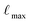Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
High-performance training and inference for deep equivariant interatomic potentials
Chuin Wei
Tan†
 a,
Marc L.
Descoteaux†
a,
Marc L.
Descoteaux†
 a,
Mit
Kotak
a,
Mit
Kotak
 b,
Gabriel
de Miranda Nascimento
c,
Seán R.
Kavanagh
b,
Gabriel
de Miranda Nascimento
c,
Seán R.
Kavanagh
 d,
Laura
Zichi
a,
Menghang
Wang
d,
Laura
Zichi
a,
Menghang
Wang
 a,
Aadit
Saluja
a,
Yizhong R.
Hu
a,
Aadit
Saluja
a,
Yizhong R.
Hu
 a,
Tess
Smidt
e,
Anders
Johansson
a,
Tess
Smidt
e,
Anders
Johansson
 f,
William C.
Witt
a,
Boris
Kozinsky
f,
William C.
Witt
a,
Boris
Kozinsky
 *ag and
Albert
Musaelian
*ah
*ag and
Albert
Musaelian
*ah
aJohn A. Paulson School of Engineering and Applied Sciences, Harvard University, Cambridge, MA, USA. E-mail: bkoz@g.harvard.edu; amusaelian@alumni.harvard.edu
bCenter for Computational Science and Engineering, Massachusetts Institute of Technology, Cambridge, MA, USA
cDepartment of Materials Science and Engineering, Massachusetts Institute of Technology, Cambridge, MA, USA
dCenter for the Environment, Harvard University, Cambridge, MA, USA
eResearch Laboratory of Electronics, Massachusetts Institute of Technology, Cambridge, MA, USA
fSandia National Laboratories, Albuquerque, NM, USA
gRobert Bosch LLC Research and Technology Center, Watertown, MA, USA
hMirian Technologies Inc., Boston, MA, USA
First published on 26th March 2026
Abstract
Machine learning interatomic potentials, particularly those based on deep equivariant neural networks, have demonstrated state-of-the-art accuracy and computational efficiency in atomistic modeling tasks like molecular dynamics and high-throughput screening. The size of datasets and demands of downstream workflows are growing rapidly, making robust and scalable software essential. This work presents a major overhaul of the NequIP framework focusing on multi-node parallelism, computational performance, and extensibility. The redesigned framework supports distributed training on large datasets and removes barriers preventing full utilization of the PyTorch 2.0 compiler at train time. We demonstrate this acceleration in a case study by training Allegro models on the SPICE 2 dataset of organic molecular systems. For inference, we introduce the first end-to-end infrastructure that uses the PyTorch Ahead-of-Time Inductor compiler for machine learning interatomic potentials. Additionally, we implement a custom kernel for the Allegro model's most expensive operation, the tensor product. Together, these advancements speed up molecular dynamics calculations on system sizes of practical relevance by up to factors of 5 to 18.
1. Introduction
Machine learning interatomic potentials (MLIPs) have become an essential tool for computational materials science and chemistry.1,2 Their widespread adoption has driven the development of specialized software frameworks for their training and inference.3–11 Meanwhile, the growing availability of large and diverse datasets, such as SPICE,12 MPTrj,13 Alexandria,14 OMat24,15 MatPES,16 and OMol25,17 has enabled the development of more generalizable MLIPs capable of serving as pretrained potentials for a wide range of applications across physics and chemistry.13,18–28 In this rapidly evolving landscape, scalable, efficient, and adaptable software infrastructure is essential. Frameworks must not only handle the computational demands of large datasets but also support novel training paradigms, facilitate efficient hyperparameter optimization, and integrate with diverse hardware environments.This work presents a major revamp of the NequIP software framework for MLIPs to dramatically improve computational performance and versatility across training and inference tasks of all sizes. Within this framework, NequIP29 pioneered the development of interatomic potentials based on equivariant neural networks, which incorporate rotational, mirror, and other physical symmetries directly in the structure of the model, while Allegro30 extended this approach with a scalable design for large-scale systems.31 More generally, equivariant model architectures have been shown to improve data efficiency, enhance generalization, and achieve state-of-the-art accuracy in MLIP-based simulations,29,30,32,33 and a number of further leading models now adapt the NequIP architecture and/or codebase.21,22,34 However, equivariant neural networks pose practical computational challenges due to their use of mathematical operations for which libraries like PyTorch do not provide optimized implementations. Native PyTorch implementations of these operations tend to suffer from kernel launch overhead and the materialization of large intermediate tensors, which can limit both speed and memory efficiency. Overcoming these limitations with compiler-based optimization and custom GPU kernels is a core motivation of this work.
We begin in Section 2 by describing the methods we used to accelerate training and inference. Then, we present a case study demonstrating the performance improvements using the Allegro deep equivariant network architecture30 in Section 3.
2. Training and inference acceleration
2.1. TorchInductor compilation
The NequIP framework is built on top of the PyTorch neural network library.35 PyTorch 2.0 introduced TorchInductor, a compiler that transforms graphs of high-level PyTorch operations into highly-optimized, device-specific code.36 On GPUs, TorchInductor leverages Triton,37 a hardware-agnostic language, to generate optimized kernels, while on CPUs, it produces C++ and OpenMP code.38 Compared to the previously used TorchScript compiler (deprecated as of PyTorch 2.9), TorchInductor provides more advanced compiler optimizations, including extensive operator fusion, inlining, and loop/layout reordering. By aggressively fusing kernels, TorchInductor reduces intermediate memory reads and writes, improving computational efficiency for memory-bound workloads. These optimizations have been shown to yield significant performance gains across diverse machine learning models,39–41 and are especially advantageous for the deep learning interatomic potential models considered in this work.TorchInductor serves as the compiler backend and can be invoked through two complementary compilation modes: just-in-time (JIT) compilation via , and ahead-of-time (AOT) compilation through AOT Inductor. Both modes share almost the same underlying compiler infrastructure and optimization pipeline, differing mainly when and where compilation occurs—JIT compilation happens dynamically within Python, whereas AOT compilation produces standalone binaries for use in non-Python environments. As will be discussed, JIT compilation with
, and ahead-of-time (AOT) compilation through AOT Inductor. Both modes share almost the same underlying compiler infrastructure and optimization pipeline, differing mainly when and where compilation occurs—JIT compilation happens dynamically within Python, whereas AOT compilation produces standalone binaries for use in non-Python environments. As will be discussed, JIT compilation with 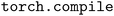 is used for training (Section 2.2) while AOT compilation is used for Python-free inference where the Python-based PyTorch JIT machinery is unavailable (Section 2.3).
is used for training (Section 2.2) while AOT compilation is used for Python-free inference where the Python-based PyTorch JIT machinery is unavailable (Section 2.3).
The main challenge in applying TorchInductor to NequIP and Allegro is the models' use of automatic differentiation to predict forces and virials as exact derivatives of the potential energy. Enforcing these derivative relationships ensures that forces remain conservative fields of the energy, which is essential for maintaining energy conservation in molecular dynamics simulations.42 Evidence suggests that using conservative forces improves the accuracy of physical property predictions,27 particularly for phonon-related properties,43 and enhances the stability of geometry relaxation tasks.44 Computing the derivative, however, is not a primitive PyTorch operation that is supported directly by the compiler machinery. To overcome this challenge, we adapted our code to trace the entire model, including the calculation of the derivative(s), into a single computational graph that is represented using PyTorch's 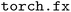 library.45 The
library.45 The 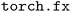 graph resulting from this procedure represents both the forward pass, which computes the potential energy, and the backward-mode automatic differentiation pass, which computes forces and other derivatives. Because this graph contains only primitive PyTorch operations, it is compatible both with train-time compilation (Section 2.2) and ahead-of-time compilation for inference (Section 2.3).
graph resulting from this procedure represents both the forward pass, which computes the potential energy, and the backward-mode automatic differentiation pass, which computes forces and other derivatives. Because this graph contains only primitive PyTorch operations, it is compatible both with train-time compilation (Section 2.2) and ahead-of-time compilation for inference (Section 2.3).
A key advantage of our TorchInductor approach is that it can accommodate dynamic tensor shapes, while many other neural network compilers enforce static tensor shapes. In MLIP-based calculations—molecular dynamics in particular—both the number of atoms and the number of pairs of neighboring atoms input to the model can vary significantly over the course of a simulation. TorchInductor generates a single set of optimized kernels that are applicable to the entire range of relevant dynamic tensor shapes.
2.2. Compiled and distributed training
We apply TorchInductor to accelerate model training by using the procedure described in Section 2.1 to convert the entire model into a computational graph containing only primitive PyTorch operations. We then apply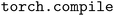 to this graph to generate optimized kernels for both the forward pass, which computes all of the model's predictions, and the train-time backward pass, which computes gradients of the loss function with respect to model parameters.
to this graph to generate optimized kernels for both the forward pass, which computes all of the model's predictions, and the train-time backward pass, which computes gradients of the loss function with respect to model parameters.
To enable training on large datasets,13–16 we have added distributed multi-GPU training to the NequIP framework. We adopt a Distributed Data Parallel (DDP) paradigm46via PyTorch Lightning.47 Instead of using PyTorch's standard DDP implementation, which employs a gradient bucketing strategy that overlaps computation with communication, we implemented a custom DDP approach that performs communications only after the full backwards pass has completed. This design choice allows TorchInductor to compile the entire backward pass, which can reduce overhead and introduce additional opportunities for fusion, rather than splitting it into multiple subgraphs with communication operations in between. Practically, omitting gradient bucketing has not imposed meaningful limitations on our MLIP training, but a gradient bucketing strategy for larger models could be reintroduced in the future if needed.
2.3. Ahead-of-time compilation for inference
It is often important to integrate MLIP models with high-performance simulation codes that are written in low-level languages, such as the leading molecular dynamics code LAMMPS,48 which is written in C++. The NequIP framework previously interfaced with LAMMPS by exporting models to the TorchScript format and providing dedicated plugins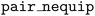 and
and 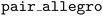 to load and call TorchScript-compiled models directly inside LAMMPS. This approach has subsequently been adopted by other deep learning interatomic potential frameworks such as MACE,32 SevenNet,22 and BAMBOO.49 The
to load and call TorchScript-compiled models directly inside LAMMPS. This approach has subsequently been adopted by other deep learning interatomic potential frameworks such as MACE,32 SevenNet,22 and BAMBOO.49 The 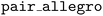 plugin also uses the Kokkos performance portability library50 to reduce overhead in the interface between Allegro and LAMMPS by eliminating CPU–GPU data transfers wherever possible.31 This feature also allows LAMMPS to use GPU-aware MPI communication.
plugin also uses the Kokkos performance portability library50 to reduce overhead in the interface between Allegro and LAMMPS by eliminating CPU–GPU data transfers wherever possible.31 This feature also allows LAMMPS to use GPU-aware MPI communication.
Here, we introduce the first end-to-end use of AOT Inductor (AOTI) compilation for MLIPs as a replacement for TorchScript export. AOTI is a variant of TorchInductor specialized to export compiled PyTorch models as self-contained native code for use in non-Python environments. AOTI compilation allows us to realize the performance benefits of MLIP compilation, including advanced fusion, inlining, and loop/layout reordering, in high-performance codes like LAMMPS without resorting to complicated, high-overhead solutions such as embedded Python interpreters.51,52 Models compiled with AOTI can also still be used in Python-based codes like the Atomic Simulation Environment (ASE)53 and TorchSim.54
2.4. Optimized tensor product kernel
Custom kernels that implement fusion, scheduling, and memory-access patterns that are more advanced than those of current compilers have demonstrated significant success in accelerating deep learning models.55,56 There have been analogous efforts for invariant neural networks57 and more recently equivariant neural networks that have focused on optimizing various tensor product operations, which are a central computational bottleneck in equivariant MLIP architectures.11,19,20,58 While Allegro's tensor product operation already has a highly optimized pure PyTorch implementation,31 it remains the most computationally expensive part of the Allegro architecture. Profiling revealed that TorchInductor currently cannot fuse the entire pure PyTorch implementation of the Allegro tensor product into a single GPU kernel. Instead, it generates two separate kernels that require costly intermediate memory reads and writes.To overcome the fusion limitations of TorchInductor, we implemented fused custom kernels for the Allegro tensor product using Triton, a cross-platform language for writing high-performance compute kernels.37 Because Triton is the native backend for TorchInductor, the kernel can be seamlessly integrated with our compilation infrastructure. We implemented kernels for the tensor product and its first derivative to accelerate the inference-time prediction of energy and its first derivatives, like forces. Importantly, custom Triton kernels integrate seamlessly with AOTI, ensuring that our custom kernel can be exported for use in LAMMPS and other inference software. The additional second-derivative kernels required for training are left for future work and would also straightforwardly integrate with train-time TorchInductor compilation.
Our custom kernel takes advantage of the mathematical structure of the tensor product by using a compressed sparse format59 to represent the combined tensor of Wigner 3-j contraction coefficients described in previous work.31 It also avoids materializing intermediate outer products between the input tensors, which results in significant memory savings. We direct to ref. 60 for more implementation details.
3. Case study: training and inference of SPICE 2 models
To demonstrate the redesigned NequIP framework, we trained Allegro models on the SPICE 2 dataset12,23 and benchmarked their performance in molecular dynamics. This dataset contains more than two million atomic configurations of organic systems, including drug-like molecules, amino acids, ions, water clusters, and their interactions, with energies and forces calculated using density functional theory (DFT) with the ωB97M-D3(BJ) functional.61,62 The dataset was designed to capture key inter- and intra-molecular interactions necessary for accurately modeling protein and drug-like systems, with molecular structures and conformations obtained by constructing chemically diverse systems, sampling from molecular dynamics at varied temperatures, relaxing DFT-sized substructures from larger complexes, and scanning specific interaction distances.12,23 Several general-purpose biomolecular MLIPs have been trained on the SPICE 2 data,19,23,63 leading to studies of the crystallization63 and hydration free energies64 of small organic molecules with higher accuracy than classical force fields. Accurate and efficient MLIPs for biomolecular systems are expected to have significant impact on drug discovery workflows.65The SPICE 2 dataset contains systems with a range of total charge states, which means that a given atomic configuration can have multiple possible DFT energy and force labels depending on the total charge used to compute them. To avoid this degeneracy, the Allegro models in this work were trained on a subset of the SPICE 2 data limited to systems with neutral total charge. This training strategy is less restrictive than that of Kovács et al.,19 who trained only on systems with neutral per-atom formal charge, but more restrictive than the approach of Eastman et al.,23 who trained on the full SPICE 2 dataset.
Using this subset of SPICE 2, we prepared three Allegro models with different cost-accuracy trade-offs and will refer to them as “small”, “medium”, and “large”. The main differences between these models are the maximum rotation order 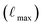 , the number of tensor features, and the number of Allegro layers, which are listed in Table 1. These hyperparameters control the cost and complexity of the Allegro tensor products that mix the models' equivariant features.30 All models employ a radial cutoff rmax of 6.0 Å, and the SI gives a full description of the hyperparameters and training procedure.
, the number of tensor features, and the number of Allegro layers, which are listed in Table 1. These hyperparameters control the cost and complexity of the Allegro tensor products that mix the models' equivariant features.30 All models employ a radial cutoff rmax of 6.0 Å, and the SI gives a full description of the hyperparameters and training procedure.
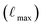 , number of tensor features, and number of Allegro layers
, number of tensor features, and number of Allegro layers
3.1. Model accuracy
To contextualize the differences in computational cost between the three models, we evaluate their accuracies using two benchmark datasets: the official SPICE 2 test set23 and the TorsionNet 500 benchmark66 recomputed at the SPICE 2 level of theory by Eastman et al.23Designed to assess the generalizability of MLIPs trained on SPICE 2, the SPICE 2 test set contains systems distinct from those in the training set, and it is divided into four categories: small ligands, large ligands, pentapeptides, and dimer pairs.23 To understand how our models (trained on neutral-total-charge data) perform on atomic configurations with different total- and per-atom-charge states, we evaluate the Allegro models on three subsets of the SPICE 2 test set: an unrestricted set that includes all systems in the test set regardless of charge state, a subset restricted to systems with neutral total charge (consistent with the data used to train the Allegro models), and a more restrictive subset of systems that contain only atomic species with neutral formal per-atom charge. Fig. 1 shows a consistent trend across all system types and charge schemes where accuracy improves from the small to medium to large model. While energy errors are large on the unfiltered test set, which includes molecular charge states that are entirely absent from the training data, the models perform well on both the neutral total charge subset and the neutral per-atom formal charge subset. Force errors on the unfiltered test set are only slightly larger than on the two filtered subsets. This disparity in performance across the three subsets is attributed to the fact that the present models have no explicit mechanism for representing global charge state. Approaches that incorporate the total charge as an input feature17 or that employ global charge-equilibration schemes that enforce global charge conservation67,68 could be integrated in future work for the models to distinguish geometrically similar configurations with different global charges.
 | ||
| Fig. 1 Allegro model results for the SPICE 2 test set.23 Energy and force mean absolute error (MAE) of the three Allegro models on three different subsets of the SPICE 2 test set: (1) the original unrestricted test set, (2) a subset containing systems with neutral total charge, and (3) a subset of systems that contain only atomic species with neutral per-atom formal charge. The error metrics are grouped by system type: large ligands, small ligands, peptides, and dimers. | ||
The second benchmark dataset, TorsionNet 500, also considered by Kovács et al.19 and Eastman et al.,23 assesses an MLIP's ability to predict the relative energy differences between molecular conformers. It contains 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 atomic configurations scanning through different values for the torsion angles along one bond in 500 drug-like molecules.66 A key property of a torsion angle scan is the height of its energy barrier, which controls the likelihood of a conformational change involving that torsional angle. Table 2 shows that the three Allegro models have barrier height errors comparable to those of Kovács et al.19 and Eastman et al.23 (see Table 1 from the SI for a detailed comparison between model accuracies and hyperparameters).
000 atomic configurations scanning through different values for the torsion angles along one bond in 500 drug-like molecules.66 A key property of a torsion angle scan is the height of its energy barrier, which controls the likelihood of a conformational change involving that torsional angle. Table 2 shows that the three Allegro models have barrier height errors comparable to those of Kovács et al.19 and Eastman et al.23 (see Table 1 from the SI for a detailed comparison between model accuracies and hyperparameters).
| Metric | Small model | Medium model | Large model |
|---|---|---|---|
| Barrier MAE [meV] | 22.75 | 15.42 | 11.37 |
| Barrier RMSE [meV] | 32.36 | 21.77 | 15.38 |
Additional accuracy benchmarks for a held-out 5% portion of the SPICE 2 training set, unused in our training, are presented in the SI.
3.2. Train-time acceleration
To demonstrate the efficiency and scalability of the redesigned NequIP infrastructure for MLIP training, we compared the cost to train the medium-sized Allegro model on the SPICE 2 dataset using the previous TorchScript compiler and the new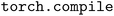 in a distributed multi-GPU setting. We performed the evaluation on both AMD MI250X GPUs using the Frontier supercomputer at the Oak Ridge Leadership Computing Facility (OLCF) and NVIDIA A100 GPUs using the Perlmutter supercomputer at the National Energy Research Scientific Computing Center (NERSC). To reflect realistic training conditions, all metrics, callbacks, logging, and other sources of overhead typical of production training runs were included in the timings.
in a distributed multi-GPU setting. We performed the evaluation on both AMD MI250X GPUs using the Frontier supercomputer at the Oak Ridge Leadership Computing Facility (OLCF) and NVIDIA A100 GPUs using the Perlmutter supercomputer at the National Energy Research Scientific Computing Center (NERSC). To reflect realistic training conditions, all metrics, callbacks, logging, and other sources of overhead typical of production training runs were included in the timings.
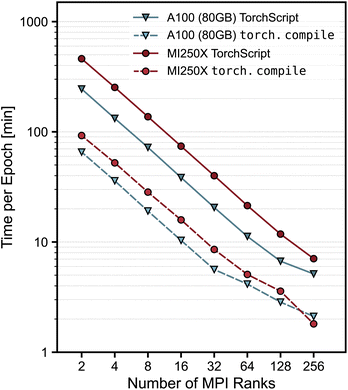
Fig. 2 presents the training performance as the number of MPI ranks increases, where each MPI rank runs a fixed per-rank local batch size of eight atomic configurations. In this scaling regime, the effective total global batch size increases with the number of ranks, which allows the entire dataset to be processed (a single “epoch”) in a smaller number of stochastic gradient descent steps, reducing the wall time per epoch. Distributed training scales well up to 128 ranks. At 256 ranks, it remains reasonably efficient and achieves parallel efficiencies of 40% and 24% on the AMD and NVIDIA systems, respectively, when running 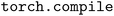 and using the 2-rank performance as a baseline. We observe that training with
and using the 2-rank performance as a baseline. We observe that training with 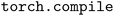 achieves between 2.4–5.0 times speedups over TorchScript on MI250X and A100 (80 GB) GPUs across the range of MPI ranks investigated.
achieves between 2.4–5.0 times speedups over TorchScript on MI250X and A100 (80 GB) GPUs across the range of MPI ranks investigated.
3.3. Inference acceleration
Next, we investigate the inference speed of the three Allegro models, highlighting how AOTI and the custom kernel provide dramatic improvements in throughput and memory efficiency. To complete these experiments, we used the new tool to portably archive the models, which were trained on Frontier, before compiling them on each inference platform using
tool to portably archive the models, which were trained on Frontier, before compiling them on each inference platform using  . All inference benchmarks were performed in LAMMPS using our Kokkos-based integration for Allegro models.
. All inference benchmarks were performed in LAMMPS using our Kokkos-based integration for Allegro models.
First, for a single AMD MI250X, NVIDIA A100 (80 GB), and NVIDIA H100 GPU, we compare molecular dynamics performance for both small and large systems using: (1) small-molecule benchmarks (Fig. 3), introduced by Eastman et al.,23 consisting of 25-, 50- and 100-atom organic molecules in vacuum, and (2) periodic boxes of water (Fig. 4) at a density of 1 g cm−3 and temperature of 300 K with system sizes ranging from 24 to 5184 atoms.
 | ||
| Fig. 3 Single-rank inference acceleration on small molecule systems. The inference speed in LAMMPS of the small, medium, and large Allegro models deployed using TorchScript, AOTI, or AOTI with the optimized tensor product kernel (AOTI + custom TP) on small molecule systems ranging from 25 to 100 atoms without periodic boundary conditions,23 for AMD MI250X, NVIDIA A100 (80 GB), and NVIDIA H100 GPUs. The inference speeds were averaged over three runs with different random seeds for the initial velocities generated by LAMMPS. One MPI rank was used. Note that one MPI rank corresponds to one of the two available graphics compute dies on an MI250X device. | ||
For all systems, AOTI exhibits a consistent performance advantage over TorchScript, and further speedups are achieved with the custom Triton tensor product. The speedup factor tends to be greater for the larger Allegro models that have correspondingly more expensive tensor product operations for the custom kernel to accelerate. Across all hardware and models, the small molecule examples (Fig. 3) see accelerations ranging from 4–18 times when comparing the TorchScript baseline to AOTI with the optimized TP; the same range of speedups is seen in the water simulations (Fig. 4).
Fig. 4 demonstrates that AOTI improves memory efficiency and that the combination of AOTI and the optimized tensor product kernel dramatically increases the maximum number of atoms that can fit on one rank. While the large model runs out of memory at water system sizes of around 100 atoms on all GPU types when using TorchScript, AOTI compilation extends this limit to 288 atoms. The custom kernel then dramatically increases the maximum system size that the large model can run on one rank to 4320 atoms on an MI250X (specifically, one of the two available graphics compute dies), 4320 atoms on an A100 (80 GB), and 5184 atoms on an H100.
We also conducted multi-GPU strong scaling tests with the medium model on two all-atom water-solvated biomolecules: the 23![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 558-atom dihydrofolate reductase (DHFR) protein system and the 408
558-atom dihydrofolate reductase (DHFR) protein system and the 408![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 609-atom cellulose sugar polymer system, both from the Amber20 benchmark.69Fig. 5 shows excellent strong scaling up to 256 nodes on both benchmark systems when run on Frontier (AMD MI250X GPUs) and Perlmutter (NVIDIA A100 40 GB GPUs). Inference throughput on Perlmutter begins to saturate between 256 and 512 nodes for AOTI-compiled models (both with and without the custom kernel), while runs on Frontier continue to improve up to 512 nodes, where the AOTI compiled model using the custom kernel achieves over 200 timesteps per second. We observe a maximum throughput of 205 timesteps per second on the 24k atom DHFR system using 256 nodes of Perlmutter with AOTI and the custom kernel. The largest observed speedup over TorchScript is 10.9 times for DHFR on 64 nodes of Perlmutter. For node counts where TorchScript inference fits in memory and can be run, we observe that the combination of AOTI and the kernel achieves an average speedup of 6.6 times on Frontier and 6.9 times on Perlmutter.
609-atom cellulose sugar polymer system, both from the Amber20 benchmark.69Fig. 5 shows excellent strong scaling up to 256 nodes on both benchmark systems when run on Frontier (AMD MI250X GPUs) and Perlmutter (NVIDIA A100 40 GB GPUs). Inference throughput on Perlmutter begins to saturate between 256 and 512 nodes for AOTI-compiled models (both with and without the custom kernel), while runs on Frontier continue to improve up to 512 nodes, where the AOTI compiled model using the custom kernel achieves over 200 timesteps per second. We observe a maximum throughput of 205 timesteps per second on the 24k atom DHFR system using 256 nodes of Perlmutter with AOTI and the custom kernel. The largest observed speedup over TorchScript is 10.9 times for DHFR on 64 nodes of Perlmutter. For node counts where TorchScript inference fits in memory and can be run, we observe that the combination of AOTI and the kernel achieves an average speedup of 6.6 times on Frontier and 6.9 times on Perlmutter.
 | ||
Fig. 5 Strong scaling of the medium Allegro model on biomolecular systems. The molecular dynamics throughput of the Allegro model deployed using TorchScript, AOTI, or AOTI with the optimized tensor product kernel (AOTI + custom TP) on the 23![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 558-atom dihydrofolate reductase (DHFR) and 408 558-atom dihydrofolate reductase (DHFR) and 408![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 609-atom cellulose systems from the Amber20 benchmark69 is measured on a number of nodes ranging from 1 to 512 on Frontier (AMD MI250X) and Perlmutter (NVIDIA A100 (40 GB)). Note that there are twice as many logical GPU devices and corresponding MPI ranks on an MI250X node (8) than on an A100 node (4). No TorchScript result is shown for cellulose on A100 GPUs because TorchScript required more GPU memory than was available on these nodes. 609-atom cellulose systems from the Amber20 benchmark69 is measured on a number of nodes ranging from 1 to 512 on Frontier (AMD MI250X) and Perlmutter (NVIDIA A100 (40 GB)). Note that there are twice as many logical GPU devices and corresponding MPI ranks on an MI250X node (8) than on an A100 node (4). No TorchScript result is shown for cellulose on A100 GPUs because TorchScript required more GPU memory than was available on these nodes. | ||
Similar to Fig. 4, the combination of AOTI compilation and the custom tensor product kernel dramatically improve memory efficiency. For the larger cellulose system, the TorchScript-compiled model requires too much GPU memory for the 40 GB A100s and fails to run even when using 512 nodes. The AOTI and custom kernel approach, in contrast, runs successfully on as few as 16 nodes, which corresponds to approximately 6384 atoms per GPU.
4. Conclusion
We redesigned the NequIP framework to meet the increasing computational demands of MLIP training and inference. By integrating PyTorch 2.0 compiler technologies, distributed training, and optimized custom Triton kernels, the new infrastructure achieves substantial performance improvements for both training and inference. We trained three Allegro models on the SPICE 2 dataset and demonstrated how the new infrastructure can support training MLIPs on large datasets.The NequIP framework implements a number of techniques that could be applied to further accelerate the Allegro models presented here, and an in-depth study of their strengths and weaknesses is an important topic for future work. In particular, the framework supports per-edge-type cutoff radii that depend on the atomic types of both the central and the neighbor atoms, which we used in ref. 31. Because the computational cost of Allegro scales linearly with the total number of edges, tuning per-edge-type cutoff radii can have a significant impact on speed. The framework also supports the faster but less numerically accurate TensorFloat32 floating-point precision for intermediate computations, which we did not enable in this work. Additionally, we observed that the throughput gains from the custom TP kernel vary across hardware platforms, and improving the performance of our custom kernels on MI250X systems is an area of ongoing optimization.
Finally, the updated NequIP framework is an extensible foundation for the development of MLIP architectures and training strategies. The modular structure it enforces facilitates the development of extension packages like Allegro and ensures that they can easily leverage both the computational acceleration techniques described here and the broader set of tools, integrations, and infrastructure offered by the framework.
Importantly, the accelerations introduced in this work are not specific to Allegro and can be applied to any model implemented within the NequIP framework. With the exception of the custom TP kernel specific to the Allegro TP, all other accelerations, including the compilation and distributed training infrastructure, are fully general. The NequIP model, while not presented in this work for clarity of focus, also benefits from these performance advances. More broadly, other recently developed architectures27,70,71 can also take advantage of these optimizations, provided they comply with the framework's infrastructural and compilation interfaces. This design makes it possible for future MLIP architectures built on top of the framework to benefit from these performance improvements with minimal additional engineering effort.
More recently, these infrastructural advancements have enabled the development of NequIP and Allegro foundation potentials pretrained on OMat24,15 and fine-tuned on MPTrj13 and Alexandria,14 now publicly available at https://www.nequip.net/and highlighted on the Matbench Discovery benchmark.72 A detailed assessment of these models will be published elsewhere; we mention them here simply to underscore the scalability of the NequIP framework to large datasets beyond the SPICE 2 case study.
Author contributions
Chuin Wei Tan: conceptualizaton; software; visualization; writing – original draft preparation; writing – review & editing. Marc Descoteaux: conceptualizaton; data curation; formal analysis; software; visualization; writing – original draft preparation; writing – review & editing. Mit Kotak: conceptualizaton; software; writing – original draft preparation. Gabriel de Miranda Nascimento: data curation; software; validation; visualization. Seán Kavanagh: software; validation; writing – original draft preparation; writing – review & editing. Laura Zichi: software; validation. Menghang Wang: validation. Aadit Saluja: software. Yizhong R. Hu: software. Tess Smidt: funding acquisition; supervision. Anders Johansson: software; supervision; writing – original draft preparation. William C. Witt: conceptualizaton; data curation; software; visualization; supervision; writing – review & editing. Boris Kozinsky: conceptualizaton; funding acquisition; resources; supervision; writing – review & editing. Albert Musaelian: conceptualizaton; project administration; software; supervision; writing – original draft preparation; writing – review & editing.Conflicts of interest
There are no conflicts to declare.Data availability
All software and code used in this work are publicly accessible. This work used PyTorch version 2.6.0. The specific software versions used in this study, v0.7.0,
v0.7.0,  v0.4.0, and
v0.4.0, and  v0.7.0, are archived on Zenodo as a curated software release (DOI: https://doi.org/10.5281/zenodo.18211443). These correspond to the tagged releases available on GitHub at https://github.com/mir-group/nequip, https://github.com/mir-group/allegro, and https://github.com/mir-group/pair_nequip_allegro. Up-to-date development versions of the software remain available at these repositories. Scripts used for data processing, model training, benchmarking, and analysis are archived on Zenodo (DOI: https://doi.org/10.5281/zenodo.18193586) and are publicly available at https://github.com/mir-group/nequip-infra-paper-scripts. Details and links to the datasets used in this work. Supplementary information: model training and inference details, and additional experiments and benchmarks. See DOI: https://doi.org/10.1039/d5dd00423c.
v0.7.0, are archived on Zenodo as a curated software release (DOI: https://doi.org/10.5281/zenodo.18211443). These correspond to the tagged releases available on GitHub at https://github.com/mir-group/nequip, https://github.com/mir-group/allegro, and https://github.com/mir-group/pair_nequip_allegro. Up-to-date development versions of the software remain available at these repositories. Scripts used for data processing, model training, benchmarking, and analysis are archived on Zenodo (DOI: https://doi.org/10.5281/zenodo.18193586) and are publicly available at https://github.com/mir-group/nequip-infra-paper-scripts. Details and links to the datasets used in this work. Supplementary information: model training and inference details, and additional experiments and benchmarks. See DOI: https://doi.org/10.1039/d5dd00423c.
Acknowledgements
We are grateful to Angela Yi and Richard Zou for discussions regarding the PyTorch compilation workflow. We also thank Jaeyeon Won and Teodoro Collins for discussions regarding the custom kernel. This work was supported by the National Science Foundation through the Harvard University Materials Research Science and Engineering Center Grant No. DMR-2011754, the Department of Navy award N00014-20-1-2418 issued by the Office of Naval Research, and the Department of Energy Office of Basic Energy Sciences Award No. DE-SC0022199. This work is also supported by Robert Bosch LLC and the National Science Foundation under Grant No. DMR-2119351. MK was supported by the National Science Foundation under Cooperative Agreement PHY-2019786 (The NSF AI Institute for Artificial Intelligence and Fundamental Interactions), and the NSF Graduate Research Fellowship program under Grant No. DGE-1745302. SRK thanks the Harvard University Center for the Environment (HUCE) for funding a fellowship. MW was supported by National Science Foundation, Office of Advanced Cyberinfrastructure (OAC), under Award No. 2118201. AJ was supported by the Laboratory Directed Research and Development program at Sandia National Laboratories, a multimission laboratory managed and operated by National Technology and Engineering Solutions of Sandia, LLC, a wholly owned subsidiary of Honeywell International, Inc., for the U.S. Department of Energy's National Nuclear Security Administration under contract DE-NA-0003525. This paper describes objective technical results and analysis. Any subjective views or opinions that might be expressed in the paper do not necessarily represent the views of the U.S. Department of Energy or the United States Government. An award of computer time was provided by the INCITE program. This research used resources of the Oak Ridge Leadership Computing Facility, which is a DOE Office of Science User Facility supported under Contract DE-AC05-00OR22725. This research also used resources of the National Energy Research Scientific Computing Center (NERSC), a DOE Office of Science User Facility supported by the Office of Science of the U.S. Department of Energy under Contract No. DE-AC02-05CH11231 using NERSC awards BESERCAP0026720 and BESERCAP0032494. Additional computations were run on the FASRC Cannon cluster supported by the FAS Division of Science Research Computing Group at Harvard University.References
- J. Behler and M. Parrinello, Phys. Rev. Lett., 2007, 98, 146401 CrossRef PubMed.
- A. P. Bartók, M. C. Payne, R. Kondor and G. Csányi, Phys. Rev. Lett., 2010, 104, 136403 CrossRef PubMed.
- K. T. Schütt, S. S. Hessmann, N. W. Gebauer, J. Lederer and M. Gastegger, J. Chem. Phys., 2023, 158, 144801 CrossRef PubMed.
- J. Zeng, D. Zhang, D. Lu, P. Mo, Z. Li, Y. Chen, M. Rynik, L. Huang, Z. Li and S. Shi, et al. , J. Chem. Phys., 2023, 159, 054801 CrossRef CAS PubMed.
- Y. Lysogorskiy, C. v. d. Oord, A. Bochkarev, S. Menon, M. Rinaldi, T. Hammerschmidt, M. Mrovec, A. Thompson, G. Csányi and C. Ortner, et al. , npj Comput. Mater., 2021, 7, 97 CrossRef CAS.
- Z. Fan, Y. Wang, P. Ying, K. Song, J. Wang, Y. Wang, Z. Zeng, K. Xu, E. Lindgren and J. M. Rahm, et al. , J. Chem. Phys., 2022, 157, 114801 CrossRef CAS PubMed.
- W. C. Witt, C. van der Oord, E. Gelžinytė, T. Järvinen, A. Ross, J. P. Darby, C. H. Ho, W. J. Baldwin, M. Sachs and J. Kermode, et al. , J. Chem. Phys., 2023, 159, 164101 CrossRef CAS PubMed.
- P. T. Salzbrenner, S. H. Joo, L. J. Conway, P. I. Cooke, B. Zhu, M. P. Matraszek, W. C. Witt and C. J. Pickard, J. Chem. Phys., 2023, 159, 144801 CrossRef CAS PubMed.
- E. Podryabinkin, K. Garifullin, A. Shapeev and I. Novikov, J. Chem. Phys., 2023, 159, 084112 CrossRef CAS PubMed.
- R. P. Pelaez, G. Simeon, R. Galvelis, A. Mirarchi, P. Eastman, S. Doerr, P. Thölke, T. E. Markland and G. De Fabritiis, J. Chem. Theory Comput., 2024, 4076–4087 CrossRef CAS PubMed.
- J. Firoz, F. Pellegrini, M. Geiger, D. Hsu, J. A. Bilbrey, H.-Y. Chou, M. Stadler, M. Hoehnerbach, T. Wang and D. Lin, et al., HPDC ’25, July 20–23, 2025, Notre Dame, IN, USA, 2025, vol. 8, pp. 1–13 Search PubMed.
- P. Eastman, P. K. Behara, D. L. Dotson, R. Galvelis, J. E. Herr, J. T. Horton, Y. Mao, J. D. Chodera, B. P. Pritchard and Y. Wang, et al. , Sci. Data, 2023, 10, 11 CrossRef CAS PubMed.
- B. Deng, P. Zhong, K. Jun, J. Riebesell, K. Han, C. J. Bartel and G. Ceder, Nat. Mach. Intell., 2023, 5, 1031 CrossRef.
- J. Schmidt, T. F. Cerqueira, A. H. Romero, A. Loew, F. Jäger, H.-C. Wang, S. Botti and M. A. Marques, Mater. Today Phys., 2024, 48, 101560 CrossRef.
- L. Barroso-Luque, M. Shuaibi, X. Fu, B. M. Wood, M. Dzamba, M. Gao, A. Rizvi, C. L. Zitnick, and Z. W. Ulissi, arXiv, 2024, preprint, arXiv:2410.12771, DOI:10.48550/arXiv.2410.12771.
- A. D. Kaplan, R. Liu, J. Qi, T. W. Ko, B. Deng, J. Riebesell, G. Ceder, K. A. Persson, and S. P. Ong, arXiv, 2025, preprint, arXiv:2503.04070, DOI:10.48550/arXiv.2503.04070.
- D. S. Levine, M. Shuaibi, E. W. C. Spotte-Smith, M. G. Taylor, M. R. Hasyim, K. Michel, I. Batatia, G. Csányi, M. Dzamba, P. Eastman, et al., arXiv, 2025, preprint, arXiv:2505.08762, DOI:10.48550/arXiv.2505.08762.
- C. Chen and S. P. Ong, Nat. Comput. Sci., 2022, 2, 718 CrossRef PubMed.
- D. P. Kovács, J. H. Moore, N. J. Browning, I. Batatia, J. T. Horton, Y. Pu, V. Kapil, W. C. Witt, I.-B. Magdău, D. J. Cole and G. Csányi, J. Am. Chem. Soc., 2025, 147(21), 17598–17611 CrossRef PubMed.
- I. Batatia, P. Benner, Y. Chiang, A. M. Elena, D. P. Kovács, J. Riebesell, X. R. Advincula, M. Asta, M. Avaylon, W. J. Baldwin, et al., arXiv, 2023, preprint, arXiv:2401.00096, DOI:10.48550/arXiv.2401.00096.
- A. Merchant, S. Batzner, S. S. Schoenholz, M. Aykol, G. Cheon and E. D. Cubuk, Nature, 2023, 624, 80 CrossRef CAS PubMed.
- Y. Park, J. Kim, S. Hwang and S. Han, J. Chem. Theory Comput., 2024, 20(11), 4857–4868 CrossRef CAS PubMed.
- P. Eastman, B. P. Pritchard, J. D. Chodera and T. E. Markland, J. Chem. Theor. Comput., 2024, 20, 8583 CrossRef CAS PubMed.
- H. Yang, C. Hu, Y. Zhou, X. Liu, Y. Shi, J. Li, G. Li, Z. Chen, S. Chen, C. Zeni, et al., arXiv, 2024, preprint, arXiv:2405.04967, DOI:10.48550/arXiv.2405.04967.
- M. Neumann, J. Gin, B. Rhodes, S. Bennett, Z. Li, H. Choubisa, A. Hussey, and J. Godwin, arXiv, 2024, preprint, arXiv:2410.22570, DOI:10.48550/arXiv.2410.22570.
- B. Rhodes, S. Vandenhaute, V. Šimkus, J. Gin, J. Godwin, T. Duignan, and M. Neumann, arXiv, 2025, preprint, arXiv:2504.06231, DOI:10.48550/arXiv.2504.06231.
- X. Fu, B. M. Wood, L. Barroso-Luque, D. S. Levine, M. Gao, M. Dzamba, and C. L. Zitnick, arXiv, 2025, preprint, arXiv:2502.12147, DOI:10.48550/arXiv.2502.12147.
- S. R. Kavanagh, arXiv, 2024, preprint, arXiv:2412.19330, DOI:10.48550/arXiv.2412.19330.
- S. Batzner, A. Musaelian, L. Sun, M. Geiger, J. P. Mailoa, M. Kornbluth, N. Molinari, T. E. Smidt and B. Kozinsky, Nat. Commun., 2022, 13, 2453 CrossRef CAS PubMed.
- A. Musaelian, S. Batzner, A. Johansson, L. Sun, C. J. Owen, M. Kornbluth and B. Kozinsky, Nat. Commun., 2023, 14, 579 CrossRef CAS PubMed.
- B. Kozinsky, A. Musaelian, A. Johansson, and S. Batzner, in Proceedings of the International Conference for High Performance Computing, Networking, Storage and Analysis, 2023, pp. 1–12 Search PubMed.
- I. Batatia, D. P. Kovacs, G. Simm, C. Ortner and G. Csányi, Adv. Neural Inf. Process. Syst., 2022, 35, 11423 Search PubMed.
- A. Bochkarev, Y. Lysogorskiy and R. Drautz, Phys. Rev. X, 2024, 14, 021036 CAS.
- I. Batatia, S. Batzner, D. P. Kovács, A. Musaelian, G. N. Simm, R. Drautz, C. Ortner, B. Kozinsky and G. Csányi, Nat. Mach. Intell., 2025, 1, 56–67 CrossRef PubMed.
- A. Paszke, S. Gross, F. Massa, A. Lerer, J. Bradbury, G. Chanan, T. Killeen, Z. Lin, N. Gimelshein and L. Antiga, et al., Proceedings of the 33rd International Conference on Neural Information Processing Systems, 2019, vol. 721, pp. 8026–8037 Search PubMed.
- J. Ansel, E. Yang, H. He, N. Gimelshein, A. Jain, M. Voznesensky, B. Bao, P. Bell, D. Berard, E. Burovski, et al., in Proceedings of the 29th ACM International Conference on Architectural Support for Programming Languages and Operating Systems, vol. 2, 2024, pp. 929–947 Search PubMed.
- P. Tillet, H.-T. Kung, and D. Cox, in Proceedings of the 3rd ACM SIGPLAN International Workshop on Machine Learning and Programming Languages, 2019, pp. 10–19 Search PubMed.
- L. Dagum and R. Menon, IEEE Comput. Sci. Eng., 1998, 5, 46 CrossRef.
- Y. Hao, X. Zhao, B. Bao, D. Berard, W. Constable, A. Aziz, and X. Liu, arXiv, 2023, preprint, arXiv:2304.14226, DOI:10.48550/arXiv.2304.14226.
- F. Zhu, A. Nowaczynski, R. Li, J. Xin, Y. Song, M. Marcinkiewicz, S. B. Eryilmaz, J. Yang, and M. Andersch, in Proceedings of the 61st ACM/IEEE Design Automation Conference, 2024, pp. 1–6 Search PubMed.
- P. Kasimbeg, F. Schneider, R. Eschenhagen, J. Bae, C. S. Sastry, M. Saroufim, B. Feng, L. Wright, E. Z. Yang, Z. Nado, et al., arXiv, 2025, preprint, arXiv:2502.15015, DOI:10.48550/arXiv.2502.15015.
- X. Fu, Z. Wu, W. Wang, T. Xie, S. Keten, R. Gomez-Bombarelli, and T. Jaakkola, arXiv, 2022, preprint, arXiv:2210.07237, DOI:10.48550/arXiv.2210.07237.
- A. Loew, D. Sun, H.-C. Wang, S. Botti, and M. A. Marques, arXiv, 2024, preprint, arXiv:2412.16551, DOI:10.48550/arXiv.2412.16551.
- F. Bigi, M. Langer, and M. Ceriotti, arXiv, 2024, preprint, arXiv:2412.11569, DOI:10.48550/arXiv.2412.11569.
- J. Reed, Z. DeVito, H. He, A. Ussery and J. Ansel, Proceedings of Machine Learning and Systems, 2022, vol. 4, pp. 638–651 Search PubMed.
- S. Li, Y. Zhao, R. Varma, O. Salpekar, P. Noordhuis, T. Li, A. Paszke, J. Smith, B. Vaughan, P. Damania, et al., arXiv, 2020, preprint, arXiv:2006.15704, DOI:10.48550/arXiv.2006.15704.
- W. Falcon and The PyTorch Lightning Team, PyTorch Lightning, 2019 Search PubMed.
- A. P. Thompson, H. M. Aktulga, R. Berger, D. S. Bolintineanu, W. M. Brown, P. S. Crozier, P. J. in 't Veld, A. Kohlmeyer, S. G. Moore, T. D. Nguyen, R. Shan, M. J. Stevens, J. Tranchida, C. Trott and S. J. Plimpton, Comput. Phys. Commun., 2022, 271, 108171 CrossRef CAS.
- S. Gong, Y. Zhang, Z. Mu, Z. Pu, H. Wang, Z. Yu, M. Chen, T. Zheng, Z. Wang, L. Chen, et al., arXiv, 2024, preprint, arXiv:2404.07181, DOI:10.48550/arXiv.2404.07181.
- C. R. Trott, D. Lebrun-Grandié, D. Arndt, J. Ciesko, V. Dang, N. Ellingwood, R. Gayatri, E. Harvey, D. S. Hollman, D. Ibanez, N. Liber, J. Madsen, J. Miles, D. Poliakoff, A. Powell, S. Rajamanickam, M. Simberg, D. Sunderland, B. Turcksin and J. Wilke, IEEE Trans. Parallel Distr. Syst., 2022, 33, 805 Search PubMed.
- Z. DeVito, J. Ansel, W. Constable, M. Suo, A. Zhang, and K. Hazelwood, arXiv, 2021, preprint, arXiv:2104.00254, DOI:10.48550/arXiv.2104.00254.
- Z. Wang and W. Yan, arXiv, 2024, preprint, arXiv:2412.18271, DOI:10.48550/arXiv.2412.18271.
- A. H. Larsen, J. J. Mortensen, J. Blomqvist, I. E. Castelli, R. Christensen, M. Dułak, J. Friis, M. N. Groves, B. Hammer, C. Hargus, E. D. Hermes, P. C. Jennings, P. B. Jensen, J. Kermode, J. R. Kitchin, E. L. Kolsbjerg, J. Kubal, K. Kaasbjerg, S. Lysgaard, J. B. Maronsson, T. Maxson, T. Olsen, L. Pastewka, A. Peterson, C. Rostgaard, J. Schiøtz, O. Schütt, M. Strange, K. S. Thygesen, T. Vegge, L. Vilhelmsen, M. Walter, Z. Zeng and K. W. Jacobsen, J. Phys.: Condens. Matter, 2017, 29, 273002 CrossRef PubMed.
- O. A. Cohen, J. Riebesell, R. Goodall, A. Kolluru, S. Falletta, J. Krause, J. Colindres, G. Ceder and A. S. Gangan, AI Sci., 2025, 1(2), 025003 CrossRef.
- T. Dao, D. Fu, S. Ermon, A. Rudra and C. Ré, Adv. Neural Inf. Process. Syst., 2022, 35, 16344 Search PubMed.
- P.-L. Hsu, Y. Dai, V. Kothapalli, Q. Song, S. Tang, S. Zhu, S. Shimizu, S. Sahni, H. Ning, and Y. Chen, arXiv, 2024, preprint, arXiv:2410.10989, DOI:10.48550/arXiv.2410.10989.
- F. Zhu, M. Futrega, H. Bao, S. B. Eryilmaz, F. Kong, K. Duan, X. Zheng, N. Angel, M. Jouanneaux, M. Stadler, et al., in Proceedings of the 52nd International Conference on Parallel Processing, 2023, pp. 274–284 Search PubMed.
- V. Bharadwaj, A. Glover, A. Buluç, and J. Demmel, in 2025 Proceedings of the Conference on Applied and Computational Discrete Algorithms (ACDA), SIAM, 2025, pp. 32–46 Search PubMed.
- J. Won, C. Hong, C. Mendis, J. Emer and S. Amarasinghe, Proceedings of Machine Learning and Systems, 2023, vol. 5, pp. 666–679 Search PubMed.
- M. Kotak, Simplifying equivariant gpu kernels through tile-based programming, 2025, available: https://mitkotak.github.io/assets/pdf/sm_thesis_v2.pdf.
- A. Najibi and L. Goerigk, J. Chem. Theory Comput., 2018, 14, 5725 CrossRef CAS PubMed.
- N. Mardirossian and M. Head-Gordon, J. Chem. Phys., 2016, 144, 214110 CrossRef PubMed.
- S. Hattori and Q. Zhu, ACS Omega, 2024, 9, 36589 CAS.
- J. H. Moore, D. J. Cole, and G. Csanyi, arXiv, 2024, preprint, arXiv:2405.18171, DOI:10.48550/arXiv.2405.18171.
- S. Barnett and J. D. Chodera, GEN Biotechnol., 2024, 3, 119 CrossRef CAS.
- B. K. Rai, V. Sresht, Q. Yang, R. Unwalla, M. Tu, A. M. Mathiowetz and G. A. Bakken, J. Chem. Inf. Model., 2022, 62, 785 CrossRef CAS PubMed.
- R. Zubatyuk, J. S. Smith, B. T. Nebgen, S. Tretiak and O. Isayev, Nat. Commun., 2021, 12, 4870 CrossRef CAS PubMed.
- M. U. Maruf, S. Kim and Z. Ahmad, J. Phys. Chem. Lett., 2025, 16, 9078 CrossRef CAS PubMed.
- The AMBER Project, Amber20 benchmark suite, 2020, available: https://ambermd.org/Amber20_Benchmark_Suite.tar.gz.
- B. Cheng, npj Comput. Mater., 2024, 10, 157 CrossRef CAS.
- Z. Yang, X. Wang, Y. Li, Q. Lv, C. Y.-C. Chen and L. Shen, npj Comput. Mater., 2025, 11, 49 CrossRef CAS.
- J. Riebesell, R. E. Goodall, P. Benner, Y. Chiang, B. Deng, G. Ceder, M. Asta, A. A. Lee, A. Jain and K. A. Persson, Nat. Mach. Intell., 2025, 7, 836 CrossRef.
Footnote |
| † Co-first author. |
| This journal is © The Royal Society of Chemistry 2026 |