Open Access Article
Open Access ArticleMSIGN: A deep learning framework based on multi-scale interaction graph neural networks for predicting binding of synthetic cannabinoids to receptors
Zhenyong
Cheng†
a,
Dinghao
Liu†
 a,
Yuanpeng
Fu
a,
Yuanpeng
Fu
 a,
Kewei
Sheng
a,
Yan
Xing
a,
Yanling
Qiao
bcd,
Shangxuan
Cai
ef,
Jubo
Wang
d,
Peng
Xu
bcd,
Bin
Di
a,
Kewei
Sheng
a,
Yan
Xing
a,
Yanling
Qiao
bcd,
Shangxuan
Cai
ef,
Jubo
Wang
d,
Peng
Xu
bcd,
Bin
Di
 cd and
Jun
Liao
cd and
Jun
Liao
 *acg
*acg
aSchool of Science, China Pharmaceutical University, Nanjing 211198, China. E-mail: liaojun@cpu.edu.cn; Tel: +086-86185160
bKey Laboratory of Drug Monitoring and Control, Drug Intelligence and Forensic Center, Ministry of Public Security, Beijing 100193, China
cOffice of China National Narcotics Control Commission, China Pharmaceutical University Joint Laboratory on Key Technologies of Narcotics Control, Beijing 100193, China
dSchool of Pharmacy, China Pharmaceutical University, Nanjing 211198, China
eState Key Laboratory of Membrane Biology, Peking University School of Life Sciences, Beijing 100871, China
fPKU-IDG/McGovern Institute for Brain Research, Beijing 100871, China
gZhejiang Lab, Hangzhou 311500, China
First published on 17th February 2026
Abstract
Deep learning-based models have been extensively applied to the task of protein–ligand binding affinity (PLA) prediction. Current 3D ligand-complex-based GNNs, though advanced, still struggle with accuracy and generalization due to their overreliance on atomic-level physical features and their neglect of chemical space dynamics, leading to data memorization rather than robust learning. To address these issues, we propose a deep learning model based on a Multi-Scale Interaction Graph Neural Network (MSIGN). By constructing ligand functional group graphs and protein amino acid graphs, we introduce chemical information features into the model, which are combined with physical features to enhance binding affinity prediction. Especially, we innovatively adopt a pre-training and fine-tuning training approach in the PLA domain to improve the model's generalization capability on downstream tasks (this study focuses on the binding affinity prediction of synthetic cannabinoids), and we validated the MSIGN model predictions with wet experiments such as SPR on three novel synthetic cannabinoids. Furthermore, we analyze the impact of different fine-tuning strategies on the model's generalization ability. Multiple results collectively demonstrate the superiority of our MSIGN model design, providing a novel approach for future PLA prediction.
1. Introduction
Driven by the exponential growth of structural and binding data, AI-based protein–ligand affinity prediction has advanced rapidly.1–5 Current representation methods are categorized into 3D-independent and 3D-dependent approaches. 3D-independent methods include 1D sequence-based models, such as DeepGLSTM,2,6,12 which capture residue-level context, and 2D graph-based representations.7–11 Among them, SIGN13 utilizes the Graph Attention Mechanism (GAT) to model atomic interactions between protein residues and ligand atoms. To better reflect binding dynamics, models like EGNA,14 further integrate spatial distances into dynamic interaction graphs, bridging the gap between topological information and 3D spatial relationships.Despite their utility, 1D and 2D representations often neglect critical 3D spatial interactions.15 To address this, 3D-dependent methods like voxel-based approaches16–20 extract features from atomic coordinates; however, they often fail to capture explicit interatomic bonding or align with real-world physical principles.21 Consequently, 3D graph structures, which enrich 2D topologies with spatial distance and angle information, have emerged as the mainstream approach for more precise feature representation.
Recent studies reveal that models trained on atomic features often capture dataset biases rather than fundamental physical principles,15 frequently over-relying on ligand structures while neglecting protein–ligand interactions. To enhance generalizability and interpretability, Moon et al.21 integrated physical priors by decomposing binding affinity into specific pairwise interactions (e.g., van der Waals forces and hydrogen bonds). Building on this, Yang et al.22 introduced a learnable bias correction term to further mitigate hidden systematic biases in the data.
While atom-centric methods provide fundamental structural insights,23 they struggle to capture higher-level functional similarities or the distribution of complexes in chemical space. To address these limitations, we propose a multi-scale model integrating physical and chemical priors. By incorporating molecular functional groups24,25 and amino acid residues,26 our model moves beyond individual atoms to capture essential chemical properties—such as reactivity, polarity, and hydrophobicity—that fundamentally govern protein–ligand interaction patterns.
Despite the success of deep learning in virtual screening, data dependency remains a significant bottleneck for targets with limited experimental data, such as G protein-coupled receptors (GPCRs).27 Cannabinoid receptor 1 (CB1R), a classical GPCR predominantly expressed in the brain, mediates critical central nervous system effects. Synthetic cannabinoids (SCs) act as potent full agonists of CB1R, exhibiting stronger psychoactive and addictive properties than their natural counterparts. Given that SC-CB1R affinity directly correlates with addictive potential, its accurate prediction is essential for pharmacological risk assessment.
To overcome the generalization limits of small-sample datasets,28 we propose a transfer learning strategy based on a pre-training and fine-tuning paradigm. A generalized MSIGN model is first pre-trained on the PDBbind dataset to capture fundamental laws of protein–ligand interactions. Subsequently, the model is fine-tuned on a specialized CB1R-SC dataset curated from BindingDB to acquire domain-specific knowledge. Wet-lab experiments, including SPR and fluorescence assays on three novel SCs, further validate that this multi-scale approach effectively bridges the gap between general physical rules and specific biological activity. In summary, the main contributions of this work are as follows:
(i) Multi-scale graph design: we propose MSIGN, a multi-scale framework that integrates atomic interaction graphs with ligand functional group and protein residue graphs. This design effectively fuses physical interaction features with chemical priors, enhancing both model accuracy and interpretability.
(ii) Systematic fine-tuning strategy: we conducted a comprehensive evaluation of transfer learning paradigms (General, No-pretraining, Frozen, and Full Fine-tuning). We demonstrate that the “pre-training + full fine-tuning” strategy is critical for overcoming domain shifts and achieving generalization in data-scarce GPCR tasks.
(iii) Experimental validation and benchmarking: we validated the model's practical utility through the wet-lab synthesis and biological assay of three novel synthetic cannabinoids. Cross-method benchmarking confirms that MSIGN achieves superior consistency with experimental results compared to physics-based docking and other state-of-the-art deep learning models.
2. Experimental
The methodological framework of our MSIGN is systematically illustrated in Fig. 1. We provide a comprehensive description of the pipeline, spanning dataset construction, feature engineering, and model architecture, with particular emphasis on the methodologically rigorous integration of physical-chemical priors and multi-scale representations.2.1 Dataset
This study utilized two public datasets, PDBbind v2020 (ref. 29) and BindingDB,30 to acquire general protein-small molecule ligand binding affinity data and CB1 receptor ligand binding affinity data, respectively. The distribution of labels for these two datasets is shown in Fig. 2A and B.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 127 complexes with experimentally determined binding affinities from the PDBbind website, wherein the binding affinities are represented as −log
127 complexes with experimentally determined binding affinities from the PDBbind website, wherein the binding affinities are represented as −log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kd, −log
Kd, −log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Ki, or −log
Ki, or −log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) IC50 (with larger values indicating stronger binding). Complexes were removed by matching PDB IDs against the CASF-2013, CASF-2016, and CASF-2019 core lists prior to dataset splitting to ensure absolutely no data leakage. We then exclude samples that cannot be parsed by RDKit and randomly partition the remaining samples into training (N = 11
IC50 (with larger values indicating stronger binding). Complexes were removed by matching PDB IDs against the CASF-2013, CASF-2016, and CASF-2019 core lists prior to dataset splitting to ensure absolutely no data leakage. We then exclude samples that cannot be parsed by RDKit and randomly partition the remaining samples into training (N = 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 904) and validation (N = 1000) sets. To test MSIGN's generalization capability, we leverage three independent external test sets: the CASF-2013 benchmark, the CASF-2016 benchmark, and the 2019 core set. It should be noted that there is no overlap among training, validation, and testing sets.
904) and validation (N = 1000) sets. To test MSIGN's generalization capability, we leverage three independent external test sets: the CASF-2013 benchmark, the CASF-2016 benchmark, and the 2019 core set. It should be noted that there is no overlap among training, validation, and testing sets.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 ratio.
2 ratio.
2.2 Feature representation
This study employs three distinct types of graph structures: the complex atomic interaction graph, the SC functional group graph, and the CB1R ligand amino acid residue graph. These graphs are designed to explore the feature space of SC-CB1R binding interactions from three perspectives: 3D structural information, physical principles, and chemical properties; all feature dimensions are shown in Tables S1–S3. The following sections provide detailed descriptions of these graph structures.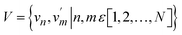 and E = {ei, ej|i, j ϵ [1, 2, …,M]}. Here, V represents the set of atoms, including SC atoms vn and CB1R atoms
and E = {ei, ej|i, j ϵ [1, 2, …,M]}. Here, V represents the set of atoms, including SC atoms vn and CB1R atoms 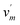 , with N being the total number of atoms in the complex. EE denotes the set of edges, comprising two types of interactions: covalent bonds ei and non-covalent bonds ej.
, with N being the total number of atoms in the complex. EE denotes the set of edges, comprising two types of interactions: covalent bonds ei and non-covalent bonds ej.
2.3 Model architecture
(1) Atomic-Level 3D interaction features: MSIGN constructs an atomic-level graph representation of the 3D complex formed by the ligand and the target protein's binding pocket. Through a message-passing network, it iteratively updates edge features by aggregating node embeddings from neighboring atoms. This process ultimately quantifies both covalent and non-covalent interactions between atoms.
(2) Functional group and amino acid-level 2D property features: in contrast to the atomic-level graph structure, proteins and ligands are modeled as 2D functional group graphs and amino acid residue graphs, respectively. By incorporating these multi-scale features, MSIGN captures both fine-grained atomic interactions and higher-level chemical characteristics, enhancing its ability to predict binding affinities accurately.
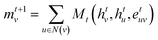 | (1) |
| ht+1v = Ut(htv, mt+1v) | (2) |
The readout phase aims to map the entire graph's features into a global feature vector that captures the graph's holistic properties. This is achieved using a readout function:
| ŷ = R({hTv|v ϵ G}) | (3) |
Specifically, to calculate the strength of covalent interactions among CB1R binding pocket atoms, we take atom vi as an example: first, we query the adjacency matrix of the graph to identify all neighboring atoms vj of vi. The feature matrices of these neighboring nodes and the edge features are processed through a message function to generate a message mvi, which is then propagated to node vj. An aggregation function combines the previous layer's hvi with the message mvi to obtain the feature H1. The mathematical representation is as follows:
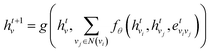 | (4) |
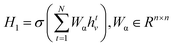 | (5) |
The covalent interactions among SC atoms are computed similarly, with the atom node set replaced by SC atoms. The equations are as follows:
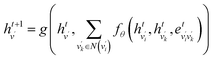 | (6) |
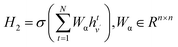 | (7) |
Due to the significantly larger number of non-covalent interactions compared to covalent interactions, we employ mean aggregation for non-covalent interaction calculations. The equations are as follows:
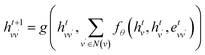 | (8) |
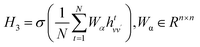 | (9) |
Similarly, to comprehensively analyze the chemical properties of the CB1R protein, we construct and incorporate an amino acid residue graph. Amino acid residues in proteins exhibit diverse chemical properties, such as polarity, hydrophobicity, and charge, which determine protein–ligand interaction patterns.
For the constructed CB1R amino acid residue graph and ligand functional group graph, we perform one iteration of message passing to aggregate features from neighboring nodes and generate messages, ultimately obtaining tensor representations of the entire protein and molecule. These features are then fed into a cross-attention layer to learn potential global correspondences. The cross-attention computation is as follows:
| Q = MfWQ, K = MaWK, V = MaWV | (10) |
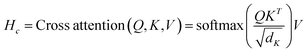 | (11) |
| ŷpred = MLP(concat(Hp, Hc)) | (12) |
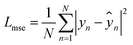 | (13) |
2.4 Implementation details
The MSIGN framework was implemented using PyTorch and PyTorch Geometric. All experiments were performed on a workstation equipped with a single NVIDIA GeForce RTX 4090 GPU. The model utilizes an Adam optimizer with a learning rate of 1 × 10−4 and a weight decay of 1 × 10−6. The batch size was set to 128, and the training was conducted for a maximum of 300 epochs with an early stopping strategy (patience set to 30 epochs) to prevent overfitting. The model architecture consists of 3 message-passing layers with a hidden feature size of 256. The pre-training process is computationally efficient, requiring approximately 5 GB of GPU memory and completing in about 3.5 hours.3. Results and discussion
3.1 Comparison of general MSIGN with baseline models
The MSIGN model architecture (as shown in Fig. 1) has demonstrated excellent performance in both the general model trained on the PDBbind dataset and the transfer learning model trained on the SC dataset. To rigorously evaluate the superiority of the general MSIGN model, we selected representative baseline models for comparison, including DeepDTA,6 MGraphDTA,41 Pafnucy,17 GIGN,3 SS-GNN,42 and EHIGN.22 Additionally, to provide a broader context, we implemented classical machine learning models, Random Forest (RF) and XGBoost, as baselines.
Table 1 and Table S4 below summarizes the performance comparison between MSIGN and other baseline models on the CASF-2013,43 CASF-2016,44 and 2019 core set benchmark test sets. The evaluation metrics used are Root Mean Square Error (RMSE) and Pearson correlation coefficient (Rp). To better quantify the model advancement and assess statistical significance, we calculated 95% confidence intervals (CI) for MSIGN and the machine learning baselines. These intervals were derived using a non-parametric bootstrap resampling procedure, as recommended for rigorous model validation.45 Specifically, prediction–target pairs were resampled with replacement 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 times, and the 2.5th and 97.5th percentiles of the resulting metric distributions were taken as the confidence bounds. The results indicate that MSIGN achieved the best performance across all metrics on all test sets. This superior performance underscores the effectiveness of MSIGN in capturing both atomic-level interactions and higher-level chemical features, demonstrating its robustness and generalizability in binding affinity prediction tasks.
000 times, and the 2.5th and 97.5th percentiles of the resulting metric distributions were taken as the confidence bounds. The results indicate that MSIGN achieved the best performance across all metrics on all test sets. This superior performance underscores the effectiveness of MSIGN in capturing both atomic-level interactions and higher-level chemical features, demonstrating its robustness and generalizability in binding affinity prediction tasks.
| Model | CASF-2013 | CASF-2016 | 2019 core set | |||
|---|---|---|---|---|---|---|
| RMSE | R p | RMSE | R p | RMSE | R p | |
a Results for DeepDTA, MGraphDTA, Pafnucy, GIGN, SS-GNN, and EHIGN were retrieved directly from the literature [ref], where confidence intervals were not reported.
b For Random Forest, XGBoost, and MSIGN, values represent the observed metric followed by the [95% Confidence Interval] in brackets. The intervals were calculated using bootstrap resampling with 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 iterations, based on the 2.5th and 97.5th percentiles of the distribution. 000 iterations, based on the 2.5th and 97.5th percentiles of the distribution.
|
||||||
| DeepDTAa | 1.651 | 0.713 | 1.363 | 0.781 | 1.503 | 0.589 |
| MGraphDTAa | 1.737 | 0.682 | 1.454 | 0.742 | 1.533 | 0.529 |
| Pafnucya | 1.523 | 0.779 | 1.438 | 0.771 | 1.441 | 0.617 |
| GIGNa | 1.384 | 0.819 | 1.188 | 0.843 | 1.399 | 0.637 |
| SS-GNNa | 1.327 | 0.826 | 1.172 | 0.848 | 1.452 | 0.629 |
| EHIGNa | 1.308 | 0.841 | 1.156 | 0.853 | 1.363 | 0.664 |
| Random forestb | 1.775 [1.538, 2.009] | 0.661 [0.533, 0.766] | 1.622 [1.478, 1.768] | 0.651 [0.571, 0.722] | 1.499 [1.467, 1.531] | 0.544 [0.521, 0.566] |
| XGBoostb | 1.588 [1.382, 1.786] | 0.749 [0.648, 0.828] | 1.448 [1.318, 1.575] | 0.742 [0.678, 0.796] | 1.466 [1.434, 1.498] | 0.580 [0.559, 0.601] |
| MSIGNb | 1.256 [1.114, 1.419] | 0.848 [0.808, 0.899] | 1.130 [1.042, 1.214] | 0.857 [0.834, 0.891] | 1.327 [1.301, 1.344] | 0.673 [0.647, 0.684] |
3.2 Ablation study and permutation analysis
To systematically evaluate the contributions of each feature component and verify whether MSIGN truly learns physical interactions rather than exploiting dataset biases, we conducted both feature ablation studies and permutation-based control tests.Feature ablation analysis: we designed four MSIGN variants by selectively removing the Atomic Interaction Graph (IG), Ligand Functional Group Graph (FG), or Amino Acid Residue Graph (AARG). As summarized in Table 2, the full MSIGN model consistently outperforms all variants. The model relying solely on IG (MSIGN_IG) shows suboptimal performance, while removing IG entirely (MSIGN_FG + AARG) leads to the worst results, confirming that atomic interactions form the physical foundation of binding. The integration of chemical priors (FG and AARG) significantly boosts performance, with the full model achieving the lowest validation RMSE and highest correlation. These results demonstrate the synergistic effect of fusing multi-scale physical and chemical features.
| Model variant/setting | IG | FG | AARG | RMSE | R p |
|---|---|---|---|---|---|
| MSIGN_IG | ✓ | ✗ | ✗ | 1.353 | 0.701 |
| MSIGN_FG + AARG | ✗ | ✓ | ✓ | 1.401 | 0.654 |
| MSIGN_IG + FG | ✓ | ✓ | ✗ | 1.263 | 0.719 |
| MSIGN_IG + AARG | ✓ | ✗ | ✓ | 1.268 | 0.723 |
| MSIGN | ✓ | ✓ | ✓ | 1.219 | 0.747 |
Permutation-based control tests: to verify that MSIGN captures genuine 3D interactions rather than dataset biases,46,47 we conducted permutation tests (Table 3). Shuffling ligand labels (ligand permutation) caused a complete performance collapse, confirming the model's sensitivity to ligand identity. In the protein permutation setting, although the model retained moderate correlation due to intrinsic ligand potency, its performance degraded significantly, dropping even below that of the ligand-only ML baselines. This observation is critical: it rules out the possibility that MSIGN acts merely as a ligand-based predictor, which would otherwise maintain baseline-level accuracy despite protein shuffling. Instead, MSIGN achieves state-of-the-art performance only when genuine protein–ligand pairs are preserved. This substantial performance gap between the permuted and original settings validates the atomic interaction graph module, confirming that the model's predictive power stems from learning specific 3D spatial complementarity rather than exploiting ligand or protein priors.
| Experimental setting | CASF-2013 | CASF-2016 | 2019 core set | |||
|---|---|---|---|---|---|---|
| RMSE | R p | RMSE | R p | RMSE | R p | |
| Ligand permutation | 3.097 | 0.0478 | 2.717 | 0.177 | 2.640 | 0.096 |
| Protein permutation | 2.126 | 0.599 | 2.040 | 0.542 | 2.278 | 0.388 |
| Joint permutation | 3.508 | −0.168 | 2.941 | −0.025 | 2.834 | 0.019 |
| Original (baseline) | 1.256 | 0.848 | 1.130 | 0.857 | 1.327 | 0.673 |
3.3 Performance of general MSIGN models on GPCR families
We first compiled 1417 GPCR entries with PDB identifiers by combining datasets from InterPro48 and GPCRdb.49 Subsequent intersection with PDBBind entries yielded 48 GPCR protein–ligand complexes, which were isolated as an external test set. The remaining data (excluding these 48 GPCR entries) was partitioned into training and validation sets at an 8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 ratio. Using identical hyperparameters to previous configurations, we trained a new MSIGN model by minimizing the RMSE on the validation set and then evaluated its performance on the GPCR test set. The first 15 results (sorted in ascending order according to affinity label size) are shown in Table 4, with the bold part indicating that the absolute value of the difference is greater than 0.5.
2 ratio. Using identical hyperparameters to previous configurations, we trained a new MSIGN model by minimizing the RMSE on the validation set and then evaluated its performance on the GPCR test set. The first 15 results (sorted in ascending order according to affinity label size) are shown in Table 4, with the bold part indicating that the absolute value of the difference is greater than 0.5.
| PDB ID | Label | Predicted value | Difference | |
|---|---|---|---|---|
| 1 | 5k5s | 2.69 | 4.94 | 2.25 |
| 2 | 3c14 | 3.52 | 4.65 | 1.13 |
| 3 | 5vex | 4.63 | 6.59 | 1.96 |
| 4 | 5vew | 5.15 | 6.98 | 1.83 |
| 5 | 3eht | 5.17 | 5.64 | 0.47 |
| 6 | 4mk0 | 5.20 | 7.75 | 2.55 |
| 7 | 1y3a | 5.41 | 7.04 | 1.63 |
| 8 | 6d1u | 5.44 | 6.44 | 1.00 |
| 9 | 1cs4 | 5.57 | 4.53 | −1.04 |
| 10 | 2g83 | 5.92 | 7.37 | 1.45 |
| 11 | 5ukl | 6.20 | 7.64 | 1.44 |
| 12 | 5ukk | 6.21 | 7.93 | 1.72 |
| 13 | 3rbq | 6.27 | 6.36 | 0.09 |
| 14 | 4wnk | 6.42 | 8.85 | 2.43 |
| 15 | 3 d7m | 6.44 | 6.32 | −0.12 |
Experimental results revealed an RMSE of 1.5626 and a correlation coefficient of 0.2705 between predicted and actual values. This demonstrates that limited training data severely impede the model's ability to capture interaction patterns between ligands and mechanistically complex protein families like GPCRs, with most predictions deviating substantially from experimental observations. To address this challenge for accurate prediction of CB1R-SC interactions, we propose implementing a pre-training and transfer learning framework for deeper investigation.
3.4 Transfer learning with MSIGN
The scarcity and specialization of synthetic cannabinoid (SC) data present a significant challenge for model generalization. To overcome this, we implemented a transfer learning paradigm (Fig. 2A and B) to bridge the gap between general physicochemical laws and domain-specific binding modes. This strategy addresses two critical bottlenecks: (i) the chemical space bias inherent in generalized models when applied directly to SCs–CB1R interactions and (ii) the high risk of overfitting when training on a limited dataset of only a few thousand entries. By first capturing fundamental interactions (e.g., hydrogen bonding and hydrophobic complementarity) from the PDBbind dataset, the model subsequently learned to adapt to unique SC-CB1R features, such as the preference of indolyl ring orientation and residue polarity distribution.Comparative analysis of different fine-tuning strategies (Fig. 2C and D and S2) highlights the necessity of this approach. Training MSIGN directly on the SCsDB without pre-training resulted in the longest training duration, significant RMSE fluctuations, and poor stability, indicating a failure to converge effectively. In contrast, while the “frozen” strategy reduced training time, the full fine-tuning strategy demonstrated superior performance. It achieved the lowest validation RMSE and reached convergence in only 124 epochs (Fig. S3 and S4). These results underscore that the “pre-training + full fine-tuning” framework not only optimizes computational efficiency but also significantly enhances the model's ability to generalize across the novel chemical spaces of synthetic cannabinoids.
3.5 Experimental verification of the reliability of model predictions
In affinity prediction, mixing Ki, Kd, and IC50 labels (converted to pK) is a common but debated practice due to potential systematic errors arising from different experimental conditions. To evaluate this effect, we compared our hybrid model (MSIGN_Mix) against sub-models trained on the Ki subset (N = 2916) and the IC50 subset (N = 1215) (Fig. S5; Kd was excluded due to N = 35). As shown in Table S5, MSIGN_Mix demonstrated the best performance, achieving the lowest RMSE and the highest correlation Rp (0.790).This suggests that label hybridization enhances performance by expanding the training data and neutralizing metric-specific noise. Interestingly, MSIGN_IC50 outperformed MSIGN_Ki in Rp, likely because IC50 experiments often use more standardized conditions (e.g., fixed enzyme/substrate concentrations), whereas Ki values may exhibit higher heterogeneity due to variable competitive inhibition assay setups. Despite these intrinsic differences, the superior accuracy of MSIGN_Mix indicates that the model effectively captures universal protein–ligand binding mechanisms that transcend specific experimental metrics.
3.6 Experimental verification of model predictions
To assess the practical reliability of the fine-tuned MSIGN model, we performed functional validation on three novel synthetic cannabinoids (SCs) through fluorescence-based sensing and SPR assays.As shown in Fig. S6 and S7, the dose–response curves obtained from these fluorescence assays serve as critical physiological “ground truth.” These results, along with the Kd values measured by SPR, were used to externally validate the predictive accuracy of our MSIGN model. The high consistency between the AI-predicted affinities and these biological responses (discussed in Section 3.7) demonstrates that MSIGN successfully captured the structural nuances required for CB1R–ligand interaction.
3.7 Full finetuning of the MSIGN model achieves optimal predictions
To evaluate the practical predictive power of MSIGN, we conducted a validation using three novel synthetic cannabinoids (SCs) confirmed by wet-lab experiments. We first compared four internal training paradigms (Fig. S8 and Tables S6 and S7). The General model showed a pronounced systematic overestimation, failing to capture the specific CB1R pocket interaction due to “domain shift.” Conversely, the no-pretraining model exhibited an underestimation trend, suggesting that small-domain datasets alone are insufficient to learn fundamental physical laws, leading to underfitting. While frozen fine-tuning partially reduced these errors, it remained constrained by parameter fixation. Ultimately, the full fine-tuning strategy achieved the highest consistency with experimental measurements, demonstrating that full-parameter updates are essential for capturing target-specific 3D spatial complementarity.We further benchmarked MSIGN against traditional ML (RF and XGBoost), state-of-the-art DL (EHIGN and SS-GNN), and physics-based docking (AutoDock Vina) (Fig. 3E and Table S8). The results reveal a substantial accuracy gap. AutoDock Vina yielded the largest deviation (highest MAE), likely due to the challenges of standard scoring functions in modeling complex GPCR thermodynamics. General structure-based models like EHIGN and SS-GNN also exhibited large overestimation errors, confirming that models lacking domain-specific fine-tuning generalize poorly to novel SCs. While ligand-based ML models performed slightly better than general DL baselines, they still failed to achieve chemical accuracy.
(A) As illustrated by the MAE comparison (Fig. 3E, left), MSIGN stands out as the only methodology providing reliable, unbiased predictions. The sharp alignment between its predictions and experimental results (Fig. 3E, right) validates our “pre-training + full fine-tuning” paradigm. These findings underscore that the synergistic integration of multi-scale interaction modeling and domain-specific knowledge transfer is the key to bridging the gap between computational screening and experimental results in data-scarce scenarios.
3.8 Limitations of the study
Despite the robust performance of MSIGN, several limitations should be acknowledged. First, while our wet-lab validation on three novel synthetic cannabinoids yielded high consistency, the sample size remains limited for a comprehensive assessment of the model's performance in broader chemical spaces. Second, experimental binding assays (SPR and fluorescence) are subject to inherent variability and environmental factors, which may introduce minor measurement uncertainties. Third, although we benchmarked against AutoDock Vina, it represents a classical scoring function; more advanced but computationally expensive methods like FEP might provide different physical insights. Finally, this study focused specifically on the CB1 receptor. Whether the multi-scale priors and fine-tuning strategy generalize as effectively to the CB2 receptor or other GPCR families remains to be explored in future work.4. Conclusion
In this study, we propose a binding affinity prediction model based on a multi-scale interaction graph neural network. Compared to other deep learning models, MSIGN achieves state-of-the-art performance across multiple evaluation metrics. This superior performance can be attributed to the feature extraction process of MSIGN, which not only incorporates physically meaningful covalent and non-covalent interactions but also integrates critical chemical prior knowledge, which is essential for accurate binding affinity prediction.Furthermore, we demonstrate the effectiveness of the pre-training and fine-tuning paradigm in the context of binding affinity prediction using a SC dataset. Moreover, the choice of fine-tuning strategy—whether full fine-tuning or frozen fine-tuning—has a substantial impact on model performance, depending on the characteristics of the downstream dataset. Specifically, on our SC dataset, full fine-tuning outperformed frozen fine-tuning, yielding more accurate predictions.
We believe that our MSIGN model provides a novel framework for researchers, offering a robust solution for achieving reliable binding affinity predictions even in scenarios where downstream task data are limited. This approach opens new avenues for advancing computational drug discovery and molecular design.
Author contributions
Jun Liao: conceptualization. Zhenyong Cheng: methodology. Dinghao Liu: software and writing – review & editing. Yuanpeng Fu: writing – original draft. Kewei Sheng: investigation. Yan Xing: formal analysis and data curation. Yanling Qiao: validation. Shangxuan Cai: validation. Jubo Wang: validation. Peng Xu: supervision and project administration.Conflicts of interest
The authors declare no competing financial interest.Data availability
The source code, trained models, and scripts used in this study are openly available on GitHub at https://github.com/Dr-F-Arthur/MSIGN/tree/master. An archived version of the repository is available on Zenodo under the identifier DOI: https://zenodo.org/records/16018515. The public datasets used for training and evaluation in this study are available from the following resources. PDBbind-v2020: https://www.pdbbind-plus.org.cn/download. CASF Benchmarks (2013/2016): https://www.pdbbind-plus.org.cn/casf. BindingDB: https://www.bindingdb.org/rwd/bind/chemsearch/marvin/Download.jsp.Supplementary information is available (SI). See DOI: https://doi.org/10.1039/d5dd00317b.
Acknowledgements
This work was supported by Zhejiang Lab (Program: Ab Initio Design and Generation of AI Models for Small Molecule Ligands Based on Target Structures, Grant No. 2022PE0AC03), the National Key R&D Program of China (Program: A Study on the Diagnosis of Addiction to Synthetic Cannabinoids and Methods of Assessing the Risk of Abuse, Grant No. 2022YFC3300905), and the National Key R&D Program of China (Program: Research on Key Technologies for Monitoring and Identifying Drug Abuse of Anesthetic Drugs and Psychotropic Substances, and Intervention for Addiction, Grant No. 2023YFC3304200).References
- M. Ragoza, J. Hochuli, E. Idrobo, J. Sunseri and D. R. Koes, Protein–Ligand Scoring with Convolutional Neural Networks, J. Chem. Inf. Model., 2017, 57(4), 942–957, DOI:10.1021/acs.jcim.6b00740.
- Z. Jin, T. Wu, T. Chen, D. Pan, X. Wang, J. Xie, L. Quan and Q. Lyu, CAPLA: Improved Prediction of Protein-Ligand Binding Affinity by a Deep Learning Approach Based on a Cross-Attention Mechanism, Bioinformatics, 2023, 39(2), btad049, DOI:10.1093/bioinformatics/btad049.
- Z. Yang, W. Zhong, Q. Lv, T. Dong and C. Yu-Chian Chen, Geometric Interaction Graph Neural Network for Predicting Protein-Ligand Binding Affinities from 3D Structures (GIGN), J. Phys. Chem. Lett., 2023, 14(8), 2020–2033, DOI:10.1021/acs.jpclett.2c03906.
- X. Zhang, H. Gao, H. Wang, Z. Chen, Z. Zhang, X. Chen, Y. Li, Y. Qi and R. Wang, PLANET: A Multi-Objective Graph Neural Network Model for Protein-Ligand Binding Affinity Prediction, J. Chem. Inf. Model., 2024, 64(7), 2205–2220, DOI:10.1021/acs.jcim.3c00253.
- Y. Zhang, S. Li, K. Meng and S. Sun, Machine Learning for Sequence and Structure-Based Protein–Ligand Interaction Prediction, J. Chem. Inf. Model., 2024, 64(5), 1456–1472, DOI:10.1021/acs.jcim.3c01841.
- H. Öztürk, A. Özgür and E. OzKirimli, DeepDTA: Deep Drug-Target Binding Affinity Prediction, Bioinformatics, 2018, 34(17), i821–i829, DOI:10.1093/bioinformatics/bty593.
- T. Nguyen; H. Le; T. P. Quinn; T. Nguyen; T. D. Le; S. VenkateshGraphDTA: Predicting Drug–Target Binding Affinity with Graph Neural Networks.
- M. Jiang, Z. Li, S. Zhang, S. Wang, X. Wang, Q. Yuan and Z. Wei, Drug–Target Affinity Prediction Using Graph Neural Network and Contact Maps, RSC Adv., 2020, 10(35), 20701–20712, 10.1039/D0RA02297G.
- T. M. Nguyen, T. Nguyen, T. M. Le and T. Tran, GEFA:Early Fusion Approach in Drug-Target Affinity Prediction, IEEE Trans. Comput. Biol. Bioinform., 2022, 19(2), 718–728, DOI:10.1109/TCBB.2021.3094217.
- M. Jiang, S. Wang, S. Zhang, W. Zhou, Y. Zhang and Z. Li, Sequence-Based Drug-Target Affinity Prediction Using Weighted Graph Neural Networks, BMC Genom., 2022, 23(1), 449, DOI:10.1186/s12864-022-08648-9.
- H. Qi, T. Yu, W. Yu and C. Liu, Drug–Target Affinity Prediction with Extended Graph Learning-Convolutional Networks, BMC Bioinf., 2024, 25(1), 75, DOI:10.1186/s12859-024-05698-6.
- Y.-B. Wang, Z.-H. You, S. Yang, H.-C. Yi, Z.-H. Chen and K. Zheng, A Deep Learning-Based Method for Drug-Target Interaction Prediction Based on Long Short-Term Memory Neural Network, BMC Med. Inform. Decis. Mak., 2020, 20(S2), 49, DOI:10.1186/s12911-020-1052-0.
- S. Li, J. Zhou, T. Xu, L. Huang, F. Wang, H. Xiong, W. Huang, D. Dou and H. XiongStructure-Aware Interactive Graph Neural Networks for the Prediction of Protein-Ligand Binding Affinity. arXiv 2021. preprint arXiv:10.48550/arXiv.2107.10670, DOI:10.48550/arXiv.2107.10670.
- C. Xia, S.-H. Feng, Y. Xia, X. Pan and H.-B. Shen, Leveraging Scaffold Information to Predict Protein–Ligand Binding Affinity with an Empirical Graph Neural Network, Briefings Bioinf., 2023, 24(1), bbac603, DOI:10.1093/bib/bbac603.
- R. Gorantla, A. Kubincová, A. Y. Weiße and A. S. J. S. Mey, From Proteins to Ligands: Decoding Deep Learning Methods for Binding Affinity Prediction, J. Chem. Inf. Model., 2024, 64(7), 2496–2507, DOI:10.1021/acs.jcim.3c01208.
- J. Jiménez, M. Škalič, G. Martínez-Rosell and G. De Fabritiis, K DEEP : Protein–Ligand Absolute Binding Affinity Prediction via 3D-Convolutional Neural Networks, J. Chem. Inf. Model., 2018, 58(2), 287–296, DOI:10.1021/acs.jcim.7b00650.
- M. M. Stepniewska-Dziubinska, P. ZielenKiewicz and P. SiedlecKi, Development and Evaluation of a Deep Learning Model for Protein-Ligand Binding Affinity Prediction, Bioinformatics, 2018, 34(21), 3666–3674, DOI:10.1093/bioinformatics/bty374.
- F. Imrie, A. R. Bradley, M. Van Der Schaar and C. M. Deane, Protein Family-Specific Models Using Deep Neural Networks and Transfer Learning Improve Virtual Screening and Highlight the Need for More Data, J. Chem. Inf. Model., 2018, 58(11), 2319–2330, DOI:10.1021/acs.jcim.8b00350.
- J. A. Morrone, J. K. Weber, T. Huynh, H. Luo and W. D. Cornell, Combining DocKing Pose Rank and Structure with Deep Learning Improves Protein–Ligand Binding Mode Prediction over a Baseline DocKing Approach, J. Chem. Inf. Model., 2020, 60(9), 4170–4179, DOI:10.1021/acs.jcim.9b00927.
- H. Hassan-Harrirou, C. Zhang and T. Lemmin, RosENet: Improving Binding Affinity Prediction by Leveraging Molecular Mechanics Energies with an Ensemble of 3D Convolutional Neural Networks, J. Chem. Inf. Model., 2020, 60(6), 2791–2802, DOI:10.1021/acs.jcim.0c00075.
- S. Moon, W. Zhung, S. Yang, J. Lim and W. Y. Kim, PIGNet: A Physics-Informed Deep Learning Model toward Generalized Drug-Target Interaction Predictions, Chem. Sci., 2022, 13(13), 3661–3673, 10.1039/d1sc06946b.
- Z. Yang, W. Zhong, Q. Lv, T. Dong, G. Chen and C. Y.-C. Chen, Interaction-Based Inductive Bias in Graph Neural Networks: Enhancing Protein-Ligand Binding Affinity Predictions From 3D Structures, IEEE Trans. Pattern Anal. Mach. Intell., 2024, 46(12), 8191–8208, DOI:10.1109/TPAMI.2024.3400515.
- W. Torng and R. B. Altman, Graph Convolutional Neural Networks for Predicting Drug-Target Interactions, J. Chem. Inf. Model., 2019, 59(10), 4131–4149, DOI:10.1021/acs.jcim.9b00628.
- R. Milo; S. Shen-Orr; S. Itzkovitz; N. Kashtan; D. ChklovsKii; U. AlonNetwork Motifs: Simple Building Blocks of Complex Networks.
- S. Han, H. Fu, Y. Wu, G. Zhao, Z. Song, F. Huang, Z. Zhang, S. Liu and W. Zhang, HimGNN: A Novel Hierarchical Molecular Graph Representation Learning Framework for Property Prediction, Briefings Bioinf., 2023, 24(5), bbad305, DOI:10.1093/bib/bbad305.
- H. Öztürk, E. OzKirimli and A. Özgür, WideDTA: Prediction of Drug-Target Binding Affinity, arXiv, 2019, preprint, DOI:10.48550/arXiv.1902.04166.
- H. T. Rube, C. Rastogi, S. Feng, J. F. Kribelbauer, A. Li, B. Becerra, L. A. N. Melo, B. V. Do, X. Li, H. H. Adam, N. H. Shah, R. S. Mann and H. J. Bussemaker, Prediction of Protein–Ligand Binding Affinity from Sequencing Data with Interpretable Machine Learning, Nat. Biotechnol., 2022, 40(10), 1520–1527, DOI:10.1038/s41587-022-01307-0.
- C. Cai, S. Wang, Y. Xu, W. Zhang, K. Tang, Q. Ouyang, L. Lai and J. Pei, Transfer Learning for Drug Discovery, J. Med. Chem., 2020, 63(16), 8683–8694, DOI:10.1021/acs.jmedchem.9b02147.
- R. Wang, X. Fang, Y. Lu, C.-Y. Yang and S. Wang, The PDBbind Database: Methodologies and Updates, J. Med. Chem., 2005, 48(12), 4111–4119, DOI:10.1021/jm048957q.
- T. Liu, L. Hwang, S. K. Burley, C. I. Nitsche, C. Southan, W. P. Walters and M. K. Gilson, BindingDB in 2024: A FAIR Knowledgebase of Protein-Small Molecule Binding Data, Nucleic Acids Res., 2025, 53(D1), D1633–D1644, DOI:10.1093/nar/gkae1075.
- T. Hua, K. Vemuri, M. Pu, L. Qu, G. W. Han, Y. Wu, S. Zhao, W. Shui, S. Li, A. Korde, R. B. Laprairie, E. L. Stahl, J.-H. Ho, N. Zvonok, H. Zhou, I. Kufareva, B. Wu, Q. Zhao, M. A. Hanson, L. M. Bohn, A. Makriyannis, R. C. Stevens and Z.-J. Liu, Crystal Structure of the Human Cannabinoid Receptor CB1, Cell, 2016, 167(3), 750–762.e14, DOI:10.1016/j.cell.2016.10.004.
- T. Hua, K. Vemuri, S. P. Nikas, Y. Wu, L. Qu, M. Pu, A. Korde, S. Jiang, J.-H. Ho, G. W. Han, K. Ding, X. Li, H. Liu, M. A. Hanson, S. Zhao, L. M. Bohn, A. Makriyannis, R. C. Stevens and Z.-J. Liu, Crystal Structures of Agonist-Bound Human Cannabinoid Receptor CB1, Nature, 2025, 646(8085), 754–758, DOI:10.1038/s41586-025-09454-5.
- X. Yang, X. Wang, Z. Xu, C. Wu, Y. Zhou, Y. Wang, G. Lin, K. Li, M. Wu, A. Xia, J. Liu, L. Cheng, J. Zou, W. Yan, Z. Shao and S. Yang, Molecular Mechanism of Allosteric Modulation for the Cannabinoid Receptor CB1, Nat. Chem. Biol., 2022, 18(8), 831–840, DOI:10.1038/s41589-022-01038-y.
- J. Lim, S. Ryu, K. Park, Y. J. Choe, J. Ham and W. Y. Kim, Predicting Drug–Target Interaction Using a Novel Graph Neural Network with 3D Structure-Embedded Graph Representation, J. Chem. Inf. Model., 2019, 59(9), 3981–3988, DOI:10.1021/acs.jcim.9b00387.
- D. Jones, H. Kim, X. Zhang, A. Zemla, G. Stevenson, W. F. D. Bennett, D. Kirshner, S. E. Wong, F. C. Lightstone and J. E. Allen, Improved Protein–Ligand Binding Affinity Prediction with Structure-Based Deep Fusion Inference, J. Chem. Inf. Model., 2021, 61(4), 1583–1592, DOI:10.1021/acs.jcim.0c01306.
- J. Gilmer; S. S. Schoenholz; P. F. Riley; O. Vinyals; G. E. DahlNeural Message Passing for Quantum Chemistry. arXiv, 2017, preprint arXiv:10.48550/arXiv.1704.01212 DOI:10.48550/arXiv.1704.01212.
- H. Cai, H. Zhang, D. Zhao, J. Wu and L. Wang, FP-GNN: A Versatile Deep Learning Architecture for Enhanced Molecular Property Prediction, Briefings Bioinf., 2022, 23(6), bbac408, DOI:10.1093/bib/bbac408.
- Z. Zheng, Y. Tan, H. Wang, S. Yu, T. Liu and C. Liang, CasANGCL: Pre-Training and Fine-Tuning Model Based on Cascaded Attention Network and Graph Contrastive Learning for Molecular Property Prediction, Briefings Bioinf., 2023, 24(1), bbac566, DOI:10.1093/bib/bbac566.
- Z. Wang, M. Liu, Y. Luo, Z. Xu, Y. Xie, L. Wang, L. Cai, Q. Qi, Z. Yuan, T. Yang and S. Ji, Advanced Graph and Sequence Neural Networks for Molecular Property Prediction and Drug Discovery, Bioinformatics, 2022, 38(9), 2579–2586, DOI:10.1093/bioinformatics/btac112.
- L. Hirschfeld, K. Swanson, K. Yang, R. Barzilay and C. W. Coley, Uncertainty Quantification Using Neural Networks for Molecular Property Prediction, J. Chem. Inf. Model., 2020, 60(8), 3770–3780, DOI:10.1021/acs.jcim.0c00502.
- Z. Yang, W. Zhong, L. Zhao, Yu-C. Chen and C. MGraphDTA, Deep Multiscale Graph Neural Network for Explainable Drug–Target Binding Affinity Prediction, Chem. Sci., 2022, 13(3), 816–833, 10.1039/D1SC05180F.
- S. Zhang, Y. Jin, T. Liu, Q. Wang, Z. Zhang, S. Zhao and B. Shan, SS-GNN: A Simple-Structured Graph Neural Network for Affinity Prediction, ACS Omega, 2023, 8(25), 22496–22507, DOI:10.1021/acsomega.3c00085.
- Y. Li, Z. Liu, J. Li, L. Han, J. Liu, Z. Zhao and R. Wang, Comparative Assessment of Scoring Functions on an Updated Benchmark: 1. Compilation of the Test Set, J. Chem. Inf. Model., 2014, 54(6), 1700–1716, DOI:10.1021/ci500080q.
- M. Su, Q. Yang, Y. Du, G. Feng, Z. Liu, Y. Li and R. Wang, Comparative Assessment of Scoring Functions: The CASF-2016 Update, J. Chem. Inf. Model., 2019, 59(2), 895–913, DOI:10.1021/acs.jcim.8b00545.
- J. R. Ash, C. Wognum, R. Rodríguez-Pérez, M. Aldeghi, A. C. Cheng, D.-A. Clevert, O. Engkvist, C. Fang, D. J. Price, J. M. Hughes-Oliver and W. P. Walters, Practically Significant Method Comparison Protocols for Machine Learning in Small Molecule Drug Discovery, J. Chem. Inf. Model., 2025, 65(18), 9398–9411, DOI:10.1021/acs.jcim.5c01609.
- G. Durant, F. Boyles, K. Birchall, B. Marsden and C. M. Deane, Robustly Interrogating Machine Learning-Based Scoring Functions: What Are They Learning?, Bioinformatics, 2025, 41(2), btaf040, DOI:10.1093/bioinformatics/btaf040.
- P. Avdiunina; S. Jamal; F. Gusev; O. IsayevAll That Glitters Is Not Gold: Importance of Rigorous Evaluation of Proteochemometric Models. January 22, 2025. doi: DOI:10.26434/chemrxiv-2025-vbmgc.
- M. Blum, A. Andreeva, L. C. Florentino, S. R. Chuguransky, T. Grego, E. Hobbs, B. L. Pinto, A. Orr, T. Paysan-Lafosse, I. Ponamareva, G. A. Salazar, N. Bordin, P. Bork, A. Bridge, L. Colwell, J. Gough, D. H. Haft, I. Letunic, F. Llinares-López, A. Marchler-Bauer, L. Meng-Papaxanthos, H. Mi, D. A. Natale, C. A. Orengo, A. P. Pandurangan, D. Piovesan, C. Rivoire, C. J. A. Sigrist, N. ThanKi, F. Thibaud-Nissen, P. D. Thomas, S. C. E. Tosatto, C. H. Wu and A. Bateman, InterPro: The Protein Sequence Classification Resource in 2025, Nucleic Acids Res., 2025, 53(D1), D444–D456, DOI:10.1093/nar/gkae1082.
- V. Isberg, S. MordalsKi, C. Munk, K. Rataj, K. Harpsøe, A. S. Hauser, B. Vroling, A. J. BojarsKi, G. Vriend and D. E. Gloriam, GPCRdb: An Information System for G Protein-Coupled Receptors, Nucleic Acids Res., 2016, 44(D1), D356–D364, DOI:10.1093/nar/gkv1178.
- D. G. Gibson, L. Young, R.-Y. Chuang, J. C. Venter, C. A. Hutchison and H. O. Smith, Enzymatic Assembly of DNA Molecules up to Several Hundred Kilobases, Nat. Methods, 2009, 6(5), 343–345, DOI:10.1038/nmeth.1318.
- K. Yusa, L. Zhou, M. A. Li, A. Bradley and N. L. Craig, A Hyperactive piggyBac Transposase for Mammalian Applications, Proc. Natl. Acad. Sci. U. S. A, 2011, 108(4), 1531–1536, DOI:10.1073/pnas.1008322108.
Footnote |
| † Zhenyong Cheng and Dinghao Liu are co-first authors. |
| This journal is © The Royal Society of Chemistry 2026 |