 Open Access Article
Open Access ArticleMolecular dynamics simulations accelerated on FPGA with high-bandwidth memory
Jing
Xiao
 abc,
Jinfeng
Chen
abc,
Jinfeng
Chen
 bcd,
Ye
Ding
bcd,
Ye
Ding
 bc,
You
Xu
bc,
You
Xu
 bc and
Jing
Huang
bc and
Jing
Huang
 *bcd
*bcd
aFudan University, Shanghai, 200433, China
bState Key Laboratory of Gene Expression, School of Life Sciences, Westlake University, Hangzhou, Zhejiang 310030, China. E-mail: huangjing@westlake.edu.cn
cWestlake AI Therapeutics Laboratory, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou, Zhejiang 310024, China
dInstitute of Biology, Westlake Institute for Advanced Study, Hangzhou, Zhejiang 310024, China
First published on 6th January 2026
Abstract
Molecular dynamics (MD) simulation is a powerful tool for investigating complex systems in physical, materials, and biological sciences. However, computational speed remains a critical bottleneck that limits its broader application. To address this challenge, we developed a dedicated hardware module based on modern field-programmable gate arrays (FPGAs) that accelerates all components of MD simulations. Our design employs pipelining strategies to optimize task execution within a fully parallel architecture, significantly enhancing performance. The latest generation of high-bandwidth memory (HBM2) is integrated and optimized to improve computational throughput. At the hardware level, we implemented an optimized register-transfer level (RTL) circuit design for a single node to maximize the efficiency of register read and write operations. Software co-design with SIMD frameworks ensures seamless integration of force calculations and system propagation. We validated the implementation across systems ranging from argon gas to solvated proteins, demonstrating stable MD trajectories and close agreement with reference energy values. This work presents a novel FPGA-based MD simulation architecture and provides a foundation for further improvements in hardware-accelerated molecular simulations.
1 Introduction
Due to their ability to provide detailed mechanistic insights into various physical interactions at the atomic level, molecular dynamics (MD) simulations have become paramount in driving the rational discovery of therapeutic agents and functional materials.1–4 However, functionally relevant processes are mostly driven by the collective motion of aggregated ensembles of molecules, which span large spatial scales and are computationally demanding to model comprehensively. Consider many-body systems interacting through classical pairwise potentials: the computational demand for integrating the equations of motion escalates rapidly with the number of interacting particles. Moreover, most biophysical activation processes of mechanistic interest occur over heterogeneous timescales.5 For example, essential cellular activities fundamental to life, such as active solute transport,6,7 can take place on sub-second timescales, while molecular vibrational motions are characterized on the femto- to picosecond scale. The multi-scale nature of these systems limits the time-step length to sufficiently short intervals (typically 0.5 to 4 femtoseconds) to ensure stable dynamical integration, rendering the computational modeling of long-timescale collective molecular behaviors highly intensive.8To extend the accessible simulation scales within the limits of modern computational hardware, contemporary MD implementations have adopted multi-CPU parallel strategies and GPU-based acceleration.9–13 However, even with GPU support, the trajectory lengths reported in most MD studies rarely exceed ten microseconds. Numerically propagating molecular trajectories for systems containing a few million particles can still require weeks of fully loaded computational power.14 While modern GPUs excel at regular computations, their architectures struggle with irregular memory-access patterns and unpredictable latency in large-scale MD simulations.
Beyond conventional CPU/GPU-based solutions, heterogeneous computing architectures have emerged as a promising paradigm for MD simulations. Recent implementations integrate microcontroller units (MCUs), field-programmable gate arrays (FPGAs),15–18 and complex programmable logic devices (CPLDs) through dedicated photonic interconnects, exemplified by the MD-GRAPE cluster architecture that achieves sub-microsecond communication latency between processing nodes.19,20 The commercial ANTON supercomputer series utilized the Application Specific Integrated Circuit (ASIC) processors to construct a specifically designed cluster with 64 or 512 nodes.21–23 However, these architectures are commercially expensive to implement, which limits their applicability for routine simulations. On the other hand, the Novo-G reconfigurable computing design leverages programmable hardware such as FPGAs to tackle the energy efficiency and performance bottlenecks, and have shown prominent potential for MD simulations.24,25
Compared to alternative computing hardware, FPGAs offer greater design flexibility and improved environmental friendliness with competitive computational efficiency.26–29 FPGAs incorporate a variety of unique production techniques, such as the flash process and the reverse fusion process, which allow for multiple reconfigurations and debugging cycles in the commercial applications, accommodating diverse experimental design requirements.30 During the chip development phase, FPGAs are often used as the core platform for prototyping, highlighting their potential for eventual translation into ASIC processes.31,32 With their high parallel capacity, reconfigurability, and excellent energy efficiency, FPGAs offer unique advantages for MD simulations. They can significantly accelerate compute-intensive floating-point operations by processing particle interaction forces, including van der Waals (vdW) and Coulomb forces, in parallel across thousands of configurable logic units. The pipelined hardware design of FPGAs enables efficient coordination of neighbor-list generation, force calculation, and integration. Moreover, FPGAs can be flexibly adapted to different potential functions and numerical integration schemes in MD simulations, thereby supporting a wide range of models from rigid bodies to coarse-grained systems.33
FPGA fabrication technologies have advanced rapidly in recent years, with process nodes reaching 7 nm and below, enabling significant performance improvements in high-performance computing. In parallel, the introduction of high-bandwidth memory (HBM) addresses the longstanding data-throughput bottleneck in MD simulations,34,35 where force calculations demand sustained memory bandwidth exceeding 100 GB s−1. Despite these advances, no existing FPGA-based MD implementations have yet taken advantage of HBM2’s unique combination of bandwidth density and near-memory compute capabilities, leaving a critical gap in scaling MD simulations beyond 10-million-atom systems.
In this study, we report the co-design of both hardware and software components based on a hybrid CPU/FPGA architecture specifically developed for MD simulations using classical force fields (FFs) such as CHARMM36m.36 To achieve optimal computational-load balancing, our design retains the bonded interaction calculations on the CPU while offloading the more computationally intensive non-bonded interaction evaluations to the FPGA. We present practical implementations of FPGA computing units specifically designed for scalable electrostatic interaction calculations based on the particle-mesh Ewald (PME) algorithm, using three-dimensional fast Fourier transform kernels. In addition, we introduce a Single Instruction, Multiple Data (SIMD) software framework developed to improve the circuit’s data processing capabilities. The rest of the paper is organized as follows: Section 2 introduces the FPGA architecture used in our design and reviews the relevant algorithms in MD simulations, focusing on their implications for achieving optimal computing performance in practice. Section 3 details the algorithmic implementation on the FPGA hardware for massively parallel non-bonded interaction computations, based on a Network-on-Chip (NoC) architecture and an HBM2 memory layout designed for efficient data dispatch with minimal memory overhead. Also presented in Section 3 is the validation of our hybrid hardware implementation on model systems with varying complexity, from argon particles to the DHFR protein in explicit solvent. Finally, Section 4 provides discussion and concluding remarks.
2 Methods
2.1 Hardware platform
This work employs the Inspur F37X FPGA accelerator card featuring the Xilinx Virtex UltraScale+ VU37P chip. Its programmable architecture enables dynamic reconfiguration of DSP blocks and hardware-level algorithm mapping, directly addressing the memory-wall bottleneck in traditional CPU architectures (via HBM2’s 460 GB s−1 bandwidth supporting dataflow parallelism) and the inflexibility of fixed instruction sets. The card delivers 28.1 TOPS INT8 compute performance at 75 W power, demonstrating 44% higher energy efficiency than the NVIDIA A100 (19.5 TOPS at 250 W),37 making it particularly suitable for deterministic-latency applications such as real-time signal processing. Comparative studies show that FPGA solutions achieve 3–8× lower tail latency than CPU/GPU hybrids for sparse matrix computations, although their stream computing advantages require careful balancing with batch-processing demands in scientific computing. This architecture ultimately addresses the joint optimization challenge of achieving high throughput and low latency through hardware-level customization (see SI Table S1 for quantified comparisons).2.2 Classical MD simulation
A classical MD program can be divided into four parts:381. Parse the specified simulation conditions, such as simulation temperature, particle number, time step, and topology of the simulation system.
2. Initialize the positions and velocities of the particles.
3. Compute the forces acting on all particles.
4. Integrate Newton’s equations to update the velocities and positions of all particles.
Steps 3 and 4 are repeated until the user-specified number of simulation steps is reached. In classical MD simulations, the force acting on each atom is typically calculated using classical force fields.39 Given the force acting on each particle, the velocity and position will be updated by an integrator algorithm, for example, the velocity Verlet methods. Most FFs model interactions between particles using bonded and non-bonded terms:
| Ftotal = Fbonded + Fnon-bonded | (1) |
The bonded force consists of bond, angle, dihedral and improper terms:
| Fbonded = Fbonds + Fangles + Fdihedrals + Fimpropers | (2) |
Most bonded terms are modeled by harmonic potentials, except for the dihedrals, which are modeled using a summation of cosine functions.36 Because the bonded terms are only calculated when bonds exists between atoms, their computational cost can be regarded as 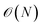 .
.
2.3 Non-bonded-force calculations
The non-bonded terms in classical FFs often consist of vdW interactions, represented by the Lennard-Jones (LJ) potential, and electrostatic interactions, modeled as Coulomb interactions between point charges: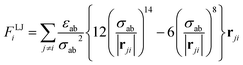 | (3) |
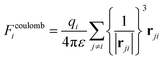 | (4) |
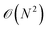 . Thus, the primary bottleneck in MD simulations lies in the evaluation of non-bonded interactions.
. Thus, the primary bottleneck in MD simulations lies in the evaluation of non-bonded interactions.
Due to the slow decay of Coulomb forces with distance, they are typically computed using the PME algorithm under periodic boundary conditions (PBC) to efficiently handle long-range interactions. The algorithm partitions ECoulomb into real-space and reciprocal-space contributions by introducing spherically distributed Gaussian charges of opposite signs. The total electrostatic energy of a system under PBC is expressed as:
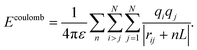 | (5) |
| ECoulomb = Edir + Erec + Ecorr, | (6) |
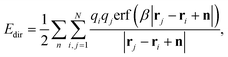 | (7) |
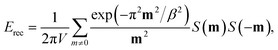 | (8) |
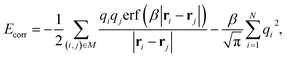 | (9) |
2.4 Neighbor list and its optimization
To improve efficiency, the calculation of short-range forces in real space sometimes uses a proximity-based lookup method to determine the spatial relationship between particles based on inter-particle distances. These relationships are stored in a neighbor list, which is periodically updated as the simulation progresses and particle positions change. For interactions within a short range (range-limited), each particle interacts only with nearby particles within a specified cutoff radius. The neighbor list is constructed by assigning each particle to a region in space and identifying neighboring particles within a cutoff distance rcut. Each particle then maintains a list of potential neighbors, which serves as input to the next computational step.In 3D periodic systems, each simulation cell has 26 adjacent cells. Newton’s third law states that action and reaction forces are equal in magnitude but opposite in direction. In MD simulations, this implies that the interaction force between particle pair (i, j) satisfies Fij = −Fji. Accordingly, to avoid redundant force calculations, only the 13 neighboring cells in the upper half of the central cell are considered when filtering particle pairs, as illustrated in Fig. S1. Essentially, one can reduce computational overhead by calculating only half of the neighbor pairs while maintaining full force accuracy.
Furthermore, we can employ a modified Verlet neighbor list that stores only unidirectional particle pairs as a strategy for optimizing the neighbor list:
| Neighbor list = {(i, j)|i < j, rij < rcut + rbuffer} | (10) |
3 Results
3.1 Proposed architecture
Based on the MD algorithm, this work proposes an FPGA architecture for accelerating MD simulations. As an example, the range-limited (RL) module, which computes the vdW interactions and the real-space component of the electrostatic interactions, is illustrated in Fig. 1. The CPU first preprocesses the simulation data by assigning a unique identifier to each particle and partitioning the dataset based on cell indices, effectively defining the spatial decomposition of the system. Particle information is then associated with the corresponding cell serial numbers. The cell size is determined by the cutoff distance, and the number of cells depends on the dimensions of the simulation box. Each cell contains information not only about its own particles but also about particles in its neighboring cells. This information is organized into structured data tables stored in memory.Each particle occupies one row in the storage space. The starting address of each data unit (cell) can be directly located through the cell serial number, and the length is allocated based on the maximum expected number of particles, with a margin reserved. An exclusion information table, containing bonded particle pairs, is also stored in the memory. This data table structure allows exclusion of the particle information in pairs, and the number and addresses of excluded particles are registered and accessed by the relevant modules during computation. The neighboring force calculation uses interpolation and coefficient tables, which are generated by the CPU and configured into the on-chip RAM of FPGA. After configuration, the CPU initiates prefetching of particle data units, and the parallel reading module supplies particle-pair information to the pipeline. Next, the multi-channel screening module processes each data unit as a main cell. It retrieves particle information from the main cell and its 13 neighboring cells to construct candidate particle pairs for distance evaluation. Specifically, each particle in the main cell is paired with particles from each of the 13 adjacent cells. The module filters out any pairs that exceed the cutoff distance. The half-shell cell-partitioning method proposed by Bowers et al. is adopted to reduce computational redundancy.40 For those within the cutoff, interpolation coefficients are computed based on squared inter-particle distances and type information. These coefficients, along with charge data, are used to evaluate electrostatic and short-range forces. All parameters required for the RL force calculation are stored in the FPGA’s on-chip RAM and passed to the distance-judgment module, ensuring that only relevant pairs proceed to the next computation stage. The range-limited force-calculation module monitors this process, updating the force-calculation result data table and overseeing the calculation progress.
3.2 Data reading and writing with HBM memory
HBM2 stores data units in a structured manner, with each unit containing multiple layers of information.34,41 The main row encodes essential metadata, including the cell serial number, particle charge, storage offset address of the particle information, and parity bits. This metadata provides the foundation for subsequent data processing. Subsequently, each quadrant uses the serialized identifier and the HBM address offset of the adjacent units to record the adjacency relationship in an orderly manner.Each spatial dimension is stored in a continuous 8-byte block, making data associations clearly traceable. Memory allocation follows a strict alignment protocol. For incomplete rows, padding is carried out to meet the requirements of the complete row boundary. Particle-specific serial numbers are stored sequentially in continuous memory segments, and the established alignment rules are maintained through terminal padding to ensure the standardization of data storage. Such a hierarchical layout not only ensures the integrity of the memory architecture but also enables efficient data access and retrieval. In contrast to the ring-based communication scheme proposed by Wu et al.42 for managing fan-in/fan-out during particle pairing, our design leverages HBM2’s multi-channel parallelism and address interleaving to achieve high-throughput access without explicit ring communication, thereby simplifying data movement while maintaining scalability. During caching, multiple channels can simultaneously read data, either at the same address or different addresses, as detailed in SI Fig. S3.
Structural omissions are made for particle items, resulting in a specialized memory topology. This layout is elaborated in detail in Fig. S3, serving as a reference structure for the computation modules of the 13 adjacent cells. The calculation of non-bonding forces uses the pre-computed interpolation coefficients and lookup tables that are loaded in the dedicated on-chip RAM resources. For particle pairs that do not require explicit calculation, identifiers of excluded bonded pairs are stored in shared memory as particle ID pairs. This arrangement ensures low-latency access during force calculations.
The particle information storage architecture establishes a dual-layer hierarchy between HomeCell and NeighborCell structures operating on 512-bit data buses, where non-functional regions undergo zero-padding optimization to maintain address alignment and facilitate efficient lookup operations as shown in Fig. 2 and 3. This unified storage schema consists of seven functional segments defined by fixed bit allocations. The initial segment uses a lookup code for rapid classification, directing processing pipelines to the appropriate computation modules based on the encoded metadata. Serial numbers are assigned globally to ensure unique identification of all simulation entities. Spatial coordinates embed precise 3D positions in the Cartesian coordinate system. Particle identifiers follow a global numbering scheme, enabling consistent tracking across the simulation. The architecture employs a hybrid addressing paradigm, combining hierarchical indexing of cell relationships with direct particle access. Register-mapped exclusion tables dynamically configure FPGA resources for force calculations. This structured approach ensures deterministic memory-access latency while maintaining compatibility with parallel processing architectures.
 | ||
| Fig. 4 Storage structure of HomeCell and NeighborCells in the particle table: 512-bit data composed of six parts. | ||
To support parallel processing, a dual-tier memory management strategy is employed: precomputed pairwise interaction data for the first 13 nearest NeighborCells are stored in on-chip registers for immediate access, while data for the remaining 13 neighbor cells are staged in 256-bit wide global memory buffers (Fig. 3). This partitioning balances latency-sensitive intra-cell computations with bandwidth-intensive inter-cell calculations and is supported by FPGA-optimized neighbor-index register files that reduce off-chip memory access. Again, the storage architecture is designed to simultaneously satisfy deterministic timing constraints for core physics kernels and maintain data-handling capacity for large-scale particle systems.
The range-limited force buffer adopts a linear addressing scheme in which entries align directly with particle enumeration, ensuring deterministic memory-access patterns essential for maintaining synchronization during large-scale N-body simulations. This structured methodology simultaneously minimizes off-chip bandwidth consumption and maximizes FPGA resource utilization through optimized register-file data structures.
Buffer management follows a two-stage protocol to handle data insertion and calculation. The dynamic memory offset is determined according to the valid-entry count. The counter is incremented after each write operation, and at the same time, the checksum for the first 16 bytes of the index row is regenerated. The dual-buffer design endows the system with the ability to resist overflow during peak calculation cycles. The IEEE 754 single-precision floating-point format is adopted to process the velocity, position, and charge components, ensuring numerical stability in the parallel processing pipeline. Compatibility with the FPGA-based DMA engine is maintained to enhance data transfer efficiency from the host to the device.
3.3 Range-limited force calculation
| FRL = Aabrij−13 + Babrij−7 + Qabrij−2 = rij × (Aabrij−14 + Babrij−8 + Qabrij−3), | (11) |
| Aab = 48εabσab12, | (12) |
| Bab = −12εabσab6, | (13) |
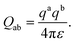 | (14) |
In this design, rij−14, rij−8 and rij−3 are required for the range-limited force computation. To reduce DSP usage, rij2 is taken as the fundamental variable, and interpolation functions for these inverse powers are precomputed. Each target function is reduced to a low-order monic polynomial in rij2, enabling reconstruction of the original values while simplifying arithmetic and lowering resource consumption. The interpolation functions are designed to balance reduced computational complexity with sufficient numerical accuracy. The performance of first-, second-, and third-order interpolation schemes was compared. By selecting an appropriate sampling interval, the derivatives of the target functions can be numerically approximated at discrete points, allowing polynomial fits valid over the chosen range. In our implementation, the second-order interpolation scheme divides the interval [0.1, 144] into eight primary segments: [0.1, 0.2], [0.2, 0.4], [0.4, 0.8], [0.8, 1.6], [1.6, 3.2], [3.2, 6.4], [6.4, 16.0], and [16, 144]). The first 7 segments are each further divided into 16 subsegments, while the last segment is split into 128 subsegments, yielding 240 interpolation functions in total. Each interpolation function requires 3 single-precision floating-point coefficients, and the full interpolation table fits within a 4 KB BRAM.
The range-limited force calculation also requires coefficient tables for Aab and Bab, where each particle-type pair has a unique set of coefficients. For N particle types, the coefficient table length is N2. The current design supports up to 32 distinct particle types, resulting in a maximum table depth of 1024 entries, sufficient for most MD systems.
In hardware floating-point computation, root operations are costly in logic resources. Therefore, rij14 and rij8 are computed via repeated multiplication from rij2, avoiding explicit square-root operations. A separate lookup table handles Coulomb forces. Parallel computations operate independently, with final results generated after floating-point multiplication and division, combined with particle parameters (Aab, Bab, etc.) fetched from memory, as illustrated in Fig. 6. Since these parameters remain constant during a simulation, the particle-pair lookup table can be precomputed, further reducing calculation logic and runtime.
Sequence control is managed through a stateful counter array (module d) that enforces strict access patterns to prevent memory-coherence violations. Data retrieval from external HBM is coordinated by a pipelined fetch unit (module e), which retrieves composite particle-attribute blocks—each home cell requires 13 cumulative data segments including self and 12 neighbor cells through burst-optimized HBM controllers. The external storage subsystem (module g) employs bank-interleaved addressing to resolve latency bottlenecks arising from concurrent access requests, while a data-fusion module (module f) aggregates force vectors from intra-cell and inter-cell interactions using SIMD-aligned accumulation registers. This architecture achieves sub-microsecond neighbor-search latencies through optimized matrix compression and pipelined dataflow.
3.4 FPGA implementation of PME for long-range electrostatics
For PME calculations of electrostatic interactions, we implemented a computationally intensive three-dimensional (3D) Fast Fourier Transform (FFT) module on FPGA, while delegating the remaining computational tasks to CPUs.29,48,49 Data exchange between the CPU and FPGA is facilitated through high-speed PCIe interfaces or optical fiber connections.50 To accelerate the calculations, we also developed a parallelized approach that leverages proximity lookup tables for Ewald summation, achieving a level of parallelism comparable to that of vdW force calculations.51 The simulation box is partitioned into cubic or rectangular grids along the X, Y, and Z axes, with grid dimensions determined by the system boundaries to optimize data alignment and hardware tiling.Charge assignment to grid points is performed using third-order B-spline interpolation. Temporal–spatial domain transformations are carried out using optimized FFT algorithms for frequency-domain computations. The hardware implementation builds upon OpenMM’s reference codebase,52 with kernel-level optimizations achieving up to a 15 times speedup in Coulomb force calculations compared to CPU-only implementations when tested on the Inspur F37X platform. Resource utilization analysis reveals 83% DSP block efficiency and 72% HBM2 bandwidth saturation when simulating a solvated protein system of approximately 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 atoms, demonstrating the feasibility of this design for large-scale MD workloads.
000 atoms, demonstrating the feasibility of this design for large-scale MD workloads.
The 3D FFT calculation proceeds sequentially along each axis (X to Y to Z), where the output of each dimension serves as the input for the next. The architectural design of this multi-stage transformation is shown in Fig. 8. Once the Z-dimension FFT is completed, the convolution module retrieves coefficients from local memory and performs convolution; for the X and Y dimensions, convolution is skipped. The processed results are then passed to the data conversion module for further handling. To improve FFT performance, we adopted the FFT 9.0 IP core.53 Among its available modes, the pipelined configuration provides the highest throughput, albeit with increased resource consumption. Performance measurements for this configuration are reported in Table 1.
The 64 × 64 × 64 FFT computation exhibits near-linear scaling compared to the 32 × 32 × 32 case, requiring approximately 8× the time—consistent with the 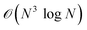 complexity of 3D FFTs. This aligns with theoretical expectations, as doubling each dimension (X, Y, Z) increases total points by 8×, while the logarithmic term accounts for the FFT’s hierarchical decomposition stages.53 Performance metrics of Anton1, though groundbreaking for its era, are now outpaced by contemporary FFT implementations. For instance, Anton3’s specialized MD simulation architecture demonstrates 100× faster throughput than generic supercomputers,23 suggesting similar leaps in FFT efficiency through domain-specific optimizations (e.g., reduced data movement and compressed encoding).
complexity of 3D FFTs. This aligns with theoretical expectations, as doubling each dimension (X, Y, Z) increases total points by 8×, while the logarithmic term accounts for the FFT’s hierarchical decomposition stages.53 Performance metrics of Anton1, though groundbreaking for its era, are now outpaced by contemporary FFT implementations. For instance, Anton3’s specialized MD simulation architecture demonstrates 100× faster throughput than generic supercomputers,23 suggesting similar leaps in FFT efficiency through domain-specific optimizations (e.g., reduced data movement and compressed encoding).
3.5 Calculation pipeline of bonded forces
Fig. 9 illustrates the cooperation mode between CPU and FPGA in the calculation of bonded forces, with data transmission between software and hardware realized through PCIe.54 Each small box in the diagram represents a circuit processing unit. The bonded forces include the bond, angle and dihedral terms in the classical FF models. In MD simulations with classical FFs, the topology, which defines how particles are connected with covalent bonds, remains fixed throughout the simulation. Particle and topology information is obtained from user input files, and the structure of the particle-information storage table can adapt to the specific topology inputs. This data is stored in a specific cache area of HBM2 using DMA technology. The FPGA data read/write submodule employs multiple channels in parallel, and is equipped with HBM2 registers, which can support multiple read and write operations simultaneously. This allows the system to efficiently read interaction data for two-body (bond), three-body (angle), and four-body (dihedral) bonded terms. The design significantly improves storage and access efficiency for large data blocks.The computation flow-control circuit manages the bonded-force calculations. Two-body force calculations are relatively straightforward and thus execute more quickly. Three-body force calculations are moderately more complex and slower, while four-body force calculations are the most computationally intensive, both in processing time and complexity, making them the primary bottleneck in the bonded-force pipeline. The covalent-bond calculation buffer is designed with these differences in mind: it incorporates specialized circuits to receive data from channels with varying access frequencies, ensuring efficient handling across all bonded-interaction types.
3.6 Performance and resource utilization of computing modules
| Modules | Latency | DSP | Resources | ||
|---|---|---|---|---|---|
| Cycles | ns | FF | LUT | ||
| First-order linear interpolation | 22 | 73.304 | 10 | 1459 | 763 |
| Second-order linear interpolation | 34 | 113.288 | 16 | 1788 | 1063 |
| Third-order linear interpolation | 45 | 148.514 | 22 | 2442 | 1452 |
On the device, the RL unit occupies only 12–18% of the total module area. A modular AXI interface enables seamless integration with the predominantly RTL design. This hierarchical approach preserves RTL’s cycle-accurate control over critical paths while leveraging HLS for rapid development of compute-intensive functions. The standardized AXI4 protocol not only decouples submodules but also establishes a scalable foundation for future multi-module chaining through configurable address mapping and clock domain crossing support.
The HLS implementation demonstrates efficient resource utilization in specific operations, such as second-order polynomial interpolation (10 DSP48 modules at 300 MHz and 22-cycle latency) and rij distance calculation (9 DSP48 and 25 cycles). In contrast, RTL-designed modules show greater potential for latency optimization of certain operations. For example, the bonded-force module designed with RTL achieves balanced resource usage (15% LUTs and 14% FFs) with only 3% DSP occupation. RTL design also allows on-chip memory balance by distributing data across LUTs, Block RAM (BRAM), and UltraRAM (URAM); in particular, URAM-based designs reduce BRAM consumption by 58% through refined memory architectures. Essentially, we adopt a hybrid approach that leverages HLS for rapid prototyping of coarse-grained parallelism while reserving RTL for latency-critical components. Experimental results confirm RTL’s consistent cycle-level advantage (e.g., 34-cycle Coulomb force vs. 45-cycle HLS interpolation), albeit at the cost of more extensive development effort. Overall, these findings underscore RTL’s adaptability for applications requiring sub-nanosecond precision, while also motivating further exploration of HLS-RTL co-design strategies.
| Resource name | Utilization% | |
|---|---|---|
| Single RL module | Double RL modules | |
| LUT | 10 | 21 |
| LUTRAM | 3 | 6 |
| FF | 13.13 | 26.4 |
| BRAM | 7 | 18.5 |
| URAM | 0.63 | — |
| DSP | 2 | 4 |
| IO | 1 | 1 |
| GT | 17 | 34 |
| BUFG | 3 | 6 |
| MMCM | 8 | 16 |
| PLL | 17 | 34 |
Expanding to two RL modules demonstrates the scalability of the architecture. Resource usage increases in a near-linear manner—LUTs rise to 21%, FFs to 26.4%, and GTs to 34%—showing that both computational and communication demands scale proportionally with the number of modules. BRAM utilization also grows from 7% to 18.5%, while URAM becomes unused, suggesting a deliberate memory-hierarchy strategy that prioritizes BRAM capacity over URAM bandwidth in multi-module deployments. This consistent scaling behavior underscores the inherently parallel nature of the design and its suitability for deployment in larger compute matrices, where dozens or hundreds of RL modules could operate concurrently with predictable performance and resource characteristics.
| Resource name | Utilization% | |
|---|---|---|
| Based on BRAM | Based on URAM | |
| LUT | 16 | 15 |
| LUTRAM | 4 | 4 |
| FF | 14 | 14 |
| BRAM | 58 | 24 |
| URAM | — | 16 |
| DSP | 7 | 7 |
| IO | 1 | 1 |
| GT | 17 | 17 |
| BUFG | 2 | 2 |
| MMCM | 8 | 8 |
| PCIe | 17 | 17 |
Logic resources (LUTs, FFs, DSPs) and I/O components (GTs, PCIe) remain modestly utilized in the range of 14–17%, leaving ample headroom for additional computational workloads despite the high resource demand of FFT operations. Nonetheless, deploying multiple FFT IPs in a single-node setup requires careful trade-off analysis, as their substantial memory and compute requirements may compete with residual BRAM (24%) and other module allocations, necessitating holistic resource optimization. We note that for the full PME pipelines, FFT modules must coexist with range-limited and bonded-force modules.
The bonded-force module, in contrast, is relatively lightweight and can be integrated efficiently into the overall design. As shown in Table S2, it demonstrates a well-balanced resource utilization on the Xilinx Virtex UltraScale+ VU37P platform. LUTs and FFs are occupied at 15% and 14%, while BRAM and URAM usages reach 24% and 13%, respectively. DSP usage is low (3%), and clock management resources (BUFG/MMCM) remain modest at 2% and 8%.
3.7 Validation: energies and forces of LJ particles
Validation of MD implementations typically involves verifying that the energies and forces are computed correctly. We applied this principle to our FPGA-based implementation of MD simulations with classical FFs. As an initial check, we considered a simple system: two LJ particles interacting with parameters σ = 3.0 Å and ε = 2.092 kJ mol−1 under periodic boundary conditions. We compared our implementation with CPU and GPU versions of OpenMM. Specifically, CPU-based reference results were obtained using an Intel i5-8400 (2.80 GHz) with OpenMM version 6.0.0, selected for its transparent implementation of 3D Fourier transforms;52,55 results were cross-checked against version 8.0 to confirm consistency. The code was compiled with the GNU C/C++ compiler 13.3.0. GPU-based results were generated on an NVIDIA GeForce GTX 1650 (driver 550.120, CUDA 12.4) using mixed-precision computation. Unless otherwise noted, subsequent benchmarks follow these same hardware and software configurations.We computed both the total energy and the force on one of the particles (symmetry makes evaluation of a single particle sufficient). The test system was placed in a 30 Å periodic box with a cutoff distance of 15 Å, and inter-particle separations ranging from 2 to 28 Å were sampled. As shown in Fig. 10, the system energy profile is symmetric about 15 Å, consistent with PBC applied. Beyond the cutoff distance, forces behave as expected, with each particle effectively interacting with the periodic image of the other across the box boundary. A further inspection between 2.5 to 8 Å further confirms that the FPGA implementation closely matches the CPU and GPU results (Fig. 10c and d). The FPGA-calculated energy exhibits a root mean square error of 1.5 × 10−5 kJ mol−1 compared to the OpenMM CPU results, demonstrating numerical equivalence. These results validate that our implementation—particularly the range-limited force module—accurately reproduces LJ interactions on FPGA hardware.
To further evaluate accuracy with more complicated systems, we extended the validation to a 15-atom argon cluster simulated in a 30 Å cubic box with PBC. Parameters from the CHARMM force field (σ = 3.8220 Å, ε = 0.9961 kJ mol−1) were employed with a 12 Å cutoff for non-bonded interactions. Five distinct configurations were extracted from the MD trajectories, and the total potential energies were compared on FPGA, GPU, and CPU platforms as described previously. The LJ potential energies (Table S3) reveal very minor inter-platform deviations, confirming the accuracy of FPGA implementation. This cross-platform agreement demonstrates that our FPGA architecture achieves high computational precision in handling LJ interactions through parallelized execution and optimized memory-bandwidth utilization.
3.8 Validation: DHFR in explicit solvent
A further validation of our design and implementation involved MD simulations of a protein solvated in an explicit water box under PBC. We chose the DHFR system, a widely used benchmark in MD studies, consisting of 159 amino acids modeled with the AMBER ff99SB protein FF56 and 7023 TIP3P water molecules, yielding a total of 23![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 558 atoms in a cubic box of 62.230 × 62.230 × 62.230 Å3. A non-bonded cutoff of 9 Å was applied, PME was used for long-range electrostatics, and 50
558 atoms in a cubic box of 62.230 × 62.230 × 62.230 Å3. A non-bonded cutoff of 9 Å was applied, PME was used for long-range electrostatics, and 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 steps were performed in the NPT ensemble with a time step of 2 fs at 300 K and 1.0 bar. Initial velocities were assigned randomly, and no additional heating or equilibrium was performed. Simulations were carried out in OpenMM on CPU and GPU (both in single-precision and double-precision). On the FPGA platform, our implementation supported full MD propagation, including all force calculations, as well as coordinate and velocity updates performed directly on the FPGA. An internal OpenMM interface was customized to enable output of the coordinates, velocities, or energies from the FPGA as needed, allowing direct comparison with CPU and GPU results.
000 steps were performed in the NPT ensemble with a time step of 2 fs at 300 K and 1.0 bar. Initial velocities were assigned randomly, and no additional heating or equilibrium was performed. Simulations were carried out in OpenMM on CPU and GPU (both in single-precision and double-precision). On the FPGA platform, our implementation supported full MD propagation, including all force calculations, as well as coordinate and velocity updates performed directly on the FPGA. An internal OpenMM interface was customized to enable output of the coordinates, velocities, or energies from the FPGA as needed, allowing direct comparison with CPU and GPU results.
The protein remained stable during 100 ps of MD simulation on FPGA, as shown in Fig. 11a, where the final snapshot from the FPGA trajectory was aligned with the initial structure. Although the simulation length is not sufficient to capture functionally relevant conformational dynamics, the heavy-atom RMSD profiles from FPGA and GPU simulations were highly similar, converging to 1.5 Å after 100 ps (Fig. 11b). We note that different random initial velocities were assigned during validation across platforms, leading to variations in the observed properties at the beginning of the nonequilibrium stage, but these quickly converge and fluctuate within the same reasonable range.
In addition, we compared the total, potential, and kinetic energies across platforms (Fig. 11). All energy profiles exhibited consistent trends and fluctuation ranges for FPGA, single-precision GPU, and double-precision GPU. The energies converged similarly within 200 steps. The last 80% of each trajectory was analyzed for average energies and fluctuations. The potential energy values were −1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 263
263![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 558.08 ± 2574.67 kJ mol−1 (single-precision GPU), −1
558.08 ± 2574.67 kJ mol−1 (single-precision GPU), −1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 262
262![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 739.74 ± 2728.18 kJ mol−1 (double-precision GPU), and −1
739.74 ± 2728.18 kJ mol−1 (double-precision GPU), and −1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 259
259![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 214.13 ± 2087.86 kJ mol−1 (FPGA). The similarity in fluctuation ranges across platforms confirms proper energy conservation. Kinetic energy values were 252
214.13 ± 2087.86 kJ mol−1 (FPGA). The similarity in fluctuation ranges across platforms confirms proper energy conservation. Kinetic energy values were 252![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 392.52 ± 1666.11 kJ mol−1 (single-precision GPU), 252
392.52 ± 1666.11 kJ mol−1 (single-precision GPU), 252![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 512.85 ± 1710.17 kJ mol−1 (double-precision GPU), and 252
512.85 ± 1710.17 kJ mol−1 (double-precision GPU), and 252![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 723.81 ± 1555.32 kJ mol−1 (FPGA). These correspond to average temperatures of 299.92 ± 1.98 K, 300.06 ± 2.03 K, and 300.31 ± 1.85 K, respectively. The observed fluctuations of 0.6% demonstrate robust temperature control across all cases, confirming that our FPGA implementation can perform MD simulations of realistic biomolecular systems with high precision.
723.81 ± 1555.32 kJ mol−1 (FPGA). These correspond to average temperatures of 299.92 ± 1.98 K, 300.06 ± 2.03 K, and 300.31 ± 1.85 K, respectively. The observed fluctuations of 0.6% demonstrate robust temperature control across all cases, confirming that our FPGA implementation can perform MD simulations of realistic biomolecular systems with high precision.
While we demonstrated that MD simulation of a solvated protein is stable with our FPGA implementation, we further examined the accuracy by comparing the energies in detail. For this we extracted five snapshots along the MD trajectories and computed energy terms under the same hardware and software environments described above (Table 5). For all five snapshots, the total energies computed on FPGA showed excellent agreement with GPU reference values, with the small deviations arising from the non-bonded-force terms. These differences most likely originated from electrostatic interactions, since the LJ terms had already been validated. The electrostatic discrepancies were traced to the 3D-FFT computation process, with two contributing factors: (1) the FPGA’s native FFT IP core supports only power-of-two matrix dimensions (e.g., 32 or 64), and (2) the FFT v9.0 IP core provides both 32-bit floating-point and fixed-point modes, each with distinct resource utilization characteristics.
| Potential energy (kJ mol−1) | ||||
|---|---|---|---|---|
| FPGA | CPU | GPU | ||
| Frame 1 | Bond | 2087.6622 | 2087.8628 | 2087.8601 |
| Angle | 5433.3571 | 5433.9008 | 5433.9029 | |
| Dihedral | 6907.1296 | 6908.4214 | 6908.4239 | |
| Non-bonded | −312![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 340.4907 340.4907 |
−312![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 345.9615 345.9615 |
−312![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 345.5637 345.5637 |
|
| Total | −297![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 910.3823 910.3823 |
−297![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 915.7766 915.7766 |
−297![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 915.3769 915.3769 |
|
| Frame 2 | Bond | 1907.2511 | 1907.1118 | 1907.1081 |
| Angle | 5511.0062 | 5511.5854 | 5511.5842 | |
| Dihedral | 6927.9415 | 6928.6778 | 6928.6803 | |
| Non-bonded | −310![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 742.7736 742.7736 |
−310![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 748.2840 748.2840 |
−310![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 747.9304 747.9304 |
|
| Total | −296![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 395.4094 395.4094 |
−296![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400.9089 400.9089 |
−296![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400.5578 400.5578 |
|
| Frame 3 | Bond | 1928.6012 | 1928.7512 | 1928.7461 |
| Angle | 5310.3633 | 5310.5181 | 5310.5246 | |
| Dihedral | 6755.2371 | 6755.8898 | 6755.8932 | |
| Non-bonded | −308![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 428.4791 428.4791 |
−308![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 434.0560 434.0560 |
−308![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 433.7031 433.7031 |
|
| Total | −294![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 433.3387 433.3387 |
−294![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 438.8969 438.8969 |
−294![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 438.5393 438.5393 |
|
| Frame 4 | Bond | 1922.3340 | 1922.4822 | 1922.4813 |
| Angle | 5359.9207 | 5360.0319 | 5360.0258 | |
| Dihedral | 7027.4043 | 7026.7016 | 7026.7046 | |
| Non-bonded | −310![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 148.6395 148.6395 |
−310![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 154.0210 154.0210 |
−310![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 153.6645 153.6645 |
|
| Total | −295![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 839.4261 839.4261 |
−295![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 844.8054 844.8054 |
−295![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 844.4528 844.4528 |
|
| Frame 5 | Bond | 1947.4269 | 1947.2788 | 1947.2756 |
| Angle | 5342.7208 | 5342.2733 | 5342.2762 | |
| Dihedral | 7001.9265 | 7001.2728 | 7001.2758 | |
| Non-bonded | −310![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 775.9111 775.9111 |
−310![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 781.2307 781.2307 |
−310![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 780.8614 780.8614 |
|
| Total | −296![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 485.0920 485.0920 |
−296![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 490.4058 490.4058 |
−296![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 490.0338 490.0338 |
|
Despite these architectural constraints, our results confirm that the overall implementation is correct and produces physically meaningful outcomes. The observed deviations fall well within acceptable ranges for MD simulations and can be attributed to known hardware limitations. Taken together, these comprehensive results with a realistic protein system validate that our FPGA implementation is accurate and suitable for production runs. The achieved accuracy, combined with the intrinsic speed and efficiency advantages of FPGA hardware, provides a strong foundation for future performance optimizations and large-scale applications.
4 Discussion and conclusions
In this work, we presented the co-design of hardware and software facilities for MD simulations with classical FFs on FPGA accelerators. Our implementation covers all major force components, including range-limited, bonded, and long-range electrostatics with PME, integrated within a hybrid CPU/FPGA architecture. To further enhance data processing capabilities, we developed a SIMD software framework that complements the hardware modules. Through careful optimization of data flow, memory hierarchy, and compute pipelines, we achieved stable and accurate simulations ranging from simple LJ particles to a solvated protein system. Validation against CPU and GPU platforms showed close numerical agreement, and most notably, we successfully simulated DHFR in explicit water on FPGA, producing stable MD trajectories. Together, these results establish FPGAs as a feasible and energy-efficient platform for production-level MD simulations.In terms of performance, the current FPGA implementation operates at 250 MHz, comparable to CPU execution but well below the device’s theoretical ceiling of 500 MHz. This gap arises primarily from frequency limitations in the critical logic paths and bandwidth bottlenecks between compute units and the storage subsystem. Several avenues exist for improvement. One strategy is adopting a mixed-precision architecture, where non-critical computations are performed at lower precision to ease timing constraints and raise the overall clock frequency. Additional gains may be achieved by reconfiguring data paths to shorten logic depth on critical paths, thereby reducing latency, and by introducing a NoC design to improve on-chip communication efficiency among computing modules. Together, these optimizations could significantly increase sustained throughput and bring the FPGA design closer to its theoretical performance potential.
Compared with existing approaches, our FPGA implementation offers distinct advantages. Unlike general-purpose CPUs and GPUs, the FPGA’s reconfigurable architecture enables tailored pipelines for particle–particle interactions, achieving near-linear scalability when modules are duplicated. Resource utilization studies reveal that critical logic and memory resources scale predictably with system size, while careful balancing of BRAM and URAM ensures efficient use of on-chip memory. Although the performance gap relative to highly specialized ASICs such as Anton and the most-advanced GPU remains substantial, FPGA-based designs provide a unique balance between configurability, efficiency, and accuracy that makes them a possible solution for future heterogeneous HPC platforms. In principle, FPGA architectures offer deterministic latency, high energy efficiency, and customizable data paths that are potentially advantageous for irregular workloads in large-scale MD simulations. However, the current implementation is not yet fully optimized—particularly in I/O coordination and memory scheduling—and future iterations will focus on these aspects to further improve sustained throughput and close the performance gap with advanced platforms.
Several limitations also remain. Our current design is constrained by vendor FFT IPs, which support only power-of-two grid dimensions and impose fixed floating-point or fixed-point operation modes, restricting flexibility in PME calculations. More broadly, the FPGA design is developed for additive FFs such as CHARMM36m, which remain the most widely used in biomolecular simulations. However, polarizable force fields are becoming increasingly important due to their ability to capture induced-dipole effects and more realistic electrostatics.57–59 Implementing such polarizable FFs on FPGA presents major challenges, particularly because induced-dipole calculations require higher precision and tighter module coupling, which place heavy demands on DSPs and memory bandwidth.60 The Drude FF, based on the classical Drude oscillator model and an extended Lagrangian formalism, could be more straightforward to implement in our FPGA design.61 Machine learning potentials or machine learning force fields (MLFFs) are also rapidly emerging,62,63 such as ANI,64,65 DeepMD,66–68 and the MACE series.69,70 However, MLFFs are still significantly slower than additive and polarizable FFs. Their implementation on FPGA remains an interesting but challenging future direction due to their irregular memory access and heavy reliance on tensor operations.
We note that in our current FPGA implementation, the computation of non-bonded interactions, particularly the PME electrostatics, remains a major bottleneck. Alternative algorithms such as the Fast Multipole Method (FMM) could offer advantages by reducing communication overhead through hierarchical decomposition.71 Beyond deterministic algorithms, stochastic approaches have recently emerged as powerful alternatives to address the communication and synchronization limitations inherent to FFT-based methods. In particular, the Random Batch Ewald (RBE) method employs stochastic mini-batch sampling to speed up the summation of the Fourier series in the calculation of Erec (eqn (8)), achieving exceptional parallel scalability on large CPU clusters.72–75 The simplified communication patterns and locality-friendly characteristics of these hierarchical and stochastic methods make them highly appealing for future implementation on specialized hardware such as FPGAs, where they may provide a transformative route to alleviating the von Neumann bottleneck in exascale MD simulations.
In conclusion, our study demonstrates that FPGA accelerators can deliver accurate, stable, and scalable MD simulations of biomolecular systems. While further optimization is needed, the demonstrated feasibility, efficiency, and adaptability of FPGA hardware establish a strong foundation for future advances in large-scale biomolecular simulations, bridging the gap between current CPU/GPU platforms and next-generation domain-specific accelerators. This proof-of-concept hybrid CPU/FPGA design highlights the feasibility of dedicated hardware for accelerating scientific computing and points toward future development of computing infrastructure that can extend the applicable scale of MD simulations.
Author contributions
Jing Xiao: methodology, investigation, software, formal analysis, validation, visualization, writing – original draft, and writing – review and editing. Jinfeng Chen: methodology, investigation, software, formal analysis, validation, visualization and writing – review and editing. Ye Ding: methodology, software, investigation, validation, and formal analysis. You Xu: methodology, supervision, and writing – review and editing. Jing Huang: conceptualization, funding acquisition, methodology, project administration, resources, supervision, and writing – review and editing.Conflicts of interest
There are no conflicts to declare.Data availability
The Verilog HDL code for molecular dynamics simulation (deployed on the Inspur F37x), together with the host-side C++ program controlling the Inspur F37x accelerator card, is publicly available as an archived and citable release on Zenodo at https://doi.org/10.5281/zenodo.18096184, which corresponds to the GitHub repository https://github.com/JingHuangLab/FPGA-MD-code-and-data. The repository includes specifications, dependencies, and detailed instructions for building, synthesizing, and executing the simulations. Representative input files and corresponding output data for key benchmarks are also provided in the GitHub repository, while larger trajectories and benchmark datasets have been deposited on Zenodo and are publicly available at https://doi.org/10.5281/zenodo.17796179.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5dd00391a.
Acknowledgements
This work was supported by the Shandong Provincial Natural Science Foundation (ZR2024LZH009), the “Pioneer” and “Leading Goose” R&D Program of Zhejiang (2023C03109, 2024SSYS0036), the National Natural Science Foundation of China (32171247), and the State Key Laboratory of Gene Expression. We thank the Westlake University Supercomputer Center and the Inspur for computational resources and related assistance.Notes and references
- L. Donaldson, Mater. Today, 2013, 16, 360 Search PubMed.
- M. De Vivo, M. Masetti, G. Bottegoni and A. Cavalli, J. Med. Chem., 2016, 59, 4035–4061 Search PubMed.
- R. Qian, J. Xue, Y. Xu and J. Huang, J. Chem. Inf. Model., 2024, 64, 7214–7237 CrossRef CAS PubMed.
- S. Jiang, J. Zhang, K. Diao, X. Liu and Z. Ding, Adv. Funct. Mater., 2025, 35, 2423566 Search PubMed.
- W. E and E. Vanden-Eijnden, Annu. Rev. Phys. Chem., 2010, 61, 391–420 CrossRef CAS PubMed.
- X. Lu and J. Huang, Biophys. J., 2024, 123, 1195–1210 Search PubMed.
- X. Li, X. Yang, X. Lu, B. Lin, Y. Zhang, B. Huang, Y. Zhou, J. Huang, K. Wu and Q. Zhou, et al. , Nat. Commun., 2025, 16, 7600 CrossRef CAS PubMed.
- G. Kumar, R. R. Mishra and A. Verma, in Introduction to Molecular Dynamics Simulations, ed. A. Verma, S. Mavinkere Rangappa, S. Ogata and S. Siengchin, Springer Nature Singapore, Singapore, 2022, pp. 1–19 Search PubMed.
- W. Hwang, S. L. Austin, A. Blondel, E. D. Boittier, S. Boresch, M. Buck, J. Buckner, A. Caflisch, H.-T. Chang, X. Cheng, Y. K. Choi, J.-W. Chu, M. F. Crowley, Q. Cui, A. Damjanovic, Y. Deng, M. Devereux, X. Ding, M. F. Feig, J. Gao, D. R. Glowacki, J. E. I. Gonzales, M. B. Hamaneh, E. D. Harder, R. L. Hayes, J. Huang, Y. Huang, P. S. Hudson, W. Im, S. M. Islam, W. Jiang, M. R. Jones, S. Käser, F. L. Kearns, N. R. Kern, J. B. Klauda, T. Lazaridis, J. Lee, J. A. Lemkul, X. Liu, Y. Luo, A. D. J. MacKerell, D. T. Major, M. Meuwly, K. Nam, L. Nilsson, V. Ovchinnikov, E. Paci, S. Park, R. W. Pastor, A. R. Pittman, C. B. Post, S. Prasad, J. Pu, Y. Qi, T. Rathinavelan, D. R. Roe, B. Roux, C. N. Rowley, J. Shen, A. C. Simmonett, A. J. Sodt, K. Töpfer, M. Upadhyay, A. van der Vaart, L. I. Vazquez-Salazar, R. M. Venable, L. C. Warrensford, H. L. Woodcock, Y. Wu, C. L. I. Brooks, B. R. Brooks and M. Karplus, J. Phys. Chem. B, 2024, 128, 9976–10042 CrossRef CAS PubMed.
- R. Salomon-Ferrer, A. W. Götz, D. Poole, S. Le Grand and R. C. Walker, J. Chem. Theory Comput., 2013, 9, 3878–3888 Search PubMed.
- J. C. Phillips, D. J. Hardy, J. D. C. Maia, J. E. Stone, J. V. Ribeiro, R. C. Bernardi, R. Buch, G. Fiorin, J. Hénin, W. Jiang, R. McGreevy, M. C. R. Melo, B. K. Radak, R. D. Skeel, A. Singharoy, Y. Wang, B. Roux, A. Aksimentiev, Z. Luthey-Schulten, L. V. Kalé, K. Schulten, C. Chipot and E. Tajkhorshid, J. Chem. Phys., 2020, 153, 044130 CrossRef CAS PubMed.
- P. Eastman, R. Galvelis, R. P. Peláez, C. R. A. Abreu, S. E. Farr, E. Gallicchio, A. Gorenko, M. M. Henry, F. Hu, J. Huang, A. Krämer, J. Michel, J. A. Mitchell, V. S. Pande, J. P. Rodrigues, J. Rodriguez-Guerra, A. C. Simmonett, S. Singh, J. Swails, P. Turner, Y. Wang, I. Zhang, J. D. Chodera, G. De Fabritiis and T. E. Markland, J. Phys. Chem. B, 2024, 128, 109–116 Search PubMed.
- M. J. Abraham, T. Murtola, R. Schulz, S. Pall, J. C. Smith, B. Hess and E. Lindahl, SoftwareX, 2015, 1–2, 19–25 CrossRef.
- D. Jones, J. E. Allen, Y. Yang, W. F. Drew Bennett, M. Gokhale, N. Moshiri and T. S. Rosing, J. Chem. Theory Comput., 2022, 18, 4047–4069 Search PubMed.
- P. Hamm, J. Chem. Phys., 2025, 162, 054108 Search PubMed.
- C. Wu, T. Geng, A. Guo, S. Bandara, P. Haghi, C. Liu, A. Li and M. Herbordt, Proceedings of the International Conference for High Performance Computing, Networking, Storage and Analysis, New York, NY, USA, 2023 Search PubMed.
- J. M. Rodriguez-Borbon, A. Kalantar, S. S. R. K. Yamijala, M. B. Oviedo, W. Najjar and B. M. Wong, J. Chem. Theory Comput., 2020, 16, 2085–2098 Search PubMed.
- C. Yang, T. Geng, T. Wang, R. Patel, Q. Xiong, A. Sanaullah, C. Wu, J. Sheng, C. Lin, V. Sachdeva, W. Sherman and M. Herbordt, Proceedings of the International Conference for High Performance Computing, Networking, Storage and Analysis, New York, NY, USA, 2019, pp. 1–31 Search PubMed.
- T. Narumi, K. Yasuoka, M. Taiji, F. Zerbetto and S. Höfinger, Comput. J., 2011, 54, 1181–1187 CrossRef.
- G. Morimoto, Y. M. Koyama, H. Zhang, T. S. Komatsu, Y. Ohno, K. Nishida, I. Ohmura, H. Koyama and M. Taijit, SC21: International Conference for High Performance Computing, Networking, Storage and Analysis, 2021, pp. 1–15 Search PubMed.
- D. E. Shaw, M. M. Deneroff, R. O. Dror, J. S. Kuskin, R. H. Larson, J. K. Salmon, C. Young, B. Batson, K. J. Bowers, J. C. Chao, M. P. Eastwood, J. Gagliardo, J. P. Grossman, C. R. Ho, D. J. Ierardi, I. Kolossváry, J. L. Klepeis, T. Layman, C. McLeavey, M. A. Moraes, R. Mueller, E. C. Priest, Y. Shan, J. Spengler, M. Theobald, B. Towles and S. C. Wang, Commun. ACM, 2008, 51, 91–97 CrossRef.
- D. E. Shaw, J. Grossman, J. A. Bank, B. Batson, J. A. Butts, J. C. Chao, M. M. Deneroff, R. O. Dror, A. Even, C. H. Fenton, A. Forte, J. Gagliardo, G. Gill, B. Greskamp, C. R. Ho, D. J. Ierardi, L. Iserovich, J. S. Kuskin, R. H. Larson, T. Layman, L.-S. Lee, A. K. Lerer, C. Li, D. Killebrew, K. M. Mackenzie, S. Y.-H. Mok, M. A. Moraes, R. Mueller, L. J. Nociolo, J. L. Peticolas, T. Quan, D. Ramot, J. K. Salmon, D. P. Scarpazza, U. B. Schafer, N. Siddique, C. W. Snyder, J. Spengler, P. T. P. Tang, M. Theobald, H. Toma, B. Towles, B. Vitale, S. C. Wang and C. Young, SC’ 14: Proceedings of the International Conference for High Performance Computing, Networking, Storage and Analysis, 2014, pp. 41–53 Search PubMed.
- D. E. Shaw, P. J. Adams, A. Azaria, J. A. Bank, B. Batson, A. Bell, M. Bergdorf, J. Bhatt, J. A. Butts, T. Correia, R. M. Dirks, R. O. Dror, M. P. Eastwood, B. Edwards, A. Even, P. Feldmann, M. Fenn, C. H. Fenton, A. Forte, J. Gagliardo, G. Gill, M. Gorlatova, B. Greskamp, J. Grossman, J. Gullingsrud, A. Harper, W. Hasenplaugh, M. Heily, B. C. Heshmat, J. Hunt, D. J. Ierardi, L. Iserovich, B. L. Jackson, N. P. Johnson, M. M. Kirk, J. L. Klepeis, J. S. Kuskin, K. M. Mackenzie, R. J. Mader, R. McGowen, A. McLaughlin, M. A. Moraes, M. H. Nasr, L. J. Nociolo, L. O’Donnell, A. Parker, J. L. Peticolas, G. Pocina, C. Predescu, T. Quan, J. K. Salmon, C. Schwink, K. S. Shim, N. Siddique, J. Spengler, T. Szalay, R. Tabladillo, R. Tartler, A. G. Taube, M. Theobald, B. Towles, W. Vick, S. C. Wang, M. Wazlowski, M. J. Weingarten, J. M. Williams and K. A. Yuh, Proceedings of the International Conference for High Performance Computing, Networking, Storage and Analysis, New York, NY, USA, 2021 Search PubMed.
- A. George, H. Lam and G. Stitt, Comput. Sci. Eng., 2011, 13, 82–86 Search PubMed.
- A. D. George, M. C. Herbordt, H. Lam, A. G. Lawande, J. Sheng and C. Yang, 2016 IEEE High Performance Extreme Computing Conference, HPEC, 2016, pp. 1–7 Search PubMed.
- A. M. Cabrera, Y. A. Yucesan, F. Y. Liu, W. Blokland and J. S. Vetter, 2023 IEEE High Performance Extreme Computing Conference, HPEC, 2023, pp. 1–6 Search PubMed.
- S. Kilts, Advanced FPGA Design: Architecture, Implementation, and Optimization, Wiley-IEEE Press, 2007 Search PubMed.
- H. M. Waidyasooriya, M. Hariyama and K. Kasahara, 2016 IEEE/ACIS 15th International Conference on Computer and Information Science (ICIS), 2016, pp. 1–5 Search PubMed.
- L. C. Stewart, C. Pascoe, E. Davis, B. W. Sherman, M. Herbordt and V. Sachdeva, 2021 IEEE 29th Annual International Symposium on Field-Programmable Custom Computing Machines (FCCM), 2021, pp. 270 Search PubMed.
- Y. Yang, W. Long, R. Kannan and V. K. Prasanna, 2023 33rd International Conference on Field-Programmable Logic and Applications (FPL), 2023, pp. 174–181 Search PubMed.
- M. Choi, X. Xu and V. Moroz, 2019 18th IEEE Intersociety Conference on Thermal and Thermomechanical Phenomena in Electronic Systems (ITherm), 2019, pp. 107–112 Search PubMed.
- U. Khan, H. Owen and J. Hughes, Sixth Annual IEEE International ASIC Conference and Exhibit, 1993, pp. 336–340 Search PubMed.
- C. Yang, T. Geng, T. Wang, R. Patel, Q. Xiong, A. Sanaullah, J. Sheng, C. Lin, V. Sachdeva, W. Sherman and M. C. Herbordt, arXiv, 2019, preprint, arXiv:1905.05359, DOI:10.48550/arXiv.1905.05359.
- N. Fujita, R. Kobayashi, Y. Yamaguchi and T. Boku, 2021 IEEE International Conference on Cluster Computing (CLUSTER), 2021, pp. 783–786 Search PubMed.
- D. Winterberg, V. Kumar, T. Chen and B. Mutnury, 2023 Joint Asia-Pacific International Symposium on Electromagnetic Compatibility and International Conference on ElectroMagnetic Interference & Compatibility, APEMC/INCEMIC, 2023, pp. 1–4 Search PubMed.
- K. Vanommeslaeghe and A. MacKerell, Biochim. Biophys. Acta Gen. Subj., 2015, 1850, 861–871 Search PubMed.
- NVIDIA Corporation, NVIDIA A100 Tensor Core GPU Architecture, NVIDIA Corporation, 2020 Search PubMed.
- D. C. Rapaport, The Art of Molecular Dynamics Simulation, Cambridge University Press, 2nd edn, 2004 Search PubMed.
- X. Lu, J. Chen and J. Huang, Structure, 2025, 33, 1138–1149 Search PubMed.
- K. J. Bowers, R. O. Dror and D. E. Shaw, J. Comput. Phys., 2007, 221, 303–329 CrossRef.
- S. Jayaraman, B. Zhang and V. Prasanna, 2022 International Conference on Field-Programmable Technology (ICFPT), 2022, pp. 1–11 Search PubMed.
- C. Wu, T. Geng, S. Bandara, C. Yang, V. Sachdeva, W. Sherman and M. Herbordt, 2021 IEEE 29th Annual International Symposium on Field-Programmable Custom Computing Machines (FCCM), 2021, pp. 142–151 Search PubMed.
- IEEE Computer Society, IEEE Standard for Floating-Point Arithmetic, IEEE Std 754-2019 (Revision of IEEE 754-2008), 2019, pp. 1–84, DOI:10.1109/IEEESTD.2019.8766229.
- E. R. Overchick and B. Abramov, 2014 IEEE 28th Convention of Electrical & Electronics Engineers in Israel (IEEEI), 2014, pp. 1–5 Search PubMed.
- J. Villarreal and W. A. Najjar, 2008 International Conference on Field Programmable Logic and Applications, 2008, pp. 667–670 Search PubMed.
- C. Yang, T. Geng, T. Wang, C. Lin, J. Sheng, V. Sachdeva, W. Sherman and M. Herbordt, 2019 IEEE 30th International Conference on Application-specific Systems, Architectures and Processors (ASAP), 2019, pp. 263–271 Search PubMed.
- M. Chiu and M. C. Herbordt, ACM Trans. Reconfigurable Technol. Syst., 2010, 3, 1–37 CrossRef PubMed.
- T. A. Darden, D. M. York and L. G. Pedersen, J. Chem. Phys., 1993, 98, 10089–10092 Search PubMed.
- Z.-H. Duan and R. Krasny, J. Chem. Phys., 2000, 113, 3492–3495 Search PubMed.
- B. S. C. Varma, K. Paul and M. Balakrishnan, 2013 26th International Conference on VLSI Design and 2013 12th International Conference on Embedded Systems, 2013, pp. 92–97 Search PubMed.
- K. Aatish and B. Zhang, Three Dimensional Fast Fourier Transform CUDA Implementation, in CSE 260: Parallel Computation (Fall 2012), University of California, San Diego, 2013, https://cseweb.ucsd.edu/~baden/classes/Exemplars/cse260_fa12/3DFFT.pdf Search PubMed.
- P. Eastman, J. Swails, J. D. Chodera, R. T. McGibbon, Y. Zhao, K. A. Beauchamp, L.-P. Wang, A. C. Simmonett, M. P. Harrigan, C. D. Stern, R. P. Wiewiora, B. R. Brooks and V. S. Pande, PLoS Comput. Biol., 2017, 13, 1–17 CrossRef PubMed.
- J. Sheng, C. Yang, A. Sanaullah, M. Papamichael, A. Caulfield and M. C. Herbordt, 2017 27th International Conference on Field Programmable Logic and Applications (FPL), 2017, pp. 1–4 Search PubMed.
- Y. Gu, T. VanCourt and M. Herbordt, International Conference on Field Programmable Logic and Applications, 2005, pp. 475–480 Search PubMed.
- P. Eastman, R. Galvelis, R. P. Peláez, C. R. A. Abreu, S. E. Farr, E. Gallicchio, A. Gorenko, M. M. Henry, F. Hu, J. Huang, A. Krämer, J. Michel, J. A. Mitchell, V. S. Pande, J. P. Rodrigues, J. Rodriguez-Guerra, A. C. Simmonett, S. Singh, J. Swails, P. Turner, Y. Wang, I. Zhang, J. D. Chodera, G. De Fabritiis and T. E. Markland, J. Phys. Chem. B, 2024, 128, 109–116 CrossRef CAS PubMed.
- V. Hornak, R. Abel, A. Okur, B. Strockbine, A. Roitberg and C. Simmerling, Proteins: Struct., Funct., Bioinf., 2006, 65, 712–725 CrossRef CAS PubMed.
- J. Deng and Q. Cui, J. Am. Chem. Soc., 2022, 144, 4594–4610 CrossRef CAS PubMed.
- J. Chen, Z. Qiu and J. Huang, ACS Omega, 2023, 8, 42936–42950 CrossRef CAS PubMed.
- Z. Pan, J. Huang and W. Zhuang, JACS Au, 2021, 1, 1788–1797 CrossRef CAS PubMed.
- Z. Huang, S. Zhao, P. Cieplak, Y. Duan, R. Luo and H. Wei, J. Chem. Theory Comput., 2023, 19, 5047–5057 CrossRef CAS PubMed.
- J. Huang, J. A. Lemkul, P. K. Eastman and A. D. MacKerell Jr, J. Comput. Chem., 2018, 39, 1682–1689 CrossRef CAS PubMed.
- Y. Ding, K. Yu and J. Huang, Curr. Opin. Struct. Biol., 2023, 78, 102502 CrossRef CAS PubMed.
- X. Feng, Y. Xu and J. Huang, Commun. Comput. Chem., 2025, 7, 152–160 CrossRef.
- J. S. Smith, O. Isayev and A. E. Roitberg, Chem. Sci., 2017, 8, 3192–3203 Search PubMed.
- C. Devereux, J. S. Smith, K. K. Huddleston, K. Barros, R. Zubatyuk, O. Isayev and A. E. Roitberg, J. Chem. Theory Comput., 2020, 16, 4192–4202 CrossRef CAS PubMed.
- H. Wang, L. Zhang, J. Han and W. E, Comput. Phys. Comm., 2018, 228, 178–184 CrossRef CAS.
- J. Zeng, D. Zhang, D. Lu, P. Mo, Z. Li, Y. Chen, M. Rynik, L. Huang, Z. Li, S. Shi, Y. Wang, H. Ye, P. Tuo, J. Yang, Y. Ding, Y. Li, D. Tisi, Q. Zeng, H. Bao, Y. Xia, J. Huang, K. Muraoka, Y. Wang, J. Chang, F. Yuan, S. L. Bore, C. Cai, Y. Lin, B. Wang, J. Xu, J.-X. Zhu, C. Luo, Y. Zhang, R. E. A. Goodall, W. Liang, A. K. Singh, S. Yao, J. Zhang, R. Wentzcovitch, J. Han, J. Liu, W. Jia, D. M. York, W. E, R. Car, L. Zhang and H. Wang, J. Chem. Phys., 2023, 159, 054801 CrossRef CAS PubMed.
- J. Zeng, D. Zhang, A. Peng, X. Zhang, S. He, Y. Wang, X. Liu, H. Bi, Y. Li, C. Cai, C. Zhang, Y. Du, J.-X. Zhu, P. Mo, Z. Huang, Q. Zeng, S. Shi, X. Qin, Z. Yu, C. Luo, Y. Ding, Y.-P. Liu, R. Shi, Z. Wang, S. L. Bore, J. Chang, Z. Deng, Z. Ding, S. Han, W. Jiang, G. Ke, Z. Liu, D. Lu, K. Muraoka, H. Oliaei, A. K. Singh, H. Que, W. Xu, Z. Xu, Y.-B. Zhuang, J. Dai, T. J. Giese, W. Jia, B. Xu, D. M. York, L. Zhang and H. Wang, J. Chem. Theory Comput., 2025, 21, 4375–4385 CrossRef CAS PubMed.
- I. Batatia, D. P. Kovacs, G. N. C. Simm, C. Ortner and G. Csanyi, Adv. Neural Inf. Process. Syst., 2022, 35, 11423–11436 Search PubMed.
- D. P. Kovacs, J. H. Moore, N. J. Browning, I. Batatia, J. T. Horton, Y. Pu, V. Kapil, W. C. Witt, I.-B. Magdau, D. J. Cole and G. Csanyi, J. Am. Chem. Soc., 2025, 147, 17598–17611 CrossRef CAS PubMed.
- B. Kohnke, C. Kutzner and H. Grubmüller, J. Chem. Theory Comput., 2020, 16, 6938–6949 Search PubMed.
- S. Jin, L. Li, Z. Xu and Y. Zhao, SIAM J. Sci. Comput., 2021, 43, B937–B960 Search PubMed.
- J. Liang, Z. Xu and Y. Zhao, J. Phys. Chem., 2022, 126, 3583–3593 Search PubMed.
- J. Liang, P. Tan, Y. Zhao, L. Li, S. Jin, L. Hong and Z. Xu, J. Chem. Phys., 2022, 156, 014114 CrossRef CAS PubMed.
- J. Liang, P. Tan, L. Hong, S. Jin, Z. Xu and L. Li, J. Chem. Phys., 2022, 157, 144102 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |










