Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
A multi-task learning approach for prediction of missing bioactivity values of compounds for the SLC transporter superfamily
Tarik
Ćerimagić
 ,
Sergey
Sosnin
,
Sergey
Sosnin
 and
Gerhard F.
Ecker
and
Gerhard F.
Ecker
 *
*
University of Vienna, Department of Pharmaceutical Sciences, Josef Holaubek Platz 2, 1090 Vienna, Austria. E-mail: gerhard.f.ecker@univie.ac.at
First published on 8th January 2026
Abstract
Solute carrier (SLC) transporters constitute the largest family of membrane transport proteins in humans. They facilitate the movement of ions, neurotransmitters, nutrients, and drugs. Given their critical role in regulating cellular physiology, they are important therapeutic targets for neurological and psychological disorders, metabolic diseases, and cancer. Inhibition of SLC transporters can modulate substrate gradients, restrict the cellular uptake of nutrients and drugs, and thereby facilitate specific pharmacological effects. Despite their pharmaceutical relevance, many SLC transporters remain understudied. Having a complete bioactivity matrix of associated compounds can expand the knowledge base of SLC ligands, enlarge the information pool to guide downstream processes, and promote informed decision-making steps in the discovery of new drug candidates for SLC transporters. To address the data sparsity of available compound-bioactivity values causing inhibitory responses for SLC transporters, we employed a multi-task learning (MTL) approach with a data imputation objective. By leveraging relationships between related tasks, deep learning has previously shown promise in imputing compound bioactivities across multiple assays. We developed a multi-task deep neural network (MT-DNN) to predict and impute missing pChEMBL (−log(IC50)) values across the SLC transporter superfamily. With a data matrix density of 2.53% and an R2 of 0.74, our model demonstrated robust predictive performance. Specifically, we predicted missing values for 9122 unique compounds across 54 SLC targets spanning various folds and subfamilies, generating 480![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 133 predictions from 12
133 predictions from 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 455 known interactions. The advantages of the multi-task learning approach were indicated in the ability of certain targets to leverage the shared representation of knowledge and acquire increased predictive accuracy over single-task learning (STL) counterparts. Despite the limitations set by low data density, activity cliffs, and inter-protein heterogeneity, the MT-DNN showed promising potential as a tool to address data sparsity within the SLC superfamily.
455 known interactions. The advantages of the multi-task learning approach were indicated in the ability of certain targets to leverage the shared representation of knowledge and acquire increased predictive accuracy over single-task learning (STL) counterparts. Despite the limitations set by low data density, activity cliffs, and inter-protein heterogeneity, the MT-DNN showed promising potential as a tool to address data sparsity within the SLC superfamily.
Introduction
SLC transporters are one of the largest groups of proteins in humans, accounting for 30% of the total proteome. The SLC superfamily consists of more than 450 members that are organized into 65 families defined by sequence homology and physiological functions.1 As membrane-bound proteins, SLC members transport different types of molecules across the membranes. These include amino acids, lipids, sugars, ions, neurotransmitters, and drugs. They exhibit different underlying transport dynamics by utilizing ion concentration gradients as symporters, translocating substrates in the opposite direction of ions as antiporters, and channel-like properties for single transported molecules as uniporters. SLC transporters are distributed across different tissues and organs, including the brain, liver, and kidneys. Being involved in the uptake and efflux of molecules relevant to phase I and phase II metabolism, they impact the absorption and elimination of drugs. Gene mutations in some SLC proteins that lead to altered or impaired ability to transport endogenous compounds contribute to the development of neurological diseases, cholesterol/bile transport defects, and cancer.2,3 Inhibition of SLC transporters can facilitate pharmacological effects with therapeutic purposes intended for treatment of different diseases.4 Targeting sugar transporters of the SLC5 family to modulate substrate translocation has been proposed for treatment of diabetes, cancer, and cardiovascular diseases.5 Transport of neurotransmitters by SLC6 family members can be disrupted by inhibitors that bind to the transporter and block the conformational changes required for substrate uptake. Such inhibition regulates neurological responses and forms the basis of several therapeutic strategies, including the treatment of depression, attention-deficit hyperactivity disorder (ADHD), and other neuropsychiatric conditions.6 Organic anion transporters (OATs) of the SLC22 family mediate the transport of negatively charged molecules such as nutrients, metabolites, toxins, and drugs. Inhibitors of OATs can modulate substrate uptake and excretion, thereby influencing the absorption and renal clearance of co-administered drugs and ultimately prolonging their exposure in patients.7 Conversely, organic cation transporters (OCTs) of the same family mediate the uptake of cationic molecules in different tissues. Inhibition of OCT3 increases extracellular levels of serotonin and norepinephrine, thereby providing an alternative therapeutic approach for the treatment of depressive disorders. However, the scarcity of specific OCT3 ligands continues to limit therapeutic development.8 Concomitant inhibition of OCTs and SLC47 members, such as multidrug and toxin extrusion protein 1 (MATE1), has been recognized as clinically relevant due to their involvement in drug–drug interactions. Successful efforts have been made with in silico methods to predict the inhibitory activity of MATE1 and aid experts in the drug discovery process.9Increased utility of artificial intelligence in drug discovery has followed technological advancements over the past years.10 Qualitative and quantitative properties of compiled datasets have a major impact on the performance of machine learning and deep learning models. The data used for training can contain various proportions of missing information due to technical errors or the intrinsic nature of the objective.11 This is evident in healthcare cases where patient information is incorrectly documented or entirely missing.12 In the context of pharmaceutical research, bioactivity information of compounds can be scarce depending on the target.13 This becomes evident when comparing industry data warehouses with public data repositories such as ChEMBL.14,15 Alternatively, there is less interest in targets that cause rare diseases, which leads to a limited information pool available on their ligands.16 Moreover, investigative studies showed that bioactivity errors and dataset size directly influence accuracy scores of machine learning models.17
Data imputation is a statistical method that is used to replace the missing values. In the context of pharmacoinformatics, it utilizes sparsely filled experimental data to impute the bioactivities or properties of compounds by leveraging the relationships between available datapoints.18 Several data imputation methods have been proposed in the cases of biological assays with sparse data matrices: (i) a single-task DNN model that was trained on a set of compounds to predict pIC50 values for each assay independently,19 (ii) a multi-task deep neural network that was trained to predict the bioactivity values across each assay simultaneously,20 (iii) feature net approach with two training steps that involve predicting activity values for each task independently in the first step and using the predicted values as features together with compound descriptors to retrain the model in the second step,19 (iv) the Alechmite DNN that uses compound descriptors and activity values as inputs; the missing activity values are substituted for the mean values, and the model is trained to iteratively update the predictions until no further improvement is observed.18,21 Additional relevant approaches to consider for data imputation are pQSAR and matrix factorization methods like Macau.22,23 The common feature among most imputation approaches that outperform single-task QSAR methods is that they establish the relationships between the tasks as endpoints to facilitate some form of knowledge sharing or transfer.24 This can lead to increased performance in sparse datasets, extend the domain of applicability for dissimilar molecules, and save computation time.25
MTL aims to generate the shared representation of knowledge by simultaneous training on multiple data domains from different tasks and, thereby, improve generalization capabilities.26 This approach can be employed to predict the bioactivity values or properties of compounds.27 Rather than focusing on activity predictions for a single protein, MTL can facilitate predictions with multiple outputs and benefit from establishing relationships between related tasks. Besides having the advantage of natural regularization, MTL models can transfer information between different tasks and increase overall accuracy. This is specifically relevant in the cases of individual proteins with scarce data, as they can potentially benefit from other similar proteins through shared knowledge. To avoid confusing the model, it is important to consider degrees of similarity and how correlated the tasks (proteins) are.28
In this study we explore MTL as a prediction tool with a data imputation objective to replace the missing pChEMBL values of 9122 unique compounds across 54 protein members of the SLC superfamily as individual tasks. We developed two MT-DNNs that were trained on different types of descriptors for comparative purposes. In addition to the overall performance, target-based performance was evaluated to inspect the abilities of individual proteins to exploit the advantages of an MTL approach. We highlight challenges associated with activity cliffs and employ dimensionality reduction methods to analyze discrepancies in the chemical space between high- and low-scoring targets. Finally, we explore comparative analysis between prediction and imputation abilities of the model. Even though protein targets belong to a single superfamily, degrees of heterology between them can vary depending on the organizational hierarchy they occupy. They belong to different families, subgroups, and folds, and have different transport mechanisms, substrates, and tissue localizations.29 Because of this, it is important to consider putative inconsistencies in degrees of correlation between the targets as individual tasks that can lead to negative transfers.30
Materials and methods
Data acquisition and processing
All data operations and experiments were conducted using the Python programming language (v3.11.5). The list of SLC transporters was acquired from the Resolute Knowledgebase31 and merged with the UniProt human-protein dataset32 to associate UniProt-ID values with the corresponding target names. The compiled SLC protein dataset contained 446 unique UniProt-ID numbers, gene names, and protein names. The list of UniProt-IDs was queried via the ChEMBL Database API (ChEMBL web services) to retrieve the corresponding ChEMBL-IDs of individual targets.33The ChEMBL-IDs of targets were imported into a Jupyter Notebook for compound retrieval from the ChEMBL33 database and processing with RDKit (v2024.03.05).14 Compounds that contained inorganic elements were excluded, stereochemistry information was removed, and standardized SMILES strings were calculated. The lowest pChEMBL value in the dataset was 4.0, corresponding to an IC50 of 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 nM. During an analysis, it could not be confirmed whether these entries at the lower end of the inhibitory potency distribution represented actual dose–response measurements or simply reflected minimum reporting thresholds (100
000 nM. During an analysis, it could not be confirmed whether these entries at the lower end of the inhibitory potency distribution represented actual dose–response measurements or simply reflected minimum reporting thresholds (100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 or 10
000 or 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 nM). Therefore, data points with IC50 values of exactly 100
000 nM). Therefore, data points with IC50 values of exactly 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 and 10
000 and 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 nM were excluded. To ensure consistency across the 54 modeled targets, entries containing the substrings “muta” or “recombi” in the assay description column were removed. Mutagenized or recombinant variants could introduce inconsistencies in target representation and confound the learning process of the multi-task model. Excluding these entries helped maintain a uniform definition of targets and improved the reliability of cross-target bioactivity estimation. Duplicate measurements of pChEMBL values from identical compound–target pairs were replaced by the mean values if the SEM (standard error of the mean) was lower than 0.2. Conversely, compound–target pairs with an SEM higher than 0.2 were excluded from the dataset. Remaining duplicate entries were consequently removed so that only one measurement was kept per compound–target pair in the dataset.
000 nM were excluded. To ensure consistency across the 54 modeled targets, entries containing the substrings “muta” or “recombi” in the assay description column were removed. Mutagenized or recombinant variants could introduce inconsistencies in target representation and confound the learning process of the multi-task model. Excluding these entries helped maintain a uniform definition of targets and improved the reliability of cross-target bioactivity estimation. Duplicate measurements of pChEMBL values from identical compound–target pairs were replaced by the mean values if the SEM (standard error of the mean) was lower than 0.2. Conversely, compound–target pairs with an SEM higher than 0.2 were excluded from the dataset. Remaining duplicate entries were consequently removed so that only one measurement was kept per compound–target pair in the dataset.
After the processing steps, the table was formatted into a data matrix intended for a multi-task imputation problem. Each compound was assigned the name of the associated target in the label column designated for the stratified split. In cases where multiple targets were measured for a single compound, the name of the target with the lowest number of compounds in the dataset was assigned. This ensured proportional representation of each target in the training, validation, and test sets. A split ratio of 80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10
10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 for training, validation, and test sets was maintained across all targets to ensure consistency. To establish valid evaluation across all endpoints, targets with fewer than ten compounds were excluded so that each modeled target contained at least one datapoint in both validation and test sets. Finally, the data matrix comprised 9122 unique compounds across 54 targets and 25 families, with a data density of 2.53% (Table 1). Two sets of descriptors were used for comparative purposes. The Continuous Data Driven Descriptors (CDDD) are calculated using an autoencoder model that was trained to represent molecular formats such as SMILES as vectors of 512 continuous values.34 The repository containing the model and instructions to calculate CDDDs is available on GitHub (https://github.com/jrwnter/cddd). A second set of descriptors was 1024 extended connectivity fingerprints with a radius of 3 (ECFP6), which were calculated with a CDK module.35 The ECFP6 are circular fingerprints that contain structural information of a given molecule in the form of the binary values (0/1).36
10 for training, validation, and test sets was maintained across all targets to ensure consistency. To establish valid evaluation across all endpoints, targets with fewer than ten compounds were excluded so that each modeled target contained at least one datapoint in both validation and test sets. Finally, the data matrix comprised 9122 unique compounds across 54 targets and 25 families, with a data density of 2.53% (Table 1). Two sets of descriptors were used for comparative purposes. The Continuous Data Driven Descriptors (CDDD) are calculated using an autoencoder model that was trained to represent molecular formats such as SMILES as vectors of 512 continuous values.34 The repository containing the model and instructions to calculate CDDDs is available on GitHub (https://github.com/jrwnter/cddd). A second set of descriptors was 1024 extended connectivity fingerprints with a radius of 3 (ECFP6), which were calculated with a CDK module.35 The ECFP6 are circular fingerprints that contain structural information of a given molecule in the form of the binary values (0/1).36
| Families | Targets | Compounds | Data density |
|---|---|---|---|
| 25 | 54 | 9122 | 2.528% |
Two different approaches, shown in Fig. 1, were chosen to split the data. (Fig. 1B) Stratified prediction split: a target-based stratified split was used to allocate 80% of the compounds to the training set and 10% to the validation and test sets, respectively. Each compound, along with its complete bioactivity profile, was treated as a single data point to ensure non-overlapping compound distributions across sets. Random imputation split (Fig. 1C): in this approach, data were split according to bioactivity measurements rather than individual compounds. Compounds with two or more available bioactivity values were divided into two data points, each representing a different subset of their bioactivity profiles. One data point was assigned to the training set, and the other was randomly allocated to either the validation or test set. This procedure artificially introduced missing values in the training data, which were then used for evaluation. Consequently, all compounds from the validation and test sets were also present in the training set but with differing bioactivity profiles. This setup was designed for imputation-specific evaluation and contrasts with the compound-based prediction split due to intentional data leakage. Although identical compounds were used during training, different output values were used to evaluate knowledge sharing/transfer across targets.
Quantitative profiles after applying two splitting approaches are shown in Table 2.
| Dataset | Stratified prediction split | Random imputation split |
|---|---|---|
| Training | 7297 | 9122 |
| Validation | 913 | 1200 |
| Test | 912 | 1200 |
Hyperparameter optimization
Hyperparameters were tuned with the Optuna library (v3.6.1) by setting the range of values or categories for the model type, quantity of hidden layers, hidden-layer sizes, sizes of task-specific layers, learning rate, and dropout rate.37 Parameters in Table 3 are shown as ranges of values in “()” and categories as “[]”. The Bayesian search determined the optimal parameters by minimizing the MSE value of the validation set calculated at the end of each training iteration (trial). A task-specific domain with two layers was conditioned with the model type being set to “shared and specific”. If the model type was set to “shared”, the task-specific layer took the role of an output layer with the size corresponding to the number of modeled tasks (targets). Analogously, the size of the third layer was conditioned by the Hidden depth being set to 3. The batch size was set to 128. After 300 iterations, the best-scoring set of parameters was selected to determine the architecture and hyperparameters of the model.| Parameter | Suggested value range-[]/category-() |
|---|---|
| Model type | (Shared, shared and specific) |
| Hidden depth | (2, 3) |
| 1st hidden layer | [100–1200] |
| 2nd hidden layer | [50–500] |
| 3rd hidden layer | [20–200] |
| 1st specific layer | [3–30] |
| 2nd specific layer | [2–50] |
| Learning rate | [0.000040–0.000170] |
| Dropout rate | [0.07–0.50] |
| Patience | [7–10] |
MT-DNN architecture
The models were built as multiple-input/multiple-output feedforward backpropagation deep neural networks with PyTorch (v2.3.0). They consist of an input layer, two or three fully connected hidden layers, and one task-specific/output layer. LeakyReLU was set as an activation function. The sizes of the input and output layers were determined by the number of descriptors and tasks, respectively. The sizes of hidden layers were determined through hyperparameter optimization steps. The loss function was calculated as the masked mean squared error (MSE) between the predicted and true values in the output layer (1). To ensure that the model parameters were exclusively updated with the MSE/loss derived from available values during training, predicted and true outputs were masked at indices where no bioactivity measurement was originally observed. Adam was selected as a method for stochastic optimization of the models. The MT-DNNs were trained on the training set with the validation MSE/loss calculated at each epoch to evaluate generalization performance and learning dynamics. To limit overfitting, a failsafe mechanism, referred to previously as the patience parameter, was set to terminate the training loop at the step where no improvement of the mean MSE validation value was observed for 7–10 consecutive epochs.Model evaluation
The model was evaluated with descriptor-type performance discrepancies, overall-prediction performance, learning dynamics, outlier vs. inlier evaluation, target-based prediction performance, visualization of the chemical space, and overall-imputation performance with benchmarking. To assess how well the entire dataset was represented by the model, 5-fold CV was implemented for the prediction splits. Every metric indicated in Table 4 followed the masked indexing principle of the MSE/loss function (1). Modules for data splitting, CV, and evaluation were adopted from scikit-learn.38 Three categories were chosen for an MTL-STL comparative analysis with aggregate task-specific evaluation: (1) unweighted: calculating mean scores and percentages by assuming that all targets are equally relevant. (2) Weighted: mean scores and percentages are adjusted by assigning higher relevance to targets with a higher number of compounds in the dataset. (3) Inversely weighted: mean scores and percentages are adjusted by assigning higher relevance to targets with a lower number of compounds in the dataset. In this evaluation step, category 3 could be considered the most representative form of quality assessment for the knowledge sharing capabilities of the multi-task imputation framework, indicating advantages/disadvantages the model has for underrepresented targets. Three targets that represent distinct performance profiles observed during target-based evaluation were further analyzed with 2D chemical space representation. Furthermore, as outlined in the section Data acquisition and processing, conceptionally different approaches for data splitting were applied: (i) in the case of the stratified prediction split, compounds are split such that entire molecules are held out from training and the task corresponds to out-of-sample prediction. In this setting, the model is evaluated on unseen compounds, and the objective is to assess generalization performance, descriptor suitability, and target-specific predictive accuracy. (ii) In random imputation split, the same MT-DNN architecture was retrained under a matrix-completion setting, where the compound set is fixed and bioactivity values for certain targets are intentionally removed during training. In this case, compounds are present as input entities during training but removed target labels do not contribute to the MSE/loss function. Model performance is then evaluated on these withheld entries. This setting corresponds to a data imputation scenario, i.e., estimating missing values within a partially observed compound–target activity matrix.Single-task models
Single-task deep neural networks (ST-DNN) and single-task random forest (ST-RF) models were generated with PyTorch and scikit-learn respectively to conduct a comparative analysis between the multi-task and single-task learning approaches. The hyperparameters for each single-task model were tuned with a Bayesian search. The hyperparameter ranges shown in SI Table 1 for the ST-DNN, as well as the model architectures, were set to reflect the MT-DNN model. The hyperparameter ranges of the ST-RF models are shown in SI Table 2. To maintain consistency across all three approaches, the optimum hyperparameters were selected based on the minimum MSE value over 300 trials with Optuna. The data were split randomly, and 5-fold CV was implemented in the evaluation step to assess the performance across the entire dataset.Chemical space
The multi-dimensional information of chemical descriptors is reduced to two or three dimensions for visual interpretation. This way, relationships between different compounds can be observed while taking into consideration the context of the entire chemical dataset. Dimensionality reduction algorithms can utilize different types of chemical descriptors to project the chemical space from a set of compounds.39 The principles behind the Uniform Manifold Approximation and Projection (UMAP) are a series of mathematical operations that reduce the dimensionality of features/descriptors while preserving the local structure of data. This method attempts to keep neighboring data points in close proximity and thereby preserve their relationship when mapping from high dimensionality in low-dimensional space. UMAP (v0.5.5) was used for visual representation of the chemical space in this study.40 A set of 9122 unique compounds with 512 CDDD descriptors was used for the chemical space analysis. After the dimensionality reduction (Fig. 2), 2D embeddings were associated with the corresponding SMILES strings. The scatterplot of the chemical space was generated with the Matplotlib library. Compounds were color-coded based on the associated targets that were selected in the evaluation. The parameters shown in SI Table 3 for the UMAP model were tuned to represent the compounds belonging to common families as neighbors in 2D and thereby form family-associated clusters. | ||
| Fig. 2 Chart showing the dimensionality reduction of 512 CDDD descriptors into 2D embeddings for an exemplary set of 2 compounds intended for chemical space visualization. | ||
Data imputation models for benchmarking
To evaluate the imputation performance of the MT-DNN, six additional imputation frameworks were adopted in this workflow for comparative analysis. The choice of benchmarking models was based on accessibility and technical reasons related to time and cost restraints. Three models were selected from the scikit-learn library (SimpleImputer, KNNImputer, and IterativeImputer) and three from the fancyimpute library (SoftImpute, IterativeSVD, and MatrixFactorization).41 Datasets obtained through the imputation split (see Fig. 1) were used to develop the models. To ensure consistency in the comparative analysis, the same data format, comprising CDDD descriptors and a sparsely populated bioactivity matrix used for MT-DNN training, was also employed to train the benchmarking imputation models. Models followed a similar strategy to the MT-DNN development by using training data for fitting, validation data to optimize hyperparameters, and test data for evaluation. Parameter ranges shown in SI Table 4 were set for a grid search of the respective imputation models.For easier comprehension of the project development pipeline, an overview of the entire workflow can be seen in Fig. 3.
Results and discussion
MT-DNN prediction performance
The parameters and architecture of the models shown in Table 5 were selected according to a Bayesian search performed with the Optuna library. Both models achieved the lowest MSE value of the validation set without the task-specific domain, indicating that the standalone shared domain of the hidden layers contributes positively to the overall performance of the model. Furthermore, the CDDD model has 2 hidden layers, whereas the ECFP6 model has 3 hidden layers in the shared domain. Considering the results from the evaluation metrics in Table 6, the CDDD model was able to achieve higher performance with less architectural complexity than the ECFP6 model. With Q2 and R2 values being closer to the MTS in Table 6, the CDDD model indicates a lower tendency to overfit to the training data. Additionally, the learning dynamics comparison in Fig. 4 shows that the MSE loss of the validation set from the CDDD model has a closer loss curve to the training data when compared to the ECFP6 model. This trend remains consistent overall and was a decisive factor for continuing to use CDDD descriptors in further development and evaluation of the MTL approach.| Hyperparameters | Models | |
|---|---|---|
| CDDD | ECFP6 | |
| a Hidden layer (HL). | ||
| Input size | 512 | 1024 |
| HL1 size | 800 | 1160 |
| HL2 size | 250 | 230 |
| HL3 size | — | 190 |
| Task-specific size | 54 | 54 |
| Learning rate | 0.000043 | 0.000046 |
| Dropout rate | 0.1 | 0.08 |
| Patience | 10 | 7 |
| Model | MTS↑ | Cross-validation | Test data | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Q 2↑ | MSE↓ | MAE↓ | RMSE↓ | R 2↑ | MSE↓ | MAE↓ | RMSE↓ | ||
| a Bold – the best value in a row. | |||||||||
| CDDD | 0.915 | 0.716 | 0.420 | 0.486 | 0.650 | 0.739 | 0.362 | 0.451 | 0.602 |
| ECFP6 | 0.952 | 0.661 | 0.501 | 0.536 | 0.708 | 0.689 | 0.432 | 0.477 | 0.657 |
Descriptor-associated discrepancy in the performance proposes higher ability by CDDD representation of compounds to capture the diversity of chemical space that was necessary to model the SLC superfamily. As only one additional descriptor type was considered, further comparative studies would be required for conclusive statements.
Activity cliffs
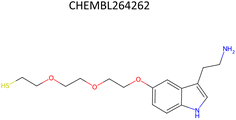
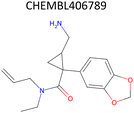
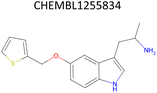
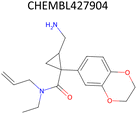
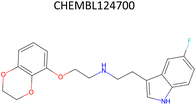
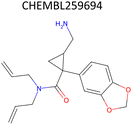
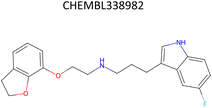
Analysis of pChEMBL predictions with the highest error values (HEV) for a given compound–target pair was conducted to identify the shortcomings of the MTL approach with CDDD descriptors. The SLC6A4 target (serotonin transporter, SERT) had instances of both HEV and low error value (LEV) predictions that showcase putative effects of activity cliffs on the performance of the model. Both examples are compared with their nearest neighbors identified via the NearestNeighbors module from scikit-learn and corresponding activity profiles. The HEV case in Table 7 shows that the predicted pChEMBL of 6.39 falls within the range of values of the nearest neighbors for the same target. However, the actual pChEMBL value of the HEV compound is 4.0. Considering that the three closest neighbors of the HEV compound in the training set had a mean pChEMBL value of 6.8, overprediction by the model could be attributed to the training-set bias. Inversely, the LEV case in Table 7 has the actual and predicted values of the test set compound falling within the bioactivity ranges of the nearest neighbors in the training set. When considering further comparison of the two examples, a higher degree of structural similarity between the LEV compound and its nearest neighbors can be observed. The challenge associated with activity cliffs could be addressed by implementing proportional distribution of biological activities across similar compounds or respective targets. This approach could ensure sufficient representation of various activity ranges between different targets and induce a balanced learning profile by the model.| HEV compound (activity cliff example) – left panel | Actual pChEMBL | Predicted pChEMBL | LEV compound – right panel | Actual pChEMBL | Predicted pChEMBL |
|---|---|---|---|---|---|
| a ChEMBL-IDs and actual pChEMBL values are annotated for each compound. Predicted pChEMBL values are annotated only for the HEV and LEV compounds in the first row because they were allocated to the test set. The nearest neighbors (rows 2–4) were allocated to the training set and, therefore, no prediction of the pChEMBL value was made during evaluation. Left panel: an activity cliff example with the HEV compound in the first row and its three nearest neighbors below. Right panel: LEV compound in the first row and its three nearest neighbors below. | |||||
|
4.0 | 6.39 |
|
6.82 | 6.82 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||
| Nearest neighbors of the HEV compound in the training set | Nearest neighbors the LEV compound in the training set | ||||
|
5.32 | — |
|
7.19 | — |
|
7.07 | — |
|
7.24 | — |
|
8.01 | — |
|
6.37 | — |
Target-based evaluation
Performance scores for individual targets were calculated as Q2 values using MT-DNN, ST-DNN, and ST-RF models during 5-fold CV and the results are shown in SI Table 5. Targets that yielded negative Q2 scores across all three models were excluded from the comparative analysis. The remaining 31 targets that were considered for the aggregate analysis comprised 8698 unique compounds, representing 95.4% of the dataset. When compared to other methods in Table 8, the MT-DNN model outperformed the single-task models in all categories, except for the aggregate standard deviation (SD) scores of the ST-RF model in the unweighted and weighted categories. The most notable improvement was observed in the inversely weighted category, indicating that underrepresented targets mostly benefited from the MT-DNN model without compromising performance in other categories. Additionally, more than 60% of the targets showed improved scores with the multi-task learning (MTL) approach.| Model | Unweighted | Weighted | Inversely weighted | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Q 2 | Q 2 − SD | TSH (%) | Q 2 | Q 2 − SD | TSH (%) | Q 2 | Q 2 − SD | TSH (%) | |
| a Every category has three scores across 31 targets: Q2 – mean of cross-validated coefficients of determination, Q2 − SD – mean of standard deviations for the Q2 value, and TSH – percentage of targets that improved with the corresponding model. | |||||||||
| MT-DNN | 0.280 | 0.267 | 61.290 | 0.521 | 0.098 | 51.556 | 0.247 | 0.326 | 70.770 |
| ST-DNN | −0.050 | 0.590 | 12.903 | 0.454 | 0.169 | 33.026 | −0.475 | 1.046 | 5.299 |
| ST-RF | 0.232 | 0.205 | 25.806 | 0.496 | 0.075 | 15.417 | 0.069 | 0.340 | 23.931 |
To showcase discrepancies between positive and negative transfers with MTL, three specific targets with different performance profiles were analyzed using a 2D representation of the chemical space in Fig. 5. The entire dataset was included in the chemical space analysis to represent relationships between individual clusters. Starting with the highest-scoring target in Table 9, SLC5A1 (sodium/glucose co-transporter, SGLT1) had a Q2 of 0.809. The chemical space of compounds associated with SLC5A1 shows a wider degree of coverage between multiple domains when compared to the other two targets in Fig. 5. The compounds of the lowest-scoring target, SLC33A1 (acetyl-coenzyme A transporter 1), with a Q2 of −2.554, contrast with the distribution pattern of the SLC5A1 chemical space by having a localized and entirely isolated cluster. Based on this representation, it is evident that no other member shares similar chemical space with SLC33A1. It cannot be excluded that the cause of the discrepancy in the performance between the two targets stems from quantitative imbalance of the dataset. SLC5A1 contributes 975 datapoints to the overall dataset, whereas SLC33A1 contributes 131 datapoints. Furthermore, SLC5A1 belongs to the SLC5 family with three additional protein members modeled as separate tasks that collectively amount to 2194 compounds in the dataset. Such instances of correlated tasks would, as stated in the introduction, be able to leverage the shared representation of knowledge generated with an MTL approach. To further support this hypothesis, a Q2 value of 0.758 is observed for the SLC5A4 (sodium/glucose cotransporter 3, SGLT3) target with 28 compounds in the dataset. Although it was represented by fewer datapoints, SLC5A4 scored higher than SLC33A1 in Table 9. As highlighted in Fig. 5, compounds of SLC5A4 share the chemical space with other targets and do not form isolated clusters. Unlike SLC33A1, which was the single member of the SLC33 family in the dataset, SLC5A4 belongs to the SLC5 family that is shared with three other members. Furthermore, comparison of the performance scores between the MT-DNN and ST-RF models in Table 9 suggests that the SLC5 members benefit from the MTL approach, whereas the SLC33A1 target was able to achieve a higher score with the ST-RF model. Understudied targets such as SLC5A4 have little information published on their active compounds when compared to some other members of the SLC superfamily. To the best of our knowledge, this MTL model presents itself currently as a unique tool for compound bioactivity prediction and imputation of the SGLT3 (SLC5A4) transporter.
 | ||
| Fig. 5 Chemical space representation with UMAP. Individual compounds are colored according to associated targets marked in the legend. | ||
| Target | Compounds | Family members | Total compounds | MT-DNN (Q2)↑ | ST-DNN (Q2)↑ | ST-RF (Q2)↑ |
|---|---|---|---|---|---|---|
| a The table shows the quantitative properties of selected targets with individual performance scores represented as Q2 and standard deviation from the MT-DNN, ST-DNN, and ST-RF models. The table additionally shows the number of compounds for each target and their respective families. Bold – the best value in a row between the three models. | ||||||
| SLC5A1 | 975 | 4 | 2194 | 0.809 ± 0.006 | 0.795 ± 0.021 | 0.799 ± 0.025 |
| SLC33A1 | 131 | 1 | 131 | −2.554 ± 1.548 | −2.866 ± 3.151 | 0.086 ± 0.198 |
| SLC5A4 | 28 | 4 | 2194 | 0.758 ± 0.107 | 0.428 ± 0.587 | 0.667 ± 0.201 |
Data imputation performance
An imputation-dedicated split was chosen to evaluate the ability of an MTL approach to predict the pChEMBL values of compounds for targets not used in the output layer during training. This means that the model is tested on compounds that are available in the training set but show different output profiles. Unlike the conventional approach where the model is tested on an entirely new and previously unseen set of compounds, this step examines how well the model performs in an imputation-driven setting by estimating pChEMBL values of known compounds for a different target.Since neither the CDDD nor ECFP6 prediction models achieved a minimum MSE validation value with the inclusion of task-specific neural network domains (Table 5), only the shared domain was incorporated as a parameter during the optimization of the CDDD imputation model. As shown in Table 10, the CDDD imputation model achieved the lowest MSE value for the validation set with higher architectural complexity than the prediction counterpart. One additional hidden layer was selected along with a higher learning rate and identical patience to counter overfitting.
| Hyperparameters | Models | |
|---|---|---|
| CDDD prediction | CDDD imputation | |
| Input size | 512 | 512 |
| H1 size | 800 | 1020 |
| H2 size | 250 | 170 |
| H3 size | — | 80 |
| Task-specific size | 54 | 54 |
| Learning rate | 0.000043 | 0.000105 |
| Dropout rate | 0.1 | 0.09 |
| Patience | 10 | 10 |
The CV step was excluded in this section due to the lack of a sufficient number of compounds with five or more bioactivity measurements for accurate allocation in each fold. The results in Table 11 show that CDDD imputation was able to score relatively close to the CDDD prediction model. A slight improvement in the prediction performance over the imputation performance could be attributed to the skewed data distribution introduced by the imputation split, which may affect generalization. The imputation split only considers compounds with two or more measurements, potentially biasing target representation, molecule types, and activity values.
| Metrics | Models | ||
|---|---|---|---|
| CDDD prediction | CDDD imputation | ||
| a Bold – the best value in a row. | |||
| MTS↑ | 0.942 | 0.940 | |
| Validation set | R 2↑ | 0.754 | 0.705 |
| MSE↓ | 0.377 | 0.429 | |
| MAE↓ | 0.465 | 0.503 | |
| RMSE↓ | 0.615 | 0.655 | |
| Test set | R 2↑ | 0.739 | 0.720 |
| MSE↓ | 0.362 | 0.407 | |
| MAE↓ | 0.451 | 0.477 | |
| RMSE↓ | 0.602 | 0.638 | |
Comparative analysis between accuracy scores on the test set in Table 12 shows that the MT-DNN outperforms all six imputation methods chosen for benchmarking. The second-highest scoring method is k-Nearest Neighbor from the sklearn.imputers module. Considering both high prediction and imputation performance, MTL presents itself as a suitable and versatile tool in bioactivity-value estimation of compounds towards SLC transporters.
| MT-DNN | sklearn.imputers | fancyimpute | |||||
|---|---|---|---|---|---|---|---|
| S-mean | KNN | Iterative | Soft | I-SVD | MF | ||
| a Scores are calculated on the test set that was derived from the random imputation split. Column 2 – MT-DNN imputation. Columns 3 to 5 – sklearn.imputers: S-mean – SimpleImputer with mean method, kNN – k Nearest Neighbor, and Iterative – IterativeImputer. Columns 6 to 8 – fancyimpute: Soft – SoftImpute, I-SVD – IterativeSVD, and MF – MatrixFactorization. | |||||||
| R 2↑ | 0.720 | 0.098 | 0.672 | 0.471 | −0.400 | 0.107 | 0.489 |
| MSE↓ | 0.407 | 1.312 | 0.477 | 0.667 | 2.033 | 1.299 | 0.744 |
Conclusion
With this study, we present an MTL-based prediction to complete a bioactivity matrix of compounds for the superfamily of SLC transporters. Besides replacing the missing values, the MT-DNN could predict bioactivity profiles for out-of-sample compounds and showcase overall satisfactory performance scores. Further results indicate that the selection of different types of descriptors can affect the ability of the model to achieve reliable predictions. In this case, the causality of descriptor-type discrepancies in the performance is not completely understood, and further studies are necessary for conclusive statements. Based on individual bioactivity-prediction error analysis and target-based evaluation, it cannot be excluded that the shortcomings of our MTL approach are associated with the presence of activity cliffs and heterogeneity between the modeled targets. The dataset contains compounds with similar structural properties but contrasting bioactivity values or sets of associated targets. If imbalanced in quantity and distribution between the training, validation, and test sets, such outlier instances designated as activity cliffs can impact the performance of the model. A similar phenomenon was observed for a target-specific outlier, where the MTL model struggled to accurately predict the bioactivity values of compounds that formed an isolated cluster in the chemical space analysis. Conversely, the performance of SLC5A4 suggests that a target with a low quantity of datapoints could benefit from the MTL approach if it shares certain qualitative and quantitative properties with other correlated targets, such as having common families or similar chemical space. In conclusion, the comparative analysis demonstrates the suitability of the multi-task model as a prediction tool intended for a data imputation task. Besides scoring highest during benchmarking against six established imputation methods, MT-DNN outperformed the single-task models by achieving higher accuracy for underrepresented targets. Despite this improvement bias, the MT-DNN model achieved comparable or slightly higher scores for most other targets with larger numbers of compounds, highlighting its strengths in handling missing bioactivity data for an imbalanced target space.Author contributions
TĆ: conceptualization, data acquisition, data processing and standardization, literature research, methodology, evaluation, visualization and validation of the results, Python code writing and documentation, and writing of the manuscript. SS: conceptualization, supervision, and draft review. GFE: conceptualization, project administration, supervision, resources, and draft review.Conflicts of interest
There are no conflicts of interest to declare.Data availability
The data used for this study are available on the Resolute Knowledgebase at: https://doi.org/10.5281/zenodo.4309586, https://re-solute.eu/knowledgebase/gene. The UniProt dataset is available at: https://doi.org/10.1093/nar/gkae1010, https://www.uniprot.org/uniprotkb?query=*%26facets=model_organism%3A9606. The intermediary datasets generated after standardization, processing, and calculating descriptors are available on the GitHub repository along with the Jupyter Notebook and Python scripts used for the development of this study at: https://doi.org/10.5281/zenodo.17991114, https://github.com/PharminfoVienna/SLC-data-imputation.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5dd00536a.
Acknowledgements
The contributions of Leo Gaskin are hereby acknowledged for his technical support during the development of the Jupyter Notebook and Python scripts. The graphical abstract was created with https://www.drawio.com/ and BioRender.com (https://app.biorender.com/illustrations/66d5af4e119d8a2615a267d8).Notes and references
- T. Xie, X. Chi, B. Huang, F. Ye, Q. Zhou and J. Huang, Rational exploration of fold atlas for human solute carrier proteins, Structure, 2022, 30, 1321–1330 CrossRef CAS PubMed.
- B. White and P. Swietach, What can we learn about acid-base transporters in cancer from studying somatic mutations in their genes?, Pflügers Archiv: European Journal of Physiology, 2024, 476, 673–688 CrossRef CAS PubMed.
- X. Liu, in Drug Transporters in Drug Disposition, Effects and Toxicity, Springer Singapore Pte. Limited, Singapore, 2019, pp. 4–6 Search PubMed.
- L. Lin, S. W. Yee, R. B. Kim and K. M. Giacomini, SLC transporters as therapeutic targets: emerging opportunities, Nat. Rev. Drug Discovery, 2015, 14, 543–560 CrossRef CAS PubMed.
- G. Gyimesi, J. Pujol-Giménez, Y. Kanai and M. A. Hediger, Sodium-coupled glucose transport, the SLC5 family, and therapeutically relevant inhibitors: from molecular discovery to clinical application, Pflügers Archiv: European Journal of Physiology, 2020, 472, 1177–1206 CrossRef CAS PubMed.
- A. B. Pramod, J. Foster, L. Carvelli and L. K. Henry, SLC6 transporters: structure, function, regulation, disease association and therapeutics, Mol. Aspects Med., 2013, 34, 197–219 Search PubMed.
- Z. Yu and G. You, Recent Advances on the Regulations of Organic Anion Transporters, Pharmaceutics, 2024, 16, 1355 CrossRef CAS PubMed.
- B. Khanppnavar, J. Maier, F. Herborg, R. Gradisch, E. Lazzarin, D. Luethi, J.-W. Yang, C. Qi, M. Holy, K. Jäntsch, O. Kudlacek, K. Schicker, T. Werge, U. Gether, T. Stockner, V. M. Korkhov and H. H. Sitte, Structural basis of organic cation transporter-3 inhibition, Nat. Commun., 2022, 13, 6714 Search PubMed.
- K. Handa, S. Sasaki, S. Asano, M. Kageyama, T. Iijima and A. Bender, Prediction of Inhibitory Activity against the MATE1 Transporter via Combined Fingerprint- and Physics-Based Machine Learning Models, J. Chem. Inf. Model., 2024, 64, 7068–7076 Search PubMed.
- A. U. Rehman, M. Li, B. Wu, Y. Ali, S. Rasheed, S. Shaheen, X. Liu, R. Luo and J. Zhang, Role of Artificial Intelligence in Revolutionizing Drug Discovery, Fundam. Res., 2024, S266732582400205X Search PubMed.
- W.-C. Lin and C.-F. Tsai, Missing value imputation: a review and analysis of the literature (2006–2017), Artificial Intelligence Review, 2020, 53, 1487–1509 CrossRef.
- J. Li, X. S. Yan, D. Chaudhary, V. Avula, S. Mudiganti, H. Husby, S. Shahjouei, A. Afshar, W. F. Stewart, M. Yeasin, R. Zand and V. Abedi, Imputation of missing values for electronic health record laboratory data, npj Digital Medicine, 2021, 4, 147 CrossRef PubMed.
- Forum on Neuroscience and Nervous System Disorders, Board on Health Sciences Policy and Institute of Medicine, Improving and Accelerating Therapeutic Development for Nervous System Disorders: Workshop Summary, National Academies Press, Washington, D.C., 2014 Search PubMed.
- A. Smajić, I. Rami, S. Sosnin and G. F. Ecker, Identifying Differences in the Performance of Machine Learning Models for Off-Targets Trained on Publicly Available and Proprietary Data Sets, Chem. Res. Toxicol., 2023, 36, 1300–1312 Search PubMed.
- D. Mendez, A. Gaulton, A. P. Bento, J. Chambers, M. De Veij, E. Félix, M. P. Magariños, J. F. Mosquera, P. Mutowo, M. Nowotka, M. Gordillo-Marañón, F. Hunter, L. Junco, G. Mugumbate, M. Rodriguez-Lopez, F. Atkinson, N. Bosc, C. J. Radoux, A. Segura-Cabrera, A. Hersey and A. R. Leach, ChEMBL: towards direct deposition of bioassay data, Nucleic Acids Res., 2019, 47, D930–D940 Search PubMed.
- L. J. Fermaglich and K. L. Miller, A comprehensive study of the rare diseases and conditions targeted by orphan drug designations and approvals over the forty years of the Orphan Drug Act, Orphanet Journal of Rare Diseases, 2023, 18, 163 CrossRef PubMed.
- F. J. Fan and Y. Shi, Effects of data quality and quantity on deep learning for protein-ligand binding affinity prediction, Bioorg. Med. Chem., 2022, 72, 117003 Search PubMed.
- B. W. J. Irwin, S. Mahmoud, T. M. Whitehead, G. J. Conduit and M. D. Segall, Imputation Versus Prediction: Applications in Machine Learning for Drug Discovery, Future Drug Discovery, 2020, 2, FDD38 CrossRef.
- S. Sosnin, D. Karlov, I. V. Tetko and M. V. Fedorov, Comparative Study of Multitask Toxicity Modeling on a Broad Chemical Space, J. Chem. Inf. Model., 2019, 59, 1062–1072 Search PubMed.
- A. Mayr, G. Klambauer, T. Unterthiner and S. Hochreiter, DeepTox: Toxicity Prediction using Deep Learning, Front. Environ. Sci., 2016, 3, 80 Search PubMed.
- T. M. Whitehead, B. W. J. Irwin, P. Hunt, M. D. Segall and G. J. Conduit, Imputation of Assay Bioactivity Data Using Deep Learning, J. Chem. Inf. Model., 2019, 59, 1197–1204 Search PubMed.
- E. J. Martin, V. R. Polyakov, L. Tian and R. C. Perez, Profile-QSAR 2.0: Kinase Virtual Screening Accuracy Comparable to Four-Concentration IC50s for Realistically Novel Compounds, J. Chem. Inf. Model., 2017, 57, 2077–2088 Search PubMed.
- A. De La Vega De León, B. Chen and V. J. Gillet, Effect of missing data on multitask prediction methods, J. Cheminf., 2018, 10, 26 Search PubMed.
- Z. Zhao, J. Qin, Z. Gou, Y. Zhang and Y. Yang, Multi-task learning models for predicting active compounds, J. Biomed. Inf., 2020, 108, 103484 Search PubMed.
- M. Walter, L. N. Allen, A. De La Vega De León, S. J. Webb and V. J. Gillet, Analysis of the benefits of imputation models over traditional QSAR models for toxicity prediction, J. Cheminf., 2022, 14, 32 Search PubMed.
- P. Knight and R. Duan, Multi-Task Learning with Summary Statistics, Advances in Neural Information Processing Systems, 2023, 36, 54020–54031 Search PubMed.
- M. Walter, J. M. Borghardt, L. Humbeck and M. Skalic, Multi-Task ADME/PK prediction at industrial scale: leveraging large and diverse experimental datasets**, Mol. Inf., 2024, 43, e202400079 Search PubMed.
- S. Allenspach, J. A. Hiss and G. Schneider, Neural multi-task learning in drug design, Nature Machine Intelligence, 2024, 6, 124–137 CrossRef.
- C. Colas, P. M.-U. Ung and A. Schlessinger, SLC transporters: structure, function, and drug discovery, MedChemComm, 2016, 7, 1069–1081 Search PubMed.
- D. Li, H. L. Nguyen and H. R. Zhang, Identification of Negative Transfers in Multitask Learning Using Surrogate Models, arXiv, 2023, preprint, arXiv:2303.14582, DOI:10.48550/ARXIV.2303.14582.
- G. Superti-Furga, D. Lackner, T. Wiedmer, A. Ingles-Prieto, B. Barbosa, E. Girardi, U. Goldmann, B. Gürtl, K. Klavins, C. Klimek, S. Lindinger, E. Liñeiro-Retes, A. C. Müller, S. Onstein, G. Redinger, D. Reil, V. Sedlyarov, G. Wolf, M. Crawford, R. Everley, D. Hepworth, S. Liu, S. Noell, M. Piotrowski, R. Stanton, H. Zhang, S. Corallino, A. Faedo, M. Insidioso, G. Maresca, L. Redaelli, F. Sassone, L. Scarabottolo, M. Stucchi, P. Tarroni, S. Tremolada, H. Batoulis, A. Becker, E. Bender, Y.-N. Chang, A. Ehrmann, A. Müller-Fahrnow, V. Pütter, D. Zindel, B. Hamilton, M. Lenter, D. Santacruz, C. Viollet, C. Whitehurst, K. Johnsson, P. Leippe, B. Baumgarten, L. Chang, Y. Ibig, M. Pfeifer, J. Reinhardt, J. Schönbett, P. Selzer, K. Seuwen, C. Bettembourg, B. Biton, J. Czech, H. De Foucauld, M. Didier, T. Licher, V. Mikol, A. Pommereau, F. Puech, V. Yaligara, A. Edwards, B. J. Bongers, L. H. Heitman, A. P. IJzerman, H. J. Sijben, G. J. P. Van Westen, J. Grixti, D. B. Kell, F. Mughal, N. Swainston, M. Wright-Muelas, T. Bohstedt, N. Burgess-Brown, L. Carpenter, K. Dürr, J. Hansen, A. Scacioc, G. Banci, C. Colas, D. Digles, G. Ecker, B. Füzi, V. Gamsjäger, M. Grandits, R. Martini, F. Troger, P. Altermatt, C. Doucerain, F. Dürrenberger, V. Manolova, A.-L. Steck, H. Sundström, M. Wilhelm and C. M. Steppan, The RESOLUTE consortium: unlocking SLC transporters for drug discovery, Nat. Rev. Drug Discovery, 2020, 19, 429–430 CrossRef CAS PubMed.
- The UniProt Consortium, A. Bateman, M.-J. Martin, S. Orchard, M. Magrane, A. Adesina, S. Ahmad, E. H. Bowler-Barnett, H. Bye-A-Jee, D. Carpentier, P. Denny, J. Fan, P. Garmiri, L. J. D. C. Gonzales, A. Hussein, A. Ignatchenko, G. Insana, R. Ishtiaq, V. Joshi, D. Jyothi, S. Kandasaamy, A. Lock, A. Luciani, J. Luo, Y. Lussi, J. S. M. Marin, P. Raposo, D. L. Rice, R. Santos, E. Speretta, J. Stephenson, P. Totoo, N. Tyagi, N. Urakova, P. Vasudev, K. Warner, S. Wijerathne, C. W.-H. Yu, R. Zaru, A. J. Bridge, L. Aimo, G. Argoud-Puy, A. H. Auchincloss, K. B. Axelsen, P. Bansal, D. Baratin, T. M. Batista Neto, M.-C. Blatter, J. T. Bolleman, E. Boutet, L. Breuza, B. C. Gil, C. Casals-Casas, K. C. Echioukh, E. Coudert, B. Cuche, E. De Castro, A. Estreicher, M. L. Famiglietti, M. Feuermann, E. Gasteiger, P. Gaudet, S. Gehant, V. Gerritsen, A. Gos, N. Gruaz, C. Hulo, N. Hyka-Nouspikel, F. Jungo, A. Kerhornou, P. L. Mercier, D. Lieberherr, P. Masson, A. Morgat, S. Paesano, I. Pedruzzi, S. Pilbout, L. Pourcel, S. Poux, M. Pozzato, M. Pruess, N. Redaschi, C. Rivoire, C. J. A. Sigrist, K. Sonesson, S. Sundaram, A. Sveshnikova, C. H. Wu, C. N. Arighi, C. Chen, Y. Chen, H. Huang, K. Laiho, M. Lehvaslaiho, P. McGarvey, D. A. Natale, K. Ross, C. R. Vinayaka, Y. Wang and J. Zhang, UniProt: The Universal Protein Knowledgebase in 2025, Nucleic Acids Res., 2025, 53, D609–D617 Search PubMed.
- M. Davies, M. Nowotka, G. Papadatos, N. Dedman, A. Gaulton, F. Atkinson, L. Bellis and J. P. Overington, ChEMBL web services: streamlining access to drug discovery data and utilities, Nucleic Acids Res., 2015, 43, W612–W620 Search PubMed.
- R. Winter, F. Montanari, F. Noé and D.-A. Clevert, Learning continuous and data-driven molecular descriptors by translating equivalent chemical representations, Chem. Sci., 2019, 10, 1692–1701 Search PubMed.
- C. Steinbeck, C. Hoppe, S. Kuhn, M. Floris, R. Guha and E. Willighagen, Recent Developments of the Chemistry Development Kit (CDK) - An Open-Source Java Library for Chemo- and Bioinformatics, Curr. Pharm. Des., 2006, 12, 2111–2120 Search PubMed.
- D. Rogers and M. Hahn, Extended-Connectivity Fingerprints, J. Chem. Inf. Model., 2010, 50, 742–754 Search PubMed.
- T. Akiba, S. Sano, T. Yanase, T. Ohta and M. Koyama, Optuna: A Next-generation Hyperparameter Optimization Framework, Proc. 25th ACM SIGKDD Int. Conf. on Knowledge Discovery & Data Mining(KDD ’19), 2019, pp. 2623–2631 Search PubMed.
- F. Pedregosa, G. Varoquaux, A. Gramfort, V. Michel, B. Thirion, O. Grisel, M. Blondel, A. Müller, J. Nothman, G. Louppe, P. Prettenhofer, R. Weiss, V. Dubourg, J. Vanderplas, A. Passos, D. Cournapeau, M. Brucher, M. Perrot and É. Duchesnay, Scikit-learn: Machine Learning in Python, DOI:10.48550/ARXIV.1201.0490.
- D. S. Karlov, S. Sosnin, I. V. Tetko and M. V. Fedorov, Chemical space exploration guided by deep neural networks, RSC Adv., 2019, 9, 5151–5157 Search PubMed.
- L. McInnes, J. Healy, N. Saul and L. Großberger, Uniform manifold approximation and projection, J. Open Source Softw. , 2018, 3, 861 CrossRef.
- A. Rubinsteyn and S. Feldman, fancyimpute: An Imputation Library for Python (v0.0.16), 2016, DOI:10.5281/zenodo.594235.
| This journal is © The Royal Society of Chemistry 2026 |