Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Physics-informed machine learning for predicting temperature-dependent chemical properties
Mahyar
Rajabi-Kochi
 ab,
Hanie
Rezaei
c,
Sartaaj Takrim
Khan
ab,
Hanie
Rezaei
c,
Sartaaj Takrim
Khan
 ab,
Bhanu
Mamillapalli
a,
Maryam
Ebrahimiazar
c,
Haoming
Ye
c,
Rose
Moosavian
c,
Mohammad
Zargartalebi
c,
David
Sinton
c and
Seyed Mohamad
Moosavi
ab,
Bhanu
Mamillapalli
a,
Maryam
Ebrahimiazar
c,
Haoming
Ye
c,
Rose
Moosavian
c,
Mohammad
Zargartalebi
c,
David
Sinton
c and
Seyed Mohamad
Moosavi
 *ab
*ab
aChemical Engineering & Applied Chemistry, University of Toronto, Toronto, ON M5S 3E5, Canada. E-mail: mohamad.moosavi@utoronto.ca
bVector Institute for Artificial Intelligence, Toronto, ON M5G 0C6, Canada
cMechanical & Industrial Engineering, University of Toronto, Toronto, ON M5S 3G8, Canada
First published on 30th March 2026
Abstract
Emerging energy and electronic systems rely on the thermodynamic properties of chemical and cooling fluids. These properties are a function of both chemical structure and temperature. For instance, the dynamic viscosity of a fluid can vary by orders of magnitude across the operating range of a cooling system. However, capturing this behavior remains a challenge for experimental and modelling approaches. Machine learning models, although powerful for fixed temperatures, fail to generalize across temperatures due to a lack of data and a lack of embedded physical constraints. Here, we introduce a physics-informed machine learning framework that incorporates established physical relationships, such as the Arrhenius equation or Clausius–Clapeyron, to capture both chemical diversity and temperature dependence. We demonstrate that decoupling chemistry from thermodynamic conditions enables accurate prediction of temperature-dependent dynamic viscosity for both pure compounds and binary mixtures, which we validated with new experimental data. Through a materials-discovery campaign for cooling applications, we show that neglecting temperature effects can cause relative efficiency errors exceeding an order of magnitude, leading to inaccurate materials ranking and suboptimal fluid selection. Finally, we extend the framework to other properties, such as vapor pressure and diffusion coefficient, highlighting a generalizable strategy for accelerating fluid property prediction and design for sustainable technologies.
Introduction
Industrial fluids play a vital role in enabling technologies across the energy, aerospace, and transportation sectors. Particularly as these industries shift toward cleaner energy sources, developing advanced fluids becomes increasingly critical, especially for electrification. For instance, thermal management in long-range electric vehicles requires coolants with low viscosity and enhanced thermal conductivity to prevent overheating and optimize performance.1,2 Increased demand for computing power is also generating additional need for immersion cooling fluids with tailored thermophysical properties.3,4 However, discovering high-performing fluids for emerging applications remains a challenge due to the vast design space of chemical formulations and the time-intensive nature of experimental fluid characterization.5–7 Recent advances have sought to address this by combining high-throughput experimentation with machine learning to accelerate property prediction by extracting relationships between chemical structures and properties.8–12 These approaches have proven to be effective across diverse chemistries, including mixtures,13 polymers,14 hydrocarbons15 and ionic liquids,16,17 and for various properties, including dynamic viscosity18,19 and heat capacity.20,21Temperature, alongside chemical composition, plays a key role in determining fluid properties. In systems like tube heat exchangers, battery immersion cooling, and electronics thermal management, properties such as viscosity can vary by orders of magnitude with temperature.22,23 Selecting suitable fluids for these applications requires accurate knowledge of thermophysical properties over a wide range of temperatures.24 However, measuring these properties across all relevant temperatures is impractical.25 Recent machine learning models attempted to address this gap by including temperature as an input variable. Yet, these models are typically constrained to the narrow ranges of temperatures of the training data, limiting their extrapolative power for use in practical engineering design.18,26 Moreover, most machine learning models are inherently agnostic to physical laws and thermodynamic constraints,27 they function as black boxes with limited interpretability, generalizability, and integration with physics-based simulators.28
To overcome these limitations, recent studies have focused on integrating physical knowledge into machine learning models.29–31 One approach, referred to as Physics-Informed Neural Networks (PINNs), incorporates governing equations such as partial differential equations (e.g., Navier–Stokes equation) into the loss function by penalizing their residuals, evaluated through automatic differentiation at collocation points.32 Similar hard-constraint approaches enforce thermodynamic consistency by incorporating relations such as the Gibbs–Duhem equation into the loss function, which has been applied to predict thermophysical properties like activity coefficients.33,34 These strategies ensure compliance with fundamental balance and consistency equations, but they do not explicitly encode constitutive relationships that govern the dependence of material properties on thermodynamic state variables (e.g., temperature, pressure). As a result, such dependencies must be learned empirically from data, which often limits generalizability beyond the training conditions, particularly for low-dimensional problems where constitutive equations are physically valid and substantially simpler to implement. An alternative approach is to leverage physics-informed equations to capture the correlation between data and temperature in machine learning to bridge between chemical structure and temperature-dependent behavior. Physics-informed correlations, such as the Arrhenius equation, are traditionally used to capture the temperature dependence of fluid properties because they are derived from fundamental thermodynamic and kinetic principles, linking molecular-level activation processes to macroscopic property variation. While these equations are valid across wide temperature ranges, their coefficients are chemistry-specific and typically obtained from curve-fitting to experimental data, which is mostly unavailable for novel compounds. This motivates a hybrid approach that can accurately predict fluid properties across diverse chemistries and temperature conditions.
In this work, we introduce a framework that encodes temperature dependence of thermophysical properties using established equations, such as the Arrhenius equation, and uses machine learning to predict the chemistry-dependent coefficients of these equations for unseen materials. We use dynamic viscosity as a case study and evaluate this approach to predict this property in industrial fluids. By compiling an experimental dataset from different sources to train this model, we show that the model achieves high accuracy and generalizability across different chemistries and temperatures. For a selection of industrial fluids relevant to cooling applications, we show that incorporating temperature dependence significantly affects key performance metrics in fluid discovery. Finally, we illustrate that this methodology is extensible to other temperature-dependent properties, such as vapor pressure and diffusivity, where well-established physical relationships exist.
Machine learning for temperature-dependent properties
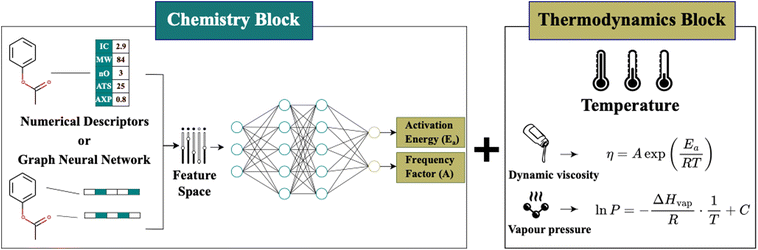
To accurately predict fluid properties across a wide range of chemistries and temperatures, we develop a machine learning framework that integrates chemical structure information with established thermodynamic equations. In engineering, physics-based models such as the Arrhenius equation and the Clausius–Clapeyron relation have long been used to describe how thermophysical properties vary with temperature. These equations isolate the effect of temperature, suggesting that temperature and chemistry can be treated as separable factors in predictive modeling. Building on this principle, we propose a hybrid modeling approach with two components: a chemistry block and a thermodynamics block (Fig. 1). The chemistry block uses state-of-the-art machine learning models to predict the coefficients of well-established temperature-dependent equations. The thermodynamics block then applies these coefficients within physics-based expressions to model how properties vary with temperature. This separation enables the model to generalize across temperatures using a small amount of data while focusing its learning capacity on capturing how chemical structure influences fluid properties.This architecture is flexible and can accommodate different temperature–property relationships, depending on the property and the application of interest. For instance, while the Arrhenius equation is suitable to predict dynamic viscosity for most industrial fluids, incorporating alternative formulations such as the Vogel–Fulcher–Tammann (VFT)35 equation can better model behavior near the glass transition temperature.
To capture the chemical similarity between fluids, we explore two approaches for encoding molecular structure: (1) Mordred numerical descriptors36 and (2) molecular graph representations based on a graph neural network (GNN) via Chemprop.37 In training these machine learning models, we split the dataset based on different scenarios to ensure generalizability. Finally, to assess prediction confidence, we adopt a bagging regressor strategy that groups multiple models together and estimates uncertainty based on the variance of their predictions. Adopting this approach provides a practical tool for identifying predictions that may be less reliable, especially for molecules dissimilar to those in the training distribution.
Case study on dynamic viscosity
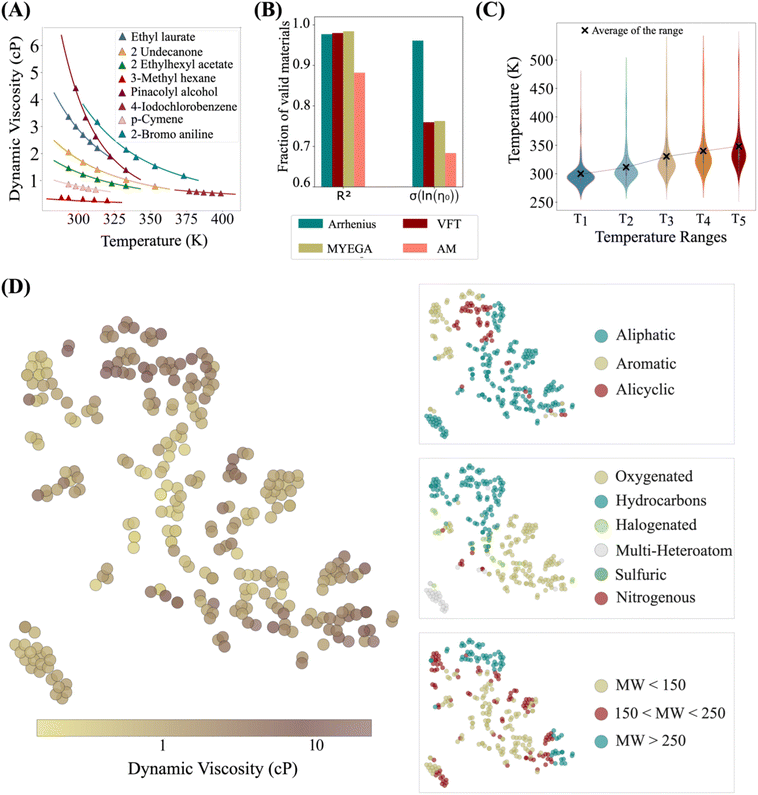
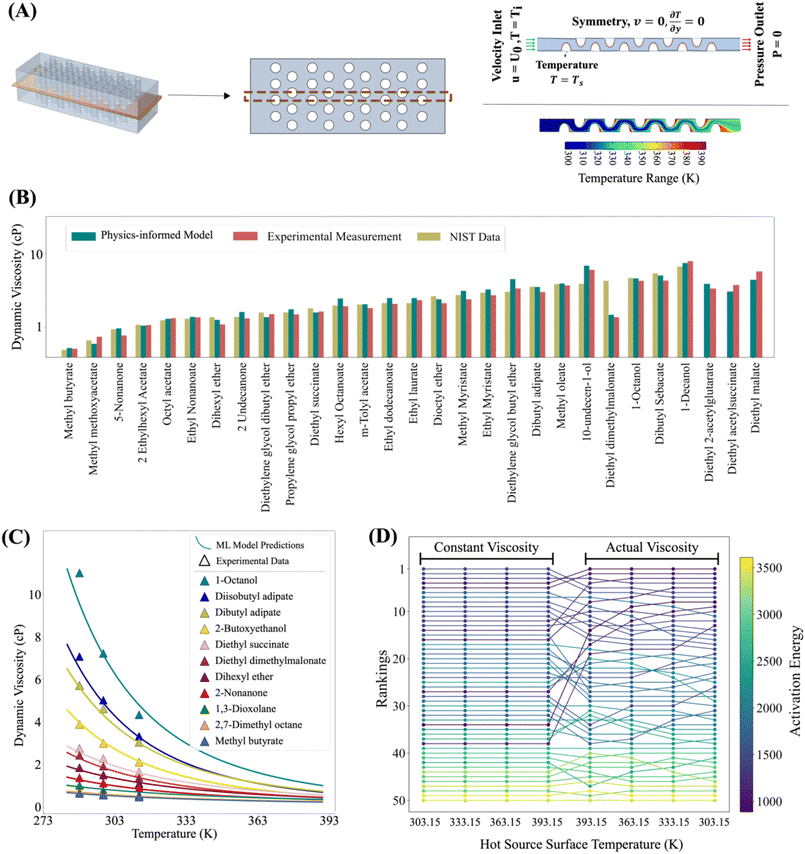
To demonstrate the effectiveness of this approach, we focus on dynamic viscosity, a key thermophysical property of fluids with broad industrial relevance ranging from electronics16 to cosmetics,38 cooling systems,39 and energy storage.40 Dynamic viscosity is an ideal test case for the model as it is strongly influenced by both temperature and chemical structure.41,42 Increasing temperature typically decreases viscosity by enhancing molecular mobility, while chemistry governs inter- and intra-molecular interactions that resist flow. Intermolecular forces, such as hydrogen bonding and van der Waals interactions, affect viscosity by modulating attraction between molecules. Intramolecular characteristics—such as rigidity and flexibility—influence how easily molecules reorient and move past one another.43To train and evaluate the model, we compiled a dataset of dynamic viscosity measurements from the National Institute of Standards and Technology (NIST) and previously published experimental studies.44–46 The dataset includes 720 organic fluids measured over a temperature range of 250 K to 550 K. As shown in Fig. 2A, viscosity exhibits substantial variation with temperature, motivating the use of physics-informed models to capture this dependence. Several equations have been proposed for this purpose, including the Arrhenius, VFT, Mauro–Yue–Ellison–Gupta–Allan (MYEGA),47 and Avramov–Milchev (AM),48 which differ in how they represent activation energy and cooperative dynamics. To evaluate which model best describes organic fluids, we performed regression analysis based on experimental data, monitoring both accuracy and the stability of shared parameters such as the pre-exponential (frequency) factor. While Arrhenius, VFT, and MYEGA each provided accurate fits for over 97% of materials, the Arrhenius model yielded more stable coefficients, indicating greater robustness (Fig. 2B). For the remaining ∼3% of materials where none of the models performed adequately, further inspection revealed that 75% of these compounds contained thiol functional groups and 88% exhibited very low viscosities (<0.5 cP), both of which may contribute to deviations from model predictions and challenges in experimental measurement.
Accordingly, we use the Arrhenius equation as the physics-informed component in the thermodynamics block of the model, while maintaining a model-agnostic design by benchmarking against the VFT, MYEGA, and AM equations. In the chemistry block, we represent molecular structure using Mordred descriptors and graphs. Because each fluid in the dataset has viscosity data reported over different temperature subranges and with varying numbers of measurements, and since the available data are concentrated at intermediate temperatures and sparse at the extremes, random sampling would bias the model toward the mid-range. To avoid this, we apply a quantile-to-median temperature selection method to achieve more uniform and representative coverage across the full temperature distribution. This method selects five data points per fluid, evenly distributed across the available temperature range, and categorizes them into five temperature bins (Fig. 2C).
The dataset spans a wide range of chemical structures and characteristics relevant to viscosity. To better characterize the chemical diversity of this dataset, we use Mordred descriptors and apply t-SNE for two-dimensional projection. As illustrated in Fig. 2D, the dataset includes compounds with molecular weights ranging from 16 to 846 g mol−1, encompassing both light, volatile species and heavier, more complex molecules. Structural diversity includes linear, branched, aromatic, and alicyclic compounds. In terms of chemical composition, the dataset contains pure hydrocarbons as well as molecules with one or more heteroatoms, such as oxygen, nitrogen, sulfur, and halogens, distributed across various functional groups. This broad chemical diversity, combined with the wide temperature range, provides a robust foundation for evaluating the model's performance across varied molecular systems and operating conditions.
Model evaluation across temperature ranges and chemical classes
For known chemicals, when experimental data is available at various temperatures, fitting the Arrhenius equation yields very high accuracy in predicting dynamic viscosity at new temperatures (MAE = 0.09, SRCC = 0.99; see Section S22 of the SI). However, we are interested in predicting the properties of materials when such data is not available. Therefore, we focus on evaluating the model's ability to predict dynamic viscosity for previously unseen chemicals. For this, we assess model performance under three distinct evaluation settings, namely new materials, new material classes, and new materials at new temperatures, designed to evaluate different aspects of generalization and extrapolation (see Methods section for details of train-test splitting approach).We first evaluate model robustness on unseen materials using a material-based train-test split. We randomly select 70% of the materials for the training set and reserve the remaining 30% for testing. The results for our model's performance are shown in Fig. 3, where we observe high accuracy in predicting the dynamic viscosity of new fluids in the test set. We compare our physics-informed model against two baseline models that do not incorporate any physical equations in the pre-final layer, all trained and evaluated using the same train and test sets. The first baseline is an isothermal model trained exclusively on data from a single temperature level (the mid-range T3), representing a limited-temperature-data scenario. The second baseline is a multitemperature model trained across all five temperature levels, where temperature is treated as an additional input variable concatenated with chemical descriptors. Neither baseline achieves satisfactory performance (Fig. 3A and B). In the absence of an explicit inductive bias49—the physics-informed equation—these models must learn the highly nonlinear temperature–viscosity relationship directly from data. Even the isothermal model, despite being trained specifically at T3, underperforms relative to the physics-informed approach at the same temperature (Fig. 3A and B). We note that in this setting, PINNs in their original format are not benchmarked, as they are designed to enforce governing partial differential equations rather than constitutive temperature–property relations, which is out of the scope of this study. Additional benchmarking results, including model selection analyses and evaluations of predictive uncertainty using calibrated metrics such as sharpness, coverage, and CRPS, are provided in Sections S12 and S15 of the SI.
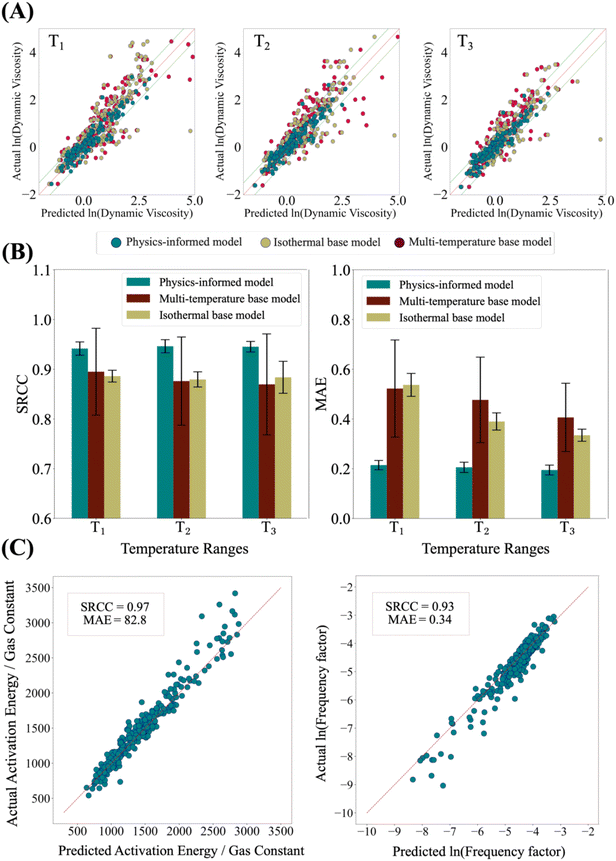 | ||
| Fig. 3 Prediction performance of the physics-informed model. (A) Parity plots for comparison of the physics-informed model and two baseline models at three temperature levels shown in Fig. 2C. (B) Comparison of prediction accuracy using SRCC and MAE. Error bars indicate the standard deviation of predictions computed from an ensemble of 20 models trained on different subset of training datasets. (C) Prediction accuracy of Arrhenius equation coefficients: the logarithmic frequency factor (right) and the activation energy divided by the gas constant (left). Ground-truth coefficients are derived from curve fitting of experimental data (see Methods for details). Additional performance metrics and parity plots across all five temperature levels are provided in the SI, Fig. S2. | ||
The high predictive performance of the physics-informed model arises from its ability to accurately predict the Arrhenius equation coefficients directly from molecular structure. As shown in Fig. 3C, the model predicts these coefficients with MAE values of 0.34 and 82.8 and SRCC values of 0.93 and 0.97, respectively, when compared against coefficients obtained from curve fitting of experimental data (see Methods for details of curve fitting approach). This approach helps the model to capture the underlying physical trends governing viscosity–temperature behavior. To further understand model performance across chemistry classes, we look into model performance for each materials class. For this, we characterize chemical diversity in the dataset using three complementary criteria: hydrocarbon content, heteroatomic composition, and molecular weight. Under this setting, the physics-informed model demonstrates strong and consistent performance across the majority of chemical classes (Table 1). Notably, performance for halogenated compounds shows a larger drop (SRCC = 0.74), which can be attributed to their limited representation in the dataset (approximately 3% of samples) and their narrow viscosity distribution. As shown in Fig. S4, halogenated compounds occupy a compact low-viscosity regime, making rank-based metrics such as SRCC particularly sensitive. Small absolute prediction errors can lead to disproportionate reductions in correlation.
| Chemical criteria | % Of total | SRCC | MAE | |
|---|---|---|---|---|
| Hydrocarbon | Alicyclic | 9.7 | 0.95 | 0.25 |
| Aliphatic | 73.4 | 0.95 | 0.21 | |
| Aromatic | 16.9 | 0.93 | 0.27 | |
| Heteroatom | Oxygenated | 48.5 | 0.9 | 0.27 |
| Nitrogenous | 2.9 | 0.94 | 0.41 | |
| Sulfuric | 7.6 | 0.96 | 0.13 | |
| Halogenated | 3.8 | 0.74 | 0.16 | |
| Hydrocarbon | 20.7 | 0.98 | 0.18 | |
| Multi-heteroatom | 16.4 | 0.98 | 0.12 | |
| Molecular weight (g mol−1) | MW < 150 | 51.9 | 0.93 | 0.23 |
| 150 < MW < 250 | 33.3 | 0.91 | 0.19 | |
| MW > 250 | 14.8 | 0.91 | 0.22 | |
It is interesting to evaluate model generalization to unseen classes of materials, in which all compounds belonging to a specific chemical class are excluded from training and used exclusively for testing. We employ a leave-one-class-out (LOCO) train-test splitting strategy and summarize the results in Table S6. We observe that while overall performance decreases relative to the previous splitting scheme, the model retains good predictive capability for most classes, highlighting its ability to extrapolate beyond training chemical classes.
To evaluate the model's ability to extrapolate simultaneously across materials and temperatures, we implement a dual-axis splitting strategy. In addition to a 70/30 material-based split, we entirely withhold the lowest temperature range (T1) from training, using only the intermediate-to-high ranges (T2–T5) for model development. We then evaluated the model's accuracy in predicting the dynamic viscosity of unseen chemistries at the withheld T1 temperature. As shown in Fig. 4A and B, the Arrhenius-based model maintains high accuracy in this scenario, confirming that the physics-informed architecture effectively generalizes across both novel chemistries and unexplored temperature ranges (A parallel analysis for the T5 temperature is available in the SI, Fig. S10).
In a practical scenario, it is essential to quantify the reliability and uncertainty of model predictions for new chemistries. We estimate predictive uncertainty by measuring the variance across an ensemble of machine learning models trained on different subsets of the data (see Methods and SI for details of uncertainty assessment). To ensure that these uncertainty estimates are valid, we evaluated them using calibrated metrics that jointly assess calibration and sharpness. These analyses confirm that our physics-informed model provides more reliable uncertainty quantification than baseline approaches (see SI – Section S15 for details). Consequently, the estimated variance can be regarded as a trustworthy indicator of predictive confidence. As shown in Fig. S5, this approach effectively assigns high uncertainty to erroneous predictions, providing a practical mechanism to flag low-confidence outputs. The combination of high predictive accuracy and robust, validated uncertainty quantification enhances the model's reliability, enabling its application to practical challenges in fluid discovery and materials design.
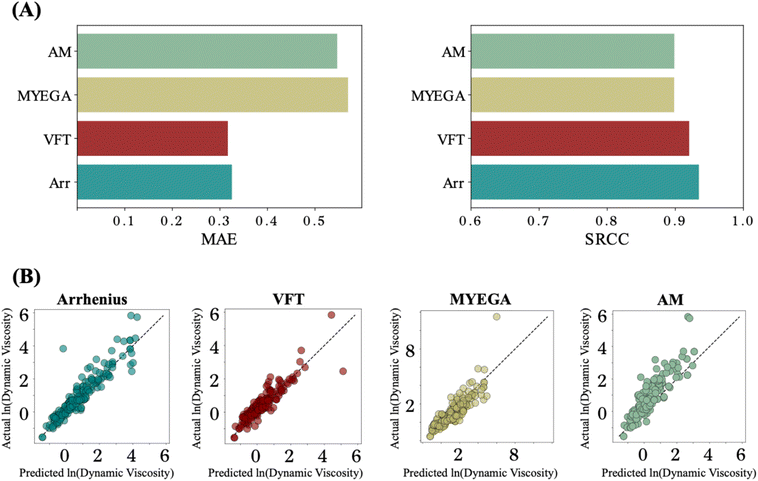
Different viscosity–temperature equations encode distinct physical assumptions, making it essential to benchmark the physics-informed framework across multiple equations to assess its generality and reliability. We construct different versions of physics-informed models that incorporate several candidate equations. In each case, the model predicts chemistry-dependent coefficients for the selected equation, which are then used to reconstruct viscosity as a function of temperature. The Arrhenius equation achieves the lowest mean absolute error and highest rank correlation, with the VFT formulation performing comparably well, whereas MYEGA and particularly the Adam–Gibbs (AM) equation exhibit reduced accuracy (Fig. 4A and B). These trends are consistent with the regression analysis in Fig. 2B, which indicates that the AM equation is less valid across the present dataset, reflecting limitations of its entropy-based formulation at low temperatures. More broadly, models with fewer free parameters (such as Arrhenius) exhibit greater stability under temperature extrapolation. In contrast, the additional temperature-dependent activation energy term in MYEGA introduces increased variability, leading to degraded predictive performance in the low-temperature regime.
Understanding physics-informed model predictions
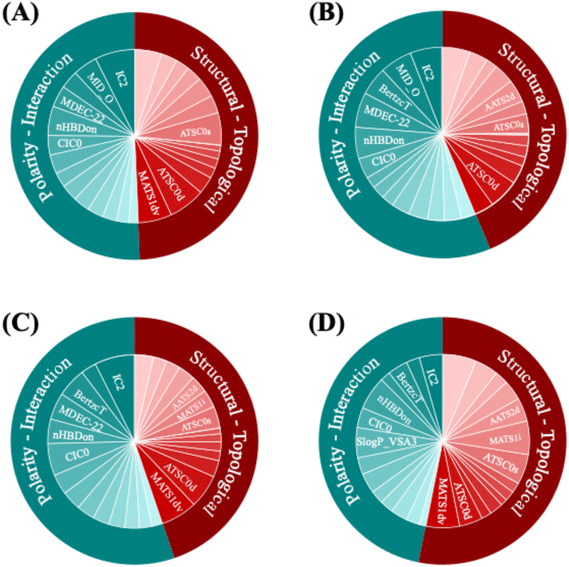
While the model achieves strong predictive performance, understanding how it interprets molecular structure provides deeper insight into the relationship between chemistry, temperature, and viscosity. A key advantage of the physics-informed approach is the interpretability of the Arrhenius equation parameters, which are directly predicted from molecular structure. This equation contains two critical coefficients: the frequency factor, which represents the dynamic viscosity at infinite temperature, and the activation energy, which determines the sensitivity of viscosity to temperature changes by reflecting the energy barrier for molecular motion.To investigate which chemical features govern the prediction of each coefficient, we categorize the Mordred descriptors into two main groups. The first group, structural–topological descriptors, summarizes the molecule's size and internal architecture (its overall extent, connectivity, and shape). The second group, polarity-interaction descriptors, captures how the molecule tends to engage with other species (its general polarity and capacity for noncovalent interactions as expressed on the molecular surface). Feature-importance analysis using SHAP values reveals distinct patterns across the two coefficients.50 For the frequency factor, structural-topological descriptors are the dominant contributors. These descriptors capture properties such as the distribution of sigma electrons (ATSC0D), valence electron counts (MATs1sv), intrinsic state indices (ATSC0s), and van der Waals volume (ATSC1v) (Fig. 5A), emphasizing the importance of molecular size, connectivity, and framework rigidity in setting baseline viscosity. In contrast, polarity-interaction descriptors play a dominant role in predicting the activation energy (Fig. 5B). These descriptors capture essential features related to electronic charge distribution, such as the presence of polar atoms (MID-O), hydrogen bond donors (nHBDon), and non-polar regions (MDEC-22), as well as global molecular complexity (BertzcT). Their prominence highlights the critical role of intermolecular forces, such as hydrogen bonding and dipole interactions, in determining the temperature dependence of viscosity.
Comparing feature importance between the physics-informed model and a baseline model that does not incorporate a physics-informed equation further illustrates this distinction. In the physics-informed model, polarity-interaction descriptors contribute over 55% to the predictions, surpassing structural-topological descriptors (Fig. 5C). In contrast, in the baseline model, their contribution decreases to less than 45% (Fig. 5D). Despite the critical importance of the polarity-interaction features for capturing temperature-dependent behavior, the baseline model fails to recognize their role. This shift underscores how embedding physical knowledge into the model architecture promotes more accurate recognition of chemically meaningful interactions, improving both interpretability and predictive robustness.
These results show that the physics-informed model does not memorize training data, but learns chemically meaningful relationships governing fluid behavior, as evidenced by its ability to generalize under leave-one-class-out evaluation (see SI, Section 20). By leveraging the synergistic influence of structural-topological, and polarity-interaction descriptors, the model captures baseline viscosity and temperature dependence that remain predictive even when entire chemical classes are excluded from training. This balanced integration enhances the model's generalization capability across a wide range of chemistries and temperatures, enabling more reliable application to new material discovery and fluid design.
Application in engineering designs
One of the practical motivations of this work is to understand the importance of accurately capturing the temperature dependence of thermophysical fluid properties for practical industrial applications. In many engineering applications, fluids undergo a wide range of operating temperatures. However, because experimental measurements across such ranges are challenging and time-consuming, it is common practice to assume constant fluid properties. This simplification, while convenient, lacks a physical foundation and can lead to significant errors in simulations, resulting in incorrect fluid rankings and suboptimal material selection.As a case study, we examine fluid performance evaluation for cooling applications, which are critical in emerging technologies such as battery immersion cooling and data center cooling. Previous studies have shown that the heat transfer coefficient as a performance criterion exhibits the highest sensitivity to dynamic viscosity.51 We focus on a representative system involving flow across staggered tube banks, commonly used in shell-and-tube heat exchangers (Fig. 6A). The temperature distribution in the flow varies between 300 K and 390 K, underscoring the need to model temperature-dependent dynamic viscosity accurately. To evaluate the impact of this dependence, we selected 50 candidate thermal fluids (see the SI for the full list). We searched the NIST Standard Reference Database 103b52 using both the PubChem name and SMILES string of each compound. For three of the fluids, dynamic viscosity data are not available at the target temperatures. So, to establish a consistent benchmark, we decided to measure the dynamic viscosity of thermal fluids experimentally at 288.15, 298.15, and 313.15 K. As shown in Fig. 6B and C, the physics-informed model provides strong predictive performance against measured data. We obtained accurate predictions for all materials and temperatures, enabling us to use the Arrhenius equation with parameters from the machine learning model to predict the dynamic viscosity across all the temperature ranges of interest, that is, 300 K to 390 K.
To assess the importance of incorporating temperature dependence, we rank the fluids under two scenarios: one assuming constant dynamic viscosity, and the other using temperature-dependent viscosity predicted by this model. Fig. 6D shows that neglecting temperature dependence leads to an incorrect fluid rankings. Fluids appearing efficient under constant viscosity assumptions are often outperformed by others when viscosity–ttemperature variations are considered. Notably, fluid selection depends strongly on operating conditions. When hot surface temperature is varied, incorporating temperature dependence leads to significant shifts in rankings (Fig. 6D), with some fluids flowing more efficiently at higher temperatures—critical insights for industrial fluid design and selection.
These findings highlight the critical role of accurately modeling temperature-dependent fluid properties in engineering design. As a proof of principle, our physics-informed ML model enables incorporation of the dynamic viscosity within COMSOL simulation across temperature ranges. Assuming constant viscosity not only introduces significant simulation errors but also risks selecting suboptimal materials. By leveraging the physics-informed model, engineers can predict property variations under realistic conditions and better understand how molecular structure governs fluid performance. Looking forward, this approach should extend to other temperature-dependent fluid properties, such as thermal conductivity and heat capacity, which also play a critical role in real-world cooling applications and would require further validation before use as an engineering design tool.
Discussion
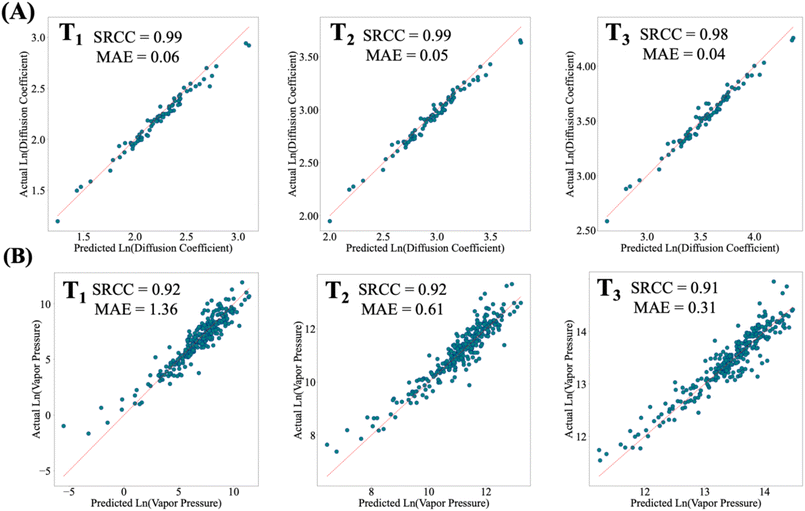
The main goal of this work was to develop a machine learning framework capable of predicting the temperature dependence of fluid properties across diverse chemistries. We hypothesized that by decoupling thermodynamic conditions from the chemical structure and encoding temperature effects through physics-informed equations such as the Arrhenius equation, accurate property predictions could be achieved across both unseen temperatures and chemistries. We therefore developed a method that leverages these physics-informed equations to capture thermodynamic state and uses machine learning to predict their chemistry-dependent coefficient. The case study on dynamic viscosity validated this approach, demonstrating high predictive accuracy while providing uncertainty estimates for each prediction. For fluids with chemistries that are quite different from the training set, the model correctly identifies unreliable predictions, guiding targeted experimental measurements. In these cases, as the physics-informed equation provides the structure for interpolations, only a few new measurements are needed to capture the general temperature-dependence trend.Beyond proof of principle on dynamic viscosity, an important question is whether the physics-informed approach generalizes to other properties, particularly those governed by different physical relationships. To explore this, we extended the framework to two additional properties: vapor pressure and infinite dilution diffusion coefficients. While an Arrhenius-type relationship captures the temperature dependence of diffusion coefficients through energy barriers within a fluid,53,54 the vapor pressure of a fluid is correlated with temperature through the Clausius–Clapeyron equation, where the coefficients represent the enthalpy of vaporization and a reference vapor pressure.55Fig. 7 shows that this physics-informed model achieves high predictive performance for both properties across different temperature ranges (see SI for dataset and model details). Moreover, in the SI, we show that models trained on graph-based molecular representations using directed message-passing neural networks37 achieve similar or superior performance compared to numerical descriptors (Fig. S9 and Table S7). This finding demonstrates that the physics-informed property prediction framework is independent of both the specific physics-informed equation and the molecular representation used. Extending this perspective, we further demonstrate that the framework is not limited to pure materials: when applied to mixture viscosity prediction across temperature, the physics-informed model remains effective. This highlights its capacity to couple learned complex chemical information with thermodynamic relationships in multi-body chemical systems (see Section 16 in the SI for additional results).
We then assessed how this approach could be used by researchers and engineers in practice. We quantified the critical impact of accurate temperature-dependent property prediction in the fluid selection, showing that traditional assumptions of constant viscosity can lead to substantial errors in performance estimation and, ultimately, to suboptimal material choices. By integrating physics-informed models into simulation workflows, it becomes possible to make better-informed decisions in materials discovery and engineering design. As materials discovery increasingly requires multi-objective optimization across several properties, the ability to predict multiple temperature-dependent behaviors becomes crucial. The physics-informed framework naturally enables multi-property prediction across both temperatures and chemistries. Equations with accurately predicted coefficients can be seamlessly integrated into physics-based simulations, supporting the discovery and optimization of new fluids for industrial and sustainability applications.
From a broader perspective, the outlined physics-informed machine learning strategy offers a simple, adaptable, and physically grounded method for predicting a wide range of temperature-dependent fluid properties. The process, starting with experimental data, verifying an appropriate physical relationship, and training models to predict coefficients with quantified uncertainty, can be applied across domains. It is not restricted to any particular equation (e.g., Arrhenius, Clausius–Clapeyron, Vogel–Fulcher–Tammann) or molecular representation (e.g., descriptors, graph neural networks), making it a versatile tool for future materials discovery and engineering design.
Future evaluation of the approach would be informative for potential application areas, including ionic liquids, complex electrolytes, and nanofluids. Implementing this methodology in practical thermal management systems requires extending the framework to other temperature-dependent properties, including thermal conductivity and heat capacity. While this work provides a successful proof-of-concept through prototype validation, further testing is necessary before it can be adopted as a standard engineering framework for complex industrial systems.
Methods
Dataset compilation
We compiled a dataset of dynamic viscosity measurements from the National Institute of Standards and Technology (NIST) and previously published studies.44–46 The dataset consists of experimental measurements for 720 organic fluids across a temperature range of 250–550 K. Each chemical, originally identified by its chemical name, was converted to a SMILES string using the Chemical Identifier Resolver (CIR) web-based service.56 These SMILES strings are subsequently input into the Mordred package36 to extract approximately 1600 molecular descriptors, covering 2D features, including basic chemical properties, topological indices, and geometric and polarity-based descriptors. To ensure a balanced representation of data points, temperature-viscosity data points are selected for each chemical using the quantile-median temperature selection (QMTS) method. Molecules with fewer than five viscosity–temperature measurements are excluded. In addition, materials for which physics-based equation fitting is inaccurate are removed. Our analysis indicates that these constitute approximately 3% of the dataset.Viscosity–temperature equations
Looking for a physics-informed equation to represent the dynamic viscosity versus temperature relationship led to a few candidates, most notably the Arrhenius, Vogel–Fulcher–Tammann (VFT), Mauro–Yue–Ellison–Gupta–Allan47 (MYEGA), and Avramov–Milchev57 (AM) equations. Each provides a distinct mathematical form and physical interpretation of how molecular mobility evolves with cooling. The Arrhenius equation assumes a constant activation energy barrier: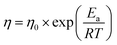 | (1) |
The VFT equation introduces a divergence at a finite temperature below the glass transition, empirically capturing the super-Arrhenius increase in viscosity associated with cooperative molecular dynamics:
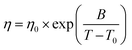 | (2) |
The MYEGA equation modifies this picture by making the activation energy explicitly temperature-dependent, yielding a smooth crossover from Arrhenius-like behavior at high temperatures to super-Arrhenius growth at lower temperatures, without a finite-temperature divergence:
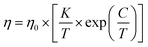 | (3) |
Similarly, the Avramov–Milchev equation links viscosity growth directly to configurational entropy (or free volume), expressing the activation barrier as a power-law function of inverse temperature, with a tunable fragility parameter:
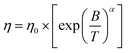 | (4) |
Together, these models represent complementary approaches: from constant-barrier kinetics (Arrhenius), to empirical divergence-based forms (VFT) to entropy-driven frameworks that incorporate temperature-dependent activation energies (MYEGA and AM). We evaluate the validity of the viscosity–temperature equations by fitting each model to the experimental datasets using nonlinear regression. We assess the goodness of fit using the coefficient of determination (R2) and the stability of the fitted coefficients. To assess stability, we apply a resampling approach: we repeatedly perturb each dataset by randomly omitting or resampling data points, refit the model to each perturbed set, and collect the resulting coefficients. We then calculate the standard deviation of these coefficients as a measure of parameter robustness for each material.
Feature engineering
Feature selection is essential for reducing the dimensionality of the feature set, thereby enhancing the model's computational efficiency while retaining the most informative predictors. A backward elimination approach is employed, beginning with the removal of descriptors that have missing values for 80% of compounds. Following this, the correlation matrix is constructed for the feature set to identify pairs of highly correlated features.58 To ensure minimal redundancy, one feature from each highly correlated pair is eliminated, resulting in a feature set where the covariance between any two selected features is below a threshold of 0.9.Subsequently, a variance threshold is applied to remove normalized descriptors with minimal variability. Descriptors with a variance below 0.005 are discarded, effectively reducing the feature set by eliminating low-variance features without significant loss of information. After reducing the feature set to 288 descriptors, three feature selection methods are employed to identify subsets ranging from 20 to 70 features, in increments of 10. These methods include a nonlinear ensemble-based approach (Random Forest),59 a sparse model technique (LASSO-CV),60 and an iterative boosting framework (XGBoost).58
To assess the importance of input features as key sources of information about the compounds, SHAP (SHapley Additive exPlanations) analysis50 is employed. This method facilitates cross-model evaluation by quantifying the contribution of each feature to the prediction of dynamic viscosity in individual models. Additionally, it highlights the interaction between uncertainty in the training data and the feature set, providing deeper insights into model behavior.
Train test splitting strategies
Fitting the Arrhenius equation shows high accuracy for predicting viscosity at unknown temperatures for known chemicals (MAE = 0.09, SRCC = 0.99) when experimental data are available (Fig. S19). However, the focus of this work is on predicting dynamic viscosity for new chemicals, a scenario in which such temperature-dependent data are unavailable. To evaluate model performance under this practically relevant setting, we assess three extrapolative scenarios:Machine learning model development
During the training process of a physics-informed ML model, the input descriptors are chemically driven and independent of thermodynamic conditions. This model features a wide and deep neural network architecture, with a 5D temperature vector bypassing the main training path and introduced at the penultimate layer. At this layer, the feed-forward network is adapted to output the coefficients of the Arrhenius equation for dynamic viscosity. The final layer of the model has fixed weights that replicate the Arrhenius equation, using the predicted frequency factor and activation energy alongside the temperature vector to predict logarithmic dynamic viscosity. The prediction performance is evaluated using mean absolute error (MAE) as the loss function, and the model is optimized through back propagation.The physics-informed model incorporates uncertainty quantification, involving an ensemble of models trained on smaller subsets of the main dataset.61 The optimal size of these training subsets is determined using a learning curve (Fig. S1), identifying the range where the validation loss function plateaus, indicating that further expansion of the training data does not significantly improve model performance.62 Afterwards, we adopt bagging (Bootstrap Aggregating) to construct 20 diverse individual models. Bagging works by training each model on different versions of the training dataset, generated through random sampling with replacement. However, the architecture of the models is the same. This ensures that each model is trained on a different subset of the data, enhancing diversity within the ensembles. Each single model predicts dynamic viscosity indirectly by estimating the coefficients of the Arrhenius equation. The final property of interest is obtained by averaging the outputs of 20 models. To quantify uncertainty, we use the standard deviation of the predicted Arrhenius frequency factor as an indicator, since it is chemistry-dependent and provides a measure of the model's confidence in viscosity predictions. Unlike viscosity, which varies strongly with temperature and can span several orders of magnitude, the frequency factor is not directly correlated with temperature, making it a more robust basis for uncertainty assessment. A material in the test dataset is flagged as “certain” if the standard deviation of the frequency factor remains below a predefined threshold (see the SI, Section S24 for sensitivity analysis of the uncertainty threshold).
Model hyperparameters, such as indicators of model complexity—including the number of hidden layers, neurons per layer, activation functions, learning rate, batch size, and regularization techniques—are optimized using the Optuna hyperparameter tuning framework.63 Each model is optimized individually, leading to scenario-specific hyperparameter configurations. However, in the case of training the physics-informed model, the natural logarithm of the frequency factor is also predicted as a helper output to regularize the coefficients of the Arrhenius equation. This helper output is useful as it helps the frequency factor converge to its actual value, thereby indirectly aiding the model in making meaningful viscosity predictions. Additionally, the weights of main and helper outputs contributing to the loss function prediction are considered as one of the hyperparameters and optimized.
Machine learning model benchmarks
For benchmarking, we employ two different approaches based on previously developed models. In both approaches, temperature is supplied as a direct input to the model—concatenated with chemical descriptors—meaning no physics-informed equation is used. In the first approach, referred to as the isothermal base model, the training dataset includes only one temperature level (here, T3 is used), under the assumption that single-temperature data is available.38 By contrast, the second approach, called the multitemperature base model, is trained on a dataset containing all five temperature levels.15,18Two accuracy matrices are utilized for model performance assessment: MAE as a measure of the differences between predicted and actual values and SRCC measuring the strength and direction of the monotonic relationship between two ranked variables are used as accuracy metrics:
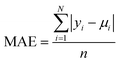 | (5) |
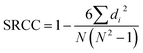 | (6) |
Accuracy measures such as MAE assess only the discrepancy between predicted means and observed values. While informative, such metrics provide no insight into the reliability of predictive uncertainty. Since our ensemble of the models predicts both a mean and an associated standard deviation for each prediction, it is essential to evaluate not only point accuracy but also whether predictive distributions are calibrated and informative. Calibration-oriented metrics are important as they allow us to quantify the alignment between predicted uncertainty and observed variability. We employ four complementary evaluation criteria: (1) Coverage Probability (CP): proportion of true outcomes (yi) falling within nominal prediction intervals (e.g., 90% or 95%) defined around the predictive mean (µi) and standard deviation (σi). Proper calibration is indicated when empirical coverage matches nominal levels:
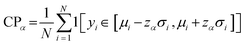 | (7) |
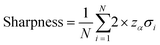 | (8) |
(3) Gaussian Negative Log-Likelihood (NLL): a scoring rule that penalizes both biased mean predictions and miscalibrated variance, thereby combining calibration and sharpness into a single objective.
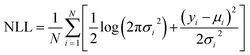 | (9) |
(4) Continuous Ranked Probability Score (CRPS): another scoring rule that measures the distance between the predictive distribution and the observation, robust to outliers and expressed in the same units as the target variable:
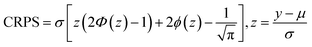 | (10) |
Cooling system simulation
To evaluate the candidate thermal fluids for the cooling device, we require viscosity data across the operational temperature range of 300–390 K. Our physics-informed ML model provides continuous viscosity predictions within this range for all candidates (solid lines in Fig. 6C). To establish a reliable benchmark and validate these predictions, we seek independent reference data. While NIST is our first source of choice, viscosity data are incomplete and inconsistent across the required temperature range for some of the candidate fluids. To fix this limitation, we measure viscosities at three representative temperatures (288.15, 298.15, and 313.15 K). The viscosity is measured through a flow-through resonance quartz sensor (LUVD1, Phase Sensors, Alberta, Canada). The working principle is based on a piezoelectric quartz tuning fork that oscillates when a certain voltage is applied. Compared to vacuum, and depending on the viscosity of the liquid, the oscillation properties (resonance frequency and peak width) of the tuning fork vary.64,65 These variations are directly correlated with the liquid viscosity. The sensor measures the dynamic viscosity at 30 seconds intervals with an accuracy of 5%, and the reported measurements are the result of at least 10 data points (∼5 minutes for a single measurement). We select these three temperatures mainly because they correspond to the practical constraints on heating the fluids.66 Agreement between the ML predictions and the experimental data at these temperatures (shown as markers in Fig. 6C) demonstrates that the predictions can remain reliable across the full operating range.In the next steps, a flow simulation across staggered tube banks is conducted in COMSOL Multiphysics version 6.2 (Fig. 6A) using the selected fluids. The best thermal fluid in terms of efficiency, defined as the ratio of heat dissipated to pump power, is identified out of these thermal fluids under two different scenarios. In the first scenario, dynamic viscosity is considered to be temperature invariant and a constant across the flow, while in the second scenario, dynamic viscosity is predicted at each temperature using the Arrhenius equation developed by the physics-informed model for each unique fluid. The temperature distribution across the geometry varies in the range of 300 K to 390 K, highlighting the importance of accurately accounting for temperature-dependent viscosity.
Author contributions
M. R. K. developed the machine learning workflow with help from S. T. K., B. M., and S. M. M.; experimental work was performed by M. E., H. Y., R. M., M. Z., and D. S.; H. R. and M. Z. performed the liquid system modelling. M. R. K., M. Z., D. S., and S. M. M. conceived the idea and designed the project. All authors contributed to analyzing the data and writing the paper.Conflicts of interest
There are no conflicts to declare.Data availability
Code and data for this project are publicly available. The full codebase is hosted on GitHub at https://github.com/AI4ChemS/thermoML and is released under the MIT License. The dataset is provided as a CSV file in the data directory of the same repository: https://github.com/AI4ChemS/thermoML/tree/master/src/thermoML/data. The archived version of the code and the full datasets used in this study are available on Zenodo at: https://doi.org/10.5281/zenodo.18929020.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5dd00489f.
Acknowledgements
The authors would like to acknowledge funding and technical support from BP Applied Sciences through BP Technology Ventures Ltd. The authors acknowledge support from the Mitacs Accelerate program, University of Toronto's Data Science Intitute and Acceleration Consortium, which receives funding from the Canada first Research Excellence Fund (CFREF), Natural Sciences and Engineering Council of Canada (NSERC), and support through the Canada Research Chairs Program (CRC-2021-00316). D. Sinton holds a Canada Research Chair in Microfluidics and Energy.References
- C. Nieto-Draghi, G. Fayet, B. Creton, X. Rozanska, P. Rotureau, J.-C. de Hemptinne, P. Ungerer, B. Rousseau and C. Adamo, Chem. Rev., 2015, 115, 13093–13164 CrossRef CAS PubMed.
- Y. Wu, Z. Huang, D. Li, H. Li, J. Peng, D. Stroe and Z. Song, Appl. Energy, 2024, 353, 122090 CrossRef CAS.
- S. Zheng, C. Su, X. Yang, Y. Zhang, K. Duan, Y. Zhang, Z. Huang, Y. Zhang, F. Liu and J. Wei, Appl. Therm. Eng., 2025, 126385 CrossRef.
- M. Azarifar, M. Arik and J.-Y. Chang, Appl. Therm. Eng., 2024, 123112 CrossRef.
- L. C. Thomas, TA Instruments, New Castle, DE, 2007 Search PubMed.
- M. Fleck, S. Darouich, J. Pleiss, N. Hansen and M. B. M. Spera, J. Chem. Inf. Model., 2025, 65(8), 3999–4009 CrossRef CAS PubMed.
- J. Götz, M. K. Jackl, C. Jindakun, A. N. Marziale, J. André, D. J. Gosling, C. Springer, M. Palmieri, M. Reck and A. Luneau, Sci. Adv., 2023, 9, eadj2314 CrossRef.
- G. Bradford, J. Lopez, J. Ruza, M. A. Stolberg, R. Osterude, J. A. Johnson, R. Gomez-Bombarelli and Y. Shao-Horn, ACS Cent. Sci., 2023, 9, 206–216 CrossRef CAS PubMed.
- A. Kazemi, M. Zargartalebi and D. Sinton, Energy Environ. Sci., 2024, 17, 813–823 RSC.
- A. E. A. Allen and A. Tkatchenko, Sci. Adv., 2022, 8, eabm7185 CrossRef CAS PubMed.
- J. Yang, L. Tao, J. He, J. R. McCutcheon and Y. Li, Sci. Adv., 2022, 8, eabn9545 CrossRef CAS.
- A. P. Soleimany, A. Amini, S. Goldman, D. Rus, S. N. Bhatia and C. W. Coley, ACS Cent. Sci., 2021, 7, 1356–1367 CrossRef CAS.
- E. M. Rajaonson, M. R. Kochi, L. M. M. Mendoza, S. M. Moosavi and B. Sanchez-Lengeling, arXiv, 2025, preprint, arXiv:2506.12231, DOI:10.48550/arXiv.2506.12231.
- A. Jain, R. Gurnani, A. Rajan, H. J. Qi and R. Ramprasad, npj Comput. Mater., 2025, 11, 42 CrossRef CAS.
- P. Panwar, Q. Yang and A. Martini, J. Chem. Inf. Model., 2023, 64(7), 2760–2774 CrossRef PubMed.
- Q.-S. Liu, J. Liu, X.-X. Liu and S.-T. Zhang, J. Chem. Thermodyn., 2015, 90, 39–45 CrossRef CAS.
- M. Mohan, K. D. Jetti, S. Guggilam, M. D. Smith, M. K. Kidder and J. C. Smith, ACS Sustain. Chem. Eng., 2024, 12, 7040–7054 CrossRef CAS.
- A. K. Chew, M. Sender, Z. Kaplan, A. Chandrasekaran, J. Chief Elk, A. R. Browning, H. S. Kwak, M. D. Halls and M. A. F. Afzal, J. Cheminf., 2024, 16, 31 Search PubMed.
- L.-Y. Yu, G.-P. Ren, X.-J. Hou, K.-J. Wu and Y. He, ACS Cent. Sci., 2022, 8, 983–995 CrossRef CAS PubMed.
- B. Sheng, Y. Zhao, X. Dong, H. Lu, W. Dai, H. Guo and M. Gong, J. Mol. Liq., 2021, 343, 117483 CrossRef CAS.
- X. Kang, X. Liu, J. Li, Y. Zhao and H. Zhang, Ind. Eng. Chem. Res., 2018, 57, 16989–16994 CrossRef CAS.
- K. R. Aglawe, R. K. Yadav and S. B. Thool, in Proceedings of the International Conference on Industrial and Manufacturing Systems (CIMS-2020) Optimization in Industrial and Manufacturing Systems and Applications, Springer, 2022, pp. 389–408 Search PubMed.
- A. Y. S. Eng, C. B. Soni, Y. Lum, E. Khoo, Z. Yao, S. K. Vineeth, V. Kumar, J. Lu, C. S. Johnson and C. Wolverton, Sci. Adv., 2022, 8, eabm2422 CrossRef CAS PubMed.
- S. Ament, M. Amsler, D. R. Sutherland, M.-C. Chang, D. Guevarra, A. B. Connolly, J. M. Gregoire, M. O. Thompson, C. P. Gomes and R. B. Van Dover, Sci. Adv., 2021, 7, eabg4930 CrossRef CAS PubMed.
- J. Deng, Z. Yang, H. Wang, I. Ojima, D. Samaras and F. Wang, Nat. Commun., 2023, 14, 6395 CrossRef CAS PubMed.
- F. Jirasek, R. A. S. Alves, J. Damay, R. A. Vandermeulen, R. Bamler, M. Bortz, S. Mandt, M. Kloft and H. Hasse, J. Phys. Chem. Lett., 2020, 11, 981–985 CrossRef CAS PubMed.
- S. Mehdi and P. Tiwary, Nat. Commun., 2024, 15, 7859 CrossRef CAS PubMed.
- A. M. Schweidtmann, J. G. Rittig, J. M. Weber, M. Grohe, M. Dahmen, K. Leonhard and A. Mitsos, Comput. Chem. Eng., 2023, 172, 108202 CrossRef CAS.
- J. G. Rittig and A. Mitsos, Chem. Sci., 2024, 15, 18504–18512 RSC.
- E. I. S. Medina, S. Linke, M. Stoll and K. Sundmacher, Digit. Discov., 2023, 2, 781–798 RSC.
- F. Jirasek and H. Hasse, Annu. Rev. Chem. Biomol. Eng., 2023, 14, 31–51 CrossRef CAS PubMed.
- M. Raissi, P. Perdikaris and G. E. Karniadakis, J. Comput. Phys., 2019, 378, 686–707 CrossRef.
- T. Specht, M. Nagda, S. Fellenz, S. Mandt, H. Hasse and F. Jirasek, Chem. Sci., 2024, 15, 19777–19786 RSC.
- J. G. Rittig, K. C. Felton, A. A. Lapkin and A. Mitsos, Digit. Discov., 2023, 2, 1752–1767 RSC.
- J. N. Al-Dawsari, A. Bessadok-Jemai, I. Wazeer, S. Mokraoui, M. A. AlMansour and M. K. Hadj-Kali, J. Mol. Liq., 2020, 310, 113127 CrossRef CAS.
- H. Moriwaki, Y.-S. Tian, N. Kawashita and T. Takagi, J. Cheminf., 2018, 10, 1–14 Search PubMed.
- E. Heid, K. P. Greenman, Y. Chung, S.-C. Li, D. E. Graff, F. H. Vermeire, H. Wu, W. H. Green and C. J. McGill, J. Chem. Inf. Model., 2023, 64, 9–17 CrossRef PubMed.
- V. Goussard, F. Duprat, J.-L. Ploix, G. Dreyfus, V. Nardello-Rataj and J.-M. Aubry, J. Chem. Inf. Model., 2020, 60, 2012–2023 CrossRef CAS.
- D. Toghraie, S. M. Alempour and M. Afrand, J. Magn. Magn. Mater., 2016, 417, 243–248 CrossRef CAS.
- A. Szczęsna-Chrzan, M. Vogler, P. Yan, G. Z. Żukowska, C. Wölke, A. Ostrowska, S. Szymańska, M. Marcinek, M. Winter and I. Cekic-Laskovic, J. Mater. Chem. A, 2023, 11, 13483–13492 RSC.
- M. Taghizadehfard, S. M. Hosseini and M. M. Alavianmehr, J. Mol. Liq., 2021, 325, 115048 CrossRef CAS.
- N. N. Matsuzawa, H. Maeshima, K. Hayashi, T. Ando, M. A. F. Afzal, K. Marshall, B. J. Coscia, A. R. Browning, A. Goldberg and M. D. Halls, Chem. Mater., 2024, 36, 11706–11716 CrossRef CAS.
- Z. Tariq, A. Hassan, U. Bin Waheed, M. Mahmoud, D. Al-Shehri, A. Abdulraheem and E. M. A. Mokheimer, J. Energy Resour. Technol., 2021, 143, 092801 CrossRef CAS.
- The National Institute of Standards and Technology (NIST), Thermophysical Properties of Fluid Systems, 2010 Search PubMed.
- C. L. Yaws, Transport properties of chemicals and hydrocarbons, William Andrew, 2014 Search PubMed.
- D. S. Viswanath, T. K. Ghosh, D. H. L. Prasad, N. V. K. Dutt and K. Y. Rani, Viscosity of liquids: theory, estimation, experiment, and data, Springer Science & Business Media, 2007 Search PubMed.
- D. R. Cassar, Acta Mater., 2021, 206, 116602 CrossRef CAS.
- J. Bradshaw, A. Zhang, B. Mahjour, D. E. Graff, M. H. S. Segler and C. W. Coley, ACS Cent. Sci., 2025, 11(4), 539–549 CrossRef CAS PubMed.
- A. Goyal and Y. Bengio, Proc. R. Soc. A, 2022, 478, 20210068 CrossRef.
- S. M. Lundberg and S.-I. Lee, Adv. Neural Inf. Process. Syst., 2017, 30 DOI:10.48550/arXiv.1705.07874.
- H. Rezaei, M. Rajabi-Kochi, M. Ebrahimiazar, M. Zargartalebi, S. M. Moosavi and D. Sinton, J. Energy Storage, 2025, 140, 119036 CrossRef CAS.
- C. D. M., V. Diky, A. Y. Smolyanitsky, A. Bazyleva, R. D. Chirico, J. W. Magee, Y. Paulechka, A. F. Kazakov, S. A. Townsend, E. W. Lemmon, M. D. Frenkel and K. G. Kroenlein, NIST/Thermodynamics Research Center (TRC), preprint, https://app.knovel.com/hotlink/toc/id:kpLTI00007/nist-standard-reference/nist-standard-reference.
- O. Großmann, D. Bellaire, N. Hayer, F. Jirasek and H. Hasse, Digit. Discov., 2022, 1, 886–897 RSC.
- A. F. F. Dias, I. Portugal, J. P. S. Aniceto and C. M. Silva, Chem. Eng. J., 2024, 153274 CrossRef CAS.
- R. Qiu, L. Li, L. Wu, E. Agathokleous, C. Liu and B. Zhang, J. Hydrol., 2022, 610, 127989 CrossRef.
- M. Sitzmann, I. V. Filippov and M. C. Nicklaus, Chemical Structure Lookup Service, 2007, https://cactus.nci.nih.gov/ Search PubMed.
- B. N. Galimzyanov and A. V Mokshin, J. Non-Cryst. Solids, 2021, 570, 121009 CrossRef CAS.
- J. Gong, S. Chu, R. K. Mehta and A. J. H. McGaughey, npj Comput. Mater., 2022, 8, 140 CrossRef CAS.
- A. L. Teixeira, J. P. Leal and A. O. Falcao, J. Cheminf., 2013, 5, 1–15 Search PubMed.
- C. Cui and D. Wang, Inf. Sci., 2016, 372, 505–517 CrossRef.
- H. Al Osman and S. Shirmohammadi, IEEE Instrum. Meas. Mag., 2021, 24, 23–27 Search PubMed.
- M. A. Ganaie, M. Hu, A. K. Malik, M. Tanveer and P. N. Suganthan, Eng. Appl. Artif. Intell., 2022, 115, 105151 CrossRef.
- T. Akiba, S. Sano, T. Yanase, T. Ohta and M. Koyama, in Proceedings of the 25th ACM SIGKDD international conference on knowledge discovery & data mining, 2019, pp. 2623–2631 Search PubMed.
- R. Blaauwgeers, M. Blazkova, M. Človečko, V. B. Eltsov, R. de Graaf, J. Hosio, M. Krusius, D. Schmoranzer, W. Schoepe and L. Skrbek, J. Low Temp. Phys., 2007, 146, 537–562 CrossRef CAS.
- L. Matsiev, in 2006 IEEE Ultrasonics Symposium, 2006, pp. 884–887 Search PubMed.
- H. Shabgard, M. J. Allen, N. Sharifi, S. P. Benn, A. Faghri and T. L. Bergman, Int. J. Heat Mass Tran., 2015, 89, 138–158 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |