 Open Access Article
Open Access ArticleMeta-learning linear models for molecular property prediction
Yulia Pimonova *a,
Michael G. Taylorb,
Alice Allen
*a,
Michael G. Taylorb,
Alice Allen bcd,
Ping Yang
bcd,
Ping Yang *b and
Nicholas Lubbers
*b and
Nicholas Lubbers *a
*a
aComputing and Artificial Intelligence Division, Los Alamos National Laboratory, Los Alamos, NM 87545, USA. E-mail: ypimonova@lanl.gov; nlubbers@lanl.gov
bTheoretical Division, Los Alamos National Laboratory, Los Alamos, NM 87545, USA. E-mail: pyang@lanl.gov
cCenter for Nonlinear Studies, Los Alamos National Laboratory, Los Alamos, NM 87545, USA
dMax Planck Institute for Polymer Research, Ackermannweg 10, 55128 Mainz, Germany
First published on 13th May 2026
Abstract
Chemists in search of structure–property relationships face great challenges due to limited high quality, concordant datasets. Machine learning (ML) has significantly advanced predictive capabilities in chemical sciences, but these modern data-driven approaches have increased the demand for data. In response to the growing demand for explainable AI (XAI) and to bridge the gap between predictive accuracy and human comprehensibility, we introduce LAMeL—a Linear Algorithm for Meta-Learning that preserves interpretability while improving the prediction accuracy across multiple properties. While most approaches treat each chemical prediction task in isolation, LAMeL leverages a meta-learning framework to identify shared model parameters across related tasks, even if those tasks do not share data, allowing it to learn a common functional manifold that serves as a more informed starting point for new unseen tasks. Our method delivers up to 60–96% reduction in MAE over standard ridge regression, depending on the domain of the dataset. While the degree of performance enhancement varies across tasks, LAMeL consistently outperforms or matches traditional linear methods, making it a reliable tool for chemical property prediction where both accuracy and interpretability are critical.
1 Introduction
Machine learning has transformed how we approach complex chemical problems, delivering accurate predictions for molecular properties and chemical reactivity while reducing the need for both costly experimental evaluations and more affordable computational methods.1–3 The intersection of machine learning and chemistry manifests in new transformative approaches to molecular property prediction, materials design, and reaction conditions optimization.4–7 Over the years, increasingly sophisticated ML algorithms have demonstrated remarkable success in predicting chemical properties, ranging from solubility and drug-likeness to atomization energies and reaction dynamics.8–11 Despite these advances, the chemistry domain, along with other physical sciences, presents unique challenges that many common ML approaches struggle to fully address.Modern chemical machine learning faces constant competition between predictive power and interpretability. Deep neural networks, graph neural networks, and other complex architectures have achieved state-of-the-art performance across numerous chemical prediction tasks.12–14 Nevertheless, these models function largely as “black boxes,” making their decision-making processes opaque to human understanding.15 This interpretability challenge is particularly acute in chemistry, where understanding the underlying structure–property relationships is essential to continuous scientific innovation. Chemists have traditionally relied on transparent and mechanistically meaningful models that reveal how specific structural features influence molecular properties.16
On the other hand, linear models are inherently interpretable, which stems from their explicit parameter weights. The coefficients in linear models directly quantify the contribution of each feature, allowing for direct interpretation of prediction results. Although linear models often lag behind neural networks in terms of performance, their transparency and ease of interpretation are compelling incentives to use them, even when they are less accurate.17,18 Recent contribution by Allen and Tkatchenko demonstrates that, with an appropriate featurization scheme, multi-linear regression can achieve performance comparable to more advanced deep learning architectures in predicting materials properties.19 Moreover, linear regression models are faster than neural networks in terms of both training speed and computational resource requirements due to their much simpler design.
The widespread application of ML in the physical sciences meets a major challenge in the pervasive data scarcity in experimental studies. Acquiring chemical data—and, more broadly, any experimental data—is resource-intensive, time-consuming, and expensive. The problem is especially pronounced in drug design20–22 but extends across many areas of chemistry.23,24 When experimental data are scarce, combining low-fidelity simulation with limited high-fidelity experiments can improve accuracy and robustness, as shown by Nevolianis et al. for toluene–water partition coefficients.25 Similarly, Eraqi et al. have demonstrated that multi-task learning over multiple sustainable aviation fuel properties provided benefits in the ultra-low data regime with as few as 29 samples.26 The low-data problem becomes particularly critical when the demand for high-accuracy prediction is high.22,27
Meta-learning has emerged as a powerful framework to address data efficiency challenges across diverse machine learning domains. Unlike methods that treat each task independently, meta-learning seeks to “learn to learn” by leveraging shared structure across related tasks.28,29 This paradigm enables models to acquire transferable knowledge that facilitates rapid adaptation to new tasks, even in low-data regimes. Meta-learning distinguishes itself from other knowledge transfer frameworks such as transfer learning and multitask learning.28 While multitask learning focuses on simultaneously learning multiple tasks to perform well on those same tasks,30 meta-learning is designed to “learn how to learn,” enabling models to quickly adapt to entirely new tasks with minimal examples. This contrasts with transfer learning, which leverages knowledge from previously learned source tasks to enhance performance on a different target task through fine-tuning.31,32 The key distinction of meta-learning lies in emphasis on rapid adaptation to new tasks rather than just applying existing knowledge (transfer learning) or handling multiple known tasks concurrently (multitask learning). This distinction is particularly relevant when tasks are conceptually related but may not share the same datapoints, motivating approaches that leverage cross-task structure without requiring aligned samples. Meta-learning develops a learning capability that allows models to efficiently learn new information with few training examples. Recent studies have demonstrated the promise of meta-learning in chemistry-related applications. For instance, Allen et al.33 showed that incorporating multiple levels of quantum chemical theory within a unified training process can enhance prediction accuracy. Building on this promise, Wang et al.34 integrated meta-learning into the design of a foundation model for chemical reactors, while Singh and Hernández-Lobato applied prototypical networks35 to improve selectivity predictions along organic reaction pathways.27
Despite these advances, most existing meta-learning approaches emphasize deep learning architectures that prioritize predictive performance at the expense of interpretability. Qian et al.36 specifically highlight the lack of interpretability as a major limitation in their few-shot molecular property prediction model. In response, several efforts have aimed to improve interpretability. One strategy involves developing interpretable models that replicate the performance of deep networks, such as the approach proposed by Fabra-Boluda et al.37 More commonly, post-hoc interpretability techniques are employed. These include symbolic metamodels layered on top of neural networks,38 analyses of specific hidden layers,39 regression models based on architectural meta-features,40 and variance decomposition methods such as Meta-ANOVA.41
This limitation highlights a critical knowledge gap: the absence of application-oriented meta-learning algorithms specifically designed for linear models. While there is growing academic interest in this area, most existing efforts remain theoretical and lack practical application to real-world problems. For instance, Tripuraneni et al. introduced a provably sample-efficient algorithm for multi-task linear regression, focusing on learning shared low-dimensional representations across tasks.42 While their contribution offers strong theoretical proof, it does not address practical deployment challenges. Similarly, Denevi et al. proposed a conditional meta-learning approach that tailors representations to individual tasks using side information, offering improved adaptation in clustered task environments, yet their method has not been tested in applied settings.43 Toso et al. extended meta-learning to linear quadratic regulators using a model-agnostic approach, demonstrating theoretical guarantees for controller stability and adaptation, but their focus remains on control theory without broader application.44 These studies underscore the need for developing meta-learning algorithms for linear models that are not only theoretically sound but also practically applicable across diverse real-world domains.
To bridge the gap in applying meta-learning to linear models for chemical domain, we introduce LAMeL—a novel algorithm that reshapes meta-learning principles specifically for linear architectures. LAMeL learns shared parameters across related support tasks, identifying a common functional manifold that serves as an informed initialization for new, unseen tasks. The familiarized starting point enables the meta-model to adapt to new tasks with only a few data points. The presented method is motivated by recent theoretical work on shared low-dimensional structure in linear regression across tasks,42,43 but is designed for applied low-data settings where heterogeneous datasets with limited data overlap, and so missing labels are common. Fig. 1 illustrates the LAMeL workflow, showcasing how meta-learning is applied to linear models for chemical property prediction by leveraging support tasks to enhance performance on a target task. In this work we provide an applied meta-learning algorithm for interpretable linear chemistry models under task structures with minimal sample overlap and missing labels. A practical advantage of LAMeL is that it can operate directly on sparse, non-aligned task data, which is common in molecular property prediction.
The primary contributions of this work include:
• The development of LAMeL, the first meta-learning algorithm specifically designed for linear models in chemistry applications.
• A comprehensive validation of LAMeL across multiple chemical domains, demonstrating performance improvements ranging from 1.1 to 25-fold over classical ridge regression.
• An investigation into the role and importance of the task similarity across support data. By providing an ML tool that preserves interpretability while working in the low-data regime, LAMeL contributes to the broader goal of making ML-acquired results more valuable for chemistry.
2 Methods
2.1 Dataset descriptions
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) S ± 0.7 demonstrates state-of-the-art performance for physics-informed solubility models.
S ± 0.7 demonstrates state-of-the-art performance for physics-informed solubility models.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 944 solubility values from 1595 studies, creating one of the largest repository for non-aqueous solubility prediction. The dataset spans 1448 unique organic solutes and 213 solvents, with temperature-dependent measurements the range of 243–425 K. Each entry contains structures of solutes and solvents (as SMILES strings), experimental solubility values (as log-values of molarity), temperature, and bibliographic information for the originating study. The breadth and diversity of BigSolDB 2.0 make it a valuable source for benchmarking ML models of solubility.
944 solubility values from 1595 studies, creating one of the largest repository for non-aqueous solubility prediction. The dataset spans 1448 unique organic solutes and 213 solvents, with temperature-dependent measurements the range of 243–425 K. Each entry contains structures of solutes and solvents (as SMILES strings), experimental solubility values (as log-values of molarity), temperature, and bibliographic information for the originating study. The breadth and diversity of BigSolDB 2.0 make it a valuable source for benchmarking ML models of solubility.2.2 Substructural fingerprints
In cheminformatics, molecular representation is a critical step in translating chemical structures into a format suitable for regression tasks. There are numerous ways to accomplish this process, commonly referred to as fingerprinting the molecules.49 Substructure-based fingerprints and other large sparse molecular representations have been central to cheminformatics since their introduction and remain widely used due to their robustness, scalability, and direct mapping to chemically meaningful motifs.50–52 In this work, we use graphlet fingerprints, a direct topological representation that has demonstrated effectiveness in molecular property prediction.53 Graphlet fingerprints have emerged as a powerful tool for molecular representation in cheminformatics, offering a balance between structural granularity and computational efficiency.53,54Graphlets operate on the molecular graph in which atoms serve as nodes and bonds as edges. Graphlets are formed from the isomorphism classes of connected subgraphs in the molecular graph; a one-node graphlet constitutes a single atom, a two-node graphlet constitutes two bonded atoms and the associated bond type, and so on for higher-sized molecular fragments. Using graphlet representations in molecular property prediction builds upon the many-body expansion principle in quantum chemistry,55 where properties are approximated as sums of contributions from increasingly complex atomic clusters. The fingerprinting process involves systematically enumerating all graphlets within a molecule up to a predefined maximum graphlet size. Fig. 2 illustrates this process for acetone, where all graphlets up to size 5 are extracted from the molecular graph. Unlike path-based56 or radial fingerprints,57 graphlets capture every kind of substructure, providing a more complete encoding of molecular topology as they identify every possible subgraph. A fast, recursive hashing procedure allows identifying the isomorphism class of each graphlet.
The set of all graphlets in a given dataset can then be assembled as a feature matrix, giving counts of each type of substructure in each molecule in the dataset. This preserves an interpretable relationship between molecular components and their contributions to predicted properties. In our meta-learning framework, model coefficients correspond directly to specific graphlet substructures, and as a result, meta-learned models preserve the interpretability of the graphlet featurization approach. It stands to reason that the structured organization of the features might facilitate knowledge transfer across tasks as it mimics the structure that human chemists use to build chemical intuition.
2.3 Meta-learning background
Meta-learning, often described as “learning to learn,” is an approach in machine learning where the goal is to develop models that can rapidly adapt to new tasks by leveraging experience gained from a collection of related tasks. In the meta-learning framework, we typically distinguish between support tasks Ti, i ∈ {1…T} – a set of training tasks used to learn meta-knowledge – and target tasks T★, which are novel tasks where the meta-learned knowledge is applied to achieve fast adaptation and improved performance.Each task Ti is characterized by its own dataset, consisting of task features XT and corresponding labels yT. The meta-learning is advantageous over other knowledge-transfer approaches as it does not need the same data point to appear across multiple tasks; rather, each task can have its own unique data distribution and labeling. This flexibility enables meta-learning to handle a diverse range of tasks with sparsely distributed data.
The meta-learning algorithm presented in this work can be considered an optimization-based approach, where the meta-learner aims to find model or initialization parameters that facilitate rapid adaptation to new tasks with minimal data. During meta-training, the model is exposed to multiple support tasks, learning to optimize its parameters such that, when presented with a target task, it can quickly fine-tune to achieve the best performance with minimal data. This approach is particularly effective in scenarios characterized by limited labeled data for new tasks. In such cases, the target task is inherently data-constrained, with only a small number of data points available for training. We refer to these data points as shots, following the convention established in the few-shot learning literature,58,59 where the objective is to achieve robust generalization from a minimal number of training examples. Implementation details, hyperparameter settings, and AutoML tuning budgets for the nonlinear (LightGBM60) and task-conditioned pooled ridge baselines are provided in the SI.
2.4 LAMeL: linear algorithm for meta-learning
Our meta-learning approach is aimed at building a linear specialization coefficients β* with predictions . The model is decomposed into parallel and perpendicular components, that is, β* = β⊥ + β‖. The component β‖ is in-plane with the T-dimensional subspace W‖ generated by the support task coefficients βτ, τ ∈ {T1…TT}, whereas β⊥ is perpendicular to this subspace. We assume that, as tasks may have some relationship to each other, they may be approximated by a lower-rank manifold.42,61 As such, we bias the specialization coefficients β* towards the manifold defined by the models built on support tasks and allow for knowledge distillation from previous learning experiences.
. The model is decomposed into parallel and perpendicular components, that is, β* = β⊥ + β‖. The component β‖ is in-plane with the T-dimensional subspace W‖ generated by the support task coefficients βτ, τ ∈ {T1…TT}, whereas β⊥ is perpendicular to this subspace. We assume that, as tasks may have some relationship to each other, they may be approximated by a lower-rank manifold.42,61 As such, we bias the specialization coefficients β* towards the manifold defined by the models built on support tasks and allow for knowledge distillation from previous learning experiences.
While many forms of bias are possible, we use sequential fitting, which has the advantage of separating out hyperparameter searches. First, we fit within the subspace W‖, and subsequently use this start space to find a residual (intuitively, smaller) component β⊥. We use a ridge loss function as a base regressor.
The first step is to build individual support models by minimizing the support task loss
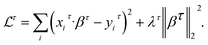 | (1) |
• 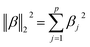 is the squared L2 norm of the regression parameters vector.
is the squared L2 norm of the regression parameters vector.
• λ ≥ 0 is the regularization parameter controlling the strength of the penalty.
• τ is each of the support tasks in {T1,…, TT}.Ridge regression encourages smaller coefficient values for model's stability and generalization.62
Next, building features which explore W‖ is a matter of dotting the specialization feature matrix X* with the model matrices, yielding new meta-features, X*·βτ, which are the prediction of the support tasks applied to the data in the target task. Thus we see explicitly how meta-learning can operate on disjoint support and target tasks: the models from a support task can still be applied to the target task, and we can use the results of these models as features for forming a new prediction. We build the meta-features using the average support task as the origin for fitting β‖. Setting an origin for fitting encourages the learned coefficients β‖ to stay close to the origin (prior) vector, effectively embedding information from previous learning experiences into the model (Fig. 3). This adjustment aligns with the principles of meta-learning, where knowledge from support tasks informs the learning process for a new task. By incorporating a task-specific or meta-learned prior βprior, this approach enforces the adaptability of linear models in scenarios with limited available data, as the prior knowledge can mitigate over-fitting and improve generalization and transferability to new tasks.
We center these features χiτ = (βτ − β)·χi using the average support model  as an origin for residual fitting. We then build β‖ by finding c ∈
as an origin for residual fitting. We then build β‖ by finding c ∈ ![[Doublestruck R]](https://www.rsc.org/images/entities/char_e175.gif) T minimizing the ridge loss function
T minimizing the ridge loss function
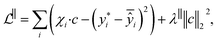 | (2) |
 . This yields the parallel component of the model,
. This yields the parallel component of the model,
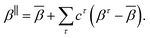 | (3) |
![[script L]](https://www.rsc.org/images/entities/char_e144.gif) ‖ is formally degenerate as there are T coefficients and T − 1 independent features – however, the resulting β‖ is well-defined.) Finally, the residual coefficient β⊥ is found by minimizing the ridge loss function of the residuals εi = yi − xiβ‖ given by
‖ is formally degenerate as there are T coefficients and T − 1 independent features – however, the resulting β‖ is well-defined.) Finally, the residual coefficient β⊥ is found by minimizing the ridge loss function of the residuals εi = yi − xiβ‖ given by
 | (4) |
1. Determine support coefficients βτ using the support task data, and construct meta-features χi by applying these models to the target task features.
2. Determine parallel coefficients β‖ using the meta-features χi.
3. Determine perpendicular coefficients β⊥ using the ordinary features xi. The final parameter vector for the specialization task after the few shot learning is:
| β* = β⊥ + β‖ | (5) |
3 Results and discussion
We evaluate the proposed meta-learning method on datasets selected to probe generalization and scalability. Our objectives are to test sample efficiency on small problems, assess robustness when the number of prediction tasks is large but per-task supervision is sparse, and analyze performance when both task count and per-task data volume are high. We consider three regimes: a small benchmark with only a few tasks;45 a large multitask benchmark in which each task provides a single label per sample;47 and a large benchmark with many tasks but limited observations per task.46 This spectrum allows us to examine how the model behaves as the task set grows, as per-task data increases or decreases, and as distributional heterogeneity increases, providing a balanced basis for the comparisons reported below. To support full reproducibility, we provide the code of LAMeL and all experiments in a public repository: https://github.com/lanl/minervachem. The repository includes a runnable Jupyter notebook demonstrating end-to-end usage (data loading, training, evaluation, and interpretation), together with scripts to reproduce the results reported in this manuscript.We evaluated a lightweight nonlinear baseline LightGBM60 and a pooled linear baseline based on joint ridge regression with task conditioning to contextualize LAMeL among other popular methods. The selected baselines represent the closest direct comparisons that can be trained on the observed data without introducing a separate missing-data model. LightGBM does not improve over LAMeL in the low-data N-shot regime, while joint task-conditioned ridge shows mixed behavior. In our experiments, the joint regression outperforms LAMeL when the target task is closely aligned with the pooled training data, as seen for acetone and methyl acetate. LAMeL is more useful when the target task is low-data or less typical of the pooled distribution. Practically, we recommend joint regression for interpolation within a family of closely related tasks, and to prefer LAMeL for few-shot transfer to new or chemically atypical tasks, especially when coefficient-level interpretability is desired. We note that joint task conditioning can shift predictive signal from chemically meaningful substructure features to task-identity features, reducing interpretability. Additionally, extending to unseen tasks requires refitting the pooled model. Full implementation details, hyperparameters, and per-task curves are reported in the SI.
3.1 Solubility database by Boobier et al.
As described above, the Boobier et al.45 dataset includes solubility data for the four solvents: water, ethanol, acetone, benzene. Given only four tasks, in our meta-learning procedure we tested each of the solvents as a target task with the remaining three being used as support tasks. An extra test was performed to ensure the target task data does not leak into the support tasks data used in the meta-learning phase. Detailed results for individual solvents at different maximum substructure sizes used for fingerprinting are provided in SI. These analyses further illustrate how molecular representation influences prediction accuracy across tasks.The results of our experiments reveal moderate improvements in prediction accuracy when employing meta-learning, with gains diminishing as the number of shots increases across all target tasks (Fig. 4). Notably, predictions for water solubility showed no improvement across all shot sizes. We primarily attribute this lack of improvement to the chemical distinctiveness of water compared to the other solvents (ethanol, acetone, and benzene), which serve as support tasks in this case. Water has the highest dielectric constant of (ε = 80.1) in comparison with the rest: ethanol (ε = 25.3), acetone (ε = 21.0) and benzene is around (ε = 2.27). Water's more polarizing nature and formation of hydrogen-bonding likely reduce its similarity to the other solvents, limiting meta-learning's ability to transfer knowledge from support to target tasks. This observation underscores a key limitation of meta-learning: task similarity among support tasks plays an important role in effective knowledge transfer. When support tasks are chemically or structurally dissimilar to the target task, meta-learning exhibits limited ability to exploit common patterns across tasks.
3.2 BigSolDB 2.0
Encouraged by the positive results obtained with the Boobier dataset, we applied meta-learning on the largest solubility dataset, BigSolDB 2.0, comprising approximately 104k datapoints for 1448 compounds across 213 solvents with temperature variation. In this work we did not want to consider temperature a variable, so in the pre-processing stage for each solvent–solute pair we kept a single entry from the temperature window between 290 K and 300 K, all other datapoints were disregarded. There are 70 unique solvents in the remaining data. Furthermore, we can filter the solvents (tasks) based on the total number of datapoints per task. To see how the total number of available solvent changes with imposed limitations on the data size, navigate to Fig. S5 in the SI. Similarly to the Boobier et al.45 dataset, water is significantly different from all other solvents with average dielectric constants differences (Δε) reaching 66 ± 12 for water-containing pairs and only 13 ± 11 for all other solvent pairs. The list of dielectric constant values for solvents in BigSolDB2.0 after preprocessing is available in Table S1.As before, in experimental setup all of the tasks that are not the target task have been used as support in the meta-training stage. We observed performance improvement with meta-learning for all but one solvent, water. To quantify the effect we calculated the relative improvement
 | (6) |
In the true few-shots regime (10–30 training datapoints for the target task) solvents exhibit substantial relative improvements, with achieving up to 60% MAE reduction. This high-variance, high-reward region demonstrates the typical few-shot learning behavior where limited data can yield significant performance gains for well-suited systems. A convergence pattern emerges as relative improvements gradually decrease and stabilize with the increasing number of training points. The high variability observed in the low-shot regime diminishes with more consistent model performance across different solvents. Most solvents converge to a plateau region with relative improvements within 15–30%. Similarly to Boobier et al. dataset, water exhibits consistently negative relative improvement (−10% to 0%) across all shot counts, suggesting fundamental incompatibility with the underlying support tasks. To investigate the individual roles of the parallel and perpendicular components of the parameter vector for the specialization task, we performed ablations which removed either perpendicular (β* = β‖) or parallel (β* = β⊥) contribution. Across solvents, β‖ captures most of the gain, while β⊥ becomes important when the target task is weakly aligned with available support tasks. Full ablation details and per-solvent curves are provided in the SI.
Additionally, we investigated the effect of the number of tasks alone by randomly selecting fixed number of support tasks from a consistent pool, details are provided in the SI. Across different numbers (3, 5, or 10) of randomly chosen tasks, LAMeL performance is very similar, suggesting that the meta-learning gains are not strongly tied to a specific choice of support-set size.
The stark contrast between the two similarity matrices—particularly water's extremely low similarity with other solvents in regression space (average of 0.07 with all support tasks)—provides quantitative support for our hypothesis regarding the role of task similarity for the effectiveness of meta-learning.
However, for the Boobier et al. dataset, it is important to consider dataset size alongside chemical similarity of solvents. The total number of data points per solvent varies (Table 2), with water being the solvent with the largest number of observations. Using water as the target reduces the total points available in its support set if compared to other targets. For example, when ethanol is the target, its support tasks contribute 2348 data points, whereas for water the corresponding total is 1611. While 1611 is substantial in many low-data settings, the smaller support set for water—together with water's chemical dissimilarity to the other solvents—likely contributes to its substantially weaker meta-learning performance. Meta-learning relies primarily on task similarity between the target and support tasks; however, when support tasks are fewer transferable common knowledge might be harder to exploit.
| Max size | Boobier et. al. | BigSolDB2.0 | QM9-MultiXC |
|---|---|---|---|
| 3 | 319 | 380 | 125 |
| 5 | 4992 | 5194 | 3280 |
| 7 | 57![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 346 346 |
58![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 365 365 |
82![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 942 942 |
| Tasks | N datapoints |
|---|---|
| Water | 1432 |
| Ethanol | 695 |
| Benzene | 464 |
| Acetone | 452 |
This dual explanation—chemical distinctiveness and limited support dataset size—provides a more nuanced understanding of why water solubility predictions fail to improve with meta-learning. It highlights two critical caveats in applying meta-learning to small-task datasets:
1. Task similarity remains a prerequisite for effective knowledge transfer.
2. Small support dataset sizes can negatively affect meta-learning performance.
3.3 QM9-MultiXC
In contrast to the experimental solubility datasets, the QM9-MultiXC data contains a large number of tasks and an abundance of datapoints per task. The computational methods here can be categorized into three groups based on the basis set used: SZ, DZ, and TZ. It is widely accepted that higher zeta levels provide more accurate results but require more computational resources.63 As the size of the investigated systems grows, high-zeta basis set calculations become increasingly inaccessible. To test the applicability of linear meta-learning to the highly localized chemical properties, we apply our LAMeL in various scenarios. First, we investigate whether a limited number of SZ functionals can provide sufficient support for LAMeL to predict SZ, DZ, and TZ targets. Additionally, we vary the depth of the fingerprinting process during the featurization step. Unlike the moderate accuracy improvements observed with solubility datasets, this setup reveals a dramatic reduction in error when applying meta-learning, as evident in Fig. 9. The most considerable improvements in accuracy were observed in extremely low-shot regimes, where as few as 10 datapoints were used for training on the target task. The color gradient in Fig. 9 highlights increasing maximum substructure size, here errors increase and R2 decreases for non-meta approaches. This behavior contrasts with previous results for solving linear models with minervachem topological fingerprinting.53 However, this is not a true contradiction and arises in this work due to the few-shot nature of experiments. As maximum substructure size increases, the feature vector size grows significantly (see Table 1), while the total number of datapoints remains unchanged. In few-shot regimes, deeper fingerprinting leads to overparameterization given the relatively small molecules in QM9 datasets. For small molecules (no more than nine heavy atoms) a deep featurization can introduce many repeating units that increase non-linearly with depth. It increases dimensionality without adding additional information and worsens ill-conditioning. In high-dimensional settings where the number of features far exceeds the number of observations, even regularized methods like ridge regression can struggle to generalize effectively. For example, training models with 125, 3280, and 82![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 942 parameters on a dataset containing only 50 datapoints leads to significant increases in test error as feature vector complexity grows. In overparameterized scenarios, the model has capacity to capture noise in the training data rather than meaningful patterns only, leading to overfitting. While ridge regression mitigates overfitting by shrinking coefficients, it does not eliminate the problem entirely. This reflects the bias–variance trade-off: with more parameters bias decreases but variance increases substantially, especially in high-dimensional spaces where small perturbations in the data can lead to large changes in predictions. Empirical studies have shown that standard regularization techniques may become less effective in these scenarios unless paired with additional strategies such as dimensionality reduction or adaptive regularization.62,64
942 parameters on a dataset containing only 50 datapoints leads to significant increases in test error as feature vector complexity grows. In overparameterized scenarios, the model has capacity to capture noise in the training data rather than meaningful patterns only, leading to overfitting. While ridge regression mitigates overfitting by shrinking coefficients, it does not eliminate the problem entirely. This reflects the bias–variance trade-off: with more parameters bias decreases but variance increases substantially, especially in high-dimensional spaces where small perturbations in the data can lead to large changes in predictions. Empirical studies have shown that standard regularization techniques may become less effective in these scenarios unless paired with additional strategies such as dimensionality reduction or adaptive regularization.62,64
The results presented in Fig. 9 highlight the interplay between feature complexity and predictive performance in few-shot meta-learning scenarios. Contrary to the non-meta approach results, the meta-learning framework maintains relatively stable error levels. For the illustrated TZ target, where high-fidelity predictions are inherently more challenging given their absence from the support tasks, meta-learning achieves comparable MAEs between different substructure depths and demonstrates some resilience against overparameterization by avoiding the sharp fall in performance observed in non-meta models. The comparative stability of error across substructure sizes in meta-learning highlights its effectiveness in balancing the bias-variance trade-off, even in few-shot regimes with high-dimensional representations.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 055 per task), providing a comprehensive basis for predictions. To further investigate the effect of support data size, we created subsets at 10, 106, 1064, 5322, 10
055 per task), providing a comprehensive basis for predictions. To further investigate the effect of support data size, we created subsets at 10, 106, 1064, 5322, 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 644, 21
644, 21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 288, 42
288, 42![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 577, 85
577, 85![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 115 datapoints sampled from the total available support data (106
115 datapoints sampled from the total available support data (106![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 444 datapoints). Interestingly, across all three target tasks (M06-L_SZ, TPSSH_DZP, and MPBE0KCIS_TZP), the relationship between meta-assisted accuracy improvement and support data size remained consistent. As illustrated in Fig. 10, the meta-learning error metrics remained relatively stable for support dataset sizes ranging from 1064 to 106
444 datapoints). Interestingly, across all three target tasks (M06-L_SZ, TPSSH_DZP, and MPBE0KCIS_TZP), the relationship between meta-assisted accuracy improvement and support data size remained consistent. As illustrated in Fig. 10, the meta-learning error metrics remained relatively stable for support dataset sizes ranging from 1064 to 106![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 444 datapoints. This observation suggests that even a small fraction of the large QM9-MultiXC dataset (approximately 1%) is sufficient to maintain meta-learning efficiency improvements. Nevertheless, when the support sample size dropped below 1%, error metrics consistently increased across all tested shot sizes for the target task. While meta-learning is robust to reductions in support data size within reasonable limits, extremely small datasets compromise its ability for knowledge extraction. Notably, these findings are different from our observations from solubility datasets, where task similarity played a significant role in determining meta-learning performance. Here, the abundance of data in QM9-MultiXC mitigates some of the challenges posed by task dissimilarity, enabling effective knowledge transfer even with limited support task similarity.
444 datapoints. This observation suggests that even a small fraction of the large QM9-MultiXC dataset (approximately 1%) is sufficient to maintain meta-learning efficiency improvements. Nevertheless, when the support sample size dropped below 1%, error metrics consistently increased across all tested shot sizes for the target task. While meta-learning is robust to reductions in support data size within reasonable limits, extremely small datasets compromise its ability for knowledge extraction. Notably, these findings are different from our observations from solubility datasets, where task similarity played a significant role in determining meta-learning performance. Here, the abundance of data in QM9-MultiXC mitigates some of the challenges posed by task dissimilarity, enabling effective knowledge transfer even with limited support task similarity.
The meta-assisted accuracy improvement depends more on the number of shots of the target task than on the size of the support data. An interesting case arises with the M06-L_SZ target task, where accuracy improvements from meta-learning show minimal sensitivity to shot size variations (apart from NS = 5, which performs noticeably worse). This behavior aligns with our hypothesis regarding task similarity: since all results for this target task were generated using five random SZ-based functionals as support tasks, their inherent similarity facilitates efficient knowledge transfer regardless of shot size.
These results highlight meta-learning's potential for enabling high-fidelity molecular energy predictions using lower-fidelity tasks as support—even under constrained data scenarios—while also emphasizing critical limitations when datasets become extremely sparse.
4 Conclusions
In this study, we developed and evaluated a linear meta-learning algorithm for molecular property prediction. Our approach leverages dispersed small-scale data through task-adaptive knowledge transfer without sacrificing interpretability. We assessed performance across three dataset regimes aligned with our objectives: small multitask datasets, large multitask datasets with small per-task data, and large datasets with many tasks and moderate per-task data. We have shown the independent linear meta-learners to use sparse data effectively and improve predictive performance when task similarity is high; under heavy per-task sparsity, meta-learning gains remain, but diminish as similarity between tasks decreases. When both task count and per-task data are high, the proposed method is competitive while retaining interpretability. Overall, our results suggest that dataset structure primarily affects the magnitude of meta-learning benefits, rather than the specific molecular properties (i.e. solubility, atomization energy).For the solubility datasets, meta-learning yielded up to a 60% increase in accuracy compared to conventional ridge regression. The magnitude of improvement was closely tied to the degree of similarity among support tasks. In both solubility datasets, water—being the most chemically distinct solvent—stood out as the sole case where meta-learning did not overcome baseline accuracy, highlighting the critical role of support task similarity for successful knowledge transfer. Our results demonstrate that the linear meta-learning framework achieves solubility prediction errors that are on par with those reported for deep learning models. For the nine most popular solvents in the BigSolDB 2.0 dataset (Fig. 6), the mean absolute error (MAE) can be consistently reduced to below 0.800 LogS units, with the lowest MAE observed at 0.683 ± 0.007 for n-propanol. This level of accuracy is comparable with literature: Ulrich et al. report an experimental uncertainty of 0.5–0.6 log units and an ML model with RMSE of 0.657 for aqueous solubility,65 MolMerger achieves an average MAE of 0.79 LogS units across solute–solvent pairs,66 AttentiveFP67 and MoGAT,68 both limited to aqueous systems, report RMSE values of 0.61 and 0.478 log units, respectively, while SolPredictor69 reaches an average RMSE of 1.09 log units for aqueous solubility. The ability of our linear meta-learning approach to deliver comparable predictive performance across a chemically diverse set of solvents supports its practical utility in real-world solubility prediction tasks with minimal available data.
For the atomization energy dataset, which involves highly localized electronic properties, linear meta-learning provided the largest relative gains, further supporting the applicability of the method to various tasks and settings. Our study demonstrates the data efficiency achieved by the meta-learning framework: accurate predictions were obtained using as little as 1% (i.e. 1064 datapoints per support task) of the full training data in the QM9-MultiXC dataset, demonstrating the potential of this method for scenarios where data collection is expensive or time-consuming. These findings suggest that meta-learning not only interpolates between tasks but also captures underlying physical and chemical principles, enabling interpretative extrapolation even in low-data regimes.
While the linear nature of the model constrains its capacity to capture complex relationships, its simplicity allows for robust and interpretable performance. LAMeL complements recent neural and symbolic meta-learners by providing a lightweight and interpretable method for small regression datasets. Future work should explore the integration of nonlinear meta-learners for interpretability, the extension of linear meta-learning approach to more chemically diverse and challenging systems, and the incorporation of active learning strategies to further enhance data efficiency and predictive power. Our method can serve as a linear baseline for benchmarking and can be combined with nonlinear learning algorithms, for example, as an interpretable linear head on learned embeddings.70,71
Overall, our results establish linear meta-learning as a powerful and computationally efficient paradigm for molecular property prediction. Beyond improving predictive performance, LAMeL provides a coefficient-level mapping that for graphlet fingerprints highlights which substructural motifs contribute most strongly to a prediction for a given task. The obtained knowledge can guide experimental prioritization by selecting candidate molecules enriched in beneficial motifs for a property of interest, and it can be used to support fragment-based molecular design.72–74 By enabling significant accuracy gains with minimal data, the presented method holds promise for accelerating high-throughput screening and materials discovery, particularly in domains where experimental resources are limited and quick adaptation is essential.
Author contributions
Yulia Pimonova: methodology, software, investigation, formal analysis, visualization, writing – original draft. Michael G. Taylor: supervision, software, writing – review & editing. Alice Allen: conceptualization, supervision, writing – review & editing. Ping Yang: conceptualization, supervision, resources, writing – review & editing, funding acquisition. Nicholas Lubbers: conceptualization, methodology, software, supervision, resources, writing – review & editing.Conflicts of interest
There are no conflicts to declare.Data availability
Data used in this study were obtained from published external datasets: by Boobier et al.,75 accessed on June 1st, 2023; BigSolDB 2.0,76 accessed on August 1st, 2025; and QM9-MultiXC,77 accessed on June 1st, 2023. All datasets are currently publicly available and free of charge. All supporting data, including detailed individual results, reference data for solvents, and classical (non-meta) performance results, are provided in the supplementary information (SI). The code for LAMeL, along with scripts for data preprocessing, has been archived at https://doi.org/10.5281/zenodo.19834714. The archived code corresponds to LAMeL version v1. The development repository is openly available at https://github.com/lanl/minervachem. Supplementary information is available. See DOI: https://doi.org/10.1039/d5dd00443h.Acknowledgements
We acknowledge the support by the U.S. Department of Energy, Office of Science, Office of Basic Energy Sciences, Heavy Element Chemistry Program at Los Alamos National Laboratory (LANL) (Y. P., M. T., P. Y., N. L.) (contract no KC0302031 LANL2023E3M2). We gratefully acknowledge the support of the U.S. Department of Energy through the LANL Laboratory Directed Research Development Program under project number 20250637DI for this work (Y. P., N. L.). This research used resources provided by the CAI-1 Darwin HPC cluster at LANL. Los Alamos National Laboratory is operated by Triad National Security, LLC, for the National Nuclear Security Administration of the U.S. Department of Energy (contract no. 89233218CNA000001). This research used resources of National Energy Research Scientific Computing Center (NERSC), a Department of Energy Office of Science User Facility using NERSC award BES-ERCAP0023367 and BES-ERCAP0033948.Notes and references
- J. A. Keith, V. Vassilev-Galindo, B. Cheng, S. Chmiela, M. Gastegger, K.-R. Müller and A. Tkatchenko, Chem. Rev., 2021, 121, 9816–9872 CrossRef CAS PubMed.
- J. Lan, X. Li, Y. Yang, X. Zhang and L. W. Chung, Acc. Chem. Res., 2022, 55, 1109–1123 CrossRef CAS PubMed.
- A. Aldossary, J. A. Campos-Gonzalez-Angulo, S. Pablo-García, S. X. Leong, E. M. Rajaonson, L. Thiede, G. Tom, A. Wang, D. Avagliano and A. Aspuru-Guzik, Adv. Mater., 2024, 36, 2402369 CrossRef CAS PubMed.
- A. N. Marimuthu and B. A. McGuire, J. Chem. Inf. Model., 2025, 65, 5424–5437 CrossRef CAS PubMed.
- Y. Zheng, H. Y. Koh, J. Ju, A. T. N. Nguyen, L. T. May, G. I. Webb and S. Pan, Nat. Mach. Intell., 2025, 7, 437–447 CrossRef.
- S.-C. Li, H. Wu, A. Menon, K. A. Spiekermann, Y.-P. Li and W. H. Green, J. Am. Chem. Soc., 2024, 146, 23103–23120 CrossRef CAS PubMed.
- D. J. Burrill, C. Liu, M. G. Taylor, M. J. Cawkwell, D. Perez, E. R. Batista, N. Lubbers and P. Yang, J. Chem. Theory Comput., 2025, 21, 1089–1097 CrossRef CAS PubMed.
- M. Amin Ghanavati, S. Ahmadi and S. Rohani, Digital Discovery, 2024, 3, 2085–2104 RSC.
- G. W. Kyro, A. Morgunov, R. I. Brent and V. S. Batista, J. Chem. Inf. Model., 2024, 64, 653–665 CrossRef CAS PubMed.
- N. Fedik, W. Li, N. Lubbers, B. Nebgen, S. Tretiak and Y. Wai Li, Digital Discovery, 2025, 4, 1158–1175 RSC.
- S. Zhang, M. Chigaev, O. Isayev, R. A. Messerly and N. Lubbers, J. Chem. Inf. Model., 2025, 65, 4367–4380 CrossRef CAS PubMed.
- Z. Yang, M. Chakraborty and A. D. White, Chem. Sci., 2021, 12, 10802–10809 RSC.
- E. Heid, K. P. Greenman, Y. Chung, S.-C. Li, D. E. Graff, F. H. Vermeire, H. Wu, W. H. Green and C. J. McGill, J. Chem. Inf. Model., 2024, 64, 9–17 CrossRef CAS PubMed.
- D. M. Anstine, R. Zubatyuk and O. Isayev, Chem. Sci., 2025, 16, 10228–10244 RSC.
- C. Rudin, Nat. Mach. Intell., 2019, 1, 206–215 CrossRef PubMed.
- G. P. Wellawatte and P. Schwaller, Commun. Chem., 2025, 8, 11 CrossRef PubMed.
- C. Ventura, D. A. R. S. Latino and F. Martins, Eur. J. Med. Chem., 2013, 70, 831–845 CrossRef CAS PubMed.
- Z. Cai, M. Zafferani, O. M. Akande and A. E. Hargrove, J. Med. Chem., 2022, 65, 7262–7277 CrossRef CAS PubMed.
- A. E. A. Allen and A. Tkatchenko, Sci. Adv., 2022, 8, eabm7185 CrossRef CAS PubMed.
- H. Altae-Tran, B. Ramsundar, A. S. Pappu and V. Pande, ACS Cent. Sci., 2017, 3, 283–293 CrossRef CAS PubMed.
- D. Vella and J.-P. Ebejer, J. Chem. Inf. Model., 2023, 63, 27–42 CrossRef CAS PubMed.
- K. Zhang, Z. Fan, Q. Wu, J. Liu and S.-Y. Huang, J. Chem. Inf. Model., 2025, 65, 7174–7192 Search PubMed.
- G. Tom, R. J. Hickman, A. Zinzuwadia, A. Mohajeri, B. Sanchez-Lengeling and A. Aspuru-Guzik, Digital Discovery, 2023, 2, 759–774 RSC.
- B. C. Haas, D. Kalyani and M. S. Sigman, Sci. Adv., 2025, 11, eadt3013 CrossRef PubMed.
- T. Nevolianis, J. G. Rittig, A. Mitsos and K. Leonhard, J. Cheminf., 2025, 17, 123 Search PubMed.
- B. A. Eraqi, D. Khizbullin, S. S. Nagaraja and S. M. Sarathy, Commun. Chem., 2025, 8, 201 CrossRef PubMed.
- S. Singh and J. M. Hernández-Lobato, Nat. Commun., 2025, 16, 3599 CrossRef CAS PubMed.
- A. Vettoruzzo, M.-R. Bouguelia, J. Vanschoren, T. Rögnvaldsson and K. C. Santosh, Advances and Challenges in Meta-Learning: A Technical Review, arXiv, 2023, preprint, arXiv:2307.04722, DOI:10.48550/arXiv.2307.04722.
- M. Huisman, J. N. van Rijn and A. Plaat, Artif. Intell. Rev., 2021, 54, 4483–4541 CrossRef.
- R. Caruana, Mach. Learn., 1997, 28, 41–75 CrossRef.
- F. Zhuang, Z. Qi, K. Duan, D. Xi, Y. Zhu, H. Zhu, H. Xiong and Q. He, Proc. IEEE, 2021, 109, 43–76 Search PubMed.
- J. S. Smith, B. T. Nebgen, R. Zubatyuk, N. Lubbers, C. Devereux, K. Barros, S. Tretiak, O. Isayev and A. E. Roitberg, Nat. Commun., 2019, 10, 2903 CrossRef PubMed.
- A. E. A. Allen, N. Lubbers, S. Matin, J. Smith, R. Messerly, S. Tretiak and K. Barros, npj Comput. Mater., 2024, 10, 1–9 Search PubMed.
- Z. Wang and Z. Wu, Towards Foundation Model for Chemical Reactor Modeling: Meta-Learning with Physics-Informed Adaptation, arXiv, 2024, preprint, arXiv:2405.11752, DOI:10.48550/arXiv.2405.11752.
- J. Snell, K. Swersky and R. Zemel, Prototypical Networks for Few-shot Learning, Advances in Neural Information Processing Systems, ed. I. Guyon, U. Von Luxburg, S. Bengio, H. Wallach, R. Fergus, S. Vishwanathan and R. Garnett, Curran Associates, Inc., 2017, vol. 30, pp. 4077–4087 Search PubMed.
- X. Qian, B. Ju, P. Shen, K. Yang, L. Li and Q. Liu, ACS Omega, 2024, 9, 23940–23948 CrossRef CAS PubMed.
- R. Fabra-Boluda, C. Ferri, J. Hernández-Orallo, M. J. Ramrez-Quintana and F. Martnez-Plumed, Intell. Data Anal., 2025, 29, 28–44 Search PubMed.
- A. M. Alaa and M. van der Schaar, Demystifying Black-box Models with Symbolic Metamodels, Advances in Neural Information Processing Systems, ed. H. Wallach, H. Larochelle, A. Beygelzimer, F. d'Alché-Buc, E. Fox and R. Garnett, Curran Associates, Inc., 2019, vol. 32, pp. 11304–11314 Search PubMed.
- X. Liu, X. Wang and S. Matwin, Interpretable Deep Convolutional Neural Networks via Meta-learning, arXiv, 2018, preprint, arXiv:1802.00560, DOI:10.48550/arXiv.1802.00560.
- G. T. Pereira, I. B. Santos, L. P. Garcia, T. Urruty, M. Visani and A. C. De Carvalho, Inf. Sci., 2023, 649, 119642 Search PubMed.
- Y. Choi, S. Park, C. Park, D. Kim and Y. Kim, J. Korean Stat. Soc., 2025, 54, 478–495 CrossRef.
- N. Tripuraneni, C. Jin and M. I. Jordan, Provable Meta-Learning of Linear Representations, arXiv, 2022, preprint, arXiv:2002.11684, DOI:10.48550/arXiv.2002.11684.
- G. Denevi, M. Pontil and C. Ciliberto, Conditional Meta-Learning of Linear Representations, arXiv, 2021, preprint, arXiv:2103.16277, DOI:10.48550/arXiv.2103.16277.
- L. F. Toso, D. Zhan, J. Anderson and H. Wang, Meta-Learning Linear Quadratic Regulators: A Policy Gradient MAML Approach for Model-free LQR, arXiv, 2024, preprint, arXiv:2401.14534, DOI:10.48550/arXiv.2401.14534.
- S. Boobier, D. R. J. Hose, A. J. Blacker and B. N. Nguyen, Nat. Commun., 2020, 11, 5753 Search PubMed.
- L. Krasnov, D. Malikov, M. Kiseleva, S. Tatarin, S. Sosnin and S. Bezzubov, Sci. Data, 2025, 12, 1236 CrossRef CAS PubMed.
- S. Nandi, T. Vegge and A. Bhowmik, Sci. Data, 2023, 10, 783 CrossRef CAS PubMed.
- R. Ramakrishnan, P. O. Dral, M. Rupp and O. A. von Lilienfeld, Sci. Data, 2014, 1, 140022 CrossRef CAS PubMed.
- D. S. Wigh, J. M. Goodman and A. A. Lapkin, Wiley Interdiscip. Rev. Comput. Mol. Sci., 2022, 12, e1603 CrossRef.
- D. Rogers and M. Hahn, J. Chem. Inf. Model., 2010, 50, 742–754 CrossRef CAS PubMed.
- A. Cereto-Massagué, M. J. Ojeda, C. Valls, M. Mulero, S. Garcia-Vallvé and G. Pujadas, Methods, 2015, 71, 58–63 CrossRef PubMed.
- T.-H. Nguyen-Vo, P. Teesdale-Spittle, J. E. Harvey and B. P. Nguyen, Memetic Comp., 2024, 16, 519–536 CrossRef.
- M. Tynes, M. G. Taylor, J. Janssen, D. J. Burrill, D. Perez, P. Yang and N. Lubbers, Digital Discovery, 2024, 3, 1980–1996 RSC.
- L. Bellmann, P. Penner and M. Rarey, J. Chem. Inf. Model., 2019, 59, 4625–4635 CrossRef CAS PubMed.
- R. M. Richard and J. M. Herbert, J. Chem. Phys., 2012, 137, 064113 CrossRef PubMed.
- G. Landrum, RDKit: Open-Source Cheminformatics, 2023, https://www.rdkit.org, release_2023.03.1 Search PubMed.
- H. L. Morgan, J. Chem. Doc., 1965, 5, 107–113 Search PubMed.
- Y. Song, T. Wang, P. Cai, S. K. Mondal and J. P. Sahoo, ACM Comput. Surv., 2023, 55, 271 CrossRef.
- H. Gharoun, F. Momenifar, F. Chen and A. H. Gandomi, ACM Comput. Surv., 2024, 56, 294 CrossRef.
- G. Ke, Q. Meng, T. Finley, T. Wang, W. Chen, W. Ma, Q. Ye and T.-Y. Liu, Proceedings of the 31st International Conference on Neural Information Processing Systems, 2017, pp. 3149–3157 Search PubMed.
- M. Forgione, A. Chakrabarty, D. Piga, M. Rufolo and A. Bemporad, Manifold meta-learning for reduced-complexity neural system identification, arXiv, 2025, preprint, arXiv:2504.11811, DOI:10.48550/arXiv.2504.11811.
- T. Hastie, R. Tibshirani and J. Friedman, The Elements of Statistical Learning, Springer, 2009 Search PubMed.
- B. Nagy and F. Jensen, in Reviews in Computational Chemistry, John Wiley & Sons, Ltd, 2017, pp. 93–149 Search PubMed.
- P. Bühlmann and S. Van De Geer, Statistics for High-Dimensional Data: Methods, Theory and Applications, Springer, 2011 Search PubMed.
- N. Ulrich, K. Voigt, A. Kudria, A. Böhme and R.-U. Ebert, J. Cheminf., 2025, 17, 55 Search PubMed.
- V. Ramani and T. Karmakar, J. Chem. Theory Comput., 2024, 20, 6549–6558 CrossRef CAS PubMed.
- W. Ahmad, H. Tayara and K. T. Chong, ACS Omega, 2023, 8, 3236–3244 CrossRef CAS PubMed.
- S. Lee, H. Park, C. Choi, W. Kim, K. K. Kim, Y.-K. Han, J. Kang, C.-J. Kang and Y. Son, Sci. Rep., 2023, 13, 957 CrossRef CAS PubMed.
- W. Ahmad, H. Tayara, H. Shim and K. T. Chong, Int. J. Mol. Sci., 2024, 25, 715 Search PubMed.
- L. Bertinetto, J. F. Henriques, P. H. S. Torr and A. Vedaldi, Meta-Learning with Differentiable Closed-Form Solvers, arXiv, 2018, preprint, arXiv:1805.08136, DOI:10.48550/arXiv.1805.08136.
- K. Lee, S. Maji, A. Ravichandran and S. Soatto, Meta-Learning with Differentiable Convex Optimization, arXiv, 2019, preprint, arXiv:1904.03758, DOI:10.48550/arXiv.1904.03758.
- S. Jinsong, J. Qifeng, C. Xing, Y. Hao and L. Wang, Commun. Chem., 2024, 7, 20 CrossRef PubMed.
- Z. Chen, M. R. Min, S. Parthasarathy and X. Ning, Nat. Mach. Intell., 2021, 3, 1040–1049 CrossRef PubMed.
- X. Liu, S. Jiang, Q. Huang, T. Xu, I. Foster, M. Wang, H. Lin and R. Stevens: FragmentGPT: A Unified GPT Model for Fragment Growing, Linking, and Merging in Molecular Design, arXiv, 2025, preprint, arXiv:2509.11044, DOI:10.48550/arXiv.2509.11044.
- BNNLab, BNNLab/Solubility_data: Leeds Solubility Data, https://zenodo.org/records/3686213, accessed: 2023-06-01 Search PubMed.
- L. Krasnov, D. Malikov, M. Kiseleva, S. Tatarin, S. Sosnin and S. Bezzubov, BigSolDB 2.0: A Comprehensive Dataset of Solubility Values for Organic Compounds in Organic Solvents and Water at Various Temperatures, https://zenodo.org/records/15094979, accessed: 2025-08-01 Search PubMed.
- A. Bhowmik, MultiXC-QM9 Data, https://data.dtu.dk/collections/MultiXC-QM9/6185986, accessed: 2023-06-01 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |










