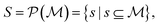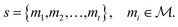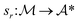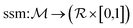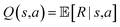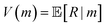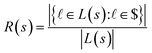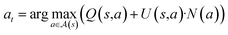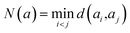Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Synthesis planning in reaction space: a study on success, robustness and diversity
Alan Kai
Hassen
 *ad,
Helen
Lai
*ad,
Helen
Lai
 b,
Samuel
Genheden
b,
Samuel
Genheden
 c,
Mike
Preuss
c,
Mike
Preuss
 d and
Djork-Arné
Clevert
d and
Djork-Arné
Clevert
 a
a
aMachine Learning Research, Pfizer Research and Development, Berlin, Germany. E-mail: AlanKai.Hassen@pfizer.com
bMolecular AI, Discovery Sciences, R&D, AstraZeneca, Cambridge, UK
cMolecular AI, Discovery Sciences, R&D, AstraZeneca, Gothenburg, Sweden
dLeiden Institute of Advanced Computer Science, Leiden University, Leiden, The Netherlands
First published on 25th February 2026
Abstract
Computer-aided synthesis planning aims to identify viable synthetic routes from a target compound to readily available building blocks by iteratively decomposing molecules into smaller precursors. Self-play search algorithms, trained with simulated experience, reach state-of-the-art performance. However, these methods typically plan in the molecular rather than the reaction space, leading to redundant or near-duplicate reaction outcomes in the search tree. In this work, we introduce a reaction-centric planning approach that measures the novelty of proposed reactions to analyze the synthesis planning search problem, constraining the search problem to genuinely unexplored disconnection ideas, i.e., unique ways of decomposing molecules using reactions. Our results show that the overall synthesis planning search space is much smaller than expected due to the absence of diverse disconnection ideas within the underlying template-based retrosynthesis model. Surprisingly, we also find that, under a reasonable time budget of less than an hour, online search algorithms outperform state-of-the-art self-play methods and are more robust to environmental changes, such as minor modifications to the available purchasable building blocks. Finally, we show that the diversity of the synthesis route solution space saturates when combining the results of different search algorithms, highlighting the importance of the single-step model in providing novel and chemically valid disconnections.
1 Introduction
Artificial intelligence is considerably changing the traditional Design–Make–Test–Analyze cycle of drug discovery. In the “Make” phase, computer-aided synthesis planning (CASP) can suggest potential synthetic routes by repeatedly deconstructing a target molecule into smaller precursors until a set of commercially available molecules, also called building blocks, is found.1,2 The ability to find these synthesis routes is a key task in modern chemistry as it allows chemists to plan the creation of molecules,3 provides a general measure of synthesizability4 and can be integrated as an objective into de novo drug design pipelines to ensure that generated compounds are synthesizable.5Modern CASP systems6–8 consist of two core components:3 first, a single-step model that encapsulates the backward reaction logic of how to disconnect a molecule into potential reactants with a neural network in a supervised learning task (e.g. ref. 9–11). Second, a search algorithm whose objective is to find a synthesis route to building blocks within the possible search space provided by the single-step model. Inevitably, these search algorithms must balance exploration and exploitation of the potential state space, as the space is too big for an exhaustive search, given that real-world synthesis routes can involve multiple steps and that many possible disconnection alternatives can be added to the search tree for each molecule. There are currently three major research branches that explore the challenge of estimating the expected reward of finding a synthesis route in a search tree for a molecule or for a reaction:
The pioneering approaches in the field rely on online value estimation in the search tree, mirroring the initialization phase of earlier AlphaGo approaches that were pre-trained with human expert games,12 for example with Monte-Carlo Tree Search (MCTS) using policy guidance,1 Depth-First Proof-Number Search,13 holistic hyper-graph exploration strategy,14 or A*-type best-first search of Retro*-0.15 However, these approaches generally underperform in comparison to methods that use offline supervised learning heuristics, which learn to estimate the expected rewards from historical synthesis data, or self-play methods, which approximate the expected rewards through simulated experiences (originating from AlphaGo variants12,16). Prominent examples that rely on historical synthesis routes to learn a guiding policy include Retro*15 or RetroGraph.17 Self-play, achieving state-of-the-art performances in the field, is applied by using simulated experience to train a value network,18 experience-guided MCTS to learn the success of using reactions19 or experience-guided Retro* to learn the success of decomposing a molecule.20 Moreover, self-play methods are used to apply an actor-critic approach,21 re-rank single-step model predictions22 or adjust the underlying reaction templates of the single-step model.23
This divide between well-performing self-play methods and less-effective online value estimation methods is a serious problem for the field of CASP because, unlike Chess or Go, CASP is not a stationary environment, making learned reward estimations less transferable and often requiring constant retraining:
• The underlying building blocks that constitute the winning states of the game can change dramatically depending on the application. For example, modern CASP tasks may rely on a specific small building block set,4 search for specific intermediate molecules24,25 or optimize multiple route characteristics simultaneously.26
• The reaction suggestions of the single-step model, which are used as the basis of the search tree, can change drastically depending on the model architecture27 or based on the reaction data that is used.28,29
• The historical reactions or simulated experiences used to train the tree policy models are generated by “playing” the game of retrosynthesis against a set of predefined molecules taken from a specific chemical space. However, as real-world reaction and molecule modalities are continually changing,30 this learned chemical space may not cover the target molecule for which synthesis planning is conducted.
To address these problems, we use reaction diversity as a guiding principle in online synthesis planning algorithms (see Fig. 1), prioritizing the exploration of novel reaction disconnection ideas inspired by prior approaches that use reaction diversity in a greedy, best-first tree search.8,31 Here, similiar reactions are pruned to reduce the search space, guiding the tree search by minimizing the distance to a provided external reaction database of literature synthesis routes,31 or expand the diversity of disconnection ideas before filtering for validity.8 Our work, beyond improving diversity in candidate synthesis route solution spaces, enables an in-depth analysis of the synthesis planning search problem. With this:
• We show that the synthesis planning search problem is computationally much more straightforward than previously assumed when using template-based single-step models as the diversity of suggested reaction disconnections is limited, leading to an effective branching factor of roughly 3 after clustering.
• We demonstrate that online value estimation in synthesis planning can outperform or match self-play methods when sufficient single-step model calls are provided within a reasonable time budget, thus allowing adaptation to non-stationary environments (where we show self-play methods underperform to adapt).
• We highlight that our reaction diversity search yields a diverse set of synthesis routes, allowing a chemist to choose from a manageable solution set by avoiding redundant analog building blocks, and provide general guidelines for successful synthesis planning.
2 Methods
2.1 Computer-aided synthesis planning
The synthesis planning problem is the task of identifying a synthesis route sr for a target molecule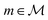 by recursively breaking down m into precursor reactants
by recursively breaking down m into precursor reactants 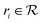 until either all reactants belong to a (commercially) available building block set $ or the search budget (e.g. time, iterations) is exhausted. The state space S for this search is defined as the power set
until either all reactants belong to a (commercially) available building block set $ or the search budget (e.g. time, iterations) is exhausted. The state space S for this search is defined as the power set 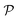 of
of 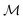 :
:where
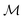 is the space of all possible molecules and each state s ∈ S represents a set of molecules:
is the space of all possible molecules and each state s ∈ S represents a set of molecules:The synthesis route sr is then formally defined as a mapping:
| sr(m) = (a1, a2, …, ak) |
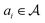 transforming a molecule mi into its precursor set ri:
transforming a molecule mi into its precursor set ri:| ai(mi) → ri. |
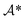 of retrosynthetic actions (a1, a2, …, ak), where each action ai transforms an intermediate molecule mi into its precursor set ri. sr is considered solved if, after applying all actions (a1, a2, …, ak) in order, the resulting set of molecules s is in the available building block set (s ⊆ $). A single-step retrosynthesis model, denoted by ssm, provides candidate precursor sets for a given molecule:
of retrosynthetic actions (a1, a2, …, ak), where each action ai transforms an intermediate molecule mi into its precursor set ri. sr is considered solved if, after applying all actions (a1, a2, …, ak) in order, the resulting set of molecules s is in the available building block set (s ⊆ $). A single-step retrosynthesis model, denoted by ssm, provides candidate precursor sets for a given molecule:where
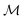 is the space of molecules,
is the space of molecules, 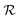 is the space of possible reactant sets, and
is the space of possible reactant sets, and 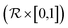 represents reactant sets paired with a likelihood score. Given m, the model outputs:
represents reactant sets paired with a likelihood score. Given m, the model outputs:| ssm (m) = {((ri)1, p1), …, ((ri)n, pn)} |
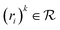 is a set of reactants forming m, and pk ∈ [0, 1] is the corresponding reaction disconnection probability. Whenever more than one candidate set of reactants is returned for a given molecule, the primary task of synthesis planning is to pick the disconnection that will most likely lead to a solved synthesis route. From a reinforcement learning perspective, the core challenge of the search algorithm is to learn or estimate the expected reward Q(s, a) given a search tree state s and a possible reaction alternative a in state-based tree representation, encapsulating the entire search problem of a possible synthesis route with all its molecules in a state s:1
is a set of reactants forming m, and pk ∈ [0, 1] is the corresponding reaction disconnection probability. Whenever more than one candidate set of reactants is returned for a given molecule, the primary task of synthesis planning is to pick the disconnection that will most likely lead to a solved synthesis route. From a reinforcement learning perspective, the core challenge of the search algorithm is to learn or estimate the expected reward Q(s, a) given a search tree state s and a possible reaction alternative a in state-based tree representation, encapsulating the entire search problem of a possible synthesis route with all its molecules in a state s:1or the expected reward of expanding a specific molecule in an AND-OR search tree V(m), representing alternative reactions as OR-nodes and all reaction precursor molecules m as AND-nodes:15
In this setting, the reward R is defined based on the set of leaf molecules L(s) that are readily available ($) in the current search tree state s:
Synthesis planning approaches can estimate Q(s, a) or V(m) “online” while running a search algorithm (e.g. ref. 1), learn a “heuristic” via supervised learning based on historical synthesis routes (e.g. ref. 15), or use “simulated experience” by repeatedly running synthesis planning against a predefined set of molecules, recording the successful synthesis pathways, and iteratively refining either Q(s, a)19 or V(m).22
2.2 Reaction importance
A key question in a tree search is which follow-up actions (in our case, reactions ak) are actually relevant for solving a search problem. In synthesis planning, the single-step model ssm provides a data-driven reaction likelihood pk derived from the reactions of historical synthesis pathways (based on previously targeted compounds, used reactions and building blocks). However, this likelihood pk is not necessarily equal to the expected reward of finding a full synthesis route in the search tree, that is, pk ≠ Q(s, a). Despite this, many online search approaches initialize Q(s, a) with pk because it is a reasonable way to prioritize historically probable reactions (e.g. ref. 6). Alternative approaches learn a re-ranking of pkvia simulated experience,22 but consequently fix the search environment to a set of targets, reactions and building blocks used to generate the simulated experience. However, if one does not wish to fix the search environment because the set of target molecules, single-step models, reactions and building blocks shifts over time (or are not fixed), we must ask what constitutes a “good” ranking to approximate Q(s, a) online using pk, given that the true tree policy *Q(s, a) (or *V(m)) remains unknown. Importantly, the single-step model ssm (m) often produces sets of very similar reactions ak, with minor variations of the resulting reactants ri (e.g. ref. 31). Consequently, not all suggested alternatives ak are equally valuable to explore, as CASP systems are known to produce many similar (analog) reactions for the same target.To address this, we propose quantifying the novelty N(a) of each reaction, measuring how much it differs from other candidate reactions instead of filtering them (e.g. ref. 31). We then incorporate N(a) into the classical exploration–exploitation trade-off in two distinct ways:
We define N(a) as the minimal distance between the current reaction aj and all previously explored reactions ai:
where
 is the set of resulting clusters. We incorporate ssmclustered(m) into both the often-used best-first search algorithm Retro*-0,15 without neural guiding policy, and into MCTS.1
is the set of resulting clusters. We incorporate ssmclustered(m) into both the often-used best-first search algorithm Retro*-0,15 without neural guiding policy, and into MCTS.1
2.3 Experimental setup
We closely follow the evaluation protocols established by Dual Value Networks22 and Syntheseus36 for the synthesis planning search problem to establish comparability between different approaches. Specifically, we use the same single-step, template-based retrosynthesis model as Retro*15 and Dual Value Networks,22 which provides up to 50 candidate retrosynthetic disconnections (reaction suggestions) per molecule, and integrate it into AiZynthfinder6 through the Models Matter single-step model adapter.29 Following Dual Value Networks, we set the number of search algorithm iterations to 500 but do not restrict the number of single-step model calls within an algorithm iteration. We deviate from this restriction because online estimation of Q(s, a) should require multiple calls for a fair algorithm comparison, especially when self-play approaches train a model using simulated experiences but exclude those training calls from the single-step inference count. Although Dual Value Networks do not enforce a strict search-depth limit (but do restrict depth in the experience-generation phase), we choose a maximum synthesis route depth of 30 to limit our search to a reasonable maximum route length. We evaluate our approach on three datasets on their provided test splits:22 USPTO-190, a widely adopted benchmark with a solved rate above 90%, which is still considered standard; ChEMBL-1000, which is highly relevant for pharmaceutical applications; and GDB17-1000, an enumerated set of molecules that presents a challenging benchmark of unexplored molecules. Notably, ChEMBL-1000 and GDB17-1000 are processed versions of subsamples from ChEMBL and GDB17, where known building blocks are removed, and molecules have at least a molecular weight, Bertz coefficient, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, and TPSA larger than the mean of the respective values in USPTO-190. Unless otherwise noted, we use the eMolecules stock37 (22,876,046 molecules), consistent with prior works.15,22 Any deviations from these default settings are explicitly stated throughout this work. We run all experiments embarrassingly parallel on an HPC cluster using only CPUs. The only exceptions are experiments regarding Dual Value Networks,22 which we train and run inference on multiple Nvidia H100 GPUs.
P, and TPSA larger than the mean of the respective values in USPTO-190. Unless otherwise noted, we use the eMolecules stock37 (22,876,046 molecules), consistent with prior works.15,22 Any deviations from these default settings are explicitly stated throughout this work. We run all experiments embarrassingly parallel on an HPC cluster using only CPUs. The only exceptions are experiments regarding Dual Value Networks,22 which we train and run inference on multiple Nvidia H100 GPUs.
3 Results
3.1 Initial experiment
The results of evaluating Distance-MCTS, Cluster-Retro*-0 and Cluster-MCTS are presented in Table 1. Distance-MCTS outperforms both Dual Value Networks22 and RetroGraph17 in terms of synthesis planning success on the pharmaceutically relevant ChEMBL-1000 dataset while being only slightly worse on the unknown enumerated chemical space of GDB17-1000 and the patent-based chemical space of USPTO-190. This result is surprising given that our approach is considerably more straightforward than the prior methods, which rely on complex reaction network modeling17 or pre-training with simulated experiences.22 Furthermore, both Cluster-Retro*-0 and Cluster-MCTS reduce the overall performance by 5–10%, compared to the non-clustered variants of Retro*-0 and Distance-MCTS, but they are still surprisingly effective. This success is unexpected, as we did not anticipate that achieving this strong synthesis planning performance with an online search algorithm would be easily attainable. These results raise several questions: (1) which factors drive the success of synthesis planning, (2) how robust are different algorithms to changing environments (given the initial assumption that online approaches would underperform self-play), and (3) how large a solution space can be explored via online search (given that the search performance seems to be comparable). We answer these questions in the following sections to clarify the strengths and limitations of online synthesis planning methods.| Source | Search structure | Tree policy | Algorithm | ChEMBL-1000 (%) | GDB17-1000 (%) | USPTO-190 (%) |
|---|---|---|---|---|---|---|
| 15 | Search tree | Online | Retro*-0 | 75.10 | 7.50 | 79.47 |
| 15 | Search tree | Heuristic | Retro* | 76.20 | 9.50 | 85.79 |
| 20 | Search tree | Self-play | Retro* + -0 | 81.10 | 15.00 | 96.32 |
| 20 | Search tree | Self-play | Retro*+ | 81.80 | 15.40 | 90.53 |
| 22 | Search tree | Self-play | PDVN + Retro*-0 | 83.50 | 26.90 | 98.95 |
| Our | Search tree | Online | Cluster-Retro*-0 | 70.70 | 7.30 | 71.58 |
| Our | Search tree | Online | Cluster-MCTS | 76.60 | 12.30 | 75.26 |
| Our | Search tree | Online | Distance-MCTS | 85.30 | 25.90 | 95.26 |
| 17 | Reaction network | Heuristic | RetroGraph | 85.20 | 21.50 | 99.47 |
| 22 | Reaction network | Self-play | PDVN + RetroGraph | 86.00 | 37.10 | 99.47 |
3.2 The success of synthesis planning
Given that Distance-MCTS achieves state-of-the-art performance on ChEMBL-1000 while remaining competitive on the other datasets, we performed an ablation study to investigate the more important question of why this happens beyond setting a new state-of-the-art. In particular, we examined how the number of single-step calls affects the overall success rate, motivated by the fact that the main difference between Distance-MCTS and the other approaches is that multiple ssm calls can be performed within a single Monte Carlo rollout iteration while a best-first search algorithm always picks the highly ranked frontier molecule for expansion. In addition to the previous experiment, we included the default implementations of MCTS1 and Retro*-0![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 15 as baselines, but increased the number of iterations to 25
15 as baselines, but increased the number of iterations to 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 for all Retro* variants (default & cluster) to match the single-step model calls used by the MCTS-based methods. We then compared these results against the self-play performance of Dual Value Networks.22
000 for all Retro* variants (default & cluster) to match the single-step model calls used by the MCTS-based methods. We then compared these results against the self-play performance of Dual Value Networks.22
Fig. 2 shows how varying the number of single-step model calls affects the synthesis planning success rate for different search algorithms. There is a clear relationship between more single-step model calls and a better solved rate for all three datasets and all online search algorithms. These results indicate that self-play variants (e.g. ref. 22) learn to find a synthesis route faster but do not perform better, unlike their motivating AlphaGo variants12,16 that outperform online search algorithms. Furthermore, the search times for online search algorithms are surprisingly short, even when using cheap CPU inference (see Fig. S1). Finally, the search space for each dataset seems to have a natural performance limit that all well-performing algorithms converge upon over time. When looking at the different algorithms, the difference between MCTS and Distance-MCTS is relatively small, even though Distance-MCTS performs slightly better. Both algorithms outperform self-play on ChEMBL-1000 and reach comparable performance on USPTO-190 and GDB17-1000. Clustering the ssm reactions decreases the performance for all algorithms by ∼20% in total, compared to the non-clustered variant, yet it retains a surprisingly high performance given that the effective search space is drastically reduced in the reaction space, from an average of 25.37 alternatives added per expansion call to 3.02 for ChEMBL-1000 and MCTS (Retro* on ChEMBL-1000: 22.54 → 2.80). This relatively low drop in performance on a much smaller clustered search space shows that the average search space for a synthesis planning problem is, in principle, much smaller than previously expected, as the diversity in suggested distinct reaction ideas is lacking, and the search space mainly consists of reactions with slightly varying reactant outcomes. Note that the width usually never reaches the ideal 50 alternatives, as direct duplicates and erroneous reactions are removed from the predictions. Finally, picking the single-step model ssm suggested alternative in a best-first search is surprisingly effective for Retro*-0 on the USPTO-190 benchmark dataset. Here, the 95% solved rate is reached within 6000 single-step model calls, and it matches previously reported state-of-the-art performance at the 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 mark. These results raise the question of how reliable the evaluation dataset is—and, consequently, the reported performance of search algorithms on it—if picking the best alternative suggested by the ssm can achieve these performance levels as quickly as our results indicate. Our results further indicate that USPTO-190 may be very much in-domain of the ssm as it is created from USPTO patent data15 and thus requires little search effort. These results are supported by the Retro*-0 performance on the GDB17-1000 dataset, where picking the best-first alternative does not work well because the enumerated molecular space is not as well known to the single-step model. In this setting, the search success rate increases only linearly after initially solving the more familiar molecules. To summarize, the computational search problem of synthesis planning is more straightforward than expected as a best-first search with enough single-step model calls is very competitive to self-play algorithms, and the effective search space after clustering is much smaller than the expected 5030.1
000 mark. These results raise the question of how reliable the evaluation dataset is—and, consequently, the reported performance of search algorithms on it—if picking the best alternative suggested by the ssm can achieve these performance levels as quickly as our results indicate. Our results further indicate that USPTO-190 may be very much in-domain of the ssm as it is created from USPTO patent data15 and thus requires little search effort. These results are supported by the Retro*-0 performance on the GDB17-1000 dataset, where picking the best-first alternative does not work well because the enumerated molecular space is not as well known to the single-step model. In this setting, the search success rate increases only linearly after initially solving the more familiar molecules. To summarize, the computational search problem of synthesis planning is more straightforward than expected as a best-first search with enough single-step model calls is very competitive to self-play algorithms, and the effective search space after clustering is much smaller than the expected 5030.1
3.3 The robustness of synthesis planning
In the last section, we showed that search algorithms' success is primarily based on the number of single-step model calls and that online search algorithms can reach the performance of self-play approaches if given enough single-step calls. However, search algorithms, both online and self-play, must be able to adapt to new pharmaceutical targets, reactions, and building blocks, which can differ substantially from the original ssm training distributions or self-play environments. Since we cannot publicly share new targets or reaction data, we change the overall synthesis planning search environment by modifying the available building blocks in eMolecules (winning states) to simulate the non-stationary nature of building block availability. For example, building block availability can vary because the desired building blocks might be out of stock or beyond the budget for a synthesis campaign. For this purpose, we remove all building blocks from our stock dataset that appear in synthesis routes found by the best-first search algorithm Retro*-0 within 500 iterations in USPTO-190, ChEMBL-1000 and GDB17-1000. We pick Retro*-0 specifically because an ideal search algorithm should be able to find synthesis routes beyond the most apparent, best-first solutions that are explored by following the ssm reactions with the highest probability within the initial search space. We remove 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 874 building blocks (0.0126%) from our initial stock dataset of 22
874 building blocks (0.0126%) from our initial stock dataset of 22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 876
876![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 046, leaving 22
046, leaving 22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 873
873![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 172 building blocks. As this amounts to only 0.0126% of the building blocks, the challenge of the search becomes to substitute these specific best-first building blocks within a large space of possible alternatives. This task could be challenging if building blocks without suitable analogs are removed or alternative reaction pathways that use entirely different building blocks must be found. We then repeat the prior experiment to measure the relationship between synthesis planning success and single-step model calls with this new building block set on USPTO-190, ChEMBL-1000 and GDB17-1000. Furthermore, we evaluate Dual Value Networks22 on this reduced set without retraining, meaning the algorithm remains pre-trained on the full eMolecules dataset, as this approach allows us to assess how well self-play methods generalize.
172 building blocks. As this amounts to only 0.0126% of the building blocks, the challenge of the search becomes to substitute these specific best-first building blocks within a large space of possible alternatives. This task could be challenging if building blocks without suitable analogs are removed or alternative reaction pathways that use entirely different building blocks must be found. We then repeat the prior experiment to measure the relationship between synthesis planning success and single-step model calls with this new building block set on USPTO-190, ChEMBL-1000 and GDB17-1000. Furthermore, we evaluate Dual Value Networks22 on this reduced set without retraining, meaning the algorithm remains pre-trained on the full eMolecules dataset, as this approach allows us to assess how well self-play methods generalize.
Surprisingly, removing 0.0126% of the best-first synthesis route building blocks leads to a large drop in synthesis planning performance for all datasets (see Fig. 3). Performance on USPTO-190 and GDB17-1000 decreases across all algorithms to less than half the initial best performance, while the drop in ChEMBL-1000 is considerably lower. In terms of algorithms, using a best-first algorithm with Retro*-0 outperforms self-play and all MCTS variants. The generalizability of self-play might be worse than expected, as these small changes in the search environment result in considerable performance drops, and the best-first search algorithm outperforms self-play in earlier iterations compared to the complete building block set. Given that we removed the building blocks of the best-first route, the task here is to find analog molecules that substitute the missing building blocks—a task that a best-first search can handle most effectively, as analogs should be highly ranked on the best-first exploration frontier. MCTS variants, however, tend to discourage the entire reaction pathway because the negative reward of the best-first route is backpropagated through the search tree and discourages the search for analogs once the original best-first route returns a negative reward. Consequently, the difference between Distance-MCTS and normal MCTS is rather small, whereas the default implementation of MCTS seems to perform slightly better as it has access to direct reaction analogs that might be discouraged by Distance-MCTS. All clustering variants that operate in reaction disconnection search space require that the newly found synthesis route differs from the best-first route; that is, the new route would need to use at least one reaction idea within the synthesis route that is not part of the best-first route, implying structurally different pathways for synthesizing the molecule beyond the best-first approach. Noteworthy, such a structurally different pathway that leads to different building blocks could be as simple as switching the order of molecule decompositions in a synthesis route or further decomposing missing building blocks (see SI Fig. S4 for an example). This usage of different reaction ideas appears not to be possible, as the synthesis route found rates decrease drastically compared to the complete building block set for USPTO-190 (Cluster-Retro*-0: 81.58% → 8.95%, Cluster-MCTS: 75.26% → 7.37%) and GDB17-1000 (Cluster-Retro*-0: 14.60% → 1.00%, Cluster-MCTS: 12.30% → 1.10%), where it seems that a specific best-first reaction route must be found. For ChEMBL-1000, however, finding alternative synthesis routes is possible (Cluster-Retro*-0: 77.60% → 39.10%, Cluster-MCTS: 76.60% → 35.00%), indicating that, in principle, there are structurally different ways of synthesizing these molecules. Notably, these algorithms are not just solving molecules that lie beyond the 500-iteration boundary for Retro*-0, as the solved rate is substantially higher than the increase in solved routes beyond 500 iterations in the first experiment (compare Fig. 2). To summarize, the ability to adapt to changes in the search environment is surprisingly limited. While a best-first search algorithm should, in principle, find solutions faster than MCTS and be more robust than a self-play algorithm, all algorithms struggle to find “meaningful” reaction alternatives when the ssm does not provide sufficiently diverse reaction disconnections, as no reaction pathway can be found that is not suggested by the ssm.
3.4 The diversity of synthesis planning
Our initial motivation for Distance-MCTS was not to develop an online search algorithm that outperforms a self-play algorithm, but to suggest more structurally distinct synthesis routes to our chemists. For this purpose, we repeat our first experiment but return all synthesis routes discovered within a 2-hour time budget to explore the possible synthesis route space. Specifically, we use 500 iterations for the MCTS variants and 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 single-step model calls for the Retro* variants. We sample 100 molecules from the ChEMBL-1000 dataset as an evaluation dataset from the set of molecules for which all online search algorithms found a synthesis route in our first experiment. We focus exclusively on solved molecules here because we are interested in how the solution space changes once a valid solution can be found, rather than whether an algorithm can solve a molecule in the first place—hence, we exclude unsolved cases from this analysis. We first evaluate the average number of found synthesis routes across evaluated molecules during the search to measure the size of the overall solution space. Additionally, we examine the average shortest route found by each algorithm. While a high number of discovered synthesis routes may indicate broad exploration of the search space, many of these routes can be close variants differing by minor modifications through the use of analog building blocks instead of genuinely different reaction pathways. Thus, the total route count alone could be misleading. Therefore, we compute the average pair-wise distance33 between the top-100 returned routes across all molecules, measuring the necessary steps to transform one synthesis route into another. A higher average distance implies that the found set of routes is more diverse, aligning with our goal of offering chemists more than one principal route rather than minor variations of essentially the same route. We do not cluster the entire route space due to computational constraints and because each algorithm should ideally return a diverse set of alternatives within a manageable top-n.
000 single-step model calls for the Retro* variants. We sample 100 molecules from the ChEMBL-1000 dataset as an evaluation dataset from the set of molecules for which all online search algorithms found a synthesis route in our first experiment. We focus exclusively on solved molecules here because we are interested in how the solution space changes once a valid solution can be found, rather than whether an algorithm can solve a molecule in the first place—hence, we exclude unsolved cases from this analysis. We first evaluate the average number of found synthesis routes across evaluated molecules during the search to measure the size of the overall solution space. Additionally, we examine the average shortest route found by each algorithm. While a high number of discovered synthesis routes may indicate broad exploration of the search space, many of these routes can be close variants differing by minor modifications through the use of analog building blocks instead of genuinely different reaction pathways. Thus, the total route count alone could be misleading. Therefore, we compute the average pair-wise distance33 between the top-100 returned routes across all molecules, measuring the necessary steps to transform one synthesis route into another. A higher average distance implies that the found set of routes is more diverse, aligning with our goal of offering chemists more than one principal route rather than minor variations of essentially the same route. We do not cluster the entire route space due to computational constraints and because each algorithm should ideally return a diverse set of alternatives within a manageable top-n.
All molecules remain solvable across all tested algorithms (see Fig. 4 and SI Table S3). Among these algorithms, the MCTS variants (default and distance-based) return the highest number of routes, averaging between 2700 and 2400. Retro*-0 and Cluster-Retro*-0 follow with around 1700 routes. The only outlier is Cluster-MCTS, which returns an average of roughly 260 routes. Nevertheless, every algorithm provides numerous routes to choose from in its respective solution space. Regarding the shortest route found per target, most algorithms produce routes that average three to four reactions. However, novelty-based algorithms tend to produce slightly longer synthesis routes, where especially Cluster-Retro*-0 returns routes that are on average more than one reaction longer. Looking more closely at the diversity of these routes, we observe that non-novelty-based algorithms tend to produce only minor variations, reflected in low distances within the top-100 returned routes and consequently suggesting a less diverse set of solutions. In contrast, Distance-MCTS produces slightly more distinct alternatives in the top-100. Finally, algorithms that rely on clustering to search in the reaction space (Cluster-Retro*-0 and Cluster-MCTS) return the most structurally distinct synthesis routes, substantially increasing the mean route distance.
In a second step, we analyze the building blocks that the respective algorithms use as end-points for their synthesis routes, as these define the possible fragmentations of the target molecules. For this purpose, we compare the set of unique building blocks found for each molecule by each algorithm against the molecule results of all other algorithms (see Fig. 5 and SI Table S4). We first assess the Building Block Coverage, defined as the percentage of a single algorithm's unique building blocks for a given molecule relative to the total set of unique building blocks found by all algorithms for that same molecule. Here, non-clustering MCTS-based methods cover the most building block space, with an average coverage of around 55%. In contrast, best-first search strategies perform worse on average (31% to 39%), and Cluster-MCTS performs the worst with roughly only 18%. These results align with the route analysis, supporting the observation that clustering methods actively remove analog building blocks from the search space. We also measured the contribution of Unique Building Blocks, which are specific to a single algorithm. Generally, most algorithms only contribute a small fraction of unique building blocks, averaging around 10% to 13%. The cluster-based methods yield the fewest unique building blocks with roughly 2% for Cluster-MCTS and 9% for Cluster-Retro*. We can also compare the building block space to the search setting in Experiment 2, where the “best-first” building blocks were removed. For each algorithm, around 36% to 40% of all unique building blocks for each molecule became unavailable, thereby reducing the average percentage of previously solved routes per molecule to below 2% for all algorithms. This decrease demonstrates that the majority of the routes identified in the standard-setting rely heavily on these “best-first” building blocks. At the molecular level, this filtering results in a vast decrease in the number of solved molecules. Solved rates drop from initially finding a synthesis route for all molecules to solving roughly 50% of the molecules for non-clustered algorithms, and only around 15% for the clustered algorithms. This disparity strongly highlights the importance of analog building blocks for successful synthesis planning in this setting, as these are precisely what the clustering methods are designed to prune. Finally, we compare the rate of molecules for which a synthesis route is found under two conditions: (1) post-run filtering of routes based on the reduced building block dataset and (2) rerunning the entire search algorithm on the reduced building block set from the start (100 ChEMBL molecule results from Experiment 2). This comparison highlights the importance of algorithm robustness to changes in the search environment. Across all search algorithms, rerunning the search on the changed environment improves the success rate by 20% to 50% in total. This improvement underscores the importance of online search algorithms in synthesis planning, which can adapt to the available building block space rather than relying on a static set of building blocks.
Following this analysis, we investigate which combinations of algorithms cover the largest unique building block space when aggregated across all molecules (see Fig. 6) using an UpSet plot.38 For our 100 ChEMBL molecules, the total unique building block space is 28![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 260. The general trend shows a logarithmic relationship: combining more algorithms increases total coverage of the building block space, but with diminishing returns for each additional algorithm when selecting the best-performing ones, which aligns with their previously discussed per-molecule coverage and uniqueness rates of different algorithms. Individually, the highest coverage is provided by the Distance and standard MCTS algorithms, each covering roughly 57% of all unique building blocks. In contrast, the clustered methods cover a much smaller portion of the space. Combining any two search algorithms, with the exception of Cluster-MCTS, greatly increases unique building block coverage to approximately 70% to 77%. A combination of three such algorithms (excluding Cluster-MCTS) already accounts for 83% to 90% of the total building block space. Finally, complete coverage of the 28
260. The general trend shows a logarithmic relationship: combining more algorithms increases total coverage of the building block space, but with diminishing returns for each additional algorithm when selecting the best-performing ones, which aligns with their previously discussed per-molecule coverage and uniqueness rates of different algorithms. Individually, the highest coverage is provided by the Distance and standard MCTS algorithms, each covering roughly 57% of all unique building blocks. In contrast, the clustered methods cover a much smaller portion of the space. Combining any two search algorithms, with the exception of Cluster-MCTS, greatly increases unique building block coverage to approximately 70% to 77%. A combination of three such algorithms (excluding Cluster-MCTS) already accounts for 83% to 90% of the total building block space. Finally, complete coverage of the 28![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 260 building blocks is achieved only when combining results from nearly all algorithms. This finding is consistent with our earlier observation that, while most algorithms overlap, many still contribute a small percentage of unique building blocks.
260 building blocks is achieved only when combining results from nearly all algorithms. This finding is consistent with our earlier observation that, while most algorithms overlap, many still contribute a small percentage of unique building blocks.
As an additional case study, we applied our diversity search methodology to two products (A and B) previously used to evaluate guidance by an external literature database.31 Notably, we conducted this evaluation without relying on a proprietary route database or closed-source single-step retrosynthesis model, as these are publicly unavailable. Consistent with the ChEMBL100 pattern, the maximum unique building block space discovered generally follows a logarithmic scale, reaching saturation when different algorithms are combined (see SI Fig. S6 and S7). Quantitatively, we found between 500 and 1300 routes with unique building blocks for each algorithm for Product A. In contrast, Product B proved more challenging, as Distance-MCTS yielded 14 routes with unique building blocks, while Cluster-Retro* failed to find any route (see SI Table S5). Qualitatively, we conceptually reproduced the reported route for Product A using Distance-MCTS (see Fig. SI S5). However, our route is one step shorter, having eliminated a protection step by identifying better building blocks in our synthesis space exploration, a result that previously required literature guidance.31 While our approach identifies the same conceptual reaction steps, it utilizes different building blocks. For Product B, the route we identified corresponds conceptually to the slightly longer, non-literature-guided synthesis route, albeit with a premature deprotection step. Here, none of our synthesis routes utilize a [3 + 2] cycloaddition ring-forming reaction identified by literature guidance.31 This is surprising given that the building blocks required for the cycloaddition route are present in our building block set. Considering that online search algorithms achieved success rates comparable to self-play methods in solvable cases (see Fig. 2) and that coverage of the unique solution building block space generally saturates after combining different algorithms (see Fig. 6), our inability to find a route utilizing the cycloaddition is unexpected. This strongly suggests that the template-based single-step model failed to predict the specific disconnection required, highlighting the importance of valid reaction predictions from the single-step model as the limiting factor, rather than the search algorithm's coverage of the synthesis route solution space.
In summary, all algorithms return numerous potential synthesis routes, where the building block solution space can be improved when combining different algorithm results. Nevertheless, cluster-based approaches find a more manageable solution space at the cost of potentially missing a shorter route, as not all analog building blocks are available during the search. The breadth of synthesis route solutions, potentially combining results from different algorithms, enables subsequent route optimization based on desired objectives (e.g. cost, availability, safety), ensuring that chemists can choose the optimal route that meets their specific project requirements from among many route alternatives.
4 Conclusion
Using different search algorithms, we investigated the success, robustness, and route diversity of synthesis planning. We focused on increasing diversity in the search by (i) encouraging the exploration of otherwise underexplored reactions (Distance-MCTS) and (ii) reducing the search space by planning in reaction disconnection space through reaction clustering of the single-step model (Cluster-Retro-0*, Cluster-MCTS).Our results show that the synthesis planning search problem of successfully finding a synthesis route may be computationally more straightforward than previously expected, as the template-based single-step model tends to suggest only a limited number of distinct reaction disconnection ideas and consequently reducing the search space greatly from the assumed 50treedepth1 to roughly 3treedepth, when clustering, or 25treedepth, when not clustering. We also demonstrated that the main driver for success in synthesis planning is the number of single-step model calls an algorithm can use. Specifically, we observed a logarithmic relationship between increasing the number of single-step calls and the fraction of molecules for which a valid route is found. Beyond a certain threshold of single-step calls, the choice of search algorithm becomes less critical, as most methods eventually discover a possible synthesis route, indicating a natural performance plateau that all search algorithms reach over time. To our surprise, online estimation of Q(s, a) and V(m) performed competitively with state-of-the-art self-play variants.22 Those self-play methods mainly gain faster inference (in terms of single-step calls) but do not achieve better synthesis planning performance. This finding deviates from what is observed in the game of Go, which inspired most self-play variants in synthesis planning, where self-play dramatically boosts performance compared to an online search algorithm.16 Unlike Go, synthesis planning is not a stationary environment with fixed rules and unchanging winning states. We show that self-play implicitly assumes a stationary environment to function well in synthesis planning.
Nevertheless, our findings do not render self-play variants entirely ineffective. Self-play can be advantageous if the environment is effectively fixed, for example, when using a never-changing set of public reactions like USPTO39 or a small set of fixed in-house building blocks.4 It might also be required when a fast synthesis planning result is necessary, such as when synthesis planning is used as an objective in de novo drug design.4,5 However, the substantial GPU resources required by self-play raise questions about its practicality for more general applications. From a purely practical standpoint, there is little difference between finding a solution in 500 or 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 iterations using only CPU infrastructure as long as the total inference time is under an hour. Furthermore, fixing the environment offers fewer benefits when applying synthesis planning to novel targets, reactions, or building blocks.
000 iterations using only CPU infrastructure as long as the total inference time is under an hour. Furthermore, fixing the environment offers fewer benefits when applying synthesis planning to novel targets, reactions, or building blocks.
We also observe that benchmarking synthesis planning experiments exclusively on the USPTO-190 dataset can be problematic. Because USPTO-190 is in-domain for the publicly available USPTO-based single-step model, it requires no substantial search effort beyond a best-first search. In contrast, we show that planning performance on more “unknown” enumerated spaces (e.g. GDB17-1000) can be surprisingly poor, raising concerns about out-of-distribution performance for truly novel targets. We want to highlight this divergence between well-known chemical spaces (e.g. USPTO-190, ChEMBL-1000) and more unexplored ones (e.g. GDB17-1000), where current methods perform remarkably worse.
Regarding the robustness of synthesis planning, it is notable how strongly synthesis planning depends on the availability of key building blocks. We observe remarkable declines in route planning success when few key building blocks are missing, even though millions of alternative building blocks are still available. Furthermore, no alternative disconnection reaction can be found if the disconnection idea in the best-first route is unavailable. This lack of alternative reactions is problematic because a perfect synthesis planning algorithm would provide chemists with a structurally diverse set of synthesis routes, allowing them to choose the appropriate ones based on project requirements. Here, the main advantage of our clustering and diversity-oriented search approaches (Cluster-Retro*-0, Cluster-MCTS, Distance-MCTS) is that they decrease the synthesis planning solution space by directly operating in reaction disconnection space and potentially exploring a wider range of reaction ideas. Such algorithms can reduce the overall solution space by providing diverse synthesis routes for chemists to choose from, rather than slight variations of the same core building blocks when returning all found synthesis routes.
Based on our insights, we recommend (i) a best-first search with a large number of iterations (e.g. 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000) for well-known chemical spaces (e.g. USPTO-190, ChEMBL-1000) and (ii) an MCTS variant in other cases as synthesis planning search settings. In both cases, an hour of search time should suffice, provided single-step model inference is fast enough. For maximum coverage of the synthesis route solution space, we recommend combining the results of multiple search algorithms on the molecule-specific search problem defined by the project's available building blocks, rather than relying on a fixed general search environment with post-run route filtering.
000) for well-known chemical spaces (e.g. USPTO-190, ChEMBL-1000) and (ii) an MCTS variant in other cases as synthesis planning search settings. In both cases, an hour of search time should suffice, provided single-step model inference is fast enough. For maximum coverage of the synthesis route solution space, we recommend combining the results of multiple search algorithms on the molecule-specific search problem defined by the project's available building blocks, rather than relying on a fixed general search environment with post-run route filtering.
Naturally, our work has certain limitations. First, we emphasize that we used the default search algorithm benchmark with a fixed, template-based single-step model. Different single-step models27,29 could yield different reaction distributions, potentially altering the results, by using either a template-free11,40,41 or an ensemble of different single-step models.42 Here, especially Transformer-based methods could increase the diversity of the effective branching factor of the single-step model (e.g. ref. 31), which are typically limited by their beam size of 10 to 20 alternatives candidate reactions.41 Second, we only evaluate synthesis planning search algorithms on their route-finding ability, but do not evaluate follow-up questions regarding the quality of the produced routes. Instead, we mainly focus on the search success problem, as it is a requirement for further route optimization. We treat the single-step model as a benchmark-defined black-box to allow comparability with other search approaches under the same evaluation environment. However, it might be interesting to change the evaluation environment by altering the single-step model to return only reactions that satisfy the round-trip prediction by ensuring the correct product is predicted given the retrosynthesis reactants (e.g. ref. 8 and 31) or to evaluate the found routes for different algorithms with a chemist to verify their overall validity as algorithmically found synthesis routes must ultimately work in real-world laboratory. Third, our reaction clustering remains an approximation of reaction diversity as we rely on heuristic reaction fingerprints instead of using the underlying reaction mechanisms. Future work could improve reaction representations by testing alternative reaction fingerprints43,44 or clustering methods and their parameterization to enhance our approach further. Our results suggest that key advances in the synthesis planning field will originate from improving the single-step model, particularly in generating a more diverse set of reaction disconnections. Such diversity would expand the overall reaction search space and enable more innovative synthesis route solutions.
Author contributions
Alan Kai Hassen: conceptualization, data curation, formal analysis, investigation, methodology, software, validation, visualization, writing – original draft, writing – review & editing. Helen Lai: conceptualization, methodology, writing – review & editing. Samuel Genheden: conceptualization, methodology, supervision, writing – review & editing. Mike Preuss: conceptualization, methodology, funding acquisition, supervision, writing – review & editing. Djork-Arné Clevert: conceptualization, methodology, funding acquisition, resources, supervision, writing – review & editing.Conflicts of interest
There are no conflicts to declare.Data availability
The code for Synthesis Planning in Reaction Space (SPRS) is openly available on GitHub at https://github.com/AlanHassen/SPRS. The datasets generated and analyzed in this study are available in the Figshare repository at https://doi.org/10.6084/m9.figshare.29409725.Supplementary information (SI): algorithm configuration parameters, search performance statistics, reaction clustering examples, and further case study information. See DOI: https://doi.org/10.1039/d5dd00280j.
Acknowledgements
This study was partially funded by the European Union's Horizon 2020 research and innovation program under the Marie Skłodowska-Curie Innovative Training Network European Industrial Doctorate grant agreement no. 956832 “Advanced machine learning for Innovative Drug Discovery”. Large Language Models (LLMs) were used throughout the creation of this manuscript to improve spelling mistakes, grammar, and the overall reading flow. All LLM suggestions were carefully checked for correctness and refined by the authors of this work. The LLM was not used for any research-related tasks.References
- M. H. Segler, M. Preuss and M. P. Waller, Nature, 2018, 555, 604–610 CrossRef CAS PubMed.
- E. J. Corey and X.-M. Cheng, The Logic of Chemical Synthesis, John Wiley & Sons, Ltd, New York, 1989 Search PubMed.
- P. Schwaller, A. C. Vaucher, R. Laplaza, C. Bunne, A. Krause, C. Corminboeuf and T. Laino, WIREs Comput. Mol. Sci., 2022, 12, e1604 CrossRef.
- A. K. Hassen, M. Šícho, Y. J. van Aalst, M. C. W. Huizenga, D. N. R. Reynolds, S. Luukkonen, A. Bernatavicius, D.-A. Clevert, A. P. A. Janssen, G. J. P. van Westen and M. Preuss, J. Cheminf., 2025, 17, 41 Search PubMed.
- J. Guo and P. Schwaller, Chem. Sci., 2025, 16, 6943–6956 RSC.
- L. Saigiridharan, A. K. Hassen, H. Lai, P. Torren-Peraire, O. Engkvist and S. Genheden, J. Cheminf., 2024, 16, 57 Search PubMed.
- Z. Tu, S. J. Choure, M. H. Fong, J. Roh, I. Levin, K. Yu, J. F. Joung, N. Morgan, S.-C. Li, X. Sun, H. Lin, M. Murnin, J. P. Liles, T. J. Struble, M. E. Fortunato, M. Liu, W. H. Green, K. F. Jensen and C. W. Coley, Acc. Chem. Res., 2025, 58, 1764–1775 CrossRef CAS PubMed.
- D. Kreutter and J.-L. Reymond, Chem. Sci., 2023, 14, 9959–9969 RSC.
- M. H. Segler and M. P. Waller, Chem.–Eur. J., 2017, 23, 5966–5971 CrossRef CAS PubMed.
- S. Chen and Y. Jung, JACS Au, 2021, 1, 1612–1620 CrossRef CAS PubMed.
- R. Irwin, S. Dimitriadis, J. He and E. J. Bjerrum, Mach. Learn.: Sci. Technol., 2022, 3, 015022 Search PubMed.
- D. Silver, A. Huang, C. J. Maddison, A. Guez, L. Sifre, G. van den Driessche, J. Schrittwieser, I. Antonoglou, V. Panneershelvam, M. Lanctot, S. Dieleman, D. Grewe, J. Nham, N. Kalchbrenner, I. Sutskever, T. Lillicrap, M. Leach, K. Kavukcuoglu, T. Graepel and D. Hassabis, Nature, 2016, 529, 484–489 CrossRef CAS PubMed.
- A. Kishimoto, B. Buesser, B. Chen and A. Botea, Advances in Neural Information Processing Systems, 2019 Search PubMed.
- P. Schwaller, R. Petraglia, V. Zullo, V. H. Nair, R. A. Haeuselmann, R. Pisoni, C. Bekas, A. Iuliano and T. Laino, Chem. Sci., 2020, 11, 3316–3325 RSC.
- B. Chen, C. Li, H. Dai and L. Song, Proceedings of the 37th International Conference on Machine Learning, Virtual, 2020, pp. 1608–1616 Search PubMed.
- D. Silver, J. Schrittwieser, K. Simonyan, I. Antonoglou, A. Huang, A. Guez, T. Hubert, L. Baker, M. Lai, A. Bolton, Y. Chen, T. Lillicrap, F. Hui, L. Sifre, G. Van Den Driessche, T. Graepel and D. Hassabis, Nature, 2017, 550, 354–359 CrossRef CAS PubMed.
- S. Xie, R. Yan, P. Han, Y. Xia, L. Wu, C. Guo, B. Yang and T. Qin, Proceedings of the 28th ACM SIGKDD Conference on Knowledge Discovery and Data Mining, 2022, pp. 2120–2129 Search PubMed.
- J. S. Schreck, C. W. Coley and K. J. Bishop, ACS Cent. Sci., 2019, 5, 970–981 CrossRef CAS PubMed.
- S. Hong, H. H. Zhuo, K. Jin, G. Shao and Z. Zhou, Commun. Chem., 2023, 6, 120 CrossRef PubMed.
- J. Kim, S. Ahn, H. Lee and J. Shin, Proceedings of the 38th International Conference on Machine Learning, 2021, pp. 5486–5495 Search PubMed.
- Y. Yu, Y. Wei, K. Kuang, Z. Huang, H. Yao, F. Wu, Advances in Neural Information Processing Systems, 2022 Search PubMed.
- G. Liu, D. Xue, S. Xie, Y. Xia, A. Tripp, K. Maziarz, M. Segler, T. Qin, Z. Zhang and T.-Y. Liu, Proceedings of the 40th International Conference on Machine Learning, 2023, pp. 22266–22276 Search PubMed.
- X. Zhang, H. Lin, M. Zhang, Y. Zhou and J. Ma, Nat. Commun., 2025, 16, 192 CrossRef CAS PubMed.
- D. Armstrong, Z. Jončev, J. Guo and P. Schwaller, Digital Discovery, 2025, 4, 2570–2578 RSC.
- K. Yu, J. Roh, Z. Li, W. Gao, R. Wang and C. W. Coley, The Thirty-eighth Annual Conference on Neural Information Processing Systems, 2024 Search PubMed.
- H. Lai, C. Kannas, A. K. Hassen, E. Granqvist, A. M. Westerlund, D.-A. Clevert, M. Preuss and S. Genheden, Artif. Intell. Life Sci., 2025, 7, 100130 CAS.
- A. K. Hassen, P. Torren-Peraire, S. Genheden, J. Verhoeven, M. Preuss and I. V. Tetko,NeurIPS 2022 AI for Science: Progress and Promises, 2022 Search PubMed.
- S. Genheden, P.-O. Norrby and O. Engkvist, J. Chem. Inf. Model., 2023, 63, 1841–1846 CrossRef CAS PubMed.
- P. Torren Peraire, A. K. Hassen, S. Genheden, J. Verhoeven, D.-A. Clevert, M. Preuss and I. V. Tetko, Digital Discovery, 2024, 3, 558–572 RSC.
- S. Genheden and G. P. Howell, Org. Process Res. Dev., 2024, 28, 4225–4239 CrossRef CAS.
- F. Zipoli, C. Baldassari, M. Manica, J. Born and T. Laino, npj Comput. Mater., 2024, 10, 101 CrossRef.
- D. Rogers and M. Hahn, J. Chem. Inf. Model., 2010, 50, 742–754 CrossRef CAS PubMed.
- S. Genheden, O. Engkvist and E. Bjerrum, J. Chem. Inf. Model., 2021, 61, 3899–3907 CrossRef CAS PubMed.
- B. J. Frey and D. Dueck, Science, 2007, 315, 972–976 CrossRef CAS PubMed.
- F. Pedregosa, G. Varoquaux, A. Gramfort, V. Michel, B. Thirion, O. Grisel, M. Blondel, P. Prettenhofer, R. Weiss, V. Dubourg, J. Vanderplas, A. Passos, D. Cournapeau, M. Brucher, M. Perrot and E. Duchesnay, J. Mach. Learn. Res., 2011, 12, 2825–2830 Search PubMed.
- K. Maziarz, A. Tripp, G. Liu, M. Stanley, S. Xie, P. Gaiński, P. Seidl and M. H. S. Segler, Faraday Discuss., 2025, 256, 568–586 RSC.
- eMolecules, Inc., eMolecules Chemical Building Blocks, 2023, https://www.emolecules.com/products/building-blocks Search PubMed.
- A. Lex, N. Gehlenborg, H. Strobelt, R. Vuillemot and H. Pfister, IEEE Trans. Visual. Comput. Graph., 2014, 20, 1983–1992 Search PubMed.
- D. M. Lowe, PhD thesis, University of Cambridge, 2012.
- I. V. Tetko, P. Karpov, R. Van Deursen and G. Godin, Nat. Commun., 2020, 11, 5575 CrossRef CAS PubMed.
- M. Andronov, N. Andronova, M. Wand, J. Schmidhuber and D.-A. Clevert, J. Cheminf., 2025, 17, 31 CAS.
- K. Maziarz, G. Liu, H. Misztela, A. Tripp, J. Li, A. Kornev, P. Gaiński, H. Hoefling, M. Fortunato, R. Gupta and M. Segler, Chemist-Aligned Retrosynthesis by Ensembling Diverse Inductive Bias Models, arXiv, 2025, preprint, arXiv:2412.05269, DOI:10.48550/arXiv.2412.05269.
- P. Schwaller, D. Probst, A. C. Vaucher, V. H. Nair, D. Kreutter, T. Laino and J.-L. Reymond, Nat. Mach. Intell., 2021, 3, 144–152 CrossRef.
- D. Probst, P. Schwaller and J.-L. Reymond, Digital Discovery, 2022, 1, 91–97 RSC.
| This journal is © The Royal Society of Chemistry 2026 |