Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Artificial intelligence and machine learning for plasmonic and surface-enhanced sensing
Ailsa
Geddis
 ,
Hannah
Williams
,
Saba
Bashir
,
Hannah
Williams
,
Saba
Bashir
 ,
Jason
Malenfant
,
Jason
Malenfant
 ,
Caroline
Dubois
,
Caroline
Dubois
 ,
Louis
Hamlet
and
Jean-François
Masson
,
Louis
Hamlet
and
Jean-François
Masson
 *
*
Département de Chimie, Institut Courtois, Centre Interdisciplinaire de Recherche sur le Cerveau et L'apprentissage, Quebec Center for Advanced Materials, Regroupement Québécois sur les Matériaux de Pointe, Université de Montréal, C.P. 6128 Succ. Centre-ville, Montréal, Québec, H3C 3J7, Canada. E-mail: jf.masson@umontreal.ca
First published on 25th February 2026
Abstract
Plasmonic sensing is a vibrant field where the optical properties of surface plasmons are exploited to create analytical sensors for biomedical, environmental and food safety applications, among others. Upon irradiation of light on a plasmon-active nanomaterial, the enhancement of the electric field leads to augmented scattering, absorption and luminescence of molecules in, respectively, surface-enhanced Raman scattering (SERS), surface-enhanced infrared absorption (SEIRA) and metal-enhanced fluorescence (MEF) and to highly sensitive refractometric sensors with surface plasmon resonance (SPR) and localised surface plasmon resonance (LSPR). The advent of a new generation of artificial intelligence (AI) and machine learning (ML) tools provides an opportunity to further advance the design, synthesis and characterisation of plasmonic materials, improve signal processing and image analysis in plasmonic sensing experiments and to design sensors with better sensitivity, selectivity and robustness. The review will first build basic knowledge in plasmonic sensing and AI/ML, before discussing opportunities for AI/ML-augmented sensor design and data analysis, and then discuss applications where AI/ML provided added benefits in plasmonic sensing. The review will conclude with a perspective on where the field is trending.
1 Introduction
The field of plasmonics encompasses a range of optical techniques that manipulate light with optical nanoantennas, giving rise to a suite of phenomena that can catalyse reactions,1,2 enable photonic devices,3,4 and sense molecules.5 The excitation of surface plasmons in nanostructures enhances the local field and probes the dielectric environment in a nanothin layer of solution in proximity to the plasmonic material. These attributes give rise to plasmonic sensing, where the higher local field enhances vibrational spectroscopies (surface-enhanced Raman scattering (SERS) and surface enhanced infrared absorption (SEIRA)) and luminescence (metal or plasmon enhanced fluorescence (MEF)), while the ability to probe the dielectric environment leads to surface plasmon resonance (SPR) and localised surface plasmon resonance (LSPR).6 These techniques are highly sensitive, achieving detection down to single molecules in SERS,7,8 plasmon-enhanced fluorescence,9 and refractometric (and colorimetric) sensing with plasmons.9 Plasmonic nanostructures can be actively controlled and actuated to accomplish different tasks, including sensing10 and the trapping of analytes or biological entities with plasmonic tweezers.11Plasmonic sensing is closely linked to the development of nanomaterials with tailored properties, enabling the generation of strong electrical fields for enhanced vibrational and luminescence spectroscopies or high sensitivity to dielectric changes for refractometric sensors. These capabilities are largely modulated by the elemental composition, shape and structure of the plasmonic material.12 Classical SPR sensors can be designed on thin gold films13,14 or using LSPR on nanomaterials,15,16 which are also used for surface-enhanced spectroscopies. While historically nanoplasmonic sensors have been mostly created from gold and silver (nano)structures,17–19 there has been increased research into the generation of 2D materials,20,21 hybrid materials made of transition metal dichalcogenides22 and even composites with metal–organic frameworks.23 Plasmonic structures can be nanoengineered to be chiral,24,25 offering new opportunities in chiral molecule sensing.
While not exclusive, plasmonic sensing mainly relies on the principles of SPR/LSPR refractometric sensing13,15,26 or on SERS27–29 and MEF.30,31 These sensing principles are quite universal, as molecules can either be directly detected from their refractive index, vibrational or luminescence signature or indirectly by engineering plasmonic nanotags active in SPR/LSPR, SERS or MEF. Hence, plasmonic sensors have been developed for a broad range of applications,32–35 such as in clinical/health monitoring,36–38 neuroscience,39 environmental monitoring,40,41 and food monitoring, including within platforms conducive for field deployment.42,43 Specific applications can be improved through the use of optical fibres44,45 or microscopy46 and with sensing schemes involving particle etching/growth for colorimetric detection,47 sensor arrays for pattern identification,48 or analyte manipulation in conjunction with hybrid materials.48 These approaches highlight only a small subset of the diverse strategies currently being explored in plasmonic sensing.
Central to this review, artificial intelligence (AI) and machine learning (ML) play an increasing role in the discovery and engineering of photonic materials and in the optimisation of photonic devices.49 ML algorithms are especially useful to extract correlations in complex, multidimensional, and large datasets, where classical chemometric approaches fail, with implications for the synthesis of nanomaterials50 and for data processing.51–54 ML tools can also provide augmented capabilities with plasmonic property prediction and inverse design of plasmonic devices.55,56 Applications of ML in plasmonic sensing are vast, with many examples in biomedical,57–59 environmental58 and food safety applications.58
2 Principles of plasmonic sensing
Plasmonic materials originate from the concept of the plasmon wave.62 Collective movement of free electrons in a material can be quantised in a manner analogous to how light is quantised into photons. In this context, a plasmon represents a collective oscillation of the electron charge density relative to the positively charged ionic core within a metal. While plasmons are possible in bulk, the propagation or localisation of a plasmon at the metal-dielectric interface confers interesting sensing properties. Specifically, plasmonic sensing exploits the strong optical response of nanostructured and nanothin materials when interacting with light. The interaction generates stronger electromagnetic fields that enhance detection sensitivity and plasmonic sensors can be optimised through careful substrate design and integration with analytical methodologies. To understand how AI can be incorporated in plasmonics, it is necessary to grasp the fundamental physical phenomena of surface plasmons and surface plasmon resonances, and its declination in many plasmonic techniques. Discussion on the theoretical aspects of plasmonics is brief, so readers are directed to the literature for more comprehensive reviews.26,63,64Specifically, incident light excites collective oscillations of conduction electrons, generating surface plasmon (SP) modes that propagate in a nanothin film of macroscopic dimensions or are localised in a nanomaterial. This excitation leads to an exponential decay of the surface plasmon along the interface and in the metal and dielectric materials,65 confining the plasmonic wave in a small volume near the interface. The resonance conditions of these SP modes are governed by the dielectric constants of the dielectric (εs) and metal (εm). This dependence on dielectric constants is particularly important for sensing applications as, the sensor response to changes in εs is the core principle of transduction for refractive index sensitivity in SPR sensing.
| εm = ε′ + iε″ | (1) |
2.1 Surface plasmon resonance (SPR)
A nanothin and flat metal surface can support a propagating SP.74 In this configuration, light enters resonance with the SP and this wave propagates along the metal-dielectric interface. The challenge in exciting propagating SPs arises from a wavevector mismatch from incident light (k0) and SP (kSP), prohibiting direct illumination of SP.75,76 Optical coupling configurations, such as prism-based approaches (the Kretschmann configuration)77–79 or the use of gratings,80–83 makes it possible to tune the k0 (eqn (2)) to match the kSP (eqn (3)). When this occurs, light enters in resonance with the SP and a dip in the reflectance spectrum is observed at the same wavelength as the plasmon. The wavelength or incidence angle of the plasmon resonance shifts with changes in dielectric constant of a thin film of solution at the metal-dielectric interface and leads to high sensitivity of surface binding.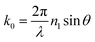 | (2) |
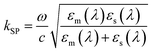 | (3) |
Different instrumental configurations are used in SPR; the main one uses monochromatic light to monitor the change in the SPR angle, while polychromatic light can also be used at a fixed excitation wavelength.79 Each scheme has benefits in terms of sensitivity, user-friendliness, and portability of sensors, necessary for the development of point-of-care (POC) devices. SPR imaging (SPRi) is also possible and builds upon these principles by recording spatially resolved reflectivity data, producing two-dimensional maps of surface binding events in real time.88 Sensor miniaturisation can be achieved on optical fibers.78,80,89,90 Common to these diverse configurations is the generation of large, multidimensional datasets with information on spatial, spectral and temporal domains. The complexity of these data, as well as configurational design, motivates the integration of AI and ML approaches to enhance signal interpretation, automate pattern recognition and accelerate discovery in plasmonic sensing (Fig. 1).
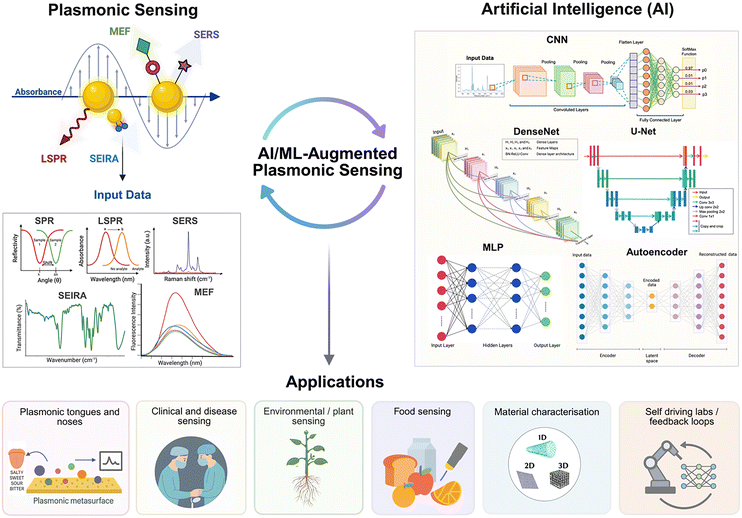 | ||
| Fig. 1 Overview of the field of AI for plasmonic sensing. Plasmonic sensing regroups a suite of techniques benefitting from the interaction of light with nanomaterials. AI and ML provide new approaches to design the plasmonic nanomaterials, to reshape how we conduct research with self-driving laboratories and improve data interpretation for a series of applications. SERS graph adapted with permission from ref. 60 from American Chemical Society, copyright 2024. Material Characterisation image is adapted with permission from ref. 61; licensed under CC-BY 4.0. Figure created in BioRender. | ||
2.2 Localised surface plasmon resonance (LSPR)
In some plasmonic materials the plasmon is not able to propagate; plasmonic nanoparticles smaller than the wavelength of light spatially confine the resonance at the nanoscopic scale.91,92 This LSPR produces a quasi-static electric field strongly confined near the nanoparticle's (NP) surface, typically within a range of 10–40 nm.91,93 This phenomenon is commonly illustrated using metallic NPs (Fig. 2) where the electric field bidirectionally oscillates in response to the incident light.94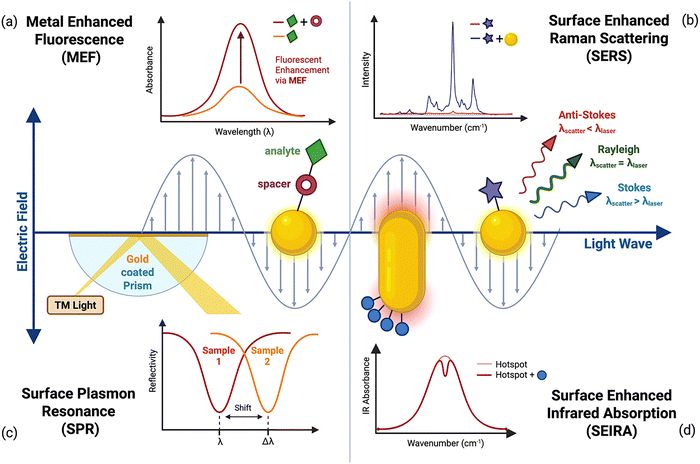 | ||
| Fig. 2 Schematic representation of plasmon-enhanced processes. Light excites surface plasmon resonances, which increases electron density at the metal surface and thereby enhances analytical signal intensity. (a) MEF relies on a spacer between the analyte and the metal surface to minimise quenching effects while enabling fluorescence enhancement. (b) SERS enhances Raman scattering when analytes are bound to or positioned near a plasmonic surface, where the light wave polarises the electron cloud. Graph adapted with permission from Hojjat Jodaylami et al.60 from American Chemical Society, copyright 2024. (c) SPR is excited by transverse magnetic light in a prism, where shifts in the reflected wavelength provide information on analyte binding. (d) SEIRA uses plasmonic nanostructures (here, nanorod arrays) to generate hotspots that amplify IR absorption signals, with decreases in absorption indicating analyte presence. Figure created in BioRender. | ||
The resulting optical response of the nanoparticles is expressed as an extinction coefficient, governed by geometrical parameters of the NP and dielectric properties (eqn (4)).63
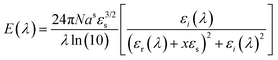 | (4) |
The design of plasmonic substrates for LSPR applications is central to enhancing performance. These techniques exploit electromagnetic field enhancements at “hot spots” regions of intensified local fields typically found at sharp tips, edges, poles, or interparticle junctions of metallic nanostructures. Precise control over nanoparticle morphology, composition, and spatial arrangement is essential for tailoring the optical response and thereby maximising signal amplification. According to eqn (4) the key factors influencing the LSPR of nanoparticles include: (i) material composition, (ii) local dielectric environment, and (iii) nanostructure morphology.
As the design of plasmonic substrates grows more complex, especially in surface-enhanced spectroscopies like SERS and SEIRA, traditional electromagnetic models such as Mie theory and Maxwell-based solvers become increasingly limited. These approaches often assume idealised conditions and quickly become computationally prohibitive for intricate or non-traditional structures. AI offers a powerful alternative. By navigating high-dimensional design spaces and identifying structure–property relationships, AI can accelerate the discovery and optimisation of LSPR substrates beyond what is feasible in conventional wet-lab or simulation-based workflows. From both analytical and physical chemistry perspectives, AI can serve as a surrogate model to compress computationally expensive simulations. Furthermore, inverse design strategies allow researchers to specify desired optical outcomes and have AI predict the corresponding material parameters, even in regions where physics-based understanding is incomplete. The integration of AI and plasmonics opens a new frontier: not only improving sensing platforms but also transforming how we approach the design and understanding of optical nanomaterials.
2.3. Surface-enhanced spectroscopies
The high electric field generated by the excitation of LSPR has led to a range of different plasmon-enhanced spectroscopies capable of detecting trace amounts of analytes with high sensitivity and specificity. For plasmonic molecular sensing applications, LSPR, SERS, SEIRA and MEF sensors have emerged, all exploiting near-field plasmonic enhancement to amplify otherwise weak spectroscopic signals. While these approaches differ in their physical readouts and the type of spectral information obtainable, they are distinguished by their reliance on near-field effects. A generalised comparative representation of these plasmon-enhanced processes is found in Fig. 2. Related strategies, including colorimetric LSPR or waveguide-based devices, generally rely on bulk optical changes rather than local field amplification and do not always require advanced ML for analysis. This section focuses on near-field-enhanced plasmonic methodologies, while sensors whose readout mechanisms fall outside this paradigm are not discussed here.Optimal enhancement typically occurs when the LSPR overlaps both the excitation and scattered Raman wavelengths, although strong SERS signals can still be observed outside this condition. Precisely adjusting the plasmon wavelength of nanoparticles for optimal enhancement could be accomplished through the use of self-driving labs (see Section 6.6). Another major source of enhancement is the presence of hot spots, nanoscale regions of intense field confinement, usually formed when interparticle gaps are less than 10 nm.126 These hot spots occupy only a small fraction of the surface but can dominate the SERS response (accounting for over 70% of overall SERS intensity)127 and are critical for single-molecule, digital and dynamic sensing applications reflected in the rich research environment around them.128–131 Nonetheless, plasmonic surfaces lacking hot spots can still provide measurable enhancement.132 The wide tuneability of LSPR, combined with engineer-able hot spots, makes SERS a uniquely powerful technique. AI approaches offer promising tools for optimising nanostructure design and predicting enhancement patterns, particularly when tailored specifically to SERS-relevant parameters such as hot spot geometry, distribution, and spectral overlap.
The resulting SERS spectra are often complicated with background signals and baseline variations that can complicate interpretation. A relatively standardised procedure exists for processing spectra,133 although specific algorithms and their sequences vary depending on the instruments and methodology used. Typical steps include x-axis calibration, cosmic ray removal, baseline subtraction, smoothing, normalisation and outlier removal. These are commonly implemented using well-known techniques such as asymmetric least-squares for baseline removal and Savitzky–Golay filtering for spectral smoothing. However, despite being a critical stage in the analysis, preprocessing is treated as routine but is susceptible to error and inconsistency. Each data processing step introduces variability, and manual tuning can introduce problems with users introducing bias. AI algorithms could bring significant benefits to this stage, automating parameter selection, reducing user bias, and improving spectral quality across acquisition conditions. This has been explored in related fields like mass spectrometry and proteomics134 and will be discussed in more detail in the review.
Two different types of SERS sensors are prevalent in the literature: indirect SERS, where Raman reporters are used to give strong, well-defined Raman signals and direct SERS measuring the analyte's Raman response. Raman reporters are highly polarisable molecules with a strong, characteristic Raman signal, normally with at least one aromatic ring and a thiol to bind to gold surfaces.135 Indirect SERS is commonly used in biosensing and for targeted detection, given low concentration detection limits from the high SERS response of the Raman dye on the NP. Direct SERS is label-free where the SERS surface is (generally) unfunctionalised (or functionalised with molecules with a small Raman cross-section) so that analytes can be detected without an added label via their fingerprint spectra. This type of sensor is most useful for the profiling, identification and classification of complex mixtures of analytes.
Historically, these complex SERS spectra have been analysed with basic peak-by-peak analysis and simple ML statistical analysis techniques such as principal component analysis (PCA), linear discriminant analysis (LDA), and other tools from the chemometrics toolbox. A recent review highlighted the potential for using AI, particularly in chemometric analysis of SERS data.136 AI is already being routinely used for classification, regression, spectral deconvolution and real-time diagnostics. While also highlighting these classical algorithms, this review aims to dive deeper into ML and focus on the possibilities of deeper learning algorithms for datasets becoming increasingly complex and large, needed to fully realise the potential of SERS in areas from environmental to biological to food sciences.
SERS enhancement can also benefit from combination with electrochemical (EC) methods (EC-SERS). This technique is rooted in the discovery of SERS on a roughened electrode, where Raman enhancement was observed.137,138 EC-SERS enables the quantitative performance of EC to be exploited while ensuring the identification of the analyte by SERS.139 Moreover, polarising the EC-SERS electrode can attract or repel molecules, which can contribute to selective SERS detection or favour the charge transfer mechanism. This attribute can be leveraged to detect multiple analytes in a complex sample,140 useful for identifying drugs,141,142 biomarkers for disease diagnostics,143–146 environmental contaminants,147 and for studying the mechanism of surface-active processes.148,149 EC can be exploited for desorbing analytes from the enhancing surface, thereby reducing fouling and providing a method to regenerate the surface.150 Though research to incorporate AI so far has been limited to regression and classification for analyte quantification,145,146,151 utilising ML can enhance the performance of EC-SERS sensors. AI represents an opportunity for time-resolved EC-SERS, where one interprets large time-correlated EC-SERS datasets.152 Alternatively, EC-SERS protocols could be optimised into an automation loop to define the best sequence of EC and SERS. AI assisted EC-SERS could facilitate analyte identification and quantification.
As with SERS, AI could play a transformative role in advancing SEIRA and MEF technologies. While substrate design stands out as a clear area of opportunity, the broader sensing performance, ranging from signal optimisation to multiplexing, may also benefit from the thoughtful application of AI. Though still largely unexplored, this intersection presents an exciting frontier for improving the accessibility, reproducibility, and overall impact of plasmon-enhanced spectroscopies.
3 Principles of AI
The potential of AI lies in its ability to uncover subtle and sometimes unexpected connections in large datasets. AI is therefore valuable for interpolating, classifying and predicting trends in complex data. However, AI is agnostic to scientific concepts, and its output depends strongly on how it is trained and applied. Different models ranging from linear regression to deep neural networks (NN) are suited for different tasks. One limitation is that none of the models can independently assess the quality and the validity of their results, making the user essential in the validation process.178 Effective use of AI, thus requires understanding of the working principles, strengths and limitations to ensure robust, interpretable and scientifically grounded outcomes.In plasmonic sensing, AI can support research at multiple levels with tools that bridge understanding of nanoscale structure, optical response, and analytical performance. A clear understanding of the key concepts of AI and how they can be applied to regression, classification, clustering, prediction and process control or optimisation is crucial for useful incorporation. The preceding terms refer to what are called the tasks, meaning the process that the user wishes to automate. In the next section, selected tasks (regression, classification, clustering, and intelligent control) and the typical models used to perform those tasks will be succinctly introduced.
3.1 AI tasks and models
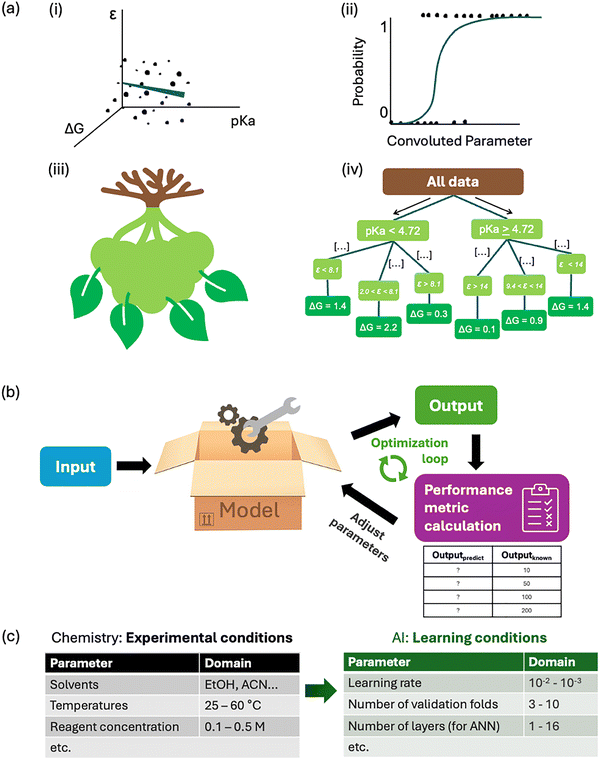
Regression tasks fit a model to describe the dependence between variables to predict, interpret or quantify your results. It correlates a numerical value from an input,179 modelled by a function (the estimator), which, in turn, is obtained by fitting the n-dimensional training data. Multivariate analysis can be seen as the most accessible example of ML.180 More complex models include multiple linear regression, polynomial regression, and regression trees, among others. The regression models fit the n-dimensional training data to a n-term linear function, or a polynomial of chosen degree, respectively. “Tree” models separate the training data in “branches”, where each branch divides a dimension in at least two segments. During the training process, these segments are optimised so that starting from the top of the tree, each branch discriminates the most data points, down to the “leaves” which contain a handful of examples. The output of a regression tree is usually an interpolation of the two leaves closest to where the point to estimate sits. In plasmonic sensing, regression tasks are useful in calibrating complex analytical signals and extracting the most important features such as the most dominant vibrational bands. Visual representations of standard regression models can be found in Fig. 3a.Categorisation of data is important, particularly in high-dimensional data. Classification tasks is one of them, aiming at identifying which category a data point belongs to. This approach is especially useful for categorising, for example, does this food contain bacteria X. Due to the discrete nature of categories, each class is encoded as a value or vector. The most common type in multi-class classification is the one-hot vector, where the category to which an example belongs to is denoted by a ‘1’ in the vector. It follows that the vector resembles, for example: [0, 1, 0, 0, 0] if the data point belongs to the second of five categories. When many categories exist in the dataset, target encoding can replace this vector with a single value, determined by the statistical impact it has on the output. Algorithms for multi-class classification include k-Nearest neighbours (kNN), decision trees, gradient boosting, and NN. In kNN, each data point is compared to the k points that are the closest to its multidimensional position, that is, its position in “feature space”. The category most prominent in the set of the k closest neighbours will determine the category predicted for the new data point.181 Decision trees and random forests (RF) separate data points based on successive quantitative discrimination criteria, as described above, from branches to leaves, where each leaf is a category. The gradient boosting technique combines a succession of tree- or forest-based algorithms to achieve better results. Finally, multi-label classification can account for the case where data points belong in more than one category, and are adaptations based on RF, decision trees, and gradient boosting.
Clustering algorithms aim to regroup similar data points together, without prior knowledge about the data, a process termed unsupervised algorithms, rather than classification tasks which are supervised. In this case, what emerges is sets of points sharing commonality in the input data. The clusters obtained have a more abstract meaning but do distinguish separate groups of points.182 The widely employed k-means algorithm places k “means” in the feature space, then partitions the space in k clusters around these means. A centroid is calculated from the data within each cluster to become the new “mean” of each cluster. This 2-step loop is repeated until the final clusters are found, with the most optimal cluster partitions. Another family of methods uses the density of points in feature space, also called packing, to group points together. The number of clusters does not need to be specified, unlike k-means clustering. Density Based Spatial Clustering of Applications with Noise (DBSCAN) is one of the only methods in this family used in the wider field of chemistry.183 They work best in datasets with well-separated groups, where a “drop in density”, or sparse boundary region exists around the clusters. Clustering tasks can identify to which general category the sensor output belongs, for example, does the patient sample have cancer or not?
The last group of AI methods is intelligent control, where a set of parameters is tweaked in a loop to explore the feature landscape and optimise a certain output value (Fig. 3b). While it is seldom applied directly in analytical methods,184,185 control theory approaches can be employed to assist in developing plasmonic sensing platforms,186 whether while connected to a reactor in real time, or used in between discrete experiments to steer development. In a typical experiment, one or more values taken from on-line analysis of a chemical species act as parameters for a model, which then calculates the next system inputs with the objective of optimising some characteristics of the product, e.g. the LSPR position. Intelligent control methods can be related to, and specific cases of, regression models that have been adapted to the practical reality of scarce data, especially at the beginning of an experiment. One example to have been applied specifically to a problem related to plasmonic sensing is the Stable Noisy Optimisation by Branch and Fit (SNOBFIT)187 used with continuous flow nanoparticle synthesis to optimise the reaction conditions.188 This topic will be discussed in detail in Section 6.6.
3.2 Training an AI model
Generally, AI models are trained by iteratively optimising their parameters until the training input gives a desired output. Performance metrics must, therefore, be utilised to steer the parameter adjustment at every training step, called epoch. Loss functions are performance metrics corresponding to the error of the model.181 Thus, when they are minimised, the model is said to have converged, and its training stops. One important issue in model training is, therefore, ensuring that the loss function arrives at the global minimum, as opposed to one of many possible local minima.For regression tasks, variants of the squared error such as mean squared error (MSE) and root mean squared error (RMSE) can be expected to provide satisfactory results in simple datasets. For classification, binary cross-entropy is a calculation of the uncertainty of the categorical identification based on a sigmoid fit of the distribution. These are simple, universal examples of loss functions, but each case should be assessed and the literature searched to find the most appropriate loss function for the problem and model at hand. As such, the dataset size required for these models can be small to moderate. With larger scale and more complex and high-dimensional datasets, these more classical models can struggle, so deep learning (DL) models such as NNs are often preferred because they can automatically learn complex non-linear relationships and hierarchical features directly from the data.
Artificial neural networks (ANNs) can be built for a large diversity of tasks and adapted to many types of datasets.189 However, they are known to be more computationally intensive, as well as require more time and effort to develop and optimise. They are also data hungry and must be developed only when large datasets are available. The benefit gained in the assembly of successive layers of neurons, through which information passes, is it allows for the emergence of rich mathematical connections, and therefore, more subtle traits of the dataset can be picked up on by the model. Central to NNs is the artificial neuron, a simple mathematical unit with modest capabilities called a “Processing Element”. It is typically composed of a weight affecting each input connected to itself, and an activation function, which regulates the net output of the neuron.190 Various activation functions can be used, and choosing the most suited one is a part of model development. Training of a NN requires the adjustment of all the weights and activation functions of all neurons involved. With many neurons and input features, this process can become extremely intensive. In the right conditions, the model converges after “learning” what the data can “teach” it. Due to ANNs being a collection of individual units, these units can be arranged in an infinitesimal number of ways; each layer can have a different number of parallel neurons (affecting the network's “width”), and the number of layers can be altered (this is the network's “depth”). DL refers to training a NN with more than two hidden layers. ANN topology development is a discipline of its own and an active field of research in computer science. For chemists, it is recommended to search for and imitate successful architectures, proven for problems like the one at hand.
Just as a chemical experiment is conducted in the laboratory with varied parameters to find the method best suited to elucidate a question, experiments are conducted in AI model training to find the settings best suited to give a proper result. In chemistry, there are theoretical parameters which will set the hypothesis boundaries and experimental parameters which will impact the conditions in which the data is collected. In an AI experiment, theoretical parameters will be set with the possible performance boundaries of the model's output, and empirical parameters, such as the number of centroids or the type of activation function, are related to the AI experiment conditions. These experimental parameters are not to be confused with a model's parameters - the individual weights and values that compose it.
A subset of AI empirical parameters are hyper-parameters. These numerical values and non-numerical settings will influence the behaviour and capacity (ability to express complex relationships) of the model. For instance, most ML and AI models have one called ‘learning rate’, which controls the degree of modification of the model's parameters for a single training step.191 In general, smaller learning rates will prolong model training but help with loss function convergence. Adaptative learning rates are sometimes available, depending on the model type and specific software package. Hyperparameters are routinely tested over a wide range of value combinations for each one in a process called tuning. Computer languages are often equipped to automatically sample given ranges of these hyperparameters to find the best ones. The more parameters, the more capacity a parametric model can develop. Non-parametric models have a capacity which is independent of the number of parameters, but that still varies with the model's functionality, sometimes called representational capacity. Examples of this in comparison to chemistry experimental conditions are explored in Fig. 3c.
Just as important as the choice of model is its validation. Like a method in analytical chemistry must be validated for a quantitative application, an AI model must be validated for a chemical application. The process generally involves: (1) visualisation of the training data;192 (2) definition of the experiment's conditions;193 (3) choosing the performance metric;179 (4) cross-validation;194 (5) calculation of the model's accuracy and robustness;179 (6) optimisation of the model's complexity to navigate the bias-variance landscape.
In AI, the bias of a model is the difference between the estimator, the global function representing the model as a whole, and the true unknowable function which exists between the variables of the dataset. Bias decreases as the complexity of the model increases because it can then capture more of the dataset's intricacies. On the other hand, increasing the model's complexity will also increase its variance on real data because the added variables will create a noisier estimator function. The situation of capturing too much noise is called overtraining, where the noise is used to create the output. The opposite situation, undertraining, is when the model's complexity is insufficient to properly capture the relationship linking the features. The optimal complexity should exist somewhere in the middle and must be found to have a good model.195 Time, effort, trials, and validation are the only remedy.
4 Sensor design
The design, synthesis, and performance optimisation of plasmonic sensors still largely depend on trial-and-error experiments and the intuition of the experimentalist. Integrating AI into sensor development design loop marks a shift from using AI solely for exploratory data analysis to employing it as a generative tool for discovery.196 While expertise remains essential in leading and curating high-quality experimental set ups, we can now rely on AI systems to rapidly identify patterns within complex datasets during optimisation to converge towards desired design methodologies.197,198Plasmonic sensors detect chemical and biological analytes with high sensitivity via surface interactions, leveraging mechanisms such as LSPR, refractive index shifts, SERS, and colorimetric changes.76,137,199 Their performance depends on finely tuned nanostructured metallic nanoparticles, patterned films, or hybrid composites whose geometry, composition, and arrangement influence optical and sensor responses.91 For example, optimising nanoparticle spacing and shape can dramatically enhance SERS signals, while tuning film thickness and the surrounding dielectric environment modulate refractive index sensitivity.26,137 Therefore, designing plasmonic nanosensors constitutes a complex, multi-objective optimisation problem, bridging fabrication parameters and physical characteristics of materials for an optimised sensing performance.56 This complexity makes the design space a challenge to navigate manually and sets up plasmonic sensors as an ideal candidate for emerging AI-driven approaches.
In plasmonic sensor development, predictive models are leveraged for their forward mapping from structure to sensor response, supporting efficient exploration of the design space.202 They implement AI in the experimental process for the prediction of how changes in experimental conditions impact the sensor response. Soft computational approaches such as NNs have been trained with NP fabrication variables such as alloy composition, annealing time, and processing conditions to predict resulting surface geometry and plasmon resonances, enabling prediction of Ag/Au layering to shifting LSPR peaks.203 Predictive models offer a path for evaluating design spaces in a more precise and forward-thinking way by linking multiparameter design spaces to material properties. A shortcoming of this approach is that experimental datasets are often small and noisy, thus simulated data can be incorporated into the training dataset, where appropriate, to improve the training of these data-hungry prediction algorithms. However, using simulated data in ML models poses a risk of poor transferability as the simulations must accurately capture the features of experimental data. Otherwise, it introduces bias and can lead to poor performance of the model. As such, the preferred route is always training the models on experimental data when possible. Including physical models in the training of the ML model can alleviate some of these challenges by considering the underlying physics of the modelled process.
Numerical simulations play a critical role in linking material properties to tuneable optical responses. By modelling the effects of shape, size, and composition on light–matter interactions, simulations facilitate the design of plasmonic structures for targeted sensing applications. For instance, DL models trained on boundary element method simulations have been employed to predict scattering spectra according to nanoparticle geometries.204 Commonly used numerical methods include the finite element method (FEM) and finite-difference time-domain (FDTD), which compute optical properties and electromagnetic (EM) field distributions across complex nanostructures.205 However, the practical application in sensor design lies in integrating FEM and FDTD simulation data on local field distributions into predictive models, enabling well-informed decisions and linking desired field distributions to optimise plasmonic sensor performance.
However, the design space for real-time optimisation of plasmonic sensors introduces an additional layer of complexity: predictive modelling becomes a continually adaptive process of discovery. Inverse-design strategies provide the foundation for this adaptivity, integrating AI-driven predictive models directly into sensor optimisation strategies (Fig. 4(i)). In this context, AI seeks to infer the material properties or synthesis pathways required to achieve a target sensor performance. Predictive, or surrogate, models approximate the forward mapping of structure to response, enabling efficient exploration of the design space.202 As such, inverse design constitutes a broader methodological framework, encompassing predictive modelling, optimisation algorithms, and experimental validation to navigate high-dimensional design spaces. In practice, plasmonic material choice is typically constrained a priori by chemical stability, operating wavelength, and biocompatibility; accordingly, AI-driven optimisation is most often applied to geometric, structural, and fabrication parameters rather than unrestricted material classification. Among the applications outlined below, the most transformative prospects lie in using AI to optimise material properties, identify synthesis strategies, and engineer plasmonic sensors for desired performance metrics. Fundamentally, these are inverse design problems, and they offer a powerful framework for the future of sensor development.
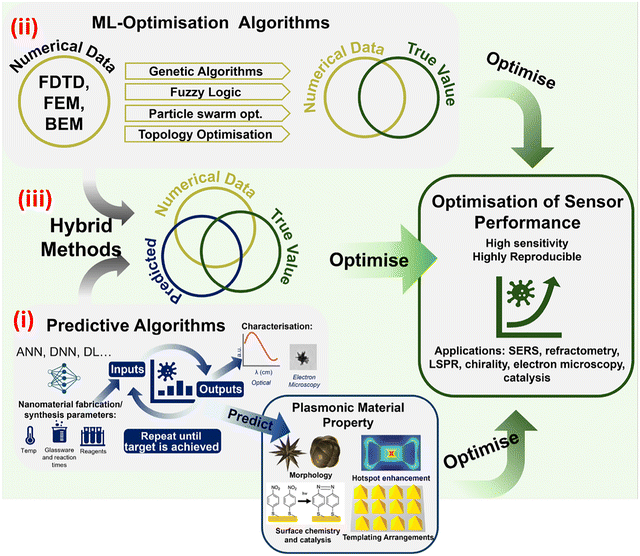 | ||
| Fig. 4 This scheme represents three approaches used for the inverse design of plasmonic sensors. In the first approach, (i) predictive algorithms map fabrication parameters to plasmonic material properties, including optical response, field enhancement, geometry, morphology, and surface interactions, enabling forward prediction and performance-guided sensor design. The second approach uses (ii) ML-based optimisation trained on numerical simulations to improve prediction accuracy and support off-bench optimisation of sensor performance. The third approach represents (iii) hybrid methods which combine predictive modelling with ML-optimisation, integrating numerical, experimental, and model-generated data into a unified framework that enhances the reliability and efficiency of inverse-designed plasmonic sensors. Morphology, surface chemistry and catalysis, and templating arrangements figures in plasmonic material property adapted with permission from ref. 200 figures licensed under AuthorChoice via CC-BY 4.0. Hotspot enhancement image in plasmonic material property adapted with permission from ref. 201 from American Chemical Society, copyright 2010. | ||
Modern inverse design of plasmonic sensors increasingly relies on hybrid frameworks with embedded optimisation loops for rapid convergence of predictive algorithms to achieve targeted performance metrics, a hallmark of inverse design. Firstly, softer modelling, such as regression modelling, has been implemented to predict optical responses of nanomaterials, for inverse morphology design of nanocylinders.206 In the case of more complex tasks, deep neural networks (DNN) have been incorporated to learn the numerical analysis of signal amplification and to predict near-field enhancement distribution across various NP geometries, enabling rapid ML analysis for comparison and design of material with specific EM field confinement.207
DNN models serve as efficient surrogates for full FDTD simulations, enabling accelerated prediction of far- and near-field optical behaviours of plasmonic NPs. This approach facilitates the rapid design and optimisation of electromagnetic field effects,208 and the inverse design of gold nanostructures with targeted transmittance profiles and LSPR positions due to the ML-informed decisions provided during ML led optimisation.209 A key step in simulation-driven design is verifying that optimised structures are experimentally feasible and stable. ML does not provide more data, but it distils existing information into more concise, high-value analyses and recommends the next experimental or design step needed to reach a target outcome. In plasmonic simulations, designs with high theoretical performance may prove impractical due to fabrication or material limitations. ML models trained on simulation data further enhance these efforts by providing strong predictive power across large, multidimensional design spaces.
As the design-space becomes increasingly complex, optimisation algorithms play a central role in navigating high-dimensional parameter landscapes to achieve target optical or sensing outcomes. Many examples of the design for highly specialised sensor design have embedded ML-optimisation algorithms in the inverse-design framework. These algorithms, such as genetic algorithms (GAs), particle swarm optimisation (PSO), and topology optimisation, are used to refine structural designs through experimental parameters, aiming to achieve optimised performance metrics derived from surrogate models of experimental and/or numerical simulations.196,210 Further, by integrating surrogates into the optimisation loop, AI can efficiently tailor sensors for specific molecular targets or optical functions, reducing trial-and-error and accelerating discovery.
GAs are implemented in plasmonic sensor design to iteratively refine candidate sensor structures according to fitness metrics derived from simulations and experimental observations.211 However, the slow convergence and susceptibility of GAs to local minima motivate the incorporation of surrogate models to improve search efficiency and enhance design robustness.196 When combined with NNs in a GA-NN framework, this approach enables rapid prediction and optimisation of structural parameters such as metal type, geometry, and layer count, to achieve targeted LSPR responses.212
Further, ANNs have been incorporated in the GA frameworks for selecting ANN-derived parameter combinations to achieve desired absorption peak locations while improving intensity, reproducibility, and Q-factor of Au coating on ZnO nanorods.213 GAs have also been applied to imaging applications, for applications such as localised plasmonic structured illumination microscopy (LPSIM), where they optimise plasmonic nanoantenna array geometries with CNNs trained on reconstruction metrics such as light intensity and noise robustness.214 The addition of GAs and ML-GA into the inverse design loop offers a powerful tool for accelerating the optimisation of plasmonic substrates, enabling rapid identification of plasmonic material parameters that yield targeted sensing responses.
ML-optimisation frameworks may also include numerical simulations for multi-objective design spaces, critical for the development of highly specialised sensing devices. For instance, DL-GA optimisers have been applied to FDTD simulations of Born-Kuhn type nanodimers to optimise their configuration for enhanced sensitivity in chiral analyte detections.215 PSO combined with FDTD simulations has been incorporated to inversely designed palladium metasurfaces for highly sensitive hydrogen detection.216 Further, surface optimisation of nanohole arrays can be achieved by predicting transmission resonances from geometric parameters using FDTD simulations, followed by the inverse design of selective LSPR arrays to enhance nanoscale sensitivity.217 Similarly, when DL is implemented with experimental conditions and FDTD-simulation reflection spectra predicted Ag coloration, inverse design of Ag arrays with precise control over structural colour formations becomes possible.218 Lastly, topology optimisation has been employed to explore and tailor EM fields in plasmonic materials, enabling targeted and maximised field enhancement with simulated data. Impressively, this technique allows for near real-time guided optimisation, which can be applied to electromagnetic localisation and sensing applications.219,220 These studies demonstrate the ability to optimise sensor-specific parameters and highlight the need for numerical simulations to obtain essential data that may not be accessible through conventional experiments.
These examples illustrate that objective-driven optimisation is central to inverse design and particularly is well suited for the multidimensional task of plasmonic sensor development. Currently, there is no standardised workflow for selecting computational or experimental strategies in plasmonic sensor design, though the main concepts discussed in this section are highlighted in Table 2. Soft computation techniques, such as GAs, and ANNs have been implemented; however, trends in recent publications suggest that deeper-learning models may be required to address high-order optimisation tasks. This challenge is twofold: it arises from the complexity of the design objectives in relation to research questions and from limitations in the type and quantity of available data. Optimisation is particularly important when the goal is to tune experimental parameters, enhance analyte sensitivity, or refine measurement protocols; as such, data collection strategies must be tailored to the specific context.
| Design stage | AI/ML approach | Role in the design pipeline | Data used | Optimised sensor parameters | Relevant plasmonic sensing modalities |
|---|---|---|---|---|---|
| Predictive models | Regression models; NNs | Forward prediction of optical and sensing response from structural parameters | Experimental measurements; simulated optical responses | Resonance position, optical response, sensing performance | SPR, LSPR, SERS, SEIRA, MEF |
| Numerical simulation driven design | DNNs | Surrogate modelling of computational EM simulations | FDTD and FEM simulation datasets | Near-field enhancement, scattering and absorption spectra | LSPR, SERS, SEIRA |
| Inverse design | CNN and DNN | Mapping desired optical response to structural design parameters | Simulated and experimental optical field distributions | Nanoantenna geometry, array configuration, signal robustness | LSPR, SERS |
| Optimisation algorithms | GAs (with optional ANN), PSOs and topology optimisation | Global and multi-objective optimisation of plasmonic structures | Fitness functions derived from EM simulations and ML predictions; FDTD simulations | Geometry, spacing, material distribution, metasurface configuration | SPR, LSPR, SERS, SEIRA |
Among the approaches explored, FDTD and FEM simulations offer high-fidelity solutions to augment sparse datasets, particularly in hybrid strategies that combine physics-informed modelling with experimental validation. While simulations are valuable for validation and surrogate training, they introduce additional steps into the sensor design workflow. A practical goal is to develop AI tools that match the predictive power of hybrid simulations and experimental methods but operate effectively on smaller datasets. Achieving this requires surrogate-based optimisation and machine-learning-guided closed-loop feedback between experiments and models (see Section 6.6), suggesting a broader shift toward adaptive, data-efficient optimisation is a critical step toward high-throughput plasmonic sensor design. In summary, a paradigm shift is taking place, as the design of sensors increasingly uses AI to refine the properties of plasmonic materials.
5 Data analysis
5.1 Raw signal processing
As plasmonic sensing techniques, particularly SERS, SPR, and LSPR, continue to advance toward higher sensitivity, miniaturisation, and multiplexing, extracting meaningful information from raw spectral data has become increasingly demanding. This challenge is particularly evident in dynamic modalities such as single-molecule SERS or live-cell SPR, where molecular orientation, binding fluctuations, and nanoscale substrate variations introduce substantial variability.51,57,221 These modalities generate inherently complex, high-dimensional and non-linear optical signals that are often compromised by low signal-to-noise ratios (SNR) due to weak analyte responses, baseline drifts caused by instrumental or environmental fluctuations, broadband fluorescence backgrounds masking vibrational features, and additional variability introduced by thermal effects or nanoscale heterogeneity of the sensing substrates.222–224 This complexity is further amplified by the heterogeneity of raw signal types across plasmonic techniques, ranging from spectral, temporal, or imaging data, each with distinct noise characteristics and structural patterns that require tailored preprocessing strategies.56 Conventional preprocessing pipelines, which typically include calibration, denoising, baseline correction, normalisation, dimensionality reduction, and validation, can mitigate noise but remain fundamentally constrained by linear assumptions and manually tuned parameters. This makes them prone to suppressing diagnostically important weak signals, especially in noisy and multiplexed plasmonic measurements.222,224,225 In contrast, AI methods, particularly DL architectures, offer adaptive and data-driven approaches that can learn noise statistics and signal patterns directly from raw data, enabling robust preprocessing, including denoising, signal extraction and feature recognition, across heterogeneous spectral datasets.226–228 Recent studies illustrate several distinct strategies for raw or near-raw signal processing with AI, which may be roughly grouped into (i) direct classification or regression from raw spectra using deep networks; (ii) integration of denoising or baseline correction modules such as autoencoders or multi-scale CNN architectures into end-to-end pipelines; and (iii) hybrid designs that retain minimal classical preprocessing but AI models are trained to accommodate residual artefacts.The first and fundamental challenge in raw signal processing is denoising. Plasmonic spectra, particularly in SERS and SPR, are often dominated by stochastic noise, detector fluctuations, and photon shot noise that obscure subtle analyte peaks. Classical approaches such as Savitzky–Golay smoothing, Wiener and Fourier filters, wavelet transforms, and polynomial curve fitting have been widely used to suppress random fluctuations and enhance peak visibility. While effective in reducing broad noise patterns, these methods depend heavily on user-defined parameters (e.g., filter size, polynomial order, or wavelet basis), and they often compromise signal fidelity by distorting peak shapes or removing diagnostically important weak vibrational features.51,225 This parameter sensitivity makes them less reliable in heterogeneous plasmonic datasets where the noise characteristics are non-stationary and sample-dependent. Recent advances in DL have introduced more adaptive denoising frameworks that learn directly from examples. CNNs,226,229,230 encoder–decoder models such as autoencoders (AEs),227 multi-scale CNNs like U-Nets,224,231–234 Residual Networks (ResNets),224 peak extraction protocol,235 and generative adversarial networks (GANs),236,237 can learn the statistical structure of noise and spectral peaks directly from raw data, enabling end-to-end reconstruction of clean spectra without manual intervention. Another approach used a noise2noise architecture based on data augmentation and a U-net, where denoising was achieved with an original signal-to-noise as low as 1.12,238,239 while a CNN-based denoising networks achieved up to 10.24 dB improvement in SNR compared to wavelet filtering in Raman spectra.226 Similarly, GAN architectures have demonstrated the ability to reconstruct high-SNR single-cell Raman spectra from short acquisitions, enabling nearly tenfold faster measurements without sacrificing spectral fidelity or identification accuracy.237 In plasmonic microscopy, DL architectures, including hybrid U-Net and residual CNN architectures, have further demonstrated the ability of DL to enhance weak nanoparticle scattering signals by leveraging spatiotemporal correlations, underscoring the adaptability of these approaches across modalities.240
Beyond noise, fluorescence and scattering backgrounds introduce slowly varying baselines that overlap with weak plasmonic peaks, complicating quantitative analysis. Traditional correction techniques include polynomial fitting,241 wavelet decomposition,242 and penalised least-squares approaches such as asymmetric reweighted penalised least squares (arPLS)243 and adaptive smoothness penalised least squares (asPLS).244 While effective for smooth baseline variations, these methods are highly sensitive to the choice of polynomial order, wavelet basis, and smoothing coefficients, which introduces subjectivity and reduces reproducibility, particularly across heterogeneous datasets.222,224 Moreover, they often risk under- or overfitting, either leaving residual baseline artefacts or inadvertently distorting spectral peaks that are critical for accurate quantification. In contrast, DL-based frameworks treat baseline subtraction as a supervised learning problem. Recent advances have introduced transformer-based models,245 triangular deep convolutional networks,223 physics-aware CNNs for parameter prediction,246 and NN-driven iterative curve-fitting approaches,247 each designed specifically for baseline correction. These architectures automatically learn and remove background trends while preserving spectral peak integrity, eliminating the need for manual parameter tuning. More advanced architectures such as cascaded CNN frameworks (ResNet, U-Net)224 and convolutional autoencoders227 have been shown to simultaneously perform denoising and baseline correction in a single inference step, improving reproducibility and throughput in biomedical Raman spectroscopy. Some AI-driven pipelines now bypass baseline correction entirely, enabling direct recognition of raw Raman and SERS spectra.248 Notably, multiscale CNN (MsCNN) has achieved over 97% accuracy in classifying hepatitis B serum spectra without any baseline subtraction,249 underscoring the potential of such architectures to handle background variability as part of their feature extraction process.
Normalisation is another critical step, which is designed to standardise sample-specific scaling effects and enable reliable comparisons across heterogeneous datasets. Classical approaches such as mean centring, orthogonal signal correction, min-max scaling, and z-score standardisation are still common but often limited by sensitivity to outliers or distributional assumptions.250 In practice, normalisation should be carefully tailored to data characteristics and analytical objectives rather than applied as a one-size-fits-all preprocessing step. Recent advances in DL have expanded this toolbox. One-dimensional CNNs (1D-CNNs) can learn modality-specific scaling functions while jointly correcting baselines and extracting features. GANs refine spectra through adversarial training, producing high-fidelity normalised signals at the expense of computational cost. Variational AEs (VAEs) harmonise scaling in latent space while quantifying reconstruction uncertainty, a valuable property for error-aware sensing, although the risk of peak over smoothing persists. By embedding normalisation into learned feature hierarchies, deep models adapt to nonlinearities, substrate heterogeneity, and inter-instrument variability more effectively than handcrafted methods.
Plasmonic signals often evolve dynamically over time because of physical and biological factors, such as analyte binding, fluctuations in local environments, or heating effects from illumination. Traditional approaches rely on baseline subtraction, curve fitting, or peak alignment algorithms to track spectral trajectories, but these often fail when confronted with heterogeneous temporal responses. This challenge is especially pronounced in dynamic sensing modalities, where analyte fluctuations occur on millisecond timescales and generate highly variable peak intensities. ML provides a fundamentally different perspective by treating signal tracking as a temporal modelling problem rather than a static correction problem. Architectures such as CNNs with dilated convolutions, recurrent neural networks (RNNs), long short-term memory (LSTM) units, and neural ordinary differential equation (neural-ODE) models have all been explored to capture temporal correlations in spectral data. These methods outperform classical drift correction by simultaneously modelling both short-term fluctuations and long-term baseline evolution, allowing automated tracking of analyte peaks even when signals are partially obscured. Another important strength of these models lies in their ability to estimate uncertainty in predictions, which helps flag unreliable intervals, a critical advantage for real-time sensing applications in biological and diagnostic sensing, where continuous feedback is essential for accurate interpretation. The integration of domain-adversarial learning has also been proposed to adapt models transferable across substrates and instruments, thereby minimising batch effects that traditionally compromise reproducibility.57,250,251
Dimensionality reduction, signal extraction and feature engineering usually mark the final step of the preprocessing pipeline. Classical statistical techniques provide interpretable axes but often struggle with overlapping peaks and nonlinear correlations. In such cases, deep models like autoencoders, VAEs, and CNN-based embeddings can capture nonlinear and hierarchical patterns more effectively than linear projections. They tend to generate latent spaces that are simultaneously denoised and baseline-corrected, which, in turn, can simplify later analysis. These representations often accelerate downstream classification and regression steps and, at the same time, enhance reproducibility across various instruments. In parallel, feature engineering shifts the focus from purely unsupervised compression toward task-specific descriptor design. Traditional strategies extract peak intensities, ratios, or widths based on chemical knowledge, whereas DL models can perform this step implicitly by learning hierarchical filters that capture both local vibrational markers and broader spectral trends.
DL methods, while computationally more demanding and requiring large datasets for training, offer superior accuracy and adaptability to complex, nonlinear spectral variations by learning features directly from data. In practice, the choice between traditional and DL approaches largely depends on the specific noise characteristics, the amount of available data, and computational resources. In recent years, there has been a gradual move towards more data-driven and adaptive methods for robust spectral analysis.51,224,227 Classical techniques still work well for small-scale, homogeneous datasets, while DL scales better to large, heterogeneous, and high-dimensional plasmonic or spectroscopic data. The ongoing convergence of these approaches suggests that hybrid workflows, where classical normalisation handles the initial correction and DL performs the fine-tuning, seem to offer a practical balance toward reproducible, high-throughput spectral analysis in noisy, heterogeneous plasmonic environments.223,224,252,253 Taken together, these approaches will favour more thorough signal preprocessing to improve data quality.
5.2 AI for classification purposes
Classification is fundamental to the analytical value of plasmonic sensors, mainly for SERS because it transforms complex vibrational fingerprints into meaningful chemical or biological insights that can guide decision-making in diagnostics and sensing. Although SERS offers exceptional sensitivity, even reaching the single-molecule regime,129 raw spectra are often marked by considerable variability arising from fluctuations in hot-spot distributions, variations in molecular orientation, and diverse environmental interferences.130 These factors compromise analytical reproducibility and make simple band assignments inadequate for reliable interpretation.254 To address these challenges, reproducible and automated classification frameworks are needed, ones capable of distinguishing highly similar spectra and extracting reliable information from complex datasets.255 Thus, classification is not merely an auxiliary step but the essential process that elevates SERS from a striking physical phenomenon into a practical and reproducible tool for chemical and biomedical sensing. Traditional chemometric approaches such as PCA, LDA, SVM, and RF have provided useful tools for dimensionality reduction and classification, yet they generally depend on extensive preprocessing, handcrafted feature extraction, and linear separability assumptions that limit robustness when confronted with noisy, overlapping, or heterogeneous spectra.254 These methods also struggle with large-scale, multi-class problems where intra-class variability rivals inter-class differences, leading to reduced robustness and reproducibility. Thus, the limitation is not only accuracy but also scalability and adaptability. For SERS, and more generally plasmonic sensing, to achieve true clinical and analytical impact, classification frameworks must advance toward architectures that are inherently self-adapting, data-driven, and capable of extracting insight from spectral heterogeneity rather than being compromised by it. In particular, AI and ML enhance the selectivity of plasmonic sensors in complex biological or environmental samples by detecting subtle, multivariate spectral patterns, learning characteristic fingerprints of individual analytes, and separating overlapping signals from backgrounds or co-existing species, thereby enabling reliable discrimination even when conventional band-based analysis fails.DL represents a decisive break from classical paradigms. Its significance lies less in marginal accuracy gains over chemometrics and more in its ability to embed preprocessing, feature extraction, and classification into a single end-to-end framework. This integrated design treats spectral variability not as an obstacle but as an informative signal, allowing the models to learn discriminative patterns even from noisy, overlapping, or fluorescence-masked spectra. CNNs have emerged as the leading DL architecture for SERS classification because their structure naturally fits the one-dimensional layout of spectral data. In Raman spectroscopy, chemical information is contained within narrow vibrational bands that must be distinguished from background signals. The convolutional filters in a CNN act as localised detectors, capturing these fine spectral features, while deeper layers integrate them into broader patterns that reflect long-range correlations across the spectrum.256 Unlike PCA-based workflows that rely on predefined dimensionality reduction and linear separation, CNNs can learn hierarchical representations directly from the data: first detecting sharp local peaks, then combining them into characteristic spectral signatures, and ultimately into global class distinctions. Studies have repeatedly shown that CNN-based classifiers outperform conventional PCA-SVM or PCA-LDA on the same SERS datasets, even when trained on minimally pre-processed spectra.257,258 1D-CNNs extend these advantages by analysing spectral sequences directly, eliminating the need for 2D conversions and reducing computational demands. Their ability to preserve subtle spectral details while maintaining high efficiency makes them particularly suited to large-scale or resource-constrained SERS applications, where they deliver reliable performance even under noisy or overlapping conditions.259,260 These types of algorithms also offer opportunities for bidirectional flow between vibrational spectra and molecular representations,261,262 where molecular structure can be predicted from the spectra and vice versa.
While conventional CNNs tend to lose efficiency as their depth increases, ResNets overcome this limitation through identity-based skip connections that allow gradients to flow directly across layers. This design mitigates the vanishing gradient problem and enables the training of much deeper architectures, which is critical for capturing the complex and high-dimensional vibrational information encoded in SERS spectra. By organising convolutional layers into residual blocks, ResNets progressively extract both local spectral patterns and broader correlations across the entire wavenumber range. As a result, they provide enhanced sensitivity to subtle spectral variations that may correspond to chemical or biological differences. Importantly, these architectures not only outperform shallow classifiers on large SERS datasets but also demonstrate strong resilience to experimental variability, including intensity fluctuations, baseline drifts, and hotspot heterogeneity, making them particularly effective for practical diagnostic and sensing applications.255,258
Another critical methodological development in advancing beyond conventional CNNs has been the introduction of transfer learning. Deep architectures, such as CNNs and ResNets, are often constrained by the limited availability of well-annotated spectral datasets, particularly in biomedical and environmental applications, where collecting thousands of spectra across diverse conditions is rarely feasible. Transfer learning mitigates this issue by leveraging models pretrained on large-scale spectral or even image datasets and adapting them to the SERS domain. In these scenarios, pretrained models on extensive Raman or hyperspectral databases can be fine-tuned to classify disease-specific signatures, thereby improving generalisation and reducing the risk of overfitting. The strength of transfer learning lies in its ability to reuse early convolutional layers that already capture universal spectral characteristics, such as peak localisation, noise suppression, and baseline trends, while allowing the deeper layers to specialise in identifying task-specific biomarkers, such as cancerous versus healthy serum, bacterial strains, or environmental contaminants.263,264
Transfer learning has enabled pretrained networks to be adapted for SERS studies with limited data, yet significant challenges remain. DL models are inherently data-intensive, whereas many SERS studies rely on relatively small or imbalanced sample sets. To mitigate this issue, researchers have begun applying data augmentation techniques such as noise injection, spectral shifting or warping, and GAN-based spectrum synthesis to expand training datasets. Oversampling strategies like the Synthetic Minority Oversampling Technique (SMOTE), although common in broader ML applications, have only recently been explored within SERS analyses.265 However, reliance on synthetic data introduces new concerns regarding model overfitting and reproducibility of results across laboratories and instrumentation. In parallel, model interpretability remains a key limitation, with current visualisation tools highlighting spectral regions that contribute most to classification decisions but not fully explaining plasmonic enhancement or analyte-substrate interactions that shape those spectral features.
Beyond data augmentation, a new direction in SERS analytics involves the use of generative modelling. GANs have been employed to synthesise realistic SERS spectra, helping to balance uneven datasets, enhancing model robustness, and reducing overfitting in studies with small or skewed sample distributions. In the DeepATsers framework, GAN-generated spectra expanded a limited dataset of 126 SARS-CoV-2 protein spectra into more than 780 synthetic samples, boosting classification accuracy from roughly 60% to nearly 98%, illustrating how generative models can effectively enrich limited spectral datasets and enhance generalisation.257 More recently, the role of generative modelling has broadened beyond augmentation towards the inverse design of SERS substrates and adaptive sensing, where synthetic spectra and physics-informed NNs are used not only to simulate experimental variability but also to guide material optimisation and data acquisition strategies.266,267
Hybrid models have emerged as an important evolution in SERS classification, designed to combine handcrafted spectral processing with DL to achieve greater robustness in complex datasets. By integrating statistical dimensionality reduction (e.g., PCA, wavelet transforms) with NNs, or combining shallow ML classifiers (SVM, RF, k-NN) with deep architectures, hybrid systems balance interpretability, robustness, and accuracy. For instance, PCA-CNN pipelines have been shown to effectively compress high-dimensional spectral inputs prior to convolutional processing, reducing overfitting and computational cost.253 Other frameworks, such as multi-feature fusion CNNs, integrate raw spectra with engineered features to balance interpretability and predictive performance, achieving diagnostic accuracies exceeding nearly 90–95% as seen in immune disease screening and multi-cancer detection.255,258 Beyond feature-level fusion, recent studies have explored multi-task hybrid networks capable of simultaneous classification, quantification, and anomaly detection, aligning decision boundaries more closely with concentration gradients in SERS datasets.268,269 Collectively, these hybrid strategies represent a practical direction for translating SERS into reliable diagnostic platforms, enabling robust feature extraction from complex spectra while preserving interpretability and generalisability across diverse biomedical and environmental applications.
DL models are often criticised as “black boxes”, achieving remarkable accuracy but offering little explanation of the spectral features that guide their predictions. This lack of interpretability limits clinical translation, where regulatory agencies and physicians require traceable connections between vibrational fingerprints and biochemical meaning to establish trust. This opacity is particularly problematic in plasmonic sensing, such as SERS and SEIRA, where diagnostic decisions must ultimately be justified in terms of molecular vibrational signatures. For such systems, high accuracy is necessary but insufficient; classifiers must also provide interpretable evidence linking their outputs to physically meaningful vibrational bands and realistic biochemical mechanisms. This requirement has catalysed a shift from pure outcome-driven modelling toward mechanistic interpretability, as emphasised in recent AI-SERS reviews and application studies that explicitly frame model transparency as a prerequisite for clinical translation and regulatory acceptance.59,270 Another approach leveraged the combination of RF for feature visualisation and a characteristic amplifier to detect weak signals.271
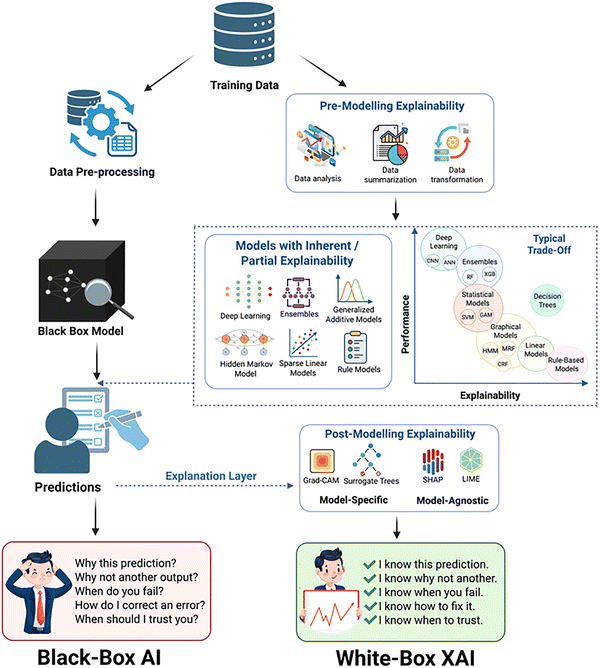
Recent efforts have begun to address this gap by integrating explainable AI (XAI) with SERS. 1D CNNs were combined with gradient-weighted class activation mapping (Grad-CAM) to classify 13 respiratory virus types with over 98% accuracy, while simultaneously highlighting Raman shift regions that consistently governed viral differentiation across solvents and clinical samples.259 Likewise, DL was employed with explainable outputs for Alzheimer's disease diagnosis, validating that the model's discriminative focus aligned with amyloid-β and metabolite peaks central to disease pathology.272 SERS-based models have begun to incorporate other formal explainability tools, including gradient-based methods such as Grad-CAM and Integrated Gradients, attention-based visualisation in convolutional and transformer architectures, and perturbation-driven schemes that probe how spectral components affect model confidence.259,272,273 More recent frameworks extend this toolbox with Local Interpretable Model-agnostic Explanations (LIME) and SHapley Additive exPlanations (SHAP) to analyse the contribution of local spectral neighbourhoods or individual wavenumber features in both classification and quantification settings.274–276 In parallel, logic explained networks (LEN) and related rule-based models have been explored as global surrogates for DL predictions, while methods such as layer-wise relevance propagation (LRP), although not yet widely deployed in SERS, are well established in other domains and illustrate how pixel/feature-level relevance maps could be extended to vibrational spectra.274,275 A comparison between regular black-box AI and explainable AI workflows can be found in Fig. 5. Across these studies, the emerging consensus is that accuracy alone is insufficient to be credible in clinical or regulatory contexts. SERS–AI systems must demonstrate that the spectral features they exploit are chemically plausible, stable across matrices, and robust under realistic experimental variability.59,270,274–276
In several SERS platforms, interpretability is no longer treated as a cosmetic, post-hoc visualisation step but as an analytics tool for interrogating and refining the sensing strategy itself. In SERS classifiers designed for multiplex viral detection, exosome-based cancer stratification, and polymicrobial infection diagnosis, Grad-CAM, Integrated Gradients, attention maps, and related saliency methods are used to assess whether DL models consistently focus on vibrational regions that match known biochemical assignments across substrates, patients, and biological replicates.259,272,273,277 Perturbation-based approaches, including spectral component filtering, feature ablation, and SHAP-derived importance scoring, quantify how removing or rescaling specific Raman bands alters model predictions, thereby distinguishing features that genuinely drive discrimination from those that reflect confounding factors or substrate artefacts.275,276 Importantly, these analyses also expose limitations such as different explainers can yield partially inconsistent maps, and highlighted “important peaks” are not always stable across cohorts or measurement conditions, raising concerns about overfitting to narrow training distributions or hidden confounders.274–276 As a result, interpretability is increasingly viewed as a design constraint that must inform substrate engineering, data acquisition, preprocessing, and architecture selection from the outset, rather than a justification applied at the end. Only through this integration can plasmonic AI systems evolve from high-performing black boxes into trustworthy diagnostic tools capable of supporting mechanistic claims, biomarker hypotheses, and the transparency demanded by clinical regulation.
5.3 Image analysis
In plasmonics, multidimensional images are relatively straightforward to acquire, yet extracting meaningful information from it presents a significant challenge. Beyond conventional 2D and 3D images, each pixel in an image can contain full spectral information, effectively adding up to a fourth dimension or even a fifth dimension (time) that extends beyond a simple binary image. When appropriately analysed, such data can support a wide range of applications, from material characterisation to the interpretation of plasmon-enhanced signals and their deployment in biomedical and other sensing contexts. Plasmonic sensors can be used for imaging where 2D, or even 3D, surfaces are scanned and analysed pixel-by-pixel to resolve spatial variations in signal intensity and distribution. This spatial mapping is key for identifying nanoparticle hotspots, characterising substrate uniformity, and visualising how analytes interact with plasmonic materials. Equally important is temporal data analysis, where spectra are collected as a function of time to capture dynamic behaviour, whether following single-molecule events, monitoring chemical reactions, or probing transient biological processes. Often, basic analytical methods like intensity maps, ratiometric comparisons, and K-means clustering are used, but there is plenty of space in this field for improvements and more in-depth analysis, which could come from the thoughtful incorporation of AI.Plasmonics offers several imaging modalities, such as hyperspectral Raman on or near SERS surfaces, plasmonic scattering imaging (PSI), and surface plasmon resonance imaging (SPRi). Traditionally, these techniques can involve long acquisition times and lower image quality compared to fluorescence imaging. However, their major benefit is the increased information richness that can be obtained from a sample, particularly in label-free and hyperspectral formats. AI can seriously improve two main aspects: image reconstruction and image analysis. Reconstruction enables higher-resolution images from low-resolution data, while AI-driven analysis, including better denoising and dimension reduction algorithms as well as more effective data upscaling, as highlighted above, can be used for better feature detection.
Image reconstruction using AI can improve the spatial resolution of images constructed from low-resolution and sparse data such as those data obtained from hyperspectral Raman (A general workflow for this can be found in Fig. 6(a)). Though this is not a very well-studied area, there are some examples where AI has been used for super-resolution plasmonic imaging, including CNNs being used in plasmonic dark-field microscopy (with U-Net based architecture, Fig. 6(b))278 and CNN in plasmonic terahertz focal-plane array279 or GAN-based models being developed for obtaining plasmonic scattering images (PSI).280 In non-plasmonic fields this is better explored, with a GAN able to reconstruct high-resolution Raman images (Fig. 6(c)).281 Other algorithms have shown good potential for use in plasmonic imaging, including more advanced CNNs,282 content aware image restoration (CARE)283–286 that can drastically improve imaging times. Super-resolution optical fluctuation imaging (SOFI)287,288 and stochastic optical reconstruction microscopy (STORM)280,289 can be combined DL algorithms for blinking experiments such as single particle imaging and digital SERS to get even higher resolutions.290 Additionally, residual dense block (RDB) based algorithms291–294 can be used for capturing deeper features, particularly in feature extraction. Transformer-based models are also becoming more popular295–297 and would be useful for reconstruction from more sparse data due to their long-range pixel interactions.
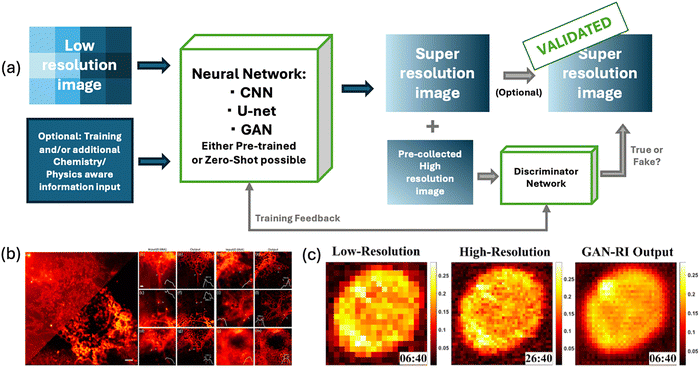 | ||
| Fig. 6 Methods for improving image resolution by image reconstruction using ML. (a) Schematic of potential processes – input low-resolution images to the trained neural network – usually a CNN, U-net or GAN. Additional information on the chemical or physical properties of the image can be input,278 or simply Zero-Shot. A super-resolution image will be output. Image validation can also be carried out – for example with a discriminator network to compare the image against a pre-collected high-resolution image.281 (b) COS-7 cell with regular plasmonic dark field image compared with network processed deep learning assisted plasmonic dark field microscopy (DAPD) image through a CNN. Figure reproduced from ref. 278; figure licensed under CC-BY-NC-ND 4.0. (c) Raman image of a murine osteosarcoma cell processed with a generative adversarial network significantly improves image quality and imaging time compared to higher-resolution Raman imaging. Figure adapted with permission from ref. 281 from the American Chemical Society, copyright 2025. | ||
Another, often less computationally intensive, methodology to improve image quality is the use of denoising algorithms. They can be carried out by spectral smoothing in the preprocessing stages (as described above in Section 5.1), or on the final image.298 Two key improvements in the denoising space have been demonstrated in the Peak2Peak algorithm238 are (i) the introduction of regularisation in the denoising module, which acts as an on/off switch for denoising when balanced against the loss function and (ii) the integration of data augmentation to improve self-supervised feature learning for peak selection from noise (including augmented false-noise examples). Additionally, the algorithm provides visualisation of these stages and can be broadly applied to other black box Zero-Shot Noise-to-Noise (N2N) denoising algorithms, which are particularly useful in this field because they require no pre-training and can be carried out on single images.238,299–301
As well as using AI to improve image quality, it can be used for image segmentation (a computer vision technique to partition images into multiple segments for object detection and medical analysis, etc.) and feature detection and therefore broader image analysis. Many algorithms have been found to be useful for this:302 Mask R-CNN (regional-based convolutional neural network) that outputs an object's class and a mask to efficiently detect objects within an image;303 and U-Net is also well suited for solving image segmentation challenges, instead of the normal pooling operators, up-sampling operators are used to increase resolution of the output.304,305 A particularly strong example of this is AtomAI, useful in electron and scanning probe microscopy data at an atomic level, and thus could be extremely useful in material characterisation tasks, with image-to-spectrum learning tasks possible.306,307 For feature extraction, RDB algorithms can be very good at finding deep connections between certain features and YOLOv8 is a leading algorithm for object detection, outperforming state-of-the-art detection models308 and have been used for detecting SERS nanoparticles and oligomerisation. This algorithm also shows promise in live imaging, nanoparticle-cell interactions and more.309 Also in plasmonics, a CNN was used for the analysis of a lateral flow immunoassay (LFIA) with SERS based on the ResNet-18 architecture,310 a CNN used to reconstruct and then identify analytes in SPRi311 and a CNN combined with an RNN that can predict the optical response of a property of a material from an image.312 Combining different algorithms of different types can improve analysis with great results.313,314
One standout use of AI in image analysis is being able to turn smartphones into portable and affordable imaging technology. One challenge with smartphones is the lower quality of the camera compared to the chilled cameras used in the higher-end systems. Temperature cooling decreases the noise level and improves image quality. Nonetheless, their use has been demonstrated several times so far with environmental315,316 and clinically relevant samples.317,318 In addition, a plasmonic biosensor for detecting pg mL−1 levels of proteins was developed using an algorithm that integrates both GAN and CNN to analyse smartphone camera images.319 Smartphone-based imaging can revolutionise POC devices by making complex data acquisition accessible, but will necessarily need AI to improve image quality and to extract meaningful results.
Beyond spatial imaging, the time dimension is crucial in any live and dynamic monitoring of a system. Data analysis becomes complex, especially when dealing with complicated spectra and intensity fluctuations that are not clearly correlated with specific molecular concentrations. In some cases, these intensity fluctuations are essential, such as super-resolution blinking experiments, where variations in intensity help construct a single high-resolution image. Often temporal resolution itself carries valuable information. Recurrent Neural Networks (RNNs), especially those incorporating LSTM cells are well suited to this type of sequential data.320 Hidden Markov models have already been applied in plasmonic imaging to monitor dynamic cell membrane cluster interactions on cell surfaces321 and show promise for broader applications. Temporal convolutional networks (TCNs) are powerful for modelling sequential signals.322 Additionally, gradient boosting, used widely in image classification,323–325 has demonstrated strong performance in time-series analysis and could be highly valuable for time-resolved plasmonic data.326
Certain imaging modalities depend on both spatial and temporal resolution. Digital SERS is a key example of this, where single molecules are detected at specific locations across an array through high intensity signal fluctuations.327 Spatial analysis reveals heterogeneity and hot spot activity, and temporal analysis provides information on transient binding events, single molecule kinetics and dynamics. To date, ML has seen limited applications in digital SERS beyond standard algorithms. One notable example is the use of Gaussian process regression (GPR) alongside a PCA-LDA algorithm for digital SERS virus monitoring.328 Beyond this, however, there remains significant potential for ML to enhance the sensitivity and specificity of digital SERS analysis, particularly with RNNs, gradient boosting and the other algorithms discussed in this section. Additionally, transformer models may also prove useful in this domain due to their application in temporal and spatiotemporal tasks, for instance, TimeSFormer329 enables spatiotemporal feature learning and could offer significant advantages in analysing digital SERS datasets.
A wide range of algorithms are now available for image analysis, as outlined above, providing a strong foundation for plasmonic imaging. However, because this field is still in its early stages, there is significant scope for creativity in selecting and adapting methods. For instance, localisation of nanoparticles in noisy images might benefit from techniques developed for challenging recognition tasks, such as exoplanet detection, where such techniques are more common.330–332 Likewise, when spatial resolution is less critical than segmentation, such as for tracking disease progression in tissue, algorithms used for large-scale environmental monitoring (e.g., satellite tracking of wildlife populations)333 could offer valuable inspiration. As AI continues to permeate every aspect of image analysis, the breadth of available tools is expanding rapidly, presenting exciting opportunities to repurpose and innovate within plasmonics.
5.4 Refractometric analysis
Refractometric sensors, such as SPR or LSPR, generate spectral data that requires analysis to generate time traces (sensorgrams) from raw signals, then, these sensorgrams are further processed to extract binding affinity or analyte concentrations. These processes are typically done in sequence, where the plasmon resonance is extracted with some algorithms finding the peak maximum (or minimum) intensity, polynomial modelling of the plasmon resonance or using centroid methods. Then, the constructed sensorgram is split in different steps (i.e. washes, binding, regeneration, etc.) and further analysed to model the response to a binding algorithm (Langmuir and Hill's models are common) to extract biomolecular interactions. ML methods have become increasingly important in refractometric sensing because they provide automated interpretation of sensor outputs and improve robustness under variable conditions.334,335 Given that responses are often nonlinear, and biomolecular interactions can diverge from binding models, AI can contribute to deciphering hidden patterns and improve data interpretation.Supervised ML methods have become particularly valuable in refractometric sensing since they can associate sensor outputs with known molecular features for the extraction of parameters such as binding kinetics. Rather than relying on a single algorithm, recent work demonstrates that different supervised models excel at different aspects of the sensing process. For instance, feedforward neural networks (FNNs) can learn complex nonlinear relationships and have been used to predict binding affinities and classify analytes based on descriptors. However, their performance typically depends on the availability of large training datasets. CNNs, by contrast, leverage their ability to capture spatial and temporal patterns within sensorgrams (kinetic behaviours), enabling tasks such as distinguishing washing steps from binding events and detecting outliers.336–338 Regression-based approaches, including Gaussian process regression (GPR), RF and support vector regression (SVR) have proven effective when the goal is to model how variables such as molecular weight and concentration influence sensor response.339 These models are particularly useful when working with small or medium-size datasets. More specialised architectures have also emerged: Context Aggregation Networks (CAN)340,341 are well adapted to refractometric methods, since it can process and filter sensor images. Meanwhile, DNN342,343 offers a flexible alternative that can be tailored in depth and neuron density to accommodate the data availability and complexity of a given sensing application.
Unsupervised ML can uncover hidden patterns in SPR sensorgrams without requiring labelled data, clustering data into categories. When combined with PCA, non-negative matrix factorisation (NMF) enables the dataset to be distilled into a lower-dimensional space that highlights the most relevant structural and temporal features in the sensor data.344 Research on this combination remains limited, though it could significantly benefit refractometric sensing by enabling cleaner isolation of kinetic contributions, reducing noise-driven variability, and uncovering low-intensity features that enhance the interpretation of complex binding events. K-means clustering is especially interesting in SPR since it can cluster analytes based on binding kinetics.345
These unsupervised strategies increasingly merge with supervised approaches to build hybrid workflows capable both of structural discovery and predictive modelling. Integrating models such as CNNs and FNNs, enables sensorgram classification and outcome prediction. FNN architectures, including spectral parameter multi-layer perceptron neural network (SPMLP), have been developed specifically for superlattice verification in plasmonic biosensors. Using them has led to improved performance in distinguishing subtle signal patterns, while CNNs remain widely adopted in optical biosensing due to their strong capabilities in pattern recognition and image-based analysis.338,343,346–349 Additional workflows further extend this hybrid strategy. Another promising workflow is the combination of PCA and NMF, as the two methods complement each other. PCA will understand the data structure, while NMF will reduce temporal features. Signals obtained from various steps can be treated as ‘regimes’ using a temporal sensorgram descriptor (TSD) coupled to a k-NN algorithm which evaluates similarity-based distances.350
Refractometric sensors have been proposed to detect a broad range of analyte in portable and POC formats.45 AI is especially useful in this context,52,340,351–353 where key attributes where AI can help are rapidity, sensitivity, portability and ease of use,45 and automated interpretation of results without requiring an expert.348 Another advantage of ML is its ability to handle outliers arising from temperature variations, sample complexity, background noise and user-dependent errors. In SPR, undesirable variations in response can come from non-specific binding, variation in ligand density, bubbles and complex sample matrices. Background refractive index changes can cause issues for the quantification of molecules. GPR offers a solution to this problem, having been shown to model refractive index changes with high sensitivity.354 Binding kinetics depend strongly on the analyte and on the surface chemistry, so ML models must disentangle true kinetic signatures from experimental artefacts when applied to refractometric sensing. This requirement becomes particularly evident in systems where data acquisition is noisier, such as smartphone-based SPR platforms.350,355,356
AI can also markedly enhance both signal processing and data classification for refractometric sensors.357,358 For instance, in fibre-based SPR sensors, incorporating Gradient Boosting Regression (GBR) has enabled the detection of cancerous cells with strong accuracy, demonstrating how ML can reinforce sensitivity in complex refractometric platforms.359 However, despite these advances, numerous refractometric biosensors still rely on conventional data processing and could greatly benefit greatly from ML frameworks designed for specific sensing architectures.360,361 This potential is evident in emerging multi-component systems such as organ-on-a-chip (OoC) devices for evaluating drug-induced liver toxicity.360 These devices combine cell culture modules with SPR-based measurements but still require a specialist operator; integrating ML could streamline the analysis and help transition such devices towards bedside implementation. A similar need appears in multiplex diagnostics, where AI can assist in complex pattern recognition, as shown by an LSPR POC setup capable of detecting and discriminating multiple cancer biomarkers in human serum.362 The same trend extends to wearable systems: encapsulated nanosensors designed for wound monitoring, for instance, illustrate how ML can support continuous, real-time interpretation of physiological signals.363
ML can be a black box, which may not always be suitable for critical decision-making contexts, where users must be able to verify data quality. This is especially critical for the translation of POC sensors, where interpretability and reliability of the results are crucial factors to consider when integrating ML models into devices operated by minimally trained users. In such settings, users must rely on AI-driven devices trusting data integrity, as such transparent ML models could help reduce scepticism and foster trust and acceptance. This need for interpretability has contributed to a growing interest in XAI algorithms.342,354 These techniques work together by linking visual features and fitting to interpretative models, enabling users to understand how specific inputs influence the output. In refractometric sensing, this combined framework has been applied to estimate and interpret metal layer thickness in a fibre-optic SPR system, demonstrating how XAI can support trustworthy analysis in practical POC configurations. Across these diverse architectures, the unifying theme is that ML does not simply ‘add another layer of analysis’ but actively reshapes how refractometric biosensors are operated, interpreted, and deployed, reducing reliance on expert users while enabling more robust and adaptable sensing.
6 Applications of AI in plasmonic sensing
6.1 Plasmonic tongues and noses
The human nose and tongue function with arrays of non-chemically selective receptors whose pattern is interpreted by our brain to be associated with flavours and smells. The concept of artificial noses and tongues mimics this concept, using an array of sensors/receptors that can be used for sensing applications, notably in the food and beverage industry;364 a concept that can be generalised for other sensing applications.365 Chemical arrays facilitate the exploration of a sensing space with little prior knowledge on the exact chemicals to be measured and leverage AI tools to extract patterns from the complex output generated by a sample on the sensor. Central to their working principles is the fabrication of individually addressable sensors that can be chemically functionalised or not, to produce a different response on each sensing element within the sample, generating a complex pattern necessitating advanced data processing algorithms to extract correlations.366While a series of optical transducers can be used in sensor arrays,367,368 many rely on plasmonic techniques. On one hand, surface-enhanced techniques such as SERS369,370 or SEIRA371 provide rich vibrational data that can help in establishing molecular patterns associated with the sample, while on the other hand, refractometric techniques such as SPR,372 LSPR,373 or SPRi374 benefit from the simplicity of image or colour analysis to create sensor arrays. Transduction in surface-enhanced plasmonic techniques originates from the complex spectral information of SERS and SEIRA. In the case of refractometric techniques, the plasmon shift can be read from the direct interaction of the analyte with the sensor array,372,373 from the aggregation of the nanoparticles in suspension,375–377 or a colour change generated indirectly by the reshaping or resizing of the metallic nanoparticle from the interaction of the analyte with a metal salt that will cause nanoparticle growth or etching.378,379 The multiple transduction possibilities of plasmonic techniques make them versatile in the design of chemical noses and tongues.
Chemical selectivity of plasmonic noses and tongues mainly originates from the combination of distinct spectral responses on the array and data processing, but chemical selectivity can be imparted on the sensor array to increase the spectral difference between each address on the sensor array with specific or non-specific interactions. For example, the surface can be modified with DNA strands,380 organic,381 inorganic,382 or metal–organic framework (MOF)370 materials and self-assembled monolayers (SAM) to provide different partition coefficients for the analytes contained in the sample on each addresses to generate different spectral signatures interpretable by the data processing algorithm. Imparting chemical selectivity allows for the design of multiple iterations of chemical noses and tongues for applications in the food industry,373,375,376,383,384 in the detection of gases and volatile compounds,370–372,381,382,385,386 metal ions,387 small organic molecules, such as organophosphates,377 drugs,388 and neurotransmitters,378 proteins,380,389 or bacteria and cells.379,389,390
Given the many confounding factors intricated in the sensor array response, the information generated is generally uninterpretable by the experimentalist, therefore must be interpreted with ML algorithms. In many applications of plasmonic noses and tongues, classification and clustering methods including PCA,372,373,384 HCA,380 and LDA389 are commonly used alone or in combination369,377,378,382,388 to separate output into distinct classes. For example, more than 50 volatile organic compounds were separated in the response space of principal components, allowing the comparison of an electronic nose to human olfaction.391 The advent of NN algorithms capable of extracting information from more complex or abstract datasets should further improve the discriminative power of plasmonic noses and tongues; exemplified recently for gas sensing.392
While the potential of plasmonic noses and tongues is already demonstrated, their translation into fully operational devices will require a series of improvements. Data processing algorithms are especially susceptible to sensor drift,366 more so if the training model only included curated data with perfectly functioning sensors. In those cases, once the sensor fails or deviates from a normal response, the algorithm may misinterpret the data and generate a false outcome. Another source of improvement concerns the orthogonality of the sensor response and its susceptibility to cross-sensitivity with multiple species. An ideal plasmonic nose or tongue response would massively differ in each sensing element to provide the high orthogonality necessary to improve discrimination.364 However, many examples of plasmonic tongues used derivatives of SAM or DNA with potentially colinear responses to chemicals. Molecular receptor selection is key to achieve high discrimination power, and AI can play a role in doing so, as shown recently for the detection of structurally similar molecules.393 Using a high number of completely orthogonal chemistries will be necessary to increase the discrimination power of plasmonic noses and tongues, though this has the potential to increase the complexity of the readout processes. A compromise could be found in extensively calibrating the sensor arrays. Additionally, ML algorithms capable of a higher level of abstraction could be used to minimise the number of orthogonally responding sensors needed to increase the discrimination power. Taken together, the current innovations and future development of plasmonic arrays should lead to a series of applications in key societal challenges related to human health, environmental sensing, food sensing and industrial applications.
6.2 Clinical and disease sensing
Plasmonic sensors have potential as clinical biosensing platforms exploiting real-time sensing and enhanced detection sensitivity for both in vitro and in vivo,394–396 as well as diagnostic assays carried out on plasmonic substrates.397 Their incorporation into lab-on-a-chip technologies and portable and wearable sensors pushes the field towards useful POC devices.398,399 Clinical diagnostics often deals with very large datasets that critically need robust statistical validation and reproducibility for accurate decision making. Of course, AI is extremely beneficial for large and complex dataset processing and live analysis and is already being used extensively in this field.400 This section aims to highlight the areas in which clinical sensing has been achieved with a focus on emerging applications facilitated by the incorporation of AI.The other major improvement needed in clinical bacterial analysis is the development of accessible and rapid testing methodologies. This is more common for single pathogen detection, and highly sensitive results can be achieved,405,406 however clinical utility would be improved with the ability to classify across a broader diagnostic panel. If classification algorithms are sufficiently sensitive and simple, reliable methodologies can be used for substrate development that keep costs low for POC use, such as a paper-based SERS chip used in combination with a multi-branch adaptive attention CNN for classification that can identify pathogens in an hour.407 In other cases, the algorithms themselves can produce sufficiently accurate results more rapidly than standard techniques,408 but require more user-friendly diagnostic platforms. Conversely some more innovative configurations, including systems that exploit the plasmonic surface's inherent bactericidal activity, would benefit from using ML for faster processing speeds and enhanced multiplexing to maximise their practical utility.409 An alternative, yet still rapid, method for classification is by analysing multidimensional images of bacteria with a spectral transformer algorithm, based on a vision transformer (ViT) and compact convolutional transformers, combined with a data augmentation algorithm.410 This was shown to improve computation time and classification accuracy compared to the CNN mentioned previously.256 Though they did not use plasmonic materials, this image-based classification algorithm410 could be readily applied to similar plasmonic datasets, and integrated into more clinically deployable instrumentation. Technology is improving rapid bacterial classification, but broader clinical accessibility remains essential. Automation of these plasmonic platforms could substantially reduce user error and bias and limit users from potential exposure to dangerous pathogens.
The extracellular environment also provides abundant biomarker information, where SERS spectra can be simplified and classified with various ML algorithms,417,418 and live monitoring made possible with dynamic SERS measurements.419 Though not currently tested directly from biofluids, a DNN has been used to both detect and quantify different aggregation states of the same structural protein biomarker related to various neurodegenerative diseases with a SEIRA enabled microfluidic device.420 As this technology progresses, it is becoming a valuable tool for drug and biomarker discovery and diagnostics.154
Extracellular vesicles (EVs), such as exosomes, are a rich source of biomarkers and other biological information that virtually all cells release into the extracellular space for cell signalling and waste removal.421 Recently, they have been proposed for diagnostic and therapeutic applications, particularly in combination with plasmonic nanosensors.397 Classification algorithms are the most broadly used for determining EV origin, and identifying their components. SVM in particular has been able to accurately detect and stage breast cancer,422 and coronary artery disease,423 and to detect early-stage lung cancer, where the combination with a CNN was able to increase classification confidence.424 In another simple but effective example, another CNN was used to distinguish between various types of cancer using the SERS fingerprint spectra of exosomes.425 After isolating the exosomes from serum, the spectra were first classified in a binary scheme (healthy vs. cancer), then analysed with an MLP network to determine the origin of the EVs, successfully distinguishing six (broad) cancer types. Building on this, another group integrated a similar workflow (binary classification followed by MLP analysis to distinguish between 10 cancer types) into a SERS chip to remove the labour- and time-intensive exosome isolation process (Fig. 7(b)).426 Their model's interpretability also enabled the discovery of a new pan-cancer biomarker demonstrating its potential for future biomarker discovery efforts. Collectively, these studies show that ML-driven EV analysis is rapidly progressing from simple binary discrimination toward multi-class diagnostic insight. Next-generation models incorporating explainability,273 and integrated sensing platforms426 will substantially expand the clinical utility of EV-based plasmonic sensors. Overall, these developments illustrate how NNs can overcome biological complexity to deliver sensitive, specific, and clinically meaningful biomarker analyses.
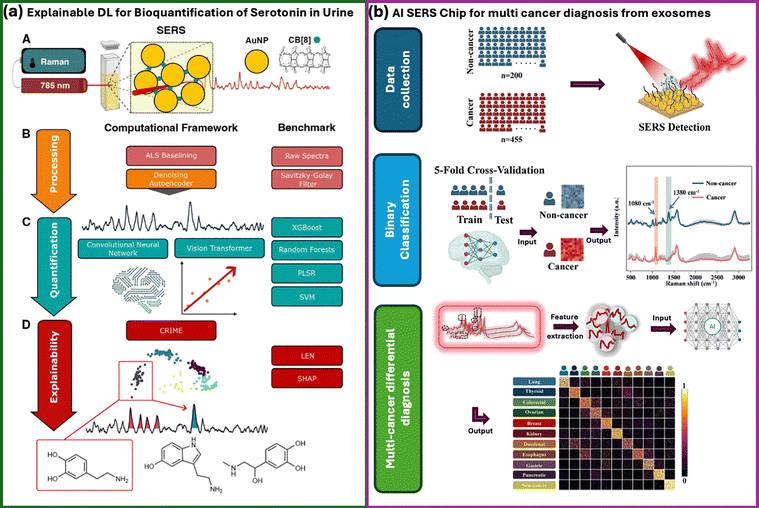 | ||
| Fig. 7 Some exemplary workflows from selected citations: (a) SERS deep learning framework development pipeline. Illustrated are the SERS measurement process applied (A) and the computational framework pipeline. Benchmark comparisons of alternative methodology are presented on the right. Preprocessing methods (B) are marked in orange and light red, quantification methods (C) are marked in blue, and explainability methods (D) are marked in dark red. Figure adapted with permission from ref. 274, licensed under CC-BY 4.0. (b) Diagram showing the process of cancer identification via AI-SERS. Comparison of average SERS spectra and heatmaps of serum exosomes from non-cancer controls and cancer patients (based on 25 spectra per group). Schematic of multi-cancer classification heatmap of the classification probabilities derived from spectra for eleven classes (ten cancer types and non-cancer group). Figure adapted with permission from ref. 426, licensed under CC-BY 4.0. | ||
AI is also increasingly influential in tissue-level imaging, particularly for image reconstruction, analysis, and segmentation as well for feature extraction for clinical imaging with plasmonic techniques. For high-quality tissue imaging (be it ex vivo, wide-area mapping, or real-time dynamic monitoring) advanced reconstruction algorithms are becoming essential. These methods compensate for weak or noisy optical signals, correct artefacts from rapid acquisition, and enhance spatial and molecular contrast that would otherwise be lost in raw data. It also provides better image stitching, where the contrast may differ at the edges of an image and mimic the image output of tissue staining. In the context of static diagnostic imaging, histopathology remains the gold standard. However, the field is beginning to incorporate spectroscopic imaging to capture molecular information that conventional staining cannot provide. Drawing inspiration from stimulated Raman scattering (SRS), this technique has been combined with a self-supervised DL algorithm to reconstruct SRS data into tissue maps, convincingly showing image output that resembles H&E staining.432 Alternatively, a DenseNet-based learning method was used for spectral feature extraction-based image segmentation at subcellular resolution,433 and Raman used for tissue age analysis with DL combined with MCR-ALS.434 SERS holds the potential to further improve the multiplexing capacity of spectral imaging to reveal more complex biochemical information.435 Alternatively, SEIRA is emerging as a viable clinical approach,436 and could similarly benefit from strategies that have advanced conventional IR, such as DL based models used to predict breast cancer reoccurrence from mid-IR histopathological imaging.437 Although the field is not yet at the point of widespread clinical deployment, the pace and direction of development indicate that such capabilities are becoming increasingly attainable.
Recent advances in combining plasmonic techniques with ML are beginning to overcome longstanding barriers to clinical adoption. The introduction of automation in analysis could be a game changer for POC applications. Another challenge in clinical sensing is clinical sample availability, currently limiting the size of datasets necessary for accurate model training. To circumvent that, data augmentation has been proposed,410,441 demonstrated with a NN to extract SERS spectra of viruses over a range of concentrations, which the dataset was then augmented by adding realistic residuals to the extracted SERS spectra.442 The authors used this augmented dataset to train their classification model and then quantified viruses in saliva samples with 91.9% accuracy. Carefully using NN-based data augmentation could improve model performance, provided the data remains representative, as there is a risk of introducing unrealistic features in the training dataset. Another limitation of NNs is the inability to interpret which spectral features drive classification, which could be overcome with more explainable AI. To support this, the quantification of serotonin was shown in urine samples using SERS with a CNN and vision transformer model. This model revealed the features at the origin of classification decisions using a new Context Representative Interpretable Model Explanation (CRIME), based on a LIME framework (Fig. 7(a)).274 The implementation of explainable AI could drastically reduce the black-box limitations of current ML models, an essential step for clinical decision-making. Looking ahead, advances in data augmentation, the development of larger and more representative Raman datasets, improved multiplexing strategies, and integration into self-driving or real-time analytical frameworks will be key for enabling genuinely translational, AI-driven plasmonic diagnostics.
6.3 Environmental and plant sensing
Environmental science is another field where the coupling of AI with plasmonic analytical techniques holds potential in monitoring aquatic environments, air quality, agriculture, among others. Continuous or frequent monitoring of molecular parameters can provide detailed information on the impact of pollutants and stressors on the environment. The high volume of data generated by real-time monitoring, imaging, and sensing benefits greatly from ML and AI. To exemplify that, multiple studies have recently used these algorithms for the observation of microplastics,443–445 of pollutants,446,447 for soil analysis,448–450 and in large panel pesticide detection.451–454 Some fundamental examples of this are demonstrated in Fig. 8.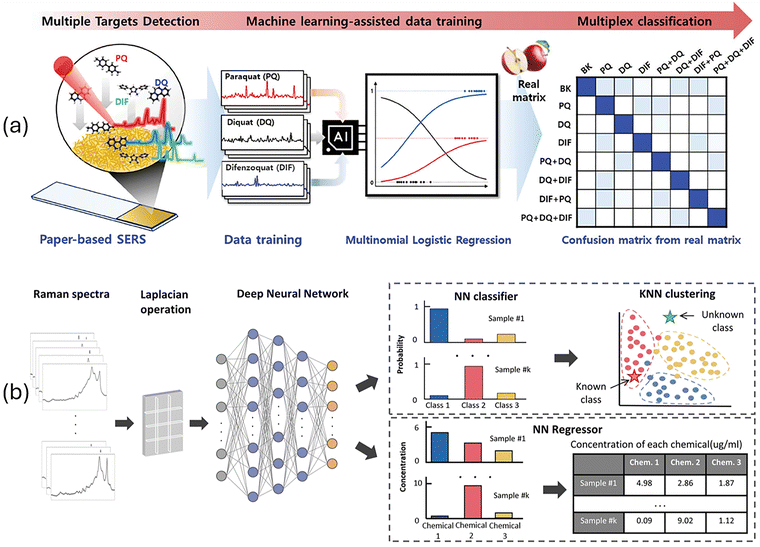 | ||
| Fig. 8 Selected figures from cited articles demonstrating use of ML for environmental classification (a) Schematic illustration of a plasmonic sensor deployed for the multiplex classification of 4 pesticides in a ground apple matrix. It features as its ML method the multinomial logistic regression, a variant of logistic regression adapted for cases where more than two discrete states or categories must be represented. Reproduced with permission from ref. 454 licensed under CC-BY 4.0. (b) The schematic diagram of a spectra processing pipeline centred around a DNN used to both identify and quantify organic pollutants in water. Figure adapted with permission from ref. 446 licensed under CC-BY 4.0. | ||
Several key factors must be considered for the application of AI in environmental and plant sensing. The selection of the appropriate algorithm and complexity level depends on the task at hand. Simpler, well-proven statistical methods such as ANOVA or ANCOVA combined with discriminant analysis can yield quality results for simpler datasets,455 which can be beneficial for the field deployment of plasmonic sensors. It is thus common to see chemometrics methods used hand in hand with AI models, which have been exemplified by a PCA456 upstream of a classification model for the detection of trace explosives.457 Others have shown that multi-layer perceptrons can also be considered for simple data modelling in both classification and regression tasks before applying DL models.458 However, if resources are to be invested in training a more complex model, there is an opportunity to make certain models serve double duty. As a recent example, a SERS sensor leveraged a single support vector-based model for both classifying and quantifying pesticides.459
Models of increasing complexity, such as CNNs, are now commonly employed in the classification of environmental spectral data and are often combined with other metrics, such as the spectral angle, which maps information to a reference spectrum,460,461 similarity indices, which measure the difference in spectral intensities,462 or normalised weighted cross-correlation.463 One notable example of a regression task in environmental analysis was the quantitative analysis of chlormequat and paraquat residues, rendered viable at low concentrations with AI.464 In that study, SVR and k-NNR were compared with two ANNs (MobileNet465 and CNNAC466), the ANNs afforded more consistent performance, exemplifying that DL models are more suited for large and complex spectral datasets. Interestingly, the best performing model was interestingly the implementation of a MobileNet,464 a family of networks initially created for computer vision. This aligns with the growing trend of utilising computationally intensive models for spectral processing.
Plant life observation is an emergent field of sensing; therefore, protocols must be developed to translate the vast knowledge gathered from human health sensing to plant monitoring. Easing this transition is the fact that AI algorithms are agnostic to the type of cell or organism sensed. A recent review discusses the main families of Raman spectroscopic processing for agricultural applications.467 Since then, avenues to push existing sensing methods and extract more information from datasets made with current methods have appeared. There have been many novel approaches in plant contaminant analysis that were made possible by AI.468 One of the opportunities demonstrated is the integration of capillary force-enhanced dynamic SERS,469 and the implementation of 3D SERS imaging.470 Dynamic SERS and imaging have also been adapted for plant health monitoring, where it was used for the detection of plant infections471–473 and to monitor specific indicators of general plant health based on plasmonic sensing,474,475 representing important developments.
Despite these advances, there are still areas where the community would benefit from adding AI to their panel of tools. The most important is in the determination of specific and intrinsic characteristics of plants.476 Successful studies of sex determination,477 ecological character,478 and intercommunication479 were reported, while transposition to phenotyping and genotyping is expected for important plant species, as it was recently done for mammalian cells.430
Because plant cells represent a different environment than animal cells, adapting technologies developed for other applications is a central avenue to enabling plasmonic techniques to become a pillar in plant science. Challenges with the use of nanoparticles in plants have yet to be fully solved. The main one is optimising the nanoparticles themselves for uptake by various parts of the organism,480 and ensuring colloidal stability in plants for the duration of the experiment.481 There is still a need to develop NP surface chemistry specifically tuned for minimising nonspecific interactions in plants. As our collective understanding of the biofouling of nanoparticles in all living tissues expands, it now appears that plant cells have different requirements than animal cells in terms of effective surface chemistry to avoid fouling from plant biomolecules such as proteins.482 Efficacious strategies to mitigate biofouling specifically in plant environments remain to be achieved.
Raman-based techniques are more present in the current literature than other approaches, such as SPR in plant analysis, since vibrational spectra inherently contain more information. Though instrumental design tends to restrict its use for whole plant organ measurements, there is great potential for fast, specific analysis of biomolecules to be found with SPR sensing. The utilisation of SPR sensors to measure species of interest to botanists represents a certain gap in current plant sensing technology.
To summarise, investigative studies of the molecular makeup of plants using plasmonic sensors are a current frontier where AI is being applied with increasing frequency.483 As the field is nascent, it leaves ample opportunities to develop sensors for environmental and plant analysis. This growth will require the implementation of AI in plasmonic environmental sensing, as it remains sparsely used. Early studies have shown its undeniable potential,484 which will only be expanded by further implementing AI and ML in plasmonic sensing for environmental and plant analysis.
6.4 Food sensing
Food sensing with plasmonics has benefitted from the integration of AI, reshaping the ability to monitor safety, authenticity, and quality across many food systems. Plasmonic techniques provide extraordinary sensitivity by amplifying weak vibrational and refractive signals, yet their spectra are often obscured by noise, fluorescence, and overlapping peaks arising from heterogeneous food matrices. Traditionally, chemometric approaches such as PCA, PLSR, SVM, and RFR have formed the analytical foundation for plasmonic sensing of food. These models effectively reduce spectral complexity and build calibration models that link signal intensity to analyte concentration. They are valued for their interpretability, simplicity, and low computational demand, and have been widely applied to SERS-based studies ranging from milk adulteration, pesticide detection, and other food safety challenges. However, their reliance on feature selection and linear assumptions constrains their ability to handle the nonlinearities and high dimensionality intrinsic to plasmonic spectral data. This limitation underscores the growing need for advanced AI models capable of autonomously learning complex spectral representations and improving the robustness and generalisability of food analysis with plasmonics.51,57A dominant trend in this field has been the use of DL to push detection limits into ultra-trace regimes while preserving robustness against complex matrix interferences. End-to-end CNNs trained directly on SERS spectra have achieved quantitative detection of pesticides, veterinary drugs, and mycotoxins at ppb and sub-ppb levels across diverse food matrices, including fruits, vegetables, oils, fish, and grains, without requiring extensive preprocessing.269,485–488 These models autonomously learn discriminative spectral features directly from raw data and remain robust even under significant background interference. This data-driven adaptability has enabled researchers to demonstrate detection of thiram in fruit peels,489 malachite green in fish,487 and aflatoxins in oils486 at concentrations aligned with or below regulatory thresholds. More recently, transformer-based and attention-based architectures have further extended these capabilities by capturing long-range dependencies across spectra, improving model generalisation when peaks broaden or overlap, and enhancing interpretability through saliency mapping that highlights chemically meaningful vibrational bands.269,488,490
Beyond achieving low concentration sensitivity, DL has also enabled multiplexing and the ability to detect and quantify multiple analytes in a single measurement. Food samples often contain mixtures of pesticides, antibiotics, and preservatives that generate overlapping spectral signatures, where traditional chemometric approaches often collapse to disentangle component contributions. In contrast, DL architectures demonstrated remarkable proficiency in resolving these convoluted spectra. For instance, pseudo-Siamese CNNs have been used to quantify mixed antibiotics in human milk at ppb levels,491 while hybrid ensemble models combining CNNs with neural regularisation strategies have successfully resolved ratios of co-occurring antibiotics in aquatic foods.492 These data-driven frameworks advance plasmonic sensing toward comprehensive multi-contaminant profiling without extensive preprocessing or sequential testing. Similarly, decision-level fusion integrating SERS and fluorescence data analysed by CNNs delivered R2 exceeding 0.99 for simultaneous determination of preservatives and heavy metals in mushroom samples, substantially outperforming single-modality baselines.493
Pathogen detection represents another frontier where DL has transformed plasmonic sensing. Bacterial and fungal spectra acquired through plasmonic techniques are often subtle, masked by background signals, and difficult to classify using classical tools. DL overcomes these challenges by automatically extracting subtle spectral-biochemical correlations that differentiate closely related pathogens with high accuracy. CNN-based classifiers trained on thousands of SERS spectra from swabs and contaminated food items have achieved near-perfect accuracy in identifying common pathogens such as E. coli, Salmonella, and Listeria.494 More advanced architectures, such as NAS-optimised U-Nets augmented with attention mechanisms, have further expanded this capability, accurately distinguishing dozens of subspecies across multiple genera with accuracies exceeding 90%.495 Beyond accuracy, these models enhance interpretability by mapping Raman features linked to biochemical markers of specific pathogens. The implications for food safety are substantial: DL models trained on large, heterogeneous datasets can be deployed in portable SERS platforms, enabling real-time pathogen surveillance across supply chains.496 Although SERS remains the most widely studied modality, related plasmonic techniques, including SEIRA, LSPR, and SPR, have also incorporated DL models, enabling accurate sugar quantification, milk adulteration detection, and improved mycotoxins or adulterants analysis in milk through data-driven spectral interpretation.497,498
Beyond pathogen detection, DL also strengthens applications in authenticity and adulteration monitoring. Multiplex SERS fingerprinting combined with NN models has been employed to authenticate agricultural products,499 and to identify dye adulteration in herbal products such as saffron and curcuma500 with high reliability. The development of flexible plasmonic substrates further enhances this synergy by ensuring conformal contact with irregular food surfaces, thereby improving spectral reproducibility and enabling DL-based regression models to achieve high predictive accuracy.489,501 These examples illustrate the co-evolution of substrate engineering and AI in enhancing both sensitivity and robustness. In parallel, DL has accelerated multimodal sensing, where plasmonic techniques are coupled with complementary modalities such as fluorescence spectroscopy and Fourier-transform near-infrared (FT-NIR) spectroscopy. By extracting synergistic spectral features from heterogeneous datasets, DL models deliver more reliable predictions than single-modality systems.493,502 This integrative capability underscores AI's emerging role as a unifying analytical layer, making multimodal plasmonic sensing practical for real-world monitoring where traditional single-modality approaches often fail.
Equally important are the modelling strategies that make DL truly deployable in real-world food monitoring scenarios. Multitask architectures have been introduced in SERS analysis to simultaneously handle detection, classification, and quantification, aligning decision boundaries with concentration gradients.269,488 To enhance model resilience, data augmentation techniques, including spectral shifting, noise injection, and band superposition, are increasingly used to improve CNN robustness against instrumental drift and environmental variability.499 Transfer learning and domain adaptation further reduce data demands by enabling pretrained spectral models to be fine-tuned on a limited number of new samples,503 while ensemble strategies add an additional layer of reliability through uncertainty quantification, flagging low-confidence predictions in heterogeneous food matrices.51 Collectively, these developments represent a pivotal shift in plasmonic food sensing, from proof-of-concept laboratory studies toward practical, deployable diagnostic platforms. DL not only reduces reliance on expert-driven preprocessing but also allows the development of portable and flexible sensing systems that can operate directly in the field. DL models trained on large, augmented datasets overcome this variability, enabling accurate detection without manual corrections.
Looking ahead, key challenges remain, particularly in model interpretability, calibration transfer across substrates and instruments, and the need for large, standardised spectral repositories. While attention-based tools and data augmentation offer partial solutions, emerging paradigms such as physics-informed NNs and federated learning are likely to play a critical role in transparent, transferable, and trustworthy models. AI is no longer peripheral but central to food sensing with plasmonics, driving the transition from laboratory studies to field-ready diagnostics that meet global food safety and quality demands.57
6.5 Materials characterisation
Extensive characterisation of plasmonic nanomaterials is required to ensure consistent synthesis and integrity of the nanoparticles. Electron microscopy (EM) is the benchmark for imaging plasmonic nanomaterial morphology, but its reliance on intensive sample prep, expensive equipment, slow imaging and experimentalist-centred image analysis makes it impractical for real time or dynamic environments. AI-driven approaches are emerging as faster, more scalable alternatives, offering predictive insights without the constraints of traditional EM protocols. For example, DL has been applied to denoise EDX tomography data, enabling high-resolution elemental mapping of nanomaterials with reduced radiation dose and acquisition time.504 DL can denoise electron microscopy images by enhancing contrast and resolution with NNs, and is especially useful for nanostructure imaging with low-energy electron microscopy, where resolution can be improved to sub nanometre, comparable with STEM.505 In addition to EM, optical techniques are frequently used to characterise plasmonic nanoparticles. Among them, dark-field imaging is commonly employed in plasmonics as the scattering of nanoparticles can be measured in real time, which can be facilitated by a CNN for rapid structural characterisation.506 Similarly, NNs have been trained to process the LSPR signals from differential wide-field imaging (DWI) in complex biological environments.507ML analysis of imaged nanostructures can reduce reliance on computationally heavy physics-based simulations. For example, DL has been used to efficiently evaluate plasmon coupling of electrostatically deposited NP, offering insights into surface interaction and correlating plasmonic properties with observed colour features on the deposited surfaces.508 Such approaches bring forward this reduced reliance on simulation-heavy spectral analysis, making plasmon coupling a more accessible and insightful approach to characterising nanomaterials. Furthermore, AI enables the use of alternative characterisation techniques that rely on complex image patterns, such as the CLoCK method. This technique allows high-throughput analysis of single-particle dimensions such as; length, width, and aspect ratio of gold nanorods (AuNRs), with accuracy approaching that of electron microscopy.509
Beyond physical characterisation, applications exist for the use of plasmonic imaging methods coupled with AI. One of them concerns the characterisation of plasmonic NP arrays, which is especially important for developing functional optical tags, such as those used in anti-counterfeit labelling. NNs are applied to dark-field imaging data to accurately identify and authenticate specific NP arrays. This is achieved by extracting key structural parameters such as array size, well width, well spacing, and cell count from the dark field images. In this case ML provides a robust framework for translating these features into reliable tag verification, enabling practical and scalable array characterisation.510
AI is also being applied to extract physical parameters from spectral data and particle motion, expanding the scope of material characterisation. ML and NNs have been used to analyse spectral data from techniques such as XANES and extinction spectroscopy, enabling precise size and shape characterisation of nanoparticles.206,511,512 This is particularly valuable, as traditional spectral interpretation often relies on empirical judgments that can lack precision. AI also facilitates the extraction of dynamic properties. For instance, rotational diffusion coefficients of single nanorods can be determined using frequency-domain DL methods, which are applicable to viscosity measurements in biomolecular condensates.513 Collectively, these approaches underscore the growing role of AI in enabling scalable, real time, and cost-effective optical characterisation, offering viable alternatives to traditional EM and opening new avenues for single-particle analysis in complex environments.
6.6 Feedback loops to self-driving labs
Feedback loops in plasmonic sensing represent a frontier in adaptive experimentation. Traditionally, scientists have relied on intuition-driven trial-and-error experiments guided by manual observation, or one-factor at a time experimentation.514 A simple illustrative example is the iterative gold salt adjustment in Turkevich–Frens nanoparticle synthesis, where subsequent additions of gold salt are informed from the feedback of the UV-Vis-NIR until a desired size is reached.105,515 However, as the design space expands from material synthesis to substrate preparation and sensing, human-intuition guided loops become increasingly complex, constraining interpretation and optimisation. To overcome this bottleneck, AI-driven systems offer a path toward scalable, autonomous feedback, especially in the context of emerging self-driving labs (SDL) with chemical sensing in this loop.516 In the context of plasmonic sensing, these feedback loops could be beneficial in optimising various aspects of sensor design (transducer properties, surface chemistry, performance optimisation), in quality control of sensors, and while sensors are being used to ensure proper functioning and calibration. Different approaches developed in other fields include self-optimisation, Bayesian optimisation (BO) and reinforcement learning (RL) could also be applicable to plasmonic sensing and researchers in the field could draw inspiration from them.BO stands out as a prominent self-optimising strategy designed to minimise both experimental effort and material consumption.518 Based on Bayesian statistics, the predictive model estimates the system's behaviour iteratively based on past observations, and through pattern recognition from the acquisition function BO will continually refine the certainty of predictions to steer experimentation toward optimal conditions.520,521 This balance is achieved through the selection of appropriate surrogate models, which mimic the design space and the selection of proper infill points for new experimental conditions to be predicted for model accuracy improvement to converging desired reward.50 The success of BO relies on several factors, including the quality of the acquisition function, the smoothness of the predictive model (or response surface) and the dimensional complexity of the design space.522 Uncertainty guided response surfaces play a crucial role, as they help BO in balancing exploration of unknown regions with exploitation of known promising predictions. BO could be a powerful experimentation tool for guiding plasmonic sensing discovery and optimisation, particularly due to its efficiency in handling sparse datasets and streamlining experimental decision making. This sets the stage for BO's implementation across diverse plasmonic sensing/sensing-related workflow, as illustrated in the following examples.523
On the side of material development, BO was integrated to deepen the understanding of gold nanorod (AuNR) growth dynamics and reactions.524 This self-optimisation strategy incorporates consulting the BO model to design the next set of experiments for converging a desired UV-Vis-NIR spectrum. Using a Gryffin framework, they explored the growth parameters of AuNS, and targeted different methodologies for synthesising particles with similar UV-Vis-NIR spectra. Five growth protocol parameters were added as inputs and the UV-Vis-NIR spectra as the output. This strategy uncovered additional growth pathways under conditions that may have been traditionally seen as unfavourable (such as high ascorbic acid concentrations and high temperatures). Hence, BO was used as a predictive tool to reveal previously overlooked parameters and alternative growth mechanisms.
An exemplary application of BO in plasmonic sensing is demonstrated by Giordano et al., who employed a multi-objective BO framework to fabricate high-performance SERS substrates.525 Their approach used BO to guide the silver coating on SERS substrate, gold nanostars, to maximise SERS intensity and signal uniformity across the substrate. By incorporating spin speed and blade velocities as inputs, the BO converged towards target outputs of high SERS signal intensity and reproducibility. The authors accelerated the optimisation processes for two different coating protocols of spin speed and flow coating, by factors of 8.8x and 1.6x, respectively. In this case BO accelerated the optimisation of SERS substrates with high enhancement and reproducibility, underscoring its value as an intelligent substrate tuning strategy.
Despite its strength in iterative feedback, BO faces limitations in high-dimensional spaces, a challenge that shapes its practical utility in complex plasmonic sensing schemes. BO acts as a high bias/low variance classifier, reducing the need for extensive hyperparameter tuning and large datasets.186 Both examples succeeded as they worked in low-dimensional design spaces, but in the case of larger design spaces, BO's effectiveness tends to decline due to increasing computational costs and model complexity.520,522 Therefore, such problems which may need higher dimensionality design spaces, such as those involving simultaneous control of substrate methodology, inline material characterisation, and inline sensitivity optimisation, may benefit from alternative algorithms better suited to navigating expansive design landscapes.
RL is a dynamic decision-making problem-solving paradigm built for real-time decision-making, for systems that require continuous interaction with complex environments.526 RL typically operates within the framework of Markov Decision Processes (MDPs), or trial-and-error exploration, where intelligent agents interact with the environment by performing actions and observing the resulting changes in system states. These agents evaluate how closely each new state aligns with a desired reward; thus, the agent refines future actions to converge on a reward function.527–529 A key strength of RL is its ability to exploit delayed rewards, where agents can associate current actions with outcomes that occur much later.528 This temporal flexibility enables RL to navigate high-dimensional, complex tasks, which pose challenges for traditional ML feedback-loop algorithms. RL's decision-making architecture makes it particularly well suited for integration within the context of a closed-loop laboratory.528 Moreover, RL's real-time adaptability provides a powerful framework for autonomous learning and discovery, allowing agents to explore and refine behaviours in real time for efficient learning and discovery.
A compelling example of RL in autonomous nanomaterial discovery is AlphaFlow.528 AlphaFlow is a closed-loop platform designed to optimise the synthesis of core–shell semiconductor nanoparticles. By integrating modular microfluidic processors, AlphaFlow autonomously designs and executes experiments without prior knowledge of the system or reaction conditions. It performs reactions in 10 µL microdroplets and continuously monitors them using inline spectral techniques, such as photoluminescence and absorption spectroscopy. AlphaFlow explored over 1012 possible reaction sequences and autonomously performed hundreds of microdroplet reactions per day over a 30-day continuous run, more than what 100 human chemists could perform in that time. As a result, AlphaFlow can efficiently and iteratively refine synthesis strategies, uncover novel synthesis routes, and eliminate unnecessary steps. Crucially, it leverages RL's delayed reward system, where the agent evaluates long-term outcomes of sequential actions, learning to associate early decisions with final product quality and system performance.
Although AlphaFlow was originally designed for quantum dots, its methodology and integration with RL offer a powerful framework for optimising plasmonic nanomaterials, particularly in real-time material tuning and sensing applications. Inline detectors serve as real-time environment assessment, guiding the selection of the most promising next steps toward a desired optical outcome.56 This technology allows RL agents to learn experimental parameters to maximise a reward's link to the detector's readout for characterising the nanomaterial's properties. In plasmonic sensing, RL's strength lies in planning sequences of decisions (or actions) that optimise the final performance of the sensor, including parameters such as selectivity and sensitivity. Further, RL can be leveraged for optimising sensor readouts by incorporating the sensor readout into the RL feedback loop. RL has the potential to associate NP methodologies, sample preparation and instrument optimising strategies to design sensors with optimal behaviour.530 Fault detection is another possible use of RL in sensing, where the algorithm monitors the sensor output to detect abnormalities. While it has not been shown yet in plasmonic sensing, other types of sensors have integrated this approach.531 This provides groundwork from performance prediction to material design for plasmonic nanosubstrates, and potential full automation with self-driving labs for chemical sensing.
Fluidic hardware coupled with (semi)-closed-loop workflows has been key to accelerating SDL and NP discovery.516 Continuous flow synthesis enables precise reagent control, real-time characterisation, and adaptive decision-making.516 Fluidic hardware streamlines experimentation through integration into closed-loop optimisation, linking growth reagent parameters to optical properties via inline detectors. This feedback-driven automation allows SDLs to autonomously tune reagents and refine colloidal NP synthesis strategies. For instance, one platform employed a continuous-flow reactor combined with inline UV-Vis-NIR spectroscopy to autonomously explore multiple synthesis routes for gold nanoparticles.535 Using ML models, including RF, Gaussian Process, and SVR, the SDL established predictive relationships between reaction conditions and optical outcomes, accelerating convergence toward a target LSPR. Building on these capabilities, the AFION (Autonomous Fluidic Identification and Optimisation Nanochemistry) platform further exemplifies the SDL paradigm by integration microfluidic with controlled photochemical nanoparticle tuning.536 AFION targets specific optical properties in real time, guiding synthesis pathways dramatically. In just 30 experiments over 30 hours, AFION successfully produced eight distinct nanoparticle morphologies, including gold nanorod, Au/Ag core-shells, and tetrapods. Thus, underscoring its strength in autonomous feedback driven synthesis and high-throughput morphology exploration.
Robotics have also been integrated to autonomously synthesise gold NPs. For example a system equipped with automated liquid handling, inline UV–Vis spectroscopy and ML for synthesis, was able to autonomously discover five AuNP morphologies.537 Through their ML-driven decision making, the investigators combined the analysis of extinction spectrum simulations with experimental data to train predictive models that could forecast the outcomes of new synthesis conditions. The robot algorithms-controlled reaction conditions and performed ∼1000 experiments. This system brings to light the opportunity of robot lab assistants to perform more labour-intensive experiments.
Tracking the evolution of autonomy of SDLs from modular, piece-wise systems to integrated closed-loop workflows reveals new opportunities for sensing. Advanced SDL for sensing could orchestrate substrate synthesis, sample-substrate preparation, and application-specific optimisation within a unified framework. A representative example is a two-step workflow: (i) automation of the synthesis of NP and inline characterisation, followed by (ii) inline characterisation analysis for the selection of new growth parameters for material discovery. Spectral features, such as LSPR shape and intensity, are directly correlated to the size, distribution and morphology of the colloidal NP, allowing for close to immediate interpretation, feedback and refinement. To push the boundaries of SDL capabilities, multi-step workflows offer tandem exploration, moving beyond NP colloidal optimisation toward integration into sensing applications.56 SDLs enable rapid exploration of multidimensional experimental spaces and can be evolved from property tuning to functional material discovery for chemical sensing. By controlling every step, from sensor fabrication to deployment, SDLs offer the potential for on-demand sensor production, addressing challenges in stability, reproducibility and calibration issues encountered with batch-prepared sensors with limited shelf-life.
Future research should bridge the spectrum of automation, with (semi) closed-loop workflows serving as a critical launch pad toward fully autonomous self-motivated discovery platforms.533 To extend towards self-motivated system, SDLs must expand how input-data is accessed. This could involve incorporating toolset, such as LLMs to extract metadata from literature figures, experimental protocols, and open-access repositories538–540 and implementing this in fully integrated SDLs.541 Establishing common data schemas and validation protocols will ensure accessible inputs are machine-ready and interoperable across discovery tools. Closing the loop on autonomous discovery requires feeding these insights back into the system for iterative optimisation without manual intervention.
The ability to implement SDLs in sensing research workflows will require a series of advances, as only a few examples have been reported with plasmonic sensing. The first one will be to design sensors compatible with integrated computing workflows, where the sensor output can be fed in the algorithm. Not all plasmonic sensing technologies have reached this level of computer automation and it poses a limitation in the adoption of SDL and other ML-driven optimisation protocols. Establishing boundaries remains necessary, as there are technical limitations to the various instruments used in plasmonic sensing. For example, pumps have limited ranges of delivery volumes and flow, lasers function well in a range of powers and detectors have limitations in acquisition rates, wavelength ranges and so on. Implementing these boundaries is necessary to avoid optimisation in a space that is technologically impossible to achieve. Finally, algorithms will need to be openly available to the community to increase the accessibility of these tools to researchers in the field. The potential of these approaches is vast, and one can envision fully autonomous decision-making plasmonic sensors, similar to what is now possible with unmanned aerial vehicles for gas detection.542
7 Outlook
The present review offers a concise overview of the opportunities emerging from the integration of sensor design with advanced, AI-driven data-processing methodologies across a diverse set of applications. Data processing and sensing are inherently interconnected, as the overall performance of a sensor relies on the accurate extraction of information from its response. This interdependence has shown limitations, as recent advances in sensor design have led to larger and more complex datasets that challenge the capabilities of classical algorithms. Taking examples from SERS, where spectral wandering and intensity fluctuations are distinct spectral patterns in single-molecule spectroscopy that cannot be efficiently captured by traditional methods. Similarly, the substantial data volumes generated in SERS imaging experiments now exceed the practical limits of traditional data-processing approaches. In this context, the emergence of ML and AI provides timely and powerful alternatives to classical data processing methods.Despite these promising developments, AI-enabled plasmonic sensing remains in its early stages, with substantial room for advancement. Many of the studies highlighted in this review are still at the proof-of-principle level, and significant progress will be necessary to translate these approaches into practical tools for end users. Such translation will require improvements in sensitivity and specificity to increase the performance of the sensors; enhanced explainability of the spectral features used for decision-making as AI algorithms can be somewhat of a black box; and improved integration of AI can be achieved with sensor-design workflows. Another challenge is the detection of weak signals in a large background. ML excels at analysing strong spectral features, but sometimes the analytical signal of interest is not the dominant feature in the spectra. A recent approach proposed spectral unmixing, with a self-supervised algorithm, spectral separate network (SSNet),543 which was then applied for the detection of antibiotics.544 At the same time, these are merely examples of conceptual and methodological gaps that need to be filled.
Other research avenues could benefit from AI and ML in plasmonic sensing. A major and current challenge in the field is achieving reliable calibration in complex and dynamic environments. The community increasingly aims to develop sensors capable of operating in clinical biofluids or in environmental samples, enabling continuous monitoring of diseases or environmental stressors, among other promising applications. Across these use cases, sensors are routinely exposed to matrices and environmental conditions that can induce fouling, temperature fluctuations, or degradation of sensor components, all of which contribute to drift in the sensor response. These factors presently limit the deployment of sensors for long-term monitoring, as conventional calibration schemes often fail once the sensor response deviates from its laboratory-derived calibration. Because the stressors affecting sensor performance are multifactorial, traditional calibration approaches are often inadequate for compensating such drift. ML, by contrast, is well suited to capturing complex relationships between environmental perturbations and sensor responses, offering powerful, yet still relatively underexplored, strategies to correct drift and enhance calibration robustness.
Another fundamental challenge arises when advanced ML algorithms are repurposed for quantitative measurements. Many of these algorithms were originally developed for classification or decision-making tasks, and their direct application to quantitative sensing introduces significant risks. Improper model selection, inadequate training, or overfitting can lead to erroneous sensor outputs. Exacerbating this issue is the limited interpretability of many ML models that may prevent users from recognising such errors, thereby undermining the reliability of quantitative decisions. Potential solutions include embedding quantitative constraints directly into ML architectures or employing hybrid approaches in which ML models extract features or preprocess data prior to applying established quantitative algorithms. While these strategies are promising, the deployment of ML for quantitative sensing remains an area requiring substantial further development.
Decision-making capabilities in ML are expected to become increasingly important as plasmonic sensors move toward real-world deployment. One envisioned application is the continuous monitoring of drug concentrations, where closed-loop systems could automatically adjust dosages in response to sensor outputs and thereby improve disease management. However, ensuring data quality is particularly challenging when ML-based systems are involved, and maintaining proper sensor function will be critical for safe operation. A faulty sensor response that mimics a low drug concentration, for instance, could lead to inappropriate dose adjustment and potentially dangerous overdoses. Consequently, the integration of explainable AI into sensor design and data-processing pipelines will be essential to evaluate response quality, detect anomalies, and ensure trustworthy decision-making. Decision making is also a hallmark of sensor development. This is an area where SDL can assist the scientist by using AI-driven optimisation algorithms for maximising performance of plasmonic sensors by integrating experiments and feedback loops.
While AI offers significant opportunities for plasmonic sensing, several limitations remain. Many current sensor scientists lack substantial training in chemometrics and ML, and programming expertise is often minimal. Consequently, improper model selection and training can compromise the quality and reliability of sensor outputs. Moreover, ML approaches typically require large, high-quality datasets, yet DL algorithms are sometimes inappropriately applied to small datasets, increasing the risk of overfitting. Another common misconception is that ML can compensate for low-quality data. While ML can extract meaningful correlations from low signal-to-noise datasets, performance is substantially improved when high-quality data are available. Generating such data generally requires measurements conducted across multiple days, instruments, reagents, and operators to capture typical sources of analytical variability. These standard practices, well established in analytical method development, enhance the robustness and translatability of ML models to practical applications. Fig. 9 presents a conceptual workflow to guide critical evaluation of whether DL is appropriate for a given plasmonic data analysis task.
In summary, the adoption of AI in plasmonic sensing is poised to reshape research practices and is likely to be needed for the translation of plasmonic technologies.545 This paradigm shift has the potential to redefine how research is conducted, for example through the emergence of self-driving laboratories, which may transition the role of the sensor scientist from predominantly hands-on experimental work to one centred on data interpretation and critical analysis. Beyond enabling hardware control and task automation in sensor design and operation, AI has already transformed data interpretation within the field of plasmonic sensing. While its use was relatively marginal less than a decade ago, a substantial proportion of recent publications now incorporate AI or ML techniques for data processing. This trend appears irreversible, and it is likely that the vast majority of future contributions to the field will rely on such methods. However, this integration must be approached rigorously. Like any analytical tool, AI and ML possess inherent limitations, and understanding these constraints is essential for their responsible and effective use. When supported by appropriate training data, rigorous controls, and thorough method validation, these approaches offer considerable opportunities to advance plasmonic sensing. As the ML field continues to evolve, researchers will need to remain attentive to emerging algorithms (such as LLM)546 and methodological advances that may further enhance sensor capabilities. Ultimately, although AI has the capacity to automate numerous tasks, human intuition, creativity, and critical reasoning will remain indispensable to the continued advancement of plasmonic sensing.
Conflicts of interest
JFM declares a conflict of interest with Affinité Instruments, a manufacturer of surface plasmon resonance instruments.Data availability
This article does not contain new data. As such, no data can be made available by the authors and the readers are referred to the authors of the referenced papers.Supplementary information (SI): reference tables are provided for the synthesis of silver nanoparticles and for applications of ML in different fields. See DOI: https://doi.org/10.1039/d5cs01522g.
Acknowledgements
The authors thank the financial support of the Natural Science and Engineering Research Council (NSERC) of Canada (grant RGPIN-2021-03114). The authors acknowledge the use of AI language polishing tools after the initial draft for some sections of the paper, which was proofread for the accuracy of the language.References
- A. Gellé, T. Jin, L. de la Garza, G. D. Price, L. V. Besteiro and A. Moores, Chem. Rev., 2020, 120, 986–1041 CrossRef PubMed.
- Z. L. Zhang, C. Y. Zhang, H. R. Zheng and H. X. Xu, Acc. Chem. Res., 2019, 52, 2506–2515 CrossRef CAS PubMed.
- W. L. Barnes, A. Dereux and T. W. Ebbesen, Nature, 2003, 424, 824–830 CrossRef CAS PubMed.
- D. K. Gramotnev and S. I. Bozhevolnyi, Nat. Photonics, 2010, 4, 83–91 CrossRef CAS.
- H. Altug, S. H. Oh, S. A. Maier and J. Homola, Nat. Nanotechnol., 2022, 17, 5–16 CrossRef CAS PubMed.
- M. Li, S. K. Cushing and N. Q. Wu, Analyst, 2015, 140, 386–406 RSC.
- S. M. Nie and S. R. Emery, Science, 1997, 275, 1102–1106 CrossRef CAS PubMed.
- K. Kneipp, Y. Wang, H. Kneipp, L. T. Perelman, I. Itzkan, R. R. Dasari and M. S. Feld, Phys. Rev. Lett., 1997, 78, 1667–1670 CrossRef CAS.
- A. B. Taylor and P. Zijlstra, ACS Sens., 2017, 2, 1103–1122 CrossRef CAS PubMed.
- N. N. Jiang, X. L. Zhuo and J. F. Wang, Chem. Rev., 2018, 118, 3054–3099 CrossRef CAS PubMed.
- Y. Q. Zhang, C. J. Min, X. J. Dou, X. Y. Wang, H. P. Urbach, M. G. Somekh and X. C. Yuan, Light Sci. Appl., 2021, 10, 59 CrossRef CAS PubMed.
- Y. J. Hang, A. Y. Wang and N. Q. Wu, Chem. Soc. Rev., 2024, 53, 2932–2971 RSC.
- J. Homola, Chem. Rev., 2008, 108, 462–493 CrossRef CAS PubMed.
- J. Homola, S. S. Yee and G. Gauglitz, Sens. Actuators, B, 1999, 54, 3–15 CrossRef CAS.
- J. N. Anker, W. P. Hall, O. Lyandres, N. C. Shah, J. Zhao and R. P. Van Duyne, Nat. Mater., 2008, 7, 442–453 CrossRef CAS PubMed.
- K. M. Mayer and J. H. Hafner, Chem. Rev., 2011, 111, 3828–3857 CrossRef CAS PubMed.
- K. S. Lee and M. A. El-Sayed, J. Phys. Chem. B, 2006, 110, 19220–19225 CrossRef CAS PubMed.
- M. C. Daniel and D. Astruc, Chem. Rev., 2004, 104, 293–346 CrossRef CAS PubMed.
- A. Loiseau, V. Asila, G. Boitel-Aullen, M. Lam, M. Salmain and S. Boujday, Biosensors, 2019, 9, 78 CrossRef CAS PubMed.
- A. Elbanna, H. Jiang, Q. D. Fu, J. F. Zhu, Y. D. Liu, M. Zhao, D. J. Liu, S. M. Lai, X. W. Chua, J. S. Pan, Z. X. Shen, L. Wu, Z. Liu, C. W. Qiu and J. H. Teng, ACS Nano, 2023, 17, 4134–4179 CrossRef CAS PubMed.
- S. Kasani, K. Curtin and N. Q. Wu, Nanophotonics, 2019, 8, 2065–2089 Search PubMed.
- X. Tang, Q. Hao, X. Y. Hou, L. L. Lan, M. Z. Li, L. Yao, X. Zhao, Z. H. Ni, X. C. Fan and T. Qiu, Adv. Mater., 2024, 36, 2312348 CrossRef CAS PubMed.
- K. Chang, Y. J. Zhao, Z. X. Xu, L. Zhu, L. H. Xu and Q. Z. Wang, Chem. Eng. J., 2023, 459, 141539 CrossRef CAS.
- L. A. Warning, A. R. Miandashti, L. A. McCarthy, Q. F. Zhang, C. F. Landes and S. Link, ACS Nano, 2021, 15, 15538–15566 CrossRef CAS PubMed.
- G. C. Zheng, J. J. He, V. Kumar, S. L. Wang, I. Pastoriza-Santos, J. Pérez-Juste, L. M. Liz-Marzán and K. Y. Wong, Chem. Soc. Rev., 2021, 50, 3738–3754 RSC.
- M. Hojjat Jodaylami, J.-F. Masson and A. Badia, Nat. Rev. Methods Primers, 2025, 5, 47 CrossRef CAS.
- X. X. Han, R. S. Rodriguez, C. L. Haynes, Y. Ozaki and B. Zhao, Nat. Rev. Methods Primers, 2022, 1, 87 CrossRef.
- J. Langer, D. J. D. Aberasturi, J. Aizpurua, R. A. Alvarez-Puebla, B. Auguié, J. J. Baumberg, G. C. Bazan, S. E. J. Bell, A. Boisen, A. G. Brolo, J. Choo, D. Cialla-May, V. Deckert, L. Fabris, K. Faulds, F. J. G. D. Abajo, R. Goodacre, D. Graham, A. J. Haes, C. L. Haynes, C. Huck, T. Itoh, M. Käll, J. Kneipp, N. A. Kotov, H. Kuang, E. C. L. Ru, H. K. Lee, J.-F. Li, X. Y. Ling, S. A. Maier, T. Mayerhöfer, M. Moskovits, K. Murakoshi, J.-M. Nam, S. Nie, Y. Ozaki, I. Pastoriza-Santos, J. Perez-Juste, J. Popp, A. Pucci, S. Reich, B. Ren, G. C. Schatz, T. Shegai, S. Schlücker, L.-L. Tay, K. G. Thomas, Z.-Q. Tian, R. P. V. Duyne, T. Vo-Dinh, Y. Wang, K. A. Willets, C. Xu, H. Xu, Y. Xu, Y. S. Yamamoto, B. Zhao and L. M. Liz-Marzán, ACS Nano, 2019, 14, 28–117 CrossRef PubMed.
- J. Yi, E.-M. You, R. Hu, D.-Y. Wu, G.-K. Liu, Z.-L. Yang, H. Zhang, Y. Gu, Y.-H. Wang, X. Wang, H. Ma, Y. Yang, J.-Y. Liu, F. R. Fan, C. Zhan, J.-H. Tian, Y. Qiao, H. Wang, S.-H. Luo, Z.-D. Meng, B.-W. Mao, J.-F. Li, B. Ren, J. Aizpurua, V. A. Apkarian, P. N. Bartlett, J. Baumberg, S. E. J. Bell, A. G. Brolo, L. E. Brus, J. Choo, L. Cui, V. Deckert, K. F. Domke, Z.-C. Dong, S. Duan, K. Faulds, R. Frontiera, N. Halas, C. Haynes, T. Itoh, J. Kneipp, K. Kneipp, E. C. L. Ru, Z.-P. Li, X. Y. Ling, J. Lipkowski, L. M. Liz-Marzán, J.-M. Nam, S. Nie, P. Nordlander, Y. Ozaki, R. Panneerselvam, J. Popp, A. E. Russell, S. Schlücker, Y. Tian, L. Tong, H. Xu, Y. Xu, L. Yang, J. Yao, J. Zhang, Y. Zhang, Y. Zhang, B. Zhao, R. Zenobi, G. C. Schatz, D. Graham and Z.-Q. Tian, Chem. Soc. Rev., 2025, 54, 1453–1551 Search PubMed.
- Y. Jeong, Y.-M. Kook, K. Lee and W.-G. Koh, Biosens. Bioelectron., 2018, 111, 102–116 CrossRef CAS PubMed.
- J. F. Li, C. Y. Li and R. F. Aroca, Chem. Soc. Rev., 2017, 46, 3962–3979 RSC.
- D. Cialla-May, A. Bonifacio, A. Markin, N. Markina, S. Fornasaro, A. Dwivedi, T. Dib, E. Farnesi, C. Liu, A. Ghosh, M. Schmitt and J. Popp, TrAC, Trends Anal. Chem., 2024, 181, 117990 Search PubMed.
- M. K. Fan, G. F. S. Andrade and A. G. Brolo, Anal. Chim. Acta, 2020, 1097, 1–29 CrossRef CAS PubMed.
- J. F. Li, Y. J. Zhang, S. Y. Ding, R. Panneerselvam and Z. Q. Tian, Chem. Rev., 2017, 117, 5002–5069 CrossRef CAS PubMed.
- C. L. Lin, Y. Y. Li, Y. S. Peng, S. Zhao, M. M. Xu, L. X. Zhang, Z. R. Huang, J. L. Shi and Y. Yang, J. Nanobiotechnol., 2023, 21, 149 CrossRef PubMed.
- M. A. Islam and J. F. Masson, ACS Sens., 2025, 10, 577–601 CrossRef CAS PubMed.
- L. Vazquez-Iglesias, G. M. S. Casagrande, D. Garcia-Lojo, L. F. Leal, T. A. Ngo, J. Perez-Juste, R. M. Reis, K. Kant and I. Pastoriza-Santos, Bioact. Mater., 2024, 34, 248–268 CAS.
- J. F. Masson, ACS Sens., 2017, 2, 16–30 CrossRef CAS PubMed.
- R. Boudries, H. Williams, S. Paquereau-Gaboreau, S. Bashir, M. H. Jodaylami, M. Chisanga, L. Trudeau and J. F. Masson, ACS Nano, 2024, 18, 22620–22647 Search PubMed.
- J. M. Chen, Y. J. Huang, P. Kannan, L. Zhang, Z. Y. Lin, J. W. Zhang, T. Chen and L. H. Guo, Anal. Chem., 2016, 88, 2149–2155 CrossRef CAS PubMed.
- S. T. R. Pang, T. X. Yang and L. L. He, TrAC, Trends Anal. Chem., 2016, 85, 73–82 CrossRef CAS.
- S. Balbinot, A. M. Srivastav, J. Vidic, I. Abdulhalim and M. Manzano, Trends Food Sci. Technol., 2021, 111, 128–140 CrossRef CAS.
- L. Y. Liu, Y. Wang, Z. H. Xue, B. Peng, X. H. Kou and Z. X. Gao, Trends Food Sci. Technol., 2024, 148, 104487 Search PubMed.
- Y. J. Hang, J. Boryczka and N. Q. Wu, Chem. Soc. Rev., 2022, 51, 329–375 RSC.
- J. F. Masson, Analyst, 2020, 145, 3776–3800 RSC.
- X. L. Zhou, Y. Z. Yang, S. P. Wang and X. W. Liu, Angew. Chem., Int. Ed., 2020, 59, 1776–1785 CrossRef CAS PubMed.
- Z. Y. Zhang, H. Wang, Z. P. Chen, X. Y. Wang, J. Choo and L. X. Chen, Biosens. Bioelectron., 2018, 114, 52–65 CrossRef CAS PubMed.
- J. W. Sun, Y. X. Lu, L. Y. He, J. W. Pang, F. Y. Yang and Y. Y. Liu, TrAC, Trends Anal. Chem., 2020, 122, 115754 CrossRef CAS.
- H. K. Lee, Y. H. Lee, C. S. L. Koh, C. P. Q. Gia, X. M. Han, C. L. Lay, H. Y. F. Sim, Y. C. Kao, Q. An and X. Y. Ling, Chem. Soc. Rev., 2019, 48, 731–756 RSC.
- H. Tao, T. Wu, M. Aldeghi, T. C. Wu, A. Aspuru-Guzik and E. Kumacheva, Nat. Rev. Mater., 2021, 6, 701–716 CrossRef.
- F. Lussier, V. Thibault, B. Charron, G. Q. Wallace and J.-F. Masson, TrAC, Trends Anal. Chem., 2020, 124, 115796 CrossRef CAS.
- D. D. Stuart, W. Van Zant, S. Valiulis, A. S. Malinick, V. Hanson and Q. Cheng, Anal. Bioanal. Chem., 2024, 416, 5221–5232 CrossRef CAS PubMed.
- H. Zhou, L. G. Xu, Z. H. Ren, J. Q. Zhu and C. K. Lee, Nanoscale Adv., 2023, 5, 538–570 RSC.
- S. Srivastava, W. Wang, W. Zhou, M. Jin and P. J. Vikesland, Environ. Sci. Technol., 2024, 58, 20830–20848 CrossRef CAS PubMed.
- X. K. Xu, D. Aggarwal and K. Shankar, Nanomaterials, 2022, 12, 633 CrossRef CAS PubMed.
- J.-F. Masson, S. J. Biggins and E. Ringe, Nat. Nanotechnol., 2023, 18, 111–123 CrossRef CAS PubMed.
- M. Chisanga and J. F. Masson, Annu. Rev. Anal. Chem., 2024, 17, 313–338 CrossRef CAS PubMed.
- Y. L. Dong, J. Y. Hu, J. L. Jin, H. B. Zhou, S. Y. Jin and D. T. Yang, TrAC, Trends Anal. Chem., 2024, 180, 117974 CrossRef CAS.
- A. Horta-Velázquez, F. Arce, E. Rodríguez-Sevilla and E. Morales-Narváez, TrAC, Trends Anal. Chem., 2023, 169, 117378 CrossRef.
- M. H. Jodaylami, O. Y. Sanvi, R. L. Rungta, A. Kolta and J.-F. Masson, ACS Nano, 2024, 18, 35075–35087 CrossRef PubMed.
- H. Mohammed, M. F. Mia, J. Wiggins and S. Desai, Molecules, 2025, 30, 883 CrossRef CAS PubMed.
- D. Pines and D. Bohm, Phys. Rev., 1952, 85, 338–353 CrossRef CAS.
- K. A. Willets and R. P. V. Duyne, Annu. Rev. Phys. Chem., 2007, 58, 267–297 CrossRef CAS PubMed.
- C. L. Haynes, A. D. McFarland and R. P. V. Duyne, Anal. Chem., 2005, 77, 338–346 CrossRef.
- H. Raether, Surface Plasmons on Smooth and Rough Surfaces and on Gratings, Springer, 1988 Search PubMed.
- J. Zhou, Y. Wang, L. Zhang and X. Li, Chin. Chem. Lett., 2018, 29, 54–60 CrossRef CAS.
- D. Barchiesi, T. Gharbi, D. Cakir, E. Anglaret, N. Fréty, S. Kessentini and R. Maâlej, Photonics, 2022, 9, 104 CrossRef CAS.
- L. T. Quynh, C.-W. Cheng, C.-T. Huang, S. S. Raja, R. Mishra, M.-J. Yu, Y.-J. Lu and S. Gwo, ACS Nano, 2022, 16, 5975–5983 CrossRef CAS PubMed.
- P. Ares and K. S. Novoselov, Nano Mater. Sci., 2022, 4, 3–9 CrossRef CAS.
- R. Alharbi, M. Irannejad and M. Yavuz, Sensors, 2019, 19, 862 CrossRef PubMed.
- D. T. Nurrohman and N.-F. Chiu, Nanomaterials, 2021, 11, 216 CrossRef CAS PubMed.
- L. Wu, H. S. Chu, W. S. Koh and E. P. Li, Opt. Express, 2010, 18, 14395–14400 CrossRef CAS PubMed.
- P. Pillai, B. Rai and P. Pal, Sci. Rep., 2023, 13, 3536 CrossRef CAS PubMed.
- R. H. Ritchie, Phys. Rev., 1957, 106, 874–881 CrossRef CAS.
- A. Otto, Z. Phys. A:Hadrons Nucl., 1968, 216, 398–410 CrossRef CAS.
- E. R. Kretschmann and H. Notizen, Z. Naturforsch., 1968, 23, 2135–2136 CrossRef CAS.
- N. Shukla, P. Chetri, R. Boruah, A. Gogoi and G. A. Ahmed, in Recent Advances in Plasmonic Probes, Lecture Notes in Nanoscale Science and Technology, ed. R. Biswas and N. Mazumder, Springer, Cham, 2022, pp. 191–222 Search PubMed.
- B. D. Gupta and R. K. Verma, J. Sens., 2009, 979761 CrossRef.
- M. Piliarik, H. Vaisocherová and J. Homola, Biosens. Biodetect., 2009, 65–88, DOI:10.1007/978-1-60327-567-5_5.
- B. Prabowo, A. Purwidyantri and K.-C. Liu, Biosensors, 2018, 8, 80 CrossRef CAS PubMed.
- V. Yesudasu, H. S. Pradhan and R. J. Pandya, Heliyon, 2021, 7, e06321 CrossRef CAS PubMed.
- D.-S. Wang and S.-K. Fan, Sensors, 2016, 16, 1175 CrossRef PubMed.
- D. Murugan, M. Tintelott, M. S. Narayanan, X.-T. Vu, T. Kurkina, C. Rodriguez-Emmenegger, U. Schwaneberg, J. Dostalek, S. Ingebrandt and V. Pachauri, Adv. Opt. Mater., 2024, 12, 2401862 CrossRef CAS.
- M. Couture, S. S. Zhao and J.-F. Masson, Phys. Chem. Chem. Phys., 2013, 15, 11190 RSC.
- D. Capelli, V. Scognamiglio and R. Montanari, TrAC, Trends Anal. Chem., 2023, 163, 117079 CrossRef CAS.
- R. B. M. Schasfoort, in Handbook of Surface Plasmon Resonance, ed. R. B. M. Schasfoort, The Royal Society of Chemistry, 2nd edn, 2017, ch. 1, pp. 1–26 Search PubMed.
- X. Caide and S.-F. Sui, Sens. Actuators, B, 2000, 66, 174–177 CrossRef CAS.
- Z. Huo, Y. Li, B. Chen, W. Zhang, X. Yang and X. Yang, Talanta, 2023, 255, 124213 CrossRef CAS PubMed.
- Q. Wang, D. Zhang, Y. Qian, X. Yin, L. Wang, S. Zhang and Y. Wang, Photonic Sens., 2024, 14, 240201 CrossRef.
- H.-M. Kim, H.-J. Kim, J.-H. Park and S.-K. Lee, Anal. Chim. Acta, 2022, 1213, 339960 CrossRef CAS PubMed.
- K. L. Kelly, E. Coronado, L. L. Zhao and G. C. Schatz, J. Phys. Chem. B, 2003, 107, 668–677 CrossRef CAS.
- A. J. Haes, C. L. Haynes, A. D. McFarland, G. C. Schatz, R. P. Van Duyne and S. Zou, MRS Bull., 2005, 30, 368–375 CrossRef CAS.
- P. L. Stiles, J. A. Dieringer, N. C. Shah and R. P. Van Duyne, Annu. Rev. Anal. Chem., 2008, 1, 601–626 CrossRef CAS PubMed.
- M. E. Stewart, C. R. Anderton, L. B. Thompson, J. Maria, S. K. Gray, J. A. Rogers and R. G. Nuzzo, Chem. Rev., 2008, 108, 494–521 CrossRef CAS PubMed.
- G. V. Naik, V. M. Shalaev and A. Boltasseva, Adv. Mater., 2013, 25, 3264–3294 CrossRef CAS PubMed.
- J.-F. Li, Z.-L. Yang, B. Ren, G.-K. Liu, P.-P. Fang, Y.-X. Jiang, D.-Y. Wu and Z.-Q. Tian, Langmuir, 2006, 22, 10372–10379 CrossRef CAS PubMed.
- Z.-Q. Tian and B. Ren, Annu. Rev. Phys. Chem., 2004, 55, 197–229 CrossRef CAS PubMed.
- K. M. Kosuda, J. M. Bingham, K. L. Wustholz, R. P. Van Duyne and R. J. Groarke, in Comprehensive Nanoscience and Nanotechnology, ed. D. L. Andrews, R. H. Lipson and T. Nann, Academic Press, Oxford, 2nd edn, 2016, pp. 117–152 DOI:10.1016/B978-0-12-803581-8.00611-1.
- B. C. Mei, E. Oh, K. Susumu, D. Farrell, T. J. Mountziaris and H. Mattoussi, Langmuir, 2009, 25, 10604–10611 CrossRef CAS PubMed.
- H. Kang, J. T. Buchman, R. S. Rodriguez, H. L. Ring, J. He, K. C. Bantz and C. L. Haynes, Chem. Rev., 2019, 119, 664–699 CrossRef CAS PubMed.
- L. M. Liz-Marzán, Langmuir, 2006, 22, 32–41 CrossRef PubMed.
- M. Rycenga, C. M. Cobley, J. Zeng, W. Li, C. H. Moran, Q. Zhang, D. Qin and Y. Xia, Chem. Rev., 2011, 111, 3669–3712 CrossRef CAS PubMed.
- N. D. Burrows, W. Lin, J. G. Hinman, J. M. Dennison, A. M. Vartanian, N. S. Abadeer, E. M. Grzincic, L. M. Jacob, J. Li and C. J. Murphy, Langmuir, 2016, 32, 9905–9921 CrossRef CAS PubMed.
- I. B. Becerril-Castro, I. Calderon, N. Pazos-Perez, L. Guerrini, F. Schulz, N. Feliu, I. Chakraborty, V. Giannini, W. J. Parak and R. A. Alvarez-Puebla, Anal. Sens., 2022, 2, e202200005 CAS.
- J. Turkevich, P. C. Stevenson and J. Hillier, Discuss. Faraday Soc., 1951, 11, 55–75 RSC.
- M. Wuithschick, A. Birnbaum, S. Witte, M. Sztucki, U. Vainio, N. Pinna, K. Rademann, F. Emmerling, R. Kraehnert and J. Polte, ACS Nano, 2015, 9, 7052–7071 CrossRef CAS PubMed.
- C. Deraedt, L. Salmon, S. Gatard, R. Ciganda, R. Hernandez, J. Ruiz and D. Astruc, Chem. Commun., 2014, 50, 14194–14196 RSC.
- N. G. Bastús, J. Comenge and V. Puntes, Langmuir, 2011, 27, 11098–11105 CrossRef PubMed.
- H.-L. Wu, C.-H. Kuo and M. H. Huang, Langmuir, 2010, 26, 12307–12313 CrossRef CAS PubMed.
- C.-J. Huang, Y.-H. Wang, P.-H. Chiu, M.-C. Shih and T.-H. Meen, Mater. Lett., 2006, 60, 1896–1900 CrossRef CAS.
- J. E. Millstone, G. S. Métraux and C. A. Mirkin, Adv. Funct. Mater., 2006, 16, 1209–1214 CrossRef CAS.
- T. H. Ha, H.-J. Koo and B. H. Chung, J. Phys. Chem. C, 2007, 111, 1123–1130 CrossRef CAS.
- N. R. Jana, L. Gearheart and C. J. Murphy, Adv. Mater., 2001, 13, 1389–1393 CrossRef CAS.
- J. Gao, C. M. Bender and C. J. Murphy, Langmuir, 2003, 19, 9065–9070 CrossRef CAS.
- B. Nikoobakht and M. A. El-Sayed, Chem. Mater., 2003, 15, 1957–1962 CrossRef CAS.
- M. S. Verma, P. Z. Chen, L. Jones and F. X. Gu, RSC Adv., 2014, 4, 10660–10668 RSC.
- C. Song, F. Li, X. Guo, W. Chen, C. Dong, J. Zhang, J. Zhang and L. Wang, J. Mater. Chem. B, 2019, 7, 2001–2008 RSC.
- S. Atta, M. Beetz and L. Fabris, Nanoscale, 2019, 11, 2946–2958 RSC.
- R. M. Pallares, T. Stilson, P. Choo, J. Hu and T. W. Odom, ACS Appl. Nano Mater., 2019, 2, 5266–5271 CrossRef.
- H. Yuan, C. G. Khoury, H. Hwang, C. M. Wilson, G. A. Grant and T. Vo-Dinh, Nanotechnology, 2012, 23, 075102 CrossRef PubMed.
- Y. Liu, L. Yang and Y. Shen, J. Mater. Res., 2018, 33, 2671–2679 CrossRef CAS.
- Y. Yu, Y. Xie, P. Zeng, D. Zhang, R. Liang, W. Wang, Q. Ou and S. Zhang, Nanomaterials, 2019, 9, 1202 CrossRef CAS PubMed.
- H. P. Liang, L. J. Wan, C. L. Bai and L. Jiang, J. Phys. Chem. B, 2005, 109, 7795–7800 CrossRef CAS PubMed.
- L. M. Liz-Marzán, M. Giersig and P. Mulvaney, Langmuir, 1996, 12, 4329–4335 CrossRef.
- S. Cong, X. Liu, Y. Jiang, W. Zhang and Z. Zhao, Innovation, 2020, 1, 100051 CAS.
- R. C. Maher, in Raman Spectroscopy for Nanomaterials Characterization, ed. C. S. S. R. Kumar, Springer-Verlag, Berlin Heidelberg, 2012, ch. 10, pp. 215–260 Search PubMed.
- Y. Fang, N.-H. Seong and D. D. Dlott, Science, 2008, 321, 388–392 CrossRef CAS PubMed.
- N. C. Lindquist, A. T. Bido and A. G. Brolo, J. Phys. Chem. C, 2022, 126, 7117–7126 CrossRef CAS.
- M. M. Schmidt, A. G. Brolo and N. C. Lindquist, ACS Nano, 2024, 18, 25930–25938 CAS.
- Y. Zou, H. Jin, Q. Ma, Z. Zheng, S. Weng, K. Kolataj, G. Acuna, I. Bald and D. Garoli, Nanoscale, 2025, 17, 3656–3670 RSC.
- M. Hardy and P. G. Oppenheimer, Nanoscale, 2024, 16, 3293–3323 RSC.
- E. C. L. Ru, C. Galloway and P. G. Etchegoin, Phys. Chem. Chem. Phys., 2006, 8, 3083–3087 RSC.
- R. Gautam, S. Vanga, F. Ariese and S. Umapathy, EPJ Tech. Instrum., 2015, 2, 8 CrossRef.
- K. R. Khondakar, H. Mazumdar, S. Das and A. Kaushik, Adv. Colloid Interface Sci., 2025, 344, 103594 CrossRef CAS PubMed.
- S. E. J. Bell, G. Charron, E. Cortés, J. Kneipp, M. L. D. L. Chapelle, J. Langer, M. Procházka, V. Tran and S. Schlücker, Angew. Chem., Int. Ed., 2020, 59, 5454–5462 CrossRef CAS PubMed.
- Y. Xu, W. Aljuhani, Y. Zhang, Z. Ye, C. Li and S. E. J. Bell, Chem. Soc. Rev., 2025, 54, 62–84 RSC.
- M. Fleischmann, P. J. Hendra and A. J. McQuillan, Chem. Phys. Lett., 1974, 26, 163–166 CrossRef CAS.
- C. L. Brosseau, A. Colina, J. V. Perales-Rondon, A. J. Wilson, P. B. Joshi, B. Ren and X. Wang, Nat. Rev. Methods Primers, 2023, 3, 79 CrossRef CAS.
- A. Lipovka, M. Fatkullin, A. Averkiev, M. Pavlova, A. Adiraju, S. Weheabby, A. Al-Hamry, O. Kanoun, I. Pašti and T. Lazarevic-Pasti, Crit. Rev. Anal. Chem., 2024, 54, 110–134 CrossRef CAS PubMed.
- R. Moldovan, E. Vereshchagina, K. Milenko, B.-C. Iacob, A. E. Bodoki, A. Falamas, N. Tosa, C. M. Muntean, C. Farcău and E. Bodoki, Anal. Chim. Acta, 2022, 1209, 339250 CrossRef CAS PubMed.
- C. E. Ott, A. Burns, E. Sisco and L. E. Arroyo, Forensic Chem., 2023, 34, 100492 CrossRef CAS.
- M. Haroon, I. Abdulazeez, T. A. Saleh and A. A. Al-Saadi, Electrochim. Acta, 2021, 387, 138463 CrossRef CAS.
- B. Greene, D. Alhatab, C. Pye and C. Brosseau, J. Phys. Chem. C, 2017, 121, 8084–8090 CrossRef CAS.
- H. Zhou and J. Kneipp, Vib. Spectrosc., 2024, 131, 103661 CrossRef CAS.
- M. Chen, G. Liu, L. Wang, A. Zhang, Z. Yang, X. Li, Z. Zhang, S. Gu, D. Cui and H. Haick, Anal. Chem., 2025, 97, 4397–4406 CrossRef CAS PubMed.
- T. Jones, D. Zhou, J. Liu, I. P. Parkin and T.-C. Lee, J. Mater. Chem. B, 2024, 12, 10563–10572 RSC.
- X. Wang, Y. Yan, H. Zhou, X. Liu, R. Zhang, D. Wang, P. Schaaf, G. Guo and X. Wang, Small, 2025, 21, e03894 CrossRef CAS PubMed.
- X. Zhang, Y. Wang, Y. Zhang, K. Zhang, R. Chu, F. Zhang and X. Zhao, J. Raman Spectrosc., 2025, 56, 165–175 CrossRef CAS.
- J. Zhang, Y.-X. Yuan, J.-W. Yan, B.-W. Mao, J.-L. Yao and Z.-Q. Tian, ACS Appl. Mater. Interfaces, 2024, 16, 50054–50060 CrossRef CAS PubMed.
- M. E. Blaha, J. Schwieger, R. Warias, A. Das, M. Polack and D. Belder, Anal. Chem., 2025, 97, 13628–13636 CrossRef CAS PubMed.
- T. Cooman, C. E. Ott and L. E. Arroyo, J. Forensic Sci., 2023, 68, 1520–1526 CrossRef CAS PubMed.
- D. Martín-Yerga, A. Pérez-Junquera, M. B. González-García, J. V. Perales-Rondon, A. Heras, A. Colina, D. Hernández-Santos and P. Fanjul-Bolado, Electrochim. Acta, 2018, 264, 183–190 CrossRef.
- Y. Gao, D. E. Aspnes and S. Franzen, J. Phys. Chem. A, 2022, 126, 341–351 CrossRef CAS PubMed.
- D. Kavungal, E. Morro, S. T. Kumar, B. Dagli, H. A. Lashuel and H. Altug, Adv. Sci., 2025, 12, e00320 CrossRef CAS PubMed.
- M. Wagner, A. Seifert and L. M. Liz-Marzán, Nanoscale Horiz., 2022, 7, 1259–1278 RSC.
- J. Brandt, K. Mattsson and M. Hassellöv, Anal. Chem., 2021, 93, 16360–16368 CrossRef CAS PubMed.
- A. Tittl, A. John-Herpin, A. Leitis, E. R. Arvelo and H. Altug, Angew. Chem., Int. Ed., 2019, 58, 14810–14822 CrossRef CAS PubMed.
- K. Aslan, I. Gryczynski, J. Malicka, E. Matveeva, J. R. Lakowicz and C. D. Geddes, Curr. Opin. Biotechnol, 2005, 16, 55–62 CrossRef CAS PubMed.
- M. Ganguly, C. Mondal, J. Chowdhury, J. Pal, A. Pal and T. Pal, Dalton Trans., 2014, 43, 1032–1047 RSC.
- D. Semeniak, D. F. Cruz, A. Chilkoti and M. H. Mikkelsen, Adv. Mater., 2023, 35, 2107986 CrossRef CAS PubMed.
- S. M. Sundaresan, S. M. Fothergill, T. A. Tabish, M. Ryan and F. Xie, Appl. Phys. Rev., 2021, 8, 041311 CAS.
- M. Wang, M. Wang, G. Zheng, Z. Dai and Y. Ma, Nanoscale Adv., 2021, 3, 2448–2465 RSC.
- C. D. Geddes, Phys. Chem. Chem. Phys., 2013, 15, 19537 RSC.
- Z. Yan, W. Yao, K. Mai, J. Huang, Y. Wan, L. Huang, B. Cai and Y. Liu, RSC Adv., 2022, 12, 8202–8210 RSC.
- D. Hong, E.-J. Jo, D. Bang, C. Jung, Y. E. Lee, Y.-S. Noh, M. G. Shin and M.-G. Kim, ACS Nano, 2023, 17, 16607–16619 CrossRef CAS PubMed.
- D.-D. Xu, B. Zheng, C.-Y. Song, Y. Lin, D.-W. Pang and H.-W. Tang, Sens. Actuators, B, 2019, 282, 650–658 CrossRef CAS.
- D. D. Xu, Y. L. Deng, C. Y. Li, Y. Lin and H. W. Tang, Biosens. Bioelectron., 2017, 87, 881–887 CrossRef CAS PubMed.
- Z. Mei and L. Tang, Anal. Chem., 2017, 89, 633–639 CrossRef CAS PubMed.
- E. Lucas, R. Knoblauch, M. Combs-Bosse, S. E. Broedel and C. D. Geddes, Spectrochim. Acta, Part A, 2020, 228, 117739 CrossRef CAS PubMed.
- K. Wang, J. Liao, X. Yang, M. Zhao, M. Chen, W. Yao, W. Tan and X. Lan, Biosens. Bioelectron., 2015, 63, 172–177 CrossRef CAS PubMed.
- X. Li, T. Kuznetsova, N. Cauwenberghs, M. Wheeler, H. Maecker, J. C. Wu, F. Haddad and H. Dai, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 7089–7094 CrossRef CAS PubMed.
- D.-D. Xu, C. Liu, C.-Y. Li, C.-Y. Song, Y.-F. Kang, C.-B. Qi, Y. Lin, D.-W. Pang and H.-W. Tang, ACS Appl. Mater. Interfaces, 2017, 9, 37606–37614 CrossRef CAS PubMed.
- C. Han, R. Chen, X. Wu, N. Shi, T. Duan, K. Xu and T. Huang, Anal. Chim. Acta, 2021, 1187, 339160 CrossRef CAS PubMed.
- R. M. Khozani, S. Abbasi-Moayed and M. R. Hormozi-Nezhad, Chemosphere, 2024, 357, 141966 CrossRef CAS PubMed.
- T. Pang, X. Tao, P. Zhuang, M. Wu, J. Li, H. Huang, J. Sun and J. Liu, J. Fluoresc., 2025, 35, 9761–9771 CrossRef CAS PubMed.
- J. Chen, S. Gao, L. Fan, Y. Wang, Z. Xiao, R. Shen, J. Xu, H. Cui and G. Xu, Talanta, 2026, 298, 128829 CrossRef CAS PubMed.
- K. J. Squire, Y. Zhao, A. Tan, K. Sivashanmugan, J. A. Kraai, G. L. Rorrer and A. X. Wang, Sens. Actuators, B, 2019, 290, 118–124 CrossRef CAS PubMed.
- R. Houhou and T. Bocklitz, Anal. Sci. Adv., 2021, 2, 128–141 CrossRef PubMed.
- I. Goodfellow, Y. Bengio and A. Courville, Deep Learning, MIT Press, 2016 Search PubMed.
- Y.-F. Shi, Z.-X. Yang, S. Ma, P.-L. Kang, C. Shang, P. Hu and Z.-P. Liu, Engineering, 2023, 27, 70–83 CrossRef CAS.
- R. T. Trevor Hastie and J. Friedman, The Elements of Statistical Learning, 2009.
- A. Burkov, The Hundred-Page Machine Learning Book, 2019.
- M. Liu, T. Wang, Q. Zhang, C. Pan, S. Liu, Y. Chen, D. Lin and S. Feng, Anal. Methods, 2024, 16, 846–855 RSC.
- D. Chen, Y. Li, Z. Tan, Z.-X. Huang, J. Zong and Q.-F. Li, Int. Dairy J., 2019, 96, 132–137 CrossRef CAS.
- S. O'Hagan, W. B. Dunn, M. Brown, J. D. Knowles and D. B. Kell, Anal. Chem., 2004, 77, 290–303 CrossRef PubMed.
- X. Chen and H. Lv, NPG Asia Mater., 2022, 14, 69 CrossRef.
- W. Huyer and A. Neumaier, ACM Trans. Math. Softw., 2008, 35, 9 CrossRef.
- B. L. Hall, C. J. Taylor, R. Labes, A. F. Massey, R. Menzel, R. A. Bourne and T. W. Chamberlain, Chem. Commun., 2021, 57, 4926–4929 RSC.
- E. Kocer, T. W. Ko and J. Behler, Annu. Rev. Phys. Chem., 2022, 73, 163–186 CrossRef CAS PubMed.
- M. Nielsen, Neural Networks and Deep Learning, 2019.
- A. Géron, Hands-On Machine Learning with Scikit-Learn, Keras, and TensorFlow, O'Reilly Media, 2017 Search PubMed.
- D. Lafuente, B. Cohen, G. Fiorini, A. A. García, M. Bringas, E. Morzan and D. Onna, J. Chem. Educ., 2021, 98, 2892–2898 CrossRef CAS.
- A. I. Cowen-Rivers, W. Lyu, R. Tutunov, Z. Wang, A. Grosnit, R. R. Griffiths, A. M. Maraval, H. Jianye, J. Wang, J. Peters and H. Bou-Ammar, J. Artif. Intell. Res., 2022, 74, 1269–1349 CrossRef.
- I. V. Tetko, D. J. Livingstone and A. I. Luik, J. Chem. Inf. Comput. Sci., 2002, 35, 826–833 CrossRef.
- M. Höge, T. Wöhling and W. Nowak, Water Resour. Res., 2018, 54, 1688–1715 CrossRef.
- H. Das and M. Sharma, ACS Appl. Electron. Mater., 2025, 7, 5757–5787 CrossRef CAS.
- K. Guo, Z. Yang, C.-H. Yu and M. J. Buehler, Mater. Horiz., 2021, 8, 1153–1172 RSC.
- X. Pengcheng, M. Yingying, L. Wencong, L. Minjie, Z. Wenyue and D. Zhilong, J. Mater. Inf., 2025, 5, 26 Search PubMed.
- C. A. Mirkin, R. L. Letsinger, R. C. Mucic and J. J. Storhoff, Nature, 1996, 382, 607–609 CrossRef CAS PubMed.
- J.-F. Masson, G. Q. Wallace, J. Asselin, A. Ten, M. H. Jodaylami, K. Faulds, D. Graham, J. S. Biggins and E. Ringe, ACS Appl. Mater. Interfaces, 2023, 15, 46181–46194 CrossRef CAS PubMed.
- N. A. Hatab, C.-H. Hsueh, A. L. Gaddis, S. T. Retterer, J.-H. Li, G. Eres, Z. Zhang and B. Gu, Nano Lett., 2010, 10, 4952–4955 CrossRef CAS PubMed.
- S. Lu, Q. Zhou, X. Chen, Z. Song and J. Wang, Natl. Sci. Rev., 2022, 9, nwac111 CrossRef PubMed.
- R. Koziol, M. Lapinski, P. Syty, W. Sadowski, J. E. Sienkiewicz, B. Nurek, V. A. Maraloiu and B. Koscielska, Appl. Surf. Sci., 2021, 567, 150802 CrossRef CAS.
- Q. Du, Q. Zhang and G. H. Liu, Nanotechnology, 2021, 32, 505607 CrossRef CAS PubMed.
- A. Amirjani and S. K. Sadrnezhaad, J. Mater. Chem. C, 2021, 9, 9791–9819 RSC.
- L. S. P. Maia, D. A. Barroso, A. B. Silveira, W. F. Oliveira, A. Galembeck, C. A. R. Fernandes, D. G. C. Bandeira, B. Cluzel, A. R. Alexandria and G. F. Guimaraes, Photonics, 2025, 12, 572 CrossRef CAS.
- Y. Blechman, S. Tsesses, M. Feinstein, G. Bartal and E. Almeida, ACS Photonics, 2023, 10, 2494–2501 CrossRef CAS.
- J. He, C. He, C. Zheng, Q. Wang and J. Ye, Nanoscale, 2019, 11, 17444–17459 RSC.
- X. B. Wang, Y. Cao, F. Feng, Z. Y. Wang and Y. C. Cao, J. Appl. Phys., 2025, 137, 063102 CrossRef CAS.
- B. Madika, A. Saha, C. Kang, B. Buyantogtokh, J. Agar, C. M. Wolverton, P. Voorhees, P. Littlewood, S. Kalinin and S. Hong, ACS Nano, 2025, 19, 27116–27158 CrossRef CAS PubMed.
- R. L. Johnston, Dalton Trans., 2003, 4193–4207, 10.1039/B305686D.
- Y. K. Liao, Y. S. Lai, F. Pan and Y. H. Su, J. Mater. Chem. A, 2023, 11, 11187–11201 RSC.
- S. C. Yen, Y. L. Chen and Y. H. Su, APL Mater., 2020, 8, 091109 Search PubMed.
- Q. Y. Wu, Y. H. Xu, J. X. Zhao, Y. M. Liu and Z. W. Liu, Nano Lett., 2024, 24, 11581–11589 CrossRef CAS PubMed.
- J. H. Han, Y. C. Lim, R. M. Kim, J. Lv, N. H. Cho, H. Kim, S. D. Namgung, S. W. Im and K. T. Nam, ACS Nano, 2023, 17, 2306–2317 Search PubMed.
- F. A. A. Nugroho, P. Bai, I. Darmadi, G. W. Castellanos, J. Fritzsche, C. Langhammer, J. G. Rivas and A. Baldi, Nat. Commun., 2022, 13, 5737 CrossRef CAS PubMed.
- C. Liu, J. L. Zhang, Y. P. Zhao and B. Ai, Adv. Intell. Syst., 2023, 5, 2300121 CrossRef.
- J. Baxter, A. Calà Lesina, J. M. Guay, A. Weck, P. Berini and L. Ramunno, Sci. Rep., 2019, 9, 8074 CrossRef PubMed.
- Y. Q. Chen, Y. Q. Hu, J. Y. Zhao, Y. S. Deng, Z. L. Wang, X. Cheng, D. Y. Lei, Y. B. Deng and H. G. Duan, Adv. Funct. Mater., 2020, 30, 2000642 CrossRef CAS.
- Z. Zeng, P. K. Venuthurumilli and X. F. Xu, ACS Photonics, 2021, 8, 1489–1496 CrossRef CAS.
- M. M. Schmidt, E. A. Farley, M. A. Engevik, T. N. Adelsman, A. Tuckmantel Bido, N. D. Lemke, A. G. Brolo and N. C. Lindquist, ACS Nano, 2023, 17, 6675–6686 CrossRef CAS PubMed.
- S. He, W. Zhang, L. Liu, Y. Huang, J. He, W. Xie, P. Wu and C. Du, Anal. Methods, 2014, 6, 4402–4407 RSC.
- T. Chen, Y. Son, C. Dong and S.-J. Baek, Analyst, 2025, 150, 2653–2660 RSC.
- M. Kazemzadeh, M. Martinez-Calderon, W. Xu, L. W. Chamley, C. L. Hisey and N. G. Broderick, Anal. Chem., 2022, 94, 12907–12918 CrossRef CAS PubMed.
- C. Yan, iScience, 2025, 28, 112759 CrossRef PubMed.
- L. Pan, P. Pipitsunthonsan, P. Zhang, C. Daengngam, A. Booranawong and M. Chongcheawchamnan, in 13th International Symposium on Computational Intelligence and Design (ISCID), Hangzhou, China, 2020, pp. 159–163.
- M. Han, Y. Dang and J. Han, Sensors, 2024, 24, 3161 CrossRef PubMed.
- J.-W. Tang, Q. Yuan, L. Zhang, B. J. Marshall, A. C. Y. Tay and L. Wang, TrAC, Trends Anal. Chem., 2025, 184, 118135 CrossRef CAS.
- Y. Zeng, Z.-q Liu, X.-g Fan and X. Wang, Microchem. J., 2023, 191, 108777 CrossRef CAS.
- M. Lionts, E. Haugen, A. Mahadevan-Jansen and Y. Huo, in Proceedings Volume 13118. Emerging Topics in Artificial Intelligence, SPIE, San Diego, 2024.
- C. C. Horgan, M. Jensen, A. Nagelkerke, J.-P. St-Pierre, T. Vercauteren, M. M. Stevens and M. S. Bergholt, Anal. Chem., 2021, 93, 15850–15860 CrossRef CAS PubMed.
- M. T. Gebrekidan, C. Knipfer and A. S. Braeuer, J. Raman Spectrosc., 2021, 52, 723–736 CrossRef CAS.
- S. Wu, Y. Zhang, C. He, Z. Luo, Z. Chen and J. Ye, Anal. Chem., 2024, 96, 17476–17485 CrossRef CAS PubMed.
- G. Yang, H. Xiao, H. Gao, B. Zhang, W. Hu, C. Chen, Q. Qiao, G. Zhang, S. Feng, D. Liu, Y. Wang, J. Jiang and Y. Luo, J. Am. Chem. Soc., 2024, 146, 28491–28499 CAS.
- S.-H. Luo, X. Wang, G.-Y. Chen, Y. Xie, W.-H. Zhang, Z.-F. Zhou, Z.-M. Zhang, B. Ren, G.-K. Liu and Z.-Q. Tian, Anal. Chem., 2021, 93, 8408–8413 CrossRef CAS PubMed.
- C. Bench, M. S. Bergholt and M. A. Al-Badri, arXiv, 2023, preprint, arXiv:2307.00513 DOI:10.48550/2307.00513.
- X. Ma, K. Wang, K. C. Chou, Q. Li and X. Lu, Anal. Chem., 2022, 94, 577–582 Search PubMed.
- S.-H. Luo, S.-Q. Pan, G.-Y. Chen, Y. Xie, B. Ren, G.-K. Liu and Z.-Q. Tian, Anal. Chem., 2024, 96, 4086–4092 CrossRef CAS PubMed.
- S.-H. Luo, X.-J. Zhao, M.-F. Cao, J. Xu, W.-L. Wang, X.-Y. Lu, Q.-T. Huang, X.-X. Yue, G.-K. Liu, L. Yang, B. Ren and Z.-Q. Tian, Anal. Chem., 2024, 96, 6550–6557 CrossRef CAS PubMed.
- J. Wang, Y. Sun, Y. Yang, C. Zhang, W. Zheng, C. Wang, W. Zhang, L. Zhou, H. Yu and J. Li, Adv. Sci., 2025, 12, e2407432 CrossRef PubMed.
- C. A. Lieber and A. Mahadevan-Jansen, Appl. Spectrosc., 2003, 57, 1363–1367 CrossRef CAS PubMed.
- Y. Hu, T. Jiang, A. Shen, W. Li, X. Wang and J. Hu, Chemom. Intell. Lab. Syst., 2007, 85, 94–101 CrossRef CAS.
- S.-J. Baek, A. Park, Y.-J. Ahn and J. Choo, Analyst, 2015, 140, 250–257 RSC.
- F. Zhang, X. Tang, A. Tong, B. Wang, J. Wang, Y. Lv, C. Tang and J. Wang, Spectrosc. Lett., 2020, 53, 222–233 CrossRef CAS.
- J. Zhao, T. Woznicki and K. Kusnierek, Spectrochim. Acta, Part A, 2025, 330, 125679 CrossRef CAS PubMed.
- P. D. Aradhye, S. Mandal, R. D. Gray and C. J. Campbell, Anal. Chem., 2025, 97, 26708–26719 CrossRef CAS PubMed.
- S. Dong, Y. Liu, H. Yu, Y. Wang and J. Wu, Appl. Spectrosc., 2024, 78, 111–119 CrossRef CAS PubMed.
- Y. Liu, J. Wu, Y. Wang and S. Dong, AIP Adv., 2022, 12, 085212 CrossRef CAS.
- J. Cheng, L. Yu, S. Tian, X. Lv and Z. Zhang, Spectroscopy, 2022, 37, 18–27 Search PubMed.
- S. Helal, H. Sarieddeen, H. Dahrouj, T. Y. Al-Naffouri and M.-S. Alouini, IEEE Signal Process. Mag, 2022, 39, 42–62 Search PubMed.
- P. Freire, E. Manuylovich, J. E. Prilepsky and S. K. Turitsyn, Adv. Optics Photonics, 2023, 15, 739–834 Search PubMed.
- S. B. Belhaouari, A. Talbi, M. Elgamal, K. A. Elmagarmid, S. Ghannoum, Y. Yang, Y. Zhao, S. M. Zughaier and H. Bensmail, Heliyon, 2025, 11, e42550 CrossRef CAS PubMed.
- T. A. Saifuzzaman, K. Y. Lee, A. R. M. Radzol, P. S. Wong and I. Looi, in 42nd Annual International Conference of the IEEE Engineering in Medicine & Biology Society, Montreal, 2020.
- X. Fan, W. Ming, H. Zeng, Z. Zhang and H. Lu, Analyst, 2019, 144, 1789–1798 RSC.
- Y. Lin, Q. Zhang, H. Chen, S. Liu, K. Peng, X. Wang, L. Zhang, J. Huang, X. Yan, X. Lin, U. M. D. Hasan, M. Sarwara, F. Fu, S. Feng and C. Wang, BMC Med., 2025, 23, 97 CrossRef CAS PubMed.
- C.-S. Ho, N. Jean, C. A. Hogan, L. Blackmon, S. S. Jeffrey, M. Holodniy, N. Banaei, A. A. E. Saleh, S. Ermon and J. Dionne, Nat. Commun., 2019, 10, 4927 CrossRef PubMed.
- A. Nyamdavaa, K. Kaladharan, E.-O. Ganbold, S. Jeong, S. Paek, Y. Su, F.-G. Tseng and T.-O. Ishdorj, Sci. Rep., 2025, 15, 12245 Search PubMed.
- J. Yang, X. Chen, C. Luo, Z. Li, C. Chen, S. Han, X. Lv, L. Wu and C. Chen, Sci. Rep., 2023, 13, 15719 CrossRef CAS PubMed.
- H. Kang, J. Lee, S. H. Lee, J. Jeon, C. Mun, J.-Y. Yang, D. Seo, H.-J. Kwon, I.-C. Lee, S. Kim, E.-K. Lim, J. Jung, Y. Jung, S.-G. Park, S. Ryu and T. Kang, Biosens. Bioelectron., 2025, 289, 117891 CrossRef CAS PubMed.
- C.-L. Chin, C.-E. Chang and L. Chao, ACS Sens., 2025, 10, 2652–2666 Search PubMed.
- X.-Y. Lu, H.-P. Wu, H. Ma, H. Li, J. Li, Y.-T. Liu, Z.-Y. Pan, Y. Xie, L. Wang, B. Ren and G.-K. Liu, Anal. Chem., 2024, 96, 7959–7975 CrossRef CAS PubMed.
- T. Hu, Z. Zou, B. Li, T. Zhu, S. Gu, J. Jiang, Y. Luo and W. Hu, J. Am. Chem. Soc., 2025, 147, 27525–27536 CrossRef CAS PubMed.
- E. X. Tan, J. R. T. Chen, D. W. C. Pang, N. S. Tan, I. Y. Phang and X. Y. Ling, Angew. Chem., 2025, e202508717 CAS.
- J. Hu, Y. Zou, B. Sun, X. Yu, Z. Shang, J. Huang, S. Jin and P. Liang, Spectrochim. Acta, Part A, 2022, 265, 120366 Search PubMed.
- C. Pan, K. Peng, T. Chen, G. Chen, Y. Lin, Q. Zhang, M. Liu, D. Lin, T. Wang and S. Feng, Adv. Intell. Syst., 2023, 5, 2300006 CrossRef.
- D. Zhu, PhD thesis, Georgia Institute of Technology, 2023.
- S. M. Quarin, D. Vang, R. I. Dima, G. Stan and P. Strobbia, npj Biosens., 2025, 2, 9 Search PubMed.
- H. Chen, H. Liu, L. Xing, D. Fan, N. Chen, P. Ma and X. Zhang, ACS Sens., 2025, 10, 2872–2882 CrossRef CAS PubMed.
- A. Hegde, M. Hajikhani, J. Snyder, J. Cheng and M. Lin, ACS Appl. Mater. Interfaces, 2024, 17, 2018–2031 Search PubMed.
- X. Bi, X. Ai, Z. Wu, L. L. Lin, Z. Chen and J. Ye, Anal. Chem., 2025, 97, 6826–6846 Search PubMed.
- S.-h Luo, W.-l Wang, Z.-f Zhou, Y. Xie, B. Ren, G.-k Liu and Z.-q Tian, Anal. Chem., 2022, 94, 10151–10158 CrossRef CAS PubMed.
- M. Kim, S. Huh, H. J. Park, S. H. Cho, M.-Y. Lee, S. Jo and Y. S. Jung, Biosens. Bioelectron., 2024, 251, 116128 CrossRef CAS PubMed.
- H. Chen, L. Wang, D. Fan, P. Ma, X. Zhang and K. Lin, Analyst, 2025, 150, 4332–4341 RSC.
- J. K. Zaki, J. Tomasik, J. A. McCune, S. Bahn, P. Lio and O. A. Scherman, ACS Sens., 2025, 10, 6597–6606 Search PubMed.
- B. Xue, X. Bi, Z. Dong, Y. Xu, M. Liang, X. Fang, Y. Yuan, R. Wang, S. Liu, R. Jiao, Y. Chen, W. Zu, C. Wang, J. Zhang, J. Liu, Q. Zhang, Y. Yuan, M. Xu, Y. Zhang, Y. Wang, J. Ye and C. Jin, Nat. Mach. Intell., 2025, 7, 743–757 Search PubMed.
- J. Jeon, C. Mun, D. Kang, J. Na, H.-J. Park, J.-J. Rha, J.-W. Kim and S.-G. Park, J. Hazard. Mater., 2025, 500, 140584 Search PubMed.
- Z. Shen, L. Xie, Y. Hou, J. Liang, Y. Jia, H. Zhang, Z. Sun, J. Du, Z. He, C. Liu and W. Liu, Adv. Sci., 2025, 46, e13502 Search PubMed.
- M. Lei, J. Zhao, A. Z. Sahan, J. Hu, J. Zhou, H. Lee, Q. Wu, J. Zhang and Z. Liu, Nano Lett., 2024, 24, 15724–15730 Search PubMed.
- X. Li, D. Mengu, N. T. Yardimci, D. Turan, A. Charkhesht, A. Ozcan and M. Jarrahi, Nat. Photonics, 2024, 18, 139–148 CrossRef.
- A. P. Olson, C. T. Ertsgaard, S. N. Elliott and N. C. Lindquist, ACS Photonics, 2016, 3, 329–336 CrossRef CAS.
- J. Xu, H. An, X. Kong, Z. Zhang, Q. Liu, J. Li, J. Qin, I. A. Bratchenko and S. Wang, Anal. Chem., 2025, 97, 17121–17131 CrossRef CAS PubMed.
- Y. Cui, W. Ren, X. Cao and A. Knoll, IEEE Trans. Pattern Anal. Mach. Intell., 2024, 46, 9423–9438 Search PubMed.
- T. Boothe, M. Ivankovic, M. Grohme, M. Markus, C. Dullin, X. Xu and J. Rink, Commun. Biol., 2023, 6, 518 CrossRef PubMed.
- T. Buchholz, A. Krull, R. Shahidi, G. Pigino, G. Jékely and F. Jug, Three-Dimensional Electron Microscopy, 2019, vol. 152, pp. 277–289 Search PubMed.
- M. Weigert, U. Schmidt, T. Boothe, A. Müller, A. Dibrov, A. Jain, B. Wilhelm, D. Schmidt, C. Broaddus, S. Culley, M. Rocha-Martins, F. Segovia-Miranda, C. Norden, R. Henriques, M. Zerial, M. Solimena, J. Rink, P. Tomancak, L. Royer, F. Jug and E. W. Myers, Nat. Methods, 2018, 15, 1090–1097 CrossRef CAS PubMed.
- X. Xiao, W. Zhang, Y. Chang, S. Cao, W. He, H. Fang and L. Yan, IEEE Trans. Geosci. Remote Sens., 2023, 61, 5524119 Search PubMed.
- W. Vandenberg, M. Leutenegger, S. Duwé and P. Dedecker, Opt. Express, 2019, 27, 25749–25766 Search PubMed.
- F. Xue, W. He, D. Peng, H. You, M. Zhang and P. Xu, Fundamental Res., 2025, 5, 1025–1033 CrossRef PubMed.
- A. Hainsworth, S. Lee, P. Foot, A. Patel, W. Poon and A. Knight, Neuropathol. Appl. Neurobiol., 2018, 44, 417–426 CrossRef CAS PubMed.
- R. Chen, X. Tang, Y. Zhao, Z. Shen, M. Zhang, Y. Shen, T. Li, C. H. Y. Chung, L. Zhang, J. Wang, B. Cui, P. Fei, Y. Guo, S. Du and S. Yao, Nat. Commun., 2023, 14, 2854 CrossRef CAS PubMed.
- J. Chen, M. Guang, G. Chen, X. Yao, T. Tassew, Z. Li, Z. Li and H. Zhang, Int. J. Imag. Syst. Technol., 2025, 35, e70022 Search PubMed.
- D. Liu, J. Li and Q. Yuan, IEEE Trans. Geosci. Remote Sens., 2021, 59, 7711–7725 Search PubMed.
- Y. Zhang, Y. Tian, Y. Kong, B. Zhong and Y. Fu, IEEE Trans. Pattern Anal. Mach. Intell., 2021, 43, 2480–2495 Search PubMed.
- D. Zhu and D. Qiu, Comput. Methods Programs Biomed., 2021, 209, 106330 Search PubMed.
- Y. Li, Q. Yang, F. Zhao, J. Deng, Q. Ren and Y. Pan, Biomed. Signal Process. Control, 2025, 110, 108126 Search PubMed.
- C. Zhao, Z. Xu, X. Wang, S. Tao, W. MacDonald, K. He, A. Poholek, K. Chen, H. Huang and W. Chen, Briefings Bioinf., 2024, 25, bbae052 Search PubMed.
- J. Li, J. Chen, Y. Tang, C. Wang, B. A. Landman and S. K. Zhou, Med. Image Anal., 2023, 85, 102762 Search PubMed.
- K. Zhang, W. Zuo and L. Zhang, IEEE Trans. Image Process., 2017, 27, 4608–4622 Search PubMed.
- Y. Mansour and R. Heckel, arXiv, 2023, preprint, arXiv:2303.11253 DOI:10.48550/arXiv.2303.11253.
- C. Qiao, Y. Zeng, Q. Meng, X. Chen, H. Chen, T. Jiang, R. Wei, J. Guo, W. Fu, H. Lu, D. Li, Y. Wang, H. Qiao, J. Wu, D. Li and Q. Dai, Nat. Commun., 2024, 15, 4180 Search PubMed.
- S. Wang, C. Zhao, Q. Wang, M. Liu, C. Mou and F. Xu, Pattern Recogn., 2025, 167, 111779 Search PubMed.
- S. Minaee, Y. Boykov, F. Porikli, A. Plaza, N. Kehtarnavaz and D. Terzopoulos, IEEE Trans. Pattern Anal. Mach. Intell., 2020, 44, 3523–3542 Search PubMed.
- K. He, G. Gkioxari, P. Dollár and R. Girshick, IEEE Trans. Pattern Anal. Mach. Intell., 2020, 42, 386–397 Search PubMed.
- Z. Zhou, M. Siddiquee, N. Tajbakhsh and J. Liang, Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support, 2018, 11045, pp. 3–11.
- O. Ronneberger, P. Fischer and T. Brox, arXiv, 2015, preprint, arXiv:1505.04597 DOI:10.48550/arXiv.1505.04597.
- P. Vashishtha, H. G. Kattamuri, N. Thawari, M. Amirthalingam and R. Batra, Nat. Mach. Intell., 2025, 7, 79–84 CrossRef.
- M. Ziatdinov, A. Ghosh, C. Y. T. Wong and S. V. Kalinin, Nat. Mach. Intell., 2022, 4, 1101–1112 CrossRef.
- M. Safaldin, N. Zaghden and M. Mejdoub, IEEE Access, 2024, 12, 59782–59806 Search PubMed.
- A. Mohsin and S. Choudhury, ACS Omega, 2025, 10, 862–870 CrossRef CAS PubMed.
- S. Zhao, M. Xu, C. Lin, W. Zhang, D. Li, Y. Peng, M. Tanemura and Y. Yang, Biosensors, 2025, 15, 458 CrossRef CAS PubMed.
- J. Lee, G. Moon, S. Ka, K. Toh and D. Kim, Sensors, 2023, 23, 8100 CrossRef PubMed.
- I. Sajedian, J. Kim and J. Rho, Microsyst. Nanoeng., 2019, 5, 27 Search PubMed.
- Q. Cheng, X. Fu, Z. Zhang, Y. Cao, Y. Sun and H. Zhang, ASENS '24: Proceedings of the International Conference on Algorithms, Software Engineering, and Network Security, Association for Computing Machinery, New York, NY, United States, 2024, pp. 440–444 Search PubMed.
- B. Li, S. Liu, F. Wu, G. Li, M. Zhong and X. Guan, Int. J. Intell. Syst., 2022, 37, 8565–8582 CrossRef.
- Y.-C. Wu, A. Shiledar, Y.-C. Li, J. Wong, S. Feng, X. Chen, C. Chen, K. Jin, S. Janamian, Z. Yang, Z. S. Ballard, Z. Göröcs, A. Feizi and A. Ozcan, Light: Sci. Appl., 2017, 6, e17046 CrossRef CAS PubMed.
- F. Dhawi, A. Ghafoor, N. Almousa, S. Ali and S. Alqanbar, Front. Plant Sci., 2025, 16, 1594728 CrossRef PubMed.
- R. D. Uthoff, B. Song, S. Sunny, S. Patrick, A. Suresh, T. Kolur, G. Keerthi, O. Spires, A. Anbarani, P. Wilder-Smith, M. A. Kuriakose, P. Birur and R. Liang, PLoS One, 2018, 13, e0207493 CrossRef PubMed.
- B. Song and R. Liang, Biosens. Bioelectron., 2025, 271, 116982 CrossRef CAS PubMed.
- K. Behrouzi, Z. Khodabakhshi Fard, C.-M. Chen, P. He, M. Teng and L. Lin, Nat. Commun., 2025, 16, 4597 CrossRef CAS PubMed.
- E. Longato, B. Di Camillo, G. Sparacino, A. Avogaro and G. Fadini, Cardiovasc. Diabetol., 2022, 21, 159 CrossRef CAS PubMed.
- Q. Yang, J. Shang, Y. Chen, D. Tang, Y. Ouyang, B. Xiong and X. Zhang, Anal. Chem., 2021, 93, 16571–16580 CrossRef CAS PubMed.
- S. Bai, J. Nico Kolter and V. Koltun, arXiv, 2018, arXiv:1803.01271 DOI:10.48550/arXiv.1803.01271.
- P. Raja and K. Ramanan, in Advances in Data Science and Management. Lecture Notes on Data Engineering and Communications Technologies, ed. S. Borah, V. Emilia Balas and Z. Polkowski, Springer, Singapore, 2020, vol 37, pp. 395–409 Search PubMed.
- A. Tahmassebi, G. Wengert, T. Helbich, Z. Bago-Horvath, S. Alaei, R. Bartsch, P. Dubsky, P. Balizen, P. Clausen, P. Kapetas, E. Morris, A. Meyer-Baese and K. Pinker, Invest. Radiol., 2019, 54, 110–117 CrossRef PubMed.
- S. Xu, S. Liu, H. Wang, W. Chen, F. Zhang and Z. Xiao, Entropy, 2021, 23, 20 CrossRef PubMed.
- X. Jiang, H. Du, S. Gao, S. Fang, Y. Gong, N. Han, Y. Wang and K. Zheng, GISci. Remote Sens., 2024, 61, 2367806 CrossRef.
- D. dos Santos, M. Temperini and A. Brolo, Acc. Chem. Res., 2019, 52, 456–464 CrossRef CAS PubMed.
- A. Garg, S. Hawks, J. Pan, W. Wang, N. Duggal, L. Marr, P. Vikesland and W. Zhou, Biosens. Bioelectron., 2024, 247, 115946 CrossRef CAS PubMed.
- G. Bertasius, H. Wang and L. Torresani, arXiv, 2021, preprint, arXiv:2102.05095v4 DOI:10.48550/arXiv.2102.05095.
- M. Bonse, T. Gebhard, F. Dannert, O. Absil, F. Cantalloube, V. Christiaens, G. Cugno, E. Garvin, J. Hayoz, M. Kasper, E. Matthews, B. Schoelkopf and S. Quanz, Astron. J., 2025, 169, 194 CrossRef.
- S. Cuéllar, P. Granados, E. Fabregas, M. Curé, H. Vargas, S. Dormido-Canto and G. Farias, PLoS One, 2022, 17, e0268199 CrossRef PubMed.
- E. O. Garvin, M. J. Bonse, J. Hayoz, G. Cugno, J. Spiller, P. A. Patapis, D. P. D. D. L. Roche, R. Nath-Ranga, O. Absil, N. F. Meinshausen and S. P. Quanz, Astron. Astrophys., 2024, 689, A143 CrossRef CAS.
- Z. Wu, C. Zhang, X. Gu, I. Duporge, L. F. Hughey, J. A. Stabach, A. K. Skidmore, J. G. C. Hopcraft, S. J. Lee, P. M. Atkinson, D. J. McCauley, R. Lamprey, S. Ngene and T. Wang, Nat. Commun., 2023, 14, 3072 CrossRef CAS PubMed.
- M. Mohseni-Dargah, Z. Falahati, B. Dabirmanesh, P. Nasrollahi and K. Khajeh, in Artificial Intelligence and Data Science in Environmental Sensing, ed. M. Asadnia, A. Razmjou and A. Beheshti, Academic Press, 2022, pp. 269–298 DOI:10.1016/B978-0-323-90508-4.00012-5.
- G. Goumas, E. N. Vlachothanasi, E. C. Fradelos and D. S. Mouliou, Diagnostics, 2025, 15, 1037 CrossRef CAS PubMed.
- B. V. S. Chauhan, S. Verma, B. M. A. Rahman and K. P. Wyche, Atmosphere, 2025, 16, 359 CrossRef CAS.
- M. Goswami, P. Khare and S. Shakya, Plasmonics, 2024, 19, 363–377 CrossRef.
- K. Thadson, S. Sasivimolkul, P. Suvarnaphaet, S. Visitsattapongse and S. Pechprasarn, Sci. Rep., 2022, 12, 2052 CrossRef CAS PubMed.
- N. Islam, M. M. Shoaib Hasan, I. Hossain Shibly, M. B. Rashid, M. A. Yousuf, F. Haider, R. Ahmmed Aoni and R. Ahmed, Opt. Express, 2024, 32, 34184–34198 Search PubMed.
- B. Ahmed Taha, A. C. Kadhim, A. J. Addie, A. J. Haider, A. S. Azzahrani, P. Raizada, S. Rustagi, V. Chaudhary and N. Arsad, Microchem. J., 2024, 205, 111307 CrossRef CAS.
- K. Thadson, S. Visitsattapongse and S. Pechprasarn, Sci. Rep., 2021, 11, 16289 CrossRef CAS PubMed.
- Y.-F. Chang, Y.-C. Wang, T.-Y. Huang, M.-C. Li, S.-Y. Chen, Y.-X. Lin, L.-C. Su and K.-J. Lin, Anal. Chim. Acta, 2025, 1341, 343640 CrossRef CAS PubMed.
- Y. Liu, Q. Geng, W. Zhan and Z. Geng, Eng. Appl. Artif. Intell., 2025, 144, 110172 CrossRef.
- A. L. Blee, J. C. C. Day, P. E. J. Flewitt, A. Jeketo and D. Megson-Smith, J. Raman Spectrosc., 2021, 52, 1135–1147 CrossRef CAS.
- Q. Chen, T. Yamada, H. Santo, E. Murakami, A. Murata, Y. Matsushita and K. Nakatani, Adv. Intell. Discovery, 2025, 1, 2400020 Search PubMed.
- L. C. Oliveira, A. Q. Sales, M. V. L. Lopes and A. M. N. Lima, IEEE Trans. Instrum. Meas., 2025, 74, 1–10 Search PubMed.
- S. Jobst, P. Recum, Á. Écija-Arenas, E. Moser, R. Bierl and T. Hirsch, ACS Sens., 2023, 8, 3530–3537 CrossRef CAS PubMed.
- C. Xiao, J. Eriksson, A. Suska, D. Filippini and W. C. Mak, Anal. Chim. Acta, 2022, 1201, 339606 CrossRef CAS PubMed.
- Q. Li, H. Fan, Y. Bai, Y. Li, M. Ikram, Y. Wang, Y. Huo and Z. Zhang, New J. Phys., 2022, 24, 063005 CrossRef CAS.
- J. C. M. Gomes, L. C. Souza and L. C. Oliveira, Biosens. Bioelectron., 2021, 172, 112760 CrossRef CAS PubMed.
- Y.-T. Chen, Y.-C. Lee, Y.-H. Lai, J.-C. Lim, N.-T. Huang, C.-T. Lin and J.-J. Huang, Biosensors, 2020, 10, 209 CrossRef CAS PubMed.
- B. A. Taha, N. M. Ahmed, R. K. Talreja, A. J. Haider, Y. Al Mashhadany, Q. Al-Jubouri, A. B. Huddin, M. H. H. Mokhtar, S. Rustagi, A. Kaushik, V. Chaudhary and N. Arsad, ACS Synth. Biol., 2024, 13, 1600–1620 CrossRef CAS PubMed.
- F. Y. Cui and H. S. S. Zhou, Biosens. Bioelectron., 2020, 165, 112349 CrossRef CAS PubMed.
- Y. S. Dwivedi, R. Singh, A. K. Sharma and A. K. Sharma, Optic. Fiber Technol., 2024, 85, 103801 Search PubMed.
- G. P. Singh and N. Sardana, Plasmonics, 2022, 17, 1869–1888 CrossRef PubMed.
- D. Liu, J. Wang, L. Wu, Y. Huang, Y. Zhang, M. Zhu, Y. Wang, Z. Zhu and C. Yang, TrAC, Trends Anal. Chem., 2020, 122, 115701 CrossRef CAS.
- M. Aslan, E. Seymour, H. Brickner, A. E. Clark, I. Celebi, M. B. Townsend, P. S. Satheshkumar, M. Riley, A. F. Carlin, M. S. Ünlü and P. Ray, Biosens. Bioelectron., 2025, 269, 116932 CrossRef CAS PubMed.
- T. Wasilewski, W. Kamysz and J. Gębicki, Biosensors, 2024, 14, 356 CrossRef CAS PubMed.
- M. T. R. Sobur, M. M. Hasan, M. R. Khatun, M. M. Mia, W. Islam, M. S. Iqbal and M. M. Hossain, Opt. Laser Technol., 2025, 192, 113796 CrossRef CAS.
- J.-W. Yang, D. Khorsandi, L. Trabucco, M. Ahmed, A. Khademhosseini, M. R. Dokmeci, J. Y. Ye and V. Jucaud, Small, 2024, 20, 2403560 CrossRef CAS PubMed.
- Y. Park, B. Ryu, S. J. Ki, M. Chen, X. Liang and K. Kurabayashi, Nano Lett., 2023, 23, 98–106 CrossRef CAS PubMed.
- O. Yavas, S. S. Aćimović, J. Garcia-Guirado, J. Berthelot, P. Dobosz, V. Sanz and R. Quidant, ACS Sens., 2018, 3, 1376–1384 CrossRef CAS PubMed.
- M. M. Safaee, M. Gravely and D. Roxbury, Adv. Funct. Mater., 2021, 31, 2006254 CrossRef CAS.
- W. J. Peveler, ACS Sens., 2024, 9, 1656–1665 Search PubMed.
- Y. Geng, W. J. Peveler and V. M. Rotello, Angew. Chem., Int. Ed., 2019, 58, 5190–5200 CrossRef CAS PubMed.
- I. Calderon, I. B. Becerril-Castro, T. Zorlu, B. Özdemir, E. García-Rico, V. A. Baulin and R. A. Alvarez-Puebla, ChemPlusChem, 2024, 89, e202400210 CrossRef CAS PubMed.
- Z. Li, J. R. Askim and K. S. Suslick, Chem. Rev., 2019, 119, 231–292 CrossRef CAS PubMed.
- A. Bigdeli, F. Ghasemi, H. Golmohammadi, S. Abbasi-Moayed, M. A. F. Nejad, N. Fahimi-Kashani, S. Jafarinejad, M. Shahrajabian and M. R. Hormozi-Nezhad, Nanoscale, 2017, 9, 16546–16563 RSC.
- N. Kim, M. R. Thomas, M. S. Bergholt, I. J. Pence, H. Seong, P. Charchar, N. Todorova, A. Nagelkerke, A. Belessiotis-Richards, D. J. Payne, A. Gelmi, I. Yarovsky and M. M. Stevens, Nat. Commun., 2020, 11, 207 CrossRef CAS PubMed.
- C. S. L. Koh, H. K. Lee, X. M. Han, H. Y. F. Sim and X. Y. Ling, Chem. Commun., 2018, 54, 2546–2549 RSC.
- J. S. Xie, Z. H. Ren, H. Zhou, J. K. Zhou, W. X. Liu and C. K. Lee, Adv. Opt. Mater., 2024, 12, 2418705 Search PubMed.
- S. Tabassum, R. Kumar and L. Dong, IEEE Sens. J., 2017, 17, 6210–6223 Search PubMed.
- E. A. R. Macias, J. T. R. Sperling, L. Peveler, E. A. Burley, T. E. Neale and A. Clark, Nanoscale, 2019, 11, 15216–15223 Search PubMed.
- L.-A. Garçon, M. Genua, Y. Hou, A. Buhot, R. Calemczuk, T. Livache, M. Billon, C. Le Narvor, D. Bonnaffé, H. Lortat-Jacob and Y. Hou, Sensors, 2017, 17, 1046 CrossRef PubMed.
- S. Forest, J. Coutu, J. M. Montiel-Leon, I. Beniani, Z. S. Yu, M. Craig and J. F. Masson, ACS Food Sci. Technol., 2023, 3, 635–647 Search PubMed.
- S. Forest, T. Théorêt, J. Coutu and J. F. Masson, Anal. Methods, 2020, 12, 2460–2468 Search PubMed.
- N. Fahimi-Kashani and M. R. Hormozi-Nezhad, Anal. Chem., 2016, 88, 8099–8106 Search PubMed.
- S. Jafarinejad, M. Ghazi-Khansari, F. Ghasemi, P. Sasanpour and M. R. Hormozi-Nezhad, Sci. Rep., 2017, 7, 8266 CrossRef PubMed.
- M. Morelos-Pacheco, M. A. Arellano-Alcántara, J. Olivares-Peralta, R. Colmenero-Solís, J. M. Saniger-Blesa, J. C. Cancino-Díaz, J. Widén, B. Chávez-Ramírez and J. S. Mendoza-Figueroa, Phytopathol. Res., 2025, 7, 36 Search PubMed.
- J. Mao, Y. Lu, N. Chang, J. Yang, S. Zhang and Y. Liu, Biosens. Bioelectron., 2016, 86, 56–61 CrossRef CAS PubMed.
- R. A. Potyrailo, M. Larsen and O. Riccobono, Angew. Chem., Int. Ed., 2013, 52, 10360–10364 Search PubMed.
- N. A. Joy, M. I. Nandasiri, P. H. Rogers, W. L. Jiang, T. Varga, S. Kuchibhatla, S. Thevuthasan and M. A. Carpenter, Anal. Chem., 2012, 84, 5025–5034 CrossRef CAS PubMed.
- J. Lee, D. Kim, G. Kim, J. H. Han and H. H. Jeong, ACS Appl. Mater. Interfaces, 2024, 16, 16622–16629 CrossRef CAS PubMed.
- M. Genua, L.-A. Garçon, V. Mounier, H. Wehry, A. Buhot, M. Billon, R. Calemczuk, D. Bonnaffé, Y. Hou and T. Livache, Talanta, 2014, 130, 49–54 CrossRef CAS PubMed.
- C. Qu, H. Fang, F. F. Yu, J. N. Chen, M. K. Su and H. L. Liu, Chem. Eng. J., 2024, 482, 148773 CrossRef CAS.
- X. G. Zhang, Z. Q. Liu, X. Y. Zhong, J. Liu, X. H. Xiao and C. Z. Jiang, J. Phys. D:Appl. Phys., 2021, 54, 255306 Search PubMed.
- Y. X. Pan, X. Liu, L. B. Qian, Y. X. Cui, X. B. Zheng, Y. R. Kang, X. Fu, S. P. Wang, P. Wang and D. Wang, Sens. Actuators, B, 2022, 352, 130971 CrossRef CAS.
- N. Mohseni, M. Bahram and T. Baheri, Sens. Actuators, B, 2017, 250, 509–517 CrossRef CAS.
- D. Li, Y. Dong, B. Li, Y. Wu, K. Wang and S. Zhang, Analyst, 2015, 140, 7672–7677 RSC.
- X. Yang, J. Li, H. Pei, Y. Zhao, X. Zuo, C. Fan and Q. Huang, Anal. Chem., 2014, 86, 3227–3231 CrossRef CAS PubMed.
- A. Fournel, M. Mantel, M. Pinger, C. Manesse, R. Dubreuil, C. Herrier, T. Rousselle, T. Livache and M. Bensafi, Sens. Actuators, B, 2020, 320, 128342 CrossRef CAS.
- K. C. Lee, I. C. Cho, M. G. Kang, J. Jeong, M. Choi, K. Y. Woo, K. J. Yoon, Y. H. Cho and I. Park, ACS Nano, 2023, 17, 539–551 CrossRef CAS PubMed.
- E. X. Tan, Y. X. Leong, S. H. Lim, M. W. K. Chng, I. Y. Phang and X. Y. Ling, Nat. Commun., 2025, 16, 7095 CrossRef CAS PubMed.
- E. K. Herkert and M. F. Garcia-Parajo, ACS Photonics, 2025, 12, 1259–1275 CrossRef CAS PubMed.
- S. H. Cho, S. Choi, J. M. Suh and H. W. Jang, J. Mater. Chem. C, 2025, 13, 6484–6507 RSC.
- H. Chang, W. Hur, H. Kang, B.-H. Jun, H. Chang, W. Hur, H. Kang and B.-H. Jun, Light: Sci. Appl., 2025, 14, 1 Search PubMed.
- S. Lee, N. A. M. Moussa and S. H. Kang, Nanomaterials, 2025, 15, 1153 CrossRef CAS PubMed.
- S. Sloan-Dennison, K. M. Scullion, B. Clark, P. Fineran, J. Mair, S. Laing, N. C. Shand, C. Rathmell, D. Creasey, D. Bingemann, J. Faircloth, M. Zieg, E. Varghese, C. J. Weir, J. W. Dear, K. Faulds and D. Graham, Nat. Commun., 2025, 16, 6223 CrossRef CAS PubMed.
- C. Jin, Z. Wu, J. H. Molinski, J. Zhou, Y. Ren and J. X. J. Zhang, Mater. Today Bio, 2022, 14, 100263 CrossRef CAS PubMed.
- S. Subburaj, C. Liu and T. Xu, Chem. Commun., 2025, 61, 18464–18489 Search PubMed.
- E. X. Tan, S. X. Leong, W. A. Liew, I. Y. Phang, J. Y. Ng, N. S. Tan, Y. H. Lee and X. Y. Ling, Nat. Commun., 2024, 15, 2582 Search PubMed.
- S. X. Leong, E. X. Tan, X. Han, I. Luhung, N. W. Aung, L. B. T. Nguyen, S. Y. Tan, H. Li, I. Y. Phang, S. Schuster and X. Y. Ling, ACS Nano, 2023, 17, 23132–23143 Search PubMed.
- J. R. T. Chen, E. X. Tan, J. Tang, S. X. Leong, S. K. X. Hue, C. S. Pun, I. Y. Phang and X. Y. Ling, J. Am. Chem. Soc., 2025, 147, 6654–6664 CrossRef CAS PubMed.
- D. Kalatzis, A. I. Katsafadou, D. Chatzopoulos, C. Billinis and Y. Kiouvrekis, Micro, 2025, 5, 46 CrossRef.
- W. Sun, J. Nan, H. Xu, L. Wang, J. Niu, J. Zhang and B. Yang, Nano Lett., 2024, 24, 8784–8792 CrossRef CAS PubMed.
- G. Rong, Y. Xu and M. Sawan, Biosensors, 2023, 13, 860 CrossRef PubMed.
- L. Bi, H. Zhang, C. Mu, K. Sun, H. Chen, Z. Zhang and L. Chen, J. Hazard. Mater., 2025, 494, 138694 Search PubMed.
- Y. Liu, Y. Gao, R. Niu, Z. Zhang, G.-W. Lu, H. Hu, T. Liu and Z. Cheng, Anal. Chim. Acta, 2024, 1332, 343376 CrossRef CAS PubMed.
- Y. Wu, J. Chen, L. Bi, Z. Zhang, X. Wang, L. Fu, Q. Yang, J. Choo and L. Chen, Adv. Sci., 2025, 13, e16240 CrossRef PubMed.
- B. L. Thomsen, J. B. Christensen, O. Rodenko, I. Usenov, R. B. Grønnemose, T. E. Andersen and M. Lassen, Sci. Rep., 2022, 12, 16436 CrossRef CAS PubMed.
- L. Bai, G. Fang, J. Li, W. Hasi and S. Han, Spectrochim. Acta, Part A, 2026, 346, 126768 Search PubMed.
- X. Xie, W. Yu, L. Wang, J. Yang, X. Tu, X. Liu, S. Liu, H. Zhou, R. Chi and Y. Huang, Spectrochim. Acta, Part A, 2024, 314, 124181 CrossRef CAS PubMed.
- A. S. Malinick, D. D. Stuart, A. S. Lambert and Q. Cheng, Biosens. Bioelectron.: X, 2022, 10, 100127 CAS.
- H. Julkunen, A. Cichońska, M. Tiainen, H. Koskela, K. Nybo, V. Mäkelä, J. Nokso-Koivisto, K. Kristiansson, M. Perola, V. Salomaa, P. Jousilahti, A. Lundqvist, A. J. Kangas, P. Soininen, J. C. Barrett and P. Würtz, Nat. Commun., 2023, 14, 604 CrossRef CAS PubMed.
- S. Alseekh, A. Aharoni, Y. Brotman, K. Contrepois, J. DAuria, J. Ewald, J. C. Ewald, P. D. Fraser, P. Giavalisco, R. D. Hall, M. Heinemann, H. Link, J. Luo, S. Neumann, J. Nielsen, L. Perez de Souza, K. Saito, U. Sauer, F. C. Schroeder, S. Schuster, G. Siuzdak, A. Skirycz, L. W. Sumner, M. P. Snyder, H. Tang, T. Tohge, Y. Wang, W. Wen, S. Wu, G. Xu, N. Zamboni and A. R. Fernie, Nat. Methods, 2021, 18, 747–756 CrossRef CAS PubMed.
- S. Qiu, J. Guo, Z. Zhang, H. Liang, H. You, Y. Hu, G. Liu and Y. Wang, Nat. Commun., 2025, 16, 11272 CrossRef CAS PubMed.
- J. Plou, P. S. Valera, I. García, D. Vila-Liarte, C. Renero-Lecuna, J. Ruiz-Cabello, A. Carracedo and L. M. Liz-Marzán, Small, 2023, 19, 2207658 CrossRef CAS PubMed.
- K. Echeverría-Altamar, C. Barreto-Gamarra, M. Domenech-García and P. Resto-Irizarry, Biosens. Bioelectron., 2025, 283, 117528 CrossRef PubMed.
- F. Lussier, D. Missirlis, J. P. Spatz and J.-F. Masson, ACS Nano, 2019, 13, 1403–1411 CAS.
- D. Kavungal, P. Magalhães, S. T. Kumar, R. Kolla, H. A. Lashuel and H. Altug, Sci. Adv., 2023, 9, eadg9644 CrossRef CAS PubMed.
- R. P. Carney, R. R. Mizenko, B. T. Bozkurt, N. Lowe, T. Henson, A. Arizzi, A. Wang, C. Tan and S. C. George, Nat. Nanotechnol., 2024, 20, 14–25 CrossRef PubMed.
- T. Wang, K. Zhu, J. Wei, Y. Zhao, Z. Xu, S. Zong, L. Wu, K. Yang, H. Wang and Z. Wang, ACS Sens., 2025, 10, 6840–6848 Search PubMed.
- X. Huang, B. Liu, S. Guo, W. Guo, K. Liao, G. Hu, W. Shi, M. Kuss, M. J. Duryee, D. R. Anderson, Y. Lu and B. Duan, Bioeng. Transl. Med., 2022, 8, e10420 CrossRef PubMed.
- H.-S. Liu, K.-W. Ye, J. Liu, J.-K. Jiang, Y.-F. Jian, D.-M. Chen, C. Kang, L. Qiu and Y.-J. Liu, Theranostics, 2025, 15, 7545–7566 Search PubMed.
- H. Shin, B. H. Choi, O. Shim, J. Kim, Y. Park, S. K. Cho, H. K. Kim and Y. Choi, Nat. Commun., 2023, 14, 1644 Search PubMed.
- C. Zhang, W.-H. Zhao, D. Zuo, T. Zhou, W. Hou, L. Wang, S. Shi, Y. Yang, Y. Liu, S.-K. Sun, L. Ren, Z. Ye, D. Liu, D. Li, X. Chen and J. Hao, Adv. Sci., 2025, 13, e16298 Search PubMed.
- L. Rodríguez-Lorenzo, A. Garrido-Maestu, A. K. Bhunia, B. Espiña, M. Prado, L. Diéguez and S. Abalde-Cela, ACS Appl. Nano Mater., 2019, 2, 6081–6086 Search PubMed.
- Z.-H. Fong, C.-H. Wang, C.-Y. Yang, H.-C. Kan, Y.-J. Lin, Y.-L. Chen, Y.-C. Shen, Y.-C. Yu, L.-K. Chau, C.-W. Io and S.-C. Wang, Microchem. J., 2025, 212, 113436 CrossRef CAS.
- D. P. Ng, P. D. Simonson, A. Tarnok, F. Lucas, W. Kern, N. Rolf, G. Bogdanoski, C. Green, R. R. Brinkman and K. Czechowska, Cytometry, Part B, 2024, 106, 228–238 CrossRef CAS PubMed.
- Y. Zhang, K. Chang, B. Ogunlade, L. Herndon, L. F. Tadesse, A. R. Kirane and J. A. Dionne, ACS Nano, 2024, 18, 18101–18117 CrossRef CAS PubMed.
- S. Park, K. Kim, A. Go, M.-H. Lee, L. Chen and J. Choo, ACS Sens., 2025, 10, 1217–1227 CrossRef CAS PubMed.
- Z. Wang, K. Han, W. Liu, Z. Wang, C. Shi, X. Liu, M. Huang, G. Sun, S. Liu and Q. Guo, J. Imag. Inform. Med., 2024, 37, 1160–1176 Search PubMed.
- J. Zhang, J. Zhao, H. Lin, Y. Tan and J.-X. Cheng, J. Phys. Chem. Lett., 2020, 11, 8573–8578 Search PubMed.
- W. Jiang, Y. Su, M. Guo, X. Wang, H. Liu, X. Liu and Y. Zhang, Spectrochim. Acta, Part A, 2026, 345, 126832 CrossRef CAS PubMed.
- Q. Huang, H. Guo, W. Wang, S. Kang and P. J. Vikesland, ACS Environ. Au, 2025, 5, 342–362 Search PubMed.
- T. Tanaka, T.-a Yano and R. Kato, Nanophotonics, 2022, 11, 2541–2561 CrossRef CAS PubMed.
- A. Keogan, T. N. Q. Nguyen, P. Bouzy, N. Stone, K. Jirstrom, A. Rahman, W. M. Gallagher and A. D. Meade, npj Precis. Oncol., 2025, 9, 18 CrossRef PubMed.
- Z. Jin, Q. Yue, W. Duan, A. Sui, B. Zhao, Y. Deng, Y. Zhai, Y. Zhang, T. Sun, G.-P. Zhang, L. Han, Y. Mao, J. Yu, X.-Y. Zhang and C. Li, Adv. Sci., 2022, 9, 2104935 CrossRef PubMed.
- K. Ember, F. Dallaire, A. Plante, G. Sheehy, M.-C. Guiot, R. Agarwal, R. Yadav, A. Douet, J. Selb, J. P. Tremblay, A. Dupuis, E. Marple, K. Urmey, C. Rizea, A. Harb, L. McCarthy, A. Schupper, M. Umphlett, N. Tsankova, F. Leblond, C. Hadjipanayis and K. Petrecca, Sci. Rep., 2024, 14, 13309 CrossRef CAS PubMed.
- G. Sheehy, F. Picot, F. Dallaire, K. J. Ember, T. Nguyen, K. Petrecca, D. Trudel and F. Leblond, J. Biomed. Opt., 2023, 28, 025002 Search PubMed.
- N. Blake, R. Gaifulina, L. D. Griffin, I. M. Bell and G. M. H. Thomas, Diagnostics, 2022, 12, 1491 CrossRef PubMed.
- Y. Liu, Y. Yang, H. Lu, J. Cui, X. Chen, P. Ma, W. Zhong and Y. Zhao, ACS Sens., 2025, 10, 3941–3952 Search PubMed.
- X. Dong, X. Zhao, F. Zheng, G. Xu, T. Zhang, M. Zhou, J. Zhou, Y. Liu and G. Wang, Microchem. J., 2025, 212, 113465 Search PubMed.
- M. Seggio, F. Arcadio, E. Radicchi, N. Cennamo, L. Zeni and A. M. Bossi, ACS Omega, 2024, 9, 18984–18994 CrossRef CAS PubMed.
- J. Y. Kim, E. H. Koh, J. Y. Yang, C. Mun, S. Lee, H. Lee, J. Kim, S. G. Park, M. Kang, D. H. Kim and H. S. Jung, Adv. Funct. Mater., 2023, 34, 2307584 CrossRef.
- J. Zhou, Z. Wu, C. Jin and J. X. J. Zhang, npj Clean Water, 2024, 7, 3 CrossRef CAS.
- Y. Lu, Y. Qiao, H. Bao, K. Chen, Y. Wei, Q. Zhao, G. K. Leon, H. Zhang, X. Y. Ling and W. Cai, Anal. Chem., 2025, 97, 8537–8544 CrossRef CAS PubMed.
- R. Qin, Y. Zhang, S. Ren and P. Nie, Int. J. Mol. Sci., 2022, 23, 10404 Search PubMed.
- H. Wang, L. Gamage, J. Li and H. Wei, Environ. Sci.: Nano, 2025, 12, 3468–3475 RSC.
- M. Dong, F. Ding, Y. Jin, C. Li, X. Lin and S. Lin, Anal. Chem., 2025, 97, 10452–10462 Search PubMed.
- G. Fang, W. Hasi, X. Lin and S. Han, J. Hazard. Mater., 2024, 474, 134814 Search PubMed.
- F. Sahin, N. Celik, A. Camdal, M. Sakir, A. Ceylan, M. Ruzi and M. S. Onses, ACS Appl. Nano Mater., 2022, 5, 13112–13122 Search PubMed.
- K. V. Serebrennikova, N. S. Komova, A. V. Zherdev and B. B. Dzantiev, Biosensors, 2024, 14, 573 CrossRef CAS PubMed.
- H. J. Seo, J. Y. Kim, J. Y. Yang, C. Mun, S. Lee, E. H. Koh, V. T. N. Linh, M. Kang and H. S. Jung, Adv. Sens. Res., 2024, 3, 2400030 Search PubMed.
- A. Lowe, N. Harrison and A. P. French, Plant Methods, 2017, 13, 80 CrossRef PubMed.
- I. T. Jolliffe and J. Cadima, Philos. Trans. A Math. Phys. Eng. Sci., 2016, 374, 20150202 Search PubMed.
- J. Olenik, V. Shvalya, M. Modic, D. Vengust, U. Cvelbar and J. L. Walsh, ACS Sens., 2025, 10, 387–397 CrossRef CAS PubMed.
- F. Rosenblatt, Psychol. Rev., 1958, 65, 386–408 CAS.
- Y. Wan, Q. Wei, H. Sun, H. Wu, Y. Zhou, C. Bi, J. Li, L. Li, B. Liu, D. Wang, X. Wang, C. Wang and W. Liu, Chem. Eng. J., 2025, 507, 160813 CrossRef CAS.
- J. W. Boardman, AVIRIS, 1993 Search PubMed.
- F. A. Kruse, A. B. Lefkoff, J. W. Boardman, K. B. Heidebrecht, A. T. Shapiro, P. J. Barloon and A. F. H. Goetz, Remote Sens. Environ., 1993, 44, 145–163 CrossRef.
- S. S. Khan and M. G. Madden, Chemom. Intell. Lab. Syst., 2012, 114, 99–108 CrossRef CAS.
- J. Malenfant, L. Kuster, Y. Gagne, K. Signo, M. Denis, S. Canesi and M. Frenette, Chem. Sci., 2024, 15, 701–709 RSC.
- Q. Wang, S. Zheng, M. Qiu and D. Hu, in Proceedings of the 2025 2nd International Conference on Generative Artificial Intelligence and Information Security, Association for Computing Machinery, 2025, pp. 359–366 DOI:10.1145/3728725.3728783.
- M. Z. Andrew, G. Howard, B. Chen, D. Kalenichenko, W. Wang, T. Weyand, M. Andreetto and H. Adam, MobileNets: Efficient Convolutional Neural Networks for Mobile Vision Applications, https://arxiv.org/abs/1704.04861.
- S. Weng, L. Tang, M. Qiu, J. Wang, Y. Wu, R. Zhu, C. Wang, P. Li, W. Sha and D. Liang, Anal. Chim. Acta, 2023, 1262, 341264 CrossRef CAS PubMed.
- C. Farber and D. Kurouski, Front. Plant Sci., 2022, 13, 887511 CrossRef CAS PubMed.
- D. Sarma, M. R. Marak, I. Chetia, L. S. Badwaik and P. Nath, Phys. Scr., 2024, 99, 026006 CrossRef CAS.
- X. He, S. Chao, S. Zhou, R. Zhao, C. Valsecchi, Y. Lin, H. Yang and M. Fan, ACS Agric. Sci. Technol., 2025, 5, 414–422 CrossRef CAS.
- X. Wang, X. Sun, Z. Liu, Y. Zhao, G. Wu, Y. Wang, Q. Li, C. Yang, T. Ban, Y. Liu, J. A. Huang and Y. Li, Adv. Sci., 2024, 11, e2405416 CrossRef PubMed.
- S. Fang, Y. Zhao, Y. Wang, J. Li, F. Zhu and K. Yu, Front. Plant Sci., 2022, 13, 802761 CrossRef CAS PubMed.
- L. N. Kissell, H. Liu, M. Sheokand, D. Vang, P. Kachroo and P. Strobbia, ACS Sens., 2024, 9, 514–523 CrossRef CAS PubMed.
- L. Javaid, N. Iralu, S. Wani and A. Hamid, in Detection of Plant Viruses, ed. A. Hamid, G. Ali, A. Shikari, S. Saleem, S. H. Wani and S. Wani, Springer Protocols Handbooks, Humana, New York, 2025 DOI:10.1007/978-1-0716-4390-7_38.
- K. Thakkar, V. Singh, P. Sharma, P. Jain, A. Jyoti, A. Kumar, S. Mehta, A. Singh, M. Singh and J. Saxena, Appl. Biochem. Biotechnol., 2025, 197, 5956–5991 CrossRef CAS PubMed.
- J. Li, Z. Yang, Y. Zhao and K. Yu, Measurement, 2024, 224, 113911 CrossRef.
- S. Ali, A. Amin, M. S. Akhtar and W. Zaman, Nanomaterials, 2025, 15, 899 CrossRef CAS PubMed.
- C. Wang, P. Xu, H. Wang, F. Luo, Q. Tu, Y. Fang, R. You, Y. Yang and Y. Lu, J. Food Compos. Anal., 2025, 140, 107242 CrossRef CAS.
- C. Zhong, L. Li and Y.-Z. Wang, Microchem. J., 2024, 206, 111447 CrossRef CAS.
- Y. S. Choi, W. K. Son, H. Kwak, J. Park, S. Choi, D. Sim, M. G. Kim, H. Kimm, H. Son, D. H. Jeong and S. Y. Kwak, Adv. Sci., 2024, 12, e2412732 CrossRef PubMed.
- J. Lv, P. Christie and S. Zhang, Environ. Sci.: Nano, 2019, 6, 41–59 RSC.
- Y. Su, V. Ashworth, C. Kim, A. S. Adeleye, P. Rolshausen, C. Roper, J. White and D. Jassby, Environ. Sci.: Nano, 2019, 6, 2311–2331 RSC.
- E. Voke, R. L. Pinals, N. S. Goh and M. P. Landry, ACS Sens., 2021, 6, 2802–2814 CrossRef CAS PubMed.
- S. X. Leong, Y. X. Leong, C. S. L. Koh, E. X. Tan, L. B. T. Nguyen, J. R. T. Chen, C. Chong, D. W. C. Pang, H. Y. F. Sim, X. Liang, N. S. Tan and X. Y. Ling, Chem. Sci., 2022, 13, 11009–11029 RSC.
- C. Yang, Y. Zhao, S. Jiang, X. Sun, X. Wang, Z. Wang, Y. Wu, J. Wu and Y. Li, Mikrochim. Acta, 2024, 191, 286 CrossRef CAS PubMed.
- J. Zhu, X. Jiang, Y. Rong, W. Wei, S. Wu, T. Jiao and Q. Chen, Food Chem., 2023, 414, 135705 CrossRef CAS PubMed.
- D. Wang, T. Ahmad, S. A. Khalid, A. S. A. Dena and Y. Liu, LWT, 2025, 223, 117738 CrossRef CAS.
- Z. Zhang, H. Li, L. Huang, H. Wang, H. Niu, Z. Yang and M. Wang, Spectrochim. Acta, Part A, 2024, 320, 124655 CrossRef CAS PubMed.
- M. Hajikhani, A. Hegde, J. Snyder, J. Cheng and M. Lin, J. Hazard. Mater., 2024, 470, 134208 CrossRef CAS PubMed.
- H. Wang, Z. Bian, Y. Wang, H. Niu, Z. Yang and H. Li, Anal. Methods, 2025, 17, 1884–1891 RSC.
- O. C. Koyun, R. K. Keser, S. O. Şahin, D. Bulut, M. Yorulmaz, V. Yücesoy and B. U. Töreyin, ACS Omega, 2024, 9, 23241–23251 CrossRef CAS PubMed.
- J.-Y. Mou, M. Usman, J.-W. Tang, Q. Yuan, Z.-W. Ma, X.-R. Wen, Z. Liu and L. Wang, Food Chem.: X, 2024, 22, 101507 CAS.
- Q. Yuan, L.-F. Yao, J.-W. Tang, Z.-W. Ma, J.-Y. Mou, X.-R. Wen, M. Usman, X. Wu and L. Wang, J. Adv. Res., 2025, 69, 61–74 CrossRef CAS PubMed.
- Y. Jin, C. Li, Z. Huang and L. Jiang, Foods, 2023, 12, 4267 CrossRef CAS PubMed.
- H. Kang, J. Lee, J. Moon, T. Lee, J. Kim, Y. Jeong, E. K. Lim, J. Jung, Y. Jung and S. J. Lee, Small, 2024, 20, 2308317 CrossRef CAS PubMed.
- Z. Wang, P. Liang, J. Zhai, B. Wu, X. Chen, F. Ding, Q. Chen and B. Sun, J. Hazard. Mater., 2025, 489, 137581 CrossRef CAS PubMed.
- H. Ma, G. Li, H. Zhang, X. Wang, F. Li, J. Yan, L. Hong, Y. Zhang and Q. Pu, Sens. Actuators, B, 2025, 436, 137646 CrossRef CAS.
- S. M. Eid, S. El-Shamy and M. A. Farag, Microchim. Acta, 2022, 189, 301 CrossRef CAS PubMed.
- E. Corcione, D. Pfezer, M. Hentschel, H. Giessen and C. Tarín, Sensors, 2021, 22, 7 CrossRef PubMed.
- X. Wang, F. Li, L. Wei, Y. Huang, X. Wen, D. Wang, G. Cheng, R. Zhao, Y. Lin and H. Yang, Anal. Chem., 2024, 96, 4682–4692 CrossRef CAS PubMed.
- L. Zhang, C. Zhang, W. Li, L. Li, P. Zhang, C. Zhu, Y. Ding and H. Sun, Foods, 2023, 12, 4124 CrossRef CAS PubMed.
- S. Wang, D. Yang, X. Wu, Z. Du, X. Zhang, X. Zhao, J. Wang and Y. Zhang, J. Photochem. Photobiol., A, 2025, 472, 116760 CrossRef.
- C. Mei, Y. Xue, Q. Li and H. Jiang, Infrared Phys. Technol., 2024, 140, 105402 CrossRef CAS.
- A. Tan, Y. He, B. Shi, H. Mu, Y. Zhang and Y. Zhao, Talanta, 2025, 298, 128852 CrossRef PubMed.
- A. Skorikov, W. Heyvaert, W. Albecht, D. M. Pelt and S. Bals, Nanoscale, 2021, 13, 12242–12249 RSC.
- E. M. Mikmeková, J. Materna, I. Konvalina, S. Mikmeková, I. Müllerová and T. Asefa, Ultramicroscopy, 2024, 262, 113965 CrossRef PubMed.
- Z. Q. Wu, Y. P. Ma, H. Liu, C. Z. Huang and J. Zhou, Anal. Chem., 2023, 95, 15375–15383 CrossRef CAS PubMed.
- M. K. Song, S. X. Chen, P. P. Hu, C. Z. Huang and J. Zhou, Anal. Chem., 2021, 93, 2619–2626 CrossRef CAS PubMed.
- V. D. Babaylova, V. S. Tuchin, N. S. Petrov, A. V. Kochakov, A. A. Starovoytov, I. A. Gladskikh and D. R. Dadadzhanov, Photonics, 2025, 12, 619 CrossRef CAS.
- Z. J. O'Dell, M. Knobeloch, S. E. Skrabalak and K. A. Willets, Nano Lett., 2024, 24, 7269–7275 CrossRef PubMed.
- M. Ibrar, S. Y. Huang, Z. McCurtain, S. Naha, D. J. Crandall, S. C. Jacobson and S. E. Skrabalak, Adv. Funct. Mater., 2024, 34, 2400842 CrossRef CAS PubMed.
- J. Timoshenko, S. Roese, H. Hövel and A. I. Frenkel, Radiat. Phys. Chem., 2020, 175, 108049 CrossRef CAS.
- K. Y. Bi, L. Lv, D. Su, S. J. Wang, X. Y. Zhang and T. Zhang, Langmuir, 2024, 40, 19412–19422 CrossRef CAS PubMed.
- J. F. Xue, Z. Wang, H. Zhang and Y. He, J. Phys. Chem. B, 2022, 129, 7541–7551 CrossRef PubMed.
- C. J. Taylor, A. Pomberger, K. C. Felton, R. Grainger, M. Barecka, T. W. Chamberlain, R. A. Bourne, C. N. Johnson and A. A. Lapkin, Chem. Rev., 2023, 123, 3089–3126 CrossRef CAS PubMed.
- J. Kimling, M. Maier, B. Okenve, V. Kotaidis, H. Ballot and A. Plech, J. Phys. Chem. B, 2006, 110, 15700–15707 CrossRef CAS PubMed.
- M. Abolhasani and E. Kumacheva, Nat. Synth., 2023, 2, 483–492 CrossRef CAS.
- J. Schmidt, M. R. G. Marques, S. Botti and M. A. L. Marques, npj Comput. Mater., 2019, 5, 83 CrossRef.
- A. D. Clayton, J. A. Manson, C. J. Taylor, T. W. Chamberlain, B. A. Taylor, G. Clemens and R. A. Bourne, React. Chem. Eng., 2019, 4, 1545–1554 RSC.
- Z. S. Ballard, D. Shir, A. Bhardwaj, S. Bazargan, S. Sathianathan and A. Ozcan, ACS Nano, 2017, 11, 2266–2274 CrossRef CAS PubMed.
- B. Shahriari, K. Swersky, Z. Wang, R. P. Adams and N. D. Freitas, Proc. IEEE, 2016, 104, 148–175 Search PubMed.
- E. Brochu, V. M. Cora and N. de Freitas, arXiv, 2010, preprint, arXiv:1012.2599 DOI:10.48550/arXiv.1012.2599.
- B. J. Shields, J. Stevens, J. Li, M. Parasram, F. Damani, J. I. M. Alvarado, J. M. Janey, R. P. Adams and A. G. Doyle, Nature, 2021, 590, 89–96 CrossRef CAS PubMed.
- F. Mekki-Berrada, Z. Ren, T. Huang, W. K. Wong, F. Zheng, J. Xie, I. P. S. Tian, S. Jayavelu, Z. Mahfoud, D. Bash, K. Hippalgaonkar, S. Khan, T. Buonassisi, Q. Li and X. Wang, npj Comput. Mater., 2021, 7, 55 CrossRef CAS.
- A. Rao and M. Grzelczak, Chem. Mater., 2024, 36, 2577–2587 CrossRef CAS PubMed.
- A. N. Giordano, S. Franqui-Rios, S. M. Quarin, D. Vang, D. R. Austin, A. G. Doyle, L. A. Baldwin, P. Strobbia and R. Rao, ACS Appl. Nano Mater., 2025, 8, 11930–11939 CrossRef CAS.
- Y. Xie, K. Sattari, C. Zhang and J. Lin, Prog. Mater. Sci., 2023, 132, 101043 CrossRef.
- A. G. Barto, AI Mag., 2019, 40, 3–15 Search PubMed.
- A. A. Volk, R. W. Epps, D. T. Yonemoto, B. S. Masters, F. N. Castellano, K. G. Reyes and M. Abolhasani, Nat. Commun., 2023, 14, 1403 CrossRef CAS PubMed.
- K. Arulkumaran, M. P. Deisenroth, M. Brundage and A. A. Bharath, IEEE Signal Process. Mag., 2017, 34, 26–38 Search PubMed.
- L. Yang, H. Wang, D. Leng, S. Fang, Y. Yang and Y. Du, Chem. Eng. J., 2024, 500, 156687 CrossRef CAS.
- E. Noorani, C. Somarakis, R. Goyal, A. Feldman and S. Rane, in 2022 IEEE 17th International Conference on Control & Automation (ICCA), Naples, Italy, 2022, pp. 338–345 DOI:10.1109/ICCA54724.2022.9831833.
- G. Tom, S. P. Schmid, S. G. Baird, Y. Cao, K. Darvish, H. Hao, S. Lo, S. Pablo-García, E. M. Rajaonson, M. Skreta, N. Yoshikawa, S. Corapi, G. D. Akkoc, F. Strieth-Kalthoff, M. Seifrid and A. Aspuru-Guzik, Chem. Rev., 2024, 124, 9633–9732 CrossRef CAS PubMed.
- A. A. Volk and M. Abolhasani, Nat. Commun., 2024, 15, 1378 CrossRef CAS PubMed.
- K. Darvish, M. Skreta, Y. Zhao, N. Yoshikawa, S. Som, M. Bogdanovic, Y. Cao, H. Hao, H. Xu, A. Aspuru-Guzik, A. Garg and F. Shkurti, Matter, 2025, 8, 101897 CrossRef.
- H. Tao, T. Wu, S. Kheiri, M. Aldeghi, A. Aspuru-Guzik and E. Kumacheva, Adv. Funct. Mater., 2021, 31, 2106725 CrossRef CAS.
- T. Wu, S. Kheiri, R. J. Hickman, H. Tao, T. C. Wu, Z.-B. Yang, X. Ge, W. Zhang, M. Abolhasani, K. Liu, A. Aspuru-Guzik and E. Kumacheva, Nat. Commun., 2025, 16, 1473 CrossRef CAS PubMed.
- Y. Jiang, D. Salley, A. Sharma, G. Keenan, M. Mullin and L. Cronin, Sci. Adv., 2022, 8, eabo2626 CrossRef CAS PubMed.
- S. X. Leong, S. Pablo-García, B. Wong and A. Aspuru-Guzik, Matter, 2025, 8, 102331 CrossRef.
- A. Mirza, N. Alampara, S. Kunchapu, M. Ríos-García, B. Emoekabu, A. Krishnan, T. Gupta, M. Schilling-Wilhelmi, M. Okereke, A. Aneesh, M. Asgari, J. Eberhardt, A. M. Elahi, H. M. Elbeheiry, M. V. Gil, C. Glaubitz, M. Greiner, C. T. Holick, T. Hoffmann, A. Ibrahim, L. C. Klepsch, Y. Köster, F. A. Kreth, J. Meyer, S. Miret, J. M. Peschel, M. Ringleb, N. C. Roesner, J. Schreiber, U. S. Schubert, L. M. Stafast, A. D. D. Wonanke, M. Pieler, P. Schwaller and K. M. Jablonka, Nat. Chem., 2025, 17, 1027–1034 CrossRef CAS PubMed.
- K. M. Jablonka, L. Patiny and B. Smit, Nat. Chem., 2022, 14, 365–376 CrossRef CAS PubMed.
- T. Song, M. Luo, X. Zhang, L. Chen, Y. Huang, J. Cao, Q. Zhu, D. Liu, B. Zhang, G. Zou, G. Zhang, F. Zhang, W. Shang, Y. Fu, J. Jiang and Y. Luo, J. Am. Chem. Soc., 2025, 147, 12534–12545 CrossRef CAS PubMed.
- H. J. Byun and H. Nam, Electron. Lett., 2022, 58, 423–425 CrossRef.
- S.-H. Luo, J. Xu, W.-L. Wang, C.-R. Xiong, L.-P. Wang, Z.-Q. Tian and G.-K. Liu, J. Am. Chem. Soc., 2025, 147, 43964–43972 CrossRef CAS PubMed.
- J. Xu, S.-H. Luo, C.-R. Xiong, J.-H. Liu, Z. Zhao, W.-Q. Lin, W. Zhang, P. Guo, C. Pan, Q. Li, Z.-Q. Tian, B. Ren and G.-K. Liu, Anal. Chem., 2025, 97, 26141–26150 CrossRef CAS PubMed.
- J. Yi, E.-M. You, G.-K. Liu and Z.-Q. Tian, Nat. Nanotechnol., 2024, 19, 1758–1762 CrossRef CAS PubMed.
- L. Duponchel, R. R. D. Oliveira and V. Motto-Ros, Anal. Chem., 2025, 97, 6956–6961 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |