 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
ToPolyAgent: AI agents for coarse-grained bead-spring topological polymer simulations
Lijie
Ding
 *a,
Jan-Michael
Carrillo
*a,
Jan-Michael
Carrillo
 b and
Changwoo
Do
b and
Changwoo
Do
 *a
*a
aNeutron Scattering Division, Oak Ridge National Laboratory, Oak Ridge, TN 37831, USA. E-mail: dingl1@ornl.gov; doc1@ornl.gov
bCenter for Nanophase Materials Sciences, Oak Ridge National Laboratory, Oak Ridge, TN 37831, USA
First published on 27th January 2026
Abstract
We introduce ToPolyAgent, a multi-agent AI framework for performing coarse-grained molecular dynamics (MD) simulations of topological polymers through natural language instructions. By integrating large language models (LLMs) with domain-specific computational tools, ToPolyAgent supports both interactive and autonomous simulation workflows across diverse polymer architectures, including linear, ring, brush, and star polymers, as well as dendrimers. The system consists of four LLM-powered agents: a Config Agent for generating initial polymer–solvent configurations, a Simulation Agent for executing LAMMPS-based MD simulations and conformational analyses, a Report Agent for compiling markdown reports, and a Workflow Agent for streamlined autonomous operations. Interactive mode incorporates user feedback loops for iterative refinements, while autonomous mode enables end-to-end task execution from detailed prompts. We demonstrate ToPolyAgent's versatility through case studies involving diverse polymer architectures under varying solvent conditions, thermostats, and simulation lengths. Furthermore, we highlight its potential as a research assistant by directing it to investigate the effect of interaction parameters on the linear polymer conformation, and the influence of grafting density on the persistence length of the brush polymer. By coupling natural language interfaces with rigorous simulation tools, ToPolyAgent lowers barriers to complex computational workflows and advances AI-driven materials discovery in polymer science. It lays the foundation for autonomous and extensible multi-agent scientific research ecosystems.
1 Introduction
Topological polymers,1–3 characterized by their diverse architectures such as linear,4,5 ring,6,7 brush,8–11 star,12,13 and dendrimer14 structures, exhibit unique physical properties that make them critical in applications ranging from drug delivery15–19 to advanced materials.20,21 These properties arise from their complex molecular configurations, which influence their conformational dynamics, phase behavior, and interactions with solvent.22–27 Understanding these behaviors often requires molecular dynamics (MD) simulations,3,28–30 which provide detailed insights into the polymer structure and dynamics at scales from atomistic to coarse-grained.31–34 However, despite the development of professional simulation software such as LAMMPS,35 GROMACS,36 AMBER,37 and Desmond,38 the computational complexity of simulating topological polymers, even in coarse-grained models, still demands significant expertise, limiting their accessibility to researchers without extensive computational backgrounds.Recent advances in large language models (LLMs)39–41 and agentic AI42,43 have opened new avenues for automating and simplifying complex scientific workflows.44 LLMs, with their ability to process and generate natural language, can interpret user instructions, orchestrate tasks, and integrate domain-specific tools to perform specialized computations.45,46 Agentic AI systems, composed of multiple AI agents with distinct roles, further enhance this capability by enabling collaborative and autonomous task execution. These developments have the potential to transform computational materials science by making advanced simulations more intuitive and efficient, while greatly lowering the barrier to using professional computational tools like MD software. The application of AI agent started with software development,47,48 and has been quickly expanding to various areas of scientific research.49–51 Recent efforts have begun to apply AI agents to molecular dynamics workflows. For example, LUNAR52 automates LAMMPS input generation and post-processing for reactive simulations using traditional scripting tools, while fine-tuned large language models53 are used for iterative script generation and refinement, primarily targeting thermodynamic parameter exploration in all-atom and coarse-grained systems. These approaches significantly reduce the effort required to produce valid simulation inputs. However, they focus predominantly on code and script generation and do not provide fully autonomous or interactive execution of the complete workflow—from topology-specific initial configuration generation, through simulation running and trajectory analysis, to final reporting.
In this work, we address the aforementioned challenges by introducing ToPolyAgent, a multi-agent AI framework designed to perform coarse-grained molecular dynamics (MD) simulations of topological polymers through natural language interfaces. The coarse-grained MD simulation workflow is based on a tutorial supported by the U.S. National Science Foundation (NSF).54,55 ToPolyAgent integrates large language models (LLMs) with domain-specific computational tools to support simulations of diverse polymer architectures under varying solvent conditions and simulation parameters. The system operates in two modes: an interactive mode, which incorporates user feedback for iterative refinement, and an autonomous mode, which performs end-to-end simulations from detailed prompts.
We demonstrate ToPolyAgent's capabilities through case studies that highlight its potential as a research assistant, including investigations of the effect of solvent quality on linear polymer conformation and the influence of grafting density on the persistence length of brush polymer backbones. By bridging natural language processing with rigorous computational methods, ToPolyAgent lowers the barrier to performing complex polymer simulations and establishes a foundation for scalable, AI-driven materials research.
The rest of this paper is organized as follows: in Section 2, we describe the system design of ToPolyAgent, and provide details on the tools been used and technical detail on the MD simulation. Section 3 presents a series of examples and use cases of the ToPolyAgent simulating different types of topological polymer under various conditions, while demonstrating the capability of ToPolyAgent as a research assistant. Finally, we summarize this work and discuss potential future directions in Section 4.
2 Method
2.1 System design
ToPolyAgent is designed not only to perform coarse-grained MD simulations for various types of topological polymers directly from detailed prompts but also able to interact with users and adjust simulation settings based on their feedback. We implement this as a multi-agent system with two operational modes: interactive mode and autonomous mode. Fig. 1 illustrates the overall system design. Four distinct agents support these two modes in conducting MD simulations for topological polymers. The interactive mode is supported by the Config Agent, Simulation Agent, and Report Agent, while the autonomous mode relies on the Workflow Agent and Report Agent.In the interactive mode, the first agent to interact with the user is the Config Agent, whose task is to generate the initial configuration data file containing the positions and connectivities of all polymer beads, as well as the initial positions of all solvent particles. The Config Agent employs six different tools to perform its tasks. Five of these tools are dedicated to generating five types of topological polymers—linear, ring, brush, star, and dendrimer—each with specific input parameters that control properties such as size, shape, and solvent concentration. The remaining tool, known as the human tool, prompts the user for additional feedback so that the Config Agent can iteratively modify the system configuration until the user is satisfied and agrees to proceed to the next phase.
The second phase of the interactive mode is handled by the Simulation Agent, which interprets user prompts and runs molecular dynamics (MD) simulations for the previously generated polymer configurations using appropriate parameters. In addition, the Simulation Agent performs data analysis on the simulation output and presents figures for user evaluation. Similar to the Config Agent, the Simulation Agent also has access to the human tool and will request user feedback to adjust and rerun simulations as needed.
Once the user approves the simulation results, the interactive mode proceeds to the third phase, in which all results—including simulation data, parameters, and interaction logs—are compiled by the Report Agent. The Report Agent uses tools to read logs and files, then generates a comprehensive simulation report in markdown format.
In the autonomous mode, the agents expect the user to provide more detailed prompts so that the simulations can be carried out with greater specificity. The main difference between the autonomous and interactive modes is the absence of a human feedback loop in the former. In this mode, the Workflow Agent is responsible for generating the initial system configuration, performing the MD simulation of the polymer–solvent system, and conducting data analysis. The resulting outputs are then passed to the same Report Agent, which compiles them into a final report.
In practice, we use CrewAI to orchastrate the multi-agent system and facilitate agent-to-agent collaboration, as well as to maintain the agents' memory systems. The LLM is powered by OpenRouter's API, which enables flexible selection of the underlying model. And without loss of generality, we use OpenAI's GPT-4o-mini56 for the results presented in this study.
2.2 Tools for agents
To enable the different agents to accomplish their designated tasks, we develope toolsets tailored to each agent's tasks. The configuration generation toolset creates the initial configurations for LAMMPS to read, simulate, and visualize through plots. The simulation toolset runs the MD simulations and performs conformational analysis using the dump files produced by LAMMPS. The human tool connects the agents to the user via command-line inputs. Finally, the report toolset allows agents to read the files generated during the simulations and compile reports in markdown format.For system configuration generation, five tools were developed, each corresponding to a specific type of topological polymer. These tools read predefined input parameters from docstrings, generate the polymer configurations based on user instructions, add solvent to the simulation box, save the resulting LAMMPS data file, and visualize the configurations. All tools require the simulation box size B and solvent density ns as inputs, with additional topology-specific parameters. GenerateLinearPolymer and GenerateRingPolymer require the chain length N. GenerateBrushPolymer requires the backbone length Nb, grafting density σg, and side chain length Ns. GenerateStarPolymer requires the arm length Na and number of arms m. GenerateDendrimer requires the number of generations G, branching factor b, and spacer length Ns.
For the simulation tools set, the runLAMMPS tool executes the MD simulation using the LAMMPS data file generated by the Config Agent. It takes as input the interaction parameters: polymer–polymer εpp, solvent–solvent εss, and polymer–solvent εsp, along with the thermostat type (Langevin or Nose–Hoover) and the number of simulation steps. Details of the MD simulations are provided in the following subsection. Additionally, the ComformationAnalysis tool analyzes the dump files produced by the LAMMPS simulations.
The following characteristic variables of polymer conformation are calculated:57,58 the square of the radius of gyration, Rg2 = 〈(ri − rc)2〉i, where 〈…〉i denotes the average over all beads on the polymer, and rc = 〈ri〉i is the center of mass of the polymer. The mean square displacement (MSD) curve is given by MSD(Δt) = 〉|rc(t + Δt) − rc(t)|2〈t, where 〉…〈t denotes the time average, and the diffusion coefficient D is obtained from the relation MSD = 6DΔt.
We also calculate the end-to-end distance, Ree = |rN − r1|, defined as the distance between the first and last beads on the polymer chain, and the persistence length lp, obtained from the bond–bond correlation function 〉cos![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) θ(s)〈 = e−s/lp, where θ(s) is the angle between tangent vectors of the chain separated by contour length s. The persistence length and end-to-end distance are not calculated for dendrimers, as these quantities are not well defined, and are evaluated only for the backbone in brush polymers and for a single arm in star polymers. The end-to-end distance is also not calculated for ring polymers.
θ(s)〈 = e−s/lp, where θ(s) is the angle between tangent vectors of the chain separated by contour length s. The persistence length and end-to-end distance are not calculated for dendrimers, as these quantities are not well defined, and are evaluated only for the backbone in brush polymers and for a single arm in star polymers. The end-to-end distance is also not calculated for ring polymers.
Additionally, we calculate the polymer form factor and the radial distribution function. The form factor is given by the isotropic intra-polymer structure factor:59,60
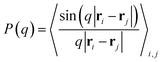 | (1) |
| g(r) = 〈δ(|ri − rj| − r)〉i,j | (2) |
Finally, the human tool and report tool set are straightforward: the human tool allows the agent to receive natural-language feedback from the user, while the report tool reads and writes markdown files to the local directory.
2.3 Molecular dynamics
The main functionality of ToPolyAgent is to perform MD simulations of coarse-grained polymer models, in which the polymer is represented by a number of beads connected in different configurations. The simulations are carried out using LAMMPS. A truncated and shifted Lennard-Jones (LJ) potential is applied between every pair of beads:28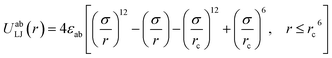 | (3) |
We use the finite extensible nonlinear elastic (FENE) bond potential for the connected beads on the polymer, given by:61
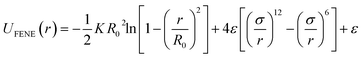 | (4) |
We implement both the Langevin and Nosé–Hoover thermostats in the simulations. The equation of motion for the Langevin thermostat is given by:62,63
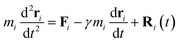 | (5) |
The Nosé–Hoover thermostat maintains a constant temperature by introducing a fictitious degree of freedom that couples the system to a heat bath, thereby generating an NVT ensemble. The equations of motion are given by:64,65
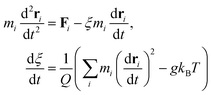 | (6) |
3 Results
We begin by demonstrating examples of using ToPolyAgent to perform MD simulations of topological polymers with various architectures, including the interactive mode, which allows revisions during the process, and the autonomous mode, which executes the workflow to completion. We then showcase ToPolyAgent as a research assistant, capable of running multiple MD simulations through natural language interaction and compiling research findings for two representative cases: linear polymers and brush polymers.3.1 Interactive mode
In the interactive mode, ToPolyAgent engages with the user throughout the entire simulation process. We present two examples: one for a brush polymer, where multiple revisions are made based on user requests, and another for a star polymer, where the simulation proceeds in a streamlined manner.Fig. 2 illustrates the workflow of the dialogue between the user and ToPolyAgent during the simulation of a brush polymer in solvent. Beginning with the generation of the brush polymer configuration and solvent packing shown in Fig. 2(a), the Config Agent first produces a brush polymer using default settings. The user then provides feedback requesting an increase in grafting density. The Config Agent interprets this feedback, retrieves the previously used parameters for brush polymer generation from memory, and re-runs the configuration tool to adjust only the grafting density while keeping other parameters unchanged.
After the user approves the generated system configuration, the Simulation Agent takes over, as shown in Fig. 2(b). It begins by asking the user for additional specifications of the simulation parameters, then performs the MD simulation and corresponding polymer conformation analysis using the simulation tools, presenting the final system configuration and analysis results to the user. Subsequently, as shown in Fig. 2(c), the user requests the Simulation Agent to extend the simulation to a longer run, which the agent executes accordingly. Upon final approval from the user, the Report Agent compiles a comprehensive report containing the simulation context, parameters, and conformation analysis results, with plots embedded in a markdown file.
The final simulation corresponds to a brush polymer with backbone length Nb = 20, grafting density σg = 0.6, and side chain length Ns = 5, within a simulation box of size B = 20 and solvent number density ns = 0.3. The interaction parameters are εpp = εss = 0.3 and εsp = 1.5. The system is simulated using a Langevin thermostat for 90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 steps with a time step of dt = 0.01. The complete conformation analysis results for the final run are shown in Fig. 3.
000 steps with a time step of dt = 0.01. The complete conformation analysis results for the final run are shown in Fig. 3.
Finally, in Fig. 4, we present an example of agent–user communication in which the user does not request any revisions during the process. The Config Agent selects its own parameters according to the system prompt defined in the tool's docstring and generates a system consisting of a star polymer with arm length Na and number of arms m = 6 in a simulation box of size B = 20, filled with solvent at a number density of ns = 0.2. The Simulation Agent then takes over and runs the simulation using user-specified parameters, including εpp = 0.5, εss = 0.7, and εsp = 1.0, for 40![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 steps under a Langevin thermostat. Finally, the Report Agent compiles the results and generates a markdown report.
000 steps under a Langevin thermostat. Finally, the Report Agent compiles the results and generates a markdown report.
3.2 Autonomous mode
In the autonomous mode, ToPolyAgent automatically performs all stages of the topological polymer simulation and provides the user with the final reports. Compared with the interactive mode, which allows for detailed user tuning throughout the process, the autonomous mode streamlines the entire workflow and benefits from a more detailed initial prompt, enabling both the configuration generation and MD simulation steps to reflect user-specified parameters.
Fig. 5 shows three examples of ToPolyAgent operating in autonomous mode, demonstrating simulations of a linear polymer, a ring polymer, and a dendrimer. In Fig. 5(a), the user instructs ToPolyAgent to simulate a linear polymer, specifying the polymer length, thermostat type (Nosé–Hoover), and a run length of 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 steps. ToPolyAgent correctly applies the user-specified parameters, performs the simulation, and provides the corresponding analysis results. In Fig. 5(b), the user requests a ring polymer simulation, specifying the polymer size, solvent density, three interaction parameters for polymer–solvent interactions, and the total number of simulation steps. The agent accurately assigns these parameters and executes the simulation. The conformation analysis excludes the end-to-end distance, as it is not defined for ring polymers. Finally, in Fig. 5(c), the agent receives detailed instructions for generating a dendrimer configuration, including solvent properties, simulation box size, and simulation length, and successfully performs the simulation using the correct parameter values. For dendrimers, since both the end-to-end distance Ree and persistence length lp are not well defined, only the other conformational characteristics are presented.
000 steps. ToPolyAgent correctly applies the user-specified parameters, performs the simulation, and provides the corresponding analysis results. In Fig. 5(b), the user requests a ring polymer simulation, specifying the polymer size, solvent density, three interaction parameters for polymer–solvent interactions, and the total number of simulation steps. The agent accurately assigns these parameters and executes the simulation. The conformation analysis excludes the end-to-end distance, as it is not defined for ring polymers. Finally, in Fig. 5(c), the agent receives detailed instructions for generating a dendrimer configuration, including solvent properties, simulation box size, and simulation length, and successfully performs the simulation using the correct parameter values. For dendrimers, since both the end-to-end distance Ree and persistence length lp are not well defined, only the other conformational characteristics are presented.
With the autonomous mode, ToPolyAgent can serve as a self-contained module within a larger system in which individual components communicate with one another through natural language, similar to human interaction. Despite relying on language-based inputs and outputs, ToPolyAgent employs rigorous scientific tools internally, producing more reliable and reproducible results than commercially available, general-purpose AI chatbots.
3.3 ToPolyAgent as research assistant
Similar to a research assistant who conducts experiments and compiles reports based on instructions from an advisor, ToPolyAgent can play the same role—translating natural-language instructions into simulation results and reporting them back to the instructor. These results can then contribute to new research findings. Fig. 6 illustrates this process. More importantly, because ToPolyAgent accepts natural-language input, the instructor does not necessarily have to be human—it could also be another AI agent. To demonstrate the potential of ToPolyAgent as an AI research assistant, we present two example use cases: one involving a linear polymer and another involving a brush polymer. | ||
| Fig. 6 Illustration of ToPolyAgent acting as a research assistant specialized in coarse-grained simulations of topological polymers. The instructor can be either a human or another AI agent. | ||
For the linear polymer, we use ToPolyAgent to study the effect of solvent quality on conformational changes, particularly on the polymer's radius of gyration. Although the simulation module supports explicit solvent particles, we simplify the study by using an implicit solvent model, where solvent effects are represented by the polymer–polymer interaction parameter εpp, and the explicit solvent particle density is set to zero. We then provide ToPolyAgent with a series of prompt variations to perform simulations of linear polymers with different values of εpp.
Prompt:
Simulate a linear polymer consisting of 40 beads without solvent, using a polymer–polymer interaction parameter pp = {epsilon_pp}. Set the box size to 40, apply a Langevin thermostat, and run the simulation for 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/i_char_2009.gif) 000 steps.
000 steps.
We then compile all simulation results to investigate the effect of solvent quality—quantified by the polymer–polymer interaction parameter εpp—on the size of the polymer chain. As shown in Fig. 7, the polymer chain collapses as the interaction strength increases. Fig. 7(a) shows the squared radius of gyration Rg2 as a function of εpp. As expected, increasing attractive interactions lead to a reduction in overall chain size, resulting in smaller Rg2 values. Moreover, Fig. 7(b) characterizes the chain conformation through the slope of the form factor P(q). The slope of P(q) versus q in the log–log scale increases with εpp, indicating a transition from a self-avoiding walk at low εpp, to an ideal random walk at intermediate εpp, and finally to a collapsed globule at high εpp, resembling a fuzzy sphere. A small-angle neutron scattering (SANS) data for polystyrene solution is added for comparison. The SANS data is normalized using contour length 32.3 nm based on flexible cylinder model fitting. The SANS data is representative for good solvent with low εpp.
For the brush polymer, we use ToPolyAgent to study the effect of grafting density on the bending stiffness, characterized by the persistence length of the backbone. Similar to the linear polymer study, we provide ToPolyAgent with a series of variations of the following prompt to simulate brush polymers in a solvent bath with different grafting densities:
Prompt:
Simulate a brush polymer with 20 beads on the backbone, grafting density {grafting_density}, and side chain length 5, in a good solvent with number density 0.2. Use a box size of 30, apply a Langevin thermostat, and run the simulation for 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/i_char_2009.gif) 000 steps.
000 steps.
Fig. 8 shows the effect of grafting density σg on the conformation of the brush polymer. As the grafting density increases, the region near the backbone becomes increasingly crowded, resulting in reduced backbone flexibility. As shown in Fig. 8(a), the persistence length lp of the backbone chain increases with σg. Meanwhile, the form factor of the entire polymer, shown in Fig. 8(b), decreases more rapidly at higher grafting densities, indicating a transition from an extended polymer-like structure to a more compact, sphere-like morphology due to the dense side chains. This conformational transformation is illustrated by the snapshots of polymers with different grafting densities shown at the top of Fig. 8.
4 Summary
In this work, we introduce ToPolyAgent, a multi-agent AI system specialized in MD simulations of coarse-grained topological polymers. The ToPolyAgent system consists of four LLM-powered agents, each assigned specific tasks and equipped with dedicated tools to execute different stages of the simulation workflow. These agents are: the Config Agent, which generates the initial system configuration including the polymer and solvent; the Simulation Agent, which performs MD simulations and conducts conformational analyses of the results; the Report Agent, which collects logs and generated files from the simulation workflow and compiles a report in markdown format; and the Workflow Agent, which integrates the functions of the Config Agent and Simulation Agent.ToPolyAgent operates in two modes: interactive and autonomous. In the interactive mode, the Config Agent, Simulation Agent, and Report Agent are activated, with the Config and Simulation Agents equipped with a human-interaction tool that enables real-time communication with the user for revision and clarification. In the autonomous mode, the Workflow Agent and Report Agent automatically produce the final report directly from the user's initial prompt, providing a more streamlined simulation process.
We present examples of both interactive and autonomous modes, covering five common types of topological polymers: linear polymers, ring polymers, brush polymers, star polymers, and dendrimers. Finally, we demonstrate the potential of ToPolyAgent as a research assistant capable of handling more complex research tasks and automating parts of the scientific discovery process.
Building on the results demonstrated in this work, ToPolyAgent represents a new direction in computational materials science by seamlessly integrating LLM-enabled agentic AI systems with rigorous simulation tools. This integration not only extends the capabilities of existing AI frameworks but also enhances the practical utility of the simulation tools developed. By enabling researchers to interact with the system through natural language, ToPolyAgent significantly lowers the technical barriers to configuring and executing complex polymer simulations, thereby making advanced computational tools accessible to a broader range of users.
More importantly, ToPolyAgent's potential as a research assistant paves the way for autonomous scientific research in computational materials science. By combining ToPolyAgent with other specialized research agents, it becomes possible to construct a virtual research group composed entirely of AI agents with complementary expertise. For example, a discovery agent could identify research gaps addressable through topological polymer simulations by reviewing relevant literature; a planning agent could design a research plan involving specific sets of simulations, which are then executed by ToPolyAgent; and a writing agent could compile a well-structured research paper based on the simulation results. Human researchers would oversee and refine this process, analogous to ToPolyAgent's interactive mode, ensuring alignment with scientific objectives. By enabling scalable, collaborative, and automated research workflows, ToPolyAgent has the potential to accelerate discovery in polymer science and beyond, constrained only by available computational resources.
Looking forward, the development of ToPolyAgent opens several promising avenues for advancing computational materials science and AI-driven research. First, the modular design enables straightforward extensions to more complex polymer systems, including support for multiple chains in dense melts (e.g., pure polymer systems without solvent), multiple bead types with diverse interaction parameters for copolymers, and alternative potential forms (e.g., 10–12 Lennard-Jones potentials suitable for protein modeling). Integrating additional molecular simulation tools such as CHARMM66 and CHARMM-GUI,67 which also support coarse-grained modeling, could further broaden its applicability to real-world materials design challenges. These improvements will greatly increase the capability of the ToPolyAgent. Additionally, incorporating advanced input-generation capabilities from frameworks such as LUNAR52 and fine-tuned LLM53 could further enhance flexibility of the simulation workflow. Second, incorporating advanced analysis methods, such as machine learning models for interpreting polymer properties from simulation data via simulation-based inference,68–75 as well as adding simulation diagnosis tool to determine whether a system is sufficiently equilibrated could further streamline the research process by providing predictive insights alongside simulation results. Third, integrating LLM-based AI agents with simulation frameworks supporting Monte Carlo methods such as GOMC,76 SPARKS77 and HOOMD-blue,78 would further enhance the system's versatility. Finally, expanding ToPolyAgent's interoperability within a modular AI research ecosystem represents an important future direction. For example, coupling ToPolyAgent with agents specialized in experimental data analysis could enable direct comparison between simulated and experimental results, fostering deeper integration between computation and experiment.
Author contributions
LD, JMC and CD conceived this work; LD designed, implemented, and tested the agentic AI system; LD, JMC and CD wrote and edited the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The code for this work is available at the GitHub repository https://github.com/ljding94/ToPolyAgent with DOI: https://doi.org/10.5281/zenodo.18077373.Acknowledgements
This research was performed at the Spallation Neutron Source, which is a DOE Office of Science User Facilities operated by Oak Ridge National Laboratory. The beam time was allocated to EQ-SANS on proposal number IPTS-32932. The authors thank Polyxeni Angelopoulou Xanthopoulou, Logan Kearney, and Amit K. Naskar for providing the SANS data of the polystyrene solution. This research was sponsored by the Laboratory Directed Research and Development Program of Oak Ridge National Laboratory, managed by UT-Battelle, LLC, for the US DOE. Portions of the computational aspect of this research were supported by the Center for Nanophase Materials Sciences (CNMS), which is a U.S. Department of Energy Office of Science User Facility at Oak Ridge National Laboratory.Notes and references
- Y. Tezuka and H. Oike, Prog. Polym. Sci., 2002, 27, 1069–1122 CrossRef CAS.
- T. Yamamoto and Y. Tezuka, Polym. Chem., 2011, 2, 1930–1941 Search PubMed.
- K. Binder, Monte Carlo and molecular dynamics simulations in polymer science, Oxford University Press, 1995 Search PubMed.
- E. F. Casassa and Y. Tagami, Macromolecules, 1969, 2, 14–26 Search PubMed.
- J. Baschnagel, K. Binder, W. Paul, M. Laso, U. Suter, I. Batoulis, W. Jilge and T. Bürger, J. Chem. Phys., 1991, 95, 6014–6025 CrossRef CAS.
- B. J. Braams and D. E. Manolopoulos, J. Chem. Phys., 2006, 125, 124105 CrossRef PubMed.
- S. Habershon, D. E. Manolopoulos, T. E. Markland and T. F. Miller III, Annu. Rev. Phys. Chem., 2013, 64, 387–413 CrossRef CAS PubMed.
- M. Murat and G. S. Grest, Macromolecules, 1989, 22, 4054–4059 Search PubMed.
- C. Feng and X. Huang, Acc. Chem. Res., 2018, 51, 2314–2323 CrossRef CAS PubMed.
- S. T. Milner, T. A. Witten and M. E. Cates, Macromolecules, 1988, 21, 2610–2619 CrossRef CAS.
- S. T. Milner, Science, 1991, 251, 905–914 Search PubMed.
- J. M. Ren, T. G. McKenzie, Q. Fu, E. H. Wong, J. Xu, Z. An, S. Shanmugam, T. P. Davis, C. Boyer and G. G. Qiao, Chem. Rev., 2016, 116, 6743–6836 Search PubMed.
- G. S. Grest, K. Kremer and T. Witten, Macromolecules, 1987, 20, 1376–1383 Search PubMed.
- D. A. Tomalia, Sci. Am., 1995, 272, 62–66 Search PubMed.
- P. Kesharwani, K. Jain and N. K. Jain, Prog. Polym. Sci., 2014, 39, 268–307 Search PubMed.
- Y. K. Sung and S. W. Kim, Biomater. Res., 2020, 24, 12 CrossRef CAS PubMed.
- W. B. Liechty, D. R. Kryscio, B. V. Slaughter and N. A. Peppas, Annu. Rev. Chem. Biomol. Eng., 2010, 1, 149–173 Search PubMed.
- L. Y. Qiu and Y. H. Bae, Pharm. Res., 2006, 23, 1–30 Search PubMed.
- O. Pillai and R. Panchagnula, Curr. Opin. Chem. Biol., 2001, 5, 447–451 Search PubMed.
- C. S. Brazel and S. L. Rosen, Fundamental principles of polymeric materials, John Wiley & Sons, 2012 Search PubMed.
- L. J. Tan, W. Zhu and K. Zhou, Adv. Funct. Mater., 2020, 30, 2003062 Search PubMed.
- M. Fixman and W. H. Stockmayer, Annu. Rev. Phys. Chem., 1970, 21, 407–428 Search PubMed.
- R. Koningsveld, W. H. Stockmayer and E. Nies, Polymer phase diagrams: a textbook, OUP Oxford, 2001 Search PubMed.
- M. Watzlawek, C. N. Likos and H. Löwen, Phys. Rev. Lett., 1999, 82, 5289 Search PubMed.
- F. S. Bates, Science, 1991, 251, 898–905 Search PubMed.
- M. Müller, Phys. Rev. E, 2002, 65, 030802 CrossRef PubMed.
- T.-L. Liu, L.-Y. Liu, F. Ding and Y.-Q. Li, Chin. J. Polym. Sci., 2022, 40, 834–842 Search PubMed.
- M. P. Allen and D. J. Tildesley, Computer simulation of liquids, Oxford university press, 2017 Search PubMed.
- S. A. Hollingsworth and R. O. Dror, Neuron, 2018, 99, 1129–1143 Search PubMed.
- T. Hansson, C. Oostenbrink and W. van Gunsteren, Curr. Opin. Struct. Biol., 2002, 12, 190–196 CrossRef CAS PubMed.
- L. Ding, C.-H. Tung, B. G. Sumpter, W.-R. Chen and C. Do, J. Chem. Theory Comput., 2024, 20, 10697–10702 Search PubMed.
- F. Müller-Plathe, ChemPhysChem, 2002, 3, 754–769 Search PubMed.
- J. Zhao, L. Wu, C. Zhan, Q. Shao, Z. Guo and L. Zhang, Polymer, 2017, 133, 272–287 CrossRef CAS.
- B. Dünweg and K. Kremer, J. Chem. Phys., 1993, 99, 6983–6997 Search PubMed.
- A. P. Thompson, H. M. Aktulga, R. Berger, D. S. Bolintineanu, W. M. Brown, P. S. Crozier, P. J. In’t Veld, A. Kohlmeyer, S. G. Moore and T. D. Nguyen, et al. , Comput. Phys. Commun., 2022, 271, 108171 CrossRef CAS.
- D. Van Der Spoel, E. Lindahl, B. Hess, G. Groenhof, A. E. Mark and H. J. Berendsen, J. Comput. Chem., 2005, 26, 1701–1718 Search PubMed.
- R. Salomon-Ferrer, D. A. Case and R. C. Walker, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2013, 3, 198–210 CAS.
- E. Chow, C. A. Rendleman, K. J. Bowers, R. O. Dror, D. H. Hughes, J. Gullingsrud, F. D. Sacerdoti and D. E. Shaw, Simulation, 2008, 1, 1–14 Search PubMed.
- T. Brown, B. Mann, N. Ryder, M. Subbiah, J. D. Kaplan, P. Dhariwal, A. Neelakantan, P. Shyam, G. Sastry and A. Askell, et al. , Adv. Neural Inf. Process. Syst., 2020, 33, 1877–1901 Search PubMed.
- A. Radford, J. Wu, R. Child, D. Luan, D. Amodei, I. Sutskever, et al., OpenAI Blog, 2019, vol. 1, p. 9, https://storage.prod.researchhub.com/uploads/papers/2020/06/01/language-models.pdf.
- H. Naveed, A. U. Khan, S. Qiu, M. Saqib, S. Anwar, M. Usman, N. Akhtar, N. Barnes and A. Mian, ACM Trans. Intell. Syst. Technol., 2025, 16, 1–72 Search PubMed.
- D. B. Acharya, K. Kuppan and B. Divya, IEEE Access, 2025, 18912–18936 Search PubMed.
- M. Gridach, J. Nanavati, K. Z. E. Abidine, L. Mendes and C. Mack, arXiv, 2025, preprint arXiv:2503.08979, DOI:10.48550/arXiv.2503.08979.
- D. A. Boiko, R. MacKnight, B. Kline and G. Gomes, Nature, 2023, 624, 570–578 Search PubMed.
- Z. Shen, arXiv, 2024, preprint arXiv:2409.18807, DOI:10.48550/arXiv.2409.18807.
- T. Masterman, S. Besen, M. Sawtell and A. Chao, arXiv, 2024, preprint arXiv:2404.11584, DOI:10.48550/arXiv.2404.11584.
- J. Yang, C. E. Jimenez, A. Wettig, K. Lieret, S. Yao, K. Narasimhan and O. Press, Adv. Neural Inf. Process. Syst., 2024, 37, 50528–50652 Search PubMed.
- Y. Liu, P. Gao, X. Wang, J. Liu, Y. Shi, Z. Zhang and C. Peng, arXiv, 2024, preprint arXiv:2409.00899, DOI:10.48550/arXiv.2409.00899.
- S. Schmidgall, Y. Su, Z. Wang, X. Sun, J. Wu, X. Yu, J. Liu, M. Moor, Z. Liu and E. Barsoum, arXiv, 2025, preprint arXiv:2501.04227, DOI:10.48550/arXiv.2501.04227.
- O. A. Mendible-Barreto, M. Díaz-Maldonado, F. J. C. Esteva, J. E. Torres, U. M. Córdova-Figueroa and Y. J. Colón, Mol. Syst. Des. Eng., 2025, 10(7), 585–598 RSC.
- L. Ding and C. Do, arXiv, 2025, preprint arXiv:2509.05363, DOI:10.48550/arXiv.2509.05363.
- J. Kemppainen, J. R. Gissinger, S. Gowtham and G. M. Odegard, J. Chem. Inf. Model., 2024, 64, 5108–5126 CrossRef CAS PubMed.
- Z. Shi, C. Xin, T. Huo, Y. Jiang, B. Wu, X. Chen, W. Qin, X. Ma, G. Huang and Z. Wang, et al. , Sci. Rep., 2025, 15, 10295 Search PubMed.
- J.-M. Carrillo, Molecular Dynamics Simulation of Polymers, 2018, https://www.youtube.com/watch?v=cxwrdl6VbHM, Tutorial video with accompanying source code available at https://code.ornl.gov/jyw/cyber-training-summer-school, Supported by the U.S. National Science Foundation (NSF).
- J. Giedt, NSF Award Number 1841768. Directorate for Computer and Information Science and Engineering, 2018, vol. 18, p. 41768 Search PubMed.
- J. Achiam, S. Adler, S. Agarwal, L. Ahmad, I. Akkaya, F. L. Aleman, D. Almeida, J. Altenschmidt, S. Altman, S. Anadkatet al., arXiv, 2023, preprint arXiv:2303.08774, DOI:10.48550/arXiv.2303.08774.
- P.-G. De Gennes, Scaling concepts in polymer physics, Cornell university press, 1979 Search PubMed.
- M. Rubinstein and R. H. Colby, Polymer physics, Oxford university press, 2003 Search PubMed.
- S.-H. Chen, Annu. Rev. Phys. Chem., 1986, 37, 351–399 CrossRef CAS.
- P. Lindner and J. Oberdisse, Neutrons, X-rays, and light: scattering methods applied to soft condensed matter, Elsevier, 2024 Search PubMed.
- K. Kremer and G. S. Grest, J. Chem. Phys., 1990, 92, 5057–5086 Search PubMed.
- A. Brünger, C. L. Brooks III and M. Karplus, Chem. Phys. Lett., 1984, 105, 495–500 Search PubMed.
- B. Dünweg and W. Paul, Int. J. Mod. Phys. C, 1991, 2, 817–827 CrossRef.
- D. J. Evans and B. L. Holian, The Nose–Hoover thermostat, J. Chem. Phys., 1985, 83(8), 4069–4074 CrossRef CAS.
- M. E. Tuckerman, J. Alejandre, R. López-Rendón, A. L. Jochim and G. J. Martyna, J. Phys. A: Math. Gen., 2006, 39, 5629 Search PubMed.
- B. R. Brooks, C. L. Brooks III, A. D. Mackerell Jr, L. Nilsson, R. J. Petrella, B. Roux, Y. Won, G. Archontis, C. Bartels and S. Boresch, et al. , J. Comput. Chem., 2009, 30, 1545–1614 Search PubMed.
- S. Jo, X. Cheng, J. Lee, S. Kim, S.-J. Park, D. S. Patel, A. H. Beaven, K. I. Lee, H. Rui and S. Park, et al. , J. Comput. Chem., 2017, 38, 1114–1124 Search PubMed.
- L. Ding and C. Do, APL Mach. Learn., 2025, 3, year Search PubMed.
- L. Ding, Y. Chen and C. Do, Appl. Crystallogr., 2025, 58, 992–999 Search PubMed.
- L. Ding, C.-H. Tung, B. G. Sumpter, W.-R. Chen and C. Do, J. Appl. Crystallogr., 2025, 58, 1526–1532 CrossRef CAS PubMed.
- L. Ding, C.-H. Tung, J. M. Y. Carrillo, W.-R. Chen and C. Do, Digit. Discov., 2025, 4(8), 2075–2082 RSC.
- L. Ding, C.-H. Tung, Z. Cao, Z. Ye, X. Gu, Y. Xia, W.-R. Chen and C. Do, Digit. Discov., 2025, 4, 1570–1577 RSC.
- L. Ding, C.-H. Tung, B. G. Sumpter, W.-R. Chen and C. Do, J. Chem. Theory Comput., 2025, 21, 4176–4182 CrossRef CAS PubMed.
- C.-H. Tung, L. Ding, M.-C. Chang, G.-R. Huang, L. Porcar, Y. Wang, J.-M. Y. Carrillo, B. G. Sumpter, Y. Shinohara and C. Do, et al. , J. Chem. Phys., 2025, 162, 074106 Search PubMed.
- C.-H. Tung, L. Ding, G.-R. Huang, L. Porcar, Y. Shinohara, B. G. Sumpter, C. Do and W.-R. Chen, Appl. Crystallogr., 2025, 58, 523–534 Search PubMed.
- Y. Nejahi, M. S. Barhaghi, J. Mick, B. Jackman, K. Rushaidat, Y. Li, L. Schwiebert and J. Potoff, SoftwareX, 2019, 9, 20–27 Search PubMed.
- J. A. Mitchell, F. Abdeljawad, C. Battaile, C. Garcia-Cardona, E. A. Holm, E. R. Homer, J. Madison, T. M. Rodgers, A. P. Thompson and V. Tikare, et al. , Model. Simulat. Mater. Sci. Eng., 2023, 31, 055001 CrossRef.
- J. A. Anderson, J. Glaser and S. C. Glotzer, Comput. Mater. Sci., 2020, 173, 109363 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |







