Open Access Article
Open Access ArticleExploring the deviation from Nernst–Einstein conductivity in ionic liquids using machine learning
Aditi
Seshadri
 a,
Lyndon T. M.
Hess
a,
Lyndon T. M.
Hess
 b and
Shuwen
Yue
b and
Shuwen
Yue
 *a
*a
aRobert F. Smith School of Chemical and Biomolecular Engineering, Cornell University, Ithaca, NY 14853, USA. E-mail: shuwen.yue@cornell.edu
bDepartment of Chemistry and Chemical Biology, Cornell University, Ithaca, NY 14853, USA
First published on 9th March 2026
Abstract
Ionic liquids (ILs) are promising candidates for safer battery electrolytes due to their low flammability, but their low ionic conductivity constrains battery performance. Given the large chemical space of potential ILs, machine learning (ML) approaches are essential for accelerated screening. However, previous studies have shown that ML models trained on common computational descriptors to predict molar ionic conductivity have accuracies comparable to the Nernst–Einstein equation [Umaña, Cashen, Zavala and Gebbie, Digital Discovery, 2025, 4, 1423–1436]. Furthermore, experimental measurements show that the ionic conductivity of many ILs deviates substantially from that predicted by the Nernst–Einstein equation. While several mechanisms have been proposed, the structural origins of these deviations are not well understood. In this study, we develop ML models to predict the deviation of the ionic conductivity of ILs from Nernst–Einstein behavior using charge-based descriptors for individual ions. We observed that ML models trained using a smaller set of sigma profile-based descriptors had similar performance to those trained using RDKit cheminformatic descriptors. Additionally, we found that models trained to predict this deviation could serve as a correction factor to the Nernst–Einstein equation and resulted in more accurate conductivity predictions compared to models that were designed to directly predict the molar ionic conductivity of ILs. We applied feature importance rankings to gain insight into model predictions and identified features relating to the cation alkyl chain length and the cation and anion polarity as being influential.
1 Introduction
Ionic liquids (ILs) are considered safer alternatives to organic battery electrolytes due to their thermal stability and low flammability. However, their widespread use is limited by the fact that many ILs have a low ionic conductivity, which constrains the battery charge/discharge rates.2–4 Due to the large chemical space of potential ILs, it is impractical to rely solely on experimental methods for tackling this problem.5 In recent years, machine learning (ML) models have bridged this gap and been used to predict properties of ILs, including their viscosity, melting point, and ionic conductivity in a computationally efficient manner.1,6–12Despite these advances, accurately predicting ionic conductivity remains challenging because conductivity depends not only on molecular structure, but also on temperature, ion–ion correlations, and collective transport mechanisms that are not easily captured by generic molecular descriptors.
Multiple studies have trained ML models to predict the ionic conductivity of ILs.1,8–10,12–15 However, recently, Umaña et al.1 found that ML models trained solely on RDKit cheminformatic descriptors had accuracies comparable to the Nernst–Einstein equation, and that an Arrhenius-type fit to conductivity data led to a greater accuracy. This result suggests that generic cheminformatic descriptors cannot fully capture the physics of ion transport, which is better described by thermally activated processes and electrostatic interactions. The study also showed that deviations from Nernst–Einstein behavior are particularly challenging to model with such descriptors and suggests the need for physically motivated features that encode charge distribution effects.
The Nernst–Einstein equation (eqn (1)) can be used to estimate ionic conductivity and assumes ions move independently and neglects ion–ion interactions. While this equation is useful in screening chemical systems, the experimentally measured ionic conductivity of ILs often deviates from that estimated using the Nernst–Einstein equation.1,13,16Fig. 1 shows that while some ILs follow Nernst–Einstein behavior, many exhibit large systematic deviations that cannot be explained by independent-ion transport. This deviation can be expressed as the ratio of the measured molar ionic conductivity (σ) and the Nernst–Einstein conductivity (σNE), and has been referred to in the literature as ionicity, the inverse Haven ratio, and the Nernst–Einstein ratio (eqn (2)).1,13,17,18 Additionally, the ratio σ/σNE, which we shall refer to as ionicity after this point, is thought to encode information relating to the extent of ion dissociation, charge transfer, correlated ion motion, and presence of alternative ion transport mechanisms.1,13,16,17,19–22 Although the term ionicity has been used as a measure of the extent of ion dissociation, one limitation of this definition is that the meaning of ionicity values above one, which has been observed for some ILs, is unclear.13,17,18 In this study, our use of the term ionicity is in reference to its mathematical definition given in eqn (2) and can encompass multiple interpretations of why ILs deviate from Nernst–Einstein behavior, including ion dissociation, charge transfer effects, and correlated ion motion. Being able to predict when an IL will deviate from Nernst–Einstein behavior and what molecular features drive this deviation can provide insight into ion transport, indicate when additional measurements are needed for accurate molar conductivity estimates, and ultimately guide the accelerated design of high-performance ILs.
 | ||
| Fig. 1 Comparison between molar ionic conductivities estimated using the Nernst–Einstein equation and experimentally measured molar ionic conductivities for all ILs in this study. For some ILs, the Nernst–Einstein equation is a reasonable approximation (corresponding to points along the ideal y = x dashed line) and in other cases, it is not. Each data point corresponds to a specific IL at a given temperature and pressure. Experimental data were obtained from the NIST ILThermo dataset (Section 2.1)23,24 | ||
Cashen et al.13 built a linear model trained on experimental and computational descriptors to predict ionicity, and they identified density, molar concentration, and ion geometry as influential features. In addition to Nernst–Einstein based definitions, ionicity has also been estimated empirically using Walden plots, which relate conductivity to reciprocal viscosity and classify ILs relative to a dilute KCl reference line.17,18,25,26 Makarov et al.27 built ML models to classify “high” and “low” ionicity ILs using Walden plots.27 However, some limitations with the Walden plot approach relate to it being empirical and that the KCl reference has been found to be non-ideal.13,17,18,28
Although the influence of charge transfer on the ionicity of ILs has been investigated from a theoretical perspective, these studies are difficult to integrate into a high-throughput screening workflow.19,20,22 Computing partial charges or other quantum mechanical properties related to ion–ion interactions in ILs is computationally expensive and sensitive to the theoretical approach used.19,22 Thus, we explore whether a ML model can learn information intrinsic to the IL (i.e. cation–anion pair) from sigma profile-based descriptors for isolated individual ions, which are relatively inexpensive to compute but encode polarity and charge distribution features that could indirectly inform ion–ion interactions. Past studies have found that ML models trained on individual ion properties, rather than the combined ion pair, could predict IL melting points.11
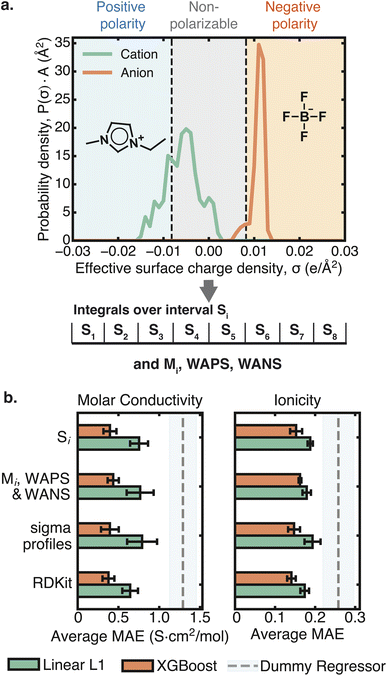
Sigma profiles describe the surface charge density of a molecule and have been used to predict IL properties, including viscosity, ionic conductivity, and melting point.6,7,10–12,29–31 Often in ML studies, descriptors are derived from the area under the sigma profile curve in various regions.6,10,12,32 These regions can be grouped into positive polarity (σ < −0.082e/Å2), negative polarity (σ > 0.082e/Å2), and nonpolarizable (σ ∈ (−0.082e/Å2, 0.082e/Å2)) domains (Fig. 2a).33 Additional interpretable descriptors can be derived from the sigma profile that quantify properties such as the extent of charge delocalization in cations and anions, polarity, and strength of hydrogen-bonding interactions.34–36
In this study, we evaluate whether sigma profile-derived descriptors can be used in ML models to predict ionicity and, through feature-importance analysis, identify which ion properties may govern deviations from Nernst–Einstein behavior. By focusing on ionicity rather than conductivity, our goal is not only to improve prediction but also to extract interpretable chemical insights that can accelerate IL design.
To the best of our knowledge, this is the first study to apply sigma profile-derived descriptors to predict ionicity in ILs, thereby linking these quantum mechanical-based descriptors to deviations from ideal transport.
2 Methods
2.1 ILThermo dataset preprocessing
We trained ML models to predict both the ionicity and molar conductivity of ILs using data from the NIST ILThermo dataset.23,24 This database contains experimental measurements of density, viscosity, and electrical (ionic) conductivity of pure, single-component ILs at different temperatures and pressures. The ILThermoPy Python package was used to interface with the NIST ILThermo dataset and obtain SMILES strings for each IL.37 We applied a multi-step data cleaning and preprocessing procedure based on workflows developed in earlier studies that extracted consistent subsets of data from the NIST ILThermo database for machine learning and structure-property analysis.1,8,17 We first removed data points where the IL was not in the liquid phase or the temperature and/or pressure conditions for the conductivity, viscosity, and density measurements were not provided. Additionally, we excluded entries where the pressure was outside the atmospheric pressure range (101.325 ± 5 kPa). When multiple experimental values existed for the same IL at the same thermodynamic conditions, we discarded measurements with reported uncertainties greater than 15% or with values that deviated by more than 15% from the mean. If multiple entries still remained, we kept the result with the lowest reported uncertainty. Lastly, any outlier ILs that contained transition metals or had an estimated ionicity above 10 were removed (SI S2). After preprocessing, the dataset contained 337 unique ILs and 2936 total data points at different temperatures and pressures.2.2 Ionicity estimation
The Nernst–Einstein equation (eqn (1)) assumes integer charges on the ions and does not account for ion–ion interactions.1,13,17,19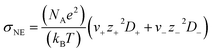 | (1) |
Ionicity is defined as the ratio of the measured molar conductivity (σ) to that estimated using the Nernst–Einstein equation (σNE).
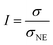 | (2) |
As self-diffusivity data was not directly available for some cations and anions in the NIST ILThermo database, the Nernst–Einstein conductivity of every IL was estimated using the Stokes–Einstein equation and viscosity data (eqn (3)).1,13,17,38
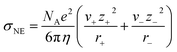 | (3) |
In order to calculate the Nernst–Einstein conductivity using eqn (3), the ion radii need to be estimated. After comparing different approaches for estimating the cation and anion radii, we selected the ratio of the ion volume to surface area (SI S2.2). To obtain these quantities, we first conducted geometry optimizations of the ions using the MMFF94 force field and then estimated the ion volumes and surface areas using the RDKit cheminformatics package.39 Additional details regarding the ionic radii estimation methods compared can be found in SI S2.2.
2.3 Descriptor calculation
As it is not known a priori if a given IL will deviate from the Nernst–Einstein behavior, we aimed to build ML models that could (a.) predict the ionicity of ILs and (b.) be used to identify molecular features that are important in predicting IL ionicity. Thus, we chose features that were (a.) computationally accessible and compatible with a high-throughput screening workflow, and (b.) interpretable and examined the hypothesis that charge-based descriptors are sufficient for predicting ionicity. Sigma profiles were calculated for each cation and anion separately using openCOSMO-RS and ORCA.40,41 To obtain more interpretable descriptors from the sigma profiles, the zeroth, first, second, and third moments of the sigma profile, which relate to the ion surface area, charge, polarity, and skewness of the sigma profile, were calculated.36,40 Also, the anion weighted average positive sigma (WAPS) and cation weighted average negative sigma (WANS) were calculated. These descriptors have been proposed to quantify the anion and cation charge localization.34,35 Lastly, the area under the sigma profile curve over eight different intervals was calculated. Additional details regarding the sigma profile descriptors can be found in SI S3.2.4 Machine learning model development
Due to the small size of the dataset, as well as the aim of developing interpretable models, traditional ML models (i.e. linear and decision tree-based models) were used rather than deep learning approaches.42 To identify the optimal model hyperparameters, 5-fold cross-validation was used. During the cross-validation procedure, only the training dataset (90% of the entire dataset) was used for training and evaluating the model. The training dataset was divided into five sets. For each hyperparameter choice, models were trained on different combinations of four of these sets (80% of the training dataset) and then evaluated on the remaining fifth set (20% of the training dataset). For the linear models, the regularization strength was varied, and for the decision tree-based models, hyperparameters corresponding to the maximum depth and number of estimators were varied. Based on the cross-validation performance of the regularized linear models (with L1 or L2 regularization) and decision tree-based models (random forest or XGBoost), we selected linear models with L1 regularization (hereafter referred to as linear L1 models) and XGBoost models for further study.43The optimal regularization strength for the linear L1 models ranged from 0.0001 to 0.01 depending on the prediction task (molar conductivity or ionicity) and the input set of descriptors (area under the cation and anion sigma profile curves (Si) descriptors, RDKit descriptors, etc.) to the model (SI S4, Tables S3 & S5). These optimal regularization strengths fall within the range of values tested for the linear L1 models, 10−6 to 1050. For the XGBoost models, the maximum depth values tested ranged from [1, 40] and the number of estimators ranged from [1, 1000]. When predicting ionicity, the optimal XGBoost models had a maximum depth between 8 and 23 along with the number of estimators ranging from 70 to 160 (SI S4, Table S6). For molar conductivity prediction, the optimal number of estimators in the XGBoost models were generally greater than those used when predicting ionicity (optimal values ranging from 720 to 1000), while the optimal maximum depth values ranged from 3 to 13 (SI S4, Table S4). The optimal hyperparameter choices for each input descriptor set and prediction task for the linear L1 and XGBoost models can be found in the SI S4.
All features were scaled to fall within a range of zero and one using scikit-learn's MinMaxScaler, and highly correlated features (|correlation coefficient| ≥ 0.9) were removed.44 The dataset was split such that 90% of the data was used for training the models and 10% was used for the test set. It was also ensured that no IL was present in both the training and test datasets.
The ML model performance using sigma profiles and sigma profile-derived descriptors were compared to a baseline model trained using RDKit descriptors, a common set of cheminformatic descriptors, that were derived from the 2D ion structures.39 In every input feature set, the temperature and pressure were included. All ML models were also compared to a dummy regressor model that outputted the mean of the training data. Additional details regarding the machine learning model development can be found in the SI S4.
2.5 Feature importance analysis
To gain insight into the IL-ionicity relationship, different feature importance analysis methods were compared. We focused on the linear L1 models as they were less prone to overfitting. The three feature analysis methods were: (1) analyzing the coefficients from the linear model, (2) permutation feature importance rankings, and (3) SHAP values.45 The implementation of permutation feature importance in scikit-learn was used in this work, with nrepeats (the number of times the values of a given feature are permuted) set to 10.44 By comparing multiple feature importance methods, we can overcome some of the potential limitations associated with each method. We also grouped ILs by cation families (SI S2.1) and the longest alkyl chain length in the cation to identify trends relating to the impact of each feature on the overall ionicity or molar conductivity of an IL in that group.3 Results and discussion
3.1 Models trained using charge-based descriptors perform as well as those trained using 2D cheminformatic features
We first compared the linear L1 and XGBoost model performance from 5-fold cross-validation across different input descriptor sets (Fig. 2b, SI S6.1). Although the XGBoost model had a lower mean absolute error (MAE), it also exhibited a greater degree of overfitting, with larger train-test discrepancies (SI S6.1). On the test set, the linear L1 model was slightly less accurate but appeared to be more generalizable (i.e. similar performance between train and test sets) (SI S6.1). For this reason, we use the linear L1 model for all further analyses.The models trained on sigma profile-based descriptors performed comparably to those trained on the initially larger set of RDKit cheminformatic descriptors for both ionicity and molar conductivity prediction (Fig. 2, SI S6.1). This finding is notable because the sigma profile-based descriptors are a compact, physically grounded descriptor set derived from the cation and anion charge distributions, whereas the RDKit descriptors used represent cheminformatic features spanning a range of physicochemical properties, fragment counts, and topological indices from 2D structures.39
Although the initial RDKit descriptor set (153 RDKit descriptors in addition to temperature and pressure) used to train the models is much greater than the number of sigma profile-related descriptors (ex: 15 Si or area under the sigma profile curve descriptors in addition to temperature and pressure), the number of descriptors with nonzero coefficients for the linear models with L1 regularization was much lower. Specifically, for the linear L1 models trained to predict ionicity, there were 46 descriptors with nonzero coefficients when the RDKit input descriptor set was used and 16 descriptors with nonzero coefficients when the Si input descriptor set was used. For molar conductivity prediction, there were 17 descriptors with nonzero coefficients when the RDKit input descriptor set was used and 12 descriptors with nonzero coefficients when the Si input descriptor set was used (SI S4, Tables S7 & S8). Despite the fact that the final number of descriptors with nonzero coefficients in the linear models are of similar orders of magnitude for the RDKit and sigma-profile based sets, the interpretability of many of the RDKit descriptors with nonzero coefficients is less clear. Some RDKit descriptors, such as the number of benzene rings, are easily interpretable in the context of the chemical structure of the IL cations and anions. However, with many other RDKit descriptors, such as those that relate to atomic contributions to molar refractivity (SMR_VSA descriptors), it is more difficult to interpret their relation to the chemical structure.46
The fact that the RDKit and sigma profile-based descriptor sets yielded models with similar predictive accuracy suggests that much of the relevant information for transport properties may be encoded in charge-related features rather than 2D topological or fragment-based structural descriptors. From a practical standpoint, this means that sigma profiles can replace a larger, and potentially more redundant feature set with a small number of interpretable, physically meaningful quantities without having to apply extensive feature selection. Their connection to ion polarity and charge localization also provides a more direct link to the molecular origins of deviations from Nernst–Einstein behavior, while still being efficient enough for high-throughput screening.
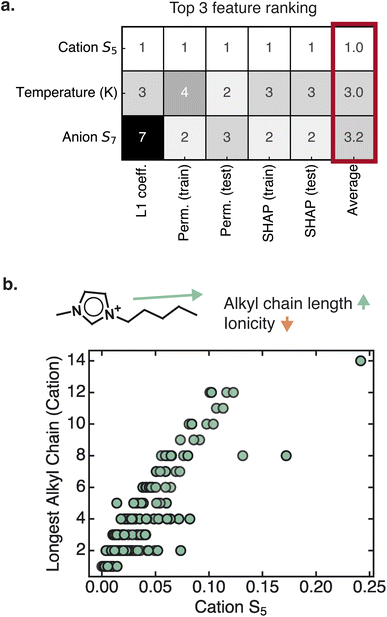
3.2 Cation nonpolar surface area and anion H-bond acceptor strength dominate feature importance
We next examined which of the sigma profile-based descriptors most strongly influence ionicity predictions. After comparing the results from different feature importance methods (linear model coefficients, permutation feature importance rankings, and mean SHAP values), we focused on the descriptors that consistently appeared near the top of the rankings across all methods (Fig. 3, SI S7). Focusing on the consensus of these feature importance rankings increases our confidence in identifying robust trends in the data.Among the Si descriptors, the cation S5 descriptor emerged as the strongest predictor of ionicity. By definition, S5 quantifies the area of the sigma profile curve in half of the nonpolarizable region and relates to the cation nonpolar surface area. Furthermore, greater cation S5 values tend to result in lower ionicity and conductivity predictions, as indicated by the negative linear model coefficient and mean SHAP value for this descriptor. This trend is physically intuitive – expanded nonpolar surface area introduces nonpolar domains that disrupt the continuity of ionic transport pathways. To further interpret this trend, we examined structural correlations and found that the cation S5 descriptor scales with the length of the longest alkyl chain in the cation (Fig. 3b). This observed trend is consistent with past experimental studies showing that increasing the alkyl chain length of the cation tends to lead to a decrease in the ionicity and molar conductivity of ILs.13,17,18
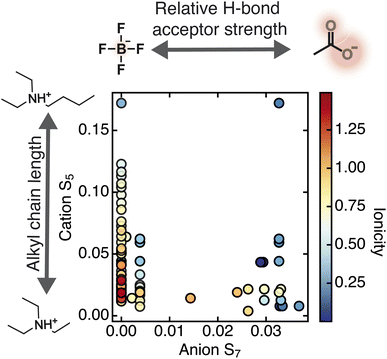
After the cation nonpolar surface area, the anion hydrogen bond acceptor strength also emerged as an important feature. The anion S7 descriptor, which measures the area of the sigma profile in the negative polarity region corresponding to stronger hydrogen bond acceptors, frequently appeared among the top-ranked features (SI S7). To further interpret this observation, we examined trends within specific cation families. We found that for the ammonium cation family, ILs with higher anion S7 values and lower cation S5 values tended to have a lower ionicity (Fig. 4, SI S7). This result suggests that for ammonium ILs, strongly hydrogen bond-accepting anions may promote ion association and correlated motion, which could enhance deviations from Nernst–Einstein behavior. Since the dataset contains more ammonium ILs with high rather than low ionicity, the generalizability of this trend remains uncertain (SI S7). Additionally, there exists some unexplored parameter space with regards to the anions paired with ammonium ILs, as highlighted by the fewer number of ILs with anion S7 values between approximately 0.01 and 0.02 (Fig. 4, SI S7). Designing ILs with anions with these intermediate hydrogen bond-accepting strengths would provide more insight into this observed trend. Although this relationship between ionicity and the cation S5 and anion S7 descriptors was not observed for imidazolium-based ILs (SI S7), future work studying other cation families will be needed to determine whether similar interactions emerge across the chemical space of ILs.
While the Si descriptors capture how much the ion surface charge falls into nonpolar or polar regions, they may miss global features of the charge distribution. To evaluate the influence of such descriptors on ionicity predictions, we trained models using sigma profile moments (Mi) and charge localization metrics (WAPS/WANS) (SI S3). The moments relate to the ion surface area (M0), ion charge (M1), ion polarity (M2), and sigma profile skewness (M3).36,40 The cation and anion M2 descriptors, in particular, consistently appeared as important predictors. Furthermore, based on the sign of the linear model coefficients and SHAP values for these descriptors, a greater cation or anion polarity (larger M2 values) may lead to lower ionicity predictions. This result is consistent with the idea that ILs with highly polar ions may have stronger Lewis acid-base interactions and greater ion–ion correlations. Stronger ion correlations can lead to a greater deviation of the IL ionic conductivity from ideal Nernst–Einstein behavior and a lower ionicity.17,18
The anion WAPS and cation WANS descriptors were used in this study to quantify the extent of charge localization,34,35 and the cation WANS descriptor occasionally emerged among the top features (SI S7, Fig. S17). This result suggests that cation charge localization may contribute to deviations from Nernst–Einstein behavior. Interestingly, although both the anion WAPS and cation WANS descriptors have been proposed as a measure of the charge localization on the ion, and were hypothesized to be important in predicting ionicity, only the cation WANS descriptor appears among the top influential features. However, there could be a nonlinear relationship between these descriptors and ionicity, and such a relationship would not be captured by the linear model.
3.3 Additional feature relationships and functional group analysis
Temperature was among the top features in the feature importance rankings, consistent with the well known impact on ionic conductivity, though its influence was less pronounced for ionicity (SI S7). This result is expected, as the temperature dependence of conductivity is often captured by the Vogel–Tammann–Fulcher relationship, which links log(conductivity) to inverse temperature.47,48 In addition, temperature is directly coupled to diffusion and viscosity, both of which correlated with ionic conductivity.49,50 Compared to the conductivity prediction models, temperature was less influential when predicting ionicity. This weaker effect can be rationalized by the definition of ionicity as a ratio, where both the measured molar conductivity and the Nernst–Einstein conductivity vary with temperature, hence much of the dependence cancels out (eqn (2)).To evaluate collinearity, variance inflation factors (VIFs) for the Si (area under the sigma profile curve) and the Mi, WAPS, WANS descriptors that had nonzero coefficients in the optimal linear L1 models used to predict ionicity or molar ionic conductivity were calculated (SI S9). In this analysis, we considered VIF values above 10 to indicate a large amount of collinearity.51 There did not appear to be a large amount of collinearity present in the set of descriptors used in the optimal linear L1 models for predicting molar ionic conductivity (i.e. no VIF values above 10). However, there was a large amount of collinearity among the descriptors that were used in the optimal linear L1 models for predicting ionicity. Specifically, the anion S6, anion S7 (Si descriptor set), and the cation M3, cation M2, and cation MHB,acceptor (Mi, WAPS, WANS descriptor set) descriptors had VIF values above 10. The presence of collinearity indicates that there may be additional influential descriptors in predicting ionicity that were not identified in the feature importance analysis conducted in this study. Future work is recommended to investigate the impact of the removal of descriptors based on their VIF values on ionicity model performance and feature importance rankings.
To probe whether specific chemistries were common in high (above 1.1) or low (below 0.9) ionicity ILs, we analyzed functional group combinations across the dataset (SI S8). A Tanimoto similarity test using functional group fingerprints showed that ILs with high ionicity (0.6 ± 0.2) and low ionicity (0.5 ± 0.2) were no more structurally similar to each other than the full dataset as a whole (0.6 ± 0.2).52 However, we did observe that ILs with a carboxylate anion paired with a nitrogen-based cation nearly always had a low ionicity. As most ILs in this dataset belong to the imidazolium family and thus have a nitrogen-containing cation, further analysis is needed to determine whether this effect extends to other cation families.
3.4 ML-predicted ionicity improves conductivity estimates
We next evaluated whether ML models trained to predict ionicity could be used as correction factors to improve conductivity estimates. In this approach, the ML-predicted ionicity was multiplied by the Nernst–Einstein conductivity for each IL, effectively using the model to correct the baseline. As shown in Fig. 5, this approach led to an improved performance on the test set compared to models trained to directly predict molar conductivity.There are at least two potential reasons for this improved performance. First, the Nernst–Einstein equation encodes known physics relating conductivity to ion size, charge, and viscosity (eqn (3)), so incorporating it provides a strong functional form for the baseline. By predicting deviations (ionicity) rather than conductivity itself, the ML model would only need to learn the residual effects rather than the full transport variable. Another reason is that the distribution of ionicity values is much narrower than that for molar conductivity (SI S5), which may make the regression task simpler. For instance, dummy regressors that output the mean of the training data have a relatively greater accuracy when predicting ionicity compared to molar conductivity (Fig. 2, SI S6.1). Taken together, these points suggest that predicting ionicity can simplify the regression problem and highlight residual physical effects, although the extent of the accuracy gain remains uncertain.
Beyond improved accuracy, using ionicity predictions as a correction factor provides greater interpretability. ML-predicted ionicity provides a direct measure of how well the Nernst–Einstein equation captures ion transport in a given IL and whether it overestimates or underestimates the ionic conductivity. Ionicity values greater than one suggest super-hydrodynamic transport and the presence of alternative mechanisms.1,13 On the other hand, ionicity values below one could indicate a greater degree of ion-pairing or ion–ion interactions.17,18 Thus, the ML models not only correct the Nernst–Einstein baseline but also classify ILs into physically meaningful transport regimes. From a design standpoint, this can enable a practical workflow: calculate the Nernst–Einstein conductivity from viscosity measurements and estimated ionic radii, apply an ML-predicted ionicity correction, and obtain a more accurate and interpretable estimate.
Additional results relating to ML model performance, feature importance rankings, along with dataset analysis can be found in the SI S6–S8.
4 Conclusions
In this work, we systematically studied the influence of different computationally derived features on predicting ionicity. We trained ML models on sigma profile-based features of IL cations and anions and found that they predict ionicity with accuracies comparable to models trained using cheminformatic descriptor sets, while providing clearer physical interpretability. Importantly, ionicity predictions from our ML models can be combined with Nernst–Einstein conductivities to provide corrected conductivity estimates, which demonstrates a practical way to integrate physics-based baselines with machine learning.Feature importance analysis highlighted several interpretable relationships. Among the Si descriptors, the cation S5 descriptor (relating to nonpolar surface area) emerged as the dominant predictor of ionicity and recapitulated the well-established experimental trend that longer alkyl chains reduce ionic conductivity. The anion S7 descriptor (corresponding to hydrogen bond acceptor strength) also appeared influential, particularly within ammonium-based ILs where higher S7 values were associated with reduced ionicity. Collectively, these results demonstrate that sigma profile descriptors serve as chemically intuitive tools for rationalizing ionic transport.
There still remains some limitations in our analysis. Linear models capture only linear structure-property trends, and nonlinear effects are likely underrepresented. The dummy regressor results also highlight that part of the apparent performance gain may arise from the narrower statistical distribution of ionicity rather than from richer structure-property learning. Future work should explore more flexible yet interpretable approaches, such as generalized additive models (GAMs), to capture nonlinearities while preserving insight.53–55 In addition, further analysis of descriptor interdependencies can improve the robustness of the feature importance analysis and may lead to the identification of additional features that are influential in predicting ionicity. Expanding the dataset to include a broader range of cation/anion chemistries will also be critical for testing the generality of the trends observed here. Lastly, future studies can use the same ML approaches trained on simulation obtained viscosity or diffusivity (similar to the approaches of Greathouse and co-workers49), and subsequently use these models to predict Nernst–Einstein conductivities. In this manner, ionic conductivity predictions can be made with any experimental data input.
Overall, this study demonstrates that sigma profile-based ML models can provide both predictive and mechanistic insight into ionicity and conductivity of ILs. By linking cation nonpolar surface area, anion hydrogen bond acceptor strength, and ion polarity to deviations from Nernst–Einstein behavior, these models provide a foundation for more interpretable, physics-informed screening of ILs.
Author contributions
A. S. developed the method framework, performed the data analysis, and wrote the initial draft. L. T. M. H. contributed to the data analysis. S. Y. supervised the project, provided input on analysis, and revised the manuscript. All authors reviewed and approved the final version.Conflicts of interest
There are no conflicts to declare.Data availability
This study was carried out using publicly available data from NIST ILThermo v2.0 at https://ilthermo.boulder.nist.gov (access date 2/25/2025). All code needed to reproduce the analysis is available in the GitHub repository https://github.com/YueGroup/IL-Ionicity-Paper. An archived version of the repository associated with this publication is available via Zenodo at: https://doi.org/10.5281/zenodo.18765223.Supplementary information (SI): additional details regarding the methods used and supplementary figures. See DOI: https://doi.org/10.1039/d5dd00414d.
Acknowledgements
This material is based upon work supported by the National Science Foundation Graduate Research Fellowship Program under Grant No. DGE-2139899. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation. This work was performed using compute resources from the Cornell University Center for Advanced Computing (CAC).Notes and references
- J. E. Umaña, R. K. Cashen, V. M. Zavala and M. A. Gebbie, Digital Discovery, 2025, 4, 1423–1436 RSC
.
- J. Kalhoff, G. G. Eshetu, D. Bresser and S. Passerini, ChemSusChem, 2015, 8, 2154–2175 CrossRef CAS PubMed
.
- R. Tiwari, D. Kumar, D. K. Verma, K. Parwati, P. Ranjan, R. Rai, S. Krishnamoorthi and R. Khan, J. Energy Storage, 2024, 81, 110361 CrossRef CAS
.
- X. Tian, Y. Yi, B. Fang, P. Yang, T. Wang, P. Liu, L. Qu, M. Li and S. Zhang, Chem. Mater., 2020, 32, 9821–9848 CrossRef CAS
.
- N. V. Plechkova and K. R. Seddon, Chem. Soc. Rev., 2008, 37, 123–150 RSC
.
- T. Lemaoui, T. Eid, A. S. Darwish, H. A. Arafat, F. Banat and I. AlNashef, Mater. Sci. Eng. R Rep., 2024, 159, 100798 CrossRef
.
- M. Mohan, K. D. Jetti, S. Guggilam, M. D. Smith, M. K. Kidder and J. C. Smith, ACS Sustain. Chem. Eng., 2024, 12, 7040–7054 CrossRef CAS
.
- P. Dhakal and J. K. Shah, Fluid Phase Equilib., 2021, 549, 113208 CrossRef CAS
.
- P. Dhakal and J. K. Shah, Mol. Syst. Des. Eng., 2022, 7, 1344–1353 RSC
.
- O. Nordness, P. Kelkar, Y. Lyu, M. Baldea, M. A. Stadtherr and J. F. Brennecke, J. Mol. Liq., 2021, 334, 116019 CrossRef CAS
.
- K. Low, R. Kobayashi and E. I. Izgorodina, J. Chem. Phys., 2020, 153, 104101 CrossRef CAS PubMed
.
- Z. Chen, J. Chen, Y. Qiu, J. Cheng, L. Chen, Z. Qi and Z. Song, ACS Sustain. Chem. Eng., 2024, 12, 6648–6658 CrossRef CAS
.
- R. K. Cashen, M. M. Donoghue, A. J. Schmeiser and M. A. Gebbie, J. Phys. Chem. B, 2022, 126, 6039–6051 CrossRef CAS PubMed
.
- C. Song, C. Wang, F. Fang, G. Zhou, Z. Dai and Z. Yang, J. Chem. Eng. Data, 2024, 69, 4310–4319 CrossRef CAS
.
- R. Datta, R. Ramprasad and S. Venkatram, J. Chem. Phys., 2022, 156, 214505 CrossRef CAS PubMed
.
- M. Kar, N. V. Plechkova, K. R. Seddon, J. M. Pringle and D. R. MacFarlane, Aust. J. Chem., 2019, 72, 3 CrossRef CAS
.
- O. Nordness and J. F. Brennecke, Chem. Rev., 2020, 120, 12873–12902 CrossRef CAS PubMed
.
- M. Watanabe, Electrochemistry, 2016, 84, 642–653 CrossRef CAS
.
- F. Philippi, D. Rauber, M. Springborg and R. Hempelmann, J. Phys. Chem., 2019, 123, 851–861 CrossRef CAS PubMed
.
- F. Philippi, K. Goloviznina, Z. Gong, S. Gehrke, B. Kirchner, A. A. H. Pádua and P. A. Hunt, Phys. Chem. Chem. Phys., 2022, 24, 3144–3162 RSC
.
- H. K. Kashyap, H. V. R. Annapureddy, F. O. Raineri and C. J. Margulis, J. Phys. Chem. B, 2011, 115, 13212–13221 CrossRef CAS PubMed
.
- O. Hollóczki, F. Malberg, T. Welton and B. Kirchner, Phys. Chem. Chem. Phys., 2014, 16, 16880–16890 RSC
.
- Q. Dong, C. D. Muzny, A. Kazakov, V. Diky, J. W. Magee, J. A. Widegren, R. D. Chirico, K. N. Marsh and M. Frenkel, J. Chem. Eng. Data, 2007, 52, 1151–1159 CrossRef CAS
.
- A. Kazakov, J. W. Magee, R. D. Chirico, E. Paulechka, V. Diky, C. Muzny, K. Kroenlein and M. Frenkel, NIST Standard Reference Database 147, NIST Ionic Liquids Database – (ILThermo), 2024, https://ilthermo.boulder.nist.gov.
- K. Ueno, H. Tokuda and M. Watanabe, Phys. Chem. Chem. Phys., 2010, 12, 1649 RSC
.
- W. Xu, E. I. Cooper and C. A. Angell, J. Phys. Chem. B, 2003, 107, 6170–6178 CrossRef CAS
.
- D. M. Makarov, Y. A. Fadeeva and L. E. Shmukler, J. Mol. Liq., 2023, 391, 123323 CrossRef CAS
.
- C. Schreiner, S. Zugmann, R. Hartl and H. J. Gores, J. Chem. Eng. Data, 2010, 55, 1784–1788 CrossRef CAS
.
- A. Klamt, J. Phys. Chem., 1995, 99, 2224–2235 CrossRef CAS
.
- E. Mullins, R. Oldland, Y. A. Liu, S. Wang, S. I. Sandler, C.-C. Chen, M. Zwolak and K. C. Seavey, Ind. Eng. Chem. Res., 2006, 45, 4389–4415 CrossRef CAS
.
- D. O. Abranches, Y. Zhang, E. J. Maginn and Y. J. Colón, Chem. Commun., 2022, 58, 5630–5633 RSC
.
- J. Palomar, J. S. Torrecilla, V. R. Ferro and F. Rodríguez, Ind. Eng. Chem. Res., 2008, 47, 4523–4532 CrossRef CAS
.
- J. Palomar, J. S. Torrecilla, J. Lemus, V. R. Ferro and F. Rodríguez, Phys. Chem. Chem. Phys., 2010, 12, 1991 RSC
.
- K. Kaupmees, I. Kaljurand and I. Leito, J. Phys. Chem., 2010, 114, 11788–11793 CrossRef CAS PubMed
.
- K. Kaupmees, I. Kaljurand and I. Leito, J. Solution Chem., 2014, 43, 1270–1281 CrossRef CAS
.
-
A. Klamt, COSMO-RS: from quantum chemistry to fluid phase thermodynamics and drug design, Elsevier, 1st edn, Amsterdam, 2005 Search PubMed
.
-
I. Chernyshov, ILThermoPy, 2023, https://github.com/IvanChernyshov/ILThermoPy Search PubMed
.
- O. Nordness, L. D. Simoni, M. A. Stadtherr and J. F. Brennecke, J. Phys. Chem. B, 2019, 123, 1348–1358 CrossRef CAS PubMed
.
- G. Landrum, P. Tosco, B. Kelley, R. Rodriguez, D. Cosgrove, R. Vianello, S. Riniker, P. Gedeck, G. Jones, NadineSchneider, E. Kawashima, D. Nealschneider, A. Dalke, M. Swain, B. Cole, S. Turk, A. Savelev, A. Vaucher, M. Wójcikowski, I. Take, V. F. Scalfani, R. Walker, K. Ujihara, D. Probst, A. Pahl, J. Lehtivarjo, F. Bérenger, 2024, DOI:10.5281/zenodo.13469390.
- S. Müller, T. Nevolianis, M. Garcia-Ratés, C. Riplinger, K. Leonhard and I. Smirnova, Fluid Phase Equilib., 2025, 589, 114250 CrossRef
.
- F. Neese, WIREs Comput. Mol. Sci., 2022, 12, e1606 CrossRef
.
- A. Y.-T. Wang, R. J. Murdock, S. K. Kauwe, A. O. Oliynyk, A. Gurlo, J. Brgoch, K. A. Persson and T. D. Sparks, Chem. Mater., 2020, 32, 4954–4965 CrossRef CAS
.
-
T. Chen and C. Guestrin, Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco California USA, 2016, pp. 785–794 Search PubMed
.
- F. Pedregosa, G. Varoquaux, A. Gramfort, V. Michel, B. Thirion, O. Grisel, M. Blondel, P. Prettenhofer, R. Weiss, V. Dubourg, J. Vanderplas, A. Passos, D. Cournapeau, M. Brucher, M. Perrot and E. Duchesnay, J. Mach. Learn. Res., 2011, 12, 2825–2830 Search PubMed
.
- S. M. Lundberg and S.-I. Lee, Adv. Neural Inf. Process. Syst., 2017, 30 Search PubMed
.
- P. Labute, J. Mol. Graph. Model., 2000, 18, 464–477 CrossRef CAS PubMed
.
- J. Vila, P. Ginés, J. Pico, C. Franjo, E. Jiménez, L. Varela and O. Cabeza, Fluid Phase Equilib., 2006, 242, 141–146 CrossRef CAS
.
- X. Wang, Y. Chi and T. Mu, J. Mol. Liq., 2014, 193, 262–266 CrossRef CAS
.
- N. S. Bobbitt, J. P. Allers, J. A. Harvey, D. Poe, J. D. Wemhoner, J. Keth and J. A. Greathouse, Mol. Syst. Des. Eng., 2023, 8, 1257–1274 RSC
.
- H. Liu and E. Maginn, J. Chem. Phys., 2011, 135, 124507 CrossRef PubMed
.
-
G. James, D. Witten, T. Hastie, R. Tibshirani and J. Taylor, An Introduction to Statistical Learning: with Applications in Python, Springer International Publishing, Cham, 2023 Search PubMed
.
-
T. T. Tanimoto, An elementary mathematical theory of classification and prediction, 1958 Search PubMed
.
-
Y. Lou, R. Caruana and J. Gehrke, Proceedings of the 18th ACM SIGKDD international conference on Knowledge discovery and data mining, Beijing China, 2012, pp. 150–158 Search PubMed
.
- P. Zschech, S. Weinzierl, N. Hambauer, S. Zilker and M. Kraus, GAM(e) changer or not? An evaluation of interpretable machine learning models based on additive model constraints, 2022, https://arxiv.org/abs/2204.09123.
- J. A. Esterhuizen, B. R. Goldsmith and S. Linic, Chem, 2020, 6, 3100–3117 CAS
.
| This journal is © The Royal Society of Chemistry 2026 |