 Open Access Article
Open Access ArticleSiamese graph neural networks for melting temperature prediction of molten salt eutectics
Nila
Mandal
 ,
James
Maniscalco
,
Mark
Aindow
,
James
Maniscalco
,
Mark
Aindow
 and
Qian
Yang
and
Qian
Yang
 *
*
University of Connecticut, Storrs, CT, USA. E-mail: qyang@uconn.edu
First published on 2nd March 2026
Abstract
High-throughput screening enabled by structure–property prediction models is a powerful approach for accelerating materials discovery. However, while machine learning of structure–property models have become widespread, its application to mixtures remains limited due to increased complexity and the scarcity of available data. Machine learning methods for high-throughput screening of eutectic mixtures have been proposed in recent years, but there remain challenges due to the lack of diverse, open-access datasets and the need for feature engineering based on chemical knowledge. To overcome these limitations, we propose a method using Siamese graph neural networks trained solely on structural information, without requiring any prior chemical descriptors, to predict eutectic melting temperatures. We demonstrate on a dataset of molten salt eutectics that this approach can reach similar performance to chemistry-based models that require significantly more prior knowledge. We show that lower-order mixtures may be used to augment data on higher-order mixtures. Interestingly, our model trained on inorganic molten salts seems to learn information about the ideal mixture model. We also evaluate the efficacy of using our inorganic molten salt model for transfer learning with a variety of organic eutectic mixtures.
1 Introduction
Mixture properties, such as eutectic melting temperatures, are critical for materials design. For example, molten salt eutectic mixtures have the potential to be highly effective in several sustainable energy applications. They have gained attention in recent years in research on battery electrolytes,1 solar power,2 thermal energy storage materials,2 and nuclear reactors.3These important applications are hindered by drawbacks of existing computational and experimental methods for determining mixture properties, which are often slow and resource-intensive. A machine learning approach for high-throughput screening of eutectic melting temperatures is thus highly desirable. However, such approaches are often hindered by the lack of diverse, publicly available experimental datasets and the computational cost of simulation methods for generating data. As a result, most recent studies rely on classical machine learning algorithms with highly informative but expensive engineered features that can be effectively trained on smaller datasets. In this work, we demonstrate that a deep learning-based approach leveraging only structural information is sufficient to train effective models for predicting eutectic melting temperatures of binary molten salt mixtures, using a dataset of 2244 data points. Our approach utilizes Siamese graph neural networks4 and incorporates ideas from Janossy pooling5 to effectively handle mixtures. We also demonstrate that individual components' melting point data can be used to augment the mixture datasets, and produce models that can extrapolate from inorganic to organic materials (Fig. 1).
Our architecture includes several novel contributions, including the ability to learn melting points of binary eutectic mixtures from structural data alone, and the ability to learn a model correlated with the ideal thermodynamic model without requiring single component melting points or enthalpy values, or eutectic compositions xe values. Although single component melting points are not required, we also demonstrate that by optionally using them as additional datapoints for data augmentation rather than features, we can further improve our model's predictive performance as well as achieve good predictions for previously unknown single component melting temperatures.
2 Background
2.1 Melting point prediction
A material's melting point is an important property to consider for any application requiring a solvent or electrolyte, and melting point prediction has long been a goal of computational chemistry. Molecular dynamics simulations can be used to estimate materials' melting points; however, simulations of a solid material melting tend to result in overestimates of melting point, while simulations of a liquid material solidifying tend to underestimate the melting point. In addition, defects can cause materials to melt at lower temperatures than would be expected of a perfect material as represented in a simulation.6Much work has been done on melting point prediction for ionic liquids, including comparisons of methods such as k-nearest neighbor regression, gradient boosting, random forests, support vector machines, and graph neural networks.7,8 Structural descriptors such as molecule fingerprints, Coulomb matrices, or other engineered features can be used to improve prediction accuracy. Fingerprints which encode functional groups have been found to be particularly effective.9 However, Acar et al.10 argues that all of these fingerprint methods are expensive to compute, and not practical for either large datasets or datasets which include large molecules. Instead, they test a very simple fully connected neural network with one dropout layer, and compare the correlation between melting point and the various engineered features they use. Although these engineered features may be less computationally complex than the fingerprint methods evaluated by Low et al.,9 expertise in chemistry is necessary in order to understand and generate these descriptors for any dataset.
2.2 Eutectic materials
Eutectic mixtures are mixtures which have a lower melting point than either of its individual mixture components; as such, they are often desirable for applications that require low-melting point materials. In our work we focus on binary eutectics. We refer to a mixture's minimum eutectic melting point as Te, and refer to the mixture ratio as xe (Fig. 2).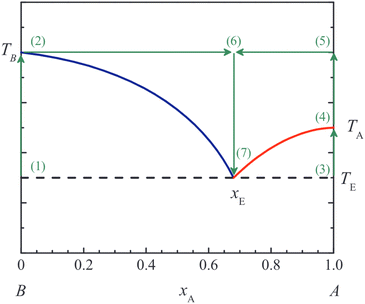 | ||
| Fig. 2 From ref. 11, a phase diagram illustrating the relationships between Ta, Tb, xe, and Te. Eutectic mixtures are mixtures with a melting point lower than that of the individual components of the mixture. The mixture proportion with the lowest possible melting temperature is referred to as xe, and that melting temperature is referred to as Te.† | ||
Determining eutectic melting points is challenging for materials screening because it is infeasible to manually test every possible mixture of candidate components. Research into data-driven approaches to Te prediction has been significantly hindered by the lack of large, publicly available datasets. Although published eutectic datasets exist, they are often small (100 examples or less) and focused on a highly specific family of materials (for example, fatty acids or organic explosives). Due to these limited dataset sizes, machine learning approaches to eutectic melting point prediction typically rely on highly informative chemical descriptors such as individual melting points, enthalpies of melting, and xe values, which are themselves difficult to obtain or compute.
If all of those values are available, Te can be computed using the ideal thermodynamic model (ITM). However, if any of these values are unavailable in existing literature for a given material, it is impossible for the ITM to compute Te for any mixture which includes that material. Even when all necessary values are known, the predicted Te values from the ITM are not perfect. For example, Ravichandran et al.1 found that for their data, the ITM predictions had a root mean squared error (RMSE) of 86.6 K. The correlation between Te and the individual melting points of the mixture components has been used to estimate how much melting point depression the eutectic mixture will have;12 however, even this extremely rough estimation requires knowledge of the melting points of each individual component, which is itself challenging to obtain for new materials exploration.
Ravichandran et al.1 approach eutectic Te prediction for molten salts by using an ensemble model composed of the ideal thermodynamic model, a gradient boosting model, and a Roost model. Of these, the Roost model is the only part of the ensemble which does not require expensive engineered features. The ensemble model, referred to in the text as the “mean model,” takes the average of the three models' predictions to determine its final output. They consider mixtures of up to six eutectic components, and their data is a subset of the experimentally-determined molten salt eutectic mixtures, proportions, and Te values compiled by Janz et al.13 Our work considers a different subset of mixtures from the same compilation.
There are several smaller datasets we are aware of for different families of organic eutectic mixtures. Kahwaji et al.11 published a set of fatty acid eutectics as well as a computational method that they designed to support eutectic phase change materials exploration. Guendouzi et al.14 published a set of melting points for individual fatty acids, which overlap with Kahwaji's individual components. Luu et al.15 and Lavrinenko et al.16 both published sets of deep eutectic solvents drawn from literature, with substantial overlap between the mixtures they compiled. More recently, databases for binary and ternary DES and corresponding ML models have also been developed.17
2.3 Graph neural networks
Graph neural networks provide a compelling approach to structure–property prediction because they can learn directly from structural data without the need for complex engineered features. As such, they have become a popular way to learn structure–property relationships for molecules and materials. A graph neural network (GNN) takes as input one or more graphs, which are made up of nodes and edges, and can solve problems either at the node level or the graph level. For example, predicting the melting point of a single component molecule from its graph structure is a graph-level problem. Most GNNs follow the message-passing framework,18 in which each layer of the neural network computes a function of each node and its one-hop neighbor's features, including edge features if present. After one layer, each node has aggregated a representation of its entire one-hop neighborhood; after two layers each node has aggregated a representation of its two-hop neighborhood, and so on.19AttentiveFP,20 which is used in this work, is one such message passing graph neural network architecture, with the addition of an attention mechanism to help represent intramolecular interactions beyond each node's one-hop neighborhood. The architecture has a series of layers for atom embedding, followed by a series of layers for molecule embedding. For each graph, AttentiveFP generates a “state node” which is connected to every node in the graph; this is used to aggregate information from the whole graph to compute a learned representation of the entire molecule.
A significant challenge in GNNs is identifying an appropriate pooling method. Graph pooling should generally be order-invariant, because the nodes in a graph typically are not meant to relate to each other in any sequential ordering. However, simple order-invariant methods like mean or max pooling fail to capture important structural information that is not explicitly represented by node features.21 Learned pooling methods, such as hierarchical pooling or Janossy pooling5 may produce better pooled representations.
In order to handle mixtures, we incorporate ideas from Janossy pooling,5 which refers to building permutation-invariant functions by taking the average, or an approximation of the average, of a permutation-sensitive function's output over every possible permutation. In the context of graph neural networks, this refers to permutations of the nodes in the graph. In the general case, an approximation method is necessary for tractability; as such, Murphy et al.5 present three categories of approximation methods. However, in problem domains where the number of possible permutations per input is small, as in this work, it may be reasonable to compute this naively.
2.4 Siamese neural networks
In a Siamese neural network, both elements of a pair are passed through a series of identical neural network layers; all weights are shared between both branches. This means that, at each layer, the weights and biases of the two neural networks are exactly equal. This type of neural network architecture is an intuitive solution to pairwise problems.Siamese neural networks4 were originally developed to measure the similarity or difference between pairs of input data. Since determination of binary eutectic Te values is an inherently pairwise problem, and Te and xe are correlated with the difference between the melting points of the mixture components,12 Siamese neural networks are well-suited to our problem.
Siamese neural networks have been applied to many problems in chemistry and materials science due to their efficacy in one- or few-shot learning and similarity comparison. In particular, they have been widely used to screen molecules for drug discovery, predict molecule solubility or toxicity, and predict drug response similarity.4,22–24 Siamese graph neural networks have also been used with engineered graph level features for drug-responsiveness prediction on cancer treatments.25
3 Methods
3.1 Graph representation and features
In this work, we develop an order-invariant architecture to predict eutectic mixture Te values with minimal input data. First, we generate a graph representation of each molecule, based on structure data drawn from PubChem. Each atom in a molecule is represented as a node in a graph, with node features consisting of the group and period number of the corresponding atom's element. Bonds are represented by edges in the graph; we chose not to include edge features. No additional node- or graph-level features are needed. Our methods do not require any engineered features, and do not require attributes such as individual components melting points or mixture proportions; this means that these methods can be applied to any eutectic data for which the individual components' structures are known (Table 1).3.2 Inorganic data
We provide a curated dataset of 913 individual molecules paired into 2244 molten salt eutectic pairs and their corresponding experimentally determined melting points. Of those individual components, we have individual melting point values for 149 of them. These mixture pairs and Te values were drawn from Janz et al.,13 which compiled several thousand experimentally-determined molten salt eutectic mixtures, proportions, and Te values from publications prior to 1978. We manually transcribed the molecules listed in this text, and cross-referenced them with PubChem27 to generate graph representations of each mixture component's solid state structure. The box-and-whisker plot in Fig. 4 depicts the first quartile, median, and third quartile of melting points for the individual components and the mixtures, showing that the dataset is skewed towards lower temperatures. We take a stratified partition of the pairs and individual components to generate a 10-fold cross validation set and a test set.3.3 Organic data
We use a smaller organic dataset comprised of 88 individual components and 239 unique deep eutectic solvent (DES) mixtures drawn from ref. 15–17 for transfer learning experiments to examine our model's ability to extrapolate from inorganic to organic data (Fig. 5). This dataset consisting of both individual component melting temperatures and binary mixture eutectic melting temperatures is partitioned into a 5-fold cross-validation set and a test set for our experiments.We further use a set of fatty acids,11 a set of explosives,12 and a set of quinones28 which form eutectic mixtures as small test sets for evaluating zero-shot transfer from inorganic to organic eutectics (Table 2).
| Dataset | Tasks | Number of data points | Avg. atoms per molecule |
|---|---|---|---|
| Molten salts A,B data | Training from scratch | 2244 | 5 |
| Molten salts A,A data | Training from scratch | 149 | 5 |
| DES A,B data | Transfer learning | 239 | 24 |
| DES A,A data | Transfer learning | 88 | 24 |
| Fatty acids | Inference only | 102 | 43 |
| Explosives | Inference only | 74 | 21 |
| Quinones | Inference only | 26 | 18 |
3.4 Handling noisy duplicates
Several of the mixture pairs in this dataset have duplicate entries with differing values. Some of these are due to the mixture pair forming multiple eutectic points, whereas others are noisy experimental data. Given the advancements in experimental technologies between 1978 and 2023, some of these noisy mixtures may now have more definitive reference values in other literature.For the purposes of this work, we focus on the minimum Te value for each unique mixture pair represented, because we are interested in predicting the lowest melting point achievable by any given pair. All temperature values presented here are in Kelvin unless otherwise stated.
After filtering duplicates, we take a stratified sample of mixtures (which we refer to as A,B pairs) and individual components (A,A pairs) into the training and test set (90% and 10%, respectively). We then divide the molten salt training set into 10 stratified folds for cross-validation.
3.5 Architecture
For each eutectic mixture pair, both molecules' graphs are passed through a set of identical AttentiveFP GNN layers; these combine a message passing GNN framework with an attention mechanism to account for intramolecular interactions beyond each node's one-hop neighborhood.20 These layers output a feature embedding for each molecule (referred to as F(A) and F(B) in Fig. 6).The interactions between the two molecules in the mixture influence its melting temperature, so rather than computing a similarity or difference metric between the two, we concatenate the two molecule representations so that the following layers can learn a function of the molecules' interactions. By concatenating A and B, it is possible for the fully connected layers in the Siamese branches to learn something about the interactions between the two molecules, as opposed to only learning their difference.
However, concatenating them implies an ordering, and because we are not incorporating information about individual melting temperatures or mixture proportions, we cannot enforce a meaningful ordering between the two. We address this by utilizing the central idea from Janossy pooling to enforce order invariance by concatenating the representations in every possible ordering, i.e. (A, B) and (B, A). One might extend to higher-order mixtures by implementing a permutation-sampling approximation of Janossy pooling, selecting a value n as the maximum number of permutations to be sampled per mixture.
These concatenated molecule representations are passed through Siamese branches consisting of fully connected layers, with dropout layers in between. The output of these branches are mean-pooled to generate a final order-invariant representation of the mixture pair, which is then passed through the output layer to predict Te. A diagram of the architecture is depicted in Fig. 6, and all selected hyperparameters and optimization methods are detailed in Table 3.
| Hyperparameter | Selected value |
|---|---|
| Batch size | 256 |
| Learning rate | 0.001 |
| LR Scheduler start factor | 0.350 |
| LR Scheduler iterations | 481 |
| Embedding size | 128 |
| Hidden channels | 256 |
| AttentiveFP layers | 6 |
| AttentiveFP timesteps | 4 |
| Dropout 1 | 0.383 |
| Fully connected layer 1 size | 8 |
| Dropout 2 | 0.002 |
| Fully connected layer 2 size | 8 |
3.6 Transfer learning
Because we have much less organic data than we do inorganic data, rather than training a model from random initialization for our organic datasets, we use transfer learning to see if our learned molecule features from AttentiveFP in the inorganic model can be applied to eutectic melting point prediction for organic molecules. We extract features from the AttentiveFP layers for the organic DES molecules, perform principal component analysis (PCA) to reduce their dimensionality, and then train a kernel ridge regression (KRR) model for DES eutectics on this reduced set of features.4 Experiments and results
First, we perform experiments training our architecture on binary molten salt eutectic mixtures and molten salt individual components. The eutectic datasets we use come from ref. 11, 13 and 15–17. Then we perform transfer learning experiments with organic eutectic datasets to see how well our model can extrapolate. We evaluate model performance in terms of root mean squared error (RMSE), mean absolute error (MAE), mean absolute percentage error (MAPE), and coefficient of determination (R2). The DES, fatty acids, explosives, and quinones datasets all consist of organic molecules, whereas our main dataset of molten salts is inorganic. All molecules' atom and bond structures, and melting points for the 149 molten salt individual components, are sourced from Pubchem.27 The data from Lavrinkenko et al.16 also includes individual component melting points.4.1 Molten salts experiments
The first experiment is to train a model on the molten salt eutectics. We use Bayesian Optimization with Hyperband (BOHB)29 for hyperparameter tuning. This method combines two existing hyperparameter optimization methods, Bayesian optimization and Hyperband, in order to achieve strong performance, scalability, and effective use of parallel resources, where the two methods individually had significant trade-offs between these.We train some models using only the molten salt eutectic mixtures, which we refer to as A,B pairs. We also train some models using both the molten salt eutectic mixtures and the individual components for which we have melting point values, in order to test whether augmenting the dataset with these individual components may improve model performance. The individual components are represented as pairs in which both components are identical, which we call A,A pairs. We compare the performance of models trained in these two ways (Fig. 7), and evaluate whether the individual component data is useful for data augmentation.
4.2 Molten salts results
Overall, our model is able to achieve a R2 of 0.93 in predicting eutectic melting temperatures for binary molten salt eutectics. Models trained with A,A pairs included in the training data perform better than those trained only on binary mixtures. Although the performance difference on the full test set is small, the difference appears much more significant when looking at the subset of data with Te ≤ 500 K, where we achieve an RMSE of 59.53 K. We emphasize this subset because mixtures with low melting points are of greatest interest when screening eutectic materials. The training set only included 134 individual component data points, so it seems unlikely that this improvement in performance can be attributed entirely to the greater quantity of data. This suggests future experiments in using lower-order mixture data to augment training data on higher-order mixtures, and vice versa.We emphasize that these single component melting points are used as data points, not features. When we compare these results to the exact same architecture and feature representation trained on A,B pairs only, we see that the gain in performance is approximately 4.5° on the overall set, and 12.5° in the low temperature subset. Table 4 shows that the model trained on A,A and A,B pairs have improved performance on all test sets compared to the model trained only on A,B pairs.
| Dataset | Model | RMSE | MAE | MAPE | R 2 | Std. dev. | Baseline MAE | Baseline MAPE |
|---|---|---|---|---|---|---|---|---|
| Full dataset from ref. 1* | ITM | 86.6 | — | — | — | — | — | — |
| Test set from ref. 1* | Mean model | 65.4 | — | — | — | — | — | — |
| Random partition from ref. 1 | A,B only, trained on1 | 95.45 ± 13.76 | 68.90 ± 10.29 | 0.09 | 0.92 | 309.86 | 220.23 | 0.85 |
| A,B test set | A,A + A,B | 103.08 ± 20.05 | 73.04 ± 9.09 | 0.10 | 0.93 | 369.9 | 267.5 | 0.46 |
| A,B test set | A,B only | 107.55 ± 11.79 | 77.74 ± 9.42 | 0.11 | 0.92 | 369.9 | 267.5 | 0.46 |
| A,B where Te < 500 K | A,A + A,B | 59.53 ± 10.33 | 47.73 ± 9.27 | 0.13 | 0.60 | 78.41 | 63.11 | 0.19 |
| A,B where Te < 500 K | A,B only | 72.04 ± 20.03 | 49.76 ± 15.03 | 0.15 | 0.48 | 78.41 | 63.11 | 0.19 |
| A,A test set | A,A + A,B | 231.61 ± 86.17 | 189.56 ± 69.85 | 0.44 | 0.89 | 629.34 | 443.35 | 0.81 |
| A,A test set | A,B only | 372.26 ± 169.92 | 275.45 ± 129.39 | 0.67 | 0.64 | 629.34 | 443.35 | 0.81 |
As shown in Fig. 8, our model trained on A,B and A,A pairs also achieves prediction on individual components' melting points with a R2 of 0.89, and a RMSE of 231.61 K. In the context of the standard deviation of the A,A dataset, this prediction error is one-third of the RMSE of a baseline trivial model (the best possible constant model), indicating that despite the relatively large numerical value of the RMSE, the model has learned significant predictive information. This suggests that by including only small quantities of A,A data in training, the final model can learn to estimate individual component melting points while also achieving improved prediction performance on A,B mixtures.
A recent work by Ravichandran et al.1 also used subsets of Janz et al.13 to train machine learning models for eutectic melting temperature prediction; their selection of binary mixtures is a subset of those used in our work. While their experiments resulted in lower RMSE values than ours, their proposed methods require all individual component melting points and enthalpies (ITM method), manually curated engineered features (GBM method), or xe values (Roost method), none of which are required by our method. For comparison, took a random training and test partition of the binary mixtures used in their dataset, and trained our architecture on that partition using only structural information. The results, as shown in Table 4, indicate that our model can achieve performance close to the ITM without requiring individual component melting points and enthalpies.
We note that differences in performance may also be in part due to the difference in our datasets; they considered a narrower selection of binary mixtures than ours, and included other n-ary mixtures which we did not.
4.3 Deep eutectic solvents transfer learning experiments
We next examine the trained model's ability to extrapolate to organic mixtures. We use 88 individual components and 239 unique mixtures drawn from ref. 15–17, to partition into training and test sets for transfer learning experiments. These data sources included some fatty acids, however we excluded these from the dataset because we planned to test extrapolation to a different fatty acid dataset. The training set had 78 single components and 190 mixtures; the test set had 10 single components and 49 mixtures.First, we select the molten salts model which performs best on our DES training set. Organic training data including both A,A and A,B pairs were input into our architecture, and then the molecule features output by the AttentiveFP layers (F(A) and F(B)) were extracted. We scale the features and then use (PCA) to reduce them from 256 features per molecule to 5 features per molecule. We then concatenate each molecule pair in both possible orders, and fit a kernel ridge regression model to the data. We use a grid search to determine the optimal kernel ridge regression hyperparameters. We evaluate the best model on the DES test set, as well as evaluate zero-shot predictions on other smaller organic datasets.
4.4 Deep eutectic solvents results
The zero-shot predictions from the molten salt A,A + A,B model chosen for transfer learning on organic A,B pairs can be seen in Fig. 9–11. | ||
| Fig. 11 Prediction results on fatty acids. (a) presents 0-shot predictions from the model trained on molten salts, while (b) presents predictions after extracting DES features from the A,A + A,B model, fitting a kernel ridge regression model, and then testing on fatty acids. Models from our experiments frequently made predictions on fatty acid eutectic mixtures which had a strong linear correlation with the target values. Mixtures including oleic acid consistently “branched” off from the rest of the group. This data was computed using an ideal thermodynamic model by Kahwaji et al.11 This suggests our architecture is able to learn a function correlated with the ideal thermodynamic model without having information about individual components' melting points or enthalpies of melting. (a) Predictions from molten salts model. (b) Predictions from transfer learning KRR with DES. | ||
Some models trained on molten salts are able to assign organic materials to the correct average melting temperatures of their corresponding molecule family (e.g. fatty acids) despite having never seen an organic molecule in training. These models are also able to assign families of closely-related molecules to similar values; for example, all quinone mixtures receive similar predictions, and fatty acid mixtures' predictions form distinct patterns. This indicates that our model is learning similar feature embeddings for mixtures that we expect to be similar to one another. Therefore, the molecule representation learned by the GNN layers of our architecture from inorganic data can be used for transfer learning to organic DES data to achieve improved predictions on DES mixtures. However, our KRR model transfers less well to both the explosives and quinone datasets.
Mixtures from the same family of materials have predictions that are close together; fatty acids, in particular, seem to follow a distinct trend. This suggests that mixtures that belong to the same family are being assigned similar learned feature embeddings; these are the molecule graph features that we extract from our GNN layers. Then we scale, apply PCA, and fit the kernel ridge regression model.
Through this process, we are able to predict DES Te values with an RMSE of 37.57 K. We are also able to predict melting temperatures of the individual components from the DES dataset with an RMSE of 67.30 K (Table 5).
| Dataset | RMSE | MAE | MAPE | R 2 | Std. dev. | Baseline MAE | Baseline MAPE |
|---|---|---|---|---|---|---|---|
| 0-shot prediction from best molten salt model | |||||||
| DES test set | 90.26 ± 16.20 | 71.64 ± 16.33 | 0.25 | 0.05 | 69.96 | 51.7 | 0.12 |
| Fatty acids test set | 43.97 ± 5.23 | 36.73 ± 5.02 | 0.12 | 0.29 | 20.15 | 17.25 | 0.06 |
| Explosives12 | 63.19 ± 13.31 | 51.32 ± 10.31 | 0.14 | 0.04 | 35.32 | 25.73 | 0.07 |
| Quinones28 | 75.31 ± 13.22 | 67.21 ± 13.34 | 0.21 | 0.33 | 19.62 | 12.95 | 0.04 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||||
| 0-shot prediction from molten salt model chosen for transfer learning | |||||||
| DES test set | 55.79 ± 13.25 | 40.58 ± 10.84 | 0.13 | 0.0 | 69.96 | 51.7 | 0.12 |
| Fatty acids test set | 20.73 ± 2.87 | 16.56 ± 2.45 | 0.06 | 0.38 | 20.15 | 17.25 | 0.06 |
| Explosives12 | 66.69 ± 12.41 | 56.37 ± 8.89 | 0.16 | 0.03 | 35.32 | 25.73 | 0.07 |
| Quinones28 | 31.30 ± 10.02 | 24.97 ± 8.16 | 0.07 | 0.01 | 19.62 | 12.95 | 0.04 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||||
| KRR transfer learning predictions | |||||||
| DES test set | 37.57 ± 4.55 | 31.21 ± 4.29 | 0.11 | 0.41 | 69.96 | 51.7 | 0.12 |
| DES AA test set | 67.30 ± 24.77 | 47.17 ± 20.09 | 0.12 | 0.62 | 77.05 | 61.32 | 0.17 |
| 0-shot prediction from KRR model | |||||||
| Fatty acids test set | 18.11 ± 2.64 | 13.34 ± 1.73 | 0.05 | 0.39 | 20.15 | 17.25 | 0.06 |
| Explosives12 | 75.53 ± 7.80 | 62.91 ± 7.47 | 0.17 | 0.02 | 35.32 | 25.73 | 0.07 |
| Quinones28 | 48.60 ± 5.01 | 44.08 ± 5.36 | 0.14 | 0.0 | 19.62 | 12.95 | 0.04 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||||
| Other works for comparison | |||||||
| Odegova et al.17 | 41.00 | — | — | 0.78 | 77 | — | — |
The most relevant work for comparison is Odegova et al.,17 from which we obtain the majority of our DES dataset. Their work achieves an RMSE of 41 K, but requires much more expensive features than our work, including the individual component melting points and corresponding xe values. However, their overall goal is to predict the melting points at any given mixture ratio, which differs from our work which is trying to predict the minimum reachable eutectic melting point regardless of ratio.
4.5 Testing on fatty acids and other organics
Our zero-shot prediction results on fatty acids across several models frequently have a strong linear correlation with the target values, and when plotted, appear to form two branches in which mixtures with and without oleic acid appear to follow two different linear slopes. Some fatty acid mixtures are themselves deep eutectic solvents, but these correlations also occur in models trained only on molten salts, before any DES transfer learning. We are not aware of any fundamental chemical relationship between molten salts and fatty acids that would explain this.The KRR model trained on DES data has slightly better zero-shot performance on fatty acids than zero-shot predictions directly from the molten salts model. The molten salt A,A + A,B model zero-shot predictions for fatty acids follow a trend roughly aligned with the x = y line with performance approximately equal to trivial baselines, while after transfer learning and KRR, all error values are slightly better than the trivial baselines. This result is meaningful: achieving baseline-level zero-shot performance indicates that the trained model correctly identifies the approximate average eutectic melting temperature of fatty acids, which exhibits a standard deviation of only about 20 K across the test set. Given that the molten salt training data have a standard deviation roughly an order of magnitude larger, this represents a significant achievement.
Both the zero-shot predictions from the molten salts model and the KRR model that was trained with deep eutectic solvent data are able to perform well on the fatty acids. The fatty acids are the only computationally generated dataset we examined in this work, as Kahwaji et al.11 used the ideal thermodynamic model to compute these. Furthermore, fatty acids are themselves a type of deep eutectic solvent, although we ensured that the fatty acids in this set were not present in the DES training or test sets. Both of these factors could potentially make this set an easier extrapolation problem than the explosives or quinone sets.
We also include test results on explosive eutectics12 and quinones.28 Our predictive performance on these datasets is poor in the context of their respective baselines. However, the neural network and KRR predictions are informative with regards to our models' ability to recognize which mixtures belong to similar families.
5 Discussion
5.1 Learning Te without engineered features
Our architecture is the only method we are aware of that uses only structural information to predict mixture Te values. Although methods which require individual components' melting points, enthalpy values, or xe values may achieve lower prediction error than our method, those same requirements present serious barriers to actually using those methods for high-throughput screening. In contrast, element composition and graph structure as described in Fig. 3 are easily accessible for any known molecule.5.2 Data augmentation with single components
Our experiments which include A,A pairs in training data suggest that we can augment binary mixture data with individual component data to improve performance. Doing so also enables us to use the same trained model to make melting point predictions for new individual components; here we demonstrate performance significantly better than the trivial baselines in all metrics that we evaluated. This also suggests future work in which we augment n-ary mixture data with any lower-order mixture data.5.3 Transfer learning and extrapolation
Typically when training on inorganic materials and testing on organic materials, we expect to see overprediction of melting temperatures. This is because materials with more ionic bonds tend to have higher melting temperatures; as a result, models trained on inorganic materials, where ionic bonds are very common, often learn to map all inputs to higher values than they would if they had been trained on organic materials. However, the model selected for transfer learning did not display this overpredicting behavior. This is a desirable property in a model to be used for transfer learning, since the organic mixtures' predicted melting points are already close to the correct values.Feature extraction and KRR, using DES training data, results in improved predictions on the DES test set, as expected. However, this model still appears unable to extrapolate to other families of organic mixtures, such as explosives and quinones. The same model performs well on fatty acids, which are a type of deep eutectic solvent not seen during model training. Therefore it is possible that the difficulty of extrapolating to explosives and quinones is because these families of materials are not similar enough to deep eutectic solvents.
Our results on the computationally-generated fatty acids dataset suggest that our model is learning a function correlated with the ideal thermodynamic model despite not requiring any individual component melting point or enthalpy data.
6 Conclusions
In this work, we develop a GNN that learns an order-invariant model of melting points for binary eutectic mixtures using minimal data. By using Siamese neural network branches to compute both possible orderings of concatenated molecule representations, we are able to leverage information about the interactions between the two molecules in predicting the melting point of the mixtures.Our architecture is able to learn eutectic melting points from only the graph structure and element composition of the molecules in the mixture, and to learn models that are correlated with the ideal thermodynamic model without requiring the xe value, individual components' melting points, or enthalpies. As a result, this architecture can be used to screen a much wider variety of materials than comparable methods, drastically reducing the barriers to high-throughput screening for eutectic mixtures. This has the potential to speed up materials design for sustainable energy applications such as development of battery electrolytes, thermal energy storage materials, and molten salt nuclear reactors.
This method is also able to use individual components' melting temperature data to augment the eutectic mixture data, leading to both improvements in the prediction of melting temperatures for mixtures, as well as the ability to predict additional individual components' melting temperatures.
We also publish a curated dataset of 2244 molten salt eutectic pairs, including their structural information and eutectic melting points, in order to promote further research by the community.
Conflicts of interest
There are no conflicts to declare.Data availability
All data, data sources, and code repository are available at DOI: https://doi.org/10.5281/zenodo.5483596.Code repository is also available at: https://github.com/nilamandal/SGNN_Eutectic_Mixtures/.
Acknowledgements
This work was supported by the NASA CT Space Grant and the National Science Foundation under Grant No. DMR-2102406. We thank Dr Antonio Baclig for helpful insights and discussion during this project.Notes and references
- A. Ravichandran, S. Honrao, S. Xie, E. Fonseca and J. W. Lawson, J. Phys. Chem. Lett., 2024, 15, 121–126 CrossRef CAS PubMed.
- P. Bhatnagar, S. Siddiqui, I. Sreedhar and R. Parameshwaran, Int. J. Energy Res., 2022, 46, 17755–17785 CrossRef CAS.
- Y. Wang, C. Zhu, M. Zhang and W. Zhou, Nuclear Power Reactor Designs, Elsevier Inc, 2024, pp. 163–183 Search PubMed.
- D. Chicco, in Siamese Neural Networks: An Overview, ed. H. Cartwright, Springer US, New York, NY, 2021, pp. 73–94 Search PubMed.
- R. L. Murphy, B. Srinivasan, V. A. Rao and B. Ribeiro, Janossy Pooling: Learning Deep Permutation-Invariant Functions for Variable-Size Inputs, ICLR, 2019 Search PubMed.
- Y. Zhang and E. J. Maginn, J. Chem. Phys., 2012, 136, 144116 CrossRef PubMed.
- V. Venkatraman, S. Evjen, H. K. Knuutila, A. Fiksdahl and B. K. Alsberg, J. Mol. Liq., 2018, 264, 318–326 CrossRef CAS.
- G. Sivaraman, N. E. Jackson, B. Sanchez-Lengeling, Á. Vázquez-Mayagoitia, A. Aspuru-Guzik, V. Vishwanath and J. J. De Pablo, Mach. Learn.: Sci. Technol., 2020, 1, 025015 Search PubMed.
- K. Low, R. Kobayashi and E. I. Izgorodina, J. Chem. Phys., 2020, 153, 1–13 CrossRef PubMed.
- Z. Acar, P. Nguyen and K. C. Lau, Appl. Sci., 2022, 12, 2408 CrossRef CAS.
- S. Kahwaji and M. A. White, Thermochim. Acta, 2018, 660, 94–100 CrossRef CAS.
- R. D. Chapman and J. W. Fronabarger, Propellants, Explos., Pyrotech., 1998, 23, 50–55 CrossRef CAS.
- G. J. Janz, C. B. Allen, J. R. Downey and R. P. Tomkins, National Standard Reference Data Series, 1978, pp. 1–243 Search PubMed.
- A. Guendouzi and S. M. Mekelleche, Chem. Phys. Lipids, 2012, 165, 1–6 CrossRef CAS PubMed.
- R. K. Luu, M. Wysokowski and M. Buehler, Appl. Phys. Lett., 2023, 122, 234103 CrossRef CAS.
- A. K. Lavrinenko, I. Y. Chernyshov and E. A. Pidko, ACS Sustain. Chem. Eng., 2023, 11, 15492–15502 CrossRef CAS.
- V. Odegova, A. Lavrinenko, T. Rakhmanov, G. Sysuev, A. Dmitrenko and V. Vinogradov, Green Chem., 2024, 26, 3958–3967 RSC.
- J. Gilmer, S. S. Schoenholz, P. F. Riley, O. Vinyals and G. E. Dahl, International Conference on Machine Learning, 2017 Search PubMed.
- W. L. Hamilton, Graph Representation Learning, 1st edn, Springer ChamCham, 2020 Search PubMed.
- Z. Xiong, D. Wang, X. Liu, F. Zhong, X. Wan, X. Li, Z. Li, X. Luo, K. Chen, H. Jiang and M. Zheng, J. Med. Chem., 2020, 63(16), 8749–8760 CrossRef CAS PubMed.
- K. Xu, W. Hu, J. Leskovec and S. Jegelka, How Powerful Are Graph Neural Networks?, ICLR, 2019 Search PubMed.
- M. Jeon, D. Park, J. Lee, H. Jeon, M. Ko, S. Kim, Y. Choi, A.-C. Tan and J. Kang, Bioinformatics, 2019, 35, 5249–5256 CrossRef CAS PubMed.
- H. Altae-Tran, B. Ramsundar, A. S. Pappu and V. Pande, ACS Cent. Sci., 2017, 3, 283–293 CrossRef CAS PubMed.
- L. Torres, N. Monteiro, J. Oliveira, J. Arrais and B. Ribeiro, 2020 IEEE 20th International Conference on Bioinformatics and Bioengineering (BIBE), 2020, pp. 168–175 Search PubMed.
- C. Fotis, N. Meimetis, A. Sardis and L. G. Alexopoulos, Mol. Omics, 2021, 17, 108–120 CrossRef CAS PubMed.
- L. Ward, A. Agrawal, A. Choudhary and C. Wolverton, npj Comput. Mater., 2016, 2, 16028 CrossRef.
- S. Kim, J. Chen, T. Cheng, A. Gindulyte, J. He, S. He, Q. Li, B. A. Shoemaker, P. A. Thiessen, B. Yu, L. Zaslavsky, J. Zhang and E. E. Bolton, Nucleic Acids Res., 2022, 51, D1373–D1380 CrossRef PubMed.
- E. Penn, A. Baclig, D. Ganapathi and W. C. Chueh, Chem. Mater., 2023, 35, 5255–5266 CrossRef CAS.
- S. Falkner, A. Klein and F. Hutter, Proceedings of the 35th International Conference on Machine Learning, ICML 2018, Stockholmsmässan, Stockholm, Sweden, 2018, pp. 1436–1445 Search PubMed.
Footnote |
| † Reprinted from Thermochimica Acta, 660, S. Kahwaji and M. A. White, Prediction of the properties of eutectic fatty acid phase change materials, 94–100, 2018, with permission from Elsevier. |
| This journal is © The Royal Society of Chemistry 2026 |









