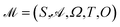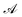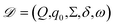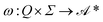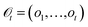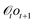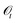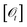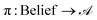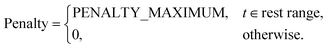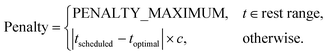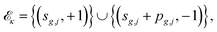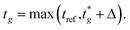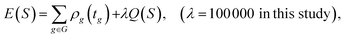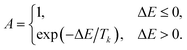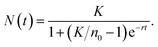Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
GEMS: a deterministic finite automaton framework for adaptive laboratory automation
Yuya
Tahara-Arai
 *ab,
Akari
Kato
*ab,
Akari
Kato
 c,
Koji
Ochiai
c,
Koji
Ochiai
 d,
Kazuya
Azumi
d,
Kazuya
Azumi
 d,
Koichi
Takahashi
d,
Genki N.
Kanda
*ef and
Haruka
Ozaki
d,
Koichi
Takahashi
d,
Genki N.
Kanda
*ef and
Haruka
Ozaki
 *ab
*ab
aLaboratory of Bioinformatics, University of Tsukuba, 1-1-1 Tennōdai, Tsukuba, Ibaraki 305-8577, Japan. E-mail: arai.yuya.qa@alumni.tsukuba.ac.jp
bLaboratory for AI Biology, RIKEN Center for Biosystems Dynamics Research, 6-7-1 Minatojima Minamimachi, Chuo-ku, Kobe, Hyogo, Japan. E-mail: ai-biology@ml.riken.jp
cResearch DX Foundation Team, RIKEN Data and Computational Sciences Integration Research Program, 6-7-1 Minatojima Minamimachi, Chuo-ku, Kobe, Hyogo, Japan
dLaboratory for Biologically Inspired Computing, RIKEN Center for Biosystems Dynamics Research, 6-7-1 Minatojima Minamimachi, Chuo-ku, Kobe, Hyogo, Japan
eMedical Research Laboratory, Institute of Integrated Research, Institute of Science Tokyo, 1-5-45 Yushima, Bunkyo-ku, Tokyo, 113-8519, Japan. E-mail: genki.kanda@tmd.ac.jp
fRobotics Innovation Center, Research Infrastructure Management Center, Institute of Science Tokyo, Japan
First published on 10th March 2026
Abstract
Laboratory automation increasingly requires handling complex, condition-dependent protocols that combine sequential, branching, and iterative operations. Many systems use task-oriented models, where control proceeds task to task through a predefined list. These are effective for linear, static protocols but are poorly suited to adapting to changing sample conditions or representing loops and conditional branches. We introduce the General Experimental Management System (GEMS), which instead adopts a sample-centred approach, progressing state to state, with each state defining both the operations to perform and the rules for transitioning based on observations. By formalising every experimental protocol as a partially observable Markov decision process (POMDP) and expressing its deterministic execution logic as a deterministic finite automaton (DFA), GEMS can represent heterogeneous workflow structures within a single, coherent framework and enable direct compilation into instrument-executable workflows. Its architecture includes a hierarchical experiment model, a penalty-aware scheduler combining a greedy baseline with simulated annealing refinement, and a file-based interface for instrument-agnostic control. We demonstrate GEMS in two contrasting cases: (i) fully automated Bayesian optimisation of liquid mixtures using a pipetting robot and imaging, and (ii) dynamic, long-term scheduling of multiple mammalian cell cultures executed by a LabDroid robot with autonomous imaging, passaging, medium exchange, and fault recovery. In both, GEMS maintained protocol constraints while adapting schedules in real time, showing that a state-to-state, sample-centred model provides an abstraction that maintains protocol constraints while adapting schedules in real time across heterogeneous workflows.
Introduction
Laboratory automation is undergoing rapid expansion across biology, chemistry, and materials science, driven by the demand for higher efficiency in research and development.1–6 While these technologies have enabled higher throughput and reproducibility, most automated workflows are described as non-adaptive execution plans, such as fixed step-by-step scripts or static dependency graphs, which cannot incorporate new observations once execution begins. Such static workflows are poorly suited to experiments that require conditional branching, iterative loops, or dynamic rescheduling in response to ongoing measurements. In practice, however, so-called “care” tasks still fall to human operators, posing a major bottleneck to scaling complex workflows.7 This limitation becomes particularly acute in long-term or condition-dependent studies, where the true state of a sample cannot be observed directly, and decisions must be made from partial information. These challenges are further amplified by the emergence and vision of facility-scale cloud laboratories, which are accelerating the demand for automating ever more complex and wide-ranging experimental protocols.8,9Existing laboratory automation platforms typically represent experimental workflows within one of four broad paradigms. First, timeline-based systems, such as Clarity LIMS and Autoprotocol, encode protocols as a fixed sequence of steps, making them straightforward to read but inherently unsuitable for conditional branching.10,11 In this work, we treat Autoprotocol as a timeline-style description standard rather than a prescriptive programming language; accordingly, our comparisons focus on representational scope (fixed step lists), not on execution semantics.11 Second, static directed acyclic graph (DAG) schedulers, exemplified by Green Button Go and SAMI EX, capture task dependencies in a graphical form and generate a single optimised schedule prior to execution; however, the graph cannot be modified during execution.12–15 Third, dynamic flow-chart frameworks—including AlabOS, ChemOS, GLAS, and COPE—maintain tasks in a queue or database and determine the subsequent action at run time, thereby supporting closed-loop optimisation, explicit loops, and adaptive resource allocation.16–22 In COPE, for example, the Logic Builder allows loops, producing directed graphs that may contain cycles. Finally, domain-specific languages such as χDL and IvoryOS treat a protocol as source code: χDL compiles declarative scripts into hardware-independent instructions, while IvoryOS assembles Python workflows from a visual canvas; both support loops and conditional logic but delegate most scheduling decisions to the execution engine.23,24
Most existing frameworks in these categories adopt a task-oriented model, in which control moves task to task through a predefined list. While effective for linear protocols and static resource allocation, this perspective makes it difficult to track the evolving condition of individual samples or to express loops and conditional branches without regenerating the entire task list. From a theoretical standpoint, such limitations can be understood by modelling experimental procedures as partially observable Markov decision processes (POMDPs),25–27 in which an unobserved latent sample state is inferred from measurements and subsequent actions are chosen according to an evolving belief state. When the policy is deterministic and the observation set is finite, the decision process can be expressed as a deterministic finite automaton (DFA, Mealy automaton).28–30 This minimal structure makes state transitions explicit, supports formal verification, and can be compiled directly into instrument-executable workflows. Framing protocols in this way suggests a natural alternative to the task-to-task paradigm: a sample-centred model in which the primary unit is the state of each sample, and control moves state to state. Each state encapsulates both the operations to be performed and the rules for transitioning based on observations. This shift aligns naturally with condition-dependent protocols and unifies short, deterministic sequences with long-running, branching workflows in a single abstraction.
Related work spanning timeline lists, static DAG schedulers, dynamic flow-chart frameworks, and DSLs has advanced throughput and, in some cases, closed-loop control; however, task-oriented models seldom treat per-sample state and progression rules as first-class control units, and scheduling/timing often remains entangled with control logic. To address this gap, we use a sample-centred deterministic finite automaton (DFA, Mealy automaton) formulation that separates state progression from task emission while delegating timing to a penalty-aware scheduler. Task-based controllers that support branching/looping, mid-run insertion/deletion, parallelism and rescheduling have been reported (e.g., Jensen and colleagues31). Other works explicitly articulate task–sample interactions (e.g., Lapkin & Kraft;32 Gregoire & Stein33). Our approach complements these directions by placing a minimal per-sample Mealy automaton at the core of control, separating progression (state transitions) from task emission while delegating timing/resource allocation to a penalty-aware scheduler. This combination unifies short deterministic sequences and long, branch-rich protocols within a single representation and supports direct compilation to instrument-executable workflows. Our contributions are: (i) a constructive mapping from protocol policies to a machine-independent Mealy automaton; (ii) a state-centred experiment model that decouples transition rules from task emission and compiles to instrument-executable tasks; and (iii) a penalty-based scheduler that respects time windows and enables on-the-fly rescheduling.
Here we present GEMS (General Experimental Management System), a general-purpose workflow engine that represents experimental protocols as DFAs derived from POMDPs. GEMS integrates (i) state-based protocol definition, (ii) instrument-agnostic scheduling with penalty-aware optimisation, and (iii) observations that store both results and metadata for decision-making. To demonstrate the generality of this framework under diverse temporal and structural constraints, we apply GEMS in two contrasting case studies: (1) sequential parameter optimisation of liquid mixtures guided by Bayesian design, and (2) dynamic scheduling of parallel mammalian cell cultures with automated imaging and passaging. Together, these examples show that a state-to-state, sample-centred model can support robust, fully automated laboratory operation across both short deterministic sequences and long, branch-rich workflows.
Experimental
Symbols used in the text
POMDP to DFA: a sufficient representation for protocol control logic
GEMS formalises experimental protocols as partially observable Markov decision processes (POMDPs) (Fig. 1A). Latent “sample states” are the complete system states, laboratory operations are actions, and measurement outcomes are observations (Fig. 1B). When a protocol uses a deterministic policy acting on a finite observation alphabet—i.e., a fully specified, rule-based procedure—the induced decision logic is captured by a deterministic finite automaton with outputs (a Mealy automaton) (Fig. 1C). This section states and proves the sufficiency of such an automaton for representing protocol control logic.used under a deterministic, rule-based policy
 that maps any finite observation history to a (possibly empty) finite sequence of laboratory actions. We separate transition rules (stage progression) from task-generation rules (emitted operations), matching GEMS's transition function g and task function f.
that maps any finite observation history to a (possibly empty) finite sequence of laboratory actions. We separate transition rules (stage progression) from task-generation rules (emitted operations), matching GEMS's transition function g and task function f.
that reproduces exactly the observation–action behaviour of.

 iff for all w ∈ Σ*,
iff for all w ∈ Σ*,  . Let
. Let  be the set of equivalence classes and q0 = [ε]. Define
be the set of equivalence classes and q0 = [ε]. DefineBy construction, for any history  and next symbol o, the action sequence prescribed by π equals the output emitted by.
and next symbol o, the action sequence prescribed by π equals the output emitted by. 
 This makes states and progression conditions explicit while keeping task generation independent of transitions, and supports formal checks and direct compilation to instrument-executable workflows.
This makes states and progression conditions explicit while keeping task generation independent of transitions, and supports formal checks and direct compilation to instrument-executable workflows.
Software architecture of GEMS and the definition of an experiment
GEMS models an experiment as a POMDP and represents discrete state transitions using a DFA. Because the latent state of a sample cannot be observed directly, each decision is based on a belief state inferred from the observations.The structure of GEMS comprises:
• Lab: the top-level collection. Each experiment runs independently and specifies a directory for saving its results.
• Experiment: the digital representation of a protocol. Internally, this is a DFA composed of a set of states, a transition function (edges), and a observations that record all outcomes (Fig. 2A).
• Machines: physical instruments available in the laboratory. Each record stores a textual description and the associated machine type defining the operations that the instrument can perform.
• State: a stage of the experiment. Each state contains (i) a task function that generates operations for that stage, and (ii) a transition function that determines the next state (Fig. 2A).
• Task: a data structure specifying the operation to be performed, its optimal start time, a penalty function for quantifying the cost of deviation from the optimal start time, and the acceptable machine type (Fig. 2B). In GEMS, all task start times are expressed on a user-defined integer timeline (relative time), not wall-clock timestamps.
• Task group: an ordered sequence of tasks with fixed intervals. Once the start time of the first task is determined, the remaining tasks are scheduled automatically.
• Task function (user-defined in Python): a function that receives the observations and emits the operations required in the current state (Fig. 2A).
• Transition function (user-defined in Python): a function that receives the observations and returns the identifier of the next state (Fig. 2A).
• Penalty function: a numerical cost function evaluating the deviation between the optimal and scheduled times.
• File-based user interface: instead of an interactive graphical user interface, GEMS relies on the file system. At start-up, the plugin manager creates the directories experimental_setting and mode and polls them for changes. Whenever a plugin file or a command file is added or modified, the corresponding method is executed and the file is removed thereafter. This design allows both external drivers and human users to control the workflow using only a text editor.
• Dry-run simulation for capacity planning: the dry-run mode executes the DFA-driven transition function and task emission without sending commands to hardware. Users can vary the number of instruments per type and adjust penalty settings to obtain feasibility, predicted lateness and indicative utilisation before execution. This supports instrument-count selection and maintenance-window placement for larger deployments, while the per-experiment DFA remains unchanged.
Fig. 2 schematically depicts the relationships between these elements: task contains a penalty function for cost calculation during scheduling, and an executable machine type. State contains a task function and a transition function. Experiment comprises a DFA of states and the observations. Lab manages multiple independent experiments and the available machines. Here, we use independence to mean independence of decision inputs: each experiment's transition and task functions consult only its own observations and current state, not those of other experiments.
Fig. 3 outlines the execution flow: state updates in the transition function, task updates in the task function, and schedule updates in the task scheduler (Fig. 2B).
Penalty functions
Each task is assigned a penalty function that converts any deviation from its optimal start time into a scalar cost. These penalties are used by the scheduler to evaluate and optimise task placement. Five variants are currently implemented:NonePenalty Returns a constant value of 0; suitable when the timing of the task is completely flexible.
LinearPenalty Takes a single parameter, penalty_coefficient c. The penalty is
| Penalty = |tscheduled − toptimal| × c |
LinearWithRangePenalty Parameters: lower, lower_coefficient, upper, and upper_coefficient. Let Δ = tscheduled − toptimal. The penalty is
CyclicalRestPenalty Parameters: cycle_start_time, cycle_duration, and rest_time_ranges. With
| t = (tscheduled − cycle_start_time) mod cycle_duration, |
The helper function adjust_time_candidate_to_rest_range shifts a candidate time forward to the next active period.
CyclicalRestPenaltyWithLinear Extends CyclicalRestPenalty by adding a linear term when outside rest ranges:
Unified representation of protocol structures in GEMS
By formalising every experimental protocol as a DFA, GEMS is able to represent heterogeneous workflow structures within a single, coherent framework:• Linear protocols (Fig. 1D, left): each operation is represented by a distinct state and the sample progresses through these states exactly once in a fixed order, with a well-defined terminal state.
• Terminatable protocols with a loop (Fig. 1D, middle): transitions may return to a previous state, so that the sample can revisit a subset of states multiple times before eventually reaching a terminal state, as in protocols with conditional repeats.
• Open-ended protocols without a terminal state (Fig. 1D, right): the state graph contains no designated terminal state, allowing continuous or indefinitely repeated procedures such as long-term monitoring.
Scheduling algorithm
Let G = {g1, g2, …, gN} be the set of task groups and the set of machines. Each task j inside a task group g is described by its processing time pg,j, the interval dg,j from the previous task, and the required machine type µg,j.
the set of machines. Each task j inside a task group g is described by its processing time pg,j, the interval dg,j from the previous task, and the required machine type µg,j.
The scheduling procedure consists of two steps:
1. A baseline greedy scheduler that produces an initial feasible schedule for each task group to avoid machine conflicts.
2. A simulated annealing (SA) refinement that adjusts start times to improve the schedule while preserving the task order within each group.
Greedy baseline
| sg,1 = tg, sg,j = sg,j−1 + pg,j−1 + dg,j (j > 1), |
where (t, + 1) denotes the start of a task and (t, − 1) its completion. Sort events by time, giving precedence to task starts in the case of ties, and sweep a counter oκ. A conflict occurs if and only if oκ exceeds the available number of machines cκ.
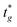 ), where
), where 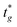 is the user-specified optimal start time for group g. For each group, iteratively try integer offsets Δ ∈ {0, 1, − 1, 2, − 2, … } until a conflict-free placement is found, and assign the group's start time as
is the user-specified optimal start time for group g. For each group, iteratively try integer offsets Δ ∈ {0, 1, − 1, 2, − 2, … } until a conflict-free placement is found, and assign the group's start time asHere, tref denotes the current reference time on the same user-defined integer timeline; only times ≥ tref are eligible for placement. For optimisation we work in a reference-centred coordinate (e.g., offsets such as Δt = tg − tref or
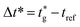 ) and then map results back by adding tref; wall-clock conversion is deferred to the display/dispatch stage.
) and then map results back by adding tref; wall-clock conversion is deferred to the display/dispatch stage.
| S = {tg|g ∈ G} |
where λ is a large constant that heavily penalises any schedule with machine conflicts.
![[t with combining tilde]](https://www.rsc.org/images/entities/i_char_0074_0303.gif) g = tg + δ, and then wrap it as follows:
g = tg + δ, and then wrap it as follows:Here, the helper function adjust_time_candidate_to_rest_range snaps the candidate start time to the nearest valid slot that is outside any rest period window.
Executing GreedySchedule followed by SimulatedAnnealing produces start times that satisfy machine capacity constraints while minimising deviation from the user-specified optimal start times. In this study, the number of improvement steps (Niter) for the annealing method was fixed at 100, 000. For the formulation in ILP (integer linear programming) and computational complexity analysis of the Greedy baseline, see Appendix A.
RGB colour optimisation of liquid mixtures
We designed a red–green–blue (RGB) colour optimisation task as a toy problem that mirrors parameter search in chemistry and materials science. In this study, the optimisation was carried out over three consecutive experiments (see Fig. 4C).Reagents
• Red acidic solution: 0.50 g citric acid and 0.10 g red food dye in 50 mL water.• Basic clear solution: 0.50 g sodium bicarbonate in 50 mL water.
• Bromothymol blue: 0.04% (w/v).
The initial volume ratio of the three stock solutions (acidic red solution, basic clear solution, and bromothymol blue) was set to 0.2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.3
0.3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.5, and a target RGB value was defined. Batch Bayesian optimisation (q-Expected Improvement policy) was carried out with BoTorch34,35 over three rounds. In each round, eight candidate mixing ratios (conditions) were generated, and each condition was tested in four adjacent wells (four technical replicates), occupying a total of 32 wells on a 96-well plate (Fig. 4E). Dispensing was performed on an Opentrons OT-2 robot (Opentrons Labworks Inc.) in descending order of component ratio to minimise mixing variance; the final volume in each well was 100 µ L. Python scripts in the Opentrons Protocol API format were auto-generated by the OT-2 driver and executed on the instrument.
0.5, and a target RGB value was defined. Batch Bayesian optimisation (q-Expected Improvement policy) was carried out with BoTorch34,35 over three rounds. In each round, eight candidate mixing ratios (conditions) were generated, and each condition was tested in four adjacent wells (four technical replicates), occupying a total of 32 wells on a 96-well plate (Fig. 4E). Dispensing was performed on an Opentrons OT-2 robot (Opentrons Labworks Inc.) in descending order of component ratio to minimise mixing variance; the final volume in each well was 100 µ L. Python scripts in the Opentrons Protocol API format were auto-generated by the OT-2 driver and executed on the instrument.
After dispensing, a Logitech c920n webcam (Logitech c920n; Logitech International S.A.) captured images of the plate. Images were processed on a 14-inch MacBook Pro (Apple, California, US): each well region was cropped, and the average RGB value was computed. A white sheet placed beneath the plate minimised background variation. A target mixture was prepared in all 96 wells at the same ratio, and the RGB values of each well were measured. To reduce variation due to the imaging environment, for each cropped well the Manhattan distance between its measured RGB and the RGB of the corresponding target well was calculated. This Manhattan distance was used as the objective for the next optimisation round. GEMS defined distinct states for mixing, imaging, and evaluation, linked by one-to-one transitions.
Cell-culture experiments
Prior to robotic culture, cells were prepared manually: they were seeded at the A1 well position in six-well plates, washed once with phosphate-buffered saline (PBS) (10010023; Thermo Fisher Scientific; Lot: 2412443]), detached with 0.05% trypsin (25300054; Thermo Fisher Scientific; Lot: 2713076]) by gentle pipetting after 2 min at room temperature, and replated at 1 × 105 and 2 × 105 cells per well. The plates were then transferred to the CO2 incubator in the LabDroid booth unit.
Day −14: plate coating and thawing. Six-well plates were coated with iMatrix-511 (385-07361; Matrixome Inc., Lot: 24B215]) at 0.5 µ g cm−2 in PBS (−) and incubated for at least 60 min at 37 °C with 5% CO2. After aspirating the coating solution, each well received 0.75 mL of hiPSC maintenance medium (StemFit AK03N; Ajinomoto; Lot: 20240527A). Cryovials of frozen hiPSCs were thawed in a 37 °C water bath, transferred into 5 mL maintenance medium containing Rho-kinase inhibitor (final 10 µ M) (Y-27632; 035-24593; FUJIFILM Wako Pure Chemical; Lot: CKK5331), and pelleted by centrifugation at 160×g for 4 min at 22 °C. The supernatant was removed, and cells were resuspended in fresh maintenance medium with 10 µ M inhibitor. After counting, 43
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300–45,000 cells were seeded per well in 1.5 mL medium.
300–45,000 cells were seeded per well in 1.5 mL medium.
Day −13 to day −8: maintenance. On Day −13, the medium was changed to maintenance medium without Rho-kinase inhibitor. From Days −12 to −8, the same medium was refreshed every 24–72 h.
Day −7: Harvest and suspension preparation. Cells were washed once with 2 mL PBS (−), then incubated with 1 mL of 0.5× TrypLE Select CTS (A1285901; Thermo Fisher Scientific, Lot: 2561785]) in 0.5 mM ethylenediaminetetraacetic acid (EDTA)/PBS (−) (13567-84; Nacalai Tesque Inc., Lot: L4G6700]) at 37 °C with 5% CO2 for 10–20 min. Detached cells were gently pipetted into a 50 mL tube, supplemented with 1 mL maintenance medium and 3 mL PBS, and centrifuged at 160×g for 4 min at 22 °C. The pellet was resuspended in 0.75 mL maintenance medium containing 10 µM inhibitor, filtered through a 40 µm cell strainer (Corning, #352340), and counted. Final suspensions were adjusted to 133
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 cells per 20 mL medium in eight 50 mL tubes.
400 cells per 20 mL medium in eight 50 mL tubes.
Meanwhile, eight new six-well plates were coated with iMatrix-511 at 0.5 µg cm−2 (Lot: 24B215]) as described above, incubated for at least 60 min at 37 °C with 5% CO2, and prepared for plating the cell suspensions.
Two cell lines with different characteristics—hiPSCs and HEK293A cells—were used to evaluate the loop structure.
HEK293A. The HEK workflow comprised four states: HEKFirstGetImageAfterPassageState, HEKGetImageState, HEKPassageState, and HEKSamplingState (see Fig. 5B). For clarity, the schematic diagram in Fig. 5C shows only three representative states, omitting HEKFirstGetImageAfterPassageState. Imaging and density estimation (getImage) were performed 24 h after passage and subsequently every 12 h. When a lineage was predicted to reach 0.80 confluence within 12 h, passage was triggered. After two passages, imaging continued every 12 h; samples predicted to reach 0.40 confluence were collected during daytime (10:00 am to 4:00 pm).
hiPSCs. The hiPSCs workflow comprised five states: PassageInitialState, MediumChange1State, MediumChange2State, PassageState, and SamplingState (see Fig. 5B). For clarity, the state diagram in Fig. 5C omits the PassageInitialState and displays only four states. After passage, getImage was followed by a first medium change (MediumChange1) at 24 h and a standard medium change (MediumChange2) at 48 h. The cycle getImage + MediumChange2 was then repeated every 24 h. When a lineage was predicted to reach the target confluence (0.30) within 24 h, plate coating was initiated 1 h in advance, and passage was executed at the predicted time. After two passages, imaging continued every 24 h; samples predicted to reach 0.30 confluence were collected during daytime (10:00 am to 4:00 pm).
Five hiPSCs lineages were started initially; four days later, five HEK lineages were added. All operations were performed by a LabDroid Maholo (Robotic Biology Institute Inc., Japan), using the same setup as described by Ochiai et al.40 Schedules were rebuilt dynamically in GEMS whenever new HEK lineages were introduced or removed. Two hiPSCs plates were lost owing to device failure; the corresponding series were deleted from the observations, and the remaining schedule was updated automatically.
Logistic model for optimal passage timing
To estimate optimal passage times, we implemented a logistic growth model in both the task and transition functions. The observed density yj at time tj is assumed to followThe estimation proceeded as follows:
1. Data grouping—each experimental result was assigned a passage index gj (passage_group). The initial density n0,i for passage i was set to the observation result in that passage.
2. Combined model—
3. Weighted least squares—The weight of observation j was wj = 2−gj. SciPy's curve_fit function was called with:
4. Time-to-target—Given a target density Nt, the time for passage i was calculated as
The helper function calculate_optimal_time_from_df applied this formula to the latest measurements.
Cell density was following the method described by Ochiai et al.40 Multiple microscope images were stitched, the well region was selected, and cell areas Ac were extracted using the Canny algorithm via the Cell Density Calculator. Using the pixel-count function f, density C was defined as
Results
We evaluate GEMS empirically in two settings: (i) sequential RGB colour optimisation of liquid mixtures, and (ii) dynamic, parallel culture of multiple mammalian cell lines. Formal definitions and software architecture are provided in Experimental (§Experimental—POMDP to DFA: a sufficient representation for protocol control logic, §Experimental—Software architecture of GEMS and the definition of an experiment); this section therefore reports quantitative outcomes and operational behaviour.Sequential parameter optimisation expressed with GEMS
Sequential parameter optimisation, widely used in chemistry and materials engineering, iteratively updates mixing ratios to approach a target property. As a case study, we performed an RGB-optimised liquid-mixing experiment controlled by GEMS. Experimental settings and hardware are in Experimental (§RGB colour optimisation of liquid mixtures). For context, the workflow overview, software connections, and DFA mapping are shown in Fig. 4A–C; the remainder of this section presents the outcomes.The best ratio was obtained in Round 3, with a distance of 61.62—a 30% improvement over the best condition (88.13) in the random initial round (Fig. 4D–F). By Round 2, the distance had already been reduced to 72.13. Each optimisation cycle—from proposal generation to execution, image processing, scoring, and scheduling of the next protocol—was completed within 60 min. In total, 96 mixtures (36 unique compositions) were tested, and all steps were executed deterministically under version control. These results demonstrate that integrating GEMS with a Bayesian optimiser enables efficient, fully automated optimisation experiments under routine laboratory conditions.
Dynamic scheduling of parallel cultures of multiple cell lines
Cell culture is a common iterative experiment in biomedical research, requiring regular monitoring of culture state (e.g., cell density), periodic medium exchanges, and passaging to maintain the density within a target range (Fig. 5A). To evaluate GEMS in a long-term, heterogeneous setting, we performed parallel execution of cell-culture protocols for two distinct cell types: human embryonic kidney cells (HEK293A) and human induced pluripotent stem cells (hiPSCs). The hardware configuration is depicted in Fig. 5B; operational rules for both lineages are in Experimental (§Cell culture experiments by GEMS). The two cell lines were implemented in GEMS as state-based protocols encoded as DFAs, enabling condition-dependent operations while preserving protocol structure (Fig. 5C).The experiment timeline is summarised in Fig. 5D. Fig. SF1 (Table ST2) provides an experiment-level verification of the orchestration by visualising the per-lineage state transitions interpreted by GEMS; detailed execution timelines are provided in SI Video S1 (operation-level Gantt) and SI Video S2 (experiment-level Gantt), with the underlying request, scheduled, and actual start timestamps summarised in SI Table ST1. SI Video S3 provides a step-by-step visualisation of the state-transition history. In this run, we deliberately challenged the system by adding new experiments while it was already managing ongoing cultures. We initiated five hiPSC series on Day 0 (14 Nov 2024). While the system was actively controlling these, we introduced a second set by adding five HEK293A series on Day 4 (18 Nov 2024) through the instantiation of new experiment objects. Later in the run, two hiPSC series were lost owing to unrelated incubator failures on 1 Dec and 5 Dec. After removing the affected series, GEMS automatically recomputed the schedule, reallocated machine time, and continued all remaining experiments without interruption. These mid-run additions and removals implicitly stress-tested scalability. GEMS re-optimised schedules on the fly as instrument availability and workload shifted, without modifying the per-experiment DFA.
Over the 36 days run, the system executed ten HEK passages, five HEK samplings, nine hiPSC passages, and three hiPSC samplings (Fig. 5E). At HEK passage, the mean density was 0.82 ± 0.04 (target 0.80); at HEK sampling, 0.39 ± 0.11 (target 0.40). For hiPSCs, the mean density was 0.32 ± 0.06 at passage (target 0.30) and 0.33 ± 0.03 at sampling (target 0.30). Despite differences in growth rates and handling schedules, both cell types consistently reached their targets with minimal overshoot. These results demonstrate that GEMS can autonomously maintain multiple mammalian cell cultures at user-defined density targets, dynamically reschedule tasks in response to the introduction of new experiments as well as unexpected failures, and support robust long-term biological experiments without human intervention.
Discussion
This study demonstrates that a deterministic finite automaton (DFA, Mealy automaton30) can make both state and transition relations explicit, while observations store experimental data together with associated meta-information. By combining these elements, GEMS realises a sample-centred, state-to-state model in which instrument-agnostic scheduling and condition-dependent control are unified within a single framework. In timeline-style description standards such as Autoprotocol and LIMS tools such as Clarity LIMS, protocols are encoded as fixed step lists for readability and interoperability rather than as prescriptive control languages; consequently, branches and loops are typically realised by template expansion or by generating new scripts/workflows.10,11 DAG schedulers such as Green Button Go can visualise resource conflicts, yet loop structures rely on template expansion and the graph size can grow rapidly.12 Dynamic domain-specific languages such as χDL and IvoryOS provide hardware-independent syntax but do not retain a sample's progress on the execution side, leaving branch management to higher-level scripts.23,24 GEMS complements these approaches by using the DFA's self-transitions and explicit state memory to decide the next action without the need for an external orchestrator.Tangible gains of adopting this sample-centred representation are threefold. First, explicit per-sample state memory and progression rules (Mealy automaton) allow the next action to be selected without regenerating global task lists or re-authoring workflows. Second, timing is decoupled from control logic via a penalty-aware scheduler, enabling on-the-fly rescheduling, maintenance windows, and capacity changes without altering the state–transition diagram. Third, operational robustness improves: mid-run insertion or removal of experiments and recovery from failures are handled by updating the active set, after which GEMS automatically rebuilds the schedule (as evidenced in the parallel HEK/hiPSC run, Fig. 5D–E). Throughout, observations maintain a consistent record of data and metadata, preserving downstream analyses as protocols evolve.
The RGB optimisation of a three-component liquid mixture confirmed that GEMS functions effectively as a framework for sequential parameter search. Ratios proposed by batch Bayesian optimisation were passed through a task function directly into OT-2 Python protocols, and the metrics required for the next round were collected automatically. Because the experimental operation, imaging, and analysis stages are separated into individual tasks, changes to the optimisation algorithm or the evaluation function do not require modification of the state-transition diagram. Even when the number of loops or samples increases, the observations preserve data and metadata in a consistent format, ensuring that downstream analyses remain coherent.
In the cell-culture study, hiPSCs and HEK293A cells—differing in growth rate and handling frequency—were managed simultaneously on the same system across multiple lines and passages. Images captured by a LabDroid Maholo were used to update growth curves in real time, and GEMS automatically determined passaging, medium exchange, and sampling times based on the predicted density. Notably, GEMS was able to incorporate additional experiments during an ongoing run—adding a second set of cultures mid-schedule—and re-optimised the entire task allocation without disrupting current operations. When a subset of lines was lost owing to hardware failure, removing the corresponding experiment was sufficient for GEMS to rebuild the remaining schedule and continue the other experiments without interruption. These results highlight that DFA-based management can simplify recovery tasks and enable dynamic adaptation in laboratories where mid-run changes and unexpected sample losses are unavoidable.
Comparison of the RGB and cell-culture case studies reveals two extremes: the former involves a short, deterministic sequence of operations, whereas the latter contains continuous branches and loops triggered by measurements. The fact that a single software framework supports both indicates that the abstraction level of GEMS is broad enough to accommodate diverse sample types and objectives. Because a task function is loosely coupled to its device driver, replacing or adding equipment requires only minimal edits, potentially reducing refurbishment costs. Likewise, optimisation algorithms and image-analysis pipelines, implemented in Python, can be swapped rapidly—an advantage for laboratories that iterate quickly.
Several limitations remain. First, writing a transition function requires domain knowledge, and complex conditions may be challenging for novice users to encode; the development of a graphical editor for specifying state transitions would therefore be valuable. Second, the current DFA model does not natively support workflows in which a single experimental sample splits into multiple experimental samples. Although this can be circumvented by registering multiple experiment with copied observations, a generalised implementation is still needed.
In summary, GEMS provides a compact abstraction layer for managing dynamic, cross-device experiments in a sample-centred, state-to-state paradigm. Although its practicality was demonstrated in two bench-scale case studies, the deterministic state description can be scaled seamlessly to facility-level operations. Because experiment definitions are decoupled from device information, each can be updated independently, enabling new instruments or assays to be integrated without downtime. This capability aligns with the needs of cloud laboratories that already operate hundreds of instruments, as well as future laboratory automation facilities.8,9 Although this work targets representational unification rather than algorithmic speed-ups, the design scales in three complementary ways. Control-logic scale keeps protocol decisions local to each experiment through a Mealy automaton, so decision-making is independent of instrument count. Resource scale expands capacity by adding instruments of the required types; the penalty-aware scheduler allocates tasks while the per-experiment logic remains unchanged. Planning scale is supported by a dry-run mode that allows users to vary instrument counts and penalty settings to estimate lateness and utilisation before execution (see Materials, Dry-run simulation for capacity planning). Together with the dynamic rescheduling seen in the parallel HEK/hiPSC run (Fig. 5D–E), these features support progression from bench to facility level. Looking ahead, integration with a graphical state editor and high-performance schedulers could further reduce the barrier to reliable, large-scale automation.
Author contributions
Yuya Tahara-Arai: conceptualization; data curation; formal analysis; funding acquisition; investigation; methodology; project administration; software; visualization; writing – original draft; writing – review & editing. Akari Kato: funding acquisition; investigation; resources; writing – review & editing. Koji Ochiai: investigation; resources; software; writing – review & editing. Kazuya Azumi: investigation; resources; writing – original draft; writing – review & editing. Koichi Takahashi: funding acquisition. Genki N. Kanda: conceptualization; funding acquisition; project administration; resources; supervision; writing – review & editing. Haruka Ozaki: conceptualization; funding acquisition; project administration; resources; supervision; writing – review & editing.Conflicts of interest
There are no conflicts to declare.Data availability
All data and code supporting the conclusions of this article are available through the following repositories. Source code: the complete GEMS framework is available at https://github.com/bioinfo-tsukuba/GEMS-python. A versioned archive of the latest snapshot is deposited at https://doi.org/10.5281/zenodo.17534259. Data and figure-generation scripts: all raw data and the scripts used to produce the figures (including the colour–mixing and cell-culture experiments) are available at https://doi.org/10.5281/zenodo.18102976. Exact software versions used: the GEMS-python version used for the RGB optimisation experiment is archived at https://doi.org/10.5281/zenodo.17534260, and the version used for the cell-culture experiment is archived at https://doi.org/10.5281/zenodo.17534282.Supplementary information (SI): Fig. SF1, Videos S1–S3, Tables ST1 and ST2, and accompanying captions are available at https://doi.org/10.5281/zenodo.18102950. See DOI: https://doi.org/10.1039/d5dd00409h.
Acknowledgements
We thank S. Takahashi (University of Tsukuba), K. Suzuki (University of Tsukuba) and H. Kawamoto (University of Tsukuba) for helpful discussions. This study was supported by grants from the JST-Mirai Program (JPMJMI18G4 and JPMJMI20G7 to K.T. and H. O.), Grant-in-Aid for Early-Career Scientists (JP22K17992 to H. O.), Grant-in-Aid for Transformative Research Areas (A) (JP23H04149 to H. O.), Grant-in-Aid for JSPS Fellows (JP22KJ3148 to A. K.), Grant-in-Aid for Scientific Research (C) (JP23K11820 to G. N. K.), JST BOOST (JPMJBS2414 to Y. T.-A.), JST K Program (JPMJKP25V1 to G. N. K.) and JST CREST (JPMJCR2551 to G. N. K. and H. O.).References
- B. P. MacLeod, F. G. L. Parlane, A. K. Brown, J. E. Hein and C. P. Berlinguette, Nat. Mater., 2022, 21, 722–726 CrossRef CAS PubMed.
- S. ul Islam, K. Kamboj and A. Kumari, MEJAST, 2023, 06, 88–97 CrossRef.
- Y. Xie, K. Sattari, C. Zhang and J. Lin, Prog. Mater. Sci., 2023, 132, 101043 CrossRef.
- M. Abolhasani and E. Kumacheva, Nat. Synth., 2023, 2, 483–492 CrossRef CAS.
- O. Bayley, E. Savino, A. Slattery and T. Noël, Matter, 2024, 7, 2382–2398 CrossRef CAS.
- M. Eisenstein, Nature, 2025, 637, 1008–1011 CrossRef CAS PubMed.
- K. Ochiai, Y. Tahara-Arai, A. Kato, K. Kaizu, H. Kariyazaki, M. Umeno, K. Takahashi, G. N. Kanda and H. Ozaki, Digital Discovery, 2025, 4, 2285–2297 RSC.
- C. Arnold, Nature, 2022, 606, 612–613 CrossRef CAS PubMed.
- N. Yachie, Robotic Biology Consortium and T. Natsume, Nat. Biotechnol., 2017, 35, 310–312 CrossRef CAS PubMed.
- Clarity LIMS software, https://www.illumina.com/products/by-type/informatics-products/lab-management-software/clarity-lims.html, accessed: 2025-5-8.
- AUTOPROTOCOL.org, An open standard for life science experimental design and automation, https://autoprotocol.org/, 2023, accessed: 2025-4-7.
- Green Button Go Scheduler Software, https://biosero.com/software/green-button-go-scheduler/, 2021, accessed: 2025-5-8.
- SAMI EX, https://www.beckman.com/liquid-handlers/software/sami-ex, Accessed: 2025-5-8.
- T. D. Itoh, T. Horinouchi, H. Uchida, K. Takahashi and H. Ozaki, SLAS Technol., 2021, 26, 650–659 CrossRef CAS PubMed.
- Y. Arai, K. Takahashi, T. Horinouchi, K. Takahashi and H. Ozaki, SLAS Technol., 2023, 28, 264–277 CrossRef CAS PubMed.
- Y. Fei, B. Rendy, R. Kumar, O. Dartsi, H. P. Sahasrabuddhe, M. J. McDermott, Z. Wang, N. J. Szymanski, L. N. Walters, D. Milsted, Y. Zeng, A. Jain and G. Ceder, Digital Discovery, 2024, 3, 2275–2288 RSC.
- L. M. Roch, F. Häse, C. Kreisbeck, T. Tamayo-Mendoza, L. P. E. Yunker, J. E. Hein and A. Aspuru-Guzik, PLoS One, 2020, 15, e0229862 CrossRef CAS PubMed.
- M. Sim, M. G. Vakili, F. Strieth-Kalthoff, H. Hao, R. J. Hickman, S. Miret, S. Pablo-García and A. Aspuru-Guzik, Matter, 2024, 7, 2959–2977 CrossRef CAS.
- J.-C. Cousty, T. Cavagna, A. Schmidt, E. Mariano, K. Villat, F. de Nanteuil and P. Miéville, Digital Discovery, 2024, 3, 2434–2447 RSC.
- M. Kulik, J. Ochs, N. König, C. McBeth, A. Sauer-Budge, A. Sharon and R. Schmitt, Procedia CIRP, 2017, 65, 242–247 CrossRef.
- S. Jung, J. Ochs, M. Kulik, N. König and R. H. Schmitt, Procedia CIRP, 2018, 72, 1245–1250 CrossRef.
- P. Egri, B. C. Csáji, K. B. Kis, L. Monostori, J. Váncza, J. Ochs, S. Jung, N. König, R. Schmitt, C. Brecher, S. Pieske and S. Wein, Procedia CIRP, 2020, 88, 600–605 CrossRef.
- S. H. M. Mehr, M. Craven, A. I. Leonov, G. Keenan and L. Cronin, Science, 2020, 370, 101–108 CrossRef CAS PubMed.
- W. Zhang, L. Hao, V. Lai, R. Corkery, J. Jessiman, J. Zhang, J. Liu, Y. Sato, M. Politi, M. E. Reish, R. Greenwood, N. Depner, J. Min, R. El-Khawaldeh, P. Prieto, E. Trushina and J. E. Hein, Nat. Commun., 2025, 16, 5182 Search PubMed.
- K. J. Åström, JMAA, 1965, 10, 174–205 CrossRef.
- L. P. Kaelbling, M. L. Littman and A. R. Cassandra, AI, 1998, 101, 99–134 Search PubMed.
- H. Kurniawati, arXiv arXiv:2107.07599, 2021 Search PubMed.
- J. E. Hopcroft, R. Motwani and J. D. Ullman, ACM SIGACT News, 2001, 32, 60–65 CrossRef.
- M. O. Rabin and D. Scott, IBM J. Res. Dev., 1959, 3, 114–125 Search PubMed.
- G. H. Mealy, Bell Syst. Tech. J., 1955, 34, 1045–1079 Search PubMed.
- B. A. Koscher, R. B. Canty, M. A. McDonald, K. P. Greenman, C. J. McGill, C. L. Bilodeau, W. Jin, H. Wu, F. H. Vermeire, B. Jin, T. Hart, T. Kulesza, S.-C. Li, T. S. Jaakkola, R. Barzilay, R. Gómez-Bombarelli, W. H. Green and K. F. Jensen, Science, 2023, 382, eadi1407 CrossRef CAS PubMed.
- J. Bai, S. Mosbach, C. J. Taylor, D. Karan, K. F. Lee, S. D. Rihm, J. Akroyd, A. A. Lapkin and M. Kraft, Nat. Commun., 2024, 15, 462 CrossRef CAS PubMed.
- F. Rahmanian, J. Flowers, D. Guevarra, M. Richter, M. Fichtner, P. Donnely, J. M. Gregoire and H. S. Stein, Adv. Mater. Interfaces, 2022, 9, 2101987 CrossRef.
- M. Balandat, B. Karrer, D. R. Jiang, S. Daulton, B. Letham, A. G. Wilson and E. Bakshy, Adv. Neural Inf. Process. Syst., 2020, 33, 21524–21538 Search PubMed.
- S. Ament, S. Daulton, D. Eriksson, M. Balandat and E. Bakshy, arXiv, 2023, DOI:10.48550/arXiv.2310.20708.
- M. Nakagawa, M. Koyanagi, K. Tanabe, K. Takahashi, T. Ichisaka, T. Aoi, K. Okita, Y. Mochiduki, N. Takizawa and S. Yamanaka, Nat. Biotechnol., 2008, 26, 101–106 CrossRef CAS PubMed.
- M. Haruta, Y. Sasai, H. Kawasaki, K. Amemiya, S. Ooto, M. Kitada, H. Suemori, N. Nakatsuji, C. Ide, Y. Honda and M. Takahashi, Invest. Ophthalmol. Vis. Sci., 2004, 45, 1020–1025 CrossRef PubMed.
- H. Kawasaki, H. Suemori, K. Mizuseki, K. Watanabe, F. Urano, H. Ichinose, M. Haruta, M. Takahashi, K. Yoshikawa, S.-I. Nishikawa, N. Nakatsuji and Y. Sasai, Proc. Natl. Acad. Sci. U. S. A., 2002, 99, 1580–1585 CrossRef CAS PubMed.
- F. Osakada, H. Ikeda, M. Mandai, T. Wataya, K. Watanabe, N. Yoshimura, A. Akaike, Y. Sasai and M. Takahashi, Nat. Biotechnol., 2008, 26, 215–224 CrossRef CAS PubMed.
- K. Ochiai, N. Motozawa, M. Terada, T. Horinouchi, T. Masuda, T. Kudo, M. Kamei, A. Tsujikawa, K. Matsukuma, T. Natsume, G. N. Kanda, M. Takahashi and K. Takahashi, SLAS Technol., 2021, 26, 209–217 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |