 Open Access Article
Open Access ArticleInSpecLearn4SDL: interpretable spectral features predict conductivity in self-driving doped conjugated polymer labs
Ankush Kumar
Mishra
 a,
Jacob P.
Mauthe
b,
Nicholas
Luke
b,
Aram
Amassian
a,
Jacob P.
Mauthe
b,
Nicholas
Luke
b,
Aram
Amassian
 *b and
Baskar
Ganapathysubramanian
*b and
Baskar
Ganapathysubramanian
 *a
*a
aDepartment of Mechanical Engineering, Iowa State University, Ames, IA 50010, USA. E-mail: baskarg@iastate.edu
bDepartment of Materials Science and Engineering and, ORaCEL, North Carolina State University, Raleigh, NC 27695, USA. E-mail: aamassi@ncsu.edu
First published on 18th March 2026
Abstract
To accelerate materials discovery using self-driving labs (SDLs), we present a machine learning pipeline that predicts the electrical conductivity of doped conjugated polymers using rapid, non-destructive optical spectroscopy. Our approach automates spectral featurization by combining a genetic algorithm with adaptive area-under-the-curve (AUC) computations, creating a quantitative structure–property relationship (QSPR) that links optical response and processing parameters to conductivity. By incorporating SHAP-guided selection and domain-knowledge based feature expansion, the model matches expert-curated performance while theoretically reducing experimental effort by ∼33% by minimizing the need for costly direct conductivity measurements. Notably, the model recovers known physical descriptors in pBTTT and identifies informative tail-state regions correlated with polymer bleaching upon successful doping. This generic, interpretable, small–data–friendly methodology can be potentially extended to other modalities, such as Raman or FTIR, providing a framework for autonomous decision-making in SDLs.
1 Introduction
Conjugated polymers (CPs) have been investigated for a variety of organic electronics applications,1 as well as emerging uses such as neuromorphic computing2 and energy storage.3 CPs are organic macromolecules with backbones of alternating single and double bonds; the resulting delocalized π-electron cloud yields distinctive optical and electrical properties.4–6 As in inorganic semiconductors, doping is required to raise charge carrier density to useful levels.7,8 The precise introduction of charge carriers has been central to advances in silicon technologies9,10 and, in organic electronics, is used to regulate charge transport for organic photovoltaics (OPVs),11 organic thermoelectrics (OTEs),12 organic photodetectors,13 organic light-emitting diodes (OLEDs),14 and organic field-effect transistors (OFETs).15–17Successful doping of CPs requires careful selection and synthesis of both the polymer and the dopant, and processing strongly influences physical state and properties.18,19 Even within a single polymer–dopant system, numerous choices (solvents, annealing temperatures, doping times, environment) create a combinatorial design space. This combinatorial design space makes traditional experimentation resource-intensive, necessitating the use of laboratory automation and advanced statistical tools to navigate the diverse range of synthesis routes.
To systematically explore this space, scalable, automated synthesis and characterization are essential. Self-driving labs (SDLs) integrate optimization, machine learning (ML), and robotics to automate discovery.20,21 SDLs have been explored for thin-film properties,22–25 carbon nanotube synthesis,26 mechanics of additively manufactured objects,27,28 nanoparticle synthesis,29–31 yeast genetics,32 and catalyst composition,33 among other areas. SDLs address slow design-space exploration, gaps between experimental stages, and the absence of feedback to select subsequent experiments,34 using adaptive design of experiments (ADoE) to minimize experimental burden. They employ robotics for repetitive tasks and ML models as cost-effective surrogates for linking processing conditions to properties. Within SDLs, properties vary widely in evaluation cost. There is a strong interest in mapping inexpensive measurements to costly properties.35 Traditionally, surrogate features are identified by domain experts, yielding strong predictions but with system-specific, time-consuming efforts that do not readily generalize. As design complexity grows, reliance on manual intuition becomes a bottleneck.
A scalable alternative is to combine expert intuition with data-driven feature identification.36 Experts frame the physics and constraints; algorithms then explore broader candidate features, rank predictive power, and reveal non-obvious relationships. This hybrid approach leverages human insight and the speed and objectivity of ML, enabling more rapid, interpretable, and generalizable feature discovery. The value of automated discovery becomes particularly evident when comparing development timelines: expert-curated features can require a year or more of literature review, experimental validation, and domain-specific analysis.37 In contrast, data-driven methods, with automated pipelines, can identify important features in a fraction of the time, enabling rapid deployment across multiple material systems and spectroscopic modalities without requiring system-specific expertise for each new application.
For doped CPs, optical spectroscopy provides rich information before and after doping.38 Spectral signatures reflect phenomena such as polymer aggregation (linked to carrier mobility)39,40 and charge generation.41 Conductivity obeys σ = |e|µn, where σ is electrical conductivity, |e| is the elementary charge magnitude, µ the mobility, and n the carrier concentration. Spectroscopy is fast (seconds to a minute) and non-destructive, preserving samples for further processing. Thus, spectral features are attractive surrogates for building quantitative structure–property relationships (QSPRs) linking structure and processing to conductivity. QSPRs have been applied across domains.42–49
While raw, pointwise spectra are ideal in principle,50 they are often impractical in low-data regimes due to their high dimensionality. Spectral featurization is a viable alternative. For X-ray absorption near-edge spectra (XANES), prior work has used cumulative distribution function (CDF), peak-based descriptors, and wavelet transforms with dimensionality reduction (PCA, Isomap, autoencoders).51–54 For UV-vis, raw absorbance with PCA/PLS has been employed.55,56 Latent representations via autoencoders have been explored for spectrum–structure relationships in catalysts.57 Torrisi et al.58 improved interpretability by transforming X-ray absorption spectra into multiscale polynomial features that capture local trends. Yoon et al.59 used B-splines-based descriptors to featurize the UV-vis-NIR spectra and used a coefficient shrinkage regression model, LASSO, to identify important regions of the UV-vis-NIR spectra for conductivity prediction of doped conjugated polymers.
Each method has trade-offs: raw spectra are unwieldy at small dataset sizes; peak features can be sensitive to noise; and dimensionality reduction methods may lose information, typically benefiting from larger datasets. We address these challenges with a featurization strategy based on the area under the curve (AUC) combined with a genetic algorithm (GA). AUC over adaptively selected windows encodes feature magnitude and width while being more noise-robust; GA identifies informative regions for downstream modeling.
We treat the derived features as surrogates for conductivity and build a QSPR via data-driven feature engineering, benchmarking against a baseline with expert-curated features. The data-driven model matches the expert-guided model, and a hybrid (data-driven + expert) model outperforms both, highlighting the value of integrating human intuition with ML. Our methodology is generic and can identify informative regions in optical spectra and, more broadly, can be potentially applied to other spectral modalities (XANES, Raman, FTIR). These regions can then be used to predict a quantity of interest (QoI), provided the spectra are physically representative of that QoI.
Our contributions: Our key contributions to this work include the following:
• Data-driven spectral featurization: We propose a data-driven method to featurize optical spectra using the AUC with optimization (GA), and develop a QSPR model for predicting conductivity in doped conjugated polymers.
• Feature engineering: We perform feature engineering to identify key, interpretable features and demonstrate that the data-driven model achieves predictive performance comparable to models based on expert-identified features.
• Human machine learning collaboration: We combine data-driven and expert features to develop a hybrid model that outperformed individual models, demonstrating the benefit of integration human intuition with machine learning.
• Theoretical reduction in experimental time: We show that conductivity characterization accounts for a measured 33% of the total experimental time. By using optical spectra as inputs, these labor-intensive steps can be theoretically eliminated, potentially enabling a 33% reduction in the total experimental cycle time.
2 Results and discussion
2.1 Data collection
Fig. 3 illustrates the step-by-step workflow for preparing a set of 32 samples with duplicates, collecting the spectroscopy, and measuring their conductivity. The process begins with the automated mixing of pBTTT precursor solutions to give the desired co-solvent mixture using the Opentrons platform, followed by automated spin coating. Optical spectroscopy is then performed on the as-cast films, after which the samples undergo annealing. Following annealing, another round of optical spectroscopy is conducted to capture any changes in the spectroscopic signatures that may have occurred during annealing. The film is then doped using a dip-doping method and annealed again. A final spectroscopy step is performed on the doped films. Lastly, sheet resistance and thickness measurements are carried out, which are used to calculate conductivity. Three measurements were taken from both duplicate samples and averaged for statistical robustness.
We perform the experiments on 128 samples. The 128 samples are selected using Bayesian Optimization (BO) for efficient exploration of the design space. We start with 32 samples, obtained through Latin Hypercube Sampling (LHS), and fit a Gaussian process regression (GPR) between the processing conditions and conductivity. We then use the Upper Confidence Bound acquisition function to select the next batch of 32 samples. We perform 3 batches of BO to obtain a total of 128 samples (32 from LHS and 96 from BO). Further details about the BO process, collection, and sharing of data between multi-disciplinary laboratories can be found in our other papers.37,65
Fig. 3 reports the time required to process a batch of 32 samples at each step. Conductivity measurement (comprised of the sheet resistance and thickness measurements) accounts for one-third of the total experimental duration. Specifically, measuring thickness via stylus profilometry is destructive and labor-intensive, requiring manual scraping and multiple readings per sample. Successfully predicting conductivity from optical signatures could eliminate these two operational steps, theoretically reducing experimental effort by ∼33% and substantially increasing the throughput of automated experimentation.
2.2 Data partitioning: train, validation and test split
Our dataset consists of 128 samples, obtained through Bayesian exploration of the design space, each corresponding to a unique combination of processing conditions and their corresponding electrical conductivity. A common approach to splitting data is to perform a random data split between the train, validation, and test sets. However, for smaller datasets, this can lead to uneven distributions between the train, validation, and test sets, resulting in biased evaluation.To avoid this, we first cluster the data to capture its structure. We utilize K-means clustering and determine the optimal number of clusters using the elbow method. The elbow method utilizes the within-cluster sum of squares (WCSS) distance to identify the optimum number of clusters. It does so by finding the “elbow point”, which corresponds to the number of clusters that slows down the decrease in WCSS distance. The optimum number of clusters identified using the elbow method was 5, as shown in Fig. 4a. From each cluster, we randomly selected 20% of the data points, corresponding to 5 points per cluster. These 25 data points are then randomly divided into two sets: a validation set and a test set. The remaining 103 points form the training dataset. The test dataset is kept separate to prevent any data leakage in subsequent model training.
To confirm that all three sets follow the same distribution, we use the Kolmogorov–Smirnov (KS) test66 which compares their empirical distributions. The KS test evaluates the following hypotheses:
 | (1) |
From Table 5 (Appendix 4.2), we observe that all p-values are greater than the significance threshold of α = 0.05. Hence, we fail to reject the null hypothesis H0, indicating that the training, validation and test data are drawn from the same distribution. This supports the assumption that the training, validation, and test data sets should originate from the same underlying data distribution, which is central to most ML models.
2.3 Featurization of spectra and identification of optimum bin locations
To utilize the spectral data, we need to extract meaningful features from the raw spectra collected during the experimental process. These spectra represent three different physical states of the film: as-cast (or unannealed), post-annealed, and post-dope. The as-cast spectra will provide insight into the effects of co-solvent mixtures. As previously noted, the processing solvent may influence aggregation of the polymer film, resulting in noticeable changes to the polymer's absorbance spectrum, such as vibronic progressions. The post-annealed spectra will therefore be more informative about the effects that annealing has on further aggregating (or deaggregating) the polymer as a function of temperature. We expect that this will be more informative than the as-cast spectra due to the strong influence of thermal history and crystallization dynamics. Finally, we expect the post-dope spectroscopy to be informative about the doping process itself. Here we can look for differences in polymer bleaching, anion spectra, and polaron spectra that may be indicative of fluctuations in carrier concentration, which could impact the conductivity.67,68 We also preprocess the raw spectra by performing min–max normalization followed by curve smoothing using the Savitzky–Golay filter function from SciPy. The raw spectra cannot be directly used for model training due to the limited dataset size, and features based on peak or valley position, width, and intensity are highly sensitive to noise. This makes it challenging for algorithms to reliably distinguish true spectral features from noise-induced artifacts.As discussed in Section 1, AUC can serve as a robust alternative feature. Fig. 5 shows how we can use AUC features as an alternative to the identification of peaks and valleys. The AUC captures both the magnitude and the spread of spectral features, implicitly accounting for peak (and valley) intensity, width, and position, while being less sensitive to noise compared to discrete peak/valley detection. To apply this method, we divide the spectrum into a set of bins (identified by the bin locations), and the AUC within each bin is computed as a feature.
The choice of bin locations is critical; well-placed bins isolate informative spectral regions and suppress noisy or irrelevant segments. We cast bin selection as a black-box optimization problem over ordered bin boundaries. This objective is non-convex and non-differentiable; the area-under-the-curve (AUC) features change discretely as boundaries cross peaks/shoulders, and the fitness depends on downstream model training and cross-validation, making gradient-based methods ill-suited.
We therefore use a genetic algorithm (GA) to identify an optimal set of bin locations (see workflow in Fig. 6). We use the training dataset solely to identify the optimal bin locations, thereby avoiding data leakage. GA is a population-based, derivative-free global search method inspired by the principles of natural selection. Rather than following local gradients, it maintains a diverse population of candidate solutions and uses selection, crossover, and mutation to explore the search space across generations. This makes GA less prone to getting trapped in a single local minimum than single-start, gradient-driven optimizers. In our encoding, each candidate represents an ordered set of bin boundaries constrained to lie within the spectral domain; ordering is essential because AUC is computed between consecutive boundaries. We also enforce a minimum bin width to avoid degenerate intervals. The fitness of a candidate is the cross-validated predictive score obtained when AUC features from its bins (optionally combined with processing parameters) are used to train the model.
Several hyperparameters govern GA behavior. The population size controls how broadly the space is explored; the crossover probability encourages exploitation by recombining high-fitness candidates; the mutation probability injects diversity to probe new regions; and the number of generations sets the search horizon (with diminishing returns after a point). We use a population of 100, a crossover probability of 0.7, a mutation probability of 0.3, and 100 generations, following common heuristics and prior practice.69 We repeat the GA multiple times with different seeds. While the exact bin locations varied, the selected spectral regions for featurization were consistently similar.
The fitness of each solution, analogous to a loss function, is evaluated through the following process:
• For each optical spectrum, we compute the AUC under each bin of the candidate.
• We then compute the AUC for the second derivative of the spectra. The choice of the second derivative, in addition to the original spectrum, was based on domain knowledge. The second derivative is calculated from the min–max normalized raw spectra. We then use the Savitzky–Golay filter function from SciPy and set the “deriv” parameter to 2.
• Then we combine the AUC features from the original and second derivative spectra with the corresponding processing parameters. As a guiding principle, we aim to keep the total number of features for the ML model to roughly 10–15% of the training dataset size to avoid overfitting. As the training dataset size was 103, we experimented with 4, 5, and 6 bin locations – corresponding to 3, 4, and 5 bins respectively – yielding 6, 8, and 10 AUC features (from both the original and second-derivative spectra). Among these, the best model performance was observed using 5 bin locations. However, the results and the important features identified for 4 and 6 bin locations were qualitatively similar, suggesting stability in feature selection across a reasonable range of bin counts.
• After this, we train an ML regression model using the training dataset to predict conductivity. We chose a random forest regression model. A detailed discussion of the choice of regression model is presented in Section 2.4.
• Finally, we evaluate the model by computing 5-fold cross-validation root mean square error (RMSE) between predicted and true conductivity for the training dataset. RMSE is used as the fitness function to be minimized.
In each generation of GA, the creation of the population proceeds as below-
• The top p% of the current population (elite solutions) are passed unchanged to the next generation to preserve high-performing candidates. We set p = 5%.
• q% of the new population is generated using crossover and mutation. We set q = 45%:
– Tournament selection is used to choose parents for crossover and mutation. This is done by selecting multiple random candidates from the current population and choosing among them based on their fitness value. This ensures randomness while also ensuring that we choose the best parent among the random candidates.
– Crossover involves swapping portions of bin locations between two parents at a randomly selected crossover point. The resulting offspring are sorted to maintain the constraint that the bin locations in a candidate should be in increasing order.
– Mutation perturbs one or more bin locations within a solution by a random value in a user-defined range.
• The remaining (100 − p − q)% (or 50%) of the population is filled with newly generated random candidates to encourage exploration.
Fig. 7a shows the fitness value across the 100 generations using GA. The optimal bin locations in the post-anneal spectra identified by GA were [1.378, 1.828, 1.982, 2.095, 2.700] eV as shown in Fig. 7b. These bins represent energy intervals where meaningful spectral changes occur, correlating with conductivity. These bin locations contain meaningful information about the polymer's aggregation when analyzed in the right context. The low-energy bin, from 1.378–1.828 eV, lies in the sub-gap region of the absorbance spectrum and thus reflects the tail states arising from the polymer's semi-crystalline nature. The second bin, from 1.828 to 1.982 eV, contains the onset of the 0–0 vibronic peak. The AUC of this bin in the original spectrum and its second derivative will contain some information about the shifting of the peak position, reflecting potential red- or blue-shifting. The third bin, from 1.982–2.095 eV, actually contains the 0–0 vibronic transition, which corresponds to an electronic excitation without a change in the molecular vibrational state. The varying of this feature's prominence in the second derivative AUC will reflect red-shifting or blue-shifting of this low-energy transition and indicate differences in the ground-state energy, likely arising from variations in aggregation or structural order. Similarly, the AUC from the original spectrum will reflect the relative prominence of the 0–0 transition compared to other spectral features, which should correspond to the well-studied 0–0/0–1 ratio. The final bin, from 2.095–2.700 eV, contains the high-energy 0–1 and 0–2 vibronic transitions. The AUC from this region will contain information relevant to the 0–0/0–1 ratio, and the second derivative will reflect the positioning of these transition energies.
Combining all of these bins together, a detailed profile of the polymer's excited state emerges: the 0–0 transition reveals information about the ground state, the 0–1 transition elucidates the strength of electron–vibration coupling, and information from the 0–2 transition would allow for quantification of these interactions through calculation of optoelectronic parameters.40 Further, the ratio of various features, for example, the 0–0/0–1 ratio, has been previously shown to indicate exciton delocalization and the degree of solid-state ordering, which are relevant for doped carrier mobility.70 A physical explanation for each of the terms used in this paragraph has been provided in Appendix 4.6.
2.4 Intermediate QSPR model 1
Once the optimal bin locations (candidate) are identified using GA, we compute the AUC for both the optical spectra and their second derivatives using these bins. Table 1 lists all 8 features and their description. These spectral features are then combined with the corresponding processing conditions to form the complete input feature set. Using this feature set, we train a variety of regression models and evaluate their performance. We explored several categories of algorithms: linear algorithms (linear regression, LASSO, ridge), tree-based ensemble algorithms (random forest and gradient boosting), as well as support vector regression, K-nearest neighbors, and Gaussian regression. Among these, tree-based ensemble algorithms consistently provided the best predictive performance. Table 6 (Appendix 4.2) shows the performance of various algorithms.| Feature | Description |
|---|---|
| CB | % of chlorobenzene solvent (processing condition) |
| DCB | % of ortho-dichlorobenzene solvent (processing condition) |
| Tol | % of toulene solvent (processing condition) |
| annealing_temperature | Annealing temperature (°C) of as-cast film (processing condition) |
| AUC_1 | AUC of original spectra between 1.378 and 1.828 eV |
| AUC_2 | AUC of original spectra between 1.828 and 1.982 eV |
| AUC_3 | AUC of original spectra between 1.982 and 2.095 eV |
| AUC_4 | AUC of original spectra between 2.095 and 2.700 eV |
| d 2AUC_1 | AUC of second derivative of spectra between 1.378 and 1.828 eV |
| d 2AUC_2 | AUC of second derivative of spectra between 1.828 and 1.982 eV |
| d 2AUC_3 | AUC of second derivative of spectra between 1.982 and 2.095 eV |
| d 2AUC_4 | AUC of second derivative of spectra between 2.095 and 2.700 eV |
| X × Y | Product between feature X and Y. X and Y can be any of the 8 AUC features above |
Tree-based models outperformed linear alternatives by effectively capturing the nonlinear interactions and feature couplings inherent in doped conjugated polymer systems. Unlike linear models, which often require extensive feature engineering to handle complex dependencies, tree-based methods automatically learn hierarchical decision rules across categorical and continuous data. This approach is particularly advantageous in our workflow as it requires minimal preprocessing and remains robust to outliers, a critical factor given that conductivity can vary by two orders of magnitude due to processing variations.
To assess how well the model generalizes to unseen samples, we use a combination of evaluation metrics: R2, RMSE, Mean Absolute Error (MAE), Kendall Tau correlation, and Pearson correlation. Each metric provides insight into different aspects of model performance in the context of predicting electrical conductivity. R2 quantifies how well the model explains the variance in measured conductivity compared to a simple baseline that always predicts the mean conductivity. RMSE emphasizes larger errors, making it relevant for identifying whether the model fails on outlier samples, such as those samples with unusually high or low conductivity. MAE provides the average magnitude of prediction error, offering a more robust and interpretable measure of accuracy across the dataset, regardless of outliers. Kendall Tau correlation measures the agreement in ranking between predicted and true conductivity values. Pearson correlation captures the strength of the linear relationship between predicted and actual conductivity values. Together, these metrics provide a comprehensive evaluation, capturing how much variance the model explains, its sensitivity to extreme cases, and how well it preserves both the direction and scale of conductivity trends.
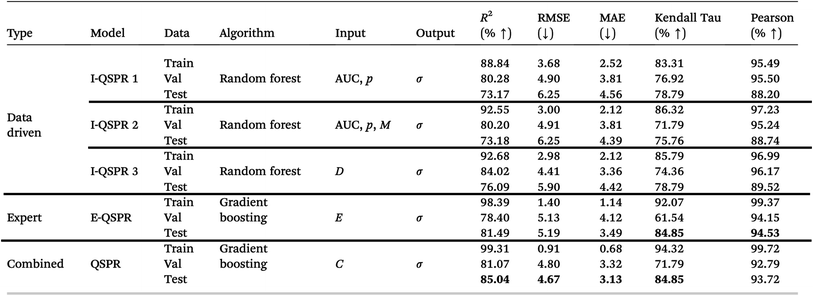
We evaluated various algorithms for intermediate QSPR 1 (Table 6). Among them, the random forest model yielded the best predictive performance. Fig. 8a shows the predicted versus true conductivity values for both the training, validation, and test sets. The performance metrics for the QSPR models are summarized in Table 2. On the test set, the model achieved an R2 score of 73.17%, indicating strong generalization and confirming the predictive capability of features derived from adaptively binned optical spectra.
| a Details: I-QSPR 1, I-QSPR 2, I-QSPR 3: intermediate models using data-driven features. E-QSPR: expert-curated model. QSPR: final model combining data-driven and expert-curated features. In the absence of expert features, I-QSPR 3 serves as the final QSPR. AUC: area-under-the-curve features from spectra and its second derivative; p: processing conditions; σ: conductivity; M: interaction products between AUC features; D: SHAP-selected data-driven subset of AUC, p, and M; E: expert-identified features; C: SHAP-selected best subset from D and E. |
|---|
|
2.5 Domain-knowledge based feature expansion – intermediate QSPR model 2
To further improve model performance, we expanded the feature set by applying simple mathematical transformations to the AUC features. Mathematical transformations, such as ratios, products, logarithms, and exponentials, could be applied to the AUC features. While a wide range of transformations could theoretically be explored, unrestricted application of all combinations would lead to a combinatorial explosion in the number of features, increasing the risk of overfitting.In our case, the selection of mathematical transformations was guided by domain knowledge. Product and ratio transformations between the AUC features were identified as meaningful. It captured the underlying physical interactions between spectral regions that influence conductivity. These derived features could be used to improve the model's predictive capability. We tested both the ratio and product mathematical transformations. We observed that for our problem, the product gave us slightly better performance compared to the ratio.
We computed the pairwise product of all combinations of AUC features. With five bin locations, this resulted in 8 primary AUC features (from the original and second-derivative spectra) and 28 interaction features (8 choose 2,  ), in addition to the 4 processing condition features, yielding a total of 40 input features.
), in addition to the 4 processing condition features, yielding a total of 40 input features.
We trained another ML model using this expanded feature set. We call this model the intermediate QSPR model 2. However, as shown in Table 2, the model's performance on the test set was similar to I-QSPR model 1. The likely reason is overfitting due to the high dimensionality of the feature space relative to the dataset size.71 The inclusion of many correlated features, especially those from the AUCs of both the original and second-derivative spectra, as well as their products, compromises generalization. Given this redundancy, feature selection becomes essential to remove irrelevant or correlated features.
2.6 SHAP-based feature selection
For feature selection, tree-based ensemble models, such as random forest and gradient boosting, provide a built-in mechanism for estimating feature importance. These models build multiple decision trees using bootstrapped samples of the data and subsets of features. During training, features are selected at splits based on how well they reduce impurity (e.g., variance or Gini index). The total reduction in impurity contributed by each feature across all trees yields a global importance score.However, tree-based feature importance has limitations. First, it is not model-agnostic. It relies on how a specific tree-based model splits the data during training. As a result, the importance scores reflect the internal structure and decision rules of that particular model, which can vary with different datasets or model configurations. Moreover, relying on tree-based methods for feature importance restricts us to tree-based models when building QSPRs. While such models performed well in our case, this may not always be the case. In certain scenarios, simpler models, such as linear regression, may offer better performance. Although linear models provide coefficients that can serve as indicators of feature importance, these can be misleading in the presence of multicollinearity or when feature scales vary. This limitation is partially addressed by LASSO regression, which applies L1 regularization to shrink irrelevant coefficients to zero, thereby enabling feature selection and enhancing interpretability. However, LASSO still assumes linear relationships and cannot capture interaction effects. Second, tree-based importance may also miss such interactions, where the relevance of one feature depends on another. Finally, these methods typically provide only global explanations, offering limited insight into individual predictions.
To address these limitations, we employ SHAP (SHapley Additive exPlanations),72 a model-agnostic method based on cooperative game theory. SHAP computes the contribution of each feature to the prediction for each individual data point, offering both global and local interpretability. The SHAP framework represents the model output as an additive model. It is mathematically represented as:
 | (2) |
SHAP values are calculated as the average marginal contribution of a feature across all possible feature subsets:
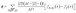 | (3) |
 is the Shapley weight and represents the probability of a particular subset S ⊆ N\{i} appearing before feature i in a random ordering of all features. This weight ensures that all possible feature orderings are fairly considered when computing the contribution of feature i. fS∪{i}(x) − fS(x), measures the marginal contribution of feature i when added to subset S. It quantifies how much the prediction changes when feature i is included, compared to using only the features in S. This captures the added value of feature i given the context of subset S. SHAP provides the average marginal contribution of each feature across all possible subset of features. It also guarantees mathematical properties, specifically, (a) efficiency: the sum of contributions of all features equals the difference between total prediction and average prediction, (b) symmetry: features that equally contribute have equal SHAP values, (c) zero contribution: if a feature does not affect the prediction, its SHAP value is zero, and (d) linearity: if two models are combined, the SHAP value for a feature in the combined model is equal to the sum of it's SHAP value in each individual model. SHAP provides an importance ranking for each feature based on its average contribution to the model.
is the Shapley weight and represents the probability of a particular subset S ⊆ N\{i} appearing before feature i in a random ordering of all features. This weight ensures that all possible feature orderings are fairly considered when computing the contribution of feature i. fS∪{i}(x) − fS(x), measures the marginal contribution of feature i when added to subset S. It quantifies how much the prediction changes when feature i is included, compared to using only the features in S. This captures the added value of feature i given the context of subset S. SHAP provides the average marginal contribution of each feature across all possible subset of features. It also guarantees mathematical properties, specifically, (a) efficiency: the sum of contributions of all features equals the difference between total prediction and average prediction, (b) symmetry: features that equally contribute have equal SHAP values, (c) zero contribution: if a feature does not affect the prediction, its SHAP value is zero, and (d) linearity: if two models are combined, the SHAP value for a feature in the combined model is equal to the sum of it's SHAP value in each individual model. SHAP provides an importance ranking for each feature based on its average contribution to the model.
We compute the mean absolute SHAP score for each input feature to evaluate its contribution to the model's predictions. We chose the random forest algorithm-based model obtained from I-QSPR 2 as it gave the best performance compared to other algorithms (Table 6). We only use the training dataset to rank the features. Table 1 lists the key for all the features, and Fig. 9 shows the SHAP scores for the most important subset of features. To provide instance-level interpretability, Fig. 18 (Appendix 4.5) presents SHAP scores across individual training samples, highlighting how each feature helps in conductivity prediction relative to the model's mean prediction. SHAP is used here to rank feature importance and to support feature selection within the trained QSPR models, rather than to infer causality. Accordingly, the SHAP-based analysis is interpreted in conjunction with established physical understanding, and no causal claims are made.
 | ||
| Fig. 9 Feature importance (SHAP score) for each feature in I-QSPR model 2 (13 features which gave the best I-QSPR model 3 shown). | ||
To identify the most important features, we use a SHAP-guided greedy forward selection strategy. Features are added one by one according to their SHAP importance ranking. At each step, models are trained on the training data and evaluated on the validation set, and the feature subset that minimizes the MAE is selected. Ties are resolved using the RMSE. MAE is chosen as the primary selection metric because it is more robust in small-dataset settings, where individual data points have a large influence on evaluation metrics. In our study, the validation and test sets contain only 13 and 12 samples, respectively, meaning that a single data point represents approximately 8–9% of the dataset. In the presence of outliers, both R2 and RMSE can vary strongly and lead to unstable feature selection. In contrast, MAE penalizes errors linearly, providing a more stable and reliable basis for model comparison. This approach allows us to identify a compact set of informative features that improves generalization while removing redundant or highly correlated features that do not contribute additional predictive value. Fig. 10 shows the validation MAE for the 40 trained models. We observe that the model with 13 features achieves the minimum MAE in the validation set. These 13 features are:
(1) d2AUC_2: AUC for the second derivative of optical spectra between (1.828, 1.982) eV.
(2) AUC_4 × d2AUC_4: product of AUC for original spectra between (2.095, 2.700) eV and AUC for the second derivative of optical spectra between (2.095, 2.700) eV.
(3) AUC_4: AUC of the optical spectra between (2.095, 2.700) eV.
(4) d2AUC_1: AUC for the second derivative of optical spectra between (1.378, 1.828) eV.
(5) d2AUC_3: AUC for the second derivative of optical spectra between (1.982, 2.095) eV.
(6) AUC_3: AUC of the optical spectra between (1.982, 2095) eV.
(7) DCB: ortho-dichlorobenzene volume fraction (%).
(8) AUC_4 × d2AUC_3: product of AUC for original spectra between (2.095, 2.700) eV and AUC for the second derivative of optical spectra between (1.982, 2.095) eV.
(9) d2AUC_4: AUC for the second derivative of optical spectra between (2.095, 2.700) eV.
(10) annealing_temperature: annealing temperature (°C).
(11) CB: chlorobenzene volume fraction (%).
(12) AUC_4 × d2AUC_2: product of AUC for original spectra between (2.095, 2.700) eV and AUC for the second derivative of optical spectra between (1.828, 1.982) eV.
(13) AUC_2 × AUC_4: product of AUC for original spectra between (1.828, 1.982) eV and (2.095, 2.700) eV.
Readers are also referred to Table 1 for descriptions of the features.
I-QSPR model 3 can serve as a surrogate for direct conductivity measurements. As shown in Fig. 3, the conductivity measurement accounts for roughly 33% of the total experimental time. By replacing it with model predictions, we can significantly reduce the experimental burden, thereby enabling higher-throughput experimentation. Moreover, in our current experimental workflow, the post-anneal spectrum is found to be the most informative. Therefore, for studies focused solely on polymer processing, theoretically, an experimental time reduction of up to 50% can be achieved by omitting post-doping steps. However, this simplification is only applicable when post-doping spectra do not provide additional relevant information. Next, we train a new model based on expert-identified features to compare against the results obtained from data-driven features.
2.7 Conductivity prediction using expert features – E-QSPR
In our related work,37 seven spectral features were identified by domain experts through an extensive literature review and validation using experimentally collected data. This effort, which involved a literature survey, prior knowledge of the conjugated polymer, and generation of spectral data from 128 individual samples, required roughly one year of dedicated analysis and resulted in a set of features highly correlated with electrical conductivity. This timeline contrasts with our automated GA-based feature extraction, which completes bin optimization and feature identification within hours, resulting in a substantial reduction in required human effort. The expert identified features are illustrated in Fig. 11 and include:• E0–0: energy corresponding to the zeroth valley in the second derivative of the post-annealed spectrum.
• E0–1: energy corresponding to the first valley in the second derivative of the post-annealed spectrum.
• E0–2: energy corresponding to the second valley in the second derivative of the post-annealed spectrum.
• A0–0/A0–1: ratio of absorbance values at E0–0 and E0–1.
• % Bleaching: ratio of ABleach (post-dope spectrum) to A0–1 (Apoly, post-anneal spectrum).
• Anion signal: ratio of AAnion to ABleach.
• Polaron signal: ratio of APolaron to ABleach.
These features are described in detail in our companion publication.37 We trained a machine learning model using these expert-curated features (referred to as E-QSPR). The model's performance was found to be slightly better than that of I-QSPR model 3, as shown in Table 2.
This result highlights the effectiveness of our data-driven feature extraction strategy, which systematically identifies informative spectral regions using AUC combined with GA. The I-QSPR model 3 achieves R2 = 76.09%, representing 93% of the expert model's performance (R2 = 81.49%) while requiring only several hours of computational time compared to approximately one year of manual analysis. Importantly, our approach is both efficient and generalizable: optimal bin selection and model training can be completed within hours and can potentially be readily applied to new polymer-dopant systems or alternative spectroscopic modalities (Raman, FTIR, XANES) without requiring considerable system-specific expertise. This demonstrates the potential of such automated strategies as a scalable alternative to traditional expert-driven analysis for deployment in self-driving laboratories where rapid, autonomous characterization is essential.
2.8 Combining data-driven features and expert-identified features – final QSPR model
We combine the data-driven features (13 in total) with expert-identified features (7 in total) to examine whether integrating expert knowledge with machine learning leads to improved model performance. A SHAP analysis is conducted to evaluate the importance of each feature, as shown in Fig. 12. Guided by the SHAP-based ranking, we apply a greedy forward-selection strategy, described in Section 2.6, to identify the most informative subset of features and the corresponding best-performing model. Fig. 13 shows the validation MAE for all 20 models. The minimum MAE on the validation set is achieved using 18 features. We also observe that the feature AUC_4 × d2AUC_3 has a perfect correlation with d2AUC_3. So, we drop the feature AUC_4 × d2AUC_3. We then further evaluate the model with the 17 features on the test set. We achieve an R2 of 85% on the test set. This represents an improvement of approximately ∼9% compared to the model built using only data-driven features and ∼4% compared to the model only using expert-identified features, highlighting the potential of combining human expertise with machine learning. Among the 17 selected features, 7 were expert-curated, and 10 were data-driven. Of the data-driven features, three corresponded to processing conditions, while the remaining seven were derived from AUC-based spectral features. A feature correlation matrix illustrating the relationship between data-driven and expert features is provided in Fig. 14. | ||
| Fig. 12 SHAP score for each sample showing directional SHAP score for data-driven features and expert-identified features. | ||
 | ||
| Fig. 14 Spearman correlation between data-driven features (y-axis) and expert-curated features (x-axis) for final QSPR. | ||
Below, we provide a brief analysis of the 7 data-driven spectral features from the combined final QSPR and their connection to the expert-identified features:
d 2AUC_2: AUC of the second derivative of optical spectra between (1.828, 1.982) eV. This feature captures the initial maximum in the second derivative spectrum, which comes from the polymer 0–0 peak onset. A high value corresponds to a red-shifted E0–0, indicative of higher aggregation, which leads to higher conductivity. This is reinforced by the strong correlations of this feature with the E0–0 and E0–1 energies, as well as 0–0/0–1 peak ratio, as shown in Fig. 14.
AUC_3: AUC of the optical spectra between (1.982, 2.095) eV. The area under the curve of this region directly reflects the prominence of the 0–0 vibronic transition relative to the other spectral regions, as well as the width/broadness of the peak onset. In pBTTT films with higher aggregation, this 0–0 peak should be more prominent; this increased aggregation tends to lead to higher mobility and thus conductivity after doping. This is confirmed by the strong correlations of this feature with the E0–0 and 0–0/0–1 ratio in Fig. 14. Interestingly, this feature is also correlated with the bleaching. This may indicate that lower energy 0–0 peaks result in a density of state more suitable for doping with F4TCNQ. This is further investigated in our companion work.
d 2AUC_3: AUC for the second derivative of optical spectra between (1.982, 2.095) eV. This feature captures the peak position of the 0–0 vibronic transition, a deep local minimum in the second derivative (leading to higher values in the SHAP analysis, Fig. 12), indicating the strength and sharpness of the 0–0 transition. This is closely tied to the order and aggregation of the polymer, as evident in the SHAP analysis, which shows high values leading to improvements in the estimated conductivity. This is reinforced by the very strong correlations of this feature with the E0–0 and E0–1 energies as well as 0–0/0–1 peak ratio as noted in Fig. 14.
d 2AUC_4: AUC for the second derivative of optical spectra between (2.095, 2.700) eV. This spectral region captures the higher energy vibronic transitions (E0–1 & E0–2). The local minima in the second derivative are conventionally used to identify these peak locations. The prominence of these minima indicates the intensity of these transitions relative to the 0–0 transition, as well as reflects the positioning of E0–1. A higher area under the curve would indicate strong 0–1 transitions, a sign of disorder and lowered aggregation in pBTTT, which would lead to decreases in conductivity. This is reinforced with the positive correlation with E0–0 and E0–1 as well as the negative correlation with the 0–0/0–1 ratio shown in Fig. 14.
d 2AUC_1: AUC for the second derivative of optical spectra between (1.378, 1.828) eV. This spectral region captures the low-energy tail states. These low-energy tail or trap states are typically found in the amorphous regions of the film and often serve as the initial doping sites. The SHAP analysis in Fig. 12 indicates that a few samples with very low values in this spectral region tend to have higher conductivity. This makes sense as the same amorphous regions that give rise to these trap states tend to have very low mobility, leading to overall lowered conductivity. This is also reinforced by the correlation with bleaching shown in Fig. 14. Notably, this feature is not correlated with any of the pre-doping spectroscopic features identified in our companion study.37
AUC_2 × AUC_4: the product of the AUC of the optical spectra for the (1.828, 1.982) eV and (2.095, 2.700) eV regions. The former region exists below the 0–0 transition and represents low-energy tail states. As previously noted these states often serve as initial doping sites in conjugated polymers though can often lead to lower mobility carriers. This is also reinforced by the correlation with bleaching shown in Fig. 14 as well as samples with low feature value having a positive SHAP value in Fig. 12. The latter spectral region captures the higher energy vibronic transitions (E0–1 & E0–2). The prominence of these transitions, particularly when considered relative to the prominence of the 0–0 transition, are a sign of heightened disorder or lowered aggregation in pBTTT, which would lead to decreases in conductivity as reinforced by the SHAP analysis. Based on the correlation analysis in Fig. 14, the component of this feature appears to be the tail states as seen with higher correlation with bleaching compared to the 0–0/0–1 peak ratio. This product feature represents how domain-knowledge-guided mathematical transformations applied to data-driven bins can encode nonlinear interaction between spectral regions.
AUC_4: AUC of the optical spectra between (2.095, 2.700) eV. This spectral region captures the higher energy 0–1 & 0–2 transitions. As noted in the previous features, this region tends to indicate enhanced disorder of the polymer when the value is high relative to the region containing the 0–0 transition. Though there is little impact in the model from low SHAP values seen in Fig. 12, the correlation analysis in Fig. 14 indicates that this feature is indeed negatively correlated with physical features associated with aggregation, such as the 0–0/0–1 peak ratio.
The overall workflow described in the paper is shown in Fig. 1. The process begins with spectral featurization using AUC combined with GA. Example graphs of the spectral featurization and of high, medium, and low conductivity samples are provided in Fig. 15. Following the data-driven featurization, domain knowledge-based features are incorporated, followed by feature engineering. Introducing additional features through simple, domain-informed mathematical operations, along with feature selection, leads to improved model performance. Further enhancement is achieved by integrating expert-curated features and refining the model, ultimately yielding the best-performing model. There is noticeable overlap in the data-driven features identified using this approach and the known materials descriptors for aggregation, tail states, and doping phenomena as highlighted in Fig. 16. The improvement in model performance upon combining data-driven and expert-curated features demonstrates the value of synergizing human expertise with machine learning.
3 Discussion
In this work, we present a data-driven framework for feature extraction from optical spectra and prediction of electrical conductivity in doped conjugated polymers. Our approach combines area-under-the-curve (AUC) features with a genetic algorithm (GA) to automatically identify informative spectral regions. The resulting QSPR model, augmented with domain-knowledge transformations and targeted feature engineering, demonstrates its advantages through three key findings.The I-QSPR model 3 achieves R2 = 76.09%, representing 93% of the expert model's performance (R2 = 81.49%) while requiring only hours of computational time compared to approximately one year of manual analysis. This efficiency gain potentially enables rapid deployment across multiple material systems, a critical requirement for self-driving laboratories. Second, the data-driven features capture both complementary and overlapping physical information relative to expert knowledge. Features such as d2AUC_1 (low-energy tail states) and product terms like AUC_2 × AUC_4 (encoding nonlinear spectral interactions) have low correlation with expert features. This suggests that the model may be capturing complementary information. Conversely, other data-driven features show high correlation with expert-identified descriptors, demonstrating that the automated method can successfully recover known physical relationships while also discovering new ones. Third, combining data-driven and expert features yields a hybrid model with R2 = 85.04%, outperforming either approach alone. This result highlights the value of human–AI synergy, where domain expertise and machine learning work together to deliver more accurate and interpretable predictors.
Because the models provide early conductivity predictions directly from post-anneal spectra, they function as a surrogate for direct conductivity measurements, theoretically reducing experimental time by approximately one-third and increasing throughput. Additional performance gains may be achievable by expanding the library of mathematical transformations and automating their composition via systematic search.
The framework also integrates naturally with multi-fidelity (Bayesian) optimization, where the QSPR acts as a low-fidelity surrogate and costly conductivity measurements are reserved for high-value candidates. Such workflows enable efficient exploration of large design spaces and support high-throughput experimentation. Overall, the hybrid strategy of combining expert knowledge with automated, data-driven analysis provides a scalable approach to accelerate materials discovery. It is well-suited to deployment in self-driving laboratories and to navigating complex design spaces in organic electronics and beyond.
This study has several limitations. First, the dataset is relatively small. This affects model complexity and limits the use of extensive cross-validation or uncertainty quantification without making performance estimates unstable. Second, the framework is shown on one material system, pBTTT: F4TCNQ. While the methodology is general, model performance and chosen features may depend on specific characteristics of this system. Third, the analysis uses only one spectral method. The approach's effectiveness with other spectroscopic techniques has not been tested and can be explored in the future. Fourth, uncertainty estimates are not reported since the analysis is based on a single train/validation/test split, not repeated resampling. Fifth, we acknowledge that some expert and data-driven features exhibit moderate-to-strong correlations, as they measure related aspects of spectra through different mathematical representations. While greedy forward selection retains only features improving validation performance, we did not explicitly assess multicollinearity or employ decorrelation strategies. Finally, the reported decrease in experimental time is a theoretical estimate based on the current workflow and has not been confirmed through closed-loop autonomous experiments. The integration of the proposed workflow in a full self-driving lab setting is an important next step to be explored in the future.
Author contributions
AKM: methodology, software, data curation, formal analysis, visualization, writing – original draft. JPM: investigation, validation, visualization, writing – original draft. NL: investigation. AA: conceptualization, resources, supervision, writing – review and editing. BG: conceptualization, resources, supervision, project administration, formal analysis, writing – review and editing.Conflicts of interest
The authors declare no competing interests.Data availability
The data and code for InSpecLearn4SDL can be found at https://github.com/ankush-kumar-mishra/InSpecLearn4SDL and https://doi.org/10.5281/zenodo.18761547. Experimental metadata, including processing conditions (solvent volume fractions and annealing temperatures) and measured electrical conductivity, are provided in a master CSV file. Additionally, the corresponding optical absorbance spectra, captured at three distinct states: as-cast, post-annealed, and post-doped, are provided as 128 × 3 individual CSV files containing wavelength and intensity data. All source code required to reproduce the results, including the genetic algorithm for spectral featurization, the QSPR model training pipeline (random forest and gradient boosting), and SHAP-guided feature selection scripts, is provided. A detailed README file explaining the functionality of each script is provided in the repository.Appendix
Appendix 1: Processing parameter selection
| Solvent | δ D (Mpa1/2) | δ P (Mpa1/2) | δ H (Mpa1/2) | Soluble | RED |
|---|---|---|---|---|---|
| Acetone | 15.5 | 10.4 | 7 | 0 | 2.986 |
| Acetonitrile | 15.3 | 18 | 6.1 | 0 | 4.748 |
| 1-Butanol | 16 | 5.7 | 15.8 | 0 | 4.076 |
| Chlorobenzene | 19 | 4.3 | 2 | 1 | 0.471 |
| Chloroform | 17.8 | 3.1 | 5.7 | 1 | 0.952 |
| o-Dichlorobenzene | 19.2 | 6.3 | 3.3 | 1 | 0.993 |
| 1,1,2,2-Tetrachloroethane | 18.8 | 5.1 | 5.3 | 1 | 0.934 |
| Tetrahydrofuran (Thf) | 16.8 | 5.7 | 8 | 0 | 1.957 |
| 1,2,4-Trichlorobenzene | 20.2 | 4.2 | 3.2 | 1 | 0.987 |
| o-Xylene | 17.8 | 1 | 3.1 | 1 | 0.753 |
| Ethyl acetate | 15.8 | 5.3 | 7.2 | 0 | 2.128 |
| Mesitylene | 18 | 0.6 | 0.6 | 1 | 0.999 |
| Toluene | 18 | 1.4 | 2 | 1 | 0.626 |
| Cyclohexane | 16.8 | 0 | 0.2 | 0 | 1.533 |
| n-Butyl acetate (nBA) | 15.8 | 3.7 | 6.3 | 0 | 1.923 |
| Material | δ D (Mpa1/2) | δ P (Mpa1/2) | δ H (Mpa1/2) | R 0 |
|---|---|---|---|---|
| pBTTT-C14 | 18.6 | 3.2 | 2.6 | 3.5 |
| F4TCNQ | 16.5 | 9.5 | 4.4 | 9.0 |
Appendix 2: Data partitioning and algorithm performance result
| Parameter | KS statistic | p-Value | Comment | ||
|---|---|---|---|---|---|
| Val | Test | Val | Test | ||
| a Hyperparameters: I-QSPR 1: n_estimators = 70, criterion = squared_error, min_samples_split = 5; I-QSPR 2: n_estimators = 50, criterion = squared_error, min_samples_split = 2; I-QSPR 3: n_estimators = 50, criterion = squared_error, min_samples_split = 2; E-QSPR: loss = squared_error, learning_rate = 0.1, n_estimators = 100, min_samples_leaf = 1; QSPR: loss = squared_error, learning_rate = 0.1, n_estimators = 150, min_samples_leaf = 5. | |||||
| % CB | 0.23 | 0.18 | 0.48 | 0.78 | Fail to reject H0 |
| % DCB | 0.21 | 0.23 | 0.56 | 0.53 | Fail to reject H0 |
| % Tol | 0.19 | 0.31 | 0.73 | 0.18 | Fail to reject H0 |
| Annealing temp (°C) | 0.15 | 0.21 | 0.92 | 0.61 | Fail to reject H0 |
| Conductivity (S cm−1) | 0.24 | 0.17 | 0.44 | 0.86 | Fail to reject H0 |
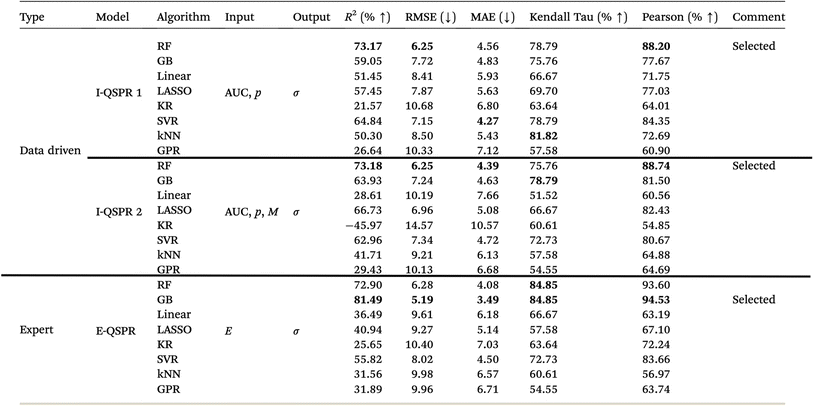
| a Details: I-QSPR 1, I-QSPR 2: intermediate models using data-driven features. E-QSPR: expert-curated model. AUC: area-under-the-curve features from spectra and their second derivative; p: processing conditions; σ: conductivity; M: interaction products between AUC features; D: SHAP-selected data-driven subset of AUC, p, and M; E: expert-identified features; C: SHAP-selected best subset from D and E. RF: random forest; GB: gradient boosting; KR: kernel ridge regression; SVR: support vector regression; kNN: k-nearest neighbor regression; GPR: Gaussian process regression. |
|---|
|
We present results for I-QSPR 1, I-QSPR 2, and E-QSPR, as these models represent different stages of feature development and provide a valuable basis for comparison (Table 6). I-QSPR 3 builds directly on I-QSPR 2. The best algorithm from I-QSPR 2 is chosen, and then we perform SHAP-based feature ranking and selection (Table 6). A similar method is employed for the final QSPR model, which combines data-driven and expert-curated features and utilizes SHAP-based feature selection again. The comparison shows that tree-based models consistently outperform other model types across all evaluation metrics. Therefore, we based the next models (I-QSPR 3 and the final QSPR) on refining the tree-based approach.
Appendix 3: Model performance under spectral noise
We added random 10% Gaussian noise to the spectral data. We use the genetic algorithm-based optimization to obtain the bin locations. The bin locations obtained were [1.966, 1.76, 2.16, 2.51, 2.88] eV. The bin locations obtained from the original data were [1.378, 1.828, 1.982, 2.095, 2.700] eV. We observe that, barring the first location of 1.378 vs. 1.966, the bins more or less cover the same spectral area. We use the bin location obtained from the noisy data and train our data-driven models. We observe that the model performance of the models based on original data was between 73 and 76%, and for the models based on bin location obtained from noisy data, it was between 74 and 77%, as shown in Table 7.| Model | Type | Algorithm | Input | Output | R 2 (% ↑) | RMSE (↓) | MAE (↓) | Kendall Tau (% ↑) | Pearson (% ↑) |
|---|---|---|---|---|---|---|---|---|---|
| a Details: I-QSPR 1, I-QSPR 2, I-QSPR 3: intermediate models using data-driven features. AUC: area-under-the-curve features from spectra and their second derivative; p: processing conditions; σ: conductivity; M: interaction products between AUC features; D: SHAP-selected data-driven subset of AUC, p, and M; E: expert-identified features; C: SHAP-selected best subset from D and E. | |||||||||
| I-QSPR 1 | Original | Random forest | AUC, p | σ | 73.17 | 6.25 | 4.56 | 78.79 | 88.20 |
| 10% noise | 77.26 | 5.75 | 4.22 | 78.79 | 91.81 | ||||
| I-QSPR 2 | Original | Random forest | AUC, p, M | σ | 73.18 | 6.25 | 4.39 | 75.76 | 88.74 |
| 10% noise | 74.80 | 6.06 | 4.13 | 78.79 | 90.49 | ||||
| I-QSPR 3 | Original | Random forest | D | σ | 76.09 | 5.90 | 4.42 | 78.79 | 89.52 |
| 10% noise | 77.87 | 5.67 | 4.01 | 72.73 | 91.26 | ||||
Appendix 4: Model performance on data with conductivity over 30 S cm−1
| Data | True conductivity (S cm−1) | I-QSPR 1 Pred (S cm−1) | I-QSPR 2 Pred (S cm−1) | E-QSPR Pred (S cm−1) |
|---|---|---|---|---|
| a Details: I-QSPR 1, I-QSPR 2: intermediate models using data-driven features. E-QSPR: expert curated model. | ||||
| Val | 32.42 | 24.17 | 25.09 | 22.10 |
| Val | 31.29 | 23.59 | 22.42 | 23.44 |
| Val | 32.65 | 25.44 | 25.85 | 25.13 |
| Val | 30.93 | 22.95 | 23.51 | 29.24 |
| Test | 49.87 | 33.48 | 32.19 | 34.35 |
| MAE | 9.51 | 9.62 | 8.58 | |
| MAE without sample 4 | 9.88 | 10.17 | 10.30 | |
Appendix 5: SHAP results for I-QSPR 3
 | ||
| Fig. 18 SHAP score for each sample in test dataset showing directional SHAP score for each feature in I-QSPR 2. | ||
Appendix 6: Expert feature terminology
Aggregation: the process by which individual polymer chains physically come together, often through π–π stacking or van der Waals forces (Fig. 17). Aggregation can lead to changes in optical properties, such as red-shifted absorption or emission, due to increased interactions between chains. Differences in aggregation arising from co-solvent and/or annealing are often reflected in the absorption spectroscopy as noted in Fig. 19.Red-shift: a shift of an absorption or emission peak to longer wavelengths (lower energy) (Table 8). Often indicative of stronger intermolecular interactions, increased conjugation length, or higher degrees of aggregation or planarity. Fig. 19 shows a red shifting resulting from annealing.
Blue-shift: a shift of an absorption or emission peak to shorter wavelengths (higher energy). Often resulting from decreased conjugation length, structural disorder, disruption of aggregation, or increased localization of the excited state.
Vibronic transition: an electronic transition that occurs along with a change in the molecule's vibrational state. Common vibronic transitions are labeled 0–0, 0–1, and 0–2, where the first number refers to the vibrational level in the ground state and the second refers to the vibrational level of the excited state. Fig. 19 inset shows how these transitions are found using the local minima in the second derivative of the absorption spectrum.
0–0 transition, 0–0 transition: a transition between the lowest vibrational level of the ground state and the lowest vibrational level of the excited state. It represents pure electronic excitation and is often the most direct indicator of the intrinsic energy gap in a conjugated polymer.
0–1 transition: a transition from the ground vibrational level of the ground electronic state to the first vibrational level of the excited electronic state.
0–2 transition: a transition from the ground vibrational level of the ground electronic state to the second vibrational level of the excited electronic state.
Structural order/disorder: refers to the degree of regularity or conformational alignment within a polymer assembly. Structural order tends to enhance electronic delocalization and sharpens optical features. Disorder often introduces broadening and increased vibronic progression.
Planarity: refers to how flat or co-planar the backbone of a conjugated polymer is. Higher planarity facilitates better π-conjugation and delocalization, leading to sharper spectral features and improved charge transport. Planarity is a factor of structural order/disorder.
Delocalization: the extent to which an electronic excitation (e.g., exciton) spreads over multiple molecular units or chains. Delocalized excitons typically result in higher 0–0 transition prominence and narrower peaks, while localized excitons show stronger 0–1 and 0–2 vibronic progression.
Electron–vibrational coupling (electron–phonon coupling): the interaction between an electron's movement and vibrations of the molecule. Strong coupling leads to vibronic progressions (e.g., prominent 0–1, 0–2 peaks) and structural relaxation in excited states.
Vibronic progression: the pattern of multiple vibronic peaks (e.g., 0–0, 0–1, 0–2…) in a spectrum that reflects the strength of vibrational coupling. A pronounced progression suggests stronger electron–vibration interactions.
Huang–Rhys factor (S): a dimensionless quantity that quantifies electron–phonon coupling of a material. A small S indicates weak coupling, often reflected in a sharp 0–0 peak, whereas a large S arises from strong coupling and is observed by more intense 0–1/0–2 transitions (Fig. 20).
Appendix 7: Correlation between data-driven and expert features
 | ||
| Fig. 20 Spearman correlation between data-driven features (first 11 features) and expert-identified features (last 7 features). | ||
Acknowledgements
We acknowledge support from ONR, United States, under award N00014-23-1-2001. J. M. and A. A. also acknowledge NC State's Data Science Academy for support toward the design and development of the materials acceleration platform used in this project. J. M., N. L., and A. A. acknowledge support by the NSF, United States, under award DMR-2523281. B. G. acknowledges partial support from NSF 2323716.References
- T.-F. Yu, H.-Y. Chen, M.-Y. Liao, H.-C. Tien, T.-T. Chang, C.-C. Chueh and W.-Y. Lee, Solution-processable anion-doped conjugated polymer for nonvolatile organic transistor memory with synaptic behaviors, ACS Appl. Mater. Interfaces, 2020, 12(30), 33968–33978 CrossRef CAS PubMed.
- S. Liu, X. Chen and G. Liu, Conjugated polymers for information storage and neuromorphic computing, Polym. Int., 2021, 70(4), 374–403 CrossRef CAS.
- Y. Liang, Z. Chen, Y. Jing, Y. Rong, A. Facchetti and Y. Yao, Heavily n-dopable π-conjugated redox polymers with ultrafast energy storage capability, J. Am. Chem. Soc., 2015, 137(15), 4956–4959 CrossRef CAS PubMed.
- H. Shirakawa, E. J. Louis, A. G. MacDiarmid, C. K. Chiang and A. J. Heeger, Synthesis of electrically conducting organic polymers: halogen derivatives of polyacetylene, (CH)x, J. Chem. Soc. Chem. Commun., 1977,(16), 578–580 RSC.
- A. H. Malik, F. Habib, M. J. Qazi, M. A. Ganayee, Z. Ahmad and M. A. Yatoo, A short review article on conjugated polymers, J. Polym. Res., 2023, 30(3), 115–130 CrossRef CAS.
- Z. Qiu, B. A. G. Hammer and K. Müllen, Conjugated polymers–problems and promises, Prog. Polym. Sci., 2020, 100, 101179 CrossRef CAS.
- P. Kar, Introduction to Doping in Conjugated Polymer, John Wiley & Sons, Ltd, 2013, ch. 1, pp. 1–18 Search PubMed.
- Y. Lu, J.-Y. Wang and J. Pei, Achieving efficient n-doping of conjugated polymers by molecular dopants, Acc. Chem. Res., 2021, 54(13), 2871–2883 CrossRef CAS PubMed.
- T. G. Allen, J. Bullock, X. Yang, A. Javey and S. De Wolf, Passivating contacts for crystalline silicon solar cells, Nat. Energy, 2019, 4(11), 914–928 CrossRef CAS.
- X. Miao, S. Tongay, M. K. Petterson, K. Berke, A. G. Rinzler, B. R. Appleton and A. F. Hebard, High efficiency graphene solar cells by chemical doping, Nano Lett., 2012, 12(6), 2745–2750 CrossRef CAS PubMed.
- R. Meerheim, C. Körner and K. Leo, Highly efficient organic multi-junction solar cells with a thiophene based donor material, Appl. Phys. Lett., 2014, 105(6), 063306 CrossRef.
- O. Bubnova, Z. U. Khan, A. Malti, S. Braun, M. Fahlman, M. Berggren and X. Crispin, Optimization of the thermoelectric figure of merit in the conducting polymer poly(3,4-ethylenedioxythiophene), Nat. Mater., 2011, 10(6), 429–433 CrossRef CAS PubMed.
- B. Siegmund, A. Mischok, J. Benduhn, O. Zeika, S. Ullbrich, F. Nehm, M. Böhm, D. Spoltore, H. Fröb and C. Körner, et al., Organic narrowband near-infrared photodetectors based on intermolecular charge-transfer absorption, Nat. Commun., 2017, 8(1), 15421 CrossRef CAS PubMed.
- S. Reineke, F. Lindner, G. Schwartz, N. Seidler, K. Walzer, B. Lüssem and K. Leo, White organic light-emitting diodes with fluorescent tube efficiency, Nature, 2009, 459(7244), 234–238 CrossRef CAS PubMed.
- T. H. Kim, J. H. Kim and K. Kang, Molecular doping principles in organic electronics: fundamentals and recent progress, Jpn. J. Appl. Phys., 2023, 62(SE), SE0803 CrossRef CAS.
- K. Pei, Recent advances in molecular doping of organic semiconductors, Surf. Interfaces, 2022, 30, 101887 CrossRef CAS.
- B. Lüssem, M. L. Tietze, H. Kleemann, C. Hoßbach, J. W. Bartha, A. Zakhidov and K. Leo, Doped organic transistors operating in the inversion and depletion regime, Nat. Commun., 2013, 4(1), 2775 CrossRef PubMed.
- J. Kim, D. Ju, S. Kim and K. Cho, Disorder-controlled efficient doping of conjugated polymers for high-performance organic thermoelectrics, Adv. Funct. Mater., 2024, 34(6), 2309156 CrossRef CAS.
- Y. Kim, K. Broch, W. Lee, H. Ahn, J. Lee, D. Yoo, J. Kim, S. Chung, H. Sirringhaus and K. Kang, et al., Highly stable contact doping in organic field effect transistors by dopant-blockade method, Adv. Funct. Mater., 2020, 30(28), 2000058 CrossRef CAS PubMed.
- J. T. Rapp, B. J. Bremer and P. A. Romero, Self-driving laboratories to autonomously navigate the protein fitness landscape, Nat. Chem. Eng., 2024, 1(1), 97–107 CrossRef CAS PubMed.
- H. G. Martin, T. Radivojevic, J. Zucker, K. Bouchard, J. Sustarich, S. Peisert, D. Arnold, N. Hillson, G. Babnigg and J. M. Marti, et al., Perspectives for self-driving labs in synthetic biology, Curr. Opin. Biotechnol., 2023, 79, 102881 CrossRef CAS PubMed.
- Y. Liu, A. N. Morozovska, E. A. Eliseev, K. P. Kelley, R. Vasudevan, M. Ziatdinov and S. V. Kalinin, Autonomous scanning probe microscopy with hypothesis learning: Exploring the physics of domain switching in ferroelectric materials, Patterns, 2023, 4(3), 100704 CrossRef PubMed.
- M. B. Rooney, B. P. MacLeod, R. Oldford, Z. J. Thompson, K. L. White, J. Tungjunyatham, B. J. Stankiewicz and C. P. Berlinguette, A self-driving laboratory designed to accelerate the discovery of adhesive materials, Digital Discovery, 2022, 1(4), 382–389 RSC.
- B. P. MacLeod, F. G. L. Parlane, T. D. Morrissey, F. Häse, L. M. Roch, K. E. Dettelbach, R. Moreira, L. P. E. Yunker, M. B. Rooney and J. R. Deeth, et al., Self-driving laboratory for accelerated discovery of thin-film materials, Sci. Adv., 2020, 6(20), eaaz8867 CrossRef CAS PubMed.
- C. Wang, Y.-J. Kim, A. Vriza, R. Batra, A. Baskaran, N. Shan, N. Li, P. Darancet, L. Ward and Y. Liu, et al., Autonomous platform for solution processing of electronic polymers, Nat. Commun., 2025, 16(1), 1498 CrossRef CAS PubMed.
- P. Nikolaev, D. Hooper, F. Webber, R. Rao, K. Decker, M. Krein, J. Poleski, R. Barto and B. Maruyama, Autonomy in materials research: a case study in carbon nanotube growth, npj Comput. Mater., 2016, 2(1), 1–6 CrossRef.
- A. E. Gongora, K. L. Snapp, E. Whiting, P. Riley, K. G. Reyes, E. F. Morgan and K. A. Brown, Using simulation to accelerate autonomous experimentation: A case study using mechanics, Iscience, 2021, 24(4), 102262 CrossRef PubMed.
- A. E. Gongora, B. Xu, W. Perry, C. Okoye, P. Riley, K. G. Reyes, E. F. Morgan and K. A. Brown, A Bayesian experimental autonomous researcher for mechanical design, Sci. Adv., 2020, 6(15), 1708 CrossRef PubMed.
- H. Zhao, W. Chen, H. Huang, Z. Sun, Z. Chen, L. Wu, B. Zhang, F. Lai, Z. Wang and M. L. Adam, et al., A robotic platform for the synthesis of colloidal nanocrystals, Nat. Synth., 2023, 2(6), 505–514 CrossRef CAS.
- A. A. Volk, R. W. Epps, D. T. Yonemoto, B. S. Masters, F. N. Castellano, K. G. Reyes and M. Abolhasani, Alphaflow: autonomous discovery and optimization of multi-step chemistry using a self-driven fluidic lab guided by reinforcement learning, Nat. Commun., 2023, 14(1), 1403 CrossRef CAS PubMed.
- R. W. Epps, M. S. Bowen, A. A. Volk, K. Abdel-Latif, S. Han, K. G. Reyes, A. Amassian and M. Abolhasani, Artificial chemist: an autonomous quantum dot synthesis bot, Adv. Mater., 2020, 32(30), 2001626 CrossRef CAS PubMed.
- R. D. King, K. E. Whelan, F. M. Jones, P. G. K. Reiser, C. H. Bryant, S. H. Muggleton, D. B. Kell and S. G. Oliver, Functional genomic hypothesis generation and experimentation by a robot scientist, Nature, 2004, 427(6971), 247–252 CrossRef CAS PubMed.
- B. Burger, P. M. Maffettone, V. V. Gusev, C. M. Aitchison, Y. Bai, X. Wang, X. Li, B. M. Alston, B. Li and R. Clowes, et al., A mobile robotic chemist, Nature, 2020, 583(7815), 237–241 CrossRef CAS PubMed.
- M. Abolhasani and E. Kumacheva, The rise of self-driving labs in chemical and materials sciences, Nat. Synth., 2023, 2(6), 483–492 CrossRef CAS.
- N. Baishnab, A. K. Mishra, O. Wodo and B. Ganapathysubramanian, Identifying representative sub-domains in 3D microstructures for accelerated structure–property mapping in organic photovoltaic, Comput. Mater. Sci., 2024, 244, 113193 CrossRef CAS.
- T. N. Narong, Z. N. Zachko, S. B. Torrisi and S. J. L. Billinge, Interpretable multimodal machine learning analysis of X-ray absorption near-edge spectra and pair distribution functions, npj Comput. Mater., 2025, 11(1), 98 CrossRef.
- J. P. Mauthe, A. K. Mishra, A. Sarkar, B. Guo, G. J. Thapa, J. Schroedl, N. Luke, J. S. Neu, S.-J. Kwon, M. Chauhan, T. Wang, T. Trapier, H. Ade, D. Ginger, W. You, R. Ghosh, B. Ganapathysubramanian and A. Amassian, AI-guided high-throughput investigation of conjugated polymer doping reveals importance of local polymer order and dopant-polymer separation, Matter, 2026, 9, 102477 CrossRef CAS.
- W. Barford and M. Marcus, Perspective: optical spectroscopy in π-conjugated polymers and how it can be used to determine multiscale polymer structures, J. Chem. Phys., 2017, 146(13), 130902 CrossRef PubMed.
- P. Boufflet, Y. Han, Z. Fei, N. D. Treat, R. Li, D.-M. Smilgies, N. Stingelin, T. D. Anthopoulos and M. Heeney, Using molecular design to increase hole transport: backbone fluorination in the benchmark material poly(2,5-bis(3-alkylthiophen-2-yl)thieno[3,2-b]-thiophene) (pBTTT), Adv. Funct. Mater., 2015, 25(45), 7038–7048 CrossRef CAS.
- S. Wang, J.-C. Tang, L.-H. Zhao, R.-Q. Png, L.-Y. Wong, P.-J. Chia, H. S. O. Chan, P. K.-H. Ho and L.-L. Chua, Solvent effects and multiple aggregate states in high-mobility organic field-effect transistors based on poly(bithiophene-alt-thienothiophene), Appl. Phys. Lett., 2008, 93(16), 162103 CrossRef.
- J. E. Cochran, M. J. N. Junk, A. M. Glaudell, P. L. Miller, J. S. Cowart, M. F. Toney, C. J. Hawker, B. F. Chmelka and M. L. Chabinyc, Molecular interactions and ordering in electrically doped polymers: blends of PBTTT and F4TCNQ, Macromolecules, 2014, 47(19), 6836–6846 CrossRef CAS.
- J. Hu, X. Zhang and Z. Wang, A review on progress in QSPR studies for surfactants, Int. J. Mol. Sci., 2010, 11(3), 1020–1047 CrossRef CAS PubMed.
- G. Fayet and P. Rotureau, How to use QSPR-type approaches to predict properties in the context of green chemistry, Biofuel Bioprod. Biorefining, 2016, 10(6), 738–752 CrossRef CAS.
- S. Kwon, H. Bae, J. Jo and S. Yoon, Comprehensive ensemble in QSAR prediction for drug discovery, BMC Bioinf., 2019, 20, 1–12 CAS.
- A. Fluetsch, E. Di Lascio, G. Gerebtzoff and R. Rodríguez-Pérez, Adapting deep learning QSPR models to specific drug discovery projects, Mol. Pharm., 2024, 21(4), 1817–1826 CrossRef CAS PubMed.
- O. Tinkov, P. Polishchuk, V. Grigorev and Y. Porozov, The cross-interpretation of QSAR toxicological models, in International Symposium on Bioinformatics Research and Applications, Springer, 2020, pp. 262–273 Search PubMed.
- S. Liu, L. Jin, H. Yu, L. Lv, C.-E. Chen and G.-G. Ying, Understanding and predicting the diffusivity of organic chemicals for diffusive gradients in thin-films using a QSPR model, Sci. Total Environ., 2020, 706, 135691 CrossRef CAS PubMed.
- T. Wang, R. Li, H. Ardekani, L. Serrano-Luján, J. Wang, M. Ramezani, R. Wilmington, M. Chauhan, R. W. Epps and K. Darabi, et al., Sustainable materials acceleration platform reveals stable and efficient wide-bandgap metal halide perovskite alloys, Matter, 2023, 6(9), 2963–2986 CrossRef CAS.
- Y. Wen, Y. Liu, B. Yan, T. Gaudin, J. Ma and H. Ma, Simultaneous optimization of donor/acceptor pairs and device specifications for nonfullerene organic solar cells using a QSPR model with morphological descriptors, J. Phys. Chem. Lett., 2021, 12(20), 4980–4986 CrossRef CAS PubMed.
- R. Zhang, R. Black, D. Sur, P. Karimi, K. Li, B. DeCost, J. R. Scully and J. Hattrick-Simpers, Editors' choice—AutoEIS: automated Bayesian model selection and analysis for electrochemical impedance spectroscopy, J. Electrochem. Soc., 2023, 170(8), 086502 CrossRef CAS.
- J. Timoshenko and A. Kuzmin, Wavelet data analysis of EXAFS spectra, Comput. Phys. Commun., 2009, 180(6), 920–925 CrossRef CAS.
- M. Munoz, P. Argoul and F. Farges, Continuous Cauchy wavelet transform analyses of EXAFS spectra: a qualitative approach, Am. Mineral., 2003, 88(4), 694–700 CrossRef CAS.
- Y. Chen, C. Chen, I. Hwang, M. J. Davis, W. Yang, C. Sun, G.-H. Lee, D. McReynolds, D. Allan and J. M. Arias, et al., Robust machine learning inference from X-ray absorption near edge spectra through featurization, Chem. Mater., 2024, 36(5), 2304–2313 CrossRef CAS.
- A. Manthiram, A reflection on lithium-ion battery cathode chemistry, Nat. Commun., 2020, 11(1), 1550 CrossRef CAS PubMed.
- R. Razavi and R. E. Kenari, Ultraviolet–visible spectroscopy combined with machine learning as a rapid detection method to the predict adulteration of honey, Heliyon, 2023, 9(10), e20973 CrossRef CAS PubMed.
- R. Gariso, J. P. L. Coutinho, T. J. Rato and M. S. Reis, A comparative analysis of deep learning and chemometric approaches for spectral data modeling, Anal. Chim. Acta, 2025, 1347, 343766 CrossRef CAS PubMed.
- P. K. Routh, Y. Liu, N. Marcella, B. Kozinsky and A. I. Frenkel, Latent representation learning for structural characterization of catalysts, J. Phys. Chem. Lett., 2021, 12(8), 2086–2094 CrossRef CAS PubMed.
- S. B. Torrisi, M. R. Carbone, B. A. Rohr, J. H. Montoya, Y. Ha, J. Yano, S. K. Suram and L. Hung, Random forest machine learning models for interpretable X-ray absorption near-edge structure spectrum-property relationships, npj Comput. Mater., 2020, 6(1), 109 CrossRef.
- J. W. Yoon, A. Kumar, P. Kumar, K. Hippalgaonkar, J. Senthilnath and V. Chellappan, Explainable machine learning to enable high-throughput electrical conductivity optimization and discovery of doped conjugated polymers, Knowl. Base Syst., 2024, 295, 111812 CrossRef.
- Y. Yamashita, J. Tsurumi, M. Ohno, R. Fujimoto, S. Kumagai, T. Kurosawa, T. Okamoto, J. Takeya and S. Watanabe, Efficient molecular doping of polymeric semiconductors driven by anion exchange, Nature, 2019, 572(7771), 634–638 CrossRef CAS PubMed.
- H. L. Yi, C. H. Wu, C. I. Wang and C. C. Hua, Solvent-regulated mesoscale aggregation properties of dilute pBTTT-C14 solutions, Macromolecules, 2017, 50(14), 5498–5509 CrossRef CAS.
- V. Pirela, A. J. Müller and J. Martín, Crystallization kinetics of semiconducting poly(2,5-bis(3-alkylthiophen-2-yl)-thieno-[3,2-b]thiophene) (PBTTT) from its different liquid phases, J. Mater. Chem. C, 2024, 12(11), 4005–4012 RSC.
- S. N. Patel, A. M. Glaudell, K. A. Peterson, E. M. Thomas, K. A. O'Hara, E. Lim and M. L. Chabinyc, Morphology controls the thermoelectric power factor of a doped semiconducting polymer, Sci. Adv., 2017, 3(6), e1700434 CrossRef PubMed.
- H.-L. Yi and C.-C. Hua, PBTTT-C16 sol–gel transition by rod associations and networking, Soft Matter, 2019, 15(40), 8022–8031 RSC.
- R. Tali, A. K. Mishra, D. Lohia, J. P. Mauthe, J. S. Neu, S.-J. Kwon, Y. Olanrewaju, A. Balu, G. Trajcevski and F. So, et al., SEARS: a lightweight fair platform for multi-lab materials experiments and closed-loop optimization, Digital Discovery, 2025, 4(11), 3126–3136 RSC.
- Wikipedia contributors, Kolmogorov–Smirnov test, https://en.wikipedia.org/wiki/Kolmogorov2025, accessed: 2025-04-15.
- S.-J. Kwon, R. Giridharagopal, J. Neu, S. Kashani, S. E. Chen, R. J. Quezada, J. Guo, H. Ade, W. You and D. S. Ginger, Quantifying doping efficiency to probe the effects of nanoscale morphology and solvent swelling in molecular doping of conjugated polymers, J. Phys. Chem. C, 2024, 128(6), 2748–2758 CrossRef CAS.
- D. Kiefer, R. Kroon, A. I. Hofmann, H. Sun, X. Liu, A. Giovannitti, D. Stegerer, A. Cano, J. Hynynen and L. Yu, et al., Double doping of conjugated polymers with monomer molecular dopants, Nat. Mater., 2019, 18(2), 149–155 CrossRef CAS PubMed.
- B. L. Greenstein, D. C. Elsey and G. R. Hutchison, Determining best practices for using genetic algorithms in molecular discovery, J. Chem. Phys., 2023, 159(9), 091501 CrossRef CAS PubMed.
- K. Zhao, H. U. Khan, R. Li, Y. Su and A. Amassian, Entanglement of conjugated polymer chains influences molecular self-assembly and carrier transport, Adv. Funct. Mater., 2013, 23(48), 6024–6035 CrossRef CAS.
- J. Hua, Z. Xiong, J. Lowey, E. Suh and E. R. Dougherty, Optimal number of features as a function of sample size for various classification rules, Bioinformatics, 2005, 21(8), 1509–1515 CrossRef CAS PubMed.
- S. M. Lundberg and S.-I. Lee, A unified approach to interpreting model predictions, Adv. Neural Inf. Process. Syst., 2017, 30, 4768–4777 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |















