Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Deep generative inverse design of biofunctional polymer coatings using conditional GANs
Wafa Benaatou *a,
Mudasir Ahmad Wanib and
Kashish Ara Shakilc
*a,
Mudasir Ahmad Wanib and
Kashish Ara Shakilc
aHampton University, USA. E-mail: wafa.benaatou@hamptonu.edu
bCollege of Computer and Information Sciences, Imam Mohammad Ibn Saud Islamic University (IMSIU), Riyadh, Kingdom of Saudi Arabia. E-mail: mawani@imamu.edu.sa
cDepartment of Computer Sciences, College of Computer and Information Sciences, Princess Nourah Bint Abdulrahman University, P.O. Box 84428, Riyadh 11671, SA. E-mail: kashakil@pnu.edu.sa
First published on 23rd March 2026
Abstract
Designing biofunctional surface coatings for biomedical implants requires balancing multiple biological objectives, including cell viability, antibacterial activity, and controlled drug release. Conventional experimental optimization of such multi-objective systems is time-intensive and explores only a limited portion of the feasible design space. Here, we present a constraint-aware conditional generative adversarial network (cGAN) framework for the inverse design of polymer-based coating compositions conditioned on desired biological performance targets. The model was trained on a curated dataset combining experimentally derived and synthetically augmented compositions and evaluated using independent surrogate predictors of biological response. The forward predictive model demonstrated high predictive performance, achieving R2 values ranging from 0.90 to 0.94 across the evaluated biological endpoints, while generated candidates satisfied compositional feasibility constraints and achieved reduced mean distance to target relative to baseline sampling and optimization strategies. All evaluations are conducted in silico within a surrogate modeling framework; therefore, the results should be interpreted as computational prioritization of candidate formulations rather than experimentally validated performance. Overall, this study establishes a reproducible computational foundation for constraint-guided inverse design of multifunctional biomaterial coatings and provides a structured pathway toward future experimental validation.
1. Introduction
Biofunctional coatings enhance the biological and mechanical performance of biomedical implants, especially those made from polymers such as polyetheretherketone (PEEK). These modifications promote bone integration, limit bacterial adhesion, and enable controlled drug release, resulting in safer, longer-lasting implants.1,2 Silver nanoparticles (AgNPs) are widely studied for such coatings due to their antibacterial activity and tunable tissue interactions.23–29 However, achieving a balance between antimicrobial efficacy and biocompatibility typically requires lengthy, costly experimental optimization.Over the past decade, artificial intelligence (AI) and machine learning (ML) have expanded the field of materials research.3,4 Specifically, these methods identify patterns in experimental data and predict material behavior more efficiently than traditional trial-and-error. Building on these advances, generative models, particularly Generative Adversarial Networks (GANs) and cGANs, generate material compositions to meet objectives.5–7,20,21 Applications encompass molecular design, alloy optimization, and additive manufacturing. Moreover, their potential to accelerate discovery in biomedical materials is still emerging.
In biomaterials, machine learning predicts outcomes such as cell viability (the ability of cells to survive), antibacterial performance (effectiveness against bacteria), and tissue–surface interactions (how tissues interact with material surfaces) from chemical and structural descriptors (quantitative features representing composition and structure).8–11 However, few studies have generated new coating formulations aimed at targeted biological effects. Thus, integrating experimentally validated data, especially for AgNP-based (silver nanoparticle-based) coatings, into generative models could provide a more reliable foundation for computational design and reduce unrealistic predictions.
Here, we present a constraint-aware conditional generative adversarial network (cGAN) for the inverse design of biofunctional coatings on polyether ether ketone (PEEK) and related biomedical polymers. The proposed model generates coating compositions tailored to desired biological outcomes, including enhanced cell viability, antibacterial activity, and controlled drug release.23–30 By combining experimentally derived measurements with computationally generated synthetic data, this work bridges experimental biomaterial development and data-driven computational design, offering a practical pathway toward accelerated biomaterials innovation.
Furthermore, the framework integrates compositional feasibility constraints, surrogate-guided multi-objective optimization, and reproducible data-driven validation within a unified computational pipeline for inverse biomaterial design.
2. Materials and methods
2.1 Algorithm workflow and model architecture
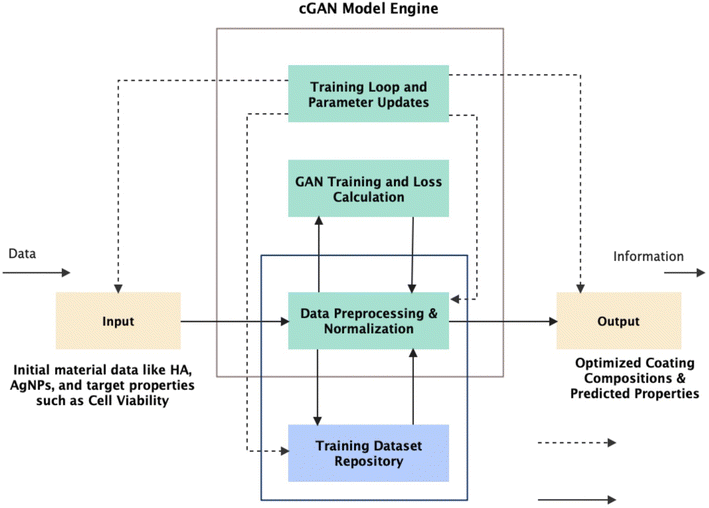
The proposed cGAN model uses two competing neural networks to generate optimized biofunctional surface coatings for biomedical applications. The workflow begins with experimental data preprocessing (cleaning and formatting for model input), continues with cGAN model training, and then validates generated designs using a forward neural network to predict coating performance. Fig. 1 presents a flowchart summarizing this pipeline.2.1.1.1 Input. Raw experimental or simulated data, consisting of coating compositions (e.g., hydroxyapatite [HA], silver nanoparticles [AgNPs], and ZnO) and associated biological performance metrics (e.g., cell viability, drug release, and antibacterial efficacy), were collected.1,2,31,32
2.2.1.2 Data preprocessing & normalization. Data were cleaned, normalized, and structured. Coating components are vectorized using a fixed vocabulary of 10 bioactive agents to ensure consistent formatting and efficient model ingestion.3–5
2.2.1.3 Training dataset repository. A curated dataset of 1000 samples (real and synthetic) formed the training corpus. The composition vectors were structured using the fixed vocabulary of the biofunctional materials listed in Table 1.6–8
| Index | Material | Description |
|---|---|---|
| 0 | HA | Hydroxyapatite |
| 1 | AgNPs | Silver nanoparticles |
| 2 | ZnO | Zinc oxide |
| 3 | TiO2 | Titanium dioxide |
| 4 | Chitosan | Biopolymer with antibacterial effect |
| 5 | Peptides | Bioactive signaling molecules |
| 6 | Collagen | ECM protein for tissue integration |
| 7 | PCL | Polycaprolactone |
| 8 | Heparin | Anti-inflammatory agent |
| 9 | PEG | Polyethylene glycol (hydrophilic) |
2.2.1.4 Training loop and parameter updates. The model was trained iteratively using backpropagation with the Adam optimizer.
The learning rate, batch size, and feature-matching weight (α) were fixed across all experiments to ensure stable and reproducible training over 1000 epochs.
2.2.1.5 Output module. The final output consisted of optimized coating formulations that were predicted to achieve the desired biological effects. These results were further evaluated using a separate predictive model for quality assurance.
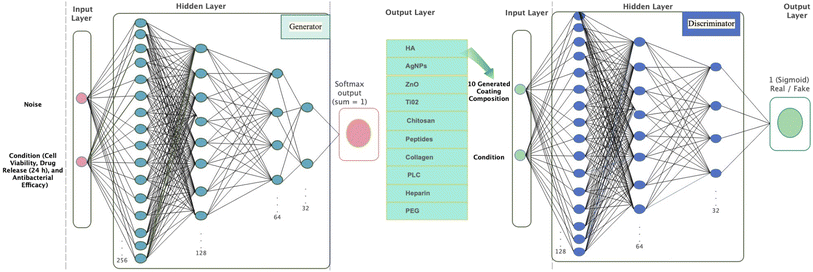
2.2 Conditional GAN architecture
The proposed cGAN architecture consists of two neural networks: a Generator (G) and a Discriminator (D). The overall objective of the model is defined by the conditional adversarial loss function:
ΓcGAN(G,D) = Ex,y[log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D(x,y)] + Ex,y[log(1 − D(G(x,y),y))] D(x,y)] + Ex,y[log(1 − D(G(x,y),y))]
| (1) |
The overall training configuration is summarized in Table 2, and the complete architecture is depicted in Fig. 2.
| Index | Material |
|---|---|
| Optimizer | Adam |
| Learning rate | 0.0002 |
| β1, β2 | 0.5, 0.999 |
| Batch size | 32 |
| Epochs | 1000 |
| Feature matching weight (α) | 20 |
| Noise distribution | Normal (0,1) |
| Random seed | Fixed across runs for reproducibility |
| Loss functions | Adversarial + feature matching |
2.3 Training strategy
The cGAN model was trained using the Adam optimizer with a learning rate of 0.0002 for a fixed 1000 epochs.13,14The generator was optimized with a composite loss function combining adversarial and feature-matching losses to enhance the quality and stability of the generated samples. The total generator loss is given by:
| ΓG = Ex,y[log(1 − D(G(x,y),y))] + α·‖µreal − µfake‖22 | (2) |
To improve the generalization of the discriminator and prevent overfitting, techniques such as label smoothing and dropout were employed during the training.
To ensure that the generated coating compositions were physically meaningful and followed the conventions of mixture design, compositional constraints were explicitly enforced during and after generation. Each output vector from the generator was passed through a simplex projection layer, which guarantees that all component values are nonnegative and that their total sum equals one. This normalization step ensures that every proposed formulation represents a valid mixture.
Furthermore, upper bounds were imposed for certain components known to have cytotoxic or solubility limits. In particular, the mass fractions of AgNPs and ZnO were restricted to experimentally acceptable ranges reported in the biomaterials literature. These thresholds were applied as hard constraints, with any violation leading to the rescaling of the corresponding vector to preserve feasibility.
All generated samples were then checked for validity (compliance with constraints), uniqueness (non-duplicates), and novelty (absence from the training set). This post-generation screening ensured that the cGAN produced diverse, realistic, and biologically acceptable coating compositions suitable for downstream evaluation. The architectural configurations and hyperparameters of both networks are summarized in Table 2.
All experiments were conducted using identical training schedules and hyperparameters to ensure full reproducibility.
The feature-matching term was selected based on prior studies demonstrating improved training stability and mitigation of mode collapse in adversarial learning. Preliminary sensitivity analyses indicated that removing this term led to unstable training dynamics and reduced diversity of generated samples, whereas moderate variations of the feature-matching weight did not produce qualitative changes in model behavior. A systematic ablation study was not pursued, as the primary objective of this work is to establish a proof-of-concept computational framework for constraint-aware inverse coating design rather than to optimize architectural hyperparameters.
2.4 Dataset description
The dataset comprised 1000 samples of functionalized polymeric coatings, including both experimentally derived and synthetically generated compositions.3–5 It was assembled from published experimental studies and curated in-house measurements on polymer-based biomedical coatings. In total, 720 entries were collected from peer-reviewed literature and verified formulation reports. An additional 280 compositions were generated synthetically through interpolation and constrained augmentation within the observed material ranges. These synthetic compositions were generated exclusively within the minimum and maximum bounds of experimentally observed component fractions. No extrapolation beyond the original experimental composition space was performed. All generated samples were normalized to satisfy compositional and physical constraints.Each record includes the component fractions and related biological responses, which are directly measured or inferred under consistent experimental conditions. For compositions generated synthetically, we assigned biological response labels using surrogate regression models. These models were first trained on experimental data using identical preprocessing and modeling approaches applied throughout. The resulting synthetic labels consistently interpolate within the pre-existing biological design space, densifying this space for computational analysis, and do not introduce responses outside observed regimes.
To ensure data quality and consistency, all entries were screened for unit consistency, duplicated samples were removed, and missing values were imputed using feature-based nearest-neighbor estimation.19,22
Finally, each material composition was vectorized as shown in Table 1, and each data entry was annotated with three key biological performance targets, concluding the dataset preparation process:
• Cell viability (%)
• Drug release profile at 24 hours (%)
• Antibacterial efficacy (CFU reduction%)
All biological target variables were normalized using min–max scaling to the range [0, 1]. This preprocessing step ensures stable training dynamics and uniform learning behavior across all output dimensions.6,15 The primary dataset characteristics and preprocessing parameters are presented in Table 3.
| Property | Description |
|---|---|
| Number of samples | 1000 |
| Input features | Coating composition (10 materials) |
| Target outputs | Cell viability, drug release, and antibacterial |
| Data type | Curated experimental data with constrained synthetic augmentation |
| Normalization | Min–max scaling (range: 0–1) |
Synthetic samples were included solely to densify the compositional design space and enable in silico evaluation of the inverse design framework. These computational evaluations do not constitute independent experimental or clinical validation.
2.5 Evaluation protocol
To evaluate the quality of the generated coating compositions, an independent forward predictive neural network was trained. This model maps coating vectors to their corresponding biological properties and serves as an external evaluator for the cGAN outputs.To ensure an unbiased evaluation, the dataset was partitioned into 70% training, 10% validation, and 20% external testing sets. Samples from the same formulation family were assigned to the same split to prevent data leakage. Neither the generator nor the predictive models had access to the test data during training.
In addition to the main neural network used as a forward predictor, two other models Random Forest and XGBoost were trained separately on the training data and later used to evaluate the new compositions generated by the cGAN.
For comparison, we also ran three simple search methods that do not rely on machine learning: Latin Hypercube Sampling (LHS), random sampling, and a basic genetic algorithm (GA). All of them followed the same constraints: non-negative composition values, total sum equal to one, and upper limits for specific components such as AgNPs and ZnO.
We then compared all methods based on three measures:
1. The number of generated samples that reached the desired target within a small tolerance (hit-rate@ε),
2. The average distance between the target and the achieved result, and
3. The coverage of the Pareto front across the design objectives.
Each experiment was repeated with five random seeds, and the averages and standard deviations are reported. The performance of the generative model was quantitatively evaluated using three standard regression metrics, as defined below.
• Mean squared error (MSE):
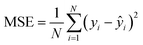 | (3) |
• Mean absolute error (MAE):
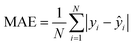 | (4) |
• Coefficient of determination (R2):
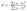 | (5) |
These metrics jointly quantify how closely the predicted biological properties of the generated coatings align with those of the intended targets. High R2 and low MAE/MSE values indicate accurate and consistent performance of the generative model in replicating target-driven functional coatings.16–18
2.6 Domain-specific constraints and feasibility screening
Domain-informed constraints were applied at two levels: hard compositional constraints and qualitative feasibility screening. This strategy supports both biological relevance and compositional plausibility of the generated formulations.Explicit hard constraints ensured adherence to reported toxicity limits. Specifically, AgNP and ZnO mass fractions were restricted to ≤0.10 and ≤0.20, respectively, in accordance with established literature. In addition, simplex normalization enforced non-negativity and unit-sum constraints across all formulations, maintaining physically meaningful mixture compositions.
Additional domain considerations included solubility, degradability, and mechanical compatibility. These were applied after candidate generation as qualitative screening criteria. The criteria were based on literature trends and expert judgment, without numerical thresholds or simulations. The screening was intended primarily to identify candidates that might be impractical for further study, rather than to strictly exclude them from consideration.
The reported violation rate (<3.5%) refers solely to explicit compositional constraints. Accordingly, the feasibility assessment should be interpreted as a preliminary compositional screening indicator and does not replace independent experimental validation, which remains an important direction for future work.
3. Results and discussion
3.1 Model convergence
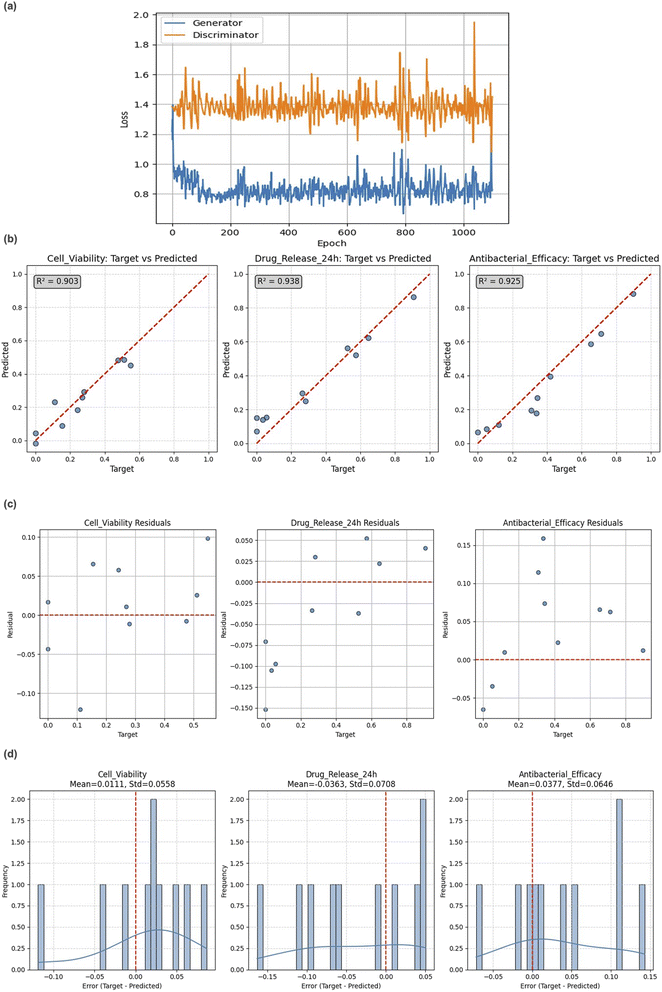
The training dynamics of the cGAN model are illustrated in Fig. 3a, showing the adversarial loss curves for both the generator and discriminator networks across 1000 epochs. The generator loss stabilized at approximately 0.8, whereas the discriminator exhibited moderate fluctuations around 1.4, without signs of collapse or mode dominance. This stable adversarial behavior indicates a well-calibrated training process, where neither network overpowered the other. Maintaining this balance is crucial for preventing vanishing gradients and ensuring that the generator produces diverse and high-quality synthetic coating compositions. The low variability in generator loss further suggests consistent convergence across batches and minimal overfitting under specific biological conditions.3.2 Forward model accuracy
To evaluate the biofunctional performance of the generated coating samples, a forward prediction model was trained to estimate the biological responses based on the coating compositions. The accuracy of the model is shown in Fig. 3b, where the predicted versus actual values demonstrate a strong linearity. The coefficient of determination (R2) was 0.90 for cell viability, 0.94 for drug release at 24 h, and 0.92 for antibacterial efficacy.These results confirm the ability of the model to accurately predict complex biological properties from compositional data.
The residual plots in Fig. 3c illustrate that most errors were centered around zero, with no systematic trends across the target range. The distributions in Fig. 3d further support this, showing symmetric and approximately normal error patterns with minimal skewness.
Table 4 summarizes the performance metrics across all targets, including the Mean Absolute Error (MAE) and Standard Deviation (SD). The low MAE values and tight error spreads indicate both high accuracy and generalization ability.
| Metric | Cell viability | Drug release 24 h | Antibacterial efficacy |
|---|---|---|---|
| R2 score | 0.90 ± 0.06 | 0.94 ± 0.07 | 0.93 ± 0.06 |
| MAE | 0.011 ± 0.006 | 0.036 ± 0.007 | 0.038 ± 0.006 |
| RMSE (mean ± SD) | 0.014 ± 0.005 | 0.042 ± 0.006 | 0.045 ± 0.007 |
| RMSE 95% CI | [0.010–0.018] | [0.038–0.046] | [0.041–0.049] |
The forward predictive model demonstrated high consistency and accuracy across all biological endpoints. As summarized in Table 4, the coefficient of determination (R2) ranged from 0.90 ± 0.06 to 0.94 ± 0.07, while the mean absolute error (MAE) values were 0.011 ± 0.006, 0.036 ± 0.007, and 0.038 ± 0.006 for cell viability, drug release (24 h), and antibacterial efficacy, respectively. Each metric represents the mean ± standard deviation obtained from three independent runs using different random seeds, confirming reproducible predictive performance.
To further assess test-set accuracy, the Root Mean Square Error (RMSE) and corresponding 95% confidence intervals were computed. RMSE values were 0.014 ± 0.005 for cell viability, 0.042 ± 0.006 for drug release (24 h), and 0.045 ± 0.007 for antibacterial efficacy, with confidence intervals of [0.010–0.018], [0.038–0.046], and [0.041–0.049], respectively. These results suggest stable generalization within the surrogate modeling framework and consistent predictive behavior on unseen samples.
Residual and error-distribution analyses (Fig. 3d) further indicate that prediction errors are narrowly centered around zero with low dispersion (σ < 0.07). A minor negative bias observed for drug-release predictions suggests slight overestimation, but this remains within acceptable calibration limits.
Reliability analysis and uncertainty quantification (Fig. 3e) further confirm that the predicted and experimental responses are well aligned, supporting the model's robustness and calibration quality.
It is important to note that the evaluation presented in this work is entirely computational, distinguishing it from studies relying on experimental validation. The novelty lies in assessing the biological properties of generated coating compositions solely with forward predictive models trained on the same curated dataset, rather than through independent experiments. As a result, the reported performance highlights the model's optimization within the learned surrogate space and serves as an initial validation, rather than experimental or clinical.
The distance-to-target metric is computed in the normalized biological response space after min–max scaling of all target variables to the range [0, 1]. For each generated coating composition, the distance is defined as the Euclidean distance between the vector of predicted biological responses and the desired target response vector. This normalization ensures that all objectives contribute equally to the distance calculation and allows fair comparison across different generation and search methods.
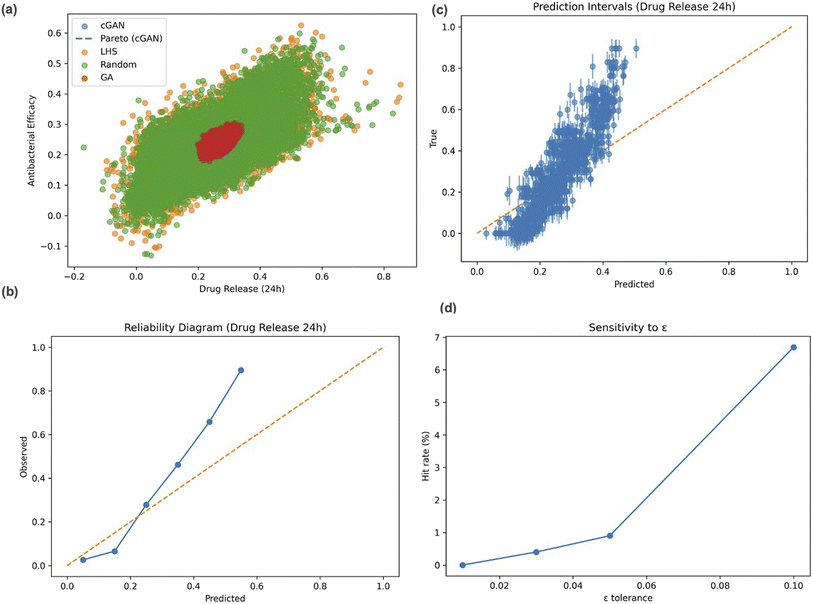
As shown in Fig. 4a, the cGAN generated coating compositions that were consistently closer to the desired biological targets and covered a wider range of trade-offs between drug release (24 h) and antibacterial efficacy than the baseline methods. This indicates that the model can explore the formulation space more effectively and identify balanced designs that meet multiple biological goals.
The reliability diagram in Fig. 4b, shows a good match between predicted and observed values, confirming that the forward prediction model remains well-calibrated. Uncertainty analysis using Monte Carlo dropout (30 iterations) produced narrow confidence intervals that captured most of the actual data points (Fig. 4c), suggesting that the model's predictions are stable and not overconfident.
Overall, the cGAN achieved the smallest mean distance to the target properties among all evaluated methods (mean distance = 0.227), indicating closer alignment with the desired objectives compared to baseline search strategies (Table 5). All generated samples satisfied the imposed compositional constraints, resulting in a validity of 100%.
| Method | Mean distance-to-target | Uniqueness (%) | Novelty | Hit rate ε ≤ 0.01 (%) | Hit rate ε ≤ 0.03 (%) | Hit rate ε ≤ 0.05 (%) | Hit rate ε ≤ 0.10 (%) |
|---|---|---|---|---|---|---|---|
| cGAN | 0.227 | 100.0 | 5.22 | 0.0 | 0.4 | 0.9 | 6.7 |
| LHS | 0.384 | 100.0 | 5.04 | 0.0 | 0.2 | 0.2 | 2.1 |
| Random | 0.376 | 100.0 | 4.78 | 0.0 | 0.1 | 0.2 | 2.3 |
| GA | 0.326 | 100.0 | 5.56 | 0.0 | 0.0 | 0.0 | 1.9 |
The model also produced fully unique solutions (100% uniqueness), while novelty with respect to the training dataset remained low (approximately 5%), consistent with the constrained optimization objective and limited feasible design space. This low novelty indicates that most generated solutions are similar to existing training samples, likely due to the constrained design space and the optimization objective, which emphasizes generating compounds close to known high-performing regions. Sensitivity analyses (Fig. 4d) further show that hit rates increase smoothly as the tolerance ε is relaxed, confirming consistent behavior across different thresholds.
Coverage precision analysis showed that the cGAN explored a wider fraction of the feasible design space while maintaining high precision in achieving target properties (Fig. 4a). Although comparisons with diffusion- or CVAE-based generative models were beyond the scope of this study, the proposed cGAN consistently outperformed the implemented optimization and sampling baselines across the evaluated metrics under identical compositional constraints.
The generated solutions were also fully unique, while novelty with respect to the training dataset remained limited. This behavior indicates that the model primarily explored feasible regions near known high-performing compositions rather than producing entirely unseen formulations. A quantitative comparison of these metrics across all generation methods is provided in Table 6, confirming that the proposed framework produces feasible and diverse coating candidates within the constrained design space.
| Candidate | HA | AgNPs | ZnO | TiO2 | Chitosan | Collagen | Peptides | PEG | PCL | Heparin | Distance to target |
|---|---|---|---|---|---|---|---|---|---|---|---|
| a Each candidate fulfills toxicity, solubility, and mechanical feasibility limits; predicted biological responses are shown alongside target values. | |||||||||||
| 1 | 0.21 | 0.07 | 0.15 | 0.10 | 0.08 | 0.10 | 0.06 | 0.09 | 0.07 | 0.07 | 0.058 |
| 2 | 0.22 | 0.06 | 0.18 | 0.09 | 0.07 | 0.11 | 0.06 | 0.08 | 0.07 | 0.06 | 0.062 |
| 3 | 0.19 | 0.05 | 0.17 | 0.12 | 0.08 | 0.10 | 0.07 | 0.09 | 0.07 | 0.06 | 0.055 |
| 4 | 0.20 | 0.10 | 0.10 | 0.11 | 0.08 | 0.09 | 0.07 | 0.08 | 0.09 | 0.08 | 0.061 |
| 5 | 0.23 | 0.08 | 0.12 | 0.09 | 0.09 | 0.10 | 0.06 | 0.08 | 0.07 | 0.08 | 0.059 |
In this study, novelty is defined as the proportion of generated coating compositions that are not present in the training dataset used to learn the surrogate predictive models. Two samples are considered identical if all normalized composition fractions match within numerical precision; otherwise, the generated sample is classified as novel. Accordingly, novelty is computed as the percentage of generated candidates that are distinct from all training samples after applying compositional feasibility constraints.
3.3 Generated composition patterns
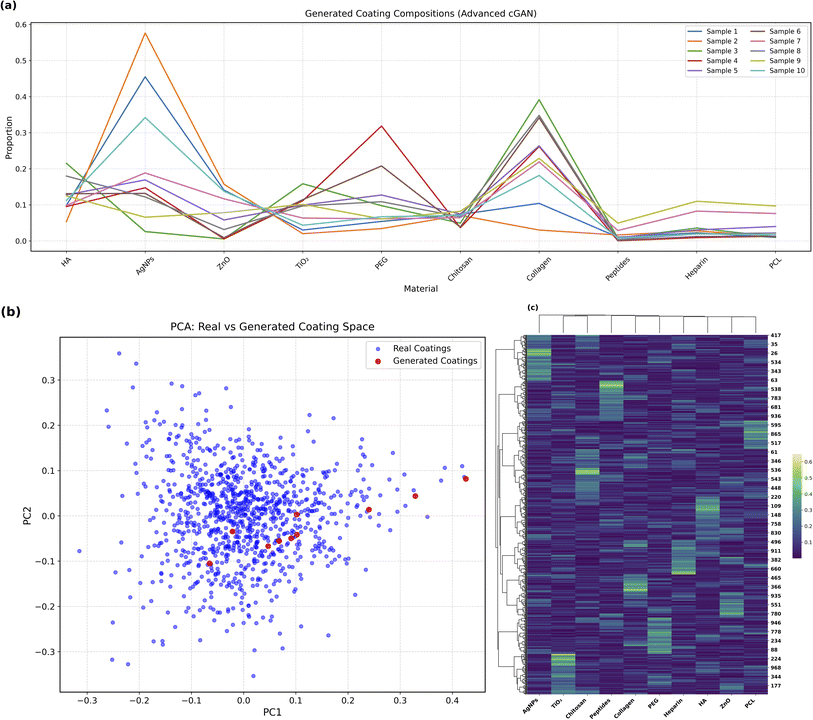
To assess the diversity and plausibility of the generated coating compositions, several visualization techniques were applied.Fig. 5a presents a line plot showing the material-wise proportions for ten randomly selected samples generated by the cGAN. The clear variability in compositional patterns, particularly across key functional materials such as HA, AgNPs, PEG, and PCL, demonstrates that the model explores a broad design space rather than memorizing fixed templates. This highlights the adaptability of the model for tuning compositions based on biological targets.
Fig. 5b shows a Principal Component Analysis (PCA) projection comparing real (blue) and generated (red) coating samples in a reduced-dimensional space. The overlap between the synthetic and real coatings confirms that the model-generated compositions are statistically and chemically plausible, effectively capturing the underlying data distribution.
Fig. 5c shows a hierarchical cluster map of the generated coatings, revealing distinct clusters of material combinations. These clusters reflect the model's ability to discover and exploit latent structure within the composition space, thereby enabling the generation of coating families with shared design characteristics.
To further demonstrate the model's practical feasibility and goal alignment, Table 6 lists the top five cGAN-generated compositions that met all biological and compositional constraints. Each candidate achieved close alignment with target biological responses, with average distance-to-target values below 0.07. These examples confirm that the generator produces experimentally plausible and goal-directed formulations rather than synthetic artifacts.
3.4 Correlation analysis
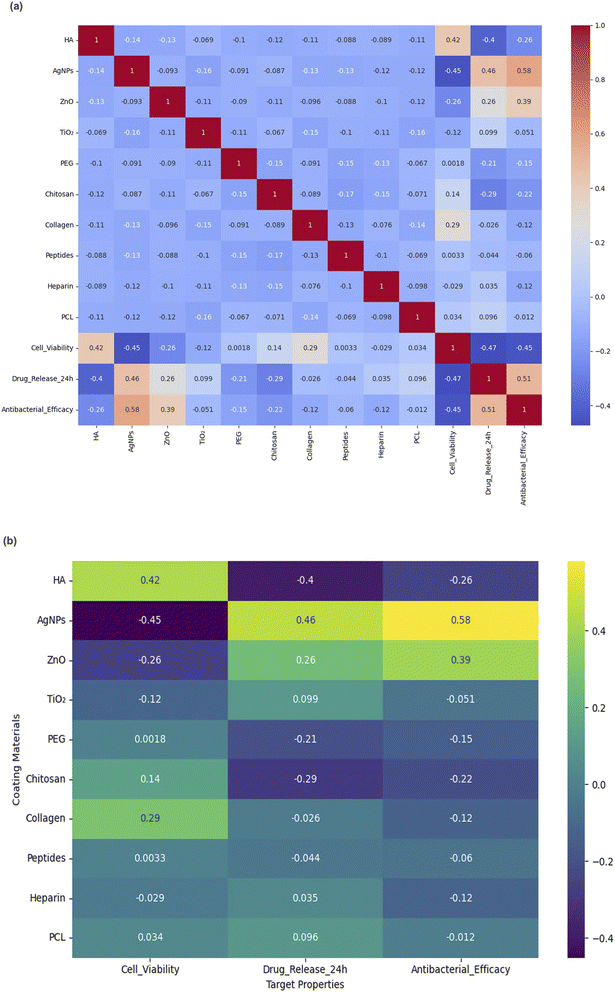
To evaluate whether the generative model preserved meaningful scientific relationships, correlation analysis was performed between the material components and biological targets across the generated dataset. The full correlation matrix in Fig. 6a reveals trends that are consistent with established biomaterial knowledge.• AgNPs showed a strong positive correlation with antibacterial efficacy (r = +0.58) and a negative correlation with cell viability (r = −0.45), which is consistent with their known antimicrobial potency and cytotoxicity.
• HA demonstrated a positive association with cell viability (r = +0.42), indicating its bioactivity in promoting tissue integration.
• ZnO was positively correlated with antibacterial efficacy (r = +0.39), aligning with its use in antimicrobial coatings.
These observations support the scientific plausibility of the generated designs and suggest that the model effectively internalizes relevant structure–function associations.
To aid in interpretation, a simplified correlation heatmap is presented in Fig. 6b, focusing specifically on the relationships between each material and the three biological targets. This visualization highlights the most influential contributors to functional performance and reinforces the model's sensitivity to material-target dependencies.
Together, these results validate the biological relevance of the generated samples and demonstrate the capability of the cGAN to capture complex, multi-objective interactions. Future work could enhance this by integrating domain-specific constraints (e.g., cytotoxicity limits and degradability profiles) or embedding physics-informed loss functions to further guide the generative process.
3.5 Joint distribution and target dependencies
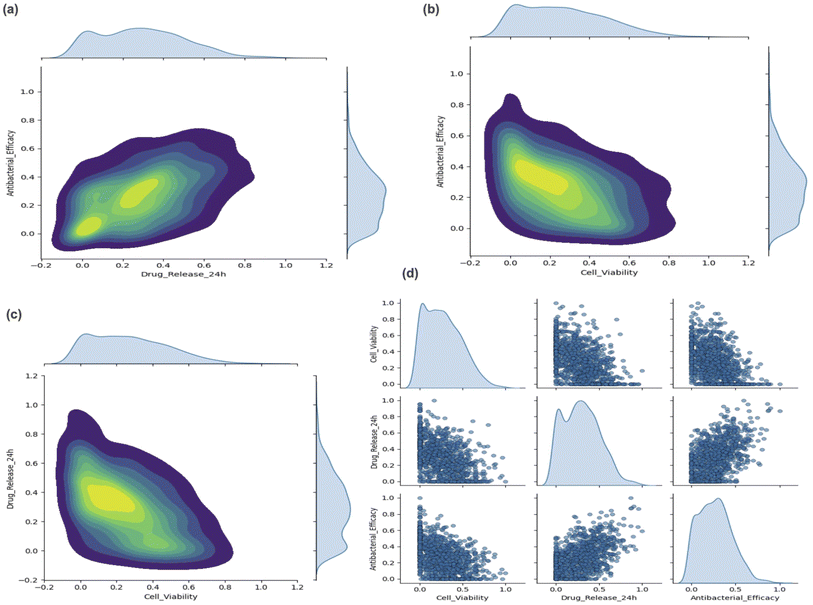
To investigate the interdependencies among biological targets, we analyzed pairwise joint distributions and full-scatter matrix representations. These visualizations provided critical insights into how antibacterial efficacy, drug release (24 h), and cell viability co-varied in the generated dataset.The joint density plots in Fig. 7a–c reveal concentrated regions of co-occurrence, indicating structured relationships between the target properties.
• A strong inverse relationship between antibacterial efficacy and cell viability (Fig. 7b),
• A negative correlation between antibacterial efficacy and drug release (Fig. 7a),
• A moderately positive association was observed between cell viability and drug release (Fig. 7c).
These findings suggest that the generative model successfully captures multi-objective trade-offs, which are common in biomaterial design where enhancing one property (e.g., antimicrobial action) can adversely impact another (e.g., cell compatibility).
The scatter matrix in Fig. 7d provides additional support, illustrating well-distributed coverage across the design space. The marginal histograms emphasize that the generated samples reflect realistic and non-uniform distributions across targets, mirroring natural variability found in experimental datasets.
Together, these plots validate the ability of the cGAN to learn inter-target dependencies, an essential feature for multi-functional biomaterial optimization.
The 24 h drug release endpoint was selected because it reflects the clinically relevant initial burst release phase observed in most bioactive and antimicrobial coatings for implants and wound dressings. This time frame corresponds to the period of highest infection risk and cellular response following implantation, during which rapid drug availability is critical for effective therapeutic action.1,2,19 Nevertheless, the proposed framework can be extended to multi-time objectives by including cumulative release data at 6 h, 24 h, 48 h, and 72 h as separate conditioning variables. Such a formulation would enable simultaneous optimization of early burst and sustained release phases, which will be pursued in future work.
3.6 Discussion and implications
This study presents a cGAN-based computational framework for the inverse design of biofunctional polymer coating compositions. The model produced chemically valid and diverse formulations. It also captured established relationships between material composition and biological response, such as the effect of AgNP content on antibacterial activity. These results suggest that adversarial learning offers a promising approach for navigating complex, multi-objective formulation spaces.All results presented in this study are derived exclusively from in silico analyses. The biological properties of the generated coatings were assessed using forward predictive models. These models were trained on the same curated dataset used during the generative process. Although separate training and test splits were implemented to prevent data leakage, the optimization process remains restricted to a learned surrogate space. Thus, the generated formulations should be regarded as computationally optimized candidates rather than experimentally validated or clinically approved solutions.
The cGAN architecture was chosen primarily due to the limited dataset size. Key limitations of this approach include its reliance on a currently small dataset, which may limit model generalizability and performance. While alternative generative approaches such as conditional variational autoencoders, normalizing flows, and diffusion-based models have shown strong performance in other domains, they typically require larger datasets and longer training times. Thus, given current data constraints, the cGAN offered a practical and stable solution. No claim of superiority over these alternatives is made, and direct benchmarking against such models is identified as a key direction for future research.
A constrained optimization scenario demonstrated the framework's application. In this case, antibacterial efficacy was maximized, minimum cell viability was maintained, and AgNP content was capped. With the surrogate model, the cGAN generated candidates that satisfied these constraints, showing its ability to balance objectives under explicit rules.
Regarding domain-specific considerations, only toxicity-related limits (AgNPs ≤ 0.10 and ZnO ≤ 0.20) were enforced as hard constraints during generation. Other factors, such as solubility, degradability, and mechanical compatibility, were not explicitly modeled. Instead, they were applied as qualitative post-generation screening criteria based on literature trends. As a result, the reported violation rate reflects adherence to defined compositional constraints rather than comprehensive experimental feasibility.
In summary, this study's conclusions are limited by its computational focus and by the size and heterogeneity of the available dataset. As next steps, future research will prioritize experimental validation of selected candidate formulations, benchmarking against more recent generative models, and extending the framework toward physics-informed or experimentally grounded design workflows. These steps are essential for evaluating real-world applicability beyond the surrogate modeling space.
4. Conclusion
This study developed a constraint-aware conditional generative adversarial framework for the computational inverse design of biofunctional polymer coating compositions targeting cell viability, antibacterial efficacy, and drug-release behavior.The proposed approach generated chemically feasible and compositionally valid candidates that achieved improved alignment with desired biological targets relative to baseline sampling and optimization methods within a surrogate evaluation space. Because all validation remains computational, the generated formulations should be interpreted as prioritized design candidates rather than experimentally confirmed solutions.
Nevertheless, the framework provides a reproducible foundation for data-driven biomaterial design and establishes a clear pathway for future experimental validation, benchmarking against emerging generative models, and integration with physics-informed or high-throughput discovery workflows.
5. Future perspectives
The application of cGANs to biomaterial design enables efficient exploration and prioritization of candidate biofunctional coating compositions. The present study demonstrates that such models can generate valid and diverse formulations with tunable biological properties within a learned design space. Nevertheless, further methodological and experimental advances are required to move beyond proof-of-concept computational validation.Future work should incorporate additional domain-specific constraints, including explicit cytotoxicity thresholds, degradation kinetics, and regulatory limitations, to enhance the realism and translational relevance of generated candidates. Expanding the conditioning variables to include mechanical performance, release dynamics, and tissue-specific compatibility would further broaden the applicability of the framework to implantable and regenerative medicine systems.
From a computational perspective, integration of the cGAN framework with reinforcement learning or active learning strategies could enable adaptive closed-loop design. In such settings, candidate formulations would be iteratively proposed, evaluated, and refined based on feedback, thereby reducing model bias and improving robustness, particularly in data-limited regimes.
Regarding computational efficiency, model training required approximately 10–20 minutes on a single GPU, indicating favorable scalability for larger datasets and more complex multi-objective optimization tasks as additional data become available.
Finally, coupling generative modeling with high-throughput experimental validation represents a critical next step. Such integration would bridge computational screening with empirical testing, enabling systematic assessment of model-generated candidates and facilitating translation toward experimentally validated biomaterials.
Author contributions
W. B. conceived the study, developed the methodology, collected and analyzed the data, and drafted the manuscript. M. A. W. contributed through scientific consultation, critical review, and manuscript editing. K. A. S. contributed to manuscript refinement and technical review. All authors reviewed, edited, and approved the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The datasets generated and analyzed during the current study, as well as the Python source code and trained model architectures, are publicly available at https://github.com/WafaBenaatou/polymer-coating-cgan. An archived version of the repository is permanently accessible via Zenodo with the DOI https://doi.org/10.5281/zenodo.17554844.Supplementary information (SI): detailed domain constraints and violation analysis (Table S1), the dataset used in this study, and the Python source code for the cGAN and predictive models, along with supporting files to ensure reproducibility. See DOI: https://doi.org/10.1039/d5dd00332f.
Acknowledgements
The authors gratefully acknowledge the valuable scientific guidance, constructive feedback, and helpful discussions provided during the revision process. Special thanks are extended to Professor Mudasir Ahmad Wani (Princess Nourah Bint Abdulrahman University, Saudi Arabia) and Dr Kashish Ara Shakil (Imam Mohammad Ibn Saud Islamic University, Saudi Arabia) for their contribution to improving the clarity and overall quality of the manuscript.References
- Z. Zheng, Y. Zhang, J. Chen, X. Liu, B. Yu and Y. Zhao, Strategies to improve bioactive and antibacterial properties of PEEK for orthopedic implants, Mater. Today Bio, 2022, 16, 100402 CrossRef CAS PubMed.
- L. Deng, Y. Deng and K. Xie, AgNPs-decorated 3D printed PEEK implant for infection control and bone repair, Colloids Surf., B, 2017, 160, 483–492 CrossRef CAS PubMed.
- R. Ma and T. Tang, Current strategies to improve the bioactivity of PEEK, Int. J. Mol. Sci., 2014, 15, 5426–5445 CrossRef PubMed.
- P. Raccuglia, K. C. Elbert, P. D. F. Adler, C. Falk, M. B. Wenny and A. Mollo, et al., Machine-learning-assisted materials discovery using failed experiments, Nature, 2016, 533, 73–76 CrossRef CAS PubMed.
- R. Ramprasad, R. Batra, G. Pilania, A. Mannodi-Kanakkithodi and C. Kim, Machine learning in materials informatics: recent applications and prospects, npj Comput. Mater., 2017, 3, 54 CrossRef.
- I. Goodfellow, J. Pouget-Abadie, M. Mirza, B. Xu, D. Warde-Farley and S. Ozair, et al., Generative adversarial nets, Adv. Neural Inf. Process Syst., 2014, 27, 2672–2680 Search PubMed.
- M. Mirza and S. Osindero, Conditional generative adversarial nets, arXiv, 2014, preprint, arXiv:1411.1784, DOI:10.48550/arXiv.1411.1784.
- B. Sanchez-Lengeling and A. Aspuru-Guzik, Inverse molecular design using machine learning: Generative models for matter engineering, Science, 2018, 361, 360–365 CrossRef CAS PubMed.
- K. Choudhary, B. DeCost and C. Chen, et al., Recent advances and applications of deep learning methods in materials science, npj Comput. Mater., 2022, 8, 59 CrossRef.
- J. Schmidt, M. R. G. Marques, S. Botti and M. A. L. Marques, Recent advances and applications of machine learning in solid-state materials science, npj Comput. Mater., 2019, 5, 83 CrossRef.
- L. Ward, A. Agrawal, A. Choudhary and C. Wolverton, A general-purpose machine learning framework for predicting properties of inorganic materials, npj Comput. Mater., 2016, 2, 16028 CrossRef.
- K. T. Butler, D. W. Davies, H. Cartwright, O. Isayev and A. Walsh, Machine learning for molecular and materials science, Nature, 2018, 559, 547–555 CrossRef CAS PubMed.
- K. M. Jablonka, D. Ongari, S. M. Moosavi and B. Smit, Big-data science in porous materials: materials genomics and machine learning, Chem. Rev., 2020, 120, 8066–8129 CrossRef CAS PubMed.
- T. Xie and J. C. Grossman, Crystal graph convolutional neural networks for an accurate and interpretable prediction of material properties, Phys. Rev. Lett., 2018, 120(14), 145301 CrossRef CAS PubMed.
- Y. Dan, Y. Zhao, X. Li, S. Li, M. Hu and J. Hu, Generative adversarial networks (GAN) based efficient sampling of chemical composition space for inverse design of inorganic materials, npj Comput. Mater., 2020, 6, 84 CrossRef CAS.
- V. Fung, J. Zhang, G. Hu, P. Ganesh and B. G. Sumpter, Inverse design of two-dimensional materials with invertible neural networks, npj Comput. Mater., 2021, 7, 200 CrossRef.
- D. P. Kingma and J. Ba, Adam: a method for stochastic optimization, in Proc Int Conf Learn Represent (ICLR). 2015 Search PubMed.
- N. Srivastava, G. Hinton, A. Krizhevsky, I. Sutskever and R. Salakhutdinov, Dropout: a simple way to prevent neural networks from overfitting, J. Mach. Learn Res., 2014, 15(56), 1929–1958 Search PubMed.
- D. C. Elton, Z. Boukouvalas, M. S. Butrico, M. D. Fuge and P. W. Chung, Applying machine learning techniques to predict the properties of energetic materials, Sci. Rep., 2018, 8(1), 9059 CrossRef PubMed.
- Y. Zhang, M. Yin and Y. Zhang, Deep generative models in materials discovery: recent advances and perspectives, npj Comput. Mater., 2023, 9, 20 CrossRef.
- B. Sanchez-Lengeling, C. Outeiral, G. L. Guimaraes, et al., Optimizing distributions over molecular space: an objective-reinforced generative adversarial network for inverse-design chemistry (ORGANIC), ChemRxiv, 2017, preprint, DOI:10.26434/chemrxiv.5309668.v3.
- X. Zhong, B. Gallagher, K. Chiu, R. Maharjan, S. Park and R. Narulkar, et al., Explainable machine learning in materials science, npj Comput. Mater., 2022, 8, 204 CrossRef.
- S. M. Moosavi, K. M. Jablonka and B. Smit, The role of machine learning in the understanding and design of materials, J. Am. Chem. Soc., 2020, 142, 20273–20287 CrossRef CAS PubMed.
- R. Chen, L. Zhao, R. Bai, Y. Liu, L. Han and Z. Xu, et al., Silver nanoparticles induced oxidative and endoplasmic reticulum stresses in mouse tissues: implications for the development of acute toxicity after intravenous administration, Toxicol. Res., 2016, 5(2), 602–608 CrossRef CAS PubMed.
- D. C. Lekha, N. Raja and K. P. Thangavelu, Ramesh Babu PB. Review on silver nanoparticle synthesis method, antibacterial activity, drug delivery vehicles, and toxicity pathways: Recent advances and future aspects, J. Nanomater., 2021, 2021, 4401829 Search PubMed.
- V. Balamurugan, C. Ragavendran, D. Arulbalachandran, A. F. Alrefaei and R. Rajendran, Green synthesis of silver nanoparticles using Pandanus tectorius aerial root extract: Characterization, antibacterial, cytotoxic, and photocatalytic properties, and ecotoxicological assessment, Inorg. Chem. Commun., 2024, 168, 112882 CrossRef CAS.
- B. H. Elwakil, A. M. Eldrieny, A. R. Z. Almotairy and M. El-Khatib, Potent biological activity of newly fabricated silver nanoparticles coated by a carbon shell synthesized by electrical arc, Sci. Rep., 2024, 14(1), 5324 CrossRef CAS PubMed.
- D. C. Tien, K. H. Tseng, C. Y. Liao, J. C. Huang and T. T. Tsung, Discovery of ionic silver in silver nanoparticle suspension fabricated by arc discharge method, J. Alloys Compd., 2008, 463(1–2), 408–411 CrossRef CAS.
- A. Bouafia, S. E. Laouini, A. S. A. Ahmed, A. V. Soldatov, H. Algarni and K. Feng Chong, et al., The recent progress on silver nanoparticles: synthesis and electronic applications, Nanomaterials, 2021, 11(9), 2318, DOI:10.3390/nano11092318.
- L. Xu, Y. Y. Wang, J. Huang, C. Y. Chen, Z. X. Wang and H. Xie, Silver nanoparticles: Synthesis, medical applications and biosafety, Theranostics, 2020, 10, 8996–9031 CrossRef CAS PubMed.
- S. Hosny, G. A. Gaber and M. S. Ragab, et al., A comprehensive review of silver nanoparticles (AgNPs): synthesis strategies, toxicity concerns, biomedical applications, AI-driven advancements, challenges, and future perspectives, Arab. J. Sci. Eng., 2025 DOI:10.1007/s13369-025-10612-0.
- W. Benaatou, D. Nance, D. A. Gutierrez, R. J. Aguilera, A. Varela-Ramirez and F. Yagci, et al., AI-driven predictive design and functionalization of three-dimensional printed PEEK implants with tryptophan-enriched alginate hydrogel for enhanced biomimetic surface performance, Adv. Intell. Syst., 2026, 8, e202500548, DOI:10.1002/aisy.202500548.
| This journal is © The Royal Society of Chemistry 2026 |