Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Enhancing predictive modeling with molecular fingerprint fusion strategies
Viktoriia
Turkina
*a,
Melanie R. W.
Messih
a,
Etienne
Kant
a,
Jelle T.
Gringhuis
a,
Annemieke
Petrignani
 a,
Garry
Corthals
a,
Jake W.
O'Brien
a,
Garry
Corthals
a,
Jake W.
O'Brien
 ab and
Saer
Samanipour
ab and
Saer
Samanipour
 *abc
*abc
aVan't Hoff Institute for Molecular Sciences (HIMS), University of Amsterdam, Amsterdam, 1090 GD, The Netherlands. E-mail: v.turkina@uva.nl; s.samanipour@uva.nl
bQueensland Alliance for Environmental Health Sciences (QAEHS), The University of Queensland, 20 Cornwall Street, Woolloongabba, QLD 4102, Australia
cUvA Data Science Center, University of Amsterdam, Amsterdam, The Netherlands
First published on 16th April 2026
Abstract
A large number of chemicals remain poorly characterized in terms of their physicochemical properties, biological activity, and environmental fate. Quantitative structure–activity relationship (QSAR) models have become indispensable tools for predicting these properties, especially for compounds that lack comprehensive experimental data. The choice of structural representation as an input to such models plays a critical role in ensuring high predictive performance and in identifying molecular features that strongly contribute to activity prediction. Both hashed and non-hashed molecular fingerprints are widely employed as inputs in QSAR modeling across various domains. While some studies have explored combining multiple fingerprints to improve molecular representation, comprehensive investigations into different fingerprint fusion strategies and the generalizability of a fused fingerprint across diverse prediction tasks remain limited. In this study, we applied low-, mid-, and high-levels fusion strategies to combine six non-hashed fingerprints and evaluated model performance across six publicly available datasets, including three regression and three classification tasks. Our results demonstrate that mid-level fusion, where fingerprint bits are selectively combined based on their importance within individual models, consistently improves predictive accuracy, as assessed by RMSE and R2 for regression, and F1-score and ROC-AUC for classification. The algorithm developed for molecular fingerprints fusion is methodologically general and can be applied to a wide range of predictive modeling problems or other non-hashed molecular fingerprints.
1 Introduction
The increase in variety and global scale of chemical production has resulted in thousands of new chemicals being introduced into the market and eventually into the environment.1,2 A number of these chemicals have been shown to have adverse impact on environmental and human health.3,4 Due to limited resources, experimental assessment of activities and properties of such a large number of chemicals has become unfeasible. According to the Registration, Evaluation, Authorisation and Restriction of Chemicals (REACH) regulation, among thousands of new chemicals produced in the European Union annually, only substances with production exceeding one tonne per year are required to be registered while the rest remain uncharacterized.5,6 To address this gap, modeling approaches become indispensable in investigating and estimating chemical properties and behavior.7–11A major part of the existing modeling approaches employ QSAR, establishing a relationship, linear or non-linear, between a chemical's structure and its activity.10 For such models, a chemical's structure can be represented in many ways, for example by molecular descriptors, molecular fingerprints (FPs), or graph-based molecular representations.10–16 Molecular descriptors are abstract numerical values that capture different properties of a molecule. Molecular fingerprints simplify the molecular structure into a series of bits or counts in a vector.17,18 In graph-based representations, each node (atom) carries certain features (such as atom type, charge, or hybridization state), and these features are aggregated and processed by computational models, such as graph neural networks (GNNs), to predict molecular properties or activities.19
Among these representations, molecular fingerprints (FPs) deserve particular attention because of their interpretability and broad applicability across machine learning models. FPs provide a standardized way to encode molecular structure into a format that is easy to process while still retaining important chemical information. Moreover, they have become a foundational tool in many cheminformatics applications, including virtual screening,17,20,21 similarity searching, and predictive modeling.
Generally, there are two ways of encoding a molecular structure into FPs: non-hashed, which uses a predefined list of molecular features, and hashed, where the structure is converted with the help of a hash function.17,22,23 The hashed algorithms are specifically tailored to capture intricate details of the structure.24 However, depending on the complexity of the structure, multiple molecular features may be represented with the same bit. This leads to a loss of one-to-one correspondence between bits and structural information.18 Such clarity in interpretation is crucial for establishing structural rules for potential hazard identification.25 While non-hashed FPs avoid this issue, the initial predefined list does not always include a comprehensive set of structural features necessary to effectively distinguish relevant variations in structures.18 This limitation can lead to the exclusion of information essential for accurate modeling.
There are several hashed and non-hashed algorithms to calculate FPs with different levels of structural information included. For example, topological FPs describe molecular features limited to a certain distance, while structure-based FPs depict different aspects of the molecular substructures.17
Although different FPs are extensively used in various fields, it has also been demonstrated that a single way to encode structure into FPs may not capture all the necessary structural information for accurate modeling.26 To address this, several studies have focused on fusing multiple types of representations to enhance predictive performance. For instance, in the context of ligand-based virtual screening, Hert et al. showed that combining similarity scores or ranked lists calculated based multiple fingerprints or reference structures improved retrieval performance and scaffold hopping.27 Fingerprint fusion has also been adopted within supervised predictive modeling frameworks. In case of the most straightforward approach, low-level fusion, multiple FPs are concatenated into a single feature vector. Xie et al., for example, combined MACCS and ECFP fingerprints through horizontal concatenation to improve predictions of log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P and protein–ligand binding affinities.28 However, this method increases feature dimensionality and may introduce redundancy and sparsity, which can challenge model training, especially with large chemical datasets.
P and protein–ligand binding affinities.28 However, this method increases feature dimensionality and may introduce redundancy and sparsity, which can challenge model training, especially with large chemical datasets.
To mitigate such issues, mid-level fusion approaches apply transformation or selection techniques to individual representations before combining them. Srisongkram et al. demonstrated improved performance on KRASG12C ligand affinity prediction by selecting 13 highly target-correlated FP bits from a conjoint PubChem and SSC representation.29 Similarly, Shen et al. applied normalization and principal component analysis (PCA) independently to ECFP4, MACCS, and 208 descriptors, then the transformed features were concatenated for classifying USP7 inhibitors.30 FP-GNN represents another mid-level strategy, integrating molecular fingerprints with graph-based embeddings in a learned latent space.31 However, such representations usually require extensive datasets, which are mostly unavailable for environmental QSAR modeling.32
At the most abstract level, high-level fusion combines predictions from multiple models trained on distinct FPs. Matsuyama and Ishida implemented this strategy using a stacking ensemble of eight different FP types, including Pharmacophore, AP2DC, Daylight, ECFP4, ECFP6, MACCS, Topotor, and Chemdesc, to predict activity across 14 PubChem targets.33
While combining multiple FPs can help capture complementary structural information, data fusion is not without trade-offs. In particular, low-level fusion can significantly expand feature space, leading to high memory usage and reduced scalability. For example, training a model on the full CompTox database34 using several concatenated FPs would require gigabytes of RAM, making it impractical for many QSAR pipelines. These challenges underscore the importance of carefully selecting fusion levels to balance informativeness, model complexity, and resource constraints.
Although fingerprint fusion was explored in several QSAR studies, the main motivation was finding the most effective structural representation to predict specific targets.28,29,35 Systematic investigations into the generalizability of fused fingerprints across multiple fusion levels and prediction tasks remain unexplored.
In this study, we systematically evaluated the effect of fingerprint fusion strategies on QSAR model performance. We selected six publicly available datasets including three for the regression and three for the classification problems. To keep the interpretability of molecular representation, we focused on a set of six non-hashed FPs: E-state, PubChem, MACCS, AtomPairCount, Substructure Count, and Klekotha Roth as structural input for QSAR models. We fused calculated fingerprints on low-, mid-, and high-levels and compared the QSAR performance based on root-mean-square error (RMSE), coefficient of determination (R2) for regression and F1-score, the area under the receiver operating characteristic curve (ROC-AUC) for classification tasks.
2 Methods
2.1 Overall workflow
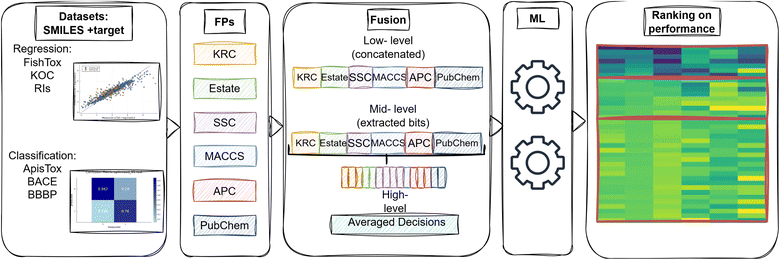
The overall workflow begins with the conversion of canonical SMILES strings36 into a set of non-hashed FPs. For each individual FP, we trained and optimized either a regression or classification model using the Random Forest (RF) algorithm, depending on the target. We then applied low-, mid-, and high-levels fusion strategies to combine the individual fingerprints and assessed the performance of models trained on both individual and fused FPs. These models were evaluated across a range of biological, ecological, and physiochemical prediction tasks (Fig. 1).2.2 Molecular fingerprints
We selected a set of various non-hashed topological and structure-based FPs including Atom Pair 2D Count (AP2DC),37 Electrotopological state (E-state),38 Klekotha-Roth Count (KRC),39 Molecular Access Systems (MACCS),40 PubChem,41 and Substructure Keys Count (SSC) FPs.42 E-state, KRC, MACCS, PubChem, and SSC FPs are substructure-based FPs and AP2DC is a topological one. Each bit within this set corresponds to an interpretable molecular feature, aiding in understanding the significance of structural information related to the targeted activity.Various FP algorithms encompass different aspects and details of molecular structure, meaning they contain different sets of substructures or molecular features in a various number of bits. For instance, AP2DC FP is a 780-bit topological FP defined by the atomic environment and the distance between atomic pairs.37 The shortest among the selected FPs is the 79-bit E-state FP, which is based on both the electronic and topological characteristics of molecules.38 In contrast, KRC FP is the largest one encoded in 4860 bits. It was originally developed to predict certain types of bioactivity through substructure screening and consists of relevant SMART strings.39 The MACCS FP, one of the most frequently used, is constructed using SMART patterns and optimized for substructure searching, resulting in 166 bits.40 Another widely adopted FP, PubChem, covers a wide range of different substructures and features recorded in the PubChem database in a 881-bit binary string.41 Finally, SSC FP consists of a count vector of 307 SMART patterns selected for functional group classification by Christian Laggner.42
The canonical SMILES were used to calculate six non-hashed FPs. All FPs except E-state were computed using the PaDEL software package, implemented via a Python 3 wrapper called PaDELPy (version 0.1.13).42 The E-state FP was calculated using the RDKit software package (version 2022.9.5), which includes Python bindings.43 SMILES for which fingerprint generation failed (e.g., due to parsing errors or unsupported structures) were excluded from subsequent analysis. Across all datasets, the fraction of removed molecules was <1.3% (Table S2). Generated fingerprints were further inspected for invalid entries. Any NaN values, missing fingerprint bits, or extreme values arising during fingerprint calculation were replaced with zero to ensure numerical stability and consistency across datasets. No imputation based on label information was performed, and all preprocessing steps were applied prior to data splitting, meaning the preprocessing steps were applied uniformly to all datasets before training, cross-validation, and test set evaluation.
2.3 Datasets
We collected six datasets from different sources: two from ecotoxicology, two related to physiological parameters, and two to physicochemical properties. Three of the datasets correspond to regression problems, FishTox,44 KOC,45 and RI,46 while the other three, ApisTox,47 BBBP,48 and BACE,49 to classification tasks.FishTox dataset contains experimentally determined acute fish toxicity (LC50[−log (mg L−1)]) for 907 chemicals. The toxicity values are defined as the concentration of the chemicals that resulted in the death of 50% of feathed minnows (Pimephales promelas) throughout 96 h (96 h LC50). These values were collected from three databases: OASIS, ECOTOX, and EAT5, and were supplied by Cassotti et al.50 The chemicals in this data set were derived from different origins, including pharmaceuticals, pesticides, conventional persistent organic pollutants, and industrial chemicals.44
KOC dataset contains experimentally measured logarithmic values of the soil organic carbon-normalized (KOC[log (L kg−1)]) for 824 organic compounds representing 31 different chemical classes. The data was compiled from EPI Suite (Version 4.1) and the literature sources and was provided by Wang et al.45 The values are measured by the standard testing guidelines, e.g. the Organization for Economic Cooperation and Development (OECD) guidelines for estimation of KOC using HPLC and the batch equilibrium method.
RIs dataset contains Retention Indices (RI) for reversed-phase liquid chromatography (RPLC), based on synthetic and naturally occurring homologous series of amphiphilic cocamide diethanolamine surfactants (C(n = 0–23)-DEA). It includes 3018 emerging pollutants, 2290 measured in positive electrospray ionization (+ESI) mode and 728 in negative (−ESI) mode. The RI values were obtained under a variety of liquid chromatography (LC) conditions, including variations in pH, column types, temperatures, flow rates, mobile phase compositions, and gradient elution programs and were supplied by Aalizadeh et al.46
ApisTox consists of 1035 compounds with their experimentally measured acute bee toxicity. The values were collected from the ECOTOX, PPDB, and BPDB databases. The dataset contains 296 toxic and 739 non-toxic molecules. U.S. EPA guidelines for honey bees were used to define toxic/non-toxic labels. Compounds with an LD50 less than or equal to a threshold 11 µg per org were classified as toxic. The dataset was compiled and provided by Adamczyk et al.47
BBBP dataset consists of 2053 molecules compiled from several publications focused on blood–brain barrier (BBB) permeability. Compounds were divided into BBB+ (able to cross the BBB) and BBB− (unable to cross) classes. Chemicals were assigned to the BBB+ class if the blood–brain partition coefficient value (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) BB) is greater than or equal to −1, and to the BBB− class if log
BB) is greater than or equal to −1, and to the BBB− class if log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) BB is less than −1. Overall, the dataset provided by Martins et al. contains 1570 BBB+ and 483 BBB− molecules.48
BB is less than −1. Overall, the dataset provided by Martins et al. contains 1570 BBB+ and 483 BBB− molecules.48
Finally, BACE dataset includes 1547 synthetic inhibitors of human β-secretase 1 (BACE-1). The dataset uses binary labels to indicate compound activity. Compounds were classified as active if their IC50 values, the concentration at which a compound inhibits 50% of its target activity, were ≤100 nM, and inactive otherwise. This dataset was compiled and provided by Subramanian et al.49
2.4 Modeling
For modeling, a Random Forest Regression (RFR) and Random Forest Classification (RFC) algorithm were implemented in Julia with scikit-learn.51 Random Forest is a supervised algorithm that constructs several unique decision trees from bootstrap data. After model development, the predictions resulting from the individual trees are averaged to produce the final RFR model prediction and from the major voting of predictions to produce the final RFC model prediction.52 The major benefit of employing the RF modeling approach lies in its ability to handle nonlinearity and noncontinuity within the data while keeping the interpretability of the results. There are several other widely applied modeling algorithms, such as Support Vector Machine (SVM) or Convolutional Neural Networks (CNNs), for this purpose. However, these algorithms tend to be very data-intensive and have low transparency. Therefore, the RF algorithm was selected to ensure the robustness and transparency of the process. Also, RF was successfully applied previously for QSAR modeling and showed accurate and robust performance.44,53–55 While this study focuses on RF models to ensure a controlled comparison of fusion strategies, the proposed fingerprint fusion framework is not tied to a specific learning algorithm and can, in principle, be combined with other commonly used algorithms, although the magnitude of performance gains may vary.To train individual fingerprint-based models, the datasets were randomly split into a training set (90%) and a test set (10%). The split was selected to maintain an independent set for testing performance while preserving sufficient training data for stable feature importance estimation, which is critical for mid-level fusion. Since such design limits the ability to quantify uncertainty in performance metrics, the observed performance should be interpreted within this limitation. The training set was employed to develop, optimize, and validate the model, while the test set was applied to evaluate the performance of the optimized models. The hyperparameters were optimized using a 3D grid with the number of trees varying from 100 to 600, the number of variables to consider being sqrt or log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2, and the minimum number of samples in each leaf ranging from 2 to 8. For classification tasks, class imbalance was handled using cost-sensitive learning by setting class_weight = “balanced” in the Random Forest classifier. The performance evaluation of the models was based on initial estimation using a training set, internal validation with 3-fold cross-validation, and external validation using a test set.
2, and the minimum number of samples in each leaf ranging from 2 to 8. For classification tasks, class imbalance was handled using cost-sensitive learning by setting class_weight = “balanced” in the Random Forest classifier. The performance evaluation of the models was based on initial estimation using a training set, internal validation with 3-fold cross-validation, and external validation using a test set.
The regression tasks were evaluated by RMSE and R2, while the classification tasks were evaluated by F1-score and ROC-AUC (eqn (S1)–(S5)). Model performance reported in the Results section refers exclusively to the external test set. Training and 3-fold cross-validation were used only for hyperparameter optimization and internal validation.
2.5 PCA
To visualize the chemical space coverage with different fingerprints, we used Principal Component Analysis (PCA) which is an unsupervised method, enabling an unbiased evaluation of the underlying trends in the data. For each dataset, we constructed a PCA model using the horizontally concatenated set of non-hashed fingerprints, representing the maximal structural information available from these fingerprints. By analyzing the resulting scores and loadings, we assessed the chemical space captured by individual fingerprints within each dataset. The algorithm was implemented in Julia using scikit-learn.512.6 HCA
To investigate the similarity and relationships between trained models based on their predictive performance, we applied Hierarchical Cluster Analysis (HCA), an unsupervised clustering technique. Models were clustered according to normalized performance metrics (R2 and F1 scores) across datasets. For each dataset, these scores, obtained from both individual and fused FPs models, were normalized and used as input for clustering. Euclidean distance was employed as the similarity metric, and the average linkage method was used to build the hierarchical clusters. HCA was implemented in Julia using the Clustering.jl package.2.7 Molecular fingerprints' fusion
We evaluated the impact of low-, mid-, and high-levels data fusion strategies on model performance. Importantly, we did not apply any explicit correlation filtering or dimensionality reduction prior to fusion or modelling. This decision was made because of models we used in this study, RF and PCA, are inherently robust to multicollinearity.For low-level fusion, we horizontally concatenated all the bits from six selected non-hashed fingerprints. In addition, we trained models using a reduced version of this fused fingerprint by retaining only those features whose cumulative importance accounted for at least 55% and up to 95% of the total, in 5% steps. The importance was calculated based on impurity-based variable importance. This variable importance metric reflects the reduction of impurity achieved by the splits on each variable during the model training process. This metric indicates the contribution of each feature to explaining the variance in the training data.
For mid-level fusion, we first trained, optimized, validated, and tested Random Forest (RF) models using each individual fingerprint. From each model, we extracted important variables based on two selection strategies: cumulative and individual importance. Specifically, we selected features with a cumulative importance between 55% and 95% (in 5% steps) or with an individual importance between 0.2% and 2.0% (in 0.2% steps). The selected features from each model were then combined to create a fused fingerprint, which served as input for final model development.
Finally, for high-level fusion, we implemented an ensemble approach by averaging the outputs of the six individual fingerprint models. In the case of classification, the final class was assigned based on the averaged class probabilities from these models.
2.8 Calculations and data availability
All calculations were performed on a personal computer (PC) with an Intel Core i7-1260P central processing unit and 32 GB of RAM operating Windows 10 Education version 22H2. All data processing and statistical analyses were performed using Julia language version 1.10.6. All datasets as well as calculated FPs in this study, and interpretation of the optimized FP can be found at https://doi.org/10.5281/zenodo.15791757. The algorithm for a molecular fingerprint fusion was converted into an open-source open-access Julia package and can be found at https://bitbucket.org/viktoriiaturkina/fp_optimization.jl/src/master/.3 Results and discussion
3.1 Individual FPs on PCA-space
To explore the chemical information captured by various FPs, we performed PCA for each dataset individually (Fig. S1–S5). Our goal was to evaluate how effectively individual FPs, as well as their combinations, represent the structural diversity of the target chemical space. If an individual FP encodes information that largely overlaps with the concatenated set of all FPs, its PCA projection should closely resemble that of the full fusion. Conversely, if the FPs capture complementary structural features, the combined coverage of their individual PCA spaces should approximate the coverage observed when all FPs are concatenated, indicating that each contributes unique and non-redundant information.We observed that concatenated FP, combining all six FPs, resulted in the most comprehensive structural coverage, while individual fingerprints captured more limited, specific subspaces. On average, the first two principal components accounted for 64% of the total variance across datasets (PC1 = 46%, PC2 = 18%). Interestingly, we observed a consistent pattern across datasets in terms of how individual FPs contributed to the coverage of PCA space. Specifically, the FPs tended to align with similar directions in PCA space regardless of the dataset, suggesting a stable, dataset-independent contribution pattern. This consistency suggests that different fingerprints systematically capture distinct structural features. Moreover, complementarity across FPs underscores the advantage of their fusion: no single fingerprint captures the full breadth of structural variability, and integrating them can yield a more holistic molecular representation.
For example, in the FishTox dataset (Fig. 2), the first two principal components explained 53.37% of the variance (PC1 = 33.89%, PC2 = 19.48%). The PCA loadings show strong orthogonal contributions from APC and KRC fingerprints along PC1 and PC2, respectively. In contrast, SSC and E-state fingerprints contributed less, while PubChem and MACCS fingerprints had the lowest influence. Notably, PubChem and MACCS are the only bit-vector fingerprints among the set, whereas the others are count-based. This difference in vector type may explain their lower variance contribution. Additionally, we did not observe a consistent relationship between fingerprint vector length and chemical space coverage.
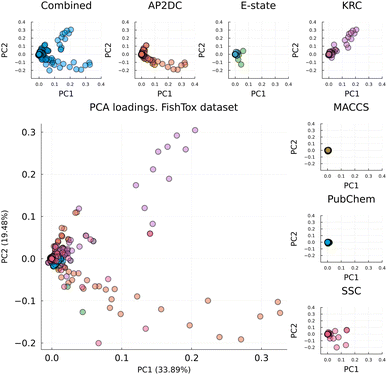 | ||
| Fig. 2 PCA loadings plot of combined fingerprint for FishTox dataset and contribution of each individual fingerprint. | ||
These six molecular FPs differ in how they encode chemical structure information. MACCS, PubChem, and SSC FPs focus on the presence of predefined chemical features or functional groups, such as rings or specific atoms. However, they complement each other because they use different sets of predefined substructures and encoding schemes, leading to a varied coverage of structural features. Atom Pair fingerprints capture the relationships between atom types and the topological distances between them, thus encoding aspects of molecular shape and connectivity. Klekota-Roth fingerprints use a much larger and more detailed set of structural fragments, offering a more comprehensive representation of molecular substructure diversity. In contrast, E-state fingerprints differ fundamentally by integrating both electronic and topological information, characterizing atoms based on their electronic environment within the molecule. Together, these fingerprints provide diverse and complementary ways to capture chemical structure.
Because PCA is unsupervised and does not depend on the prediction target, it provides an unbiased view of how different FPs encode chemical information. The fact that individual FPs showed consistent loading directions across multiple datasets indicates that each fingerprint systematically captures specific and complementary features of chemical structure. This consistency across datasets indicates that different FPs capture largely non-overlapping aspects of molecular structure, and their combination provides a more complete and structured representation of chemical space.
3.2 Individual fingerprint-based models
To evaluate the target-specific predictive performance of individual FPs, independent from their PCA space coverage, we trained, optimized, validated, and tested six RF models per dataset, each using a different FP as input. Model performance was assessed using the coefficient of determination (R2) and root mean squared error (RMSE) for regression tasks, and area under the curve (AUC) and F1-score for classification (Table 1).| Dataset | Metric | Fingerprints | |||||
|---|---|---|---|---|---|---|---|
| KRC | E-state | SSC | MACCS | APC | PubChem | ||
| Regression | |||||||
| FishTox | R 2 | 0.396 | 0.494 | 0.484 | 0.418 | 0.482 | 0.527 |
| RMSE | 1.291 | 1.182 | 1.194 | 1.267 | 1.196 | 1.142 | |
| Koc | R 2 | 0.528 | 0.700 | 0.679 | 0.695 | 0.748 | 0.770 |
| RMSE | 0.910 | 0.726 | 0.752 | 0.732 | 0.664 | 0.635 | |
| RIs | R 2 | 0.656 | 0.654 | 0.692 | 0.651 | 0.695 | 0.672 |
| RMSE | 189.3 | 189.8 | 179.0 | 190.7 | 178.1 | 184.6 | |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||||
| Classification | |||||||
| ApisTox | F 1 | 0.723 | 0.711 | 0.720 | 0.723 | 0.622 | 0.681 |
| AUC | 0.830 | 0.841 | 0.868 | 0.867 | 0.820 | 0.854 | |
| BACE | F 1 | 0.840 | 0.872 | 0.867 | 0.814 | 0.861 | 0.794 |
| AUC | 0.916 | 0.945 | 0.930 | 0.90 | 0.939 | 0.911 | |
| BBBP | F 1 | 0.935 | 0.936 | 0.932 | 0.929 | 0.954 | 0.942 |
| AUC | 0.947 | 0.943 | 0.950 | 0.949 | 0.947 | 0.958 | |
In contrast to the unsupervised PCA analysis, which provided insight into chemical space coverage, the supervised RF models did not reveal a consistent trend in predictive performance based on the type of FP (bit vector vs. count vector). PubChem FP, a bit vector, outperformed the rest of the individual non-hashed FPs for FishTox and Koc datasets. For RI dataset, the best performance was achieved with the SSC and APC, count vector FPs. For ApisTox, strong performance came from KRC, SSC, count vectors, and MACCS, a short 166 bit vector. For the BACE dataset, E-state, the shortest count vector, and SSC showed the best predictive performance. Finally, for BBBP the best results were archived with APC, a count vector, and PubChem, a bit vector. However, it is important to note that overall predictive performance remained modest across all models. These results indicate that the level of structural detail or FP format, count or bit vectors, encoded does not consistently translate to better predictive performance. Furthermore, no association was found between fingerprint length and model performance, implying that not all features encoded by a fingerprint are relevant to specific activity predictions. Instead, irrelevant or redundant structural features may introduce noise into the model, ultimately reducing its predictive power.
Moreover, no single FP is sufficient to fully represent molecular structure, as each captures distinct aspects of chemical information. However, the features identified as important across different FPs are often complementary. For example, the APC FP encodes atom pair occurrences (e.g., C–C, C–X, C–Cl, C–O, C–N, N–O, O–O) at various distances, but does not capture bond types. In contrast, the MACCS fingerprint emphasizes bond types rather than atom identity. The predictive power of the E-state fingerprint primarily stems from its encoding of electronic and topological features related to aromaticity, which are less emphasized in other FPs. KRC highlights diverse oxygen-containing substructures, while SSC-derived models rely heavily on features such as the number of chiral centres and rotatable bonds.
It is important to note that all the FPs used in this study are based on 2D molecular representations and thus do not capture 3D conformational or stereochemical information. This limitation may constrain their ability to model QSAR for targets sensitive to stereochemistry.
3.3 Fingerprints fusion performance
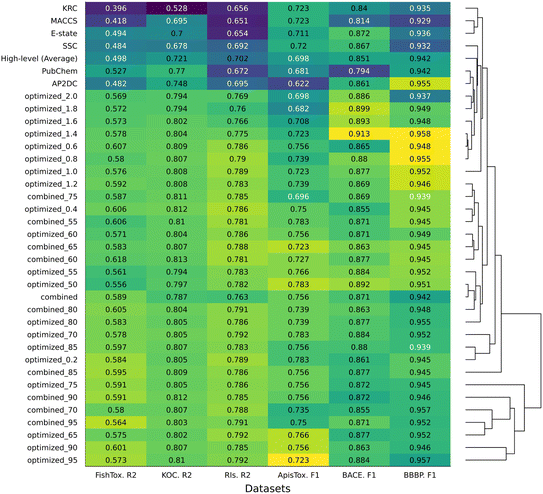
We investigated how different fusion strategies (low-, mid-, and high-levels) affect the predictive power of models built using non-hashed fingerprints (FPs) (Fig. 3). Performance differences were interpreted in terms of consistency and relative trends rather than statistical significance.Mid-level fusion, particularly with features selected based on 95% cumulative importance, yielded the best average performance across tasks. Low-level fusion showed improvements over individual FPs in several datasets. However, its performance was on average lower than that of the best mid-level fused models. This may be due to the inclusion of redundant features, as no feature selection was applied prior to concatenation. In contrast, high-level fusion, which aggregates predictions from individual models, was generally less effective. On average, it was outperformed by individual FPs such as APC and PubChem. Since high-level fusion works at the decision level, this completely misses potential synergies or interactions between features across different FPs. Thus, the simple averaging form of high-level fusion likely underperformed because it could not exploit cross-FP feature interactions, but also treated weak models equally.
In regression tasks, fused FPs showed clear advantages. For instance, in the FishTox dataset, PubChem reached an R2 of 0.53, while the 95% mid-level fused FP achieved 0.62. In the Koc dataset, the top individual R2 was 0.77 (PubChem), compared to 0.81 for a fused FP using a 0.4% individual importance threshold. Similarly, for the RI dataset, APC yielded 0.70, while the fused model reached 0.79.
Classification tasks showed a similar trend. In ApisTox, the best individual F1 score was 0.72 (KRC and MACCS), while the fused 95% model reached 0.81. For BACE, E-state and SSC achieved F1 scores of 0.87, while the fused 1.8% model improved to 0.90. In the BBBP dataset, where individual models already performed strongly (e.g., APC with an F1 of 0.95), fusion strategies offered only marginal improvements, with F1 values ranging from 0.94 to 0.96. Such improvements should be interpreted cautiously, as they may fall within the expected variability of model training and data splitting. In some cases, fused models matched or slightly underperformed compared to the top individual FPs. For example, in ApisTox, fused variants with 1.4%, 70%, 90%, and 95% importance matched the best F1 of 0.72, while others (1.8%, 2.0%) were slightly lower.
HCA of normalized metrics of models performance reveals two main clusters (Fig. 3). The first one is a group comprising the better-performing fused models specifically, including mid-level fused optimized FP from 75% and 95% cumulative importance. While the second is a broader cluster, including individual FPs, high-level fusion, and low-importance threshold optimized models.
Among all models, the optimized 95% fused FP performed best on average across the six datasets (Fig. 3). While low- and high-levels fusion methods slightly improved over individual FPs, mid-level fusion proved to be the most effective approach, improving model performance in most cases. These findings highlight the value of combining complementary features, particularly through selective fusion strategies.
However, no single fusion strategy proved universally optimal across all datasets, which shows the need for case specific optimization approach. Furthermore, while RF algorithm was selected to ensure interpretability and enable controlled comparison of fusion strategies, the outcome of the fused FPs is specific to tree-based QSAR workflows and may not be generalizable to other machine learning algorithms (e.g., SVM, neural networks).
3.4 Feature composition of optimized fingerprints
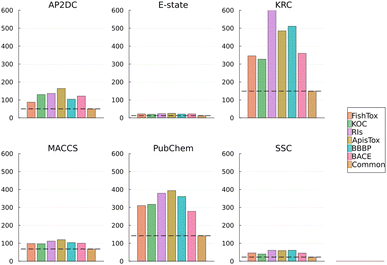
Despite the variability in performance across individual tasks, a mid-level fused FP, composed of features cumulatively accounting for 95% of model importance, achieved the best average performance across datasets. While it was not the top-performing strategy for every case, its overall robustness suggests it can serve as a strong baseline for fusion design. We further investigated its feature composition across different prediction tasks. The goal was to examine how shared and task-specific features contribute to predictive performance, and to identify structural patterns that are consistently informative across different prediction tasks.From the total number of 7800 features from a cumulative set of six FPs, a subset of 444 important bits were selected for each target: 50 from APC, 12 from E-state, 149 from KRFPC, 68 from MACCS, 142 from PubChem, and 23 from SubFPs. Among these, 96 bits specifically describe different aspects of the carbon backbone. These include 8 bits from APC, 7 from E-state, 45 from KRFPC, 16 from MACCS, 14 from PubChem, and 6 from SubFPs. Each individual FP contributed unique representations of carbon-based features. For example, KRFPC encodes different carbon chain lengths with specific bits such as KRFPC3224 (“CC”), KRFPC3640 (“CCC”), and KRFPC3692 (“CCCC”). In contrast, MACCS and SubFPs represent carbon branching characteristics: SubFPC4 for primary carbons and SubFPC2 for secondary carbons. PubChem fingerprints capture information with bits like PubChemFP696, which represents long continuous carbon chains such as “C–C–C–C–C–C–C–C”. E-state descriptors provide information on electronic environments, such as bit 19 for ssssC (quaternary carbon) and bit 13 for sssCH (tertiary carbon with one hydrogen). APC captures the presence of C–C bonds at various topological distances, specifically distances 1 through 5 and 9. The remaining bits describe features involving aromaticity and heteroatoms, such as oxygen, nitrogen, chlorine, sulfur, and fluorine in a variety of substructural contexts. These include aromatic and saturated ring systems, heterocycles, and functional groups. Some fingerprints, such as MACCS, include more generalized representations. For instance, MACCSFP107 (“XA(A)A”) describes the structure where “X” represents any halogen atom (F, Cl, Br, I), and “A” denotes any atom type.
On the other hand, the extracted bits which correspond to prediction of specific target focus more on specific functional groups or heteroatoms. For example, the FishTox model highlights halogenated features such as C–Cl, O–Cl, and Cl–Cl, along with sulfur-containing atoms and aromatic chlorine or bromine substructures, as well as nitro groups, phenols, diaryl ethers, and contributions from long alkyl chains. The KOC model emphasizes halogenation: particularly C–Br and polychlorinated aromatic rings along with large aromatic systems, carboxylic acids, and ethers. The ApisTox model shows signals for phosphorus-based substructures (e.g., C–P, N–P, O–P), as well as N–O and N–S linkages. The RI model is influenced by nitro groups, amines, and long alkyl chains, with features including N–O, O–O, and C–N bonds, high hydrogen and carbon content, and aromatic amines. The BACE model highlights fluorinated and nitrogen-rich substructures such as C–F, N–F, N–N, and N–O, along with amines and fluorinated aromatics. Finally, the BBBP model shares several features with the BACE model, particularly nitrogen- and oxygen-containing substructures (e.g., C–N, N–O, C–S), but also includes peptide-like and carboxylic acid substructures. Overall, these models reflect distinctive substructure patterns associated with their respective prediction tasks.
To further evaluate the impact of individually selected fingerprint bits, we trained models using a shared subset of common bits selected across all tasks, referred to as common bits. Their performance was then compared to that of models trained on task-specific fingerprints optimized at the 95% cumulative importance level. Across the six datasets, the optimized 95% fingerprint consistently demonstrated either improved or comparable predictive performance (Table S3). The most notable gain was observed in the ApisTox dataset, where the F1-score increased from 0.696 to 0.809. FishTox also exhibited modest improvements, while performance for KOC, RIs, and BBBP remained almost unchanged. These results indicate that the addition of specific to individual tasks fingerprint bits can enhance predictive accuracy. Importantly, the common fingerprint bits were selected based on shared importance across all six datasets, suggesting that this representation captures structural features broadly relevant to diverse prediction tasks (Fig. 4).
3.5 Limitations
Although we combined molecular FPs on different fusion level, certain important structural information for target prediction could not be fully captured. For example, most existing fingerprinting algorithms do not encode three-dimensional (3D) molecular characteristics, such as chirality. This limits the ability of QSAR models to differentiate the activity of stereoisomers. On the other hand, features like charge distribution are not explicitly represented in standard FPs, although they may be indirectly captured through correlated structural elements.Furthermore, even the most carefully curated combination of fingerprint keys has limited structural coverage. As a result, highly specific or uncommon substructures associated with niche activity classes may still be underrepresented. This implies that expanding the diversity of the training dataset could enrich the final fused FP, allowing for broader coverage of molecular features relevant to specific targets. Alternatively, ML-based FPs (e.g. MolBERT56), for instance, the ones generated by graph neural networks, can automatically learn relevant structural representations from the data itself, potentially capturing rare or complex features missed by predefined fingerprints. However, this advantage comes at the cost of limited interpretability, often making it difficult or even impossible to trace specific features back to identifiable chemical substructure.
In this study, we focused exclusively on non-hashed fingerprints to preserve interpretability and ensure consistent bit-to-substructure alignment across molecules. This alignment is critical for traditional machine learning models such as Random Forests and SVMs, where each feature (bit) is assumed to have a consistent and interpretable meaning. Hashed fingerprints, in contrast, compress structural information into fixed-length vectors using hashing functions, which can lead to bit collisions (i.e., different substructures mapping to the same bit). This results in a loss of alignment and can introduce noise, reducing model reliability and interpretability. As a result, hashed fingerprints are often better suited for deep learning architectures, which can learn hierarchical representations even when individual features lack clear chemical meaning. Nevertheless, alternative molecular representations, such as circular fingerprints (e.g., ECFP), physicochemical descriptors, or graph-based encodings may still offer complementary information. Integrating these with our current approach could improve predictive performance and offer new opportunities for multi-level fingerprint fusion.
Although our study included six diverse datasets and targets, results may vary for other tasks or datasets with different chemical distributions. Therefore, the conclusions regarding fusion strategies should be interpreted with respect to the data and endpoints investigated here. Moreover, like many widely used benchmarks the selected datasets may have their limitations. Importantly, in this study we do not propose a new state-of-the-art predictor any of the targets, but rather analyze the relative behavior of fingerprint fusion strategies across a range of diverse supervised predictive tasks.
Lastly, the study was limited to the use of RF models to ensure interpretability and a controlled comparison of fusion strategies. Although the proposed fingerprint fusion framework is not algorithm-specific and can be combined and applied to other learning algorithms, the observed performance gains may not directly generalize across different approaches.
4 Conclusions
Molecular representations play a crucial role in achieving the best accuracy of predictive modeling. Non-hashed molecular fingerprints, in particular, offer consistent, human-readable, and computationally efficient structural descriptors, making them valuable tools for assessing chemical behavior. However, individual fingerprint representations may not fully capture all relevant structural information needed for robust QSAR performance. To address this, we explored the impact of various fingerprint fusion strategies, including low-, mid-, and high-level, on QSAR model performance.Our assessment demonstrated that all fusion approaches, except high-level fusion, outperformed models trained on individual fingerprints. This confirms that combining complementary structural features enhances predictive capability. Among the tested strategies, mid-level fusion resulted in the best average performance across six diverse datasets and tasks. In this approach, fingerprint bits with cumulative importance up to 95% were selected from each individual fingerprint and then concatenated to form a conjoint representation. This strategy not only preserved the interpretability of features but also ensured that the most informative structural elements were retained. The mid-level fusion algorithm successfully captured relevant molecular features, particularly for predicting toxicity-related endpoints. Moreover, its flexibility allows application to a wide range of case studies, although the magnitude of performance improvement remains dataset- and endpoint-dependent. By focusing on interpretability and predictive strength, the proposed algorithm represents a practical and generalizable solution for enhancing QSAR modeling through optimized fingerprint fusion.
Author contributions
V. T.: conceptualization, methodology, investigation, visualization, formal analysis, writing – original draft; M. R. W. M.: investigation; E. K.: investigation; J. T. G.: investigation; A. P.: funding acquisition, review and editing; G. G.: review and editing; J. W. O. B.: conceptualization, funding acquisition, supervision, review and editing; S. S.: conceptualization, supervision, methodology, investigation, resources, project administration, funding acquisition, review and editing.Conflicts of interest
The authors declare no conflicts of interest.Data availability
The datasets used for model development, training, and evaluation can be accessed via the following link: https://doi.org/10.5281/zenodo.15791757. The molecular fingerprints, models, figures, and summaries generated during the study are also included in the repository.The source code used for fingerprints calculation, model training, and data-fusion analysis is via the same Zenodo repository (https://doi.org/10.5281/zenodo.15791757). The latest development version of the code, including future updates, is available at https://bitbucket.org/viktoriiaturkina/fp_optimization.jl/src/master/.
Supplementary information (SI): additional details on selected non-hashed molecular fingerprints (Table S1), evaluation metrics (eqn (S1)–(S5)), PCA loadings and score plots for each dataset (Fig. S1–S5), and the performance of the model trained only on common bits (Table S2). See DOI: https://doi.org/10.1039/d5dd00302d.
Acknowledgements
The authors express their gratitude to all the members of Environmental Modeling & Computational Mass Spectrometry (https://www.emcms.info). They are also thankful to Marco Federici, Denice van Herwerden, and Adan Rotteveel for their kind help and contribution to this study. S. S., A. P., and V. T. thank the ChemistryNL TKI and the UvA Data Science Centre for their funding support (projects EDIFIED and SCOPE). J. W. O. B. is the recipient of a National Health and Medical Research Council (NHMRC) Investigator Grant (EL12009209) funded by the Australian Government. S. S. also acknowledges financial support from the Australian National Health and Medical Research Council (NHMRC; APP1185347). The Queensland Alliance for Environmental Health Sciences (QAEHS), The University of Queensland (UQ), gratefully acknowledges the financial support of Queensland Health. For the purposes of open access, the author has applied a Creative Commons Attribution (CC BY) license to any Author Accepted Manuscript version arising from this submission.Notes and references
- H. P. H. Arp, D. Aurich, E. L. Schymanski, K. Sims and S. E. Hale, Environ. Sci. Technol., 2023, 57, 6355–6359 CrossRef CAS PubMed.
- Z. Wang, G. W. Walker, D. C. Muir and K. Nagatani-Yoshida, Environ. Sci. Technol., 2020, 54, 2575–2584 CrossRef CAS PubMed.
- B. I. Escher, H. M. Stapleton and E. L. Schymanski, Science, 2020, 367, 388–392 CrossRef CAS PubMed.
- R. Vermeulen, E. L. Schymanski, A. L. Barabási and G. W. Miller, Science, 2020, 367, 392 CrossRef CAS PubMed.
- C. Rudén and S. O. Hansson, Environ. Health Perspect., 2010, 118, 6 CrossRef PubMed.
- D. C. Muir and P. H. Howard, Environ. Sci. Technol., 2006, 40, 7157–7166 CrossRef CAS PubMed.
- M. T. Cronin, J. S. Jaworska, J. D. Walker, M. H. Comber, C. D. Watts and A. P. Worth, Environ. Health Perspect., 2003, 111, 1391 CrossRef CAS PubMed.
- P. H. Howard and D. C. Muir, Environ. Sci. Technol., 2011, 45, 6938–6946 CrossRef CAS PubMed.
- K. T. Butler, D. W. Davies, H. Cartwright, O. Isayev and A. Walsh, Nature, 2018, 559(7715), 547–555 CrossRef CAS PubMed.
- M. Sigurnjak Bureš, M. Cvetnić, M. Miloloža, D. Kučić Grgić, M. Markić, H. Kušić, T. Bolanča, M. Rogošić and Š. Ukić, Environ. Chem. Lett., 2021, 19, 1629–1655 CrossRef.
- K. Roy, R. N. Das and P. L. Popelier, Chemosphere, 2014, 112, 120–127 CrossRef CAS PubMed.
- M. Karelson, V. S. Lobanov and A. R. Katritzky, Chem. Rev., 1996, 96, 1027–1043 CrossRef CAS PubMed.
- H. Liu, E. Papa and P. Gramatica, Chem. Res. Toxicol., 2006, 19, 1540–1548 Search PubMed.
- P. Gedeck, B. Rohde and C. Bartels, J. Chem. Inf. Model., 2006, 46, 1924–1936 CrossRef CAS PubMed.
- F. Melnikov, J. Kostal, A. Voutchkova-Kostal, J. B. Zimmerman and P. T. Anastas, Green Chem., 2016, 18, 4432–4445 RSC.
- J. Mao, J. Akhtar, X. Zhang, L. Sun, S. Guan, X. Li, G. Chen, J. Liu, H. N. Jeon, M. S. Kim, K. T. No and G. Wang, iScience, 2021, 24, 103052 CrossRef CAS PubMed.
- A. Cereto-Massagué, M. J. Ojeda, C. Valls, M. Mulero, S. Garcia-Vallvé and G. Pujadas, Methods, 2015, 58–63 CrossRef PubMed.
- F. Grisoni, D. Ballabio, R. Todeschini and V. Consonni, Methods Mol. Biol., 2018, 1800, 3–53 CrossRef CAS PubMed.
- M. Boulougouri, P. Vandergheynst and D. Probst, Nat. Mach. Intell., 2024, 6, 754–763 CrossRef.
- I. Muegge and P. Mukherjee, Expert Opin. Drug Discovery, 2016, 11, 137–148 CrossRef CAS PubMed.
- W. M. Czarnecki, S. Podlewska and A. J. Bojarski, J. Cheminf., 2015, 7, 1–15 Search PubMed.
- G. Huang, J. Li and C. Zhao, Molecules, 2018, 23, 954 CrossRef PubMed.
- D. Warszycki, L. Struski, M. Smieja, R. Kafel and R. Kurczab, J. Chem. Inf. Model., 2021, 61, 5054–5065 CrossRef CAS PubMed.
- D. Rogers and M. Hahn, J. Chem. Inf. Model., 2010, 50, 742–754 CrossRef CAS PubMed.
- M. T. Cronin, F. J. Bauer, M. Bonnell, B. Campos, D. J. Ebbrell, J. W. Firman, S. Gutsell, G. Hodges, G. Patlewicz, M. Sapounidou, N. Spînu, P. C. Thomas and A. P. Worth, Regul. Toxicol. Pharmacol., 2022, 135, 105249 CrossRef CAS PubMed.
- Y. J. Tseng, A. J. Hopfinger and E. X. Esposito, J. Comput. Aided Mol. Des., 2012, 26, 39–43 CrossRef CAS PubMed.
- J. Hert, P. Willett, D. J. Wilton, P. Acklin, K. Azzaoui, E. Jacoby and A. Schuffenhauer, J. Chem. Inf. Model., 2006, 46, 462–470 CrossRef CAS PubMed.
- L. Xie, L. Xu, R. Kong, S. Chang and X. Xu, Front. Pharmacol, 2020, 11, 606668 CrossRef PubMed.
- T. Srisongkram, P. Khamtang and N. Weerapreeyakul, J. Mol. Graph. Model., 2023, 122, 108466 CrossRef CAS PubMed.
- W. f. Shen, H. w. Tang, J. b. Li, X. Li and S. Chen, J. Cheminf., 2023, 15, 1–16 Search PubMed.
- H. Cai, H. Zhang, D. Zhao, J. Wu and L. Wang, Briefings Bioinf., 2022, 23, 1–12 CAS.
- K. V. Chuang, L. M. Gunsalus and M. J. Keiser, J. Med. Chem., 2020, 63, 8705–8722 CrossRef CAS PubMed.
- Y. Matsuyama and T. Ishida, Lecture Notes in Computer Science, Springer Verlag, 2018, vol. 10955, pp. 279–288 Search PubMed.
- A. J. Williams, C. M. Grulke, J. Edwards, A. D. McEachran, K. Mansouri, N. C. Baker, G. Patlewicz, I. Shah, J. F. Wambaugh, R. S. Judson and A. M. Richard, J. Cheminf., 2017, 9, 1–27 Search PubMed.
- W. Ding, Y. Nan, J. Wu, C. Han, X. Xin, S. Li, H. Liu and L. Zhang, Comput. Biol. Med., 2022, 144, 105390 CrossRef PubMed.
- D. Weininger, J. Chem. Inf. Comput. Sci., 1988, 28, 31–36 CrossRef CAS.
- D. H. Smith, R. E. Carhart and R. Venkataraghavan, J. Chem. Inf. Comput., 1985, 25, 64–73 CrossRef.
- L. H. Hall and L. B. Kier, J. Chem. Inf. Comput., 1995, 35, 1039–1045 CrossRef CAS.
- J. Klekota and F. P. Roth, Bioinformatics, 2008, 24, 2518–2525 CrossRef CAS PubMed.
- J. L. Durant, B. A. Leland, D. R. Henry and J. G. Nourse, J. Chem. Inf. Comput., 2002, 42, 1273–1280 CrossRef CAS PubMed.
- S. Kim, J. Chen, T. Cheng, A. Gindulyte, J. He, S. He, Q. Li, B. A. Shoemaker, P. A. Thiessen, B. Yu, L. Zaslavsky, J. Zhang and E. E. Bolton, Nucleic Acids Res., 2021, 49, D1388–D1395 CrossRef CAS PubMed.
- C. W. Yap, J. Comput. Chem., 2011, 32, 1466–1474 CrossRef CAS PubMed.
- RDKit: Open-source cheminformatics Software(accessed 23/01/2024), 2024, https://www.rdkit.org/.
- S. Samanipour, J. W. O'Brien, M. J. Reid, K. V. Thomas and A. Praetorius, Environ. Sci. Technol., 2023, 57, 17950–17958 CrossRef CAS PubMed.
- Y. Wang, J. Chen, X. Yang, F. Lyakurwa, X. Li and X. Qiao, Chemosphere, 2015, 119, 438–444 CrossRef CAS PubMed.
- R. Aalizadeh, V. Nikolopoulou and N. S. Thomaidis, Anal. Chem., 2022, 94, 15987–15996 CrossRef CAS PubMed.
- J. Adamczyk, J. Poziemski and P. Siedlecki, Sci. Data, 2025, 12, 1–15 Search PubMed.
- I. F. Martins, A. L. Teixeira, L. Pinheiro and A. O. Falcao, J. Chem. Inf. Model., 2012, 52, 1686–1697 CrossRef CAS PubMed.
- G. Subramanian, B. Ramsundar, V. Pande and R. A. Denny, J. Chem. Inf. Model., 2016, 56, 1936–1949 CrossRef CAS PubMed.
- M. Cassotti, D. Ballabio, R. Todeschini and V. Consonni, SAR QSAR Environ. Res., 2015, 26, 217–243 CrossRef CAS PubMed.
- F. Pedregosa, V. Michel, O. Grisel, M. Blondel, P. Prettenhofer, R. Weiss, J. Vanderplas, D. Cournapeau, F. Pedregosa, G. Varoquaux, A. Gramfort, B. Thirion, O. Grisel, V. Dubourg, A. Passos, M. Brucher, M. Perrot and E. Duchesnay, J. Mach. Learn. Res., 2011, 12, 2825–2830 Search PubMed.
- V. Svetnik, A. Liaw, C. Tong, J. Christopher Culberson, R. P. Sheridan and B. P. Feuston, J. Chem. Inf. Comput., 2003, 43, 1947–1958 CrossRef CAS PubMed.
- D. van Herwerden, J. W. O'Brien, S. Lege, B. W. Pirok, K. V. Thomas and S. Samanipour, Anal. Chem., 2023, 95, 12247–12255 CrossRef CAS PubMed.
- F. Yang, D. van Herwerden, H. Preud’homme and S. Samanipour, Molecules, 2022, 27(19), 6424 CrossRef CAS PubMed.
- P. G. Polishchuk, E. N. Muratov, A. G. Artemenko, O. G. Kolumbin, N. N. Muratov and V. E. Kuz’min, J. Chem. Inf. Model., 2009, 49, 2481–2488 CrossRef CAS PubMed.
- N. Wen, G. Liu, J. Zhang, R. Zhang, Y. Fu and X. Han, J. Cheminf., 2022, 14(1), 71 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |