through combined dense vector retrieval and sparse lexical retrieval. Finally, the retrieved documents serve as context for the language model to generate comprehensive answers to the user's questions, even when they are complex, based on the most relevant supporting documents.
Structured database construction from literature
The label definitions for all three stages (paragraph classification, synthesis method classification, and RE-NER) are provided in Fig. S1. Based on these definitions, all paragraph labeling and entity tagging were manually performed by a materials science graduate researcher. To ensure annotation consistency, paragraphs with ambiguous classifications were excluded from the dataset. In our pipeline, the models were chosen based on their proven performance in materials science under different learning paradigms, namely few-shot learning and fine-tuning. MatBERT is a domain-specific model that shows good performance when fine-tuned with hundreds of labeled examples,41,42 while large language models (LLMs) such as GPT-4, LLaMA 3.3, and HoneyBee achieve good performance with few-shot learning using only a few dozen examples per class.43,44 Although LLMs are also known to perform well when fine-tuned on large datasets, the high cost of such training makes them less practical. Therefore, we selected MatBERT to represent the fine-tuning approach and GPT-4 Turbo, LLaMA 3.3-70B, and HoneyBee-7B to represent the few-shot learning approach, and compared their performance under these distinct strategies. Since the two approaches rely on different training methods, we also set the number of training examples differently. Specifically, we used about 20–30 times more data for MatBERT fine-tuning compared with LLM few-shot learning. This ensured that the comparison considered not only performance but also cost within a reasonable range. Since annotating thousands of examples at each stage is prohibitively time-consuming, we determined the fine-tuning dataset size based on the point at which MatBERT achieved sufficient classification performance, which corresponded to roughly 20–30 times the data used for LLM few-shot learning.44In stage 1 and stage 2 classification tasks, MatBERT was fine-tuned with 1260 and 720 training examples, while LLMs were applied with few-shot examples (Fig. 3a) using general instruction prompts with Chain-of-Thought reasoning (Fig. S2).45 GPT-4 Turbo46 and LLaMA 3.3-70B were applied with 40 and 35 few-shot examples for paragraph and synthesis classification, respectively. For HoneyBee-7B, due to the increased context length from Chain-of-Thought prompting and its limited context window, only one example per category was used in the few-shot setting. On a test set of 240 paragraphs in stage 1, MatBERT achieved an F1-score of 0.960, GPT-4 Turbo achieved 0.950, LLaMA 3.3-70B achieved 0.932, and HoneyBee achieved 0.485 (Fig. 3b). In stage 2, MatBERT and GPT-4 Turbo both achieved the highest F1-score of 0.964 on a test set of 280 paragraphs, while LLaMA 3.3-70B achieved 0.834 and HoneyBee achieved 0.480 (Fig. 3c). These results indicate that GPT-4 Turbo performs comparably to fine-tuned MatBERT in classification tasks, while LLaMA 3.3-70B shows competitive but slightly lower performance. HoneyBee, despite being a materials science domain-specific model, showed limited performance in the few-shot setting, likely due to the restricted number of examples imposed by its context window constraints. In the RE-NER task, MatBERT was fine-tuned with 300 training examples, while LLMs were applied with few-shot examples and an additional filtering step to refine the results. To address material naming ambiguity in scientific literature, we also implemented a normalization step that converts material names to their molecular formula representations (Fig. S3). This ensures consistency across different naming conventions such as “NiFe LDH” vs. “NiFeOOH” or other abbreviations commonly used in the literature. GPT-4 and LLaMA 3.3-70B were applied with 10 few-shot examples. HoneyBee was applied with 5 few-shot examples due to its limited context window. On a test set of 390 paragraphs, MatBERT achieved an average F1-score of 0.805, while GPT-4 achieved 0.947, LLaMA 3.3-70B achieved 0.755, and HoneyBee achieved 0.452 (Fig. 3d). RE-NER is a particularly challenging task because entities must be bound to their correct targets while preserving relational coherence (Fig. S4). To address this difficulty, we applied LLMs with few-shot learning using detailed NER protocol instructions (Fig. S2). The performance gap across models can be attributed to differences in parameter capacity and instruction-following capabilities. GPT-4, with its largest parameter scale and advanced instruction-following ability, significantly outperformed all other models. LLaMA 3.3-70B showed moderate performance, outperforming MatBERT in target extraction but showing lower performance in other entity types. HoneyBee, constrained by its limited context window and fewer few-shot examples, showed the lowest performance across all entity types.
The cost analysis further highlights the trade-offs. Since MatBERT and GPT-4 Turbo achieved comparable F1-scores in the classification tasks, we focused our cost comparison on these two models. MatBERT requires a large amount of training data at the beginning, resulting in significant human labeling cost, but once trained, it incurs almost no additional cost. GPT-4, in contrast, requires only a small number of examples for setup, so the initial cost is low, but each classification generates API usage fees, causing the cost to increase progressively with more data. Given their similar performance, the choice between these two models can be determined by cost efficiency depending on dataset size. We found that GPT-4 is more cost-effective for datasets below 2500 paragraphs, with MatBERT becoming the better choice for larger volumes (Fig. S5). The resulting structured database comprises 2343 papers collected from major publishers including Elsevier, ACS, RSC, and Wiley, spanning research from 2003 to 2023 across a wide range of target metal elements (Fig. S6).
Comparison of retrieval-augmented generation methods
As discussed in the introduction, C-RAG is not well-suited for complex scientific domains. In this study, to enable rigorous quantitative evaluation, all questions were formulated to require the model to output the correct DOI. Under this DOI-based evaluation setting, C-RAG demonstrated low performance when applied to the raw database, achieving only 21.3% accuracy on water-splitting catalyst Q/A tasks (Table 1). To overcome these limitations, it was necessary to design a RAG system applicable to a specialized domains.47 To this end, we first indexed both the raw database and the structured database into a paragraph-level vector store using a GPT embedding model.48 Based on this shared database, which served as an invariant experimental variable, we compared two RAG approaches. The C-RAG directly uses the user's question and conducts document retrieval via dense retrieval (Fig. 4a). In contrast, the query reformulation RAG (QR-RAG) designed in this study applies reformulation prior to retrieval, decomposing and optimizing the user's question (Fig. 4b). Query optimization is first applied to refine the user's question by removing redundant expressions and retaining only essential condition terms. If this single-step process fails to retrieve relevant passages, a second step is applied in which the original question is decomposed into multiple sub-queries and each sub-query is then optimized (Fig. 4c). In addition, QR-RAG employs hybrid retrieval that combines dense vector and sparse lexical retrieval methods. Through this combination of query reformulation and hybrid retrieval, the system shifts away from semantically similar but contextually irrelevant passages and instead accesses passages that can directly provide answers to the user's questions.| Model | DB | Method | Single | Multiple | Avg | Cost |
|---|---|---|---|---|---|---|
| GPT-4o | HTML | C-RAG | 22.2% (24/108) | 20.2% (19/94) | 21.3% (43/202) | $29.2 |
| QR-RAG | 49.1% (53/108) | 44.7% (42/94) | 47.0% (95/202) | $56.9 | ||
| JSON | C-RAG | 81.5% (88/108) | 59.6% (56/94) | 71.3% (144/202) | $14.1 | |
| QR-RAG | 90.7% (98/108) | 79.8% (75/94) | 85.6% (173/202) | $17.8 | ||
| LLaMA 3.3 70B | HTML | C-RAG | 14.8% (16/108) | 10.6% (10/94) | 12.9% (26/202) | — |
| QR-RAG | 40.7% (44/108) | 35.1% (33/94) | 38.1% (77/202) | — | ||
| JSON | C-RAG | 70.4% (76/108) | 58.5% (55/94) | 64.9% (131/202) | — | |
| QR-RAG | 80.6% (87/108) | 70.2% (66/94) | 75.7% (153/202) | — |
Quantitative accuracy and cost of Q/A systems
The DOI-based evaluation was designed to assess the system's ability to retrieve the correct source document for a given user query. Since DOI provides a unique identifier with a single correct answer, it enables rigorous quantitative evaluation of retrieval accuracy. We measure Q/A accuracy across four settings that combine two databases and two pipelines: HTML (raw) with C-RAG, HTML with QR-RAG, JSON (structured) with C-RAG, and JSON with QR-RAG. For the Q/A evaluation, we compared GPT-4o and LLaMA 3.3-70B. MatBERT was excluded as it is not an LLM suitable for RAG-based generation, and HoneyBee was excluded due to its context window limitations that prevent RAG implementation. We conducted experiments on a total of 202 questions (Table S1). Each question was constructed by identifying paragraphs containing distinctive scientific features that uniquely characterize a specific paper. During preliminary experiments, questions for which multiple papers satisfied the given conditions were iteratively refined to ensure that each question corresponds to exactly one correct answer. These 202 questions were designed to ask for the DOI. Since general descriptive questions can have multiple valid answer formats and do not converge to a single short answer, making quantitative evaluation nearly impossible, we restricted our questions to those requesting the DOI. For GPT-4o, the average accuracy across HTML/C-RAG, HTML/QR-RAG, JSON/C-RAG, and JSON/QR-RAG is 21.3%, 47.0%, 71.3%, and 85.6%, respectively, while LLaMA 3.3-70B achieved 12.9%, 38.1%, 64.9%, and 75.7% under the same settings (Table 1). Although GPT-4o outperformed LLaMA 3.3-70B across all configurations, both models exhibited similar performance trends: structuring the database and applying query reformulation each independently improved accuracy. The pattern reveals two independent gains. First, structuring the database (JSON) improves retrieval even when the pipeline remains unchanged. Second, enhancing the pipeline with query reformulation and hybrid retrieval improves accuracy even on the HTML database. The best result comes from combining the structured database with the QR-RAG. We also compare single-condition queries that can be answered from one paragraph and multiple-condition queries that require combining results across paragraphs (Fig. S7). The structured database reduces condition mismatches by aligning retrieved passages more closely with the intended synthesis or testing context. Query reformulation reduces failures from underspecified questions by removing unnecessary words and focusing on the condition terms already present in the question (Fig. S8). In all settings, answers are grounded in retrieved passages and verified against them to confirm support.A cost analysis further reveals significant efficiency gains. The JSON database shortens average context length and increases the proportion of relevant passages, which reduces tokens and retries. Query reformulation incurs additional cost on the HTML database because the reformulation step requires extra LLM calls before retrieval. However, this cost is offset on the JSON database since fewer and better-matched passages are supplied to the generator, thereby reducing overall token usage and retries. Operating cost decreases from $29.2 for HTML with C-RAG to $17.8 for JSON with QR-RAG, representing a 39% reduction (Table 1). Cost analysis was performed only for GPT-4o, as LLaMA 3.3-70B is an open-source model that does not incur API usage fees.
To verify that the framework extends beyond DOI identification, we additionally evaluated 50 questions targeting numerical property extraction, including synthesis temperature, overpotential, and Tafel slope (Table S2). Using LLaMA 3.3-70B with the structured JSON database and QR-RAG, the system achieved 84.0% accuracy, which is higher than the DOI identification accuracy (75.7%) under the same configuration (Fig. S9). This result confirms that our approach effectively extracts specific scientific metrics from the literature.
Qualitative assessment with RAGAS
While the DOI-based evaluation focuses on retrieval accuracy, it does not fully capture the system's ability to generate comprehensive and contextually appropriate answers. To address this limitation and evaluate performance on descriptive questions commonly encountered in practical Q/A scenarios, we conducted a qualitative assessment using the RAGAS framework. Most questions that users ask in practice are free-form and descriptive. A simple accuracy score on short facts cannot fully test whether an answer is relevant,49 well-supported by evidence, and complete. Manual review would be ideal but is time-consuming and not scalable. We therefore use the RAGAS framework,50 which is widely adopted for LLM-based qualitative evaluation, to approximate human judgments at scale. RAGAS reports three scores that match our requirements: context relevance, faithfulness, and answer relevance (Fig. 5a). Context relevance measures whether the retrieved passages are relevant to the user's question by evaluating how many sentences in the context contribute to answering the query. Faithfulness assesses whether the generated answer is grounded in the retrieved context by decomposing the answer into individual claims and verifying each claim against the context, thereby detecting potential hallucinations. Answer relevance evaluates whether the answer directly addresses the user's question by generating reverse questions from the answer and measuring their semantic similarity to the original question. The detailed formulas for each metric are provided in the methods section. Fig. 5b illustrates a representative example of the RAGAS evaluation process. In this case, the query asks about the relationship between core–shell heterostructure architecture and electrocatalytic behavior. The retrieved context contains relevant information including synthesis details and OER performance metrics (270 mV at 10 mA cm−2), resulting in a high context relevance score (1.00). The generated answer accurately extracts key concepts from the context, such as “encapsulated core–shell structure enables numerous accessible reactive sites” and “facilitates charge transfer,” which are directly supported by the retrieved passages. This grounding in the context yields a high answer relevance score (0.93). The faithfulness score (0.71) indicates that while most claims in the answer are verifiable from the context, some additional interpretations were generated by the model.Based on this evaluation framework, we systematically assessed the performance across different database and pipeline configurations. We conducted experiments on a total of 202 descriptive questions (Table S3). For GPT-4o, the JSON database with the QR-RAG achieves the highest scores across all three metrics (answer relevance: 0.717, context relevance: 0.643, faithfulness: 0.662), while LLaMA 3.3-70B showed similar trends with the highest scores in the same configuration (answer relevance: 0.769, context relevance: 0.561, faithfulness: 0.558) (Fig. 6). Although GPT-4o achieved higher context relevance and faithfulness scores overall, both models consistently demonstrated that the JSON database outperformed the HTML database, and QR-RAG outperformed C-RAG across all metrics. Such qualitative evaluation is based on LLMs, so the scores themselves do not have absolute meaning. However, they are valuable as relative indicators across different configurations. Moreover, the trends are consistent with the quantitative evaluation (Table 1), showing that the qualitative and quantitative assessments complement each other.
 | ||
| Fig. 6 Qualitative evaluation of water-splitting Q/A system comparing raw (HTML)/structured (JSON) databases and RAG methods using GPT-4o and LLaMA 3.3-70B. | ||
Discussion
This study develops a domain-specific Q/A framework for water-splitting catalysis by combining a structured database with a QR-RAG pipeline. Database structuring and query reformulation each improve performance independently and together produce the best accuracy and cost efficiency. On the same query sets and model backbone, the average accuracy increases from 21.3% with a raw database and a C-RAG to 85.6% with a structured database and a QR-RAG. The same trend holds for both single-condition and multiple-condition queries and is consistent with the qualitative assessment.The effect of structuring follows from how the literature is organized. The three-stage pipeline separates text into system, performance, synthesis, and others, assigns synthesis paragraphs to seven common methods, and applies RE-NER to extract target, precursor, solvent, additive, and substrate in context. Records are stored in a structured database as hierarchical JSON, which enables retrieval to filter by section and method before retrieval ranking and to provide the generator with shorter and more focused passages. In practice, this reduces irrelevant context and improves retrieval precision. As the quantitative evaluation was based on questions restricted to requesting DOIs, once retrieval succeeded, the task reduced to locating factual information, leaving limited opportunity for generation errors. Thus, the observed accuracy gains (21.3% → 71.3%, 47.0% → 85.6%) serve as direct evidence that the structured database enhances retrieval precision, which in turn lowers token usage and increases the likelihood51 that the first retrieved set already contains the evidence needed to answer the user's question.
The QR-RAG provides an independent improvement by pairing query reformulation with hybrid retrieval, which removes unnecessary words while retaining essential condition terms in the user's question. Because results in water-splitting catalysis depend strongly on operating conditions, questions that omit condition terms can surface paragraphs that read similarly but were measured under different settings. By focusing the query on the stated conditions, retrieval performance improves consistently on both the raw and the structured databases, showing that query reformulation enhances retrieval precision regardless of database type.52 Furthermore, even when retrieved paragraphs exhibit high semantic similarity a common challenge in specialized scientific domains the system demonstrates reliable answer identification by leveraging contextual interpretation of domain-specific constraints (Fig. S10). The comparative analysis of model performance underscores the critical role of model architecture, specifically the synergy between context window length and parameter capacity, in scientific Q/A tasks. While HoneyBee-7B is specifically fine-tuned for the materials science domain, its performance was significantly lower than that of general-purpose models like GPT-4o and LLaMA 3.3-70B. This gap is primarily attributed to HoneyBee-7B's limited context window, which restricted the number of few-shot examples and necessitated the exclusion of CoT reasoning in certain tasks, alongside its smaller parameter capacity which affected instruction-following capabilities. Our results suggest that for complex domain-specific RAG systems, the advantage of a longer context window, which enables richer contextual guidance and the processing of multiple retrieved passages, combined with sufficient parameter scale, can outweigh the benefits of domain-specific fine-tuning on a smaller scale. A larger context window and higher capacity allow the model to act as a more effective reasoner over external knowledge, mitigating the need for internalizing all domain expertise within the model weights. To evaluate the generalizability of our approach, we tested the QR-RAG framework on databases of varying sizes and domains using LLaMA 3.3-70B with the structured (JSON) database. The smallest subset contained only the 123 gold-standard papers, and non-answer papers were progressively added to simulate increasing retrieval difficulty. Accuracy decreased as database size increased, dropping from 92.1% with 123 papers to 75.7% with the full 2343 papers. To further test generalizability across different research fields, we added a battery materials database (5000 papers) to the full OER database,53 resulting in a combined database of 7343 papers. Despite the significant increase in database size and domain diversity, the system achieved 75.2% accuracy, demonstrating that the benefits of structured databases and query reformulation extend to other domains and database scales (Fig. S11).
To comprehensively assess system performance, we employed three complementary evaluation approaches. DOI identification measures retrieval accuracy by testing whether the system locates the correct source literature. RAGAS-based assessment evaluates the quality of generated answers through context relevance, faithfulness, and answer relevance metrics. Numerical property extraction verifies the ability to extract precise scientific values such as synthesis temperature, overpotential, and Tafel slope with appropriate units. Each approach has inherent limitations: DOI identification does not assess answer quality, RAGAS metrics cannot validate the scientific correctness of numerical values, and numerical property extraction covers a subset of experimental parameters. Together, however, these evaluations provide complementary evidence of retrieval accuracy, answer quality, and quantitative information extraction capabilities.
While effective, several limitations suggest directions for future work. The entity extraction stage, referred to as RE-NER in this study, is implemented through relation-aware prompting but does not include an explicit relation extraction module. Extending the schema to encode links among entities could further stabilize retrieval for multi-step synthesis procedures. Additionally, the current database focuses primarily on experimental synthesis information, and theoretical data such as DFT calculations or computational screening results are not included. Since the paper collection targeted experimental literature, the current framework may have limitations in answering questions related to theoretical details such as functional choices, U values, solvation models, and adsorption energies. Incorporating theoretical fields could enhance the system's ability to answer a broader range of scientific questions, although careful design would be required to manage potential ambiguity or noise arising from the integration of heterogeneous data types. We plan to address theoretical data integration in future work. Furthermore, unit variations in scientific literature (e.g., mA cm−2vs. A g−1, mV vs. V) are not standardized in the current pipeline. When users search for specific units, the system may not retrieve all relevant results due to these variations. Comprehensive unit normalization would require complex pre- and post-processing pipelines that account for various conventions and conversion factors, which we plan to address in future work. Drawing from these current limitations and our overall development experience, we emphasize that establishing standardized normalization procedures is necessary when extracting data from materials science literature. To guide future research and database construction, we propose three key best practices for domain-specific data normalization. First, material names should be converted into standardized molecular formulas to resolve ambiguities arising from diverse naming conventions and abbreviations (e.g., standardizing terms like “NiFe LDH” and “NiFeOOH”). Second, consistent unit normalization must be implemented for quantitative metrics, as variations in reporting units (e.g., mA cm−2vs. A g−1 for current density) can easily lead to context mismatches during the retrieval stage. Finally, maintaining hierarchical relationships between materials and their experimental conditions is critical to preserve the relational coherence of the extracted data. Implementing these practices can significantly reduce noise in structured databases and provide a more reliable foundation for LLM-based scientific reasoning. The structured database contains 2343 papers after the three-stage processing and filtering. This study focuses on water-splitting catalysis, but the same design can transfer to other materials domains by adapting the label space and entity types.41
Conclusions
This work presents a domain-aligned LLM framework that combines structured knowledge representation with QR-RAG to achieve reliable, evidence-grounded Q/A for experimental systems. Structuring domain literature into a hierarchical database and incorporating query reformulation each independently enhance Q/A performance improving average accuracy from 21.3% to 85.6% and reducing operating cost by 39%.Although water-splitting catalysis was used as a representative proof-of-concept domain, the approach is general and applicable to other experimental domains that demand contextual reasoning and integration of knowledge from multiple sources. This domain-aligned LLM paradigm provides a practical route toward trustworthy domain-specialized AI assistants that can support data-driven discovery, automated literature analysis, and hypothesis generation across diverse experimental materials research fields.
Methods
Literature collection and preparation
Research papers were collected through keyword-based search using the query “OER electrocatalyst water splitting” or “OER electrocatalyst water oxidation.” The collected papers were then refined through file size filtering, title analysis, and TF-IDF analysis to remove irrelevant or low-quality documents. The refined literatures are converted to plain text and split into paragraph units. When available, section labels and document identifiers are retained for reference during retrieval. Through the three-stage database construction process described in the Results, the structured database contains 2343 papers. Structured records are stored as hierarchical JSON, and the raw paragraphs are preserved in HTML to maintain the original text.Three-stage structured database construction
The three-stage construction consists of paragraph classification, synthesis method classification, and relational named entity recognition. Stage 1 and stage 2 use gpt-4-1106-preview with few-shot learning and Chain-of-Thought prompting46 for paragraph and synthesis method classification we also train and evaluate a fine-tuned MatBERT for comparison. For additional comparison, we evaluate LLaMA 3.3-70B via Vertex AI and HoneyBee-7B with 8 bit quantization for local deployment. Hyperparameters for LLaMA 3.3-70B were set identical to those used for GPT-4. Stage 3 uses gpt-4-0613 with few-shot learning for RE-NER to bind precursor, solvent, additive, and substrate to the local target within the same paragraph.Embedding and indexing
Embeddings are generated with text-embedding-3-large with 3072 dimensions. We store both the structured database and the raw database in ChromaDB v1.0.3 and index them at the paragraph level. We index at the paragraph level for similarity search.Water-splitting Q/A with query reformulation and hybrid retrieval
Two pipelines are evaluated. The C-RAG uses vector search based solely on cosine similarity and then generates an answer from the retrieved passages. The QR-RAG uses query reformulation, consisting of query decomposition and query optimization. In query decomposition, the model is instructed to break down a complex, multi-condition scientific question into up to three independent sub-queries, each expressed in a concrete and concise manner while retaining the essential terms and domain-specific concepts. In query optimization, the model removes unnecessary words and grammatical elements while preserving technical terms critical for retrieval. Both steps requested the output in a short JSON format, ensuring the sub-queries and optimized queries could be directly used in the paragraph-level retrieval process. After reformulation, the system combines sparse lexical retrieval with dense vector retrieval in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio before generating the final answer. For Q/A answer generation and QR-RAG operations, we use gpt-4o-2024-11-20. For automatic verification of incorrect or insufficient answers, we use gpt-3.5-turbo-0125. Retrieval uses top-k = 5.
1 ratio before generating the final answer. For Q/A answer generation and QR-RAG operations, we use gpt-4o-2024-11-20. For automatic verification of incorrect or insufficient answers, we use gpt-3.5-turbo-0125. Retrieval uses top-k = 5.
Evaluation metrics
We measure Q/A performance using three metrics from the RAGAS framework: answer relevance, context relevance, and faithfulness.where q is the original user query, qgen is the query generated by the model based on the answer, h(q) and h(qgen) denote their corresponding embedding representations.
| [Context relevance(q, c) = s] |
where a is the generated answer, ai is the i-th statement in the answer, fi equals 1 if the statement is directly supported by the retrieved context and 0 otherwise, and n is the total number of statements in a. All three RAGAS metrics were computed using gpt-4o-2024-11-20 as the evaluation LLM (temperature = 0) and text-embedding-ada-002 as the embedding model for answer relevance calculation.
Author contributions
D. L. designed this study, developed the experimental pipeline, performed programming and analysis, and drafted the manuscript. J. C. and G. H. Y. contributed to the extraction and curation of data from the catalysis literature. B. L., S. S. S. and D. K. supervised the project, reviewed the manuscript and guided the research. All authors discussed the results and approved the final manuscript.Conflicts of interest
The authors declare no competing interests.Data availability
The data and all codes for data curation, database construction, the query reformulation RAG framework, and evaluation used in this work are publicly available on Zenodo. The archived version corresponding to this manuscript is available at DOI: https://doi.org/10.5281/zenodo.19676935, and the most recent version can be accessed via the Concept DOI: https://doi.org/10.5281/zenodo.19676934.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d6dd00028b.
Acknowledgements
This work was supported by the National Research Foundation of Korea funded by the Ministry of Science and ICT (NRF-2021M3A7C2089739 and RS-2024-00450102), the InnoCORE program through the National Research Foundation of Korea funded by the Ministry of Science and ICT (1.260022.01) and Institutional Projects at the Korea Institute of Science and Technology (26E0223) and at the Korea Advanced Institute of Science and Technology (G04250010). We confirm that the contribution of RS-2024-00450102 to this work is 50%.References
- O. Kononova, et al. , iScience, 2021, 24, 102155 CrossRef PubMed.
- E. A. Olivetti, et al. , Appl. Phys. Rev., 2020, 7, 041317 CAS.
- A. Green, et al., Database, 2025, 2025, baaf003 Search PubMed.
- D. Scherbakov, et al. , Digit. Discov., 2025, 4, 112–125 Search PubMed.
- J. Lála, et al. , J. Chem. Inf. Model., 2024, 64, 1120–1135 Search PubMed.
- J. D'Souza, E. K. Sander and A. Aioanei, bioRxiv, 2025, preprint, bioRxiv:2025.07.14.664755, DOI:10.1101/2025.07.14.664755.
- A. Dunn, et al., arXiv, 2022, preprint arXiv:2212.05238, DOI:10.48550/arXiv.212.05238.
- G. Wang, et al. , Annu. Rev. Mater. Res., 2025, 1, 025001 Search PubMed.
- L. Huang, et al. , ACM Trans. Inf. Syst., 2025, 43, 1–55 Search PubMed.
- Z. Ji, et al. , ACM Comput. Surv., 2023, 55, 1–38 CrossRef.
- W. Zhang and J. Zhang, Mathematics, 2025, 13, 756 CrossRef.
- P. Lewis, et al. , Adv. Neural Inf. Process. Syst., 2020, 33, 9459–9474 Search PubMed.
- A. Brown, M. Roman and B. Devereux, arXiv, 2025, preprint arXiv:2508.06401, DOI:10.48550/arXiv.2508.06401.
- X. Zhong, et al., arXiv, 2025, preprint arXiv:2505.07671, DOI:10.48550/arXiv.2505.07671.
- C. Kiss, M. Nagy and P. Szilágyi, Comput. Sci. Discov., 2025, 28, 45 CrossRef.
- H. Liu, et al. , Scientometrics, 2025, 130, 1–30 Search PubMed.
- S. Wang, et al. , ACM Comput. Surv., 2025, 57, 1–47 Search PubMed.
- Y. Zhang, et al., arXiv, 2025, preprint arXiv:2508.19667, DOI:10.48550/arXiv.2508.19667.
- Y. Luo, et al., arXiv, 2023, preprint arXiv:2308.08747, DOI:10.48550/arXiv.2308.08747.
- O. Ovadia, M. Brief, M. Mishaeli and O. Elisha, Proc. 2024 Conf. Empirical Methods Nat. Lang. Process. (EMNLP), 2024, 233–249.
- S. Wu, et al., arXiv, 2023, preprint arXiv:2303.17564, DOI:10.48550/arXiv.2303.17564.
- R. Luo, et al. , Brief. Bioinform., 2022, 23, bbac409 CrossRef PubMed.
- A. Trewartha, et al. , Patterns, 2022, 3, 100488 CrossRef PubMed.
- T. Gupta, M. Zaki and N. M. A. Krishnan and Mausam, npj Comput. Mater., 2022, 8, 102 CrossRef.
- Y. Song, S. Miret, H. Zhang and B. Liu, Findings Assoc. Comput. Linguist.: EMNLP 2023, 2023, 5724–5739.
- V. Mishra, et al. , Nat. Mach. Intell., 2026, 8, 435–448 CrossRef.
- D. Dubey, et al., arXiv, 2024, preprint arXiv:2407.21783, DOI:10.48550/arXiv.2407.21783.
- A. M. Bran, et al. , Nat. Mach. Intell., 2024, 6, 525–535 CrossRef PubMed.
- H. Zhang, Y. Song, Z. Hou, S. Miret and B. Liu, Findings Assoc. Comput. Linguist.: EMNLP 2024, 2024, 3369–3382.
- M. Zaki and N. M. A. Krishnan, Digit. Discov., 2024, 3, 313–327 RSC.
- L. Gao, X. Ma, J. Lin and J. Callan, arXiv, 2022, preprint arXiv:2212.10496, DOI:10.48550/arXiv.2212.10496.
- L. Wang, N. Yang and F. Wei, Proc. 2023 Conf. Empirical Methods Nat. Lang. Process. (EMNLP), 2023, 9414–9423.
- X. Ma, Y. Gong, P. He, H. Zhao and N. Duan, Proc. 2023 Conf. Empirical Methods Nat. Lang. Process. (EMNLP), 2023, 5303–5315.
- S. Mao, et al., Findings Assoc. Comput. Linguist.: EMNLP, 2024, 2024, 884–901.
- J. Wu, et al., arXiv, 2024, preprint arXiv:2402.11166, DOI:10.48550/arXiv.2402.11166.
- C. M. Chan, et al., arXiv, 2024, preprint arXiv:2404.00610, DOI:10.48550/arXiv.2404.00610.
- J. J. Song, et al. , Chem. Soc. Rev., 2020, 49, 2196–2214 RSC.
- N. Wen, et al. , Mater. Chem. Front., 2023, 7, 4833–4864 RSC.
- S. R. Ede and Z. P. Luo, J. Mater. Chem. A, 2021, 9, 20131–20163 RSC.
- U. Peters and B. Chin-Yee, R. Soc. Open Sci., 2025, 12, 241259 Search PubMed.
- Y. Song, S. Miret and B. Liu, Proc. 61st Annu. Meet. Assoc. Comput. Linguist. (ACL), 2023, 3621–3639.
- A. Trewartha, et al. , Patterns, 2022, 3, 100488 CrossRef PubMed.
- T. Brown, et al. , Adv. Neural Inf. Process. Syst., 2020, 33, 1877–1901 Search PubMed.
- H. Liu, et al. , Adv. Neural Inf. Process. Syst., 2022, 35, 1950–1965 Search PubMed.
- J. Wei, et al. , Adv. Neural Inf. Process. Syst., 2022, 35, 24824–24837 Search PubMed.
- J. Achiam, et al., arXiv, 2023, preprint arXiv:2303.08774, DOI:10.48550/arXiv.2303.08774.
- A. J. Oche, et al., arXiv, 2025, preprint arXiv:2507.18910, DOI:10.48550/arXiv.2507.18910.
- A. S. Kasmaee, et al., NeurIPS Efficient Natural Language and Speech Processing Workshop, 2024.
- X. Yang, et al. , Adv. Neural Inf. Process. Syst., 2024, 37, 10470–10490 Search PubMed.
- S. Es, et al., Proc. 18th Conf. Eur. Chapter Assoc. Comput. Linguist.: System Demonstrations, 2024, 150–158.
- Z. Sepasdar, et al., arXiv, 2024, preprint arXiv:2409.17580, DOI:10.48550/arXiv.2409.17580.
- R. Nogueira and K. Cho, arXiv, 2017, preprint arXiv:1704.04572, DOI:10.48550/arXiv.1704.04572.
- D. Lee, H. Mizuseki, J. Choi and B. Lee, Commun. Mater., 2025, 6, 100 CrossRef.
| This journal is © The Royal Society of Chemistry 2026 |