Open Access Article
Open Access ArticleRedox potential prediction of Fe(II)/Fe(III) complexes: a density functional theory and graph neural network approach
Fakhrul H.
Bhuiyan
 a,
Hassan
Harb
a,
Hassan
Harb
 b,
Rajeev Surendran
Assary
b,
Rajeev Surendran
Assary
 b and
Álvaro
Vázquez-Mayagoitia
b and
Álvaro
Vázquez-Mayagoitia
 *a
*a
aComputational Science Division, Argonne National Laboratory, Lemont, IL 60439, USA. E-mail: vama@alcf.anl.gov
bMaterials Science Division, Argonne National Laboratory, Lemont, IL 60439, USA
First published on 4th February 2026
Abstract
This work presents an integrated computational approach that combines tight-binding density functional theory (DFT) with standard DFT calculations to accurately compute the redox potential of micro-solvated iron-based transition metal complexes. Comparison with experimental and computational reference values confirmed the reliability of the computational approach. A comprehensive redox dataset of 2267 iron complexes was generated using the computational approach for machine learning (ML) applications. Chemical analysis of the ligand space in the dataset provided detailed insights into how different ligand classes and local Fe-coordination environments can systematically influence redox potential. Finally, a graph neural network (GNN) framework featuring automated graph data generation directly from 3D Cartesian coordinates was developed to model and predict redox potentials of metal complexes. Four distinct GNN architectures (GCN, GAT, DimeNet++, and SchNet) were evaluated using the curated redox potential dataset. The best-performing model achieved a root mean squared error of 0.24 ± 0.01 V, representing state-of-the-art performance for redox potential prediction of transition metal complexes. Feature attribution analysis using integrated gradients provided insights into the factors influencing the GNN model predictions. This combined DFT-ML workflow provides both predictive power and chemical insight, offering a scalable pathway to accelerate the discovery and optimization of transition metal complexes for any application.
Introduction
The redox potential is a useful measure of the tendency of a chemical system to gain or lose electrons and is therefore a relevant indicator of the reactivity and stability of transition metal complexes.1 The redox of iron-based metal complexes is of significant interest because of its relevance in various applications, including energy storage technologies such as redox flow batteries (RFBs),2–6 bioinorganic chemistry,7–9 and catalysis.10–12 The redox potential of iron-based metal complexes can vary significantly depending on the coordinating ligands, reflecting their tunability and potential applications.12–14 The ability to tune the redox potential of iron-based metal complexes through selective ligand complexation positions them as promising components in advanced electrochemical systems. However, the vast chemical space (billions) of possible ligand–metal combinations prohibits exhaustive experimental or accurate quantum-mechanical exploration.15,16 As a result, there is a critical need for fast, accurate, and scalable methods to predict the redox potentials of iron-based metal complexes.Experimentally, the redox potential of transition metal complexes can be measured using cyclic voltammetry,17,18 differential pulse voltammetry,19 or square-wave voltammetry.20,21 Among these techniques, cyclic voltammetry is the most commonly used electrochemical technique that measures the electric current (I) response of a redox-active material as its potential (V) is linearly swept in a cyclic manner. The experimental determination of redox potentials is time-consuming and costly, rendering these experimental techniques unsuitable for high-throughput screening of a large chemical space of metal complexes.
Computational chemistry methods, particularly density functional theory (DFT), can complement experimental methods and serve as a valuable tool for determining redox potentials.22–24 The DFT offers a good balance between computational cost and accuracy to simulate a wide range of chemical systems and their material properties. Quantum chemical calculations based on DFT have been successfully used in numerous studies to calculate standard redox potentials for complexes in aqueous solutions using implicit solvation models, often achieving good agreement with available experimental values.25–29 Despite its utility, DFT calculations can be computationally expensive for large molecular systems and for high-throughput screening efforts.
Classical molecular dynamics (MD) simulations using force fields have also been applied, though in a limited capacity, to study redox potentials. For example, molecular mechanics force fields were developed for transition-metal oxide clusters to estimate oxidation potentials.30,31 Reactive force fields (ReaxFF) have been widely employed to model diverse chemical reactions,32–34 but their lack of explicit electron treatment limits their use for redox studies. To overcome this, e-ReaxFF was introduced, enabling explicit electron treatment and applied to model redox processes such as silver halide reactions35,36 and solid-oxide electrocatalysis.37 However, e-ReaxFF remains in early development and is not yet widely adopted for redox potential calculations.
Machine learned interatomic potentials (MLIPs) can also be used to study chemical reactions in MD simulations.38 However, the use of MLIPs for calculating redox potential is rare due to the inherent difficulty in modeling long range electrostatic interactions and performing charge equilibration using currently available MLIPs.39–41 Modeling redox reactions using MLIPs require complex modification of the architecture of the MLIP model. One such modification of MLIP was recently demonstrated for the high-dimensional neural network potentials to capture the redox chemistry of Fe2+ and Fe3+ ions in water.42 Despite the recent progress, application of MLIPs for redox potential calculation in MD simulations remains challenging and not suitable for high-throughput screening purposes.
Machine learning (ML) has emerged as a powerful approach to accelerate materials discovery by providing rapid property prediction capabilities. ML models can serve as fast surrogates for computationally intensive simulations, such as DFT, predicting properties in seconds making them suitable for high-throughput screening. For instance, Gaussian Process Regression models trained using fingerprints-based graph representation of organic ligands were used to predict redox potentials of organic molecules.43 Graph neural networks (GNNs), among various ML approaches, are particularly well-suited for handling non-Euclidean data such as molecular structures.44,45 GNNs operate on graph-structured data, where molecules are represented as graphs with nodes for atoms and edges for bonds. This approach allows GNNs to capture complex relationships between atoms and their chemical environments, leading to improved performance over conventional descriptor-based models.46
Several studies have demonstrated the effectiveness of GNNs in predicting redox potentials. For organic compounds, GNNs have outperformed classical ML models and have been used to screen large molecular databases.47 A molecular graph attention network, a type of GNN, was developed to predict the redox potential of organic compounds using molecular structures, atomic properties, and bond attributes, outperforming other GNN variants.48 Graph-based methods have also been applied to transition metal complexes. For instance, a graph-based descriptor model was used to model octahedral Fe(II/III) redox couples to show that redox potential increased with increasing weight of the complex but decreased for branched structures.26 Although that work successfully demonstrated in silico discovery of metal complexes from a large chemical space of ligands, the redox calculations and predictions remained constrained in a small, concentrated dataset containing less than 50 metal complexes. Despite the advances, a general methodology for the high-throughput screening of a large and diverse set of metal complexes, including iron-based ones, is highly desired.
In this work, we developed a general methodology for modeling and predicting the redox potential of thousands of iron-based metal complexes from the transition metal quantum mechanics (tmQM) dataset using high-throughput screening.49 Calculation of the redox potentials led to the generation of a redox dataset containing 2267 iron complexes under explicit and implicit solvent effects. Extensive chemical analysis of the iron complexes in the dataset revealed key insights into the effect of ligand chemistry and iron-ion coordination in determining the redox behavior of a complex. The redox dataset was then used for training multiple GNN models through a neural network training framework developed here and model performances were compared. We introduce two edge-aware GNN models, RexGCN and RexGAT, that showed more precise predictions and reached earlier learning curve plateau than previously reported GNN models. To our knowledge this is the largest redox database for calculated redox potential for the Fe(II)/Fe(III) pair of structures observed experimentally and reported in the Cambridge Structural database. A schematic representation of our workflow is shown in the Fig. 1. Although the main focus of this work was on iron-based complexes, the methods developed here can be readily applied to model any metal complex from any dataset, suggesting the far-reaching utility of our approach.
Methodology
Data collection
The iron complexes used in this study were collected from the tmQM dataset (2024 release).49 Built based upon structures retrieved from the Cambridge Structural database, the tmQM dataset is currently one of the largest openly available datasets for transition metal complexes. The 2024 release of this dataset contains the geometries and quantum mechanical properties of ∼108![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 metal complexes including iron complexes.
000 metal complexes including iron complexes.
We first filtered the tmQM dataset to include only iron complexes containing 60 or fewer atoms. The selected iron complexes were further filtered to retain only those containing an iron ion with an oxidation state of +2 (i.e., Fe(II) complexes). We calculated the oxidation state using the following equation:
 | (1) |
Quantum chemical calculations
Each complex was evaluated with a hybrid aqueous solvation effect approach, where solvent effects are treated with inclusion of both explicit water molecules and an implicit continue model. To do so, we first microsolvated by adding 30 water molecules using the STOCASTIC algorithm in the Solvator module of ORCA 6.0 software.51,52 The number of water molecules was chosen arbitrarily based on previous studies.27 Next, each microsolvated complex geometry was optimized using the GFN2-xTB semiempirical method,53 with singlet spin multiplicity and the total charge reported in the tmQM dataset, and with the ALPB implicit solvation model.54 We performed a vibrational frequency (Hessian) calculation to confirm that each structure corresponded to a local minimum.To compute the redox potential, we evaluated single point energies (SPEs) of the microsolvated Fe(III) complexes and their reduced form, Fe(II), using the TPSSh/def2-TZVP level of theory. To account for solvent effects, we used the SMD model with water dielectric constant. This level of theory was suitable for computing approximately 2000 complexes with our computational resources.
Our selection of this method was made after a comparison of the performance between B3LYP55 and TPSSh56 exchange-correlation (xc) functionals, and basis sets def2-SVP, def2-TVZP, and def2-TZVPP, in the SI (Fig. S1). We found a linear correlation between both functionals, in particular with larger basis sets, this observation is consistent with previous reports comparing xc hybrid functionals.57,58 Particularly, Jensen59 reported that TPSSh has a good performance describing bond energies of transition metals and bioinorganic molecules. Our results show that values computed with def2-TVZP basis have a trend closer to a larger basis set, def2-TVZPP, than to the smaller def2-SVP. Semi empirical dispersion energies were not added, since we performed differences between single point energy calculations the effects of dispersion energies are canceled. Full relaxation of solvated complexes at DFT level had a prohibitive cost for the scope of this work. A systematic comparison among DFT xc functionals, basis sets and solvent models have been extensively explored in a previous studies.5,27,60
We estimated the Gibbs free energy of redox, Gred, as the difference of SPEs from electronic structure calculations, using the following equation:
| ΔGred ≈ SPEIII − SPEII | (2) |
The redox potential, Ered, was calculated using the following equation, adapted from the work of Harb27
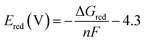 | (3) |
In Table 1, we show a comparison of the redox potentials evaluated using our methodology, previous reports by Harb27, and experimentally measured values. In most cases, our methodology provided redox values comparable to experimental values. The largest error was observed for the pair Fe(CN)6−4/−3 with a calculated redox potential of +1.29 V compared with the experimental value of +0.37 V.62,63 Nevertheless, our calculated redox potential value was positive, whereas previous studies reported negative values of the redox potential, for example −0.56 V,58 −0.34 V,64 and −0.33 V.60 Although the discrepancies of redox estimations may be larger for other complexes that are not included in Table 1, the current evaluations suggest that our dataset could be a starting point for a further more in detail analysis, both theoretical and experimental.
| Entry | Ligand | This work | Harb27 | Exp. | ||
|---|---|---|---|---|---|---|
| 1 | Maltolate | −0.30 | (0.07) | −0.57 | (0.34) | −0.23 (ref. 58) |
| 2 | Deferiprone | −0.31 | (0.27) | −0.76 | (0.18) | −0.58 (ref. 58) |
| 3 | Kojate | −0.33 | (0.20) | −0.27 | (0.14) | −0.1 (ref. 58) |
| 4 | Catecholate | −0.93 | (0.10) | −0.99 | (0.16) | −0.83 (ref. 58) |
| 5 | Salicylate | −0.67 | (0.01) | −0.84 | (0.18) | −0.66 (ref. 58) |
| 6 | Fe(CN)6 | 1.29 | (0.93) | 0.30 | (0.07) | 0.37 (ref. 62) |
| 7 | BIS-TRIS | −0.52 | (0.39) | −0.91 (ref. 65) | ||
| 8 | DIPSO | −0.74 | (0.10) | −0.84 (ref. 66) | ||
| 9 | TEA | −0.44 | (0.41) | −0.85 (ref. 6) | ||
| 10 | TIPA | −0.63 | (0.25) | −0.88 (ref. 67) | ||
| 11 | Phen | 0.64 | (0.48) | 1.12 (ref. 5) | ||
| 12 | Terpy | 0.67 | (0.46) | 1.13 (ref. 5) | ||
To perform all these atomistic simulations, we used a workflow manager based on Fireworks,68 using Jobflow to create Firetasks, and the atomic simulation environment (ASE)69 to drive calls of ORCA and xTB binaries. We used two mongoDB databases, one for managing the jobs, and a second for storing the data. All our computations used the Improv (CPU AMD Ryzen Rome) cluster at Argonne Laboratory Computing Resource Center.
GNN training datasets
Following the redox potential calculations, three distinct datasets were constructed for training the GNN models. All training datasets inherited our calculated redox potentials as the target property from the full Redox2k dataset. The three datasets differed in both the types of geometries included and the total number of structures or data points in the datasets.The first training dataset was generated by collecting desolvated iron complexes obtained after removing all explicit solvent water molecules from the optimized, microsolvated structures used in the redox potential calculations. All resulting structures were converted from Cartesian coordinates into RDKit Mol objects using a modified version of the xyz2mol_tm method, as described in the SI.50 Several filtering and preprocessing steps were required to enable reliable structure conversion, which resulted in the exclusion of some metal complexes from the original Redox2k dataset and yielded a curated set of 1450 iron complexes. A detailed description of all filtering and processing steps applied during the collection of desolvated structures is provided in the SI. This dataset, consisting of 1450 desolvated iron complex geometries, is referred to, in this text, as the GNN-Redoxdesolv-geo1450 dataset.
The second dataset was constructed using the same 1450 iron complexes as the first dataset, but with geometries obtained directly from the tmQM dataset instead of desolvated geometries. This dataset is referred to, in this text, as the GNN-RedoxtmQM-geo1450 dataset. Finally, the third dataset was created by collecting iron complex geometries directly from the tmQM dataset. No filtering or preporcessing steps were required for the third dataset since the tmQM geometries were directly used. Thus, all iron complexes in the full Redox2k dataset could be utilized. All collected iron complexes were converted from Cartesian coordinates into RDKit Mol objects using our modified xyz2mol_tm method to construct the dataset. This third dataset consisting of ∼2000 iron complexes is referred to, in this text, as the GNN-RedoxtmQM-geo2k dataset.
Graph data generation
All GNN-Redox training datasets contained RDKit mol objects to represent the iron complexes in the GNN model training process. However, some graph convolution model architectures cannot work directly with such mol objects or 3D coordinates, and instead, require graph data with handcrafted features. So, the RDKit mol objects of the iron complexes in our GNN-Redox training datasets were converted to graph representations with comprehensive node, edge, and graph-level features. A complete list of all features is provided in Table 2. Notably, we used a condensed version of the full 2048 bit Morgan fingerprint70 with a bond diameter of six bonds (i.e., ECFP6) as a graph feature. ECFP6 has been previously used in various molecular property prediction tasks71–73 and was found to yield the best overall model performance in this study (data not shown) compared to fingerprints generated with larger diameters. The condensed fingerprint comprised of the most important 64 fingerprint bits, as determined by a Random Forest Regressor model that was trained to predict the redox potential using the full Morgan fingerprint as input features. Utility of such fingerprint condensation has been demonstrated before to detect structurally diverse active compounds.74 Here, condensation of the Morgan fingerprint aimed to enhance model performance by highlighting key chemical features relevant for redox behavior of the complexes.| Item | Feature name | Description | Type | Size |
|---|---|---|---|---|
| Node features | Atomic number | Atomic number scaled by Uranium's atomic number: (atomic_number − 1)/91 | Numerical | 1 |
| Symbol | Chemical symbol of the element | One-hot vector | 92 | |
| Valence | Number of maximum valence electrons | One-hot vector | 8 | |
| Formal charge | Formal electrical charge of the atom | Numerical | 1 | |
| Radical electrons | Number of radical electrons | Numerical | 1 | |
| Hybridization | Hybridization as s, sp, sp2, sp3, sp3d, sp3d2, or unspecified | One-hot vector | 7 | |
| Aromaticity | Indicates if an atom is in an aromatic ring (1 or 0) | Numerical | 1 | |
| Hydrogens | Number of connected hydrogens, encoded as 0, 1, 2, 3, or 4 | One-hot vector | 5 | |
| Atomic mass | Scaled atomic mass: (mass − 1.008)/237.021 | Numerical | 1 | |
| Vdw radius | Scaled Van der Waals radius: (RVDW − 1.2)/1.35 | Numerical | 1 | |
| Covalent radius | Scaled covalent radius: (Rcovalent − 0.23)/1.71 | Numerical | 1 | |
| Edge features | Bond type | Bond type as single, double, triple, or dative | One-hot vector | 4 |
| Conjugated bond | Indicates if the bond is conjugated (1 or 0) | Numerical | 1 | |
| Ring | Indicates if the bond is in a ring (1 or 0) | Numerical | 1 | |
| Stereo | Stereochemistry: StereoNone, StereoAny, StereoZ, StereoE | One-hot vector | 4 | |
| Graph features | Morgan fingerprint | Condensed version of the Morgan fingerprint | Numerical | 64 |
Graph neural network models
We trained four GNN architectures to predict the redox potentials of iron complexes: graph convolutional network (GCN),75 graph attention network (GAT),76 DimeNet++,77 and SchNet.78,79 GCN and GAT are topology-centric models that learn from abstract graph connectivity and node features. The models differ, however, in their approach to neighborhood aggregation. In the message aggregation step, a GCN assigns fixed, structure-dependent weights to neighboring nodes, typically based on node degrees, without learning their relative importance. In contrast, a GAT aims to extend GCN by employing a learnable attention mechanism that dynamically assigns different weights to each neighbor based on their features. Both GCN and GAT models have been widely and successfully used for various molecular property prediction tasks, including the prediction of redox potential of organic molecules.47,76,80RexGCN
In this work, we modified the original GCN75 implementation to incorporate edge features. Addition of edge features in message construction has been demonstrated previously in many other GNN models for molecular property prediction tasks.81–83 The standard GCN formulation only considers node features during message passing, which can be defined as:mij(l) = âijxj(l)Θ(l), Â = ![[D with combining tilde]](https://www.rsc.org/images/entities/b_char_0044_0303.gif) −1/2Ã −1/2Ã![[D with combining tilde]](https://www.rsc.org/images/entities/b_char_0044_0303.gif) −1/2, Ã = A + I −1/2, Ã = A + I | (4) |
 denotes the feature vector of node j at layer l,
denotes the feature vector of node j at layer l,  is the learnable transformation matrix, Ã is the adjacency matrix with self-loops added,
is the learnable transformation matrix, Ã is the adjacency matrix with self-loops added, ![[D with combining tilde]](https://www.rsc.org/images/entities/b_char_0044_0303.gif) is its degree matrix, and
is its degree matrix, and 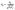 are the normalized edge weights. The GCN we used in this work, which will be referred to as the RexGCN, extends the original formulation by incorporating edge features eij into the message construction:
are the normalized edge weights. The GCN we used in this work, which will be referred to as the RexGCN, extends the original formulation by incorporating edge features eij into the message construction:| m(l)ij = âij[xj(l)Θ(l)‖eij] | (5) |
 represents the edge features between nodes i and j. Since the concatenation increases the message dimensionality from d(l+1) to d(l+1) + de, a learnable update function was introduced here to project the aggregated messages back to the desired output dimension:
represents the edge features between nodes i and j. Since the concatenation increases the message dimensionality from d(l+1) to d(l+1) + de, a learnable update function was introduced here to project the aggregated messages back to the desired output dimension: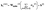 | (6) |
 denotes the updated feature (hidden representation) of node i at the output of layer l,
denotes the updated feature (hidden representation) of node i at the output of layer l, is the learnable update projection matrix, and
is the learnable update projection matrix, and  is the bias term. The complete RexGCN layer can be expressed in matrix form as:
is the bias term. The complete RexGCN layer can be expressed in matrix form as: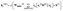 | (7) |
 is the set of neighbors of node i and σ(·) is a nonlinear activation function (e.g., ReLU). These modifications aim to create a better description of the local chemical environments and, subsequently, enhance the GCN's performance for modeling redox potential of metal complexes. To confirm that incorporating edge features improves GCN performance, we trained a baseline GCN model without edge features and compared its results with our modified RexGCN implementation. The comparison results are presented in the Results and Discussion section below.
is the set of neighbors of node i and σ(·) is a nonlinear activation function (e.g., ReLU). These modifications aim to create a better description of the local chemical environments and, subsequently, enhance the GCN's performance for modeling redox potential of metal complexes. To confirm that incorporating edge features improves GCN performance, we trained a baseline GCN model without edge features and compared its results with our modified RexGCN implementation. The comparison results are presented in the Results and Discussion section below.
RexGAT
We constructed the GAT model used in this work based on the MolGAT model, which was previously used to predict redox potential of small organic molecules.48 We implemented several modifications to enhance model robustness and performance. The primary changes include: (1) explicit self-loop handling with appropriate edge feature management, (2) implementation of residual connections with adaptive input projection, and (3) refined message passing that separates attention computation from message content.The GAT layer used here, which will be referred to as the RexGAT, begins with a linear transformation of input node features:
| hi(l),h = x(l−1)iW(l),h, h = 1,…, H | (8) |
 represents the input feature vector from the previous layer,
represents the input feature vector from the previous layer,  is the learnable weight matrix for head h, and
is the learnable weight matrix for head h, and  is the head-specific transformed feature. Similar to the MolGAT model, the attention mechanism in this work incorporates both node and edge features to compute attention coefficients:
is the head-specific transformed feature. Similar to the MolGAT model, the attention mechanism in this work incorporates both node and edge features to compute attention coefficients:| eij(l),h = LeakyReLU(a(l),h⊤[h(l),hi‖h(l),hj‖eij]) | (9) |
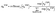 | (10) |
 is the h-th head vector for node i,
is the h-th head vector for node i,  is the edge feature between nodes i and j, ‖ denotes concatenation, and
is the edge feature between nodes i and j, ‖ denotes concatenation, and  is the learnable attention vector for head h.
is the learnable attention vector for head h.
We then constructed messages based on the following equation:
| mij(l),h = αij(l),hh(l),hj | (11) |
Here, the edge features do not directly contribute to the message content, unlike the original MolGAT where the edge features influence both attention weights and messages directly. The message construction approach used here ensures that edge features modulate the importance of neighboring nodes through the attention coefficient αij(l),h while maintaining clean node feature propagation. Next, the per-head aggregated message  is obtained by summing messages over neighbors (including self-loop):
is obtained by summing messages over neighbors (including self-loop):
 | (12) |
The final layer output incorporates multi-head averaging and a learnable update transformation:
 | (13) |
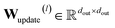 is the learnable update matrix
is the learnable update matrix  is the bias term. To improve training stability and gradient flow, residual connections are incorporated when feasible:
is the bias term. To improve training stability and gradient flow, residual connections are incorporated when feasible:| xi(l) = xi(l) + Proj(xi(l−1)) | (14) |
The attention mechanism in our RexGAT implementation remains fundamentally the same as the original MolGAT formulation, ensuring that the model can effectively capture the importance of different molecular bonds and neighboring atoms. However, the modified architecture, with its explicit self-loop handling, residual connections, and refined message passing, provides a better model implementation, enhanced training stability, and improved capacity for capturing complex atomic bonding environments of coordination complexes.
DimeNet++ and SchNet
Distinct from the topology-based GCN or GAT models, DimeNet++ and SchNet are geometry-centric models that incorporate 3D spatial information into the graph representation. SchNet uses continuous interatomic distances in its message-passing framework, while DimeNet++ further enriches this by incorporating both distances and angular relationships using radial and spherical basis functions. As a result, DimeNet++ and SchNet produce rotationally invariant representations, making them effective for tasks such as molecular property prediction where 3D geometry is critical. Although their usability for redox prediction has not been explored previously, both DimeNet++ and SchNet have been used for various molecular property prediction tasks.77,79,84–86 We created the DimeNet++ and SchNet models for redox potentials based on the same architecture and hyper-parameters used to model the QM9 dataset.77,79 Detailed model hyper-parameters are provided in the SI.General training parameters
The GNN-Redoxdesolv-geo1450 dataset containing desolvated geometries of 1450 complexes was used to train and evaluate all models. Average model performance was assessed by conducting five-fold cross validation (CV) for RexGCN, RexGAT, and DimeNet++ models, while the SchNet model was assessed using a three-fold CV. Finally, one representative model was trained from each model type with early stopping based on validation set performance for final evaluation. Before training any model, the dataset was divided into training, validation, and test sets using a 70![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 15
15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 15 split. To avoid any data leakage, the random forest regressor used to condense the fingerprints was trained only using the train split in both the CV analysis and final model training. Training epoch varied depending on the model architecture. RexGCN, RexGAT, DimeNet++, and SchNet models were trained for a maximum of 200 epochs, 200 epochs, 400 epochs, and 2000 epochs, respectively, with a maximum batch size of 128 to balance between computational cost and model performance. For early stopping, patience was set to 50 epochs for RexGCN, RexGAT, and DimeNet++ models, while patience was set to 100 for the SchNet model.
15 split. To avoid any data leakage, the random forest regressor used to condense the fingerprints was trained only using the train split in both the CV analysis and final model training. Training epoch varied depending on the model architecture. RexGCN, RexGAT, DimeNet++, and SchNet models were trained for a maximum of 200 epochs, 200 epochs, 400 epochs, and 2000 epochs, respectively, with a maximum batch size of 128 to balance between computational cost and model performance. For early stopping, patience was set to 50 epochs for RexGCN, RexGAT, and DimeNet++ models, while patience was set to 100 for the SchNet model.
Results and discussion
Redox2k dataset analysis
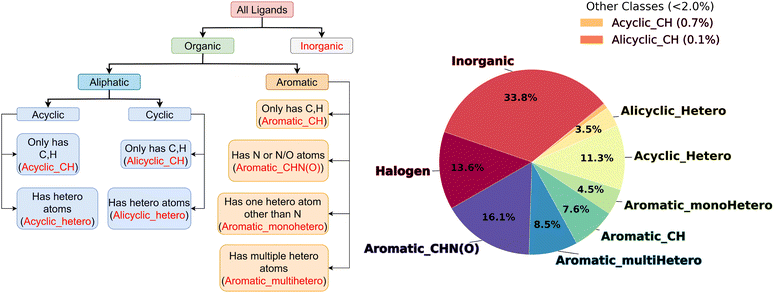
To analyze the diversity of the chemical space of the selected 2267 iron complexes in the Redox2k dataset, we systematically categorized the ligands into distinct classes. First, the ligands were classified into organic, i.e., contains C–C or C–H bonds, or inorganic ligands. Next, the organic ligands were categorized as aromatic or aliphatic groups which were then further sub-classified based on the presence of heteroatoms. The classification scheme and the distribution of ligands across these categories are illustrated through a classification tree and a pie chart in Fig. 2 for the Redox2k dataset. Examples of ligands from each class are presented in Fig. 3 and additional examples are shown in Fig. S2.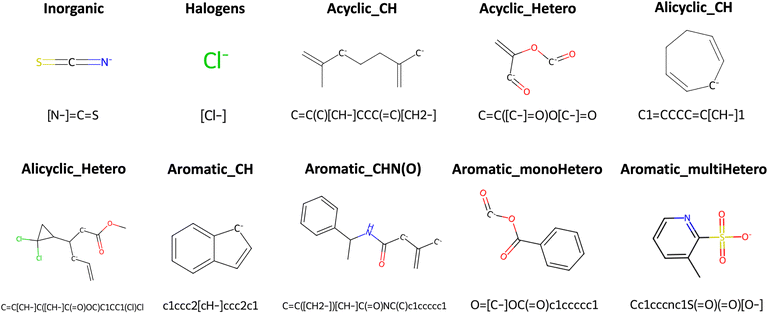 | ||
| Fig. 3 Examples of representative ligands from each various ligand classes described in Fig. 2. The ligands are accompanied by their respective SMILES string. | ||
The iron complexes in the Redox2k dataset contained a total of 7463 ligands, of which ∼37% were aromatic compounds, ∼34% were inorganic, ∼14% were halogens, and the remaining ∼15% were aliphatic groups (Fig. 2). Among the aromatic ligands, compounds containing N or both N and O heteroatoms were found to be the most frequent (∼16% of total ligands) class. In case of aliphatic ligands, with ∼11% of total ligands, acyclic compounds containing heteroatoms were the most common class of ligands. Ligand class distribution in the GNN-Redoxdesolv-geo1450 dataset (Fig. S3), which was curated from the Redox2k dataset, followed the same trend as observed in the Redox2k dataset.
Analysis of iron coordination number in the Redox2k dataset showed octahedral coordination to be the most prevalent case (Fig. S4). Analysis of ligand denticity revealed that the Redox2k dataset predominantly contained mono- and bi-dentate ligands but up to hexa-dentate ligands were observed, excluding one outlier with a higher denticity (Fig. S4). The GNN-Redoxdesolv-geo1450 dataset also followed these same trends in iron coordination number and ligand denticity as observed in the Redox2k dataset.
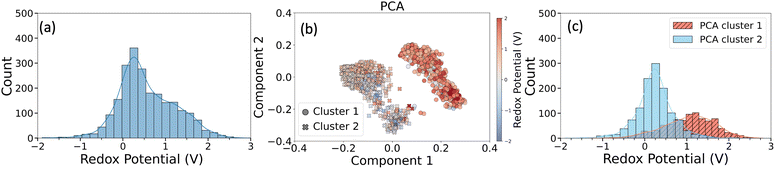
Fig. 4a shows the distribution of the computed redox potentials (vs. SHE) in the Redox2k dataset. The distribution ranged from −1.89 V to 7.47 V with an average of 0.56 V. The distribution exhibited a slight right skew, with a tail extending towards more positive potentials. Six randomly selected iron complexes with redox potentials ranging from −1 V to +1 V are shown in Fig. S5 as representative structures.
The range of redox potentials observed in Fig. 4a spanned a broad spectrum (∼−2 V to ∼+2 V) indicating the diverse set of electrochemical responses due to the presence of ligands-metal ion combinations, which could enable various redox-switch and flow-battery applications. More specificaly, our computations showed that 377 of the iron complexes had a negative redox potential. Identification of these iron complexes with negative or lower redox potentials may provide new candidate anolytes for all-soluble, all-iron redox flow batteries. This is particularly important since current anolyte options are very limited with most notable options being iron-triethanolamine and iron-nitrilotri(methylphosphonic acid) complexes.63,87
To explore this structure-redox potential relationship, Morgan fingerprints were used to represent the chemical structure of the complexes, capturing detailed substructural information about the ligands. To reduce the dimensionality and highlight the most relevant features, a random forest regressor was first trained using the full 2048 bit Morgan fingerprints as input features to predict redox potentials. Then, 64 of the most important fingerprint bits as determined by feature importance scores were selected to generate a condensed version of the fingerprint which served as input for principal component analysis (PCA). Fig. 4b displays the PCA plot based on this condensed fingerprint, with the first two principal components capturing the largest variance in the data. This dimensionality reduction led to a clearer separation of complexes in the PCA space, offering better insight into how specific chemical features influence redox properties.
The iron complexes formed two well-separated clusters in the PCA space, as shown in Fig. 4b. To formally define these groupings, a k-means clustering algorithm with k = 2 was applied. When the data points were colored according to their redox potentials, a clear distinction in redox behavior emerged between the two clusters. This is further illustrated in Fig. 4c, which presents the redox potential distributions for each cluster. Cluster 1 exhibited redox potentials ranging from −1.76 V to 2.81 V, with an average value of 1.05 V, standard deviation of 0.61 V, and median at 1.11 V. Redox potentials in Cluster 2, on the other hand, ranged from −1.90 V to 2.06 V, excluding one outlier iron complex that had 7.48 V redox potential. Both the average and median redox potential in cluster 2 was the same, 0.24 V, and the standard deviation was 0.44 V. The significant difference in redox potential distributions between the two clusters highlights the influence of underlying ligand chemistry and suggests that redox behavior in iron complexes can be effectively tuned by targeted ligand selection.
The ligand classes, as defined in Fig. 2, were analyzed to gain deeper insight into the chemical characteristics associated with each PCA-derived cluster. Fig. 5a highlights the five most common ligand classes within each cluster. In Cluster 1, inorganic ligands were the most dominant, accounting for approximately 50% of all ligands in Cluster 1 (blue bars in Fig. 5a). Different aromatic groups and acyclic compounds containing heteroatoms were the other notable classes in the remaining 50% of ligands in Cluster 1. Remarkably, all complexes in Cluster 1 contained at least one inorganic ligand (red bars in Fig. 5a). Aromatic groups containing only C and H atoms were the second most common class appearing in ∼55% of complexes. All other classes individually appeared in no more than ∼35% of the complexes. These trends suggest that inorganic ligands are the defining chemical feature of Cluster 1.
 | ||
| Fig. 5 Analysis of the two clusters observed in the PCA space. (a) Analysis of the ligand classes, as defined in Fig. 2, in the two clusters. The blue bars represent the frequency of ligand classes observed in all ligands of cluster 1 (top panel) and cluster 2 (bottom panel) transition metal complexes (TMCs). Only the five most frequent ligand classes are shown. The red bars represent the percentage of iron complexes that contain a ligand from a specific class. (b) Analysis of the direct neighbors of the iron ion in the two clusters. The bars represent the percentage of iron complexes that contain a specific element as a neighbor. | ||
In contrast, the Cluster 2 includes a more diverse ligand composition. Aromatic compounds containing N and/or O as heteroatoms (aromatic_CHN(O) class), inorganic ligands, and halogens were observed with comparable overall frequencies (blue bars in Fig. 5a). These aromatic_CHN(O) ligands were present in approximately 60% of the complexes (red bars in Fig. 5a), making them the most prevalent ligand class in Cluster 2. Notably, the aromatic_CHN(O) ligand class was absent from the top five classes in Cluster 1, strongly indicating that the presence of aromatic_CHN(O) ligands drives a complex toward Cluster 2. Additionally, the presence of halogen atoms, or combinations of inorganic ligands with halogens, also appears to favor assignment to Cluster 2.
The chemical environment surrounding the metal ion plays a critical role in shaping the redox behavior of a metal complex. Therefore, analyzing the local coordination environment around the iron ion can offer further valuable insights into the distinct redox behaviors observed between the two PCA-derived clusters. Fig. 5b presents the distribution of neighboring chemical species directly bonded to the iron ion. In Cluster 1, every iron complex had at least one carbon atom coordinated to the iron center. Phosphorus and nitrogen were similarly frequent, appearing in approximately 30% of the complexes. In contrast, Cluster 2 was characterized by a markedly different coordination environment: nearly 80% of the complexes featured nitrogen as a neighbor to the iron ion, followed by chlorine and oxygen with each having presence in ∼30% of Cluster 2 complexes. Notably, carbon was observed as a neighbor in only ∼20% of Cluster 2 complexes. These results suggest that the presence of carbon and/or phosphorus atoms in the immediate environment of the iron ion is strongly associated with Cluster 1, while coordination with nitrogen, oxygen, and/or chlorine favors assignment to Cluster 2. This distinction in local chemical environments likely plays a significant role in driving the divergence in redox behavior between the two clusters. Further, the observed relation among redox potential of iron complexes, ligand class, and local environment can be utilized using GNN models for high-throughput screening purposes.
Performance analysis of the GNN models
Four different GNN models (RexGCN, RexGAT, DimeNet++, and SchNet) were trained here using the GNN-Redoxdesolv-geo1450 dataset. Despite containing less structures, the training dataset remained a fair representation of the full Redox2k dataset since the redox potential distribution (Fig. S6a), presence of the PCA clusters (Fig. S6b), and the chemical and redox properties of the clusters (Fig. S6c and S7) in the GNN-Redoxdesolv-geo1450 dataset followed the same trends as in the full Redox2k dataset. K-fold CV analysis was carried out for all the models to evaluate their average performance. Table 3 provides a summary of model performance across their respective CV folds. Among the four models, with the lowest root mean squared error (RMSE) of 0.24 ± 0.01 V, the RexGCN model achieved the best average validation performance. The RMSE loss in the validation set across the CV folds are presented in Fig. S8 for all four models.| Model type | Folds | Best validation RMSE (standard deviation) | |
|---|---|---|---|
| RexGCN | 5 | 0.24 V | (0.01 V) |
| RexGAT | 5 | 0.27 V | (0.01 V) |
| DimeNet++ | 5 | 0.28 V | (0.01 V) |
| SchNet | 3 | 0.48 V | (0.02 V) |
To further evaluate individual model performance, a representative instance was trained for each of the four GNN model types. We trained all models using an early stopping criterion based on validation RMSE loss. The training and validation loss curves for all representative models are shown in Fig. S9. Parity plots of redox potentials for all four models on the training, validation, and test sets are shown in Fig. 6. Among the four models, the RexGCN model demonstrated the highest predictive accuracy, achieving an R2 = 0.84 and RMSE = 0.26 V on the test set. The predictive accuracy of RexGCN surpasses that of a previously developed graph-based descriptor model, which was trained to predict the redox potentials of iron complexes with nitrogen ligands and achieved an R2 score of 0.6 on the test set.26 Note that the RexGCN model discussed here incorporates both node and edge features in message construction (eqn (5)). Our implementation improves upon the baseline GCN without edge features, as shown in Fig. S10. While the baseline achieved comparable average performance in the 5-fold CV analysis (0.27 ± 0.01 V), its predictive performance on the test set was poorer, with R2 = 0.78 and RMSE = 0.30 V, compared to our edge-aware RexGCN implementation.
Although RexGCN achieved the best overall performance, both RexGAT and DimeNet++ exhibited comparable predictive accuracy. As shown in Table 3, the average validation set RMSE loss of RexGAT and DimeNet++ in CV analysis differed by ≤0.04 V compared to RexGCN. In addition, representative RexGAT and DimeNet++ models achieved test set R2 scores of ∼0.8 (Fig. 6). SchNet, in contrast, exhibited the poorest performance among all evaluated models, yielding the highest average validation RMSE of 0.48 ± 0.02 V in CV analysis (Table 3). Consequently, the representative SchNet model showed the lowest predictive accuracy on the test set, characterized by the lowest R2 score and the highest RMSE loss (Fig. 6). Notably, SchNet also required the longest training time, converging after ∼1750 epochs. While its performance could potentially be improved through extensive hyperparameter optimization, such efforts would be computationally expensive, and increasing the number of model parameters would further slowdown the training process. Considering both computational cost and predictive accuracy, we found SchNet as a suboptimal choice for modeling redox potential of metal complexes.
The four models used here differ significantly in their architectural design and the way they process molecular information. GCN and GAT are both message-passing frameworks that require explicit construction of node-level, edge-level, and graph-level descriptors.75,76 GAT aims to enhance the graph convolution of GCN by incorporating attention mechanisms to weigh neighbor contributions dynamically.48,76 In contrast, SchNet and DimeNet++ are continuous-filter convolution models that rely directly on atomic coordinates and atomic numbers to generate features dynamically during training, without the need for manual featurization.77,79 While both models encode interatomic distances, DimeNet++ introduces greater expressivity by incorporating directional information and angular terms, and employing radial and spherical basis functions. This allows DimeNet++ to capture higher-order geometric relationships, offering a more nuanced representation of molecular structure than SchNet.
The comparative performance of the models reveals that the success of a GNN in predicting redox potentials depends less on the complexity of the learning algorithm and more on the quality and completeness of the chemical representation, whether learned or engineered. These observations underscore the critical role of feature representation in determining model performance. Models with either well-engineered graph features (as in GCN and GAT) or with architectures capable of constructing rich geometric features internally (as in DimeNet++) can learn redox behavior effectively. However, once meaningful structural and chemical information is captured either through manual featurization or geometric encoding, the choice of learning algorithm has a diminishing impact on performance. Instead, the limiting factors may be the overall quality of the dataset itself, determined by the size, accuracy, diversity, and chemical richness.
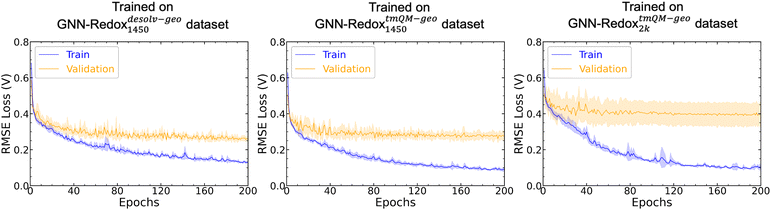
Effect of TM geometry on model performance
The quality and chemical relevance of the training dataset have a substantial impact on model performance, as illustrated in Fig. 7 which compares the average validation RMSE obtained from 5-fold CV of the RexGCN model trained on three different datasets. These datasets are: (i) the GNN-Redoxdesolv-geo1450 dataset, containing desolvated geometries of 1450 iron complexes; (ii) the GNN-RedoxtmQM-geo1450 dataset, comprising of tmQM geometries of the same 1450 complexes; and (iii) the GNN-RedoxtmQM-geo2k dataset, which includes tmQM geometries for nearly all iron complexes in the Redox2k dataset.As shown in Fig. 7, the RexGCN model achieved comparable average performance in validation set when trained on the GNN-Redoxdesolv-geo1450 (RMSE loss of 0.24 ± 0.01 V) and GNN-RedoxtmQM-geo1450 datasets (RMSE loss of 0.25 ± 0.02 V). In contrast, with an RMSE loss of 0.37 ± 0.06 V, a clear degradation in performance in validation set was observed when the model was trained on the larger GNN-RedoxtmQM-geo2k dataset.
The tmQM dataset consists of crystal geometries of iron complexes, which can differ substantially from solution-phase geometries in the presence of explicit solvent. In aqueous environments, water molecules may displace ligands and form solvation shells around the iron center, altering its coordination environment. However, the geometries in the GNN-Redoxdesolv-geo1450 and GNN-RedoxtmQM-geo1450 datasets are highly similar, as complexes that exhibited significant deviations from their tmQM geometries upon solvation were excluded during the curation of the GNN-Redoxdesolv-geo1450 dataset. As a result, these 1450 iron complexes largely retain their crystal-like coordination environments in solution, leading to comparable model performance across the two datasets. In contrast, the GNN-RedoxtmQM-geo2k dataset includes iron complexes that do not preserve their crystal geometries in solution and instead exhibit altered coordination environments around the iron center. The degradation in model performance when trained on the GNN-RedoxtmQM-geo2k dataset despite the larger dataset size suggests that the quality and consistency of structural inputs play a more critical role in model performance than simply increasing dataset size.
The importance of the immediate iron coordination environment is further highlighted by a feature attribution analysis of the RexGCN model on the test set using integrated gradients which is a gradient-based attribution method. The integrated gradients was employed to quantify the sensitivity of the predicted redox potential to perturbations in the input features of individual atoms, which in turn, elucidated the chemical rationale underlying the GNN predictions. The resulting importance scores reflect each atom's contribution to the final prediction. Atoms with high scores exert a strong influence on the predicted potential, whereas near-zero scores indicate negligible impact.
As illustrated in Fig. 8, the analysis reveals a clear and consistent pattern where the highest importance scores are assigned to one or more atoms in the immediate coordination sphere of the central iron ion, i.e., the directly bonded ligand donor atoms. More specifically, about 80% of the atoms with a high important score (top 25% importance) were within 3.5 Å distance of the central iron ion, evident from the distribution of iron-neighbor distances shown in Fig. S11. This strong localization of attribution indicates that the GNN has learned to prioritize the immediate environment of the redox-active metal center, in agreement with fundamental coordination chemistry principles, rather than relying on spurious correlations from distal atoms within the ligand framework.
Conclusions
A scalable and efficient computational approach was developed here by utilizing tight-binding DFT and all-electron DFT methods to calculate the redox potential of transition metal complexes. To accurately capture the solvent effects in the redox calculations, our computational approach included both explicit water molecules and implicit continuum in the solvation structure. Redox potentials computed using our approach closely matched previous experimental values for several iron complexes, with an average absolute error of 0.13 V, demonstrating the reliability of our approach. This computational approach was then used to calculate the redox potential of 2267 iron complexes curated from the tmQM dataset, subsequently forming the Redox2k dataset.Analysis of the Redox2k dataset revealed that ligand chemistry and the local coordination environment around the iron center are key determinants of the redox potential. Specifically, two distinct clusters with contrasting redox behavior were identified in the top two component space of the PCA analysis performed on the Morgan fingerprints of the iron complexes in the redox dataset. A closer examination of the clusters revealed that complexes with inorganic ligands tended to exhibit high and positive redox potentials, while complexes with aromatic ligands containing nitrogen and/or oxygen as hetero atoms showed lower, possibly negative, redox potentials.
Next, a GNN framework was developed here to enable high-throughput screening of the vast chemical space of possible iron complexes. The GNN framework employed the xyz2mol method with new modifications to automatically generate fully featured graph data using only the 3D coordinates of the iron complex molecules. The GNN framework was then used to train four GNN models, namely RexGCN, RexGAT, DimeNet++, and SchNet, to learn and predict the redox potential of iron complexes from the GNN-Redoxdesolv-geo1450 dataset which was curated from the full Redox2k dataset through a series of filtering steps. The RexGCN model with engineered features provided the highest prediction accuracy, achieving an RMSE of 0.24 V ± 0.01 V and an R2 of 0.84 on the test set. The RexGAT and DimeNet++ models also achieved similar predictive performance compared to the RexGCN model whereas SchNet exhibited inferior performance.
Results here indicated that the quality of input features, whether engineered or learned, was more critical to model's performance than the complexity of the GNN architecture itself. Additionally, the predictive performance of the models dropped slightly when the full redox dataset was used for training by utilizing structures taken directly from the tmQM dataset. This observation highlighted the importance of the quality and consistency of the structures in the dataset rather than the dataset size to improve model performance. Finally, feature attribution analysis using integrated gradients revealed that atoms in the immediate coordination environment of the iron center were assigned greater importance by the GNN models in redox potential prediction. The methodology presented here provide a robust framework for exploring and predicting the redox properties of any transition metal complexes. This framework can accelerate the discovery of new materials for applications in redox flow batteries and catalysis.
Author contributions
F. H. B curated the dataset for GNN model training; developed all the GNN models; prepared the figures and tables; prepared the codes and data repositories; and contributed with most of the manuscript writing. H. H. and R. S. A. calibrated the DFT protocol and validated results. A. V. M conceived this study; performed all the simulations and data collection; outlined the manuscript. All the authors reviewed the manuscript and discussed the results.Conflicts of interest
The authors declare no competing interests.Data availability
Data and models are available at https://doi.org/10.5281/zenodo.18419091, and the code is available at https://github.com/argonne-lcf/gnnredox.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5dd00431d.
Acknowledgements
This material is based upon work supported by the U.S. Department of Energy’s (DOE) Office of Science Energy Earthshot Initiative as part of the Center for Steel Electrification by Electrosynthesis (C-STEEL) at Argonne National Laboratory (ANL) under contract DE-AC02-06CH11357. We acknowledge the computing resources provided on ‘Improv’ computing clusters operated by the Laboratory Computing Resource Center at ANL. This research used resources of the Argonne Leadership Computing Facility, a U.S. DOE Office of Science User Facility at ANL, which is supported by the Office of Science of the U.S. DOE under contract DE-AC02-06CH11357. A. V. M. and F. H. B. were supported by the Office of Science of the U.S. DOE under contract DE-AC02-06CH11357.References
- P. Lemoine, Electrochemistry of transition metal clusters, Coord. Chem. Rev., 1982, 47, 55–88 Search PubMed.
- M. Mouselly, H. Alawadhi and S. T. Senthilkumar, Current status of ferro-/ferricyanide for redox flow batteries, Curr. Opin. Electrochem., 2024, 48, 101581 CrossRef CAS.
- K. L. Hawthorne, J. S. Wainright and R. F. Savinell, Studies of iron-ligand complexes for an all-iron flow battery application, J. Electrochem. Soc., 2014, 161, A1662 CrossRef CAS.
- P. Schröder, D. Obendorf and T. Bechtold, Electrochemistry of iron (II/III)-N, N’-ethylene-bis-(o-hydroxyphenylglycine) complexes in aqueous solution indicates potential for use in redox flow batteries, ChemElectroChem, 2019, 6, 3311–3318 Search PubMed.
- J. Hannonen, A. Tuna, G. Gonzalez, E. Martínez González and P. Peljo, Investigation of Fe (II) Complexes with 1, 10-Phenanthroline and 2, 2’; 6’, 2 “-Terpyridine for Aqueous Flow Battery Applications, ChemElectroChem, 2025, 12, e202400574 CrossRef CAS.
- C. Noh, Y. Chung and Y. Kwon, Organometallic redox flow batteries using iron triethanolamine and cobalt triethanolamine complexes, J. Power Sources, 2020, 466, 228333 Search PubMed.
- D. Buccella, M. H. Lim and J. R. Morrow, Metals in Biology: From Metallomics to Trafficking, Inorg. Chem., 2019, 58, 13505–13508 CrossRef CAS PubMed.
- J. Karges, R. W. Stokes and S. M. Cohen, Metal complexes for therapeutic applications, Trends Chem., 2021, 3, 523–534 Search PubMed.
- T. Inomata, Coordination Chemistry: Molecular Science of Organic–Inorganic Complexes, Royal Society of Chemistry, 2024 Search PubMed.
- V. K. Praneeth, M. R. Ringenberg and T. R. Ward, Redox-active ligands in catalysis, Angew. Chem., Int. Ed., 2012, 51, 10228–10234 CrossRef CAS PubMed.
- K. Singh, A. Kundu and D. Adhikari, Ligand-based redox: catalytic applications and mechanistic aspects, ACS Catal., 2022, 12, 13075–13107 CrossRef CAS.
- A. Fürstner, Iron catalysis in organic synthesis: a critical assessment of what it takes to make this base metal a multitasking champion, ACS Cent. Sci., 2016, 2, 778–789 Search PubMed.
- J. M. Hudson, G. W. Luther III and Y.-P. Chin, Influence of organic ligands on the redox properties of Fe (II) as determined by mediated electrochemical oxidation, Environ. Sci. Technol., 2022, 56, 9123–9132 Search PubMed.
- M. A. Rizvi, Complexation modulated redox behavior of transition metal systems, Russ. J. Gen. Chem., 2015, 85, 959–973 CrossRef CAS.
- N. Arunachalam, S. Gugler, M. G. Taylor, C. Duan, A. Nandy, J. P. Janet, R. Meyer, J. Oldenstaedt, D. B. Chu and H. J. Kulik, Ligand additivity relationships enable efficient exploration of transition metal chemical space, J. Chem. Phys., 2022, 157, 184112 Search PubMed.
- N. Rahbani, P. de Silva, C. Bellay, S. Guihéneuf, T. Godet-Bar and E. Baudrin, Screening of First-Row Transition Metal Complexes for Aqueous Redox Flow Batteries: Experimental and Density Functional Theory Approaches, Meet. Abstr., 2023, 244, 2862 Search PubMed.
- N. Elgrishi, K. J. Rountree, B. D. McCarthy, E. S. Rountree, T. T. Eisenhart and J. L. Dempsey, A practical beginner's guide to cyclic voltammetry, J. Chem. Educ., 2018, 95, 197–206 CrossRef CAS.
- P. T. Kissinger and W. R. Heineman, Cyclic voltammetry, J. Chem. Educ., 1983, 60, 702 Search PubMed.
- R. Osteryoung and J. Osteryoung, Pulse voltammetric methods of analysis, Phil. Trans. Roy. Soc. Lond. Math. Phys. Sci., 1981, 302, 315–326 Search PubMed.
- J. G. Osteryoung and R. A. Osteryoung, Square wave voltammetry, Anal. Chem., 1985, 57, 101–110 CrossRef.
- V. Mirceski, S. Skrzypek and L. Stojanov, Square-wave voltammetry, ChemTexts, 2018, 4, 1–14 CrossRef.
- J. Ho, M. L. Coote, C. J. Cramer and D. G. Truhlar, Theoretical calculation of reduction potentials, Org. Electrochem., 2015, 5, 229–259 Search PubMed.
- M.-H. Baik and R. A. Friesner, Computing redox potentials in solution: Density functional theory as a tool for rational design of redox agents, J. Phys. Chem., 2002, 106, 7407–7412 Search PubMed.
- J. Moens, P. Jaque, F. De Proft and P. Geerlings, The study of redox reactions on the basis of conceptual DFT principles: EEM and vertical quantities, J. Phys. Chem., 2008, 112, 6023–6031 CrossRef CAS PubMed.
- H. Neugebauer, F. Bohle, M. Bursch, A. Hansen and S. Grimme, Benchmark study of electrochemical redox potentials calculated with semiempirical and DFT methods, J. Phys. Chem., 2020, 124, 7166–7176 CrossRef CAS PubMed.
- J. P. Janet, T. Z. Gani, A. H. Steeves, E. I. Ioannidis and H. J. Kulik, Leveraging cheminformatics strategies for inorganic discovery: application to redox potential design, Ind. Eng. Chem. Res., 2017, 56, 4898–4910 CrossRef CAS.
- H. Harb and R. S. Assary, Systematic improvement of redox potential calculation of Fe (III)/Fe (II) complexes using a three-layer micro-solvation model, Phys. Chem. Chem. Phys., 2025, 27, 10717–10729 RSC.
- A. Masliy, T. Grishaeva and A. Kuznetsov, Standard redox potentials of Fe (III) aqua complexes included into the cavities of cucurbit [n] urils (n= 6–8): a DFT forecast, J. Phys. Chem., 2019, 123, 5341–5346 Search PubMed.
- P. Comba, D. Faltermeier and B. Martin, Computational Approaches for Redox Potentials of Iron (IV)-oxido Complexes, Z. Anorg. Allg. Chem., 2020, 646, 1839–1845 Search PubMed.
- G. Cárdenas, P. Marquetand, S. Mai and L. González, A force field for a manganese-vanadium water oxidation catalyst: redox potentials in solution as showcase, Catalysts, 2021, 11, 493 CrossRef.
- M. Boniolo, P. Chernev, M. H. Cheah, P. A. Heizmann, P. Huang, S. I. Shylin, N. Salhi, M. K. Hossain, A. K. Gupta and J. Messinger, others Electronic and geometric structure effects on one-electron oxidation of first-row transition metals in the same ligand framework, Dalton Trans., 2021, 50, 660–674 Search PubMed.
- T. P. Senftle, S. Hong, M. M. Islam, S. B. Kylasa, Y. Zheng, Y. K. Shin, C. Junkermeier, R. Engel-Herbert, M. J. Janik and H. M. Aktulga, others The ReaxFF reactive force-field: development, applications and future directions, npj Comput. Mater., 2016, 2, 1–14 Search PubMed.
- K. Chenoweth, A. C. Van Duin and W. A. Goddard, ReaxFF reactive force field for molecular dynamics simulations of hydrocarbon oxidation, J. Phys. Chem., 2008, 112, 1040–1053 Search PubMed.
- F. H. Bhuiyan, Y.-S. Li, S. H. Kim and A. Martini, Shear-activation of mechanochemical reactions through molecular deformation, Sci. Rep., 2024, 14, 2992 CrossRef CAS PubMed.
- M. M. Islam, G. Kolesov, T. Verstraelen, E. Kaxiras and A. C. Van Duin, eReaxFF: a pseudoclassical treatment of explicit electrons within reactive force field simulations, J. Chem. Theor. Comput., 2016, 12, 3463–3472 CrossRef CAS PubMed.
- B. Evangelisti, K. A. Fichthorn and A. C. Van Duin, Development and initial applications of an e-ReaxFF description of Ag nanoclusters, J. Chem. Phys., 2020, 153, 104106 CrossRef CAS PubMed.
- M. J. Hossain, P. Gaikwad, Y. K. Shin, J. A. Schulze, K. A. Penrod, M. Li, Y. Lin, G. Pawar and A. C. van Duin, eReaxFF force field development for BaZr0. 8Y0. 2O3-δ solid oxide electrolysis cells applications, npj Comput. Mater., 2024, 10, 136 Search PubMed.
- Y. Yang, S. Zhang, K. D. Ranasinghe, O. Isayev and A. E. Roitberg, Machine learning of reactive potentials, Annu. Rev. Phys. Chem., 2024, 75, 371–395 Search PubMed.
- D. M. Anstine and O. Isayev, Machine learning interatomic potentials and long-range physics, J. Phys. Chem., 2023, 127, 2417–2431 Search PubMed.
- A. Grisafi and M. Ceriotti, Incorporating long-range physics in atomic-scale machine learning, J. Chem. Phys., 2019, 151, 204105 Search PubMed.
- J. Behler and G. Csányi, Machine learning potentials for extended systems: a perspective, Eur. Phys. J. B, 2021, 94, 142 Search PubMed.
- E. Kocer, R. E. Haouari, C. Dellago, and J. Behler, Machine learning potentials for redox chemistry in solution, arXiv, 2024, preprint, arXiv:2410.03299, DOI:10.48550/arXiv.2410.03299.
- P. Gao, D. Kochan, Y.-H. Tang, X. Yang and E. G. Saldanha, Machine learning for the redox potential prediction of molecules in organic redox flow battery, J. Power Sources, 2025, 629, 236035 Search PubMed.
- G. Corso, H. Stark, S. Jegelka, T. Jaakkola and R. Barzilay, Graph neural networks, Nat. Rev. Methods Primers, 2024, 4, 17 CrossRef CAS.
- Y. Wang, Z. Li, and A. Barati Farimani, Machine learning in molecular sciences, Springer, 2023, pp 21–66 Search PubMed.
- X. Wu, H. Wang, Y. Gong, D. Fan, P. Ding, Q. Li and Q. Qian, Graph neural networks for molecular and materials representation, J. Inf. Rec. Mater., 2023, 3, N–A Search PubMed.
- L. Jia, É. Brémond, L. Zaida, B. Gaüzère, V. Tognetti and L. Joubert, Predicting redox potentials by graph-based machine learning methods, J. Comput. Chem., 2024, 45, 2383–2396 Search PubMed.
- M. D. Chaka, C. A. Geffe, A. Rodriguez, N. Seriani, Q. Wu and Y. S. Mekonnen, High-throughput screening of promising redox-active molecules with molgat, ACS Omega, 2023, 8, 24268–24278 Search PubMed.
- D. Balcells and B. B. Skjelstad, tmQM dataset—quantum geometries and properties of 86k transition metal complexes, J. Chem. Inf. Model., 2020, 60, 6135–6146 Search PubMed.
- M. H. Rasmussen, M. Strandgaard, J. Seumer, L. K. Hemmingsen, A. Frei, D. Balcells and J. H. Jensen, SMILES all around: structure to SMILES conversion for transition metal complexes, J. Cheminf., 2025, 17, 1–13 Search PubMed.
- F. Neese, Software update: the ORCA program system, version 5.0, WIRES Comput. Molec. Sci., 2022, 12, e1606 Search PubMed.
- F. Neese, F. Wennmohs, U. Becker and C. Riplinger, The ORCA quantum chemistry program package, J. Chem. Phys., 2020, 152, L224108 CrossRef PubMed.
- C. Bannwarth, S. Ehlert and S. Grimme, GFN2-xTB—An Accurate and Broadly Parametrized Self-Consistent Tight-Binding Quantum Chemical Method with Multipole Electrostatics and Density- Dependent Dispersion Contributions, J. Chem. Theory Comput., 2019, 15, 1652–1671 CrossRef CAS PubMed.
- S. Ehlert, M. Stahn, S. Spicher and S. Grimme, Robust and efficient implicit solvation model for fast semiempirical methods, J. Chem. Theory Comput., 2021, 17, 4250–4261 CrossRef CAS PubMed.
- A. D. Becke, Density-functional thermochemistry. III. The role of exact exchange, J. Chem. Phys., 1993, 98, 5648–5652 CrossRef CAS.
- J. P. Perdew, J. Tao, V. N. Staroverov and G. E. Scuseria, Meta-generalized gradient approximation: Explanation of a realistic nonempirical density functional, J. Chem. Phys., 2004, 120, 6898–6911 Search PubMed.
- Q. Zhang, A. Khetan and S. Er, Comparison of computational chemistry methods for the discovery of quinone-based electroactive compounds for energy storage, Sci. Rep., 2020, 10, 22149 Search PubMed.
- N. Rahbani, P. de Silva and E. Baudrin, Density Functional Theory-Based Protocol to Calculate the Redox Potentials of First-row Transition Metal Complexes for Aqueous Redox Targeting Flow Batteries, ChemSusChem, 2023, 16, e202300482 Search PubMed.
- K. P. Jensen, Bioinorganic chemistry modeled with the TPSSh density functional, Inorg. Chem., 2008, 47, 10357–10365 CrossRef CAS PubMed.
- G. Liang, N. J. DeYonker, X. Zhao and C. E. Webster, Prediction of the reduction potential in transition-metal containing complexes: How expensive? For what accuracy?, J. Comput. Chem., 2017, 38, 2430–2438 CrossRef CAS PubMed.
- L. E. Roy, E. Jakubikova, M. G. Guthrie and E. R. Batista, Calculation of one-electron redox potentials revisited. Is it possible to calculate accurate potentials with density functional methods?, J. Phys. Chem., 2009, 113, 6745–6750 CrossRef CAS PubMed.
- P. A. Rock, The standard oxidation potential of the ferrocyanide-ferricyanide electrode at 25 and the entropy of ferrocyanide ion, J. Phys. Chem., 1966, 70, 576–580 CrossRef CAS.
- K. Gong, F. Xu, J. B. Grunewald, X. Ma, Y. Zhao, S. Gu and Y. Yan, All-soluble all-iron aqueous redox-flow battery, ACS Energy Lett., 2016, 1, 89–93 Search PubMed.
- T. F. Hughes and R. A. Friesner, Development of accurate DFT methods for computing redox potentials of transition metal complexes: results for model complexes and application to cytochrome P450, J. Chem. Theor. Comput., 2012, 8, 442–459 CrossRef CAS PubMed.
- M. Shin, S. Oh, H. Jeong, C. Noh, Y. Chung, J. W. Han and Y. Kwon, Aqueous redox flow battery using iron 2, 2-bis (hydroxymethyl)-2, 2’, 2’-nitrilotriethanol complex and ferrocyanide as newly developed redox couple, Int. J. Energy Res., 2022, 46, 8175–8185 CrossRef CAS.
- M. Shin, C. Noh, Y. Chung and Y. Kwon, All iron aqueous redox flow batteries using organometallic complexes consisting of iron and 3-[bis (2-hydroxyethyl) amino]-2-hydroxypropanesulfonic acid ligand and ferrocyanide as redox couple, Chem. Eng. J., 2020, 398, 125631 CrossRef CAS.
- J. Yang, W. Wei, C. Zhou, H. Yan, H. Che, L. Hao, X. Tan, A. W. Robertson, T.-S. Wu and Y.-L. Soo, others High-Stable All-Iron Redox Flow Battery with Innovative Anolyte based on Steric Hindrance Regulation, Angew. Chem., 2025, 137, e202414452 Search PubMed.
- A. Jain, S. P. Ong, W. Chen, B. Medasani, X. Qu, M. Kocher, M. Brafman, G. Petretto, G.-M. Rignanese and G. Hautier, others FireWorks: a dynamic workflow system designed for high-throughput applications, Concurrency Comput. Pract. Ex., 2015, 27, 5037–5059 CrossRef.
- A. H. Larsen, J. J. Mortensen, J. Blomqvist, I. E. Castelli, R. Christensen, M. Dułak, J. Friis, M. N. Groves, B. Hammer and C. Hargus, others The atomic simulation environment—a Python library for working with atoms, J. Phys.: Condens. Matter, 2017, 29, 273002 CrossRef PubMed.
- D. Rogers and M. Hahn, Extended-connectivity fingerprints, J. Chem. Inf. Model., 2010, 50, 742–754 Search PubMed.
- M. A. Skinnider, R. G. Stacey, D. S. Wishart and L. J. Foster, Chemical language models enable navigation in sparsely populated chemical space, Nat. Mach. Intell., 2021, 3, 759–770 Search PubMed.
- J. Deng, Z. Yang, H. Wang, I. Ojima, D. Samaras and F. Wang, A systematic study of key elements underlying molecular property prediction, Nat. Commun., 2023, 14, 6395 CrossRef CAS PubMed.
- A. Mayr, G. Klambauer, T. Unterthiner, M. Steijaert, J. K. Wegner, H. Ceulemans, D.-A. Clevert and S. Hochreiter, Large-scale comparison of machine learning methods for drug target prediction on ChEMBL, Chem. Sci., 2018, 9, 5441–5451 Search PubMed.
- K. Heikamp and J. Bajorath, How do 2D fingerprints detect structurally diverse active compounds? Revealing compound subset-specific fingerprint features through systematic selection, J. Chem. Inf. Model., 2011, 51, 2254–2265 CrossRef CAS PubMed.
- T. Kipf and M. Welling, Semi-Supervised Classification with Graph Convolutional Networks, arXiv, 2016, preprint, arXiv:1609.02907, DOI:10.48550/arXiv.1609.02907.
- P. Veličković, G. Cucurull, A. Casanova, A. Romero, P. Lio, and Y. Bengio, Graph attention networks, arXiv, 2017, preprint, arXiv:1710.10903, 10.48550/arXiv:1710.10903 Search PubMed.
- J. Gasteiger, S. Giri, J. T. Margraf, and S. Günnemann, Fast and uncertainty-aware directional message passing for non-equilibrium molecules, arXiv, 2020, preprint, arXiv:2011.14115, DOI:10.48550/arXiv.2011.14115.
- K. Schütt, P.-J. Kindermans, H. E. Sauceda Felix, S. Chmiela, A. Tkatchenko and K.-R. Müller, Schnet: A continuous-filter convolutional neural network for modeling quantum interactions, Adv. Neural Inf. Process. Syst., 2017, 30 Search PubMed.
- K. T. Schütt, H. E. Sauceda, P.-J. Kindermans, A. Tkatchenko and K.-R. Müller, Schnet–a deep learning architecture for molecules and materials, J. Chem. Phys., 2018, 148 CrossRef PubMed.
- O. Wieder, S. Kohlbacher, M. Kuenemann, A. Garon, P. Ducrot, T. Seidel and T. Langer, A compact review of molecular property prediction with graph neural networks, Drug Discovery Today: Technol., 2020, 37, 1–12 CrossRef PubMed.
- J. Gilmer, S. S. Schoenholz, P. F. Riley, O. Vinyals, and G. E. Dahl, Neural message passing for quantum chemistry, International conference on machine learning, 2017, pp 1263–1272 Search PubMed.
- M. Schlichtkrull, T. N. Kipf, P. Bloem, R. Van Den Berg, I. Titov, and M. Welling, Modeling relational data with graph convolutional networks, European Semantic Web Conference, 2018, pp 593–607 Search PubMed.
- E. Isufi, F. Gama and A. Ribeiro, EdgeNets: Edge varying graph neural networks, IEEE Trans. Pattern Anal. Mach. Intell., 2021, 44, 7457–7473 Search PubMed.
- L. Ward, B. Blaiszik, I. Foster, R. S. Assary, B. Narayanan and L. Curtiss, Machine learning prediction of accurate atomization energies of organic molecules from low-fidelity quantum chemical calculations, MRS Commun., 2019, 9, 891–899 CrossRef CAS.
- P. Reiser, M. Neubert, A. Eberhard, L. Torresi, C. Zhou, C. Shao, H. Metni, C. van Hoesel, H. Schopmans and T. Sommer, others Graph neural networks for materials science and chemistry, Mater. Today Commun., 2022, 3, 93 Search PubMed.
- A. Guzman-Pando, G. Ramirez-Alonso, C. Arzate-Quintana and J. Camarillo-Cisneros, Deep learning algorithms applied to computational chemistry, Mol. Diversity, 2024, 28, 2375–2410 CrossRef CAS PubMed.
- G. S. Nambafu, A. M. Hollas, S. Zhang, P. S. Rice, D. Boglaienko, J. L. Fulton, M. Li, Q. Huang, Y. Zhu and D. M. Reed, others Phosphonate-based iron complex for a cost-effective and long cycling aqueous iron redox flow battery, Nat. Commun., 2024, 15, 2566 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |