Open Access Article
Open Access ArticleA deep learning approach to searching property spaces of materials
Robert J. Appleton a,
Brian C. Barnesb,
Steven F. Sonc and
Alejandro Strachan*a
a,
Brian C. Barnesb,
Steven F. Sonc and
Alejandro Strachan*a
aSchool of Materials Engineering and Birck Nanotechnology Center, Purdue University, West Lafayette, Indiana 47907, USA. E-mail: strachan@purdue.edu
bU.S. Army Combat Capabilities Development Command Army Research Laboratory, Aberdeen Proving Ground, Maryland 21005, USA
cSchool of Mechanical Engineering, Purdue University, West Lafayette, Indiana 47907, USA
First published on 13th April 2026
Abstract
Melt processing of molecular crystals has several advantages over alternative routes for manufacturing materials such as pharmaceuticals, organic photovoltaics, and energetic materials. The experimental characterization of the materials properties required to assess melt processability (melting temperature, boiling temperature, decomposition temperature and vapor pressure) for the 1.3 million known molecular crystals is unfeasible; in fact, our survey of the research literature and open databases resulted in only 43 molecular materials with experimentally measured properties that satisfy a common criterion for melt-casting. We developed multi-task, graph-based neural network models that simultaneously predict these properties using a molecular graph as the only input. Screening databases of known molecules with our ML model resulted 2532 melt-castable candidates, with melting temperature between 343 K and 393 K, boiling and decomposition temperature greater than 453 K, and vapor pressure less than 0.0005 mmHg. Going beyond the space of known molecules, we apply our model with a generative approach to the CHNO chemical space, we discover 55![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 745 additional novel candidates with promising melt-castable characteristics. This three-orders-of-magnitude expansion highlights the power of coupling ML screening and generative design to accelerate materials discovery.
745 additional novel candidates with promising melt-castable characteristics. This three-orders-of-magnitude expansion highlights the power of coupling ML screening and generative design to accelerate materials discovery.
1. Introduction
Melt processing techniques, such as melt-casting and melt crystallization, are widely used for manufacturing various types of materials, including metals,1 polymers,2 and molecular crystals.3 Advantages of these methods include near-final shape production that minimizes the need for extensive machining that, in turn, leads to a reduction of material waste and processing time.4,5 In addition, controlled solidification can lead to uniform microstructures with fewer defects such as porosity, inclusions, and cracks, enhancing mechanical properties.6 Finally, melt-casting enables the production of components with intricate geometries that are difficult or impossible to achieve with traditional subtractive manufacturing methods making it a desirable method for biomedical implants,7 controlled drug delivery8,9 and solid-state batteries.10 Specific melt-cast applications for organic materials include organic glasses for high-efficiency scintillators,11 organic thin-film transistors,12 small molecule organic photovoltaics13 and energetic materials.14–16 The popular over-the-counter drugs ibuprofen and acetaminophen are prime examples where melt-casting has led to enhanced pharmaceutical characteristics, enabling faster, solvent-free production while also improving drug delivery, taste-masking, and controlled release.17–21 Unfortunately, a small fraction of molecular materials are melt-castable due to the multiple constraints on underlying properties. Melt-castability is largely dependent on the thermal stability of the molten liquid. The melting temperature should be moderately low to reduce energy costs and prevent damaging other components, but it cannot be too low to prevent unwanted melting during transport, storage, or other stages of the manufacturing and transportation process. The range of melting temperatures for melt-casting can vary across applications and for this work, we use the common range of 343 K to 393 K to enable energy-efficient melting through steam heating.14,15,22 We note that this melting temperature criteria would not include the previously mentioned acetaminophen due to its higher melting temperature (441 K).23 The stability of the liquid phase must be ensured by high decomposition and boiling temperatures to prevent generating toxic fumes and reduce material loss. To ensure the thermal stability of the liquid phase, Benz et al.14 proposed a minimum stability criterion of 453 K. Surveying experimental data for melting temperature,24,25 boiling temperature,24 and decomposition temperature,26,27 we identified 329 known CHNOFCl molecules that satisfy the thermal stability criteria for melt-casting. However, the vapor pressure of the material should also be low and using a generous filter of <0.0005 mmHg reduces the list to only 43 CHNOFCl molecules with experimental measured properties that meet both the thermal stability and vapor pressure criteria. This estimate represents a lower bound as it was obtained based on available data in common public databases discussed below. We have listed the resulting known melt-castable materials from this survey in the SI. In addition, this number does not include composite formulations designed to improve or enable melt-processing.Gaps in available experimental measurements for known molecules can be bridged with machine learning (ML), and such data would be very valuable to guide experimental studies. The development of quantitative structure–property relationship (QSPR) models has been a cornerstone of chemistry research since the 1940s28 and has also played a role in other areas of the physical sciences.29–42 With the emergence of ML, QSPR modeling has expanded in scope and capability, enabling more accurate and data-driven predictions across a wider range of scientific domains. These models were originally developed using hand-crafted descriptors43,44 that attempted to capture important chemical and structural properties of a molecule that govern the corresponding property of interest.45–48 Modern deep learning approaches have implemented graph-based methods such as graph convolutional networks49–51 (GCNs) and message-passing neural networks52–57 (MPNNs) that can achieve higher accuracy for property prediction than their QSPR predecessors. In this work, we trained a directed-message passing neural network (D-MPNN)52,53 to predict fundamental properties for determining melt-castability of molecular crystals from the corresponding gas-phase molecular graph. Using our ML model, we screened molecules based on thermal stability and vapor pressure and identified 2532 known molecules predicted to be melt-castable.
Beyond screening known molecules, we utilized a generative approach to explore property spaces and discover novel melt-castable materials. Deep generative methods for inverse design in molecular and materials science have been widely studied over the last decade, including variational autoencoders (VAEs),58–68 generative adversarial networks (GANs),69–71 and diffusion models.72–75 A key challenge in molecular and materials discovery is generating candidates that are both synthesizable and processable. In metallic alloys, computational phase diagram (CALPHAD) approaches, often parameterized using large datasets of first-principles formation energies41,76 and convex hulls77–79 calculations, are routinely used to identify thermodynamically stable structures with accessible processing conditions.80 In contrast, melt-castability of molecular crystals depends on multiple thermophysical properties, motivating the deep-learning-based, multi-property screening and generative strategy adopted in this work.
Deep generative methods have proven successful at generating novel structures with ideal properties but require large datasets and expensive training. Metaheuristic optimization algorithms, such as genetic algorithms (GAs), have a longer history in molecular design.81–83 These methods can operate on the molecular graph or string representations such as simplified molecular-input line-entry system (SMILES)84,85 and typically require evolutionary operations with expert chemical rules to ensure generated molecules are valid.86–89 Self-referencing embedded strings (SELFIES),90 a recently-proposed replacement for SMILES, is a robust string representation where every SELFIES string corresponds to a valid molecule. This representation led to the development of the STONED91 method for efficient and rule-free mutation and crossover operations in chemical space. Shortly after, JANUS-GA92 was introduced, it uses these genetic operators to enable inverse molecular design that is competitive with deep generative methods without the need for extensive training. In this work we extend JANUS-GA to incorporate Pareto-awareness and apply it to the discovery of novel melt-castable materials. We generate 55![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 940 molecules predicted to be melt-castable, 55
940 molecules predicted to be melt-castable, 55![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 745 of which are not reported in the PubChem93 or CAS SciFinder94 databases and are likely new.
745 of which are not reported in the PubChem93 or CAS SciFinder94 databases and are likely new.
2. Methods
2.1 Data
We compiled a large dataset of experimental melting temperatures (Tm), boiling temperatures (Tb), decomposition temperatures (Td) and vapor pressures (Pv) for known CHNOFCl materials. Data sources include the PHYSPROP database,24 the Jean-Claude Bradley Open Melting Point Dataset (JB OMPD),25 and decomposition temperature data published in open literature,26,27 see details in Table 1.For all molecules, SMILES strings were canonicalized using RDKit.95 We removed cocrystals/mixtures and molecules with radicals. Duplicates were identified, and for multiple measurements of the same property, the median was used for training. We use the log10 of the vapor pressures to account for the fact that the values span several orders of magnitude (10−10 to 105), this is consistent with previous works.29,96
2.2 Property prediction model design
The multi-task (MT) ML model for property prediction was developed using the chemprop framework,52,53 an implementation of a directed message-passing neural network (D-MPNN) for predicting properties from the molecular graph. To evaluate the model performance, we use a 5-fold cross-validation scheme. Specifically, we trained five separate models (one for each fold) and assessed the performance of each model on a corresponding test set that was not used in its training. Upon deployment, the models are used as an ensemble where the prediction is taken as the mean of the five models, and the uncertainty is the standard deviation. The values for each property are scaled using a standard scaler before training to address the fact that the properties have different ranges of magnitude. Hyperparameters were optimized minimally by adjusting the size of the network and other training details for the D-MPNN are provided in Table S1 of the SI. Similar models built with this framework have proven successful for these properties in previous works.26,57,962.3 Genetic algorithm for molecule generation
The generative approach used in this work is built on the JANUS-GA framework.92 JANUS-GA is a GA designed for inverse molecular design, employing a parallel-tempering-inspired97,98 framework with two distinct populations: one for global exploration and another for local exploitation. We describe the algorithm briefly here and refer the readers to the original publication92 and the SI for additional details. A fitness function is used to evaluate the members of both exploration and exploitation populations and the 5 top ranked individuals in each population are shared. The explorative population is subject to both mutation and crossover operations. This step generates an “overflow of potential children” and uses a deep neural network (DNN) to down sample molecules to maintain a fixed population size. The exploitative population refines promising molecular candidates using mutations based primarily on molecular similarity. JANUS-GA leverages the SELFIES molecular representation and the STONED algorithm to generate new molecules,91 bypassing the need for manually defined structure-validation rules. We extended the JANUS-GA for dual-property optimization using both the change in area under the Pareto front and their combination into a single objective function. We study different combinations of Pareto front and single objective function as the fitness function and for overflow down-sampling, both using a pre-trained MT D-MPNN. The implemented Pareto-aware fitness evaluation is described in Algorithm 1 below.Using these modifications, we can push the 2D Pareto front equally on all fronts or along a preferred region by combining the two variables into a single objective function. This was motivated by our observation in preliminary work where Pareto optimization alone can lead to trivial solutions, Fig. S2 of the SI. Also, we implemented functions for collecting ensemble predictions from our MT D-MPNN models. The properties predicted by the D-MPNN are used for identifying the Pareto front and computing fitness functions for new molecules but can also be used by the filters. Predictions come from the five different models trained on each of the 5-folds during cross validation. We record the mean and standard deviation of each property across these models. Previous work found that using a DNN to down sample the explorative population leads to populations with higher median fitness.92 In this work, the DNN is replaced with an objective function using actual predictions from our ML models. In the original implementation of JANUS-GA,92 the DNN serves as a surrogate for the fitness function, aiming to reduce the number of potentially expensive fitness evaluations by leveraging model predictions. In contrast, our implementation uses the same ML model for both down-sampling and fitness evaluation, resulting in no computational savings since the evaluation cost remains unchanged. The source code for the modified JANUS-GA is available at https://github.com/R-applet/JANUS and a summary of the algorithm parameters used in this work is provided in Table S2 of the SI.
3. Results
3.1 Property model performance
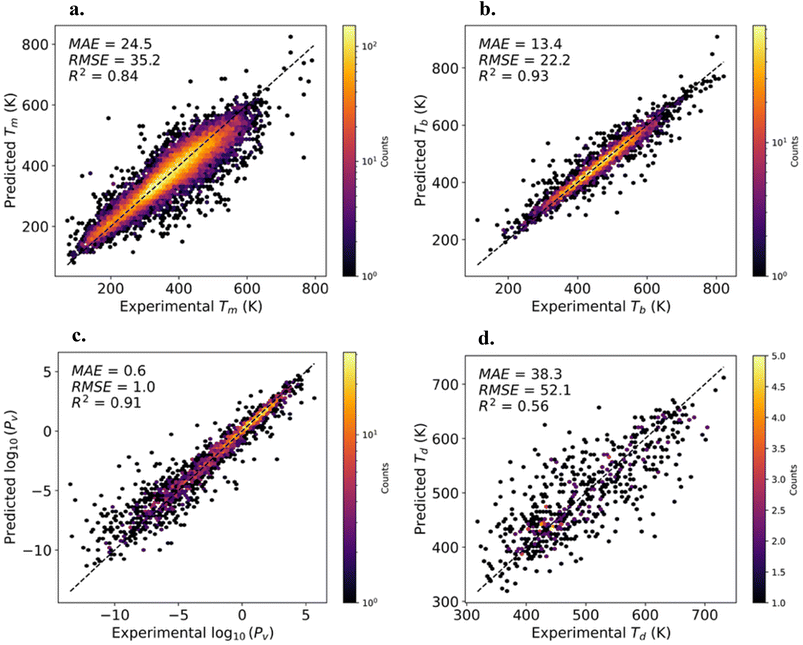
Fig. 1 shows the MT model predictions compared to the experimental values for the testing data of each property. We find that the model predictions for boiling temperatures (Tb) and vapor pressures (Pv) are in exceptional agreement with experiment with R2 > 0.9. The model predictions for melting temperatures (Tm) are slightly less accurate with R2 = 0.84 with MAE and RMSE values comparable to previous works that looked at similar datasets for slightly different chemical spaces.26,99 We note that the performance of our model on decomposition temperatures (Td) is poor with R2 = 0.56. The MAE and RMSE on Td are consistent with previous reports on this dataset.26 The limitation of our model reflects the data scarcity and for experimental Td data. In our usage of the model, we are looking to maximize the decomposition temperature, not try to predict a specific value, therefore the model predictions are strong indicators for thermal stability. For these reasons, we believe our model can be used to estimate both thermal stability and volatility. The validation learning curves for each fold are shown in Fig. S1 of the SI. Using this model to make predictions on the materials within the experimental data used to develop the model, we are able to fill the gaps for the measured properties. Screening this completed dataset, we identify 2532 known molecules predicted to be melt-castable. The list of these materials and their associated predicted properties are provided in the SI and experimental efforts to validate these predictions would be highly valuable.3.2 Generation of molecules with high melt casting potential
We use the modified JANUS-GA92 to explore property spaces of CHNO molecules to generate novel molecules with ideal properties for melt-casting. Specifically, we sought to find new molecules that maximize the liquid stability range, minimize the distance from the ideal melting temperature, and minimize the volatility. The liquid stability range (ΔTliq), is the temperature interval between the predicted melting temperature and whichever is lower, the predicted decomposition temperature or the predicted boiling temperature:| ΔTliq = min(Td,Tb) − Tm. | (1) |
The distance from the ideal melting temperature (ΔTim) is the absolute difference between the predicted melting temperature and the provided ideal melting temperature (taken as 368 K, which is the midpoint between 343 K and 393 K):
| ΔTim = |Tm − 368| | (2) |
To discover melt castable molecules, we explored two approaches: (i) in the first we consider the Pareto front formed by ΔTliq and ΔTim, (ii) the second approach considers vapor pressure and ΔTliq and limits the search to desirable values of ΔTim.
| O = ΔTliq − 3ΔTim. | (3) |
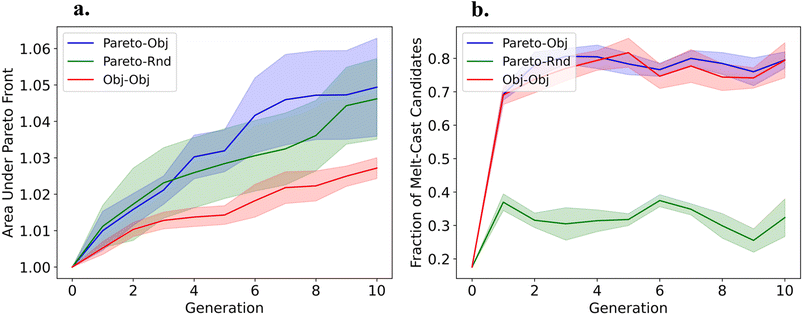
We evolved the population for 10 generations using a population size of 50 for both the exploitative and explorative populations. Each GA experiment was repeated 5 times to collect statistics. More details about the parameters of the GA are summarized in Table S2 and the results for the sampling experiments are shown in Section S4.1 of the SI. In all cases, we discovered molecules that are highly thermally stable and push the ΔTliq and ΔTim Pareto front. To track the performance of the GA we plot the area under the Pareto front from each generation (the area is normalized such that the area in generation 0 is 1), see Fig. 2(a). We find that the Pareto–Rnd GA pushes on a wide area of the Pareto front, while the Pareto–Obj and Obj–Obj GAs are highly localized with ΔTim less than ∼25 K. The advantage of using the objective function for down-sampling is that we generate more molecules that fall within the melting temperature range for melt-casting, as depicted by the fraction of melt-cast candidates per generation in Fig. 2(b). Comparing the Pareto–Obj GA to the Obj–Obj GA we find that the Pareto–Obj GA results in a more converged Pareto front.
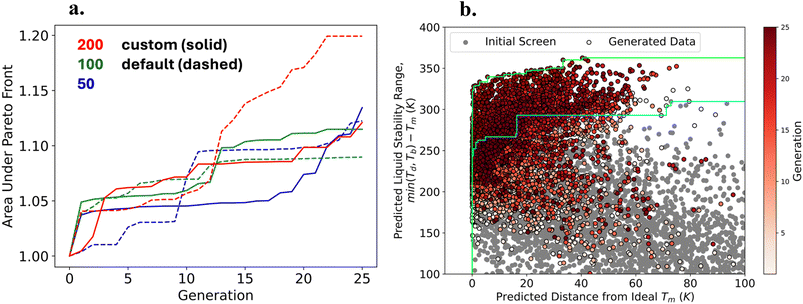
The results from the sampling experiments showed that the Pareto fronts generated were not converged and that running for more generations and larger generation sizes would continue to drive the Pareto front. On the other hand, there are hyperparameters of the GA that control the number of unique generated molecules and consequently the convergence and time for the GA to complete. We tested different hyperparameters of the JANUS-GA and evaluated how they influenced the evolution of the population. Specifically, we tested 3 different generation sizes (50, 100, and 200), using 5 random crossovers in the exploration population (default is 1), and 10% of the generation size as the number of mutations and number of random samples in the exploitation population (default is 400 for each). For each experiment, we tracked the number of unique molecules evaluated, unique molecules sampled in the population, and melt-cast candidates generated. We also report the wall time of each experiment. The results comparing the custom hyperparameters to the defaults are recorded in Table S3. Furthermore, we observed the distribution of each experiment and displayed them in Fig. S5. The convergence by comparing the evolution of the area under the Pareto front across 25 generations displayed in Fig. 3(a) and the distribution of ΔTliq and ΔTim for the generated molecules from the most converged GA is shown in Fig. 3(b). For this set of experiments, the GA with a generation size of 200 and the default hyperparameters (1 random crossover in the exploration population and 400 mutations and random samples in the exploitation population) resulted in having the largest area under the Pareto front, see Fig. 3(a).
Observing the distributions of ΔTliq and ΔTim across the hyperparameter experiments (Fig. S5), it is clear that using more mutations/random samples for the exploitative population leads to more points concentrated at the Pareto front. However, the consequence of using more mutations/random samples is that it generates a larger overflow of molecules and thus many more evaluations and a longer wall time, as shown in Table S3. We also note that even though more unique molecules were evaluated, we found fewer unique molecules were used in the population and typically less melt-castable candidates were discovered (with exemption of the runs with a population size of 100). We attribute this to the fact that more molecules are near the Pareto front and thus have high fitness leading to more individuals that are kept in the population and not replaced. In all cases, we are able to drive the Pareto front successfully and discover a large number of melt-castable candidates.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pv), we
Pv), we
O = ΔTliq − 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log10 log10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pv Pv
| (4) |
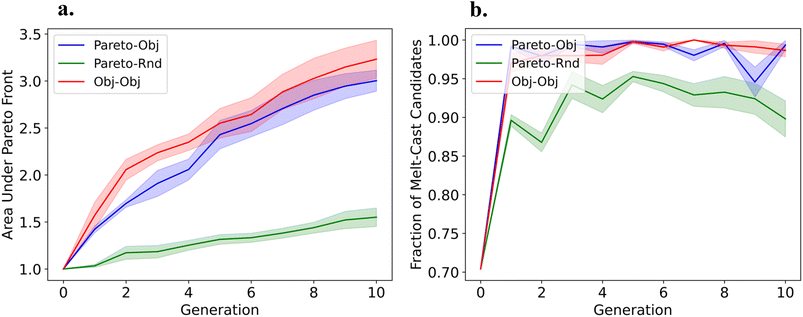
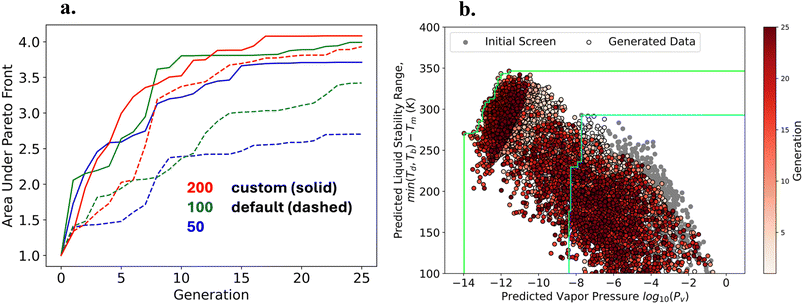
The results for the sampling experiments for this case are shown in Section 4.2 of the SI. In all cases, we can discover molecules with high thermal stability and push the Pareto front between ΔTliq and log10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pv. In these experiments, we find the Pareto–Obj and Obj–Obj GAs push the Pareto front much further than the Pareto–Rnd GA, see Fig. 4(a). Furthermore, the results for Pareto–Obj and Obj–Obj GAs are very similar for this set of experiments. Again, we find the Pareto–Obj and Obj–Obj GAs generate a larger fraction of melt-cast candidates per generation compared to Pareto–Rnd, see Fig. 4(b). A clear inverse relationship is observed between ΔTliq and log10
Pv. In these experiments, we find the Pareto–Obj and Obj–Obj GAs push the Pareto front much further than the Pareto–Rnd GA, see Fig. 4(a). Furthermore, the results for Pareto–Obj and Obj–Obj GAs are very similar for this set of experiments. Again, we find the Pareto–Obj and Obj–Obj GAs generate a larger fraction of melt-cast candidates per generation compared to Pareto–Rnd, see Fig. 4(b). A clear inverse relationship is observed between ΔTliq and log10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pv, where materials with wider liquid stability ranges generally exhibit lower vapor pressures, consistent with thermodynamic expectations from the Clausius–Clapeyron relation.100 The strong underlying correlations between ΔTliq and log10
Pv, where materials with wider liquid stability ranges generally exhibit lower vapor pressures, consistent with thermodynamic expectations from the Clausius–Clapeyron relation.100 The strong underlying correlations between ΔTliq and log10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pv makes this a much simpler optimization problem than optimizing ΔTliq and ΔTim. This explains why the difference between a Pareto–Obj driven GA and Obj–Obj driven GA is much less noticeable for the ΔTliq and log10
Pv makes this a much simpler optimization problem than optimizing ΔTliq and ΔTim. This explains why the difference between a Pareto–Obj driven GA and Obj–Obj driven GA is much less noticeable for the ΔTliq and log10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pv Pareto front. Furthermore, the Pareto–Rnd driven GA struggles to push the ΔTliq and log10
Pv Pareto front. Furthermore, the Pareto–Rnd driven GA struggles to push the ΔTliq and log10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pv Pareto front because it is not taking advantage of the strong correlations.
Pv Pareto front because it is not taking advantage of the strong correlations.
The results from the sampling experiments showed that the Pareto fronts generated were not converged and that running for more generations and larger generation sizes would continue to drive the Pareto front.
Similar to the previous set of experiments, we studied the hyperparameters and their effects on the evolution of the Pareto front. The results comparing the custom hyperparameters to the defaults are recorded in Table S4. Furthermore, we observed the distribution of each experiment and displayed them in Fig. S6. The convergence by comparing the evolution of the area under the Pareto front across 25 generations displayed in Fig. 5(a) and the distribution of ΔTliq and log10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pv for the generated molecules from the most converged GA is shown in Fig. 5(b). For this set of experiments, the GA with a generation size of 200 and the custom hyperparameters (5 random crossovers in the exploration population and 20 mutations and random samples in the exploitation population) resulted in having the largest area under the Pareto front, see Fig. 5(a).
Pv for the generated molecules from the most converged GA is shown in Fig. 5(b). For this set of experiments, the GA with a generation size of 200 and the custom hyperparameters (5 random crossovers in the exploration population and 20 mutations and random samples in the exploitation population) resulted in having the largest area under the Pareto front, see Fig. 5(a).
Observing the distributions of ΔTliq and log10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pv across the hyperparameter experiments (Fig. S6), again we find that using more mutations/random samples for the exploitative population leads to more points concentrated at the Pareto front. Again, this corresponds to a larger overflow of molecules, more evaluations, and a longer wall time, as shown in Table S4. Consistent with the previous set of experiments, despite more unique molecules being evaluated, we found less unique molecules were used in the population and less melt-castable candidates were discovered. In all cases, we are able to drive the Pareto front successfully and discover a large number of melt-castable candidates.
Pv across the hyperparameter experiments (Fig. S6), again we find that using more mutations/random samples for the exploitative population leads to more points concentrated at the Pareto front. Again, this corresponds to a larger overflow of molecules, more evaluations, and a longer wall time, as shown in Table S4. Consistent with the previous set of experiments, despite more unique molecules being evaluated, we found less unique molecules were used in the population and less melt-castable candidates were discovered. In all cases, we are able to drive the Pareto front successfully and discover a large number of melt-castable candidates.
4. Discussion
Across the populations in all simulations, we generated 62![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 082 unique molecules and 55
082 unique molecules and 55![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 940 of these have predicted thermal stability and volatility to be considered a potential melt-castable material. From the 55
940 of these have predicted thermal stability and volatility to be considered a potential melt-castable material. From the 55![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 940 molecules, 195 were found to have been synthesized by searching PubChemPy,101 CIRpy102 and SciFinder.94 We emphasize that these 195 molecules were not in the training set or initial population. These molecules confirm that our generative approach can discover real, synthesizable, molecules. Using SciFinder,94 we obtained experimental melting temperatures for 58 of the 195 rediscovered molecules and plotted them against the model predictions in Fig. S7 of the SI to provide further validation of the model accuracy on unseen molecules. We find that the vast majority (51/58) of predictions fall within the model's test set RMSE (see Fig. 1). Importantly, 61% of these molecules had experimental melting temperatures within the range to be melted via steam heating. This is further evidence that the model predictions are within the expected error.
940 molecules, 195 were found to have been synthesized by searching PubChemPy,101 CIRpy102 and SciFinder.94 We emphasize that these 195 molecules were not in the training set or initial population. These molecules confirm that our generative approach can discover real, synthesizable, molecules. Using SciFinder,94 we obtained experimental melting temperatures for 58 of the 195 rediscovered molecules and plotted them against the model predictions in Fig. S7 of the SI to provide further validation of the model accuracy on unseen molecules. We find that the vast majority (51/58) of predictions fall within the model's test set RMSE (see Fig. 1). Importantly, 61% of these molecules had experimental melting temperatures within the range to be melted via steam heating. This is further evidence that the model predictions are within the expected error.
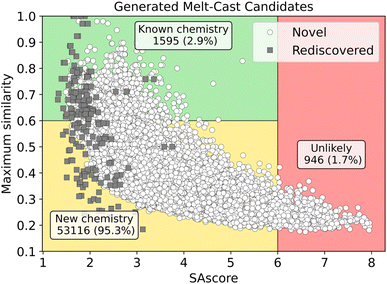
The remaining 55![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 745 molecules have not been previously reported, based on our searches using PubChemPy,101 CIRpy102 and SciFinder.94 To characterize the molecules generated, we computed the maximum Tanimoto similarity103 of each new molecule to an extensive set of known molecules93,104 and also computed the synthetic accessibility score (SAscore).105 The SAscores can be interpreted as 1 being “easy to make” and 10 being “very difficult to make” with the authors suggesting that molecules with SAscores greater than ∼6 are difficult to synthesize.105 Fig. 6 we shows these values for the novel melt-cast molecules we generated.
745 molecules have not been previously reported, based on our searches using PubChemPy,101 CIRpy102 and SciFinder.94 To characterize the molecules generated, we computed the maximum Tanimoto similarity103 of each new molecule to an extensive set of known molecules93,104 and also computed the synthetic accessibility score (SAscore).105 The SAscores can be interpreted as 1 being “easy to make” and 10 being “very difficult to make” with the authors suggesting that molecules with SAscores greater than ∼6 are difficult to synthesize.105 Fig. 6 we shows these values for the novel melt-cast molecules we generated.
Molecules with high similarity and low SAscore are classified as high priority candidates, as they are likely synthetically accessible and may follow similar synthetic routes to known compounds. Molecules with low similarity and low SAscore are considered low priority because they are still likely synthetically accessible but may require additional effort to determine viable synthesis pathways. Molecules with SAscore values above 6 are labeled as difficult due to their synthetic complexity and are unlikely to be pursued further. We provide further comparisons of the distributions of maximum similarity and SAscore for the novel generated melt-cast candidates and the rediscovered molecules in Fig. S8 of the SI. Shown in Fig. S8(a), the novel generated molecules display much lower maximum similarity scores with a mean less than 0.4, which has been used in prior works to indicate the novelty of generated molecules.92 This suggests that many of the generated melt-cast candidates are indeed novel with strong chemical distinction from known molecules. Shown in Fig. S8(b), the SAscores of the novel generated molecules are shifted higher (mean ∼ 3.5) than the rediscovered (mean ∼ 2) but we still find that 98.3% are below 6 indicating the vast majority are likely to be synthetically accessible.
We note that recent work using the JANUS-GA to explore molecules with high crystal heat of formation (ΔHf,s) and crystalline density (ρ), found that generated molecules had a distribution of SAscores shifted much higher (mean ∼ 6) than what we observe in Fig. 6 and S8.106 We speculate that because our optimization was focused on generating thermally stable molecules this may have correspondingly resulted in molecules with lower SAscore.
5. Conclusion
We developed a multi-task D-MPNN to predict fundamental properties (melting temperature, boiling temperature, decomposition temperature and vapor pressure) from the molecular graph. We deploy this model to screen known molecules and fill the gaps in the experimental data where we identify 2532 melt-castable candidates based on thermal stability and volatility. We extend an existing genetic algorithm for dual-property optimization by implementing modifications for computing the Pareto front between two properties and use a distance metric as the fitness function. Deploying our model within this generative framework we discovered 55![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 745 novel melt-castable candidates, the majority of which display SAscores that suggest they can be synthetically accessible. This workflow can be readily adapted to other material properties and optimization problems. Future efforts should investigate Pareto optimization in higher-dimensional property spaces (i.e., for n > 2) to extend beyond dual-objective fronts.
745 novel melt-castable candidates, the majority of which display SAscores that suggest they can be synthetically accessible. This workflow can be readily adapted to other material properties and optimization problems. Future efforts should investigate Pareto optimization in higher-dimensional property spaces (i.e., for n > 2) to extend beyond dual-objective fronts.
Author contributions
R. J. A.: conceptualization, data curation, formal analysis, writing – original draft. B. C. B.: conceptualization, funding acquisition, resources, writing – reviewing and editing. S. F. S.: supervision, funding acquisition, writing – reviewing and editing. A. S.: conceptualization, methodology, funding acquisition, resources, writing – original draft.Conflicts of interest
There are no conflicts to declare.Data availability
The compiled dataset of experimental measurements, model development code and scripts for running the genetic algorithm are available at GitHub https://github.itap.purdue.edu/StrachanGroup/MeltCastExplore with DOI; https://doi.org/10.5281/zenodo.18474595. The source code for the modified JANUS-GA is available at GitHub https://github.com/R-applet/JANUS with DOI; https://doi.org/10.5281/zenodo.18474741. The resulting 43 melt-cast materials from surveying experimental data, the 2532 ML screened melt-cast materials, and the novel generated melt-cast candidates with the corresponding predicted properties are provided along the manuscript as supplementary information (SI). A graphical user interface allowing users to make predictions and visualize the data is freely available at nanoHUB at https://nanohub.org/tools/mcexplorer with DOI; https://doi.org/10.21981/TTDN-JB78.Supplementary information is available. See DOI: https://doi.org/10.1039/d5dd00422e.
Acknowledgements
The authors thank Betsy M. Rice for useful discussion during the preparation of this manuscript. This research was sponsored in part by the Army Research Laboratory and was accomplished under Cooperative Agreement No. W911NF-20-2-0189. This research was sponsored in part by the Army Research Office and was accomplished under Cooperative Agreement No. W911NF-22-2-0170. This work was supported in part by a grant of computer time from the DoD High Performance Computing Modernization Program at the DEVCOM Army Research Laboratory.References
- J. Campbell, Casting, in Complete Casting Handbook, Elsevier, 2015, pp. 821–882, DOI:10.1016/B978-0-444-63509-9.00016-9.
- Z. Tadmor and C. G. Gogos, Principles of Polymer Processing, John Wiley & Sons, Inc., 2006 Search PubMed.
- H. Zhou, J. Sabino, Y. Yang, M. D. Ward, A. G. Shtukenberg and B. Kahr, Tailor-Made Additives for Melt-Grown Molecular Crystals: Why or Why Not?, Annu. Rev. Mater. Res., 2023, 53, 143–164 CrossRef CAS.
- J. T. Black and R. A. Kohser, DeGarmo's Materials and Processes in Manufacturing, Wiley, 2019 Search PubMed.
- T. A. Osswald and G. Menges, Part 2: Influence of Processing on Properties: Introduction to Processing, in Material Science of Polymers for Engineers, Hanser, 2012, pp. 161–262, DOI:10.3139/9781569905241.
- ASM Handbook, Volume 15: Casting, ed. S. Viswanathan, D. Apelian, R. J. Donahue, B. DasGupta, M. Gywn, J. L. Jorstad, R. W. Monroe, M. Sahoo, T. E. Prucha and D. Twarog, ASM International, 2008, vol. 15 Search PubMed.
- J. Hu and L. Tan, Polyurethane Composites and Nanocomposites for Biomedical Applications, in Polyurethane Polymers: Composites and Nanocomposites, Elsevier Inc., 2017, pp. 477–498, DOI:10.1016/B978-0-12-804065-2.00016-4.
- M. Y. Krasko, A. Shikanov, A. Ezra and A. J. Domb, Poly(ester anhydride)s prepared by the insertion of ricinoleic acid into poly(sebacic acid), J. Polym. Sci., Part A:Polym. Chem., 2003, 41, 1059–1069 CrossRef CAS.
- A. Basu and A. J. Domb, Recent Advances in Polyanhydride Based Biomaterials, Adv. Mater., 2018, 30, 1706815 CrossRef.
- X. Fu, T. Wang, W. Shen, M. Jiang, Y. Wang, Q. Dai, D. Wang, Z. Qiu, Y. Zhang, K. Deng, Q. Zeng, N. Zhao, X. Guo, Z. Liu, J. Liu and Z. Peng, A High-Performance Carbonate-Free Lithium|Garnet Interface Enabled by a Trace Amount of Sodium, Adv. Mater., 2020, 32, 2000575 CrossRef CAS.
- J. S. Carlson and P. L. Feng, Melt-cast organic glasses as high-efficiency fast neutron scintillators, Nucl. Instrum. Methods Phys. Res., Sect. A, 2016, 832, 152–157 CrossRef CAS.
- S. Schmid, A. K. Kast, R. R. Schröder, U. H. F. Bunz and C. Melzer, Improved thin-film transistor performance through a melt of poly(para-phenyleneethynylene), Macromol. Rapid Commun., 2014, 35, 1770–1775 CrossRef CAS.
- A. Rahmanudin, L. Yao, X. A. Jeanbourquin, Y. Liu, A. Sekar, E. Ripaud and K. Sivula, Melt-processing of small molecule organic photovoltaics: via bulk heterojunction compatibilization, Green Chem., 2018, 20, 2218–2224 RSC.
- M. Benz, A. Delage, T. M. Klapötke, M. Kofen and J. Stierstorfer, A rocky road toward a suitable TNT replacement – a closer look at three promising azoles, Propellants, Explos., Pyrotech., 2023, 48, e202300042 CrossRef CAS.
- P. Ravi, D. M. Badgujar, G. M. Gore, S. P. Tewari and A. K. Sikder, Review on melt cast explosives, Propellants, Explos., Pyrotech., 2011, 36, 393–403 CrossRef CAS.
- E. Holbrook, M. P. Kroonblawd, B. W. Hamilton, H. K. Springer and A. Strachan, Modeling Framework to Predict Melting Dynamics at Microstructural Defects in TNT-HMX High Explosive Composites, J. Phys. Chem. C, 2025, 129, 14111–14129 CrossRef CAS.
- S. P. Forster and D. B. Lebo, Continuous melt granulation for taste-masking of ibuprofen, Pharmaceutics, 2021, 13, 863 CrossRef CAS.
- Z. Yang, Y. Hu, G. Tang, M. Dong, Q. Liu and X. Lin, Development of ibuprofen dry suspensions by hot melt extrusion: Characterization, physical stability and pharmacokinetic studies, J. Drug Delivery Sci. Technol., 2019, 54, 101313 CrossRef CAS.
- M. Yang, P. Wang, C. Y. Huang, M. S. Ku, H. Liu and C. Gogos, Solid dispersion of acetaminophen and poly(ethylene oxide) prepared by hot-melt mixing, Int. J. Pharm., 2010, 395, 53–61 CrossRef CAS PubMed.
- A. R. Paradkar, M. Maheshwari, A. R. Ketkar and B. Chauhan, Preparation and evaluation of ibuprofen beads by melt solidification technique, Int. J. Pharm., 2003, 255, 33–42 CrossRef CAS.
- A. Milanovic, I. Aleksic, S. Ibric, J. Parojcic and S. Cvijic, Tableting of hot-melt coated paracetamol granules: Material tableting properties and quality characteristics of the obtained tablets, Eur. J. Pharm. Sci., 2020, 142, 105121 CrossRef CAS PubMed.
- W. J. Genck, B. Albin, F. A. Baczek, D. S. Dickey, C. G. Gilbert, T. Herrera, T. J. Laros, W. Li, P. McCurdie, J. K. McGillicuddy, T. P. McNutty, C. G. Moyers, F. Schoenbrunn, T. W. Wisdom and W. Chen, Liquid-Solid Operations and Equipment, in Perry's Chemical Engineers' Handbook, ed. Green, D. W. and Southard, M. Z., McGraw-Hill Education, New York, 2019, pp. 1895–2050 Search PubMed.
- Thermodynamics Research Center (TRC), Thermodynamics Source Database, in NIST Chemistry WebBook, ed. Linstrom, P. J. and Mallard, W. G., National Institute of Standards and Technology, Gaithersburg, MD, 2025, DOI:10.18434/T4D303.
- K. Mansouri, C. M. Grulke, A. M. Richard, R. S. Judson and A. J. Williams, An automated curation procedure for addressing chemical errors and inconsistencies in public datasets used in QSAR modelling, SAR QSAR Environ. Res., 2016, 27, 939–965 CrossRef CAS.
- J.-C. Bradley, A. Williams and A. Lang, Jean-Claude Bradley Open Melting Point Dataset, 2014, vol. 2, DOI:10.6084/m9.figshare.1031638.v1.
- C. Wespiser and D. Mathieu, Application of Machine Learning to the Design of Energetic Materials: Preliminary Experience and Comparison with Alternative Techniques, Propellants, Explos., Pyrotech., 2023, 48, e202200264 CrossRef CAS.
- J. Rein, J. M. Meinhardt, J. L. Hofstra, M. S. Sigman and S. Lin, A Physical Organic Approach towards Statistical Modeling of Tetrazole and Azide Decomposition, Angew. Chem., 2023, 62, e202218213 CrossRef CAS.
- H. Wiener, Structural Determination of Paraffin Boiling Points, J. Am. Chem. Soc., 1947, 69, 17–20 CrossRef CAS.
- K. Mansouri, C. M. Grulke, R. S. Judson and A. J. Williams, OPERA models for predicting physicochemical properties and environmental fate endpoints, J. Cheminf., 2018, 10, 10 Search PubMed.
- S. L. Dixon, J. Duan, E. Smith, C. D. Von Bargen, W. Sherman and M. P. Repasky, AutoQSAR: An automated machine learning tool for best-practice quantitative structure-activity relationship modeling, Future Med. Chem., 2016, 8, 1825–1839 CrossRef CAS PubMed.
- K. Paduszyński, K. Kłębowski and M. Królikowska, Predicting melting point of ionic liquids using QSPR approach: Literature review and new models, J. Mol. Liq., 2021, 344, 117631 CrossRef.
- X. Zang, X. Zhou, H. Bian, W. Jin, X. Pan, J. Jiang, M. Yu. Koroleva and R. Shen, Prediction and Construction of Energetic Materials Based on Machine Learning Methods, Molecules, 2022, 28, 322 CrossRef PubMed.
- D. Klinger, A. Casey, T. Manship, S. Son and A. Strachan, Prediction of Solid Propellant Burning Rate Characteristics Using Machine Learning Techniques, Propellants, Explos., Pyrotech., 2023, 48, e202200267 CrossRef CAS.
- P. Borlido, J. Schmidt, A. W. Huran, F. Tran, M. A. L. Marques and S. Botti, Exchange-correlation functionals for band gaps of solids: benchmark, reparametrization and machine learning, npj Comput. Mater., 2020, 6, 96 CrossRef CAS.
- R. Gee and R. Lindsey, Revolutionizing Energetic Materials Discovery and Design: The Role of Data Science and Machine Learning, Propellants, Explos., Pyrotech., 2023, 48, e202380431 CrossRef.
- S. Wang, S. Gong, T. Böger, J. A. Newnham, D. Vivona, M. Sokseiha, K. Gordiz, A. Aggarwal, T. Zhu, W. G. Zeier, J. C. Grossman and Y. Shao-Horn, Multimodal Machine Learning for Materials Science: Discovery of Novel Li-Ion Solid Electrolytes, Chem. Mater., 2024, 36, 11541–11550 CrossRef CAS.
- A. Mannodi-Kanakkithodi, A guide to discovering next-generation semiconductor materials using atomistic simulations and machine learning, Comput. Mater. Sci., 2024, 243, 113108 CrossRef CAS.
- J. Yang, P. Manganaris and A. Mannodi-Kanakkithodi, Discovering novel halide perovskite alloys using multi-fidelity machine learning and genetic algorithm, J. Chem. Phys., 2024, 160, 064114 CrossRef CAS.
- R. J. Appleton, P. Salek, A. D. Casey, B. C. Barnes, S. F. Son and A. Strachan, Interpretable Performance Models for Energetic Materials using Parsimonious Neural Networks, J. Phys. Chem. A, 2024, 128, 1142–1153 CrossRef CAS.
- C. Li, J. C. Verduzco, B. H. Lee, R. J. Appleton and A. Strachan, Mapping microstructure to shock-induced temperature fields using deep learning, npj Comput. Mater., 2023, 9, 178 CrossRef CAS.
- C. C. Chen, R. J. Appleton, S. Mishra, K. Nykiel and A. Strachan, Discovery of new high-pressure phases – integrating high-throughput DFT simulations, graph neural networks, and active learning, npj Comput. Mater., 2025, 11, 191 CrossRef CAS.
- Y. Zhang, R. J. Appleton, K. Lin, M. J. McCarthy, J. T. Paci, S. K. R. S. Sankaranarayanan, A. Strachan and H. D. Espinosa, Generalizable machine learning potentials for quantum-accurate predictions of non-equilibrium behavior in 2D materials, Comput. Method Appl. M., 2026, 448, 118502 CrossRef.
- C. W. Yap, PaDEL-descriptor: An open source software to calculate molecular descriptors and fingerprints, J. Comput. Chem., 2011, 32, 1466–1474 CrossRef CAS.
- H. Moriwaki, Y. S. Tian, N. Kawashita and T. Takagi, Mordred: A molecular descriptor calculator, J. Cheminf., 2018, 10, 4 Search PubMed.
- R. J. Appleton, D. Klinger, B. H. Lee, M. Taylor, S. Kim, S. Blankenship, B. C. Barnes, S. F. Son and A. Strachan, Multi-Task Multi-Fidelity Learning of Properties for Energetic Materials, Propellants, Explos., Pyrotech., 2024, 50, e202400248 CrossRef.
- Z. Boukouvalas, M. Puerto, D. C. Elton, P. W. Chung and M. D. Fuge, Independent vector analysis for molecular data fusion: application to property prediction and knowledge discovery of energetic materials, in 28th European Signal Processing Conference vols 2021-January, IEEE, 2021, pp. 1030–1034 Search PubMed.
- D. C. Elton, Z. Boukouvalas, M. S. Butrico, M. D. Fuge and P. W. Chung, Applying machine learning techniques to predict the properties of energetic materials, Sci. Rep., 2018, 8, 9059 CrossRef PubMed.
- B. C. Barnes, D. C. Elton, Z. Boukouvalas, D. E. Taylor, W. D. Mattson, M. D. Fuge and P. W. Chung, Machine Learning of Energetic Material Properties, in Proceedings of the 16th International Detonation Symposium, 2018, DOI:10.48550/arXiv.1807.06156.
- M. Sun, S. Zhao, C. Gilvary, O. Elemento, J. Zhou and F. Wang, Graph convolutional networks for computational drug development and discovery, Briefings Bioinf., 2020, 21, 919–935 CrossRef.
- K. Liu, X. Sun, L. Jia, J. Ma, H. Xing, J. Wu, H. Gao, Y. Sun, F. Boulnois and J. Fan, Chemi-net: A molecular graph convolutional network for accurate drug property prediction, Int. J. Mol. Sci., 2019, 20, 3389 CrossRef CAS.
- M. H. Rahman, P. Gollapalli, P. Manganaris, S. K. Yadav, G. Pilania, B. DeCost, K. Choudhary and A. Mannodi-Kanakkithodi, Accelerating defect predictions in semiconductors using graph neural networks, APL Mach. Learn., 2024, 2, 016122 CrossRef CAS.
- K. Yang, K. Swanson, W. Jin, C. Coley, P. Eiden, H. Gao, A. Guzman-Perez, T. Hopper, B. Kelley, M. Mathea, A. Palmer, V. Settels, T. Jaakkola, K. Jensen and R. Barzilay, Analyzing Learned Molecular Representations for Property Prediction, J. Chem. Inf. Model., 2019, 59, 3370–3388 CrossRef CAS.
- E. Heid, K. P. Greenman, Y. Chung, S. C. Li, D. E. Graff, F. H. Vermeire, H. Wu, W. H. Green and C. J. McGill, Chemprop: A Machine Learning Package for Chemical Property Prediction, J. Chem. Inf. Model., 2024, 64, 9–17 CrossRef CAS PubMed.
- J. Gasteiger, J. Groß and S. Günnemann, Directional Message Passing for Molecular Graphs, in International Conference on Learning Representations, 2020, DOI:10.48550/arXiv.2003.03123.
- S. Zhang, Y. Liu and L. XieMolecular Mechanics-Driven Graph Neural Network with Multiplex Graph for Molecular Structures, 2020, DOI:10.48550/arXiv.2011.07457.
- C. Chen, W. Ye, Y. Zuo, C. Zheng and S. P. Ong, Graph Networks as a Universal Machine Learning Framework for Molecules and Crystals, Chem. Mater., 2019, 31, 3564–3572 CrossRef CAS.
- R. J. Appleton, B. C. Barnes and A. Strachan, Data Fusion of Deep Learned Molecular Embeddings for Property Prediction, J. Chem. Inf. Model., 2025, 65, 11620–11630 CrossRef CAS.
- R. Gómez-Bombarelli, J. N. Wei, D. Duvenaud, J. M. Hernández-Lobato, B. Sánchez-Lengeling, D. Sheberla, J. Aguilera-Iparraguirre, T. D. Hirzel, R. P. Adams and A. Aspuru-Guzik, Automatic Chemical Design Using a Data-Driven Continuous Representation of Molecules, ACS Cent. Sci., 2018, 4, 268–276 CrossRef.
- W. Jin, K. Yang, R. Barzilay and T. Jaakkola, Learning Multimodal Graph-to-Graph Translation for Molecular Optimization, 2018, DOI:10.48550/arXiv.1812.01070.
- R. Assouel, M. Ahmed, M. H. Segler, A. Saffari and Y. Bengio, DEFactor: Differentiable Edge Factorization-based Probabilistic Graph Generation, 2018, DOI:10.48550/arXiv.1811.09766.
- Q. Liu, M. Allamanis, M. Brockschmidt and A. L. Gaunt Constrained Graph Variational Autoencoders for Molecule Design. in Advances in neural information processing systems, vol. 31, 2018 Search PubMed.
- W. Jin, R. Barzilay and T. Jaakkola, Junction Tree Variational Autoencoder for Molecular Graph Generation, in Proceedings of the 35th International Conference on Machine Learning, 2018, pp. 2323–2332 Search PubMed.
- T. Ochiai, T. Inukai, M. Akiyama, K. Furui, M. Ohue, N. Matsumori, S. Inuki, M. Uesugi, T. Sunazuka, K. Kikuchi, H. Kakeya and Y. Sakakibara, Variational autoencoder-based chemical latent space for large molecular structures with 3D complexity, Commun. Chem., 2023, 6, 249 CrossRef PubMed.
- M. J. Kusner, B. Paige and J. Miguel Hernández-Lobato, Grammar Variational Autoencoder, in Proceedings of the 34th International Conference on Machine Learning, 2017, pp. 1945–1954 Search PubMed.
- J. Lim, S. Y. Hwang, S. Moon, S. Kim and W. Y. Kim, Scaffold-based molecular design with a graph generative model, Chem. Sci., 2020, 11, 1153–1164 RSC.
- Y. Pathak, K. S. Juneja, G. Varma, M. Ehara and U. D. Priyakumar, Deep learning enabled inorganic material generator, Phys. Chem. Chem. Phys., 2020, 22, 26935–26943 RSC.
- J. Boitreaud, V. Mallet, C. Oliver and J. Waldispühl, OptiMol: Optimization of Binding Affinities in Chemical Space for Drug Discovery, J. Chem. Inf. Model., 2020, 60, 5658–5666 CrossRef CAS PubMed.
- Z. Yao, B. Sánchez-Lengeling, N. S. Bobbitt, B. J. Bucior, S. G. H. Kumar, S. P. Collins, T. Burns, T. K. Woo, O. K. Farha, R. Q. Snurr and A. Aspuru-Guzik, Inverse design of nanoporous crystalline reticular materials with deep generative models, Nat. Mach. Intell., 2021, 3, 76–86 CrossRef.
- Ł. Maziarka, A. Pocha, J. Kaczmarczyk, K. Rataj, T. Danel and M. Warchoł, Mol-CycleGAN: a generative model for molecular optimization, J. Cheminf., 2020, 12, 2 Search PubMed.
- N. De Cao and T. Kipf, MolGAN: an implicit generative model for small molecular graphs, 2018, DOI:10.48550/arXiv.1805.11973.
- G. L. Guimaraes, B. Sanchez-Lengeling, C. Outeiral, P. L. C. Farias and A. Aspuru-Guzik, Objective-Reinforced Generative Adversarial Networks (ORGAN) for Sequence Generation Models, 2017, DOI:10.48550/arXiv.1705.10843.
- J. Park, Y. Lee and J. Kim, Multi-modal conditional diffusion model using signed distance functions for metal-organic frameworks generation, Nat. Commun., 2025, 16, 34 CrossRef.
- N. T. Runcie and A. S. J. S. Mey, SILVR: Guided Diffusion for Molecule Generation, J. Chem. Inf. Model., 2023, 63, 5996–6005 CrossRef CAS.
- L. Huang, T. Xu, Y. Yu, P. Zhao, X. Chen, J. Han, Z. Xie, H. Li, W. Zhong, K. C. Wong and H. Zhang, A dual diffusion model enables 3D molecule generation and lead optimization based on target pockets, Nat. Commun., 2024, 15, 2657 CrossRef CAS.
- A. Alakhdar, B. Poczos and N. Washburn, Diffusion Models in De Novo Drug Design, J. Chem. Inf. Model., 2024, 64, 7238–7256 CrossRef CAS PubMed.
- C. C. Chen, R. J. Appleton, K. Nykiel, S. Mishra, S. Yao and A. Strachan, How accurate is density functional theory at high pressures?, Comput. Mater. Sci., 2025, 247, 113458 CrossRef CAS.
- F. Biermair, V. I. Razumovskiy and G. Ressel, Influence of alloying on thermodynamic properties of AlCoCrFeNiTi high entropy alloys from DFT calculations, Comput. Mater. Sci., 2022, 202, 110952 CrossRef CAS.
- A. Kumar, Z. A. Ali and B. M. Wong, Efficient predictions of formation energies and convex hulls from density functional tight binding calculations, J. Mater. Sci. Technol., 2023, 141, 236–244 CrossRef CAS.
- K. Nykiel and A. Strachan, High-throughput density functional theory screening of double transition metal MXene precursors, Sci. Data, 2023, 10, 827 CrossRef CAS PubMed.
- S. Karumuri, A. Hernandez, S. Mishra, Z. McClure, V. Tucker, J. C. Flanagan, S. Hwang, K. H. Sandhage, I. Bilionis, M. S. Titus and A. Strachan, Design of high-hardness complex concentrated alloys from physics, machine learning, and experiments, J. Appl. Phys., 2025, 138, 085106 CrossRef CAS.
- T. Slater and D. Timms, Meeting on binding sites: Characterizing and satisfying steric and chemical restraints, J. Mol. Graphics, 1993, 11, 248–251 CrossRef CAS.
- D. R. Westhead, D. E. Clark, D. Frenkel, J. Li, C. W. Murray, B. Robson and B. Waszkowycz, PRO_LIGAND: an approach to de novo molecular design. 3. A genetic algorithm for structure refinement, J. Comput.-Aided Mol. Des., 1995, 9, 139–148 CrossRef CAS PubMed.
- R. C. Glen and W. R. Payne, A genetic algorithm for the automated generation of molecules within constraints, J. Comput.-Aided Mol. Des., 1995, 9, 181–202 CrossRef CAS.
- D. Weininger, Smiles, a chemical language and information system. 1. introduction to methodology and encoding rules, J. Chem. Inf. Comput. Sci., 1988, 28, 31–36 CrossRef CAS.
- D. Weininger, A. Weininger and J. L. Weininger, Smiles. 2. algorithm for generation of unique smiles notation, J. Chem. Inf. Comput. Sci., 1989, 29, 97–101 CrossRef CAS.
- P. Polishchuk, CReM: chemically reasonable mutations framework for structure generation, J. Cheminf., 2020, 12, 28 Search PubMed.
- J. Leguy, T. Cauchy, M. Glavatskikh, B. Duval and B. Da Mota, EvoMol: a flexible and interpretable evolutionary algorithm for unbiased de novo molecular generation, J. Cheminf., 2020, 12, 55 CAS.
- Y. Kwon and J. Lee, MolFinder: an evolutionary algorithm for the global optimization of molecular properties and the extensive exploration of chemical space using SMILES, J. Cheminf., 2021, 13, 24 CAS.
- J. H. Jensen, A graph-based genetic algorithm and generative model/Monte Carlo tree search for the exploration of chemical space, Chem. Sci., 2019, 10, 3567–3572 RSC.
- M. Krenn, F. Häse, A. K. Nigam, P. Friederich and A. Aspuru-Guzik, Self-referencing embedded strings (SELFIES): a 100% robust molecular string representation, Mach. Learn. Sci. Technol., 2020, 1, 045024 CrossRef.
- A. Nigam, R. Pollice, M. Krenn, G. D. P. Gomes and A. Aspuru-Guzik, Beyond generative models: superfast traversal, optimization, novelty, exploration and discovery (STONED) algorithm for molecules using SELFIES, Chem. Sci., 2021, 12, 7079–7090 RSC.
- A. K. Nigam, R. Pollice and A. Aspuru-Guzik, Parallel tempered genetic algorithm guided by deep neural networks for inverse molecular design, Digital Discovery, 2022, 1, 390–404 RSC.
- S. Kim, J. Chen, T. Cheng, A. Gindulyte, J. He, S. He, Q. Li, B. A. Shoemaker, P. A. Thiessen, B. Yu, L. Zaslavsky, J. Zhang and E. E. Bolton, PubChem 2023 update, Nucleic Acids Res., 2023, 51, D1373–D1380 CrossRef.
- SciFinder, https://scifinder-n.cas.org/ Search PubMed.
- G. Landrum, RDKit: Open-source Cheminformatics, 2023, DOI:10.5281/zenodo.591637.
- J. L. Lansford, K. F. Jensen and B. C. Barnes, Physics-informed Transfer Learning for Out-of-sample Vapor Pressure Predictions, Propellants, Explos., Pyrotech., 2023, 48, e202200265 CrossRef CAS.
- U. H. E. Hansmann, Parallel tempering algorithm for conformational studies of biological molecules, Chem. Phys. Lett., 1997, 281, 140–150 CrossRef CAS.
- D. J. Earl and M. W. Deem, Parallel tempering: theory, applications, and new perspectives, Phys. Chem. Chem. Phys., 2005, 7, 3910–3916 RSC.
- T. Galeazzo and M. Shiraiwa, Predicting glass transition temperature and melting point of organic compounds via machine learning and molecular embeddings, Environ. Sci.: Atmos., 2022, 2, 362–374 CAS.
- D. V. Schroeder, An Introduction to Thermal Physics, Oxford University Press, 2000 Search PubMed.
- M. Swain, PubChemPy: A Python Wrapper for the PubChem API, https://pubchempy.readthedocs.io/en/latest/index.html Search PubMed.
- M. Swain, CIRpy: A Python Library for Chemical Information Retrieval, https://cirpy.readthedocs.io/en/latest/index.html Search PubMed.
- D. J. Rogers and T. T. Tanimoto, A Computer Program for Classifying Plants: The computer is programmed to simulate the taxonomic process of comparing each case with every other case, Science, 1979, 132, 1115–1118 CrossRef.
- S. Chithrananda, G. Grand and B. Ramsundar, ChemBERTa: Large-Scale Self-Supervised Pretraining for Molecular Property Prediction, 2020, DOI:10.48550/arXiv.2010.09885.
- P. Ertl and A. Schuffenhauer, Estimation of synthetic accessibility score of drug-like molecules based on molecular complexity and fragment contributions, J. Cheminf., 2009, 1, 8 Search PubMed.
- E. Antoniuk, P. Li, N. Keilbart, S. Weitzner, B. Kailkhura and A. M. Hiszpanski, Active Learning Enables Extrapolation in Molecular Generative Models, 2025, DOI:10.48550/arXiv.2501.02059.
| This journal is © The Royal Society of Chemistry 2026 |