Open Access Article
Open Access ArticleHarnessing generative AI for efficient organic materials discovery in low-data regimes
Jun Hyeong
Kim†
 a,
Kyunghoon
Lee†
a,
Kyunghoon
Lee†
 a,
Hyeonsu
Kim
ab,
MinSoo
Kang
c,
Suk-Ku
Chang
d,
Yinglan
Jin
d,
Dongwook
Kim
e and
Woo Youn
Kim
a,
Hyeonsu
Kim
ab,
MinSoo
Kang
c,
Suk-Ku
Chang
d,
Yinglan
Jin
d,
Dongwook
Kim
e and
Woo Youn
Kim
 *a
*a
aDepartment of Chemistry, KAIST, 291 Daehak-ro, Yuseong-gu, Daejeon 34141, Republic of Korea. E-mail: wooyoun@kaist.ac.kr
bSimulation Group, Samsung SDI, Samsung-ro, Yeongtong-gu, Suwon-si, 16678, Gyeonggi-do, Republic of Korea
cMaterial Development Department, SK Materials JNC, 5609 Dongtangiheung-ro, Hwa-sung City, 18469, Gyeonggi-do, Republic of Korea
dUNIPlus, 40 Omokcheon-ro 152beon-gil, Suwon-si, 16642, Gyeonggi-do, Republic of Korea
eDepartment of Chemistry, Kyonggi University, 154-42 Gwanggyosan-ro, Yeongtong-gu, Suwon-si, 16227, Gyeonggi-do, Republic of Korea. E-mail: dongwook-kim@kgu.ac.kr
First published on 2nd March 2026
Abstract
Generative AI has emerged as a powerful tool for the discovery of organic light-emitting diode (OLED) materials. However, its practical application remains underexplored due to the small datasets and difficulties in ensuring molecular synthesizability. To overcome these challenges, we introduce a building block-based autoregressive generative model. Trained on a dataset of approximately 1000 OLED molecules, the model demonstrated refined control over key thermally activated delayed fluorescence (TADF) properties, including S1 energy and the singlet–triplet energy gap ΔEST, while generating structurally novel candidates through strategic repurposing of building blocks not previously associated with TADF activity. In addition, we experimentally validated its potential by synthesizing four AI-designed green emitters and integrating them into OLED devices, achieving external quantum efficiencies of up to 11.22% at 1000 cd m−2. It achieved more than a 100-fold reduction in the computational cost of quantum chemical calculations compared to conventional heuristic methods. This work bridges the gap between generative molecular design and experimental realization, showcasing a pathway to overcome data scarcity and unlock innovative discovery of optoelectronic materials.
1 Introduction
High-throughput virtual screening (HTVS) has gained great attention as a powerful strategy for accelerating materials discovery.1–3 By rapidly exploring large chemical spaces with in silico methods, HTVS can facilitate the identification of promising candidates that would benefit from greater experimental validation.4,5 Numerous successful applications, confirmed by experiments, have demonstrated the potential of HTVS.6–11One of the critical factors that influence the HTVS process is the quality of the initial chemical library. When the library has a low density of desirable candidates, it is necessary to enlarge the library in order to identify valuable hits, which can eventually lead to substantial increase in computational costs. Unfortunately, such low densities are often inevitable when using simple strategies like random sampling, due to the complex and specific requirements of functional materials. Traditionally, these issues have been tackled by deriving expert-driven rules to capture key characteristics of materials tailored for specific applications.3 For example, photovoltaic and organic light-emitting diode (OLED) materials are typically designed through the combination of donor and acceptor building blocks to facilitate charge-transfer excitations.7,12–14 While these rule-based strategies can be effective, they are unlikely to capture all the key characteristics. Consequently, HTVS guided solely by expert heuristics still requires the screening of millions to billions of compounds to identify high-potential candidates.
In this context, generative AI has emerged as a promising alternative to overcome the limitations of expert rule-based design strategies. These models can learn the underlying distribution of training data and capture key features of data, enabling the design of libraries enriched with promising candidates.15 In recent years, generative models have achieved remarkable advancement in diverse fields, including image synthesis,16 natural language processing,17 and drug discovery.18 In materials discovery, generative AIs have been used to design novel functional materials.19–31 However, applying generative AI to materials discovery poses several unique challenges. Unlike in previous domains, where models are typically exposed to millions or even billions of data points, datasets relevant to materials discovery often contain just a few thousand, or even just a few dozen, data points.32,33 Furthermore, synthesizability must be ensured, which is less pertinent to language or image generation tasks.34,35 These requirements are further complicated by the multi-objective nature of materials discovery, including factors such as stability, functionality, and performance.36 As a result, the experimental validation of molecules designed by generative models is uncommon in the materials domain.
In this work, we present a successful application of generative AI to accelerate materials discovery, supported by experimental validation. Among various types of materials, we focused on thermally activated delayed fluorescence (TADF) materials,37–39 a domain characterized by limited data availability (approximately 1000 samples) and significant challenges when relying solely on expert-driven design rules, such as the combination of donor and acceptor building blocks.
To address these challenges, we used the Building Block–based AutoRegressive (BBAR) molecular generative model, which was originally developed for drug discovery.40 BBAR generates molecules by iteratively assembling molecular building blocks and has shown strong performance in generating candidates with desired properties given as conditions, including those that rarely appear in the training data. Furthermore, BBAR can implicitly consider synthesizability; by preparing the building blocks through retrosynthetic decomposition of known compounds and restricting fragment attachments to compatible connection sites defined by the same retrosynthetic templates, the generated molecules are more likely to be synthesizable.
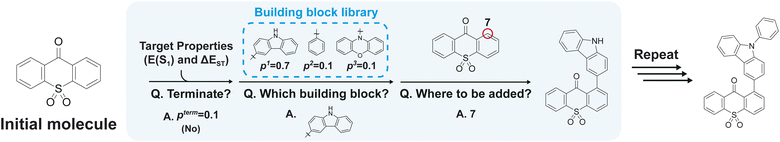
Leveraging the BBAR model, we synthesized four novel green TADF molecules and fabricated OLED devices to evaluate their performance. Fig. 1 illustrates the overall workflow of our study, which proceeded as follows. We first compiled a training set of approximately 1000 OLED molecules from various literature sources, focusing on molecular structures and their optoelectronic properties. Using this dataset, we trained the BBAR model to generate candidate molecules for red, green, and blue TADF emitters. The model produced a virtual chemical library of candidates projected to achieve significantly higher hit rates compared to traditional heuristic-based approaches, which was verified by extensive quantum chemical calculations. Targeting green emitters, we identified dozens of highly promising candidates for experimental validation. Since the BBAR model inherently considers synthesizability during generation, the selected candidates were readily synthesizable. This streamlined process culminated in four final molecules for device testing. Their device level evaluation showed EQEs of up to 11.22% at 1000 cd m−2, highlighting the potential of generative AI to bridge computational design and experimental realization, particularly in data-scarce scenarios.
2 Methodology
2.1 Preparation of the OLED dataset
To construct the training dataset for our BBAR model, we utilized molecular structures from the OLED database reported by Zhao et al.41 The dataset was originally developed for predicting thermal stability, specifically glass transition (Tg) and decomposition temperatures (Td), and thus does not include electronic properties relevant to TADF design. Therefore, we extracted the molecular structures (e.g., SMILES) and labeled two key electronic properties, the S1 energy and ΔEST, by performing quantum chemical calculations on all molecules.Here, the S1 energy represents the vertical excitation energy of a molecule transitioning from the ground state to the first excited singlet state (S1), which closely correlates with the display color range, and the ΔEST represents the energy difference between S1 and the first excited triplet state (T1), known as one of the critical factors for enhanced reverse intersystem crossing (RISC).42–46 These two electronic properties were utilized as the ground-truth generation conditions for training the BBAR model, with further details provided in Section 2.2.
To obtain these two properties, we first generated initial molecular geometries using the ETKDG algorithm,47 followed by universal force field (UFF) optimization,48 implemented in the RDKit library (RDKit: Open-source cheminformatics, https://www.rdkit.org). To reduce computational cost, these geometries were pre-optimized with the GFN2-xTB method49 and then optimized using DFT with the B3LYP functional50 and the 6-31g(d) basis set, as implemented in Gaussian 16.51 From the optimized ground-state geometries, the vertical excitation energies to the S1 state and T1 state were calculated using time-dependent DFT (TD-DFT) at the same level of theory. Out of 1182 molecules, 64 failed during the calculations, mainly due to optimization convergence failures, resulting in a final dataset of 1118 molecules. The distributions of their calculated electronic properties are provided in Section 1 of the SI.
2.2 Workflow of the BBAR model
Before presenting the main results, we briefly describe the workflow of the BBAR model used in this study (Fig. 2). The BBAR model is designed to generate candidate molecules that satisfy specified target property requirements, which are provided as input conditions. Starting from an initial building block, the model iteratively adds new building blocks until the resulting molecule achieves the desired properties. This conditional generation process involves three sequential steps. In the first step, the model determines whether the current molecule requires further modification to achieve the target properties. Multiple properties can be specified simultaneously, and BBAR considers them simultaneously during the generation process. Importantly, BBAR does not explicitly compute intermediate property values to make this decision. Instead, it predicts a termination probability, pterm, and stochastically samples whether to terminate based on this value. In the next step, if termination is not selected, the model chooses the next building block from a predefined library. Rather than treating this as a standard classification task over a fixed set of options, BBAR leverages graph representations of each building block to individually estimate likelihood scores (pi), indicating how likely each is to improve the molecule with respect to the desired properties. This workflow is a key innovation of the original BBAR model, as it allows the evaluation and incorporation of new or previously unseen building blocks during generation if they are deemed useful. This capability significantly expands the design space and opens new opportunities for innovation in materials discovery, an advantage that will be discussed in more detail below. In the final step, the model selects an atomic attachment site on the current molecule for appending the chosen building block. Specifically, it predicts a probability distribution over all candidate attachment sites conditioned on the current molecule and the target properties. One site (marked as a red circle in Fig. 2) is then selected according to this probability distribution, and the building block is attached at the selected site. The resulting molecule is then used as the input for the next iteration of the three-step cycle.The BBAR model is trained on sequences representing the stepwise assembly of molecular building blocks. These sequences are prepared by first decomposing OLED molecules in the training set using the BRICS (Breaking of Retrosynthetically Interesting Chemical Substructures) algorithm.52 Specifically, we utilized the BRICS implementation in RDKit, which applies 16 retrosynthetically inspired chemical decomposition patterns to partition molecules into synthetically meaningful units. Applying this procedure to the 1118 molecules in the training set yielded a total of 5905 initial fragments. After removing duplicates via canonicalization, we established a finalized library of 589 unique building blocks, which serves as the fundamental component set for molecular generation. Next, random assembly orders of these building blocks are enumerated. For example, if a molecule is composed of three connected fragments (A–B–C), possible training sequences might include A → A–B → A–B–C or C → B–C → A–B–C. These sequences are randomly sampled during training. The BBAR model is trained to reconstruct the original molecules by following the selected assembly sequence, conditioned on the calculated target properties (S1 energy and ΔEST). After several training epochs, the model is then used to generate new molecules tailored to specific property requirements.
We randomly split the dataset into training, validation, and test sets with a ratio of 75![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 15
15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10, and trained the model for 20 epochs. Details on the model architecture and training hyperparameters are provided in Section 2 of the SI. Additional technical information, including the training objective function, can be found in the original work by Seo et al.40
10, and trained the model for 20 epochs. Details on the model architecture and training hyperparameters are provided in Section 2 of the SI. Additional technical information, including the training objective function, can be found in the original work by Seo et al.40
3 Results
3.1 Tuning S1 energy and ΔEST
We first assessed the performance of the trained model in the conditional generation of TADF candidates. During generation, we set the target ΔEST to a low value of 0.1 eV to induce high RISC rates and set the target S1 energies to 1.87 eV, 2.23 eV, and 2.73 eV, which correspond to the red, green, and blue emission ranges, respectively. For each pair of target properties, 1000 candidate molecules were generated. These molecules were first validated using the RDKit library, and the S1 energies and ΔEST were calculated for the validated molecules using the same procedure described in Section 2.1.For comparison, we prepared a baseline set of molecules using the “random enumeration” method as described by Kim et al.5 This protocol follows the combinatorial assembly of predefined donors, linkers, and acceptors originally curated by Gómez-Bombarelli et al.,7 specifically forming donor–linkern–acceptor structures with n = 0, 1, 2. Considering all possible combinations results in a pool of approximately 5.49 million candidates, from which we randomly sampled 1000 molecules for testing. These selected molecules were then validated using the same procedure described above. This baseline set is referred to as the “Random” set.
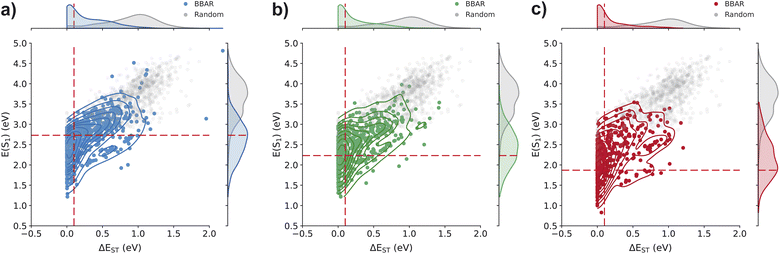
Fig. 3 shows the distributions of S1 energy and ΔEST for the generated molecules targeting blue, green, and red colors, respectively. The red dashed lines indicate the target values for each property in each task. As shown in the figure, a large portion of the randomly enumerated molecules lie far from the target values, with substantial variance. In particular, the distributions of both S1 and ΔEST are approximately Gaussian, centered away from the desired regions, reflecting the inherent randomness of the enumeration process. This not only suggests that heuristic-based design strategies alone are insufficient to guide efficient TADF discovery, but also implies that such strategies may bias exploration toward irrelevant regions of the chemical space.
In contrast, the BBAR-generated molecules exhibit property distributions that are more closely aligned with the target values. For ΔEST, the distributions consistently peak near zero across all color targets, which is a favorable condition for TADF. For S1 energy, although the distributions remain relatively broad, the peak positions shift downward from blue to red, closely tracking the respective target values. These results indicate that the trained generative model can effectively guide exploration of TADF-relevant chemical space, even when trained on a relatively small dataset of approximately 1000 molecules.
Table 1 summarizes the detailed hit rates for each generation task. We considered a molecule to be a hit if its S1 energy falls within 1.6–2.0 eV for red, 2.17–2.5 eV for green, or 2.5–2.75 eV for blue emission, and its ΔEST value is below 0.2 eV. On average, 61.4% of the BBAR-generated molecules satisfy the ΔEST threshold, while only 6.3% for the randomly enumerated molecules. This again confirms the earlier observation that heuristic donor–acceptor combinations alone are insufficient to consistently generate molecules with low ΔEST values.
| Random enumeration | BBAR | |||||
|---|---|---|---|---|---|---|
| ΔESTa | E(S1)b | Bothc | ΔESTa | E(S1)b | Bothc | |
| a Number of molecules satisfying ΔEST < 0.2 eV. b Number of molecules satisfying S1 energy criteria: 2.5–2.75 eV (blue), 2.17–2.5 eV (green), and 1.6–2.0 eV (red). c Number of molecules satisfying both ΔEST and S1 energy criteria. Percentages indicate the proportion relative to all successfully validated candidates. | ||||||
| Blue | 57 (6.3%) | 15 (1.7%) | 8 (0.9%) | 363 (53.0%) | 134 (19.6%) | 83 (12.1%) |
| Green | 57 (6.3%) | 16 (1.8%) | 11 (1.2%) | 379 (60.4%) | 147 (23.4%) | 104 (16.6%) |
| Red | 57 (6.3%) | 1 (0.1%) | 1 (0.1%) | 339 (70.8%) | 144 (30.1%) | 127 (26.5%) |
From the perspective of emission color, the difference is even more pronounced. On average, 24.4% of BBAR-generated molecules fall within the desired S1 energy ranges, whereas only around 1% of the randomly enumerated molecules meet this condition. This larger discrepancy arises because donor–acceptor heuristics are primarily effective in promoting low ΔEST (i.e., enhancing RISC), but are less suited for fine-tuning emission color. One can see that the hit rate for red-emitting molecules with the random enumeration set is particularly low, a mere 0.1%. This is likely because the donor and acceptor fragments used in the original work were primarily prepared for developing blue TADF materials. However, even for the blue emission task, the hit rate remains low at 1.7%, underscoring the limitations of heuristic-based design. On the other hand, the BBAR model has achieved substantially higher hit rates for all emission colors without access to any prior domain knowledge. When both criteria were considered simultaneously, BBAR, as expected, achieved much higher hit rate than the random enumeration method: on average, 18.4% of BBAR-generated molecules were hits, compared to just 0.73% for the random enumeration.
In addition to the hit rate analysis, we evaluated the chemical diversity of the generated molecules, as summarized in SI Tables 2 and 3 (see Section 3 of the SI). Across several metrics, the results indicate high chemical diversity. The novelty scores were close to 1, and the generated molecules exhibited a wide range of scaffolds and building blocks not prevalent in the training set. These findings confirm that BBAR effectively avoids structural redundancy while exploring a broad and diverse chemical space to generate molecules with the desired properties. We refer the reader to the SI for a detailed analysis.
3.2 Exploring TADF-relevant space with BBAR
We visualized the chemical space of the evaluated sets using the t-distributed stochastic neighbor embedding (t-SNE) algorithm.53 We used Morgan fingerprints54 with a radius of 2 and a bit length of 1024 for visualization. Fig. 4 presents the t-SNE plot of molecules generated by the BBAR model (referred to as “BBAR”) and those generated via random enumeration (referred to as “Random”). For additional context, we included two reference sets: “TADF (train),” consisting of molecules from the training set with ΔEST < 0.2 eV (indicative of TADF potential), and “TADF (ref. 7),” comprising four blue TADF emitters reported by Gómez-Bombarelli et al.7 To balance the number of molecules between the “BBAR” and “Random” sets, we additionally sampled 2000 molecules for the “Random” set.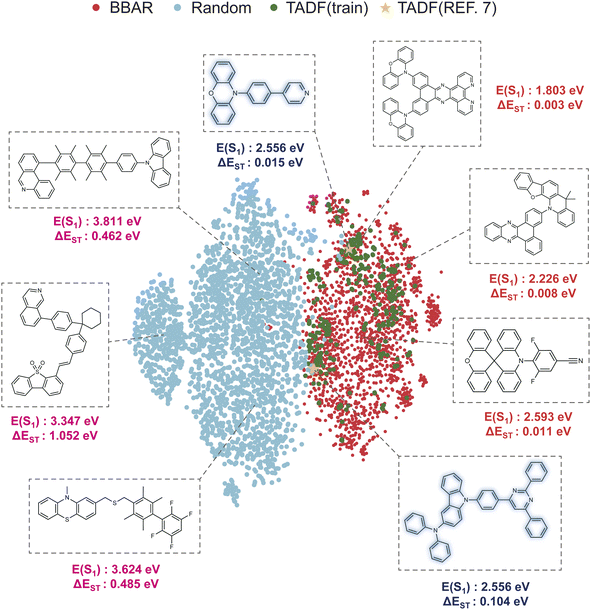 | ||
| Fig. 4 Visualization of the chemical space using t-SNE. Four sets of molecules, molecules generated by BBAR (BBAR), molecules satisfying ΔEST < 0.2 eV from the training set (TADF (train)), randomly enumerated molecules (Random), and TADF materials in Gómez-Bombarelli et al.7 (TADF (ref. 7)) are displayed in the figure. The BBAR-generated candidates near the desired TADF materials demonstrate the model's ability to focus on exploring the desired chemical space. Annotated structures highlight representative molecules with their corresponding S1 energy (E(S1)) and ΔEST values. The pink labels represent randomly enumerated molecules with high ΔEST and S1 energies outside the visible range. The dark blue labels correspond to TADF materials from ref. 7, while the red labels denote BBAR-generated molecules with very low ΔEST and S1 energies within the visible range. | ||
Fig. 4 reveals two key findings. First, the “BBAR” set shows substantial overlap with the “TADF (train)” set, significantly more than the “Random” set, indicating that the model successfully learned the property distributions of the training data. Second, the four experimentally reported TADF molecules (“TADF (ref. 7)”) are located within the same clusters as the training set, confirming that the training data is well suited for guiding the model in TADF material design. In contrast, the “Random” set (generated via donor–acceptor enumeration) shows clear separation from both the “BBAR” and “TADF (train)” clusters. This separation suggests that random heuristic methods often produce molecules with properties incompatible with viable TADF materials. For example, many “Random” molecules exhibit high ΔEST values (unfavorable for high RISC rates) and S1 energies outside the visible range (pink labels). The “BBAR” set, however, contains numerous candidates with low ΔEST and visible-range S1 energies (red labels), aligning with TADF requirements. This analysis underscores that, when well-suited building blocks are provided, BBAR can systematically explore a meaningful subset of TADF-relevant chemical space, representing a significant advancement over conventional heuristic strategies.
3.3 Qualitative observations of the generation process
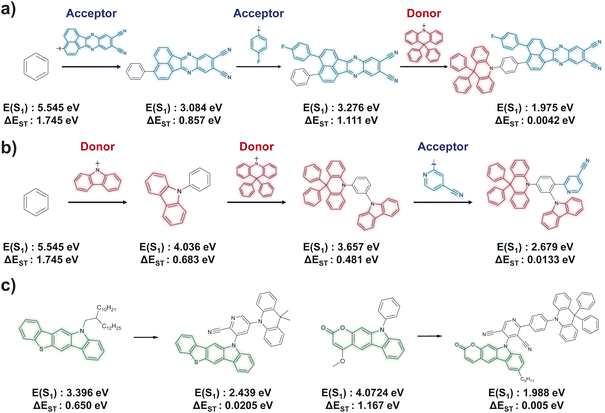
In this section, we highlight two notable behaviors of the BBAR model, observed by closely examining numerous generation processes. First, BBAR frequently adopts a donor–acceptor combination strategy. Fig. 5a and b illustrates two representative examples, where both start from a benzene scaffold but target different emission colors. When targeting a red-emitting TADF molecule (Fig. 5a), BBAR first adds two acceptor blocks (highlighted in blue), which lower the lowest unoccupied molecular orbital (LUMO) energy level. It then attaches a donor block (highlighted in red) to the opposite side, raising the highest occupied molecular orbital (HOMO) energy level. This combination decreases the S1 energy from 3.276 eV to 1.975 eV, falling within the red emission range, while simultaneously lowering ΔEST. Conversely, when designing blue-emitting TADF molecule (Fig. 5b), BBAR first introduces two donor blocks (red-colored fragments) to elevate the HOMO level, followed by an acceptor block (blue-colored fragment) with limited π-conjugation to maintain a relatively high LUMO energy level. This results in a molecule with blue emission and a low ΔEST.We believe this behavior stems from the fact that many of the OLED molecules in the training set were originally designed using conventional donor–acceptor combination strategies, which the BBAR model appears to have learned effectively. Remarkably, chemical concepts such as HOMO, LUMO, donor, or acceptor were never explicitly introduced to the model during training or generation. This suggests that BBAR was able to infer these design principles solely through its data-driven learning process. Beyond these simple heuristics, the model appears to have developed more sophisticated design strategies, as previously demonstrated by its focused exploration of TADF candidates. This distinction is likely to originate from fundamental differences in the generation process. Traditional heuristic approaches are essentially qualitative: they categorize fragments as either donors or acceptors and randomly combine one from each category, regardless of nuanced differences in electronic or structural properties. Defining more advanced design rules that account for such complexities is challenging to do manually. Therefore, these approaches often overlook the current molecular context and generate many unsuitable fragment combinations, leading to a low hit rate in TADF design.
In contrast, BBAR behaves more like a quantitative decision-making system. Given the molecular state and target properties, it flexibly evaluates both fragments and attachment positions as continuous probability values. This probabilistic reasoning allows BBAR to surpass heuristic strategies in TADF design, despite following a superficially similar donor–acceptor combination strategy. A more systematic interpretability analysis, such as statistically characterizing recurring decision patterns across generation processes, could potentially uncover new design principles beyond conventional heuristics. However, such quantitative analysis is challenging, particularly for the BBAR model, which was not designed with interpretability as a primary goal. Therefore, we leave this aspect for future work.
Another key observation is that BBAR can generate structurally novel candidates by incorporating unseen building blocks. Here, we define “unseen” as building blocks that do not appear in any TADF-active molecules within the training set, where TADF-active molecules are defined as those with ΔEST < 0.2 eV. Fig. 5c presents representative examples, where the green-highlighted components indicate such unseen blocks. Notably, these building blocks were initially present in non-TADF molecules, exhibiting large ΔEST values and S1 energies outside the visible range. However, BBAR successfully repurposes these blocks by combining them with other suitable units, shifting S1 into the visible region and significantly lowering ΔEST. This behavior highlights BBAR's ability to enhance chemical diversity by leveraging building blocks that were not previously associated with TADF-relevant properties in the training set. Conventional approaches such as random enumeration would entail substantial computational cost to achieve this, as expanding donor/acceptor fragment sets will exponentially increase the library size. Overall, these results demonstrate the dual advantages of generative AI to materials discovery: enabling efficient exploration of chemical space and facilitating the generation of structurally diverse candidates, even in low-data regimes (here, ∼1000 molecules).
3.4 Experimental validation of BBAR
Leveraging the strong performance of BBAR, we successfully synthesized four new green TADF materials. Of the 1000 BBAR-designed candidates targeting green emission, we first filtered out molecules with molecular weights above 1100 Da, leaving 518 candidates. After excluding molecules that failed quantum chemical calculations, 496 molecules remained. These were then screened using the following criteria: (1) to ensure green emission, only molecules with S1 energies between 2.17 and 2.50 eV were retained. (2) To ensure TADF-relevant properties, we selected molecules with ΔEST < 0.2 eV. (3) To confirm radiative fluorescence, we retained only those with nonzero oscillator strength (f(S1) > 0). (4) For suitable energy-level alignment in device applications, we selected molecules with HOMO energies above −6.0 eV and LUMO energies below −1.0 eV. Applying these filters yielded 72 candidates, a manageable set for human evaluation. Similarly, applying the same screening criteria to the blue and red sets yielded 48 blue and 65 red candidates, respectively.Although BBAR biases generation toward synthesizable structures by assembling building blocks, the generated molecules are not guaranteed to be readily synthesizable. This is mainly because (i) we did not fully dissect the commercial availability of all building blocks in the library, and (ii) even when the structures can be expressed as fragment assemblies, the corresponding synthetic routes may still be impractical under realistic experimental constraints. For these reasons, we performed retrosynthetic analysis on the green candidates using SciFindern55 to assess synthetic accessibility. As a result, we identified 17 viable candidates. Considering building-block availability and synthetic cost, we finally chose four candidates (G-1, G-2, G-3, and G-4) for device-level testing, as shown in Fig. 6a. Detailed synthetic procedures for each candidate are provided in the SI.
A common device architecture was used to characterize the four candidates under consistent conditions (Fig. 6b). Their photophysical and device-level properties are summarized in SI Table 4. Photoluminescence (PL) measurements in toluene confirmed that the emission wavelengths of the four candidates ranged from 515 to 542 nm, falling within the typical green emission range of 495–570 nm. The PL wavelengths were slightly red-shifted compared to the theoretical calculations, but this deviation was not significant. Consequently, BBAR was able to effectively guide the design of green-emitting TADF molecules. Transient PL decay measurements (Section 6 of the SI) further confirmed that all four candidates exhibited distinct TADF characteristics. Following device fabrication, the emission wavelengths exhibited a slight blue shift relative to their solution-state values. The corresponding electroluminescence spectra of the OLED devices are shown in Fig. 6c. Fig. 6d presents EQE as a function of current density, while Fig. 6e and f show current density and luminance as a function of voltage, respectively. The EQE values at 1000 cd m−2 were around 10% for all candidates, with G-4 achieving the highest performance at 11.22%. Remarkably, throughout the entire process, we considered only around 1000 candidates for quantum chemical evaluation, in contrast to a previous HTVS-based study7 that screened over one million molecules and performed quantum calculations on more than 400![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 to identify just four novel TADF emitters. Since model inference typically takes less than a second, which is negligible compared to DFT calculations as they often require tens to hundreds of minutes, this reduction represents more than a 100-fold improvement in discovery efficiency and underscores the potency of generative AI to accelerate materials discovery while successfully yielding experimentally validated materials with device-level performance.
000 to identify just four novel TADF emitters. Since model inference typically takes less than a second, which is negligible compared to DFT calculations as they often require tens to hundreds of minutes, this reduction represents more than a 100-fold improvement in discovery efficiency and underscores the potency of generative AI to accelerate materials discovery while successfully yielding experimentally validated materials with device-level performance.
4 Conclusions
In this work, we demonstrated the successful application of generative AI to overcome longstanding challenges in materials discovery—particularly in data-scarce domains such as thermally activated delayed fluorescence (TADF) emitters, where synthesizability and multi-property optimization pose significant hurdles. By training the BBAR model (a building block–based autoregressive framework) on OLED molecular data, we developed a generative system capable of designing TADF candidates with enhanced synthesizability and tailored optoelectronic properties. BBAR effectively optimized key TADF metrics, including S1 energy and ΔEST, achieving over a 100-fold increase in hit rate compared to expert-guided heuristic approaches. While BBAR mimics traditional donor–acceptor assembly in the generation process, it outperforms heuristic enumeration strategies by quantitatively evaluating building block suitability through data-driven likelihood estimation. Furthermore, the model could generate novel TADF candidates by repurposing building blocks not previously associated with TADF activity, expanding the accessible chemical space. With these advantages, we successfully designed four green-emitting TADF molecules at the device level, achieving external quantum efficiencies (EQEs) of up to 11.22% at 1000 cd m−2. These results underscore generative AI's transformative potential in molecular discovery: it enables efficient exploration of high-probability candidates while fostering structural diversity, even in data-limited scenarios.However, we note that our approach may still struggle in extremely low-data regimes (e.g., training sets with fewer than a couple of hundred examples) and should therefore be applied with caution in such scenarios. In these cases, we have observed that model performance can degrade substantially, even for relatively simple tasks such as log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P-conditioned molecule generation. To address such challenges, more advanced methodologies such as pre-training on large-scale datasets and employing improved model architectures will likely be necessary. We leave this as future work and consider it a key direction in materials discovery. Nonetheless, our work demonstrates a successful integration of computational design and experimental validation, highlighting its potential to accelerate the development of functional materials for next-generation optoelectronic applications.
P-conditioned molecule generation. To address such challenges, more advanced methodologies such as pre-training on large-scale datasets and employing improved model architectures will likely be necessary. We leave this as future work and consider it a key direction in materials discovery. Nonetheless, our work demonstrates a successful integration of computational design and experimental validation, highlighting its potential to accelerate the development of functional materials for next-generation optoelectronic applications.
Author contributions
J. H. K and K. L. contributed equally to this work. J. H. K. designed the methodology and experiments, performed DFT calculations, analyzed the results, and wrote the manuscript. K. L. contributed to the methodology design and experimental design, analyzed the results, and wrote the manuscript. H. K. contributed to part of the experimental design. S. C. and Y. J. synthesized the TADF candidates. M. K. fabricated the devices and measured their performance characteristics. W. Y.K. and D. K. conceived and supervised the project. All authors discussed the results and contributed to the manuscript revision.Conflicts of interest
The authors declare no competing financial interests.Data availability
The source data, source code, and scripts for TD-DFT calculations are available on GitHub at https://github.com/junhkim1226/TADF-Design and are also deposited in Zenodo under accession code https://doi.org/10.5281/zenodo.18297108.Supplementary information (SI): aditional dataset and model details, extended evaluation results, synthesis procedures for selected TADF candidates, and photophysical and device characterization data. See DOI: https://doi.org/10.1039/d5dd00463b.
Acknowledgements
This work was supported by the Technology Innovation Program (or Industrial Strategic Technology Development Program) (Grant No. 20016007, Development of phosphorescent TADF dopant with EQE 20% using AI platform) funded by the Ministry of Trade, Industry & Energy (MOTIE, Korea), and by the Basic Science Research Programs through the National Research Foundation of Korea funded by the Ministry of Science and ICT (Grant No. NRF-2023R1A2C2004376).Notes and references
- B. K. Shoichet, Nature, 2004, 432, 862–865 CrossRef CAS PubMed.
- S. Curtarolo, G. L. W. Hart, M. B. Nardelli, N. Mingo, S. Sanvito and O. Levy, Nat. Mater., 2013, 12, 191–201 CrossRef CAS PubMed.
- E. O. Pyzer-Knapp, C. Suh, R. Gómez-Bombarelli, J. Aguilera-Iparraguirre and A. Aspuru-Guzik, Annu. Rev. Mater. Res., 2015, 45, 195–216 CrossRef CAS.
- J. H. Kim, H. Kim and W. Y. Kim, Bull. Korean Chem. Soc., 2022, 43, 645–649 CrossRef CAS.
- H. Kim, K. Lee, J. H. Kim and W. Y. Kim, J. Chem. Inf. Model., 2024, 64, 677–689 CrossRef CAS PubMed.
- X. Qu, A. Jain, N. N. Rajput, L. Cheng, Y. Zhang, S. P. Ong, M. Brafman, E. Maginn, L. A. Curtiss and K. A. Persson, Comput. Mater. Sci., 2015, 103, 56–67 CrossRef CAS.
- R. Gómez-Bombarelli, J. Aguilera-Iparraguirre, T. D. Hirzel, D. Duvenaud, D. Maclaurin, M. A. Blood-Forsythe, H. S. Chae, M. Einzinger, D.-G. Ha, T. Wu, G. Markopoulos, S. Jeon, H. Kang, H. Miyazaki, M. Numata, S. Kim, W. Huang, S. I. Hong, M. Baldo, R. P. Adams and A. Aspuru-Guzik, Nat. Mater., 2016, 15, 1120–1127 CrossRef PubMed.
- Q. Yan, J. Yu, S. K. Suram, L. Zhou, A. Shinde, P. F. Newhouse, W. Chen, G. Li, K. A. Persson, J. M. Gregoire and J. B. Neaton, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 3040–3043 CrossRef CAS PubMed.
- C. Gorgulla, A. Boeszoermenyi, Z.-F. Wang, P. D. Fischer, P. W. Coote, K. M. Padmanabha Das, Y. S. Malets, D. S. Radchenko, Y. S. Moroz, D. A. Scott, K. Fackeldey, M. Hoffmann, I. Iavniuk, G. Wagner and H. Arthanari, Nature, 2020, 580, 663–668 CrossRef CAS PubMed.
- J. F. Joung, M. Han, J. Hwang, M. Jeong, D. H. Choi and S. Park, JACS Au, 2021, 1, 427–438 CrossRef CAS PubMed.
- D. H. Mok, H. Li, G. Zhang, C. Lee, K. Jiang and S. Back, Nat. Commun., 2023, 14, 7303 CrossRef CAS PubMed.
- J. Hachmann, R. Olivares-Amaya, S. Atahan-Evrenk, C. Amador-Bedolla, R. S. Sánchez-Carrera, A. Gold-Parker, L. Vogt, A. M. Brockway and A. Aspuru-Guzik, J. Phys. Chem. Lett., 2011, 2, 2241–2251 CrossRef CAS.
- J. H. Yun, J.-M. Kim, W. J. Chung, J. Lim, J. Y. Lee, Y. Lee and C. Choo, J. Mater. Chem. C, 2021, 9, 15242–15250 RSC.
- K. Thapa, J. I. Jones and L. E. Ratcliff, A High Throughput Virtual Screening Approach for Identifying Thermally Activated Delayed Fluorescence-Based Emitters, 2025, https://arxiv.org/abs/2505.10127.
- S. Bond-Taylor, A. Leach, Y. Long and C. G. Willcocks, IEEE Trans. Pattern Anal. Mach. Intell., 2022, 44, 7327–7347 Search PubMed.
- J. Li, C. Zhang, W. Zhu and Y. Ren, Ann. Data Sci., 2024, 12, 141–170 CrossRef.
- J. Li, T. Tang, W. X. Zhao, J.-Y. Nie and J.-R. Wen, ACM Comput. Surv, 2024, 56, 1–39 Search PubMed.
- A. Alakhdar, B. Poczos and N. Washburn, J. Chem. Inf. Model., 2024, 64, 7238–7256 CrossRef CAS PubMed.
- D. Schwalbe-Koda and R. Gómez-Bombarelli, Machine Learning Meets Quantum Physics, Springer International Publishing, Cham, 2020, pp. 445–467 Search PubMed.
- Y. Bian and X.-Q. Xie, J. Mol. Model., 2021, 27, 71 CrossRef CAS PubMed.
- J. Meyers, B. Fabian and N. Brown, Drug Discovery Today, 2021, 26, 2707–2715 CrossRef CAS PubMed.
- Z. Tan, Y. Li, Z. Zhang, X. Wu, T. Penfold, W. Shi and S. Yang, ACS Omega, 2022, 7, 18179–18188 CrossRef CAS PubMed.
- A. S. Fuhr and B. G. Sumpter, Front. Mater, 2022, 9, 865270 CrossRef.
- D. Menon and R. Ranganathan, ACS Omega, 2022, 7, 25958–25973 CrossRef CAS PubMed.
- C.-H. Li and D. P. Tabor, Chem. Sci., 2023, 14, 11045–11055 RSC.
- H. Park, Z. Li and A. Walsh, Matter, 2024, 7, 2355–2367 CrossRef CAS.
- M. Han, J. F. Joung, M. Jeong, D. H. Choi and S. Park, ACS Cent. Sci., 2024, 11, 219–227 CrossRef PubMed.
- S. Back, A. Aspuru-Guzik, M. Ceriotti, G. Gryn’ova, B. Grzybowski, G. H. Gu, J. Hein, K. Hippalgaonkar, R. Hormázabal, Y. Jung, S. Kim, W. Y. Kim, S. M. Moosavi, J. Noh, C. Park, J. Schrier, P. Schwaller, K. Tsuda, T. Vegge, O. A. von Lilienfeld and A. Walsh, Digital Discovery, 2024, 3, 23–33 RSC.
- K. Sattari, D. Li, B. Kalita, Y. Xie, F. B. Lighvan, O. Isayev and J. Lin, Digital Discovery, 2024, 3, 410–421 RSC.
- K. Yasuda, F. Berenger, K. Amaike, A. Ueda, T. Nakagomi, G. Hamasaki, C. Li, N. Y. Otani, K. Kaitoh, K. Tsuda, K. Itami and Y. Yamanishi, iScience, 2025, 28, 111526 Search PubMed.
- Y. Xu, Y. Luo, B. Li, W. Jiang, J. Zhang, J. Wei, H. Bai, Z. Wang, J. Ge, R. Lin, Z. Mi, H. Zhang, Y. Tang, M. S. Jones, X. Li, J. Z. Zhang and C.-W. Ju, JACS Au, 2025, 5, 3082–3091 CrossRef PubMed.
- B. Dou, Z. Zhu, E. Merkurjev, L. Ke, L. Chen, J. Jiang, Y. Zhu, J. Liu, B. Zhang and G.-W. Wei, Chem. Rev., 2023, 123, 8736–8780 CrossRef CAS PubMed.
- Q. Zhou, X. Chen and J. Wang, Acc. Mater. Res., 2025, 6, 685–694 CrossRef CAS.
- W. Gao and C. W. Coley, J. Chem. Inf. Model., 2020, 60, 5714–5723 CrossRef CAS PubMed.
- D. M. Anstine and O. Isayev, J. Am. Chem. Soc., 2023, 145, 8736–8750 CrossRef CAS PubMed.
- A. N. M. N. Abeer, N. M. Urban, M. R. Weil, F. J. Alexander and B.-J. Yoon, Patterns, 2024, 5, 101042 CrossRef PubMed.
- A. Endo, K. Sato, K. Yoshimura, T. Kai, A. Kawada, H. Miyazaki and C. Adachi, Appl. Phys. Lett., 2011, 98, 083302 CrossRef.
- H. Uoyama, K. Goushi, K. Shizu, H. Nomura and C. Adachi, Nature, 2012, 492, 234–238 CrossRef CAS PubMed.
- Q. Zhang, B. Li, S. Huang, H. Nomura, H. Tanaka and C. Adachi, Nat. Photonics, 2014, 8, 326–332 CrossRef CAS.
- S. Seo, J. Lim and W. Y. Kim, Advanced Science, 2023, 10, 2206674 CrossRef CAS PubMed.
- Y. Zhao, C. Fu, L. Fu, Y. Liu, Z. Lu and X. Pu, Mater. Today Chem., 2021, 22, 100625 CrossRef CAS.
- Y. Im, M. Kim, Y. J. Cho, J.-A. Seo, K. S. Yook and J. Y. Lee, Chem. Mater., 2017, 29, 1946–1963 CrossRef CAS.
- Z. Yang, Z. Mao, Z. Xie, Y. Zhang, S. Liu, J. Zhao, J. Xu, Z. Chi and M. P. Aldred, Chem. Soc. Rev., 2017, 46, 915–1016 RSC.
- K. Zhao, Ö. H. Omar, T. Nematiaram, D. Padula and A. Troisi, J. Mater. Chem. C, 2021, 9, 3324–3333 RSC.
- J. Kim, K. H. Lee and J. Y. Lee, Adv. Mater., 2023, 35, 2209953 CrossRef CAS PubMed.
- N. R and A. Mondal, J. Chem. Phys., 2025, 162, 144103 CrossRef CAS PubMed.
- S. Riniker and G. A. Landrum, J. Chem. Inf. Model., 2015, 55, 2562–2574 CrossRef CAS PubMed.
- A. K. Rappe, C. J. Casewit, K. S. Colwell, W. A. Goddard and W. M. Skiff, J. Am. Chem. Soc., 1992, 114, 10024–10035 CrossRef CAS.
- C. Bannwarth, S. Ehlert and S. Grimme, J. Chem. Theory Comput., 2019, 15, 1652–1671 Search PubMed.
- C. Lee, W. Yang and R. G. Parr, Phys. Rev. B:Condens. Matter Mater. Phys., 1988, 37, 785–789 CrossRef CAS PubMed.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, G. A. Petersson, H. Nakatsuji, X. Li, M. Caricato, A. V. Marenich, J. Bloino, B. G. Janesko, R. Gomperts, B. Mennucci, H. P. Hratchian, J. V. Ortiz, A. F. Izmaylov, J. L. Sonnenberg, D. Williams-Young, F. Ding, F. Lipparini, F. Egidi, J. Goings, B. Peng, A. Petrone, T. Henderson, D. Ranasinghe, V. G. Zakrzewski, J. Gao, N. Rega, G. Zheng, W. Liang, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, K. Throssell, J. A. Montgomery Jr, J. E. Peralta, F. Ogliaro, M. J. Bearpark, J. J. Heyd, E. N. Brothers, K. N. Kudin, V. N. Staroverov, T. A. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. P. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, J. M. Millam, M. Klene, C. Adamo, R. Cammi, J. W. Ochterski, R. L. Martin, K. Morokuma, O. Farkas, J. B. Foresman and D. J. Fox, Gaussiañ16 Revision B.01, Gaussian Inc. Wallingford CT, 2016 Search PubMed.
- J. Degen, C. Wegscheid-Gerlach, A. Zaliani and M. Rarey, ChemMedChem, 2008, 3, 1503–1507 CrossRef CAS PubMed.
- L. van der Maaten and G. Hinton, J. Mach. Learn. Res., 2008, 9, 2579–2605 Search PubMed.
- H. L. Morgan, J. Chem. Doc., 1965, 5, 107–113 CrossRef CAS.
- Chemical Abstracts Service (CAS), SciFinder-n, https://scifinder-n.cas.org/.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |