Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Coupled fragment-based generative modeling with stochastic interpolants
Tuan
Le
 *a,
Yanfei
Guan
*a,
Yanfei
Guan
 b,
Djork-Arné
Clevert
b,
Djork-Arné
Clevert
 a and
Kristof T.
Schütt
a and
Kristof T.
Schütt
 a
a
aPfizer Research and Development, Machine Learning and Computational Sciences, Friedrichstraße 110, 10177 Berlin, Germany. E-mail: tuan.le@pfizer.com
bPfizer Research and Development, Medicine Design Computational Chemistry, 1 Portland St, Cambridge, MA 02139, USA
First published on 5th March 2026
Abstract
Fragment-based drug design (FBDD) has become a key approach in structure-based drug discovery, allowing researchers to systematically develop molecular fragments into potent ligands. Although recent generative AI models, such as diffusion-based approaches, show great potential for designing new molecules, applying them to fragment-based methods faces challenges due to mismatches between training and inference procedures, as well as computational limitations. In this work, we develop a generative model based on stochastic interpolants that unify diffusion and flow matching paradigms, learning to create fragments through conditional training on molecular substructures. Our experiments show that models trained with explicit fragment-based conditioning perform much better than unconditional models that are adapted for fragment completion tasks. We compare diffusion models with flow matching models using identical backbone architectures and find that flow matching delivers better convergence and produces higher-quality 3D molecular poses with reduced strain energies, all while needing fewer computational steps. We test our method on standard benchmark datasets and examine different fragmentation strategies, finding that the choice of fragmentation algorithm plays an important role in model performance. Through a detailed case study on an internal PLK3 inhibitor structure, we demonstrate that our approach can generate new fragments that show computationally favorable docking scores and binding energy estimates competitive with tested internal Pfizer compounds, while also exploring regions of chemical space that go beyond existing fragment libraries. These findings establish flow matching within the stochastic interpolants framework as a promising approach for fragment-based drug design, providing both improved computational efficiency and better molecular quality for structure-based optimization.
1 Introduction
Drug discovery is a risky resource-intensive process, with early-stage candidate selection posing a significant bottleneck due to the vastness of chemical space and the high cost of synthesis and testing.1,2 Structure-based drug design (SBDD) offers a powerful framework for rational ligand design by leveraging detailed structural information about protein targets.3,4 Among SBDD approaches, fragment-based drug design (FBDD) has emerged as a particularly effective strategy, wherein small, low-molecular-weight fragments are identified as initial binders and then elaborated into higher-affinity ligands through systematic chemical expansion.5–8 In this exercise, fragments with precise structural complementarity, including shape and explicit interactions, are designed by medicinal chemists through structure–activity relationship (SAR), chemical intuition or computational tools.Conventional computational methods employ abstracted ligand–protein interaction representatives, e.g. pharmacophores, to search against a pre-defined fragment library;9,10 or draw inspiration from historical lead optimization trajectories by building transformations between matched molecular pairs.11 Those computational methods might provide practical fragment replacement ideas though, they are confined in existing chemical space and the suggested fragments replacement could suffer from lack of novelty.
As generative artificial intelligence (AI) emerges as a powerful tool in structure-based drug design, models like Pocket2Mol,12 TargetDiff,13 PILOT14 and DiffBP,15 are aware of the pocket structure and able to generate small molecules with more favorable docking scores and physically plausible binding poses compared to reference ligands. However, in real-world drug design, molecules generated from scratch are usually intractable and challenging in synthesis. From an industrial perspective, those generative AIs should have more potential in FBDD by having a known essential part of the molecule fixed. This is widely applied in Hit/lead optimization, and can be readily validated through parallel medicinal chemistry (PMC).
Generative models as mentioned above trained on whole-molecule generation task (unconditional) can be applied to completing partial molecules given the constrained environment (conditional). Unconditionally trained diffusion models can perform this kind of conditional sampling, so-called inpainting, as demonstrated by RePaint16 for image generation. This has been applied to structure-based drug design in DiffSBDD.17 Crucially, to align the inference process to the training process, a fixed context for inpainting requires to be perturbed as well. This is because the unconditionally trained model has not seen clean context input during training, aside from the protein pocket. That discrepancy between training and sampling can lead to distorted local conformation of the generated fragments, or even distortions of the fixed molecular context that lead to an invalid molecule. We hypothesize that a model conditionally trained directly on the fragments set (for inpainting) will perform better at inference, if the training was also performed in such way.
Diffusion models typically need many sampling steps and carefully tuned noise schedules, which slows inference and introduces a training–sampling mismatch. The common hundreds of sampling steps makes long sampling time and thus limits the large-scale generations with limited computation resources. Flow matching18–20 instead learns a transport velocity along a chosen probability path, usually converges faster, and enables straighter, low-curvature transports that sample in far fewer steps with better numerical conditioning.
Flow matching (FM) has been applied to molecular generation on joint discrete–continuous graphs and in 3D, often outperforming diffusion in validity and sampling speed. Mixed continuous–categorical FM jointly generates atom types, bonds, and coordinates,21 harmonic self-conditioned FM improves multi-conformer pose and strain metrics,22 and SEMLA-Flow report faster convergence and higher ligand quality than diffusion.23 FLOWR24 extends SEMLA-Flow with a pocket encoder and shows that protein–ligand interactions present in the test set are recovered by generated compounds using their multi learning approach promoting PL interactions as well as ligand inpainting. DrugFlow developed by Schneuing et al.25 includes an uncertainty measure in their generative model with the option to delete atoms through virtual nodes.
Other approaches condition ligand generation through shape constraints by incorporating 3D reference ligand representations into diffusion models. For instance, PoLiGenX26 employs latent representations of 3D reference ligands to guide the generation process. Similarly, SQUID27 represents molecular shapes as point clouds sampled from ligand surfaces, combining spatial information with chemical elements through an encoder–decoder architecture. This approach has been further extended in ShEPhERD, which incorporates additional molecular features including electrostatics and pharmacophore vectors to enhance shape-guided generation.28
Fragment-based diffusion models generate or replace substructures while keeping a scaffold or linker fixed. DiffLinker builds 3D linkers between two fragments with an E(3)-equivariant denoiser and recovers realistic linkers over graph baselines.29 Ghorbani et al.30 and Xie et al.31 formulate scaffold decoration as conditional inpainting over atoms and 3D coordinates: variable substructures are masked, attachment points and local geometry are encoded, and an E(3)-equivariant denoiser regrows fragments, yielding higher validity, better substructure retention, and improved docking/pose metrics.
In the present work, we develop diffusion and flow matching models to explicitly learn fragment generation from fragment structures curated from CrossDocked2020 and Kinodata-3D dataset.32,33 The models are trained to generate both ending fragments and linkers. We compare such explicit conditional training/sampling strategy with the “inpainting”/RePaint sampling mode of unconditionally trained models.16 Impact of different fragmentation methods on the model performance are also benchmarked and discussed. Finally we demonstrate the developed fragment generation model on a newly disclosed PLK3 protein target. Generated molecules are compared to real-world ligands as well as molecules generated using conventional library-based fragment replacement method with respect to binding affinities. The benchmarking and comparison suggests that a flow matching models that has been explicitly trained on fragment structures outperforms others and is capable of generating low-strain and high-potency molecules.
1.1 Problem formulation
We address the problem of de novo 3D molecular generation for structure-based drug design (SBDD), where the structural context is defined by a protein binding pocket and potentially additional molecular fragments. Formally, we represent a protein pocket as P = (Hp, Xp), where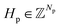 and
and 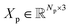 denote the atomic numbers and Cartesian coordinates of all Np atoms within the binding pocket, respectively. Similarly, we represent the ligand as M = (Hl, Xl, El), where Hl and Xl follow the same convention as the protein representation, and
denote the atomic numbers and Cartesian coordinates of all Np atoms within the binding pocket, respectively. Similarly, we represent the ligand as M = (Hl, Xl, El), where Hl and Xl follow the same convention as the protein representation, and  is an adjacency matrix encoding the molecular connectivity and bond types. In practice, we augment the atomic representations with additional chemical features such as formal charges and hybridization states to enhance molecular specificity.34 The objective of our de novo molecular generation framework is to sample complete molecules M ∼ pθ(M|P) conditioned on the protein pocket P. Additionally, our framework supports fragment-based conditional generation pθ(MF|MS, P), where molecular fragments are generated while keeping a scaffold MS fixed, enabling effective exploration of chemical space for both de novo design and lead optimization scenarios.
is an adjacency matrix encoding the molecular connectivity and bond types. In practice, we augment the atomic representations with additional chemical features such as formal charges and hybridization states to enhance molecular specificity.34 The objective of our de novo molecular generation framework is to sample complete molecules M ∼ pθ(M|P) conditioned on the protein pocket P. Additionally, our framework supports fragment-based conditional generation pθ(MF|MS, P), where molecular fragments are generated while keeping a scaffold MS fixed, enabling effective exploration of chemical space for both de novo design and lead optimization scenarios.
Molecular fragmentation through strategic bond cleavage enables the decomposition of a molecule M into multiple constituent fragments (Fig. 1), making fragment-based drug design a promising alternative to de novo molecular generation. This approach is particularly valuable during lead optimization, where specific molecular regions are systematically modified to enhance desired pharmacological properties while preserving favorable structural motifs. Scaffold decoration represents a key lead optimization strategy that maintains a core molecular framework while introducing targeted modifications through R-group substitutions. This methodology reduces the dimensionality of the design space by constraining the search to specific molecular regions, thereby limiting the solution space for fragment extensions and typically requiring the generation of smaller molecular components.
To obtain molecular partitions, we employ established fragmentation algorithms such as RECAP35 or BRICS,36 which systematically decompose molecules into collections of scaffolds and R-groups. Following fragmentation, a molecule can be represented as M = (Hl,i,Xl,i,El,i)i=1Nm, where Nm denotes the number of fragments obtained through bond cleavage. The specific fragmentation pattern depends on the chosen decomposition rules, with each molecule potentially yielding multiple R-groups and scaffolds. Fragment-based 3D generative models aim in generating molecular fragments MF ∼ pθ(MF|MS, P) with the constraint to complete the molecule M = (MS, MF), while keeping MS and P fixed as condition.
2 Experiments and results
2.1 Datasets
We evaluate our approach using the CrossDocked2020 benchmark dataset,32 employing the processed version containing 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 protein–ligand (PL) complexes as used in prior studies.12,37 The dataset was partitioned into training and test sets using MMseqs2 (ref. 38) with a 30% sequence identity threshold, yielding 100 distinct test targets. Despite its widespread adoption in the machine learning community, the processed CrossDocked2020 dataset exhibits limited chemical diversity. Specifically, the 100
000 protein–ligand (PL) complexes as used in prior studies.12,37 The dataset was partitioned into training and test sets using MMseqs2 (ref. 38) with a 30% sequence identity threshold, yielding 100 distinct test targets. Despite its widespread adoption in the machine learning community, the processed CrossDocked2020 dataset exhibits limited chemical diversity. Specifically, the 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 PL complexes contain only approximately 10
000 PL complexes contain only approximately 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 600 unique SMILES strings, indicating a high degree of cross-docking redundancy where identical ligands are paired with multiple protein targets.
600 unique SMILES strings, indicating a high degree of cross-docking redundancy where identical ligands are paired with multiple protein targets.
To learn on a broader chemical space with more drug-like compounds, we leverage the Kinodata-3D dataset compiled by Backenköhler et al.33 Kinodata-3D is a collection of kinase complexes processed in silico using cross-docking data with POSIT template docking, more closely resembling common practices in drug discovery applications. We use 104![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 850 pocket–ligand complexes for training, with 310 and 136 complexes for validation and testing, respectively, allocated through random splitting. The chemical space coverage in Kinodata-3D is substantially larger, including approximately 73
850 pocket–ligand complexes for training, with 310 and 136 complexes for validation and testing, respectively, allocated through random splitting. The chemical space coverage in Kinodata-3D is substantially larger, including approximately 73![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 560 unique compounds. Additional details are provided in the SI Section B.
560 unique compounds. Additional details are provided in the SI Section B.
2.2 Model architecture
We utilize the EQGAT message-passing neural network,39 originally employed in EQGAT-Diff34 for small molecule generation and subsequently modified in PILOT14 for target-aware ligand design. PILOT generates molecules conditioned on protein pockets using diffusion models in both unconditional and property-guided contexts.The PILOT network architecture processes ligand and protein pocket data as a full-atom heterogeneous point cloud graph. In this representation, ligand atoms are fully connected through edges, while ligand–pocket and pocket–pocket edges are constructed using a radius cutoff of 5 Å. Message-passing is performed over L = 12 layers propagating invariant scalar and equivariant vector features, and the model contains 12.4M parameters.
We train two diffusion models on the Kinodata-3D dataset33 under unconditional and conditional settings. In the conditional setting, the model is trained on randomly masked molecular fragments obtained through RECAP and BRICS. These fragments, together with the protein pocket, serve as fixed context during training. For inpainting with the unconditional model, we apply the RePaint procedure16 with r = 1 resampling steps and the same noise schedule used during training.
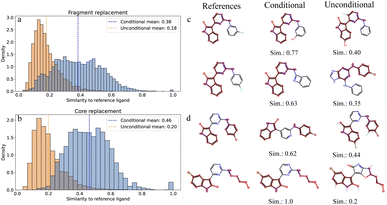
We evaluate both models on fragment replacement and core replacement tasks. In fragment replacement, a single variable fragment is attached to a given molecular context, while core replacement involves placing a molecular linker between two fixed molecular components—a generally more challenging task. For both inpainting tasks, we observe that the unconditional diffusion model achieves significantly lower molecular validity (24.75%) compared to the conditional diffusion model (87.27%). This validity gap at r = 1 is consistent with findings from Schneuing et al.,17 who reported similar validity rates (20–40%) for RePaint with minimal resampling, requiring r = 10 to achieve 60–80% validity (see Section S5.6 and Fig. 2 in their work). While higher resampling steps would likely improve validity, this comes with a proportional increase in computational cost.
As shown in the Fig. 2a and b, ligands generated by the unconditional model share lower 2D chemical similarity to reference molecules compared to those generated by the conditional model, when using binary ECFPs of length 4096 and radius 2. This dissimilarity primarily arises from the unconditional model's inability to place new atoms to the variable region near the anchor point(s), whereas the conditional model successfully achieves this localization through explicit anchor point information provided to the network. Representative examples of misplaced atoms by the unconditional model are illustrated in the third column of Fig. 2c and d. In the fragment replacement study, the unconditional model incorrectly places a fluorine atom on the pyrimidine ring (first row) and fuses an imidazole to the pyrimidine (second row), despite the requirement that new fragments should only connect to the nitrogen atom (as shown in the reference ligand). In contrast, the conditional model (second column) correctly places atoms as discrete new fragments without modifying the fixed molecular context in both inpainting tasks. While generated molecules pass validity checks, some may contain strained motifs such as the fused azete ring in Fig. 2c (conditional, second row). Post-generation filtering using synthetic accessibility scores or medicinal chemistry review is recommended in practice.
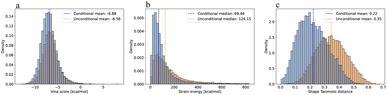
Quantitatively, the conditional model preserves the fixed substructure in 97.02% and 97.52% among the valid molecules for fragment and core replacement studies, respectively, while the unconditional model maintains substructural integrity in only 84.56% and 85.24% of valid molecules. The conditional model also generates 3D poses that are more favorable in terms of lower Vina pose scores and ligand strain energies (Fig. 3b). The conditional model achieves a median strain energy of 69.44 kcal mol−1 compared to 124.15 kcal mol−1 for the unconditional model, while maintaining a higher PoseBusters (PB) validity rate40 of 79.29% versus 68.89%, indicating more physically plausible poses. Strain energies are computed following Harris et al.41 as the difference between the internal energy of a relaxed pose and the generated pose. Both relaxation and energy calculations are performed using the Universal Force Field (UFF).42 Since newly generated fragments and cores are correctly attached to anchor points with the conditional model, we also observe reduced shape Tanimoto distances as indicated in the last panel in Fig. 3c. Therefore, we conclude that conditionally trained models on masked fragments are better suited for inpainting tasks compared to unconditional models using RePaint. Shape Tanimoto distances are computed using RDKit's 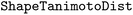 function, which evaluates geometric shape overlap on a grid. Unlike OpenEye's ROCS,43 which incorporates pharmacophore features (e.g., hydrogen bond donors/acceptors, hydrophobic regions) and electrostatic similarity, this implementation considers only molecular shape.
function, which evaluates geometric shape overlap on a grid. Unlike OpenEye's ROCS,43 which incorporates pharmacophore features (e.g., hydrogen bond donors/acceptors, hydrophobic regions) and electrostatic similarity, this implementation considers only molecular shape.
While the conditional model demonstrates superior performance in inpainting tasks, we hypothesize that further improvements are possible, particularly for 3D pose generation. Recent advances in generative modeling have leveraged the Flow Matching framework, which shares close connections with diffusion models under the unified umbrella of Stochastic Interpolants.44
We perform unconditional ligand generation on 136 test targets from the Kinodata-3D benchmark set and observe that samples generated by the flow model achieve superior performance across multiple metrics, including 2D molecular validity and 3D pose quality, compared to the diffusion model (Table 1). Higher molecular validity indicates correct alignment between atomic chemical valency and bond topology with bonded neighbors.
| Model/set | Number of atoms | Validity [%] ↑ | 3D pose | ||||
|---|---|---|---|---|---|---|---|
| Molecule | Connected components | Clashes ↓ | Vina score [kcal mol−1] ↓ | Strain energy [kcal mol−1] ↓ | PB validity [%] ↑ | ||
| Training | 31.24 ± 6.37 | 100 | 100 | 4.88 ± 9.26 | −7.62 ± 4.95 | 5.62 ± 1.81 | 97.35 |
| Test | 26.8 ± 5.1 | 100 | 100 | 3.32 ± 3.34 | −7.45 ± 1.26 | 13.84 ± 5.74 | 100 |
| Diffusion | 26.26 ± 5.97 | 89.82 ± 5.46 | 97.91 ± 2.58 | 6.64 ± 3.53 | −6.69 ± 1.78 | 45.95 ± 27.56 | 87.57 ± 8.05 |
| Flow | 26.34 ± 5.99 | 99.04 ± 1.18 | 99.95 ± 0.22 | 3.06 ± 1.81 | −8.27 ± 1.44 | 13.99 ± 8.95 | 97.90 ± 3.07 |
The flow model demonstrates significant computational advantages, requiring only 100 integration steps compared to 500 for the diffusion model, resulting in 5× faster inference speed. While efficient diffusion samplers like DDIM45 exist for continuous variables (coordinates), no such acceleration is available for discrete variables (atom types, bonds), which follow continuous-time Markov chain.46 Flow matching, in contrast, naturally supports fewer steps for both continuous and discrete components.47
Despite this efficiency gain, ligands generated by the flow model achieve superior Vina (pose evaluation) scores of −8.27 kcal mol−1 compared to −6.69 kcal mol−1 for the diffusion model. The improved pose quality is further evidenced by lower strain energies and higher PoseBusters40 validity (97.90% vs. 87.57%), which evaluates the physical plausibility of ligand poses within protein pockets.
Ligands generated by the diffusion model reveal more rotatable bonds but fewer rings compared to those in the Kinodata-3D training set, as illustrated in Fig. 4. On average, the diffusion model produces ligands with 6.7 rotatable bonds and 3.6 rings, whereas the flow model generates ligands with fewer rotatable bonds (4.5) but more rings (4.3). Despite these structural differences, ligand sizes remain consistent between the two generation approaches (Table 1, first column).
The increased occurrence of rings in ligands generated by the flow model likely stems from the deterministic ODE integration process that starts from the prior distribution. The flow matching model demonstrates superior learning of ring and rotatable bond statistics, achieving Jensen–Shannon (JS) divergences of 0.1464 and 0.0524 for rotatable bonds and rings, respectively, when compared to the training set. In contrast, ligands from the diffusion model show higher JS divergences of 0.1817 and 0.2567 for the same metrics. We list quantitative divergence metrics for various 2d descriptors in the SI Section D.1.
While the comparison between diffusion and flow matching models focuses on de novo ligand generation conditioned solely on the protein pocket, we demonstrate in the SI Section D.3 that PILOT-Flow also achieves superior evaluation metrics compared to PILOT-Diffusion for conditional inpainting tasks involving core and fragment replacement.
Different fragmentation algorithms yield varying numbers of unique fragments within a given dataset. As detailed in SI Section B, the custom cuttable fragmentation algorithm produces the largest number of unique fragments on both CrossDocked2020 and Kinodata-3D datasets. This increased fragment diversity means that, given a fixed training epoch budget, the generative model trained with custom cuttable algorithm fragments has greater exposure to fragment variety compared to models trained on BRICS or RECAP fragments.
In both fragment replacement and core replacement tasks, we observe that generative models trained on BRICS algorithm partitions tend to sample ligands with unfavorable poses, characterized by higher strain energies and reduced shape similarity to reference ligands (Fig. 5). The core replacement task proves more challenging than fragment replacement, as generating a core that connects multiple molecular arms constrains the feasible conformational space. This results in higher strain energies across all three fragmentation approaches (Fig. 5a and third column in Table 2). Notably, since all conditional models employ flow matching, they achieve higher PoseBusters validity rates compared to the conditional diffusion model reported in Section 2.2.1, supporting the conclusions from Section 2.2.2.
| Task | Fragmentation | Vina score ↓ [kcal mol−1] | Efficiency ↑ | Strain energy ↓ [kcal mol−1] | Shape Tanimoto distance ↓ | Clashes ↓ | PB validity ↑ |
|---|---|---|---|---|---|---|---|
| Core | Brics | −6.93 ± 0.92 | 0.35 ± 0.38 | 84.38 ± 49.14 | 0.16 ± 0.05 | 8.74 ± 3.68 | 0.90 ± 0.21 |
| Cuttable | −7.16 ± 0.98 | 0.49 ± 0.41 | 59.54 ± 38.90 | 0.13 ± 0.06 | 8.21 ± 3.63 | 0.89 ± 0.21 | |
| Recap | −7.12 ± 0.98 | 0.48 ± 0.40 | 62.94 ± 39.51 | 0.13 ± 0.06 | 8.08 ± 3.62 | 0.89 ± 0.20 | |
| Fragment | Brics | −7.73 ± 0.76 | 0.66 ± 0.38 | 42.07 ± 28.68 | 0.22 ± 0.05 | 6.37 ± 2.62 | 0.97 ± 0.11 |
| Cuttable | −7.80 ± 0.80 | 0.70 ± 0.38 | 35.40 ± 24.05 | 0.21 ± 0.05 | 6.02 ± 2.53 | 0.97 ± 0.11 | |
| Recap | −7.78 ± 0.77 | 0.68 ± 0.37 | 37.75 ± 25.38 | 0.21 ± 0.05 | 6.17 ± 2.62 | 0.97 ± 0.11 |
Since the custom cuttable algorithm returns more variable fragments as opposed to RECAP (see SI Section B) and the analysis favors the generative model trained via the custom fragmentation algorithm, we decide to use that model for further analysis and comparison to traditional fragment based replacement tools.
3 Fragment replacement on PLK3 target
To further assess the proposed model on unseen targets, we performed a retrospective in silico study on the PLK3 inhibitor project from Pfizer/Postera, comparing generated ligands against approximately 100 previously synthesized and experimentally tested compounds. PLK3 is one of the Polo-like kinases (PLKs) family and has a significantly role in DNA replication and is hypothesized to have a further role in mitotic progression. In this project, the team has discovered a potent lead compound, PF-07976265 (see Fig. 6a), and a subseries with about 100 compounds which carry different left-hand side R groups as shown in the box, while preserving the right-hand side structure. The binding structure of PF-07976265 in the PLK3 pocket is disclosed for the first time in the present work and the structure is depicted in Fig. 6b. PF-07976265 binds in the ATP-binding pocket, with the right-hand part siting along the p-loop and aminopyrimidine core serving as the H-bond donor towards the hinge region. The pocket for left-hand part (as highlighted by the dash box) features a couple of polar residues that could interact with the ligand. For example, in Fig. 6b, the cyclic amine forms an ionic bonding interaction with Glu78.These tested ligands, which vary in their left-hand side substituents, serve as the benchmark for evaluating ligands generated through PILOT-Flow. Structure-based GenChem AI literature usually relies on docking scores to assess generated ligands. To better capture the pocket flexibility, ligand strain energy, as well as solvation terms, we computed MM/GBSA binding energies with Prime-MMGBSA of Schrodinger suite.49 The MM/GBSA binding energy for tested ligands achieves moderate correlation with experimental potency as shown in Fig. 6a. Given the balance between accuracy and speed, herein we evaluate generated ligands with the MM/GBSA binding energy (dG) as a more reliable metrics than docking score.
In our study, we compare PILOT-Flow against BROOD, a fragment replacement and scaffold hopping tool developed by OpenEye50 which is frequently used in industry for Hit or Lead optimization. Brood enumerates fragments from a chosen library to replace selected atoms in a reference molecule. During fragment enumeration, BROOD attempts to match both the shape and electrostatic properties (termed “colors”) of the query fragment, and try to avoid clashes with protein. These color descriptors represent features such as hydrogen bond donors/acceptors, hydrophobic regions, and other pharmacophores. In this study, fragments are extracted from ChEMBL database51 version 22.
We generated 5000 ligands with PILOT-Flow and BROOD, respectively. Table 3 shows statistics from the sets obtained by BROOD and PILOT-Flow, indicating that PILOT-Flow yields slightly larger ligands on average compared to BROOD (37.22 vs. 36.30 heavy atoms) with slightly better drug-likeness (QED: 0.38 vs. 0.33) but marginally lower synthetic accessibility scores (SA: 3.56 for PILOT-Flow vs. 3.98 for BROOD). Since BROOD aims to maintain shape and electrostatic complementarity, the replaced fragments predominantly retain similar sizes, as indicated by the lower standard deviation in heavy atom counts.
| # HA | Rot. | QED ↑ | SA ↑ | PL dG ↓ | Strain ↓ | Vina ↓ | |
|---|---|---|---|---|---|---|---|
| a HA = heavy atoms, Rot. = rotatable bonds, dG/strain/Vina in kcal mol−1. | |||||||
| BROOD | 36.3 ± 0.7 | 7.3 ± 1.6 | 0.33 ± 0.06 | 3.98 ± 0.40 | −74.3 ± 5.0 | 5.2 ± 2.6 | −11.9 ± 1.0 |
| PILOT | 37.2 ± 3.5 | 5.3 ± 1.0 | 0.38 ± 0.10 | 3.56 ± 0.55 | −74.9 ± 7.0 | 5.3 ± 3.5 | −13.1 ± 1.1 |
| Tested | 33.3 ± 2.6 | 5.2 ± 1.3 | 0.44 ± 0.09 | 3.06 ± 0.27 | −76.0 ± 4.8 | 3.7 ± 1.5 | — |
We observe that samples from PILOT-Flow achieve lower Vina scores compared to BROOD (see last column in Table 3). While Vina scores evaluate generated ligand poses without re-docking in static protein pockets, they do not account for required interactions and ligand stability in the protein environment by default. To assess a more realistic scenario, we employ MM-GBSA protein–ligand energy minimization to compute ligand strain and binding energies (dG).
Ligand strain energy represents a conformational penalty for adopting the binding-competent conformation, potentially deviating from the most stable isolated conformation. With flexible atoms from both pocket (within a cutoff of 4 Å) and ligand in the MM-GBSA calculations, we observe that after minimization, samples from BROOD and PILOT-Flow reveal similar ligand strain energies (5.19 vs. 5.26 kcal mol−1) and protein–ligand binding energies (−74.29 vs. −74.92 kcal mol−1) with PILOT-Flow demonstrating a longer, more favorable tail toward lower energies. Given the limited explanatory power of MM-GBSA energies for experimental potency (R2 = 0.279, Fig. 6a), we do not interpret these similar values as evidence of equivalent binding affinity. Instead, we use MM-GBSA primarily as a computational filter to prioritize candidates for structural analysis of protein–ligand interactions.
Statistics for synthesized and tested ligands are listed in Table 3 as well. Tested ligands have a lower mean binding energy dG than generated ligands (PL dG: −75.96 for tested vs. −74.92 for PILOT-Flow and −74.29 for BROOD). Note that the statistics shown in Table 3 are for the whole generated ligands set (5000 for each of BROOD and PILOT-Flow). In practice, those 5000 generated ligands are used as the initial pool of candidates from which computational and medicinal chemists will select top-K ligands for further assessment.
We examine top-K generated ligands and compare those with the tested compounds. If the ligand generation method could render a score or ranking order, we can naturally use those to select top-K suggestions. Currently, PILOT-Flow, as well as many other GenChem AI methods, is not yet able to provide a reliable score for generated structures that could correlate with the binding strength. On the other hand, BROOD provides a score only based on the color/shape similarity against reference ligand. In a similar vein, we select the top-K ligands by Vina docking score. Binding energy distributions for top-K PILOT-Flow and BROOD ligands are plotted in Fig. 7e. We plot the distribution for all tested ligands along each top-K selection for comparison. It shows that starting from top-2000, PILOT-Flow ligands tend to have a stronger computed binding energy than tested compounds, especially the first quartile Q1 which is significantly lower than BROOD and Tested. This trend is more pronounced as K gets smaller. While these computed energies should be interpreted cautiously, this suggests that PILOT-Flow, when combined with appropriate scoring functions, can effectively prioritize candidates for further evaluation.
Beyond computed binding energies, a more reliable indicator of potential binding is the presence of key interactions with specific pocket residues. The reference ligand PF-07976265 interacts with Glu78 through ionic bonding from the piperidine amine. We observe that 398 ligands from PILOT-Flow and 1734 ligands from BROOD maintain that type of interaction towards Glu78. Distributions of binding energies (dG) vs. ligand strain for the top-300 ligands that preserve this interaction are shown in Fig. 8a. PILOT-Flow and BROOD exhibit similar medians for both binding and strain though PILOT-Flow has a longer tail towards stronger binding. We show four examples from the strong binding region (marked by stars in Fig. 8a) in Fig. 8b. All four examples feature a cyclic amine in the generated fragment that engages ionic bonding interactions towards Glu78, and a relatively rigid conjugated linker connecting to the core to stabilize the binding pose (Fig. 8c and d).
In addition to Glu78, other polar residues in the pocket could also be critical to binding, e.g., Lys145 and Arg66. PILOT-Flow generates ligands picking up interactions with those residues as well. Fig. 9a shows an example where the generated lactam serves as a H-bond acceptor to interact with Lys145. In another example, the generated cyclic sulfone moiety interacts with Arg66, and the second oxygen atom of sulfone forms a bidentate H-bond acceptor together with the lactam carbonyl group to interact with Lys145 (Fig. 9b). Both examples achieve superior computed binding energies (−90.6 and −89.2 kcal mol−1). These examples indicate that PILOT-Flow is able to discover new types of interactions through its understanding of the underlying pocket structure. This ability to identify chemically diverse fragments that engage meaningful binding interactions-rather than simply optimizing computed binding energies-represents the key practical value of the generative approach.
4 Methods
4.1 Fragmentation algorithm
We implement a custom fragmentation approach using SMARTS pattern matching to cut specific bond types. This method targets bonds that standard approaches like RECAP and BRICS typically avoid, such as amide bonds, providing more comprehensive fragmentation options.The algorithm works in two modes. Single cuts break one bond per fragmentation, creating terminal fragments suitable for scaffold decoration. Double cuts simultaneously break two bonds, extracting central fragments or linkers between cut sites. We exclude ring bonds to preserve cyclic structures.
We use the SMARTS pattern 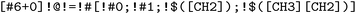 and systematically identify acyclic single bonds involving carbon atoms, including amide bonds. In single-cut mode, we cleave these bonds individually to generate terminal fragments containing either the carbonyl or amine portion. In double-cut mode, we simultaneously cut two such bonds within the same molecule, effectively isolating middle segments that may contain amide groups or serve as linkers between amide-containing regions.
and systematically identify acyclic single bonds involving carbon atoms, including amide bonds. In single-cut mode, we cleave these bonds individually to generate terminal fragments containing either the carbonyl or amine portion. In double-cut mode, we simultaneously cut two such bonds within the same molecule, effectively isolating middle segments that may contain amide groups or serve as linkers between amide-containing regions.
4.2 Stochastic interpolants
Generative models based on neural transport methods have demonstrated exceptional performance across image and text generation tasks.52–58 These approaches share a common framework: using ordinary or stochastic differential equations (ODEs/SDEs) to continuously transform samples from a simple base distribution p0 (e.g., random noise) to a complex target distribution p1 (e.g., molecular data).The seminal works of Lipman et al.,20 Albergo et al.,44 Liu et al.59 introduced novel simulation-free training objectives for continuous normalizing flows.60 These methods construct conditional probability paths connecting samples from base and target distributions, thereby circumventing the computationally expensive maximum-likelihood training over ODE trajectories. Building on this foundation, Albergo and Vanden-Eijnden19 developed the stochastic interpolants framework, which provides a unified theoretical approach encompassing both deterministic flow matching and stochastic diffusion processes.
For continuous molecular data 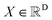 (such as atomic coordinates), we define the stochastic interpolant process It as:
(such as atomic coordinates), we define the stochastic interpolant process It as:
| It(X0,X1) = αtX0 + βtX1 + γtZ, t ∈ [0,1], | (1) |
Different choices of these coefficient functions lead to different generative models. Linear interpolation (αt = 1 − t, βt = t) yields the shortest path between source and target, as used in rectified flows.59 Adding noise with 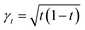 recovers the Brownian Bridge process.61
recovers the Brownian Bridge process.61
For molecular generation, we employ the cosine scheduler54 with βt = cos(0.5π(1 − t)ν) and αt = 1.0 − βt, which has proven effective for both diffusion14,62 and flow matching models21 in chemistry. This scheduler preserves more information from the target molecular data during training.
The resulting conditional probability path is Gaussian:
| pt(Xt|X0,X1,Z) = N(Xt;µt = αtX0 + βtX1,Σt = γt2I). | (2) |
For discrete molecular data X = (Xi)i=1N such as atom types and bond types, where each element Xi ∈ {1, …, D} represents one of D possible categories, we use a discrete analogue. Following recent advances in discrete flow matching,58,63 we employ continuous-time Markov chains where tokens transition between discrete states over time.
The discrete conditional probability path takes the simpler form:
| pt(Xt|X0,X1) = (1 − t)δX0 + tδX1, | (3) |
Data-dependent couplings67 go further by factorizing the coupling as p0,1(X0, X1) = p1(X1)p0(X0|X1), where the source distribution is conditioned on the target. This provides more information about the starting point X0 by leveraging knowledge of the target molecular structure X1. For fragment-based molecular design, this translates to conditioning the generation process on fixed molecular scaffolds while varying specific regions or subgraphs.
For molecular coordinates, data-dependent couplings offer particular advantages since chemically informed priors can reduce transport costs during training. Examples include centering variable molecular fragments using their center-of-mass with Gaussian noise, or preserving bond connectivity while sampling conformations.22,68
In our fragment-based approach, we decompose a ligand molecule into Nm subgraphs M = {Mi}i=1Nm and select one subgraph Mv as the variable region to be generated, e.g. see Fig. 1b and c. The model learns the conditional distribution pθ(Mv|![[M with combining tilde]](https://www.rsc.org/images/entities/i_char_004d_0303.gif) v,P), where
v,P), where ![[M with combining tilde]](https://www.rsc.org/images/entities/i_char_004d_0303.gif) v represents the fixed molecular scaffold and P is the protein pocket. Computationally, this is implemented using node and edge masks that fix the scaffold partitions
v represents the fixed molecular scaffold and P is the protein pocket. Computationally, this is implemented using node and edge masks that fix the scaffold partitions ![[M with combining tilde]](https://www.rsc.org/images/entities/i_char_004d_0303.gif) v while allowing generation of the variable fragment Mv. This conditional approach is fundamentally different from unconditional models pθ(M|P) that generate entire molecules from scratch, which face a much larger and more challenging search space.
v while allowing generation of the variable fragment Mv. This conditional approach is fundamentally different from unconditional models pθ(M|P) that generate entire molecules from scratch, which face a much larger and more challenging search space.
The denoising training objective predicts the ground-truth molecule M1 from perturbed molecular data Mt = (Ht, Xt, Et), where Ht, Xt, and Et represent noisy atom types, coordinates, and bond types at time t. The model conditions on the fixed protein pocket P and any remaining fixed molecular scaffold ![[M with combining tilde]](https://www.rsc.org/images/entities/i_char_004d_0303.gif) . The fragment-based loss function is:
. The fragment-based loss function is:
Lt = w(t) × ld(M0,pθ(Mt,t|P,![[M with combining tilde]](https://www.rsc.org/images/entities/i_char_004d_0303.gif) )), )), | (4) |
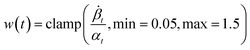 provides time-dependent loss weighting following Cremer et al.14. For continuous coordinates, we use mean-squared error loss ld, while discrete variables (atom types, hybridization states, bond types) employ cross-entropy loss.
provides time-dependent loss weighting following Cremer et al.14. For continuous coordinates, we use mean-squared error loss ld, while discrete variables (atom types, hybridization states, bond types) employ cross-entropy loss.
4.2.2.1 Masking mechanism for fragment-based training. The PILOT backbone architecture uses message-passing to update atomic coordinates. To control which atoms undergo modification, we provide a variable fragment mask as input that specifies which nodes are subject to updates during processing. This ensures context atoms maintain fixed coordinates throughout message-passing, similar to the treatment of the fixed protein pocket.
Formally, given the fixed molecular context ![[M with combining tilde]](https://www.rsc.org/images/entities/i_char_004d_0303.gif) , we construct a binary matrix F ∈ {0,1}NM×2 where NM is the total number of molecular atoms. The first column identifies fixed scaffold atoms, while the second column designates anchor/attachment points for fragment connections. For complete de novo generation, we set
, we construct a binary matrix F ∈ {0,1}NM×2 where NM is the total number of molecular atoms. The first column identifies fixed scaffold atoms, while the second column designates anchor/attachment points for fragment connections. For complete de novo generation, we set  , indicating no fixed constraints.
, indicating no fixed constraints.
4.2.2.2 Training details. We train all models from scratch for 300 epochs on CrossDocked2020 and 200 epochs on Kinodata-3D using the AdamW optimizer with AMSGrad. Hyperparameters included learning rate 2 × 10−4, weight decay 10−12, and gradient clipping for values exceeding 10. Complete training and sampling algorithms are provided in SI Section C.2.
5 Conclusion
We have presented a comprehensive framework for fragment-based drug design using stochastic interpolants that unifies diffusion and flow matching approaches for conditional molecular generation. Our work demonstrates several key advances in structure-based drug design through explicit fragment-based training. In particular, we have shown that models trained with explicit conditional fragment masking significantly outperform unconditional models adapted for inpainting tasks. The conditional approach achieves higher molecular validity (87.27% vs. 24.75%), better preservation of fixed substructures (97% vs. 85%), and generates more physically plausible poses with lower strain energies. This finding challenges the common practice of applying unconditional models to conditional tasks and highlights the importance of training-inference alignment in generative molecular design.We have found that flow matching consistently outperforms diffusion models across multiple evaluation metrics while requiring 5× fewer sampling steps. Flow matching achieves superior molecular validity (99.04% vs. 89.82%), better geometric accuracy, and generates molecules with more favorable binding energies and lower strain. The deterministic ODE integration in flow matching produces more reliable molecular poses compared to the stochastic sampling in diffusion models, making it particularly suitable for structure-based applications where pose quality is critical.
Our analysis further shows that the choice of fragmentation algorithm significantly impacts model performance. The custom cuttable fragmentation algorithm uses SMARTS pattern matching to target specific bond types, including those typically preserved by standard methods. This produces more diverse fragment libraries compared to the reaction-based rules of RECAP and BRICS. Models trained on cuttable fragments consistently generate ligands with lower strain energies and better shape similarity to reference molecules.
Finally, the PLK3 case study validates the practical applicability of our approach on an industrially relevant target not present in training data. PILOT-Flow generates fragments with binding energies comparable to experimentally tested compounds while exploring novel chemical space beyond traditional fragment libraries. The model maintains essential protein–ligand interactions while discovering new chemical scaffolds, demonstrating its potential for real-world drug discovery applications.
Our current approach has three key limitations that future work could address. First, we rely on MM-GBSA and Vina docking scores, which have known limitations in accurately predicting experimental binding affinities. Second, our conditional sampling approach generates higher ligand strain energies when fixing substructures, as the model must accommodate potentially suboptimal fixed fragments. Third, our evaluation is limited to kinase targets. Validation on diverse protein families would be valuable to assess broader applicability beyond this study's methodological focus. Future work could develop more sophisticated scoring functions that incorporate machine learning-based binding affinity predictors, protein–ligand interaction fingerprints, and physics-based free energy calculations to better guide molecular generation. Additionally, improved conditioning strategies could allow for minor adjustments to fixed regions, or employ strain-aware loss functions that explicitly penalize high-energy conformations during training. Furthermore, incorporating synthetic accessibility considerations by biasing generation toward known building blocks and established reaction pathways could ensure that generated fragments and molecules are readily synthesizable, bridging the gap between computational design and experimental implementation.
Author contributions
This work was conceptualized jointly by T. L. and Y. G., who also conducted the formal analysis. T. L. was responsible for data curation, led the investigation, developed the model methodology, and created the visualizations. Y. G. developed the fragmentation methodology and contributed to the visualization efforts. D. A. C. provided critical resources that enabled this research. The study was supervised by K. T. S., who also contributed to writing the original draft together with T. L. and Y. G. All authors participated in the proofreading process.Conflicts of interest
There are no conflicts of interest to declare.Data availability
We provide the open-source code for training and inference under the repository https://github.com/pfizer-opensource/pilot-sbdd.The publicly available raw and processed data from CrossDocked2020 and Kinodata-3D can be downloaded under the link https://doi.org/10.6084/m9.figshare.30739232, while checkpoints for trained diffusion and flow models can be downloaded under https://doi.org/10.6084/m9.figshare.30739268.
Supplementary information (SI): the stochastic interpolants framework, training and sampling algorithms, comparative flow matching vs. diffusion analyses, and the PLK3 case study. See DOI: https://doi.org/10.1039/d5dd00535c.
Acknowledgements
We thank our colleagues at Pfizer for valuable discussions and feedback throughout this project. We are grateful to Shayne Wierbowski, Christopher McClendon, Julian Cremer, Dina Sharon, Ben Burke and Yang Joy for their discussions and suggestions that improved this work. We thank Gabrielle Lovett, Jayasankar Jasti, as well as other colleagues from Pfizer and Postera who contributed to the PLK3 project for the crystal structure and potency data. This research was supported by computational resources from Pfizer's internal HPC cluster. D. A. C. acknowledges the support from the European Union's Horizon 2020 research and innovation program under the Marie Skłodowska-Curie Actions grant agreement (AiChemist; 101120466).Notes and references
- S. M. Paul, D. S. Mytelka, C. T. Dunwiddie, C. C. Persinger, B. H. Munos, S. R. Lindborg and A. L. Schacht, Nat. Rev. Drug Discovery, 2010, 9, 203–214 CrossRef CAS PubMed.
- P. G. Polishchuk, T. I. Madzhidov and A. Varnek, J. Comput.-Aided Mol. Des., 2013, 27, 675–679 CrossRef CAS PubMed.
- A. C. Anderson, Chem. Biol., 2003, 10, 787–797 CrossRef CAS PubMed.
- E. Lionta, G. Spyrou, D. K. Vassilatis and Z. Cournia, Curr. Top. Med. Chem., 2014, 14, 1923–1938 CrossRef CAS PubMed.
- M. Congreve, R. Carr, C. Murray and H. Jhoti, Drug Discovery Today, 2003, 8, 876–877 CrossRef PubMed.
- R. J. Hall, P. N. Mortenson and C. W. Murray, Prog. Biophys. Mol. Biol., 2014, 116, 82–91 CrossRef CAS PubMed.
- M. Bon, A. Bilsland, J. Bower and K. McAulay, Mol. Oncol., 2022, 16, 3761–3777 CrossRef CAS PubMed.
- W. Xu and C. Kang, J. Med. Chem., 2025, 68, 5000–5004 CrossRef CAS PubMed.
- G. Hessler and K.-H. Baringhaus, Drug Discovery Today: Technol., 2010, 7, e263–e269 CrossRef CAS.
- R. Ferreira de Freitas, R. J. Harding, I. Franzoni, M. Ravichandran, M. K. Mann, H. Ouyang, M. Lautens, V. Santhakumar, C. H. Arrowsmith and M. Schapira, J. Med. Chem., 2018, 61, 4517–4527 CrossRef CAS PubMed.
- A. Dalke, J. Hert and C. Kramer, J. Chem. Inf. Model., 2018, 58, 902–910 CrossRef CAS PubMed.
- X. Peng, S. Luo, J. Guan, Q. Xie, J. Peng and J. Ma, Proceedings of the 39th International Conference on Machine Learning, 2022, pp. 17644–17655 Search PubMed.
- J. Guan, X. Zhou, Y. Yang, Y. Bao, J. Peng, J. Ma, Q. Liu, L. Wang and Q. Gu, Proceedings of the 40th International Conference on Machine Learning, 2023, pp. 11827–11846 Search PubMed.
- J. Cremer, T. Le, F. Noé, D.-A. Clevert and K. T. Schütt, Chem. Sci., 2024, 15, 14954–14967 RSC.
- H. Lin, Y. Huang, O. Zhang, S. Ma, M. Liu, X. Li, L. Wu, J. Wang, T. Hou and S. Z. Li, Chem. Sci., 2025, 16, 1417–1431 RSC.
- A. Lugmayr, M. Danelljan, A. Romero, F. Yu, R. Timofte and L. Van Gool, Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2022, pp. 11461–11471 Search PubMed.
- A. Schneuing, C. Harris, Y. Du, K. Didi, A. Jamasb, I. Igashov, W. Du, C. Gomes, T. L. Blundell, P. Lio, M. Welling, M. Bronstein and B. Correia, Nat. Comput. Sci., 2024, 4, 899–909 CrossRef CAS PubMed.
- Q. Liu, Rectified Flow: A Marginal Preserving Approach to Optimal Transport, arXiv, 2022, preprint, arXiv:2209.14577, DOI:10.48550/arXiv.2209.14577, https://arxiv.org/abs/2209.14577.
- M. S. Albergo and E. Vanden-Eijnden, The Eleventh International Conference on Learning Representations, 2023 Search PubMed.
- Y. Lipman, R. T. Q. Chen, H. Ben-Hamu, M. Nickel and M. Le, The Eleventh International Conference on Learning Representations, 2023 Search PubMed.
- I. Dunn and D. R. Koes, Mixed Continuous and Categorical Flow Matching for 3D De Novo Molecule Generation, arXiv, 2024, preprint, arXiv:2404.19739, DOI:10.48550/arXiv.2404.19739, https://arxiv.org/abs/2404.19739.
- H. Stark, B. Jing, R. Barzilay and T. Jaakkola, Forty-first International Conference on Machine Learning, 2024 Search PubMed.
- R. Irwin, A. Tibo, J. P. Janet and S. Olsson, The 28th International Conference on Artificial Intelligence and Statistics, 2025 Search PubMed.
- J. Cremer, R. Irwin, A. Tibo, J. P. Janet, S. Olsson and D.-A. Clevert,FLOWR: Flow Matching for Structure-Aware De Novo, Interaction- and Fragment-Based Ligand Generation, arXiv, 2025, preprint, arXiv:2504.10564, DOI:10.48550/arXiv.2504.10564, https://arxiv.org/abs/2504.10564.
- A. Schneuing, I. Igashov, A. W. Dobbelstein, T. Castiglione, M. M. Bronstein and B. Correia, The Thirteenth International Conference on Learning Representations, 2025 Search PubMed.
- T. Le, J. Cremer, D.-A. Clevert and K. T. Schütt, J. Cheminf., 2025, 17, 90 CAS.
- K. Adams and C. W. Coley, The Eleventh International Conference on Learning Representations, 2023 Search PubMed.
- K. Adams, K. Abeywardane, J. Fromer and C. W. Coley, The Thirteenth International Conference on Learning Representations, 2025 Search PubMed.
- I. Igashov, H. Stärk, C. Vignac, A. Schneuing, V. G. Satorras, P. Frossard, M. Welling, M. Bronstein and B. Correia, Nat. Mach. Intell., 2024, 6, 417–427 CrossRef.
- M. Ghorbani, L. Gendelev, P. Beroza and M. J. Keiser, Autoregressive fragment-based diffusion for pocket-aware ligand design, arXiv, 2023, preprint, arXiv:2401.05370, DOI:10.48550/arXiv.2401.05370, https://arxiv.org/abs/2401.05370.
- J. Xie, S. Chen, J. Lei and Y. Yang, J. Chem. Inf. Model., 2024, 64, 2554–2564 CrossRef CAS PubMed.
- P. G. Francoeur, T. Masuda, J. Sunseri, A. Jia, R. B. Iovanisci, I. Snyder and D. R. Koes, J. Chem. Inf. Model., 2020, 60, 4200–4215 CrossRef CAS PubMed.
- M. Backenköhler, J. Groß, V. Wolf and A. Volkamer, J. Chem. Inf. Model., 2024, 64, 4009–4020 CrossRef PubMed.
- T. Le, J. Cremer, F. Noe, D.-A. Clevert and K. T. Schütt, The Twelfth International Conference on Learning Representations, 2024 Search PubMed.
- X. Q. Lewell, D. B. Judd, S. P. Watson and M. M. Hann, J. Chem. Inf. Comput. Sci., 1998, 38, 511–522 CrossRef CAS PubMed.
- J. Degen, C. Wegscheid-Gerlach, A. Zaliani and M. Rarey, ChemMedChem, 2008, 3, 1503–1507 CrossRef CAS PubMed.
- S. Luo, J. Guan, J. Ma and J. Peng, Advances in Neural Information Processing Systems, 2021, pp. 6229–6239 Search PubMed.
- M. Steinegger and J. Söding, Nat. Biotechnol., 2017, 35, 1026–1028 CrossRef CAS PubMed.
- T. Le, F. Noe and D.-A. Clevert, Proceedings of the First Learning on Graphs Conference, 2022, p. 30 Search PubMed.
- M. Buttenschoen, G. M. Morris and C. M. Deane, Chem. Sci., 2024, 15, 3130–3139 RSC.
- C. Harris, K. Didi, A. R. Jamasb, C. K. Joshi, S. V. Mathis, P. Lio and T. Blundell, Benchmarking Generated Poses: How Rational is Structure-based Drug Design with Generative Models?, arXiv, 2023, preprint, arXiv:2308.07413, DOI:10.48550/arXiv.2308.07413, https://arxiv.org/abs/2308.07413.
- A. K. Rappe, C. J. Casewit, K. S. Colwell, W. A. I. Goddard and W. M. Skiff, J. Am. Chem. Soc., 1992, 114, 10024–10035 CrossRef CAS.
- P. C. D. Hawkins, A. G. Skillman and A. Nicholls, J. Med. Chem., 2007, 50, 74–82 CrossRef CAS PubMed.
- M. S. Albergo, N. M. Boffi and E. Vanden-Eijnden, arXiv, 2023arXiv:2303.08797, DOI:10.48550/arXiv.2303.08797.
- J. Song, C. Meng and S. Ermon, International Conference on Learning Representations, 2021 Search PubMed.
- A. Campbell, J. Benton, V. De Bortoli, T. Rainforth, G. Deligiannidis and A. Doucet, Proceedings of the 36th International Conference on Neural Information Processing Systems, Red Hook, NY, USA, 2022 Search PubMed.
- Y. Lipman, M. Havasi, P. Holderrieth, N. Shaul, M. Le, B. Karrer, R. T. Chen, D. Lopez-Paz, H. Ben-Hamu and I. Gat, arXiv, 2024, preprint, arXiv:2412.06264, DOI:10.48550/arXiv.2412.06264.
- J. Degen, C. Wegscheid-Gerlach, A. Zaliani and M. Rarey, ChemMedChem, 2008, 3, 1503–1507 CrossRef CAS PubMed.
- J. Li, R. Abel, K. Zhu, Y. Cao, S. Zhao and R. A. Friesner, Proteins: Struct., Funct., Bioinf., 2011, 79, 2794–2812 CrossRef CAS.
- C. M. S. OpenEye, BROOD, 2024, https://www.eyesopen.com/.
- A. Gaulton, L. J. Bellis, A. P. Bento, J. Chambers, M. Davies, A. Hersey, Y. Light, S. McGlinchey, D. Michalovich, B. Al-Lazikani and J. P. Overington, Nucleic Acids Res., 2012, 40, D1100–D1107 CrossRef CAS PubMed.
- J. Ho, A. Jain and P. Abbeel, Advances in Neural Information Processing Systems, 2020, pp. 6840–6851 Search PubMed.
- Y. Song, J. Sohl-Dickstein, D. P. Kingma, A. Kumar, S. Ermon and B. Poole, International Conference on Learning Representations, 2021 Search PubMed.
- P. Dhariwal and A. Q. Nichol, Advances in Neural Information Processing Systems, 2021 Search PubMed.
- R. Rombach, A. Blattmann, D. Lorenz, P. Esser and B. Ommer, Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2022, pp. 10684–10695 Search PubMed.
- X. Liu, X. Zhang, J. Ma, J. Peng and Q. Liu, The Twelfth International Conference on Learning Representations, 2024 Search PubMed.
- A. Lou, C. Meng and S. Ermon, Proceedings of the 41st International Conference on Machine Learning, 2025 Search PubMed.
- I. Gat, T. Remez, N. Shaul, F. Kreuk, R. T. Q. Chen, G. Synnaeve, Y. Adi and Y. Lipman, The Thirty-eighth Annual Conference on Neural Information Processing Systems, 2024 Search PubMed.
- X. Liu, C. Gong and Q. Liu, The Eleventh International Conference on Learning Representations, 2023 Search PubMed.
- R. T. Q. Chen, Y. Rubanova, J. Bettencourt and D. K. Duvenaud, Advances in Neural Information Processing Systems, 2018 Search PubMed.
- B. Li, K. Xue, B. Liu and Y.-K. Lai, Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2023, pp. 1952–1961 Search PubMed.
- C. Vignac, N. Osman, L. Toni and P. Frossard, Machine Learning and Knowledge Discovery in Databases: Research Track, Cham, 2023, pp. 560–576 Search PubMed.
- A. Campbell, J. Yim, R. Barzilay, T. Rainforth and T. Jaakkola, Proceedings of the 41st International Conference on Machine Learning, 2024, pp. 5453–5512 Search PubMed.
- A.-A. Pooladian, H. Ben-Hamu, C. Domingo-Enrich, B. Amos, Y. Lipman and R. T. Q. Chen, Proceedings of the 40th International Conference on Machine Learning, 2023, pp. 28100–28127 Search PubMed.
- A. Tong, K. FATRAS, N. Malkin, G. Huguet, Y. Zhang, J. Rector-Brooks, G. Wolf and Y. Bengio, Transactions on Machine Learning Research, 2024 Search PubMed.
- L. Klein, A. Krämer and F. Noe, Thirty-seventh Conference on Neural Information Processing Systems, 2023 Search PubMed.
- M. S. Albergo, M. Goldstein, N. M. Boffi, R. Ranganath and E. Vanden-Eijnden, Proceedings of the 41st International Conference on Machine Learning, 2024, pp. 921–937 Search PubMed.
- B. Jing, E. Erives, P. Pao-Huang, G. Corso, B. Berger and T. Jaakkola, EigenFold: Generative Protein Structure Prediction with Diffusion Models, arXiv, 2023, preprint, arXiv:2304.02198, DOI:10.48550/arXiv.2304.02198, https://arxiv.org/abs/2304.02198.
| This journal is © The Royal Society of Chemistry 2026 |