Open Access Article
Open Access ArticleAdvances in intelligent multi-mode lateral flow assays: from multi-metallic nanomaterials to smart analytical integration
Xinyi
Liang
a,
Hayoung
Kim
cd,
Thanh Mien
Nguyen
 e,
Kun
Wang
a,
Chengcheng
Li
a,
Seunghyun
Lee
e,
Kun
Wang
a,
Chengcheng
Li
a,
Seunghyun
Lee
 *bcd,
Jingbin
Zeng
*bcd,
Jingbin
Zeng
 *a and
Jaebum
Choo
*a and
Jaebum
Choo
 *e
*e
aState Key Laboratory of Chemical Safety, College of Chemistry and Chemical Engineering, China University of Petroleum (East China), Qingdao 266580, China. E-mail: zengjb@upc.edu.cn
bDepartment of Energy and Bio Sciences, Hanyang University ERICA, Ansan 15588, South Korea. E-mail: leeshyun@hanyang.ac.kr
cDepartment of Applied Chemistry, Hanyang University ERICA, Ansan 15588, South Korea
dCenter for Bionano Intelligence Education and Research, Hanyang University ERICA, Ansan 15588, South Korea
eDepartment of Chemistry, Chung-Ang University, Seoul 06974, South Korea. E-mail: jbchoo@cau.ac.kr
First published on 7th April 2026
Abstract
Lateral flow assays (LFAs) have evolved from simple qualitative tools into intelligent, multi-modal analytical platforms that integrate rationally engineered multi-metallic nanoparticles (MMNPs) with artificial intelligence (AI)-assisted data analysis to redefine the frontier of point-of-care diagnostics. This transformation has been driven by the advent of MMNPs, which couple plasmonic, catalytic, and magnetic properties within a single nano-system to achieve the tuneable synergistic enhancement of sensitivity, specificity, and dynamic range. The rational design of alloy, core–shell, hetero-structured, and hollow MMNP architectures allows simultaneous multi-signal readouts (e.g. colourimetric, fluorescence, chemiluminescence, surface-enhanced Raman scattering, photothermal, and electrochemical), thereby enabling intrinsic cross-verification and expanding diagnostic reliability. Parallel advances in AI, smartphone integration, and the Internet of Things connectivity have further elevated LFAs into digitally networked biosensors where embedded algorithms perform automated signal interpretation, error correction, and multi-mode data fusion, while cloud-linked infrastructures enable remote monitoring and epidemiological intelligence. These developments collectively reframe LFAs as integral components of data-driven, personalised, and preventive healthcare systems. Herein, we provide a unified framework that links design-on-demand MMNP synthesis, fully automated microfluidic LFA devices, AI-enhanced clinical decision support, and regulatory standardisation, and outline strategies for translating next-generation intelligent LFAs from laboratory innovation to global medical deployment.
1. Introduction
Advances in modern medicine are inextricably linked to the ability to perform accurate and timely diagnostic measurements. Healthcare needs are shifting from reactive treatment to proactive health management, and demand for preventive, personalized, and precision medicine is on the rise. This shift necessitates the development of diagnostic technologies that are not only accurate and reliable but also readily accessible, rapid, and information rich. For decades, the gold standard for bioassays has relied on sophisticated laboratory techniques such as cell culturing, polymerase chain reaction (PCR), and enzyme-linked immunosorbent assays. Although these methods are highly sensitive and specific, they require specialised equipment, trained personnel, centralised facilities, and long turnaround times and are therefore impractical for widespread use in point-of-care testing (POCT) settings such as in community clinics, pharmacies, and remote field environments, where rapid and accessible diagnostic results are paramount.1–3Lateral flow assays (LFAs) have long been at the forefront of POCT and have a long-standing history of use in biological testing (e.g. home pregnancy testing and initial screening for respiratory virus antigens) owing to their user-friendly nature, low cost, high detection speed, and minimal instrument requirements.4–6 The LFA test strip generally consists of five main components, namely a sample pad, conjugate pad, nitrocellulose (NC) membrane, absorbent pad, and background card. The NC membrane, pre-coated with test line (T-line) and control line (C-line), serves as the reaction core and result display area. From the perspective of recognition components, LFAs can be categorised into lateral flow immunoassays, which are based on antigen–antibody specific binding, and nucleic acid LFAs, which are based on Watson–Crick base pairing.7–11 Traditional LFAs typically use Au nanoparticles (NPs) as nanotags owing to their strong visible-range absorption, which generates colour signals observable by the naked eye. However, the human eye possesses a limited colour discrimination capability and heavily depends on subjective judgement. Consequently, direct visual observation suffers from low sensitivity and tends to provide false-negative results. This method is commonly applied to qualitative and semi-quantitative detection but proves inadequate for precise quantification, particularly at very low target concentrations.12,13
To fully realise the potential of POCT, LFA platforms must strictly meet three central criteria: speed (rapid turnaround time), reliability (consistent performance and minimal false results), and accuracy (high sensitivity and specificity). Thus, the LFA field is transitioning towards intelligent multi-mode detection systems. This advancement is driven by synergistic progress in two key technological domains, namely the rational design and controlled synthesis of novel multi-functional multi-component NPs, and the integration of artificial intelligence (AI)-driven intelligent analysis systems.14–18 Within the realm of multi-component NPs, multi-metallic NPs (MMNPs) represent a potent class of multi-functional materials integrating the properties of different metals while harnessing intermetallic synergies. These features enable MMNPs to outperform conventional single-metal nanoparticles in terms of optical characteristics, catalytic activity, and structural stability. The precise structural control achievable in MMNPs induces pronounced synergistic effects that give rise to distinctively superior performance. Specifically, these advantages are manifested in enhanced localized surface plasmon resonance (LSPR) for stronger optical signals,19 excellent catalytic amplification (e.g., enhanced nanozyme-like activity),20 improved energy conversion and electron transfer efficiency,21,22 unique magnetic responsiveness,23 and remarkably enhanced chemical stability.24 When used as probe materials in LFAs, MMNPs not only improve intrinsic colourimetric detection capabilities but also enable the generation of secondary, tertiary, or even multiple readout signals within a single test strip. This multi-mode integration broadens the detection range and facilitates complementary cross-verification between different modes, markedly boosting the sensitivity, accuracy, and reliability of LFAs to accommodate diverse application scenarios.
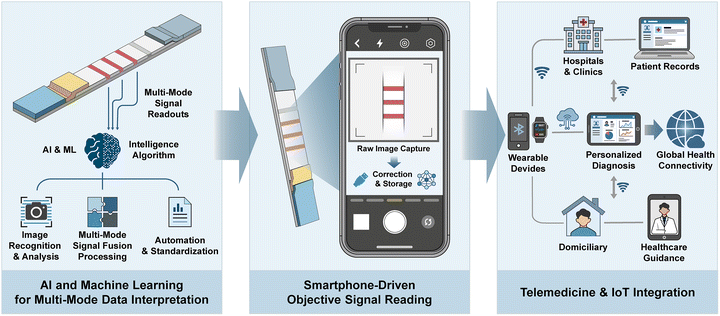
AI has experienced rapid progress, finding widespread daily-life application. The integration of LFAs with digital technology paves the way for truly intelligent POCT systems. The incorporation of portable readers, smartphones, and other connected devices renders test results more objective and quantitatively precise. The introduction of AI and machine learning (ML) algorithms enhances the analytical capabilities for processing high-dimensional and non-linear data. Intelligent algorithms can perform accurate quantitative analysis by identifying test strip characteristics, minimising human misinterpretation and adapting to environmental variations through real-time result calibration. These algorithms also enable the integrated analysis of multimodal signals, facilitate the batch processing of multi-dimensional data, and notably improve result handling efficiency. Furthermore, when combined with the Internet of Things (IoT) technology to upload test results to the cloud, LFAs can support remote medical guidance and epidemiological monitoring systems, propelling the field towards precision, automation, and full digital intelligence.
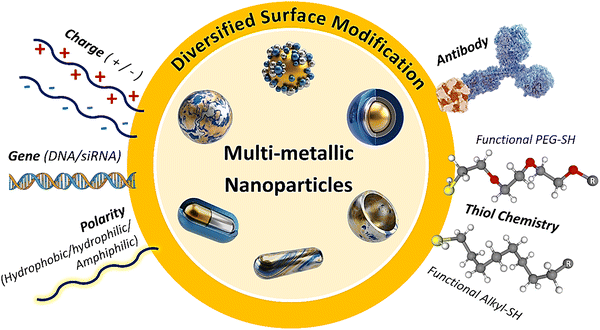
This review focuses on recent advances in intelligent multi-modal LFAs for precise diagnostics, as illustrated in Fig. 1. Beginning with the structure and properties of MMNPs, we explore how their structural synergies can be harnessed to enhance LFA sensitivity and multi-modal analytical performance, and we discuss how MMNPs enable the integration of multiple readout signals within a single LFA platform. This strategy facilitates the construction of robust multi-mode and multi-channel LFA systems, addressing the limitations of conventional single-mode LFAs to achieve greater accuracy and broader applicability (Section 2). We then examine the integration of intelligent analytics with LFAs, highlighting the role of AI-driven data processing, smartphone-based readers, and IoT-connected telemedicine technologies in enhancing their analytical performance and application scope (Section 3). Finally, we summarise the remaining challenges and future directions, outlining the development of next-generation intelligent multi-mode LFA diagnostic platforms to meet the demanding requirements of POCT requirements for “one test strip to address multiple scenarios and diverse clinical needs.” Although prior reviews have addressed nanomaterial-enhanced sensitivity and advances in LFA engineering separately, no comprehensive analysis has systematically mapped the integrated workflow from MMNP design, through multi-modal signal generation to AI-powered result interpretation within LFA systems. This review analyses the synergetic effects of material innovation, integrated multi-signal platforms, and state-of-the-art intelligent data analytics and smart devices, providing a currently lacking holistic perspective.
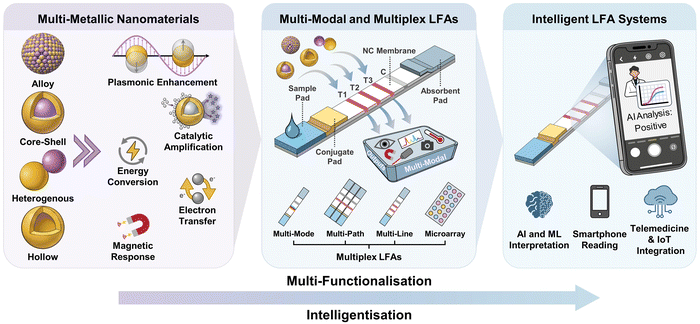 | ||
| Fig. 1 Schematic diagram of a multi-mode/multi-channel intelligent lateral flow assay system driven by MMNPs. | ||
2. MMNP-driven transition from single- to multi-modal LFAs
LFAs represent the simple, rapid, low-cost and user-friendly POCT method designed to achieve instantaneous readouts upon sample application, and are now widely applied in clinical diagnostics, food safety, environmental monitoring, and national security.25,26 Conventional LFAs typically use monodisperse AuNPs as nanoprobes for colourimetric detection. However, their intrinsic limitations in sensitivity, quantitative accuracy, and resistance to interference constrain their application in high-precision assays demanding high analytical performance, such as early disease screening or multiplex biomarker analysis.27 LFAs have experienced a continuous increase in accuracy, sensitivity, and information density,28,29 and the introduction of MMNPs has revolutionised them from simple qualitative tools into powerful quantitative analytical platforms.30 MMNPs rationally integrate two or more metallic elements within a single NP, not only effectively combining the properties of each component but also generating performance enhancements through structural synergies and even yielding in new functions. This structural synergy-driven enhancement has profoundly transformed the fundamental detection modes of LFAs, fostering their evolution from simplicity to sophistication and from single-signal detection to multi-signal integration.In this section, we systematically trace the evolution of MMNP-based LFA detection modes, focusing on how the structural and synergistic enhancement effects of MMNPs boost detection performance and broaden application scopes. Initially, we introduce the structural design and functional properties of key MMNPs. Subsequently, discussing how these multi-component nanostructures synergistically enhance the performance of various detection modes and enable new modalities that are otherwise unattainable or underutilised in conventional LFAs. Each detection mode offers advantages in sensitivity, specificity, and detection speed. However, reliance on a single mode yields one-dimensional signal output and increases susceptibility to environmental interference, thereby compromising overall accuracy and reliability. To achieve higher sensitivity, improved accuracy, and broader applicability while maximising the multi-functional integration capabilities of MMNPs, multi-mode LFAs integrating multiple detection modes have been developed. These systems enable multiple signal verifications for the same target or coverage of different sensitivity ranges within a single test. The exploitation of synergistic strengths across modes substantially enhances result reliability and detection scope. Building upon this foundation, multi-channel detection strategies that leverage the multi-functional properties of MMNPs further increase LFA throughput, allowing the parallel arrangement of multiple detection lines or reaction zones on a single test strip and enabling the simultaneous detection of different targets using minimal sample volumes. This presents innovative strategies for comprehensive pathogen profiling, infection source identification, and multiplex biomarker screening. Furthermore, aiming to bridge the gap between idealised laboratory designs and real-world POCT, we investigated how unique MMNPs and multi-mode LFAs successfully overcame severe matrix effects in complex samples, thereby validating their translational potential. Progressing from single-point detection to multi-dimensional sensing, parallel arrays, and ultimate clinical validation, the LFA technology is continuously advancing toward a more reliable, sensitive, and versatile diagnostic platform for integrated multiplex analyses.
2.1. Structurally engineered MMNPs for advanced LFA signal readout
The evolution of LFA readout from single-metal nanoparticles to structurally complex multi-metallic nanomaterials has shifted the paradigm from simple signal visualisation to mechanism-driven signal amplification. MMNPs represent a class of multi-component nanostructures formed by the integration of two or more metallic elements at the nanoscale through specific spatial arrangements. Unlike single-metal nanoparticles, MMNPs are not simple physical mixtures of different metals, but rather rationally engineered systems constructed through ordered design. This sophisticated integration not only combines the inherent properties of each component but also generates synergistic effects, endowing the NPs with superior and novel physicochemical characteristics unattainable by any single component alone.31 However, this approach also faces challenges, including the compatibility of different metallic components, uniformity of atomic distribution, stability of particle dispersion, and efficiency of molecular coupling.32,33 Moreover, the enhanced properties of MMNPs are intrinsically linked to their finely modulated structures. Structural parameters—such as diverse metallic composition, morphology, and size—determine a material's capacity to manipulate light, energy, and charge at the nanoscale, thus generating distinct enhancement effects. Consequently, strategically engineered MMNP structures utilise multiple synergistic mechanisms to effectively amplify signal intensity, lower detection limits, improve quantitative accuracy, and enhance detection stability, thereby meeting diverse environmental detection requirements. These advances address the limitations of conventional single-metal nanoprobes, such as challenges difficulties in precise quantification and low result reproducibility. Thus, a thorough understanding of the structure–property relationship of MMNPs is a fundamental prerequisite for the rationally design and construction of next-generation LFAs.This section first addresses the structural engineering of MMNPs, exploring the distinct advantages of different structural configurations for enhancing nanomaterial properties (Section 2.1.1). We then discuss rational surface modification strategies that confer biomolecular recognition capabilities, maintain colloidal stability, and minimise non-specific binding potential (Section 2.1.2), thereby enabling highly sensitive detection in complex matrices. Finally, we focus on the synergistic signal amplification mechanisms derived from the unique structure–function relationships of MMNPs (Section 2.1.3), providing a comprehensive understanding of the intricate interplay between nanostructures, enhanced properties, and signal generation/amplification. These insights offer valuable theoretical guidance for the design of multi-mode and multi-channel LFAs.
Alloy NPs. Alloy NPs are one of the most prevalent types of multi-metallic nanostructures in which two or more metallic elements are uniformly distributed at the atomic level within a single NP. This results in a uniquely tuneable electronic structure, and physicochemical properties intermediate between those of the constituent metals. This atomic level homogeneity gives rise to a LSPR peak that can be continuously and precisely tuned across a wide spectral range by adjusting the compositional ratio. For example, Au–Ag alloys bridge the LSPR peaks of Au (520 nm) and Ag (400 nm), thereby optimising absorption and scattering properties for specific light sources and producing distinct colouration.34,35 During the multi-metallic alloying process, lattice strain and ligand effects synergistically modulate the d-band centres and optimise the electronic structure of catalytic active sites, while optimising the adsorption energy of intermediates. This leads to significantly increased catalytic efficiency and conversion rates.36–38 The synergistic effects of multi-metallic alloying also reduce the electron transfer barrier, fine-tune the band structure and conductivity, and improve electrocatalytic activity and stability.39,40
Core–shell NPs. Core–shell nanostructures generate intense plasmonic coupling effects at their interfaces, exhibiting strong electromagnetic field enhancement that induces significant scattering and absorption.41,42 The LSPR peaks can be precisely tuned by manipulating the shell thickness and core–shell ratio. This not only results in vivid colour variations but also allows for the fine-tuning of the absorption profile and improving photostability to optimise the photothermal conversion performance of nanomaterials and generate stronger thermal signals.43–46 The deposition of a catalytically active shell layer onto a plasmonic core exploits both plasmon-enhanced catalysis and lattice strain effects to refine surface reactivity, thereby maximising catalytic utilisation.47–49 Additionally, core–shell MMNPs can also serve as efficient electron conductors and electrocatalysts, establishing gradient electron transport pathways that promote surface reactions at the electrode.50,51 Depositing metallic materials onto the surface of magnetic materials (such as Fe3O4) constructs a magnetic core–metal shell nanostructure, effectively preventing oxidation and corrosion of the magnetic core while simultaneously preventing self-aggregation to maintain stable magnetic properties.52,53
Heterostructures. Heterostructures such as Janus and dumbbell-like nanostructures create localised electromagnetic ‘hotspots’ at the junctions or interfaces between different metals. Within these regions, electromagnetic fields undergo exponential amplification, enabling the achievement of single-molecule detection sensitivity.54–56 Furthermore, heterogeneous MMNPs exhibit enhanced photothermal properties. They can generate coupling or superposition effects, broadening the absorption spectrum (from ultraviolet to near-infrared) to achieve a wider band of light capture.57–59 Constructing heterostructures promotes charge redistribution at interfaces, facilitates charge carrier separation, and allows the in situ integration of multi-step reactions to shorten diffusion pathways.60–63 Concurrently, the interconnection of metallic materials with distinct functionalities generates Schottky junctions or heterojunctions, driving directed electron flow and thus enhancing the electrochemical current response.64,65 Additionally, when a portion of the heterostructure comprises magnetic nanomaterials, this facilitates the spatial separation of the magnetic and plasmonic domains, thereby preventing the magnetic core from quenching optical properties, while delivering a dual-functional surface to accommodate diverse modification requirements.66–69
Hollow nanostructures. Hollow MMNPs exhibit significant red shifts and increased scattering cross-sections due to plasmonic hybridisation between the cavity's inner and outer surfaces. By adjusting shell thickness and cavity dimensions, they can provide highly tuneable plasmonic bands in the near-infrared region, yielding superior optical contrast compared to solid spheres.70,71 When hollow MMNPs absorb light energy, the heat generated by the photothermal effect is confined within the cavity and its surrounding micro-spaces through multiple internal reflections within light traps and a high surface-to-volume ratio. This enables localised micro-regions to achieve extremely high temperatures within a very short time, significantly amplifying the photothermal signal intensity.72–74 Moreover, hollow nanostructures possess ultra-high specific surface areas and internal cavities, providing abundant active sites both internally and externally for catalytic reactions. They can also confine substrates, increasing local concentrations and reaction probabilities to accelerate reaction kinetics via a caging effect.73,75,76 In addition, hollow and porous structures provide high electroactive surface areas, facilitating electrolyte permeation and charge transport, which improves the response speed and signal intensity of electrochemical detection.77,78
Thiol chemistry. The functionalisation of MMNPs is grounded in thiol chemistry, which leverages the high affinity between sulfur (S) atoms and noble metal surfaces. The bond energy of Au–S or Ag–S (approximately 40–50 kcal mol−1) approximates that of covalent bonds, allowing thiol-terminated ligands to effectively displace weakly adsorbed species like citrates and form robust self-assembled monolayers (SAM).79–81 A distinguished feature of this process is the precise control it affords ligand density and molecular orientation. Bifunctional linkers, such as SH–PEG–COOH, act as protective barriers against oxidation while the polyethylene glycol (PEG) chain length is tuned to optimize the distance between the metal surface's plasmonic field and external analytes.82 This optimization is critical for maximizing electromagnetic SERS enhancement while providing terminal carboxyl groups as chemically active sites for subsequent bioconjugation. Current strategies employ mixed SAMs composed of hydroxyl (–OH) and carboxyl (–COOH) terminated thiols to fine-tune the surface landscape, minimising non-specific adsorption while maximising target capture efficiency.83
Antibody immobilisation. The interface between MMNPs and antibodies is the primary determinant of analytical sensitivity and specificity. Passive physical adsorption often leads to protein denaturation or random orientation in which the antigen-binding sites face the particle surface, potentially resulting in the loss of 70% of the effective binding capacity.84 Furthermore, a critical spatial trade-off arises from the intrinsic dimensions of whole antibodies (approximately 10–15 nm), which often position the captured analyte beyond the effective electromagnetic near-field decay length (<10 nm) of plasmonic MMNPs.85 This distance-dependent attenuation represents a significant challenge for SERS-based detection, as it limits the achievable enhancement despite high-affinity biorecognition. Consequently, engineering the bio-interface to bridge this nanometre-scale gap has emerged as a key frontier for maximizing the sensitivity of MMNP-based optical sensors.86
To overcome these limitations, EDC/NHS carbodiimide coupling is employed to establish stable amide bonds between the primary amines (–NH2) of the antibody and the carboxyl groups on the MMNP surface. By meticulously adjusting the pH and ionic strength of the buffer, researchers can leverage the antibody's isoelectric point to induce electrostatic pre-concentration, significantly enhancing coupling efficiency. Such precision engineering is essential for the reproducible detection of ultra-low abundance biomarkers such as cardiac troponin I. Recent advancements have introduced site-specific immobilization using Protein A/G or carbohydrate moiety modification on the Fc region, ensuring that antibodies remain in a “vertical” orientation to maximize capture potential.87
Surface charge and polarity. Within the tortuous pore network of NC membranes, the transport and capture efficiency of MMNPs are fundamentally governed by the chemical identity and ionisation state of their surface ligands. Beyond a first-order DLVO description, the surface charge arises from the deprotonation of ionisable functional groups, such as carboxylates (–COO−) or sulfonates (–SO3−). For instance, the use of 11-mercaptoundecanoic acid or citrate capping agents ensures a high density of negative charges at physiological pH (pH > pKa).88 While traditional passive incubation is often used, potential-assisted deposition has emerged as a more rapid (within minutes) and more controlled method to achieve specific surface compositions of carboxylated thiols, which are otherwise difficult to incorporate into stable monolayers.89 This robust chemical stabilisation maintains a zeta potential typically below −30 mV, providing the necessary electrostatic repulsion to prevent van der Waals-mediated aggregation even in high-ionic-strength biological fluids like human serum.90
While charge ensures long-range stability, the surface polarity of MMNPs dictates their short-range interactions with the membrane and biological proteins. Non-DLVO forces, such as hydration repulsion, are modulated by the hydrophilicity of the interfacial layer. The introduction of PEGylation (via SH–PEG–COOH) or zwitterionic ligands (e.g., sulfobetaine thiols or L-cysteine) transforms the MMNP surface into a highly polar interface. Sum frequency generation vibrational spectroscopy reveals that these surfaces facilitate extensive hydrogen bonding with water molecules, forming a dense hydration shell characterized by a strong water signal at approximately 3200 cm−1.91,92 Unlike PEG, which often yields a more three-dimensional brush-like coating, zwitterionic ligands produce a thinner, monolayer-type “stealth” layer. Resistance to non-specific protein adsorption (the “protein corona”) is driven by the prevention of ion pairing between the protein and surface charges, which would otherwise release counterions and water molecules—a process that is entropically favourable for fouling.93 These polar barriers are essential for reducing background noise in SERS-based LFAs.94
However, in MMNP systems, surface charge and polarity are not uniformly distributed but are influenced by the atomic arrangement of the multi-metal surface. Unlike monometallic particles, MMNPs exhibit facet-specific chemical affinities; for example, thiol-terminated ligands bind more preferentially to gold-rich facets than to silver or platinum domains. Moreover, if mixed ligands exhibit a chain length difference of more than four carbon atoms, phase segregation can occur on the surface.95 On non-spherical morphologies such as nanorods, thermodynamically incompatible surfactants can self-organize into alternating striped patterns perpendicular to the rod axis.96 This “chemical anisotropy” leads to localized variations in surface polarity and charge density, which significantly influences hydrodynamic drag and how the particles interact with the dipolar sites of NC fibres. Consequently, achieving consistent assay reproducibility requires a strategic selection of mixed SAMs to homogenize the chemical landscape and optimise the density of bioreceptor molecules for specific capture at the test line.97
DNA/RNA immobilisation. The emerging frontier in MMNP surface engineering transcends traditional gene detection, evolving into a multifaceted functional nucleic acid (FNA) interfacing strategy. By grafting thiolated DNA or RNA probes at high densities, MMNPs are transformed into spherical nucleic acids that possess physicochemical properties far superior to their linear counterparts.98 A key advantage of this approach is the integration of aptamers—single-stranded oligonucleotides that function as “chemical antibodies”.99 Aptamers provide high-affinity, high-stability target capture for a wide range of analytes, including proteins and small molecules, while exhibiting greater resistance to thermal and chemical denaturation compared to protein-based antibodies. This ensures the structural integrity and long-term reliability of the LFA sensor in diverse environmental conditions.100 Furthermore, this strategy enables DNA-directed assembly, in which the programmable nature of DNA hybridisation is used as a molecular adhesive to create precision nano-arrays.101 By meticulously controlling inter-particle distances at the nanometre scale—often through the use of poly-A spacers—researchers can optimize the density of SERS “hotspots” and improve the overall sensitivity of the platform.102
Plasmonic enhancement. The compositional richness and morphological diversity of MMNPs confer a uniquely tuneable LSPR effect, enabling precise regulation of LSPR peaks across a broad spectral range from visible to near-infrared light. This capability is difficult to achieve with single-metal materials. Therefore, MMNPs facilitate precise LSPR tuning and maximise localised electromagnetic field intensity. This firstly yields richer, more vivid colour shifts in LFA colourimetric modes for enhanced sensitivity towards low-abundance targets.103–105 It is worth noting that Lee et al.106 recently pioneered a novel, highly sensitive and naked-eye LFA via plasmonic scattering. The excellent scattering properties of MMNPs also present significant application potential in this plasmonic light scattering mode of LFAs. Secondly, the strong localised electromagnetic fields generated by MMNPs also significantly enhance the excitation efficiency and radiative rate of nearby fluorophores, greatly improving fluorescence intensity and photostability. This phenomenon is known as metal-enhanced fluorescence (MEF).107–110 Spectral overlap and distance between the fluorophore and metal are considered key factors in the MEF effect. By modulating the SPR peaks of MMNPs through alterations in their size, shape, and composition to match the excitation/emission spectra of different fluorophores, and by precisely controlling the distance between the metallic core and the fluorophore, optimal enhancement effects can be achieved. Moreover, the plasmonic enhancement mechanism also serves as the foundation for SERS-based LFAs. This enhancement relies critically on nanoscale structures, which determine the intensity and distribution of the electromagnetic field. Specifically, features such as nanogaps, sharp tips, and structures with minimal curvature radii are pivotal for generating strong hotspots. Ordered periodic assembly of nanoparticles into two- or three-dimensional superlattices enables precise engineering of plasmonic hotspots, yielding enhanced intensity and superior spatial uniformity. MMNPs offer greater design flexibility for SERS substrates and can generate stronger, more stable enhancement effects, leading to more sensitive and reliable SERS signal readouts.111–116
Catalytic amplification. MMNPs with superior catalytic activity can serve as highly efficient nanozymes, replacing natural enzymes to catalyse reactions and achieve secondary signal amplification. First, MMNPs can be used for catalytically enhanced colourimetric mode. Upon the capture of the target by MMNPs and its immobilisation on the T-lines, additional chromogenic substrates are added to trigger highly efficient catalytic reactions, generating abundant chromogenic products. This effectively amplifies the initial colourimetric signal. The common chromogenic substrates include 3,3′,5,5′-tetramethylbenzidine,36,60,117,118 3-amino-9-ethylcarbazole,38,119–121 3,3′-diaminobenzidine,122–124 and 4-chloro-1-naphthol/3,3′-diaminobenzidine.125 Different amplification systems can be selected depending on the colourimetric response of the substrate and the specific properties of the nanoprobes used. Concurrently, it may be applied in chemiluminescent mode as a substitute for horseradish peroxidase, catalysing the oxidation of luminol by hydrogen peroxide to generate light radiation.126,127 This effectively overcomes the inherent susceptibility of natural enzymes to environmental factors such as pH and temperature, improving storage stability and batch-to-batch reproducibility.128,129
Energy conversion. The multi-component nature of MMNPs facilitates energy conversion processes, allowing excitation light energy to be transformed into detectable signals. Photothermal conversion represents a significant branch of this field, for which MMNPs exhibit strong absorption properties tuneable to the near-infrared spectrum. Harnessing their excellent photothermal effect, MMNPs can efficiently convert light energy into thermal energy. These thermal changes can be measured by high-precision temperature sensors (such as infrared thermal imagers), and quantitative detection is achieved by monitoring the relationship between local temperature changes and analyte concentration.74,130,131 Meanwhile, the heating of target metallic nanoparticles by an external laser also induces rapid thermal expansion and contraction in the surrounding medium, thereby generating acoustic signals. These sonic vibration responses can be measured by acoustic sensors, enabling quantitative detection of targets.132,133 Photothermal and photoacoustic detection modes can circumvent inherent spectroscopic interference from complex biological matrices, offering high signal-to-noise ratios and superior sensitivity. Apart from quantitative signals, the high temperature generated by the photothermal effect can also inactivate bioactive substances such as nucleic acids and proteins in bacterial cells and biofilms, and distort or even rupture bacterial cells, thereby killing bacteria and effectively preventing bacterial contamination.44,73,134 In addition, when an excited-state energy donor (such as dye molecules or quantum dots) approaches the surface of MMNPs, the donor's energy undergoes non-radiative transfer to the nanoparticles, quenching its own fluorescence. This nanometal surface energy transfer process can be controlled by adjusting the distance and dielectric environment to achieve sensitive detection.135 When an excited donor fluorophore is in extremely close proximity to an acceptor nanoparticle (typically <10 nm), sufficient overlap between the donor's emission spectrum and the acceptor's absorption spectrum allows energy to transfer via non-radiative dipole–dipole coupling from the donor to the acceptor, resulting in diminished donor fluorescence. This FRET effect is also frequently used to design LFAs.136–138
Electron transfer enhancement. Another significant advantage of MMNPs involves enhanced electron transfer. By reducing energy barriers, enhancing electron mobility, or leveraging synergistic redox activity, these MMNPs promote efficient charge transfer at the nanoparticle–electrode interface, thereby yielding a significantly enhanced electrical signal output. Incorporating electrodes into LFAs generates measurable electrical signals (current, potential), thereby enabling electrochemical detection.139 One strategy for generating electrical signal changes in test strips involves acid dissolution releasing metal ions to the electrode for redox reactions, followed by electrochemical detection.140–142 However, the addition of strong acid solution may damage the NC membranes and immune-sensing layers on the sensor surface, and direct electron transfer of electroactive species is only effective within a 1–2 nm range of the electrode. To avoid the need for manually adding toxic chemicals to dissolve metal nanoprobes, milder reagents, such as iodides, can be used to dissolve the corresponding metals.143 Another solution is to use metal nanoparticles to catalyse electroactive species, enabling redox reactions on the electrode surface to generate current, voltage, and other electrical changes.144–146 Consequently, the synergistic electronic effects of MMNPs lower the detection limit, facilitating precise quantification.
Magnetic response enhancement. The incorporation of magnetic metals (such as Fe, Co, Ni) into MMNPs confers a novel property upon the material—magnetism. This magnetic property enables manipulation and separation of the nanomaterial via external magnetic fields. It controls the flow velocity and direction of the nanoprobes, prolonging the interaction time between targets and capture probes.147–149 This facilitates efficient enrichment and purification of targets within complex samples, streamlines sample preparation, reduces matrix effects, and enhances signal-to-noise ratios.118,150–152 Simultaneously, it introduces novel magnetic response detection signals. As most biological samples exhibit magnetic transparency, the magnetic signal can generate background-free outputs, thereby circumventing optical background interference and enabling highly sensitive detection. Magnetic readout methods mainly include giant magnetoresistance,153,154 tunnelling magnetoresistance155 and anisotropic magnetoresistance156 methods based on magnetoresistance;157 magnetic particle quantification methods based on frequency mixing technology;158–161 magnetic assay reader methods based on magnetic flux;149,162,163 magnetic resonance coil magnetometer methods based on magnetic permeability;164 and atomic magnetometer methods based on the quantum optical effect.165 This dual capability makes MMNPs not only highly efficient nanoprobes but also essential tools for sample-to-result integrated detection, proving crucial for detecting trace analytes in complex environments.
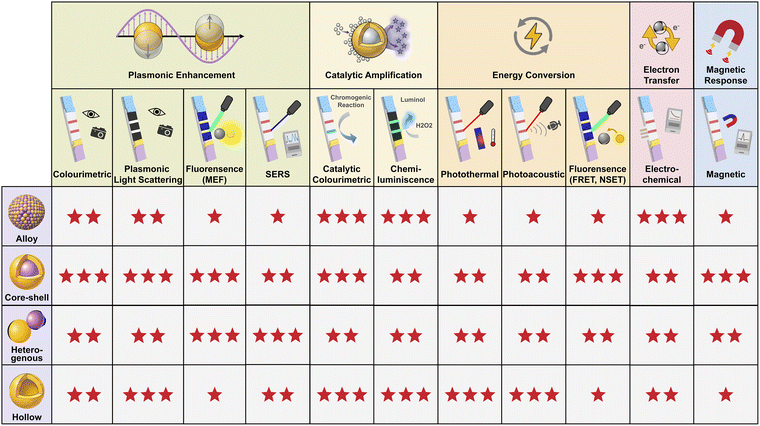
To summarise, MMNPs leverage structural features such as alloying, core–shell encapsulation, hetero-coupling, and cavity formation between different metallic components. Through synergistic enhancement mechanisms including plasmonic enhancement, catalytic amplification, energy conversion, electron transfer, and magnetic response, they exhibit significantly enhanced optical, catalytic, electrical, and magnetic properties that far exceed the simple sum of their component parts. These upgraded properties precisely activate and amplify distinct readout signals, enabling a series of enhanced detection modalities: colourimetric, fluorescent, chemiluminescent, SERS, plasmonic light scattering, photothermal, photoacoustic, electrochemical, and magnetic response (advantages and disadvantages of each detection mode are shown in Table 1). With the in-depth development and optimisation of each mode, they collectively form a robust ‘toolkit’ for LFA technology. We investigated the compatibility between different detection derived from various signal amplification mechanisms and various MMNP nanostructures and graded them accordingly. The results are summarised in Fig. 3, which employs a star rating system to identify the degree of compatibility (★ = acceptable, ★★ = good, ★★★ = most suitable). Appropriate detection modes or multiple combinations can be selected as required, laying a solid methodological and theoretical foundation for subsequent innovations in multi-mode/multi-channel detection. Nevertheless, it is important to recognise that the sophisticated structural engineering required to achieve these advanced functionalities inherently involves laborious, multi-step and technically demanding synthetic procedures.166–168 Achieving precise structural control at a commercial scale while ensuring reproducibility, minimizing batch-to-batch variability, maintaining long-term stability, and controlling production costs remains a significant challenge, which will be further discussed in detail in Section 4.
| Detection modes | Advantages | Limitations |
|---|---|---|
| Colourimetric | Simple operation, intuitive readout, and low cost. | Low sensitivity, poor quantitative accuracy, and difficulty in detecting coloured samples. |
| Catalytically enhanced colourimetric | Tuneable signal intensity and sensitivity exceeding that of direct colourimetry. | Complex operating procedures, unstable chromogenic substrates, and stringent reaction conditions. |
| Fluorescent | Accurate and highly sensitive quantitative detection. | Fluorescence reader and careful fluorescence design are required. Results can be affected by sample autofluorescence. |
| Chemiluminescent | No need for excitation light sources, reduced optical background noise, and high signal-to-noise ratios. | Cumbersome operation, short signal duration requiring timely, and limited stability. |
| SERS | High spectral resolution, sensitivity, and interference resistance. | Professional Raman spectrometer is required. Signal intensity depends on the reproducibility of nanotag preparation. |
| Plasmonic light scattering | Stable non-quenchable signals with no need for additional molecular modification steps. | New approach that requires optimisation for practical implementation. |
| Photothermal | Unaffected by matrix colour or turbidity and suitable for complex biological samples. | Laser light source and thermal signal detection equipment are required. |
| Photoacoustic | Extension of photothermal mode with high penetration depths and a signal-to-noise ratios. | Expensive and technically complex detection instrumentation is required. |
| Electrochemical | Rapid analysis, high sensitivity, and easy integration into miniaturised and automated systems. | Often shows limited reproducibility and stability, and electrode integration on strips can be complicated. |
| Magnetic | High signal-to-noise ratios, negligible magnetic background from biological samples, and magnetic enrichment for enhanced sensitivity. | Specialised magnetic sensors and trained personnel are required for operation and data analysis. |
2.2. Multi-functional platform for integrated multi-modal detection
Although single-mode detection methods each possess unique advantages, no single detection technique can achieve optimal performance across all parameters (e.g., sensitivity, specificity, interference resistance, and portability) (Table 1). To overcome the inherent limitations of a single-mode detection, two or more independent signal amplification strategies are increasingly integrated onto a single detection platform. Through synergistic and complementary interactions between detection modes, highly efficient multi-mode LFAs are established. This multi-mode integration approach represents the evolution of LFA technology towards better reliability, richer information dimensions, and stronger environmental adaptability, achieving cross-validation of detection results and more comprehensive analytical insights. MMNPs integrate diverse material properties within a single particle through rational structural design and can even generate novel functionalities owing to synergistic effects among their components. This provides a key driving force for the development of multi-mode LFAs, enabling different detection modes to be efficiently connected on a single strip.169–172 The selection of MMNPs also directly determines the available detection modes for LFAs, enabling the customisation of nanoprobes according to the characteristics of the analyte and the application scenario. Thus, this section will sequentially explore the design strategies, synergistic mechanisms, and application examples of dual-, tri- and quad-mode LFAs. It will elucidate how MMNPs can be multi-functionally combined to facilitate multi-mode integrated detection, offering powerful avenues for enhancing the accuracy, reliability, and quantitative capabilities of LFAs.To begin with, in order to attain superior naked-eye detection performance, we can build upon the traditional colourimetric mode by supplementing the nanoprobes with nanozyme components to endow LFAs with a second signal—a catalytically-enhanced colourimetric mode. The primary colourimetric signal is for direct qualitative or semi-quantitative analysis, while the secondary catalytically enhanced signal gives high-sensitivity quantitative readouts. For example, Shu and colleagues36 demonstrated a label-free dual-mode LFA for Salmonella typhimurium detection, through the strategic use of trimetallic PtMnIr nanozymes for their dual catalytic and adhesive functions. This assay demonstrated excellent analytical reliability, as evidenced by its high specificity, accurate performance in spiked sample analyses, and satisfactory recovery rates. Bai et al.47 prepared Au@Ag-Pt NPs with a rattle-like structure that possessed peroxidase-like activity while retaining plasmonic properties and intense colours within the visible spectrum, thereby constructing a dual-mode LFA (Fig. 4A). This approach delivers more sensitive and accurate colourimetric signals. However, it requires the secondary addition of reagents such as chromogenic substrates, which increases operational complexity and poses challenges for practical application. The most valuable aspect lies in augmenting the fundamental colourimetric mode with a precise quantitative method—a combination termed ‘Colourimetric Plus’. This fusion significantly enhances the sensitivity and accuracy of LFA without compromising testing convenience.
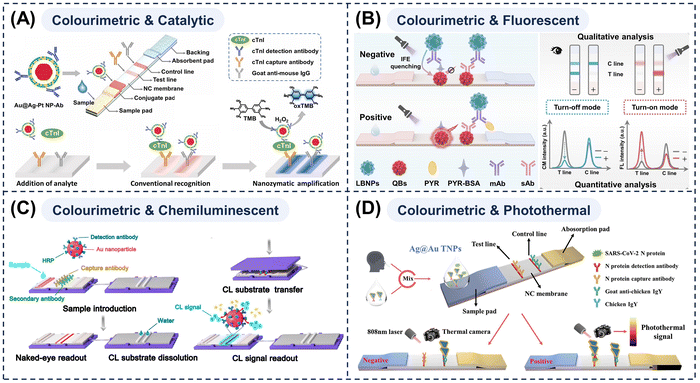 | ||
| Fig. 4 (A) Schematic of Au@Ag-Pt NP-based LFA strips and operational procedure highlighting the colourimetric and nanozyme-assiated amplification. Reprinted with permission from ref. 47. Copyright 2022, Elsevier. (B) Test principle and qualitative/quantitative readout of dual-mode LBNP-based LFA strips. Reprinted with permission from ref. 173. Copyright 2024, Elsevier. (C) Schematic operation workflow of a self-contained chemiluminescent LFA strips. Reprinted with permission from ref. 175. Copyright 2018, American Chemical Society. (D) Schematic detection principle of a colourimetric–photothermal dual-mode LFA strip employing Ag@Au triangular nanoplates. Reprinted with permission from ref. 43. Copyright 2024, American Chemical Society. | ||
First, nanomaterials that possess both intrinsic colour and fluorescence, such as noble metal core–shell nanostructures can be employed.108,173 The colourimetric–fluorescent dual-mode LFA system delivers instant qualitative colourimetric results without any equipment, as well as highly sensitive, low-background quantitative fluorescent signals to meet diverse application situations. Meanwhile, the inherent ‘signal on/off’ verification considerably reduces the probability of false positives and false negatives.174 Shu et al.173 employed novel litchi-like Au–Ag bimetallic nanospheres (LPNPs) as fluorescence quenchers (Fig. 4B). These LPNPs possessed broadband absorption with peaks that highly overlapped the excitation and emission peaks of quantum dots. This property enabled the establishment of an LFA platform with dual readouts (colourimetric and fluorescence quenching) operating in dual modes (off and on). Fluorescent detection is a mature and widely available POCT method, making colourimetric–fluorescent dual-mode LFAs well-suited for low-concentration pathogen detection.107 However, their application in complex matrices is limited by fluorescent signal instability (e.g., photobleaching and flickering) and background autofluorescence.
Chemiluminescence requires no external light source, exhibits low background, and minimises nonspecific signals, resulting in an exceptionally high signal-to-noise ratio and sensitivity. These attributes make it an ideal partner for colourimetric readouts, enabling dual-mode LFAs that provide more accurate qualitative/quantitative detection across a broader linear range. Roda et al.127 developed a dual optical/chemiluminescent LFA immunosensor for detecting SARS-CoV-2-specific immunoglobulin A in serum and saliva. They incorporated a transparent glass fibre pad pre-loaded with lyophilized chemiluminescent substrates (sodium perborate, luminol, and p-iodophenol) as described previously (Fig. 4C).175 After routine visual analysis, the pad was placed on the NC membrane and dissolved. Upon dissolution, the released substrates subsequently reacted under HRP catalysis to generate a chemiluminescent signal for quantification. Therefore, colourimetric-chemiluminescent dual-mode LFAs are well-suited for developing low-cost, integrated automated devices owing to their high sensitivity, broad linear range, and simple optics. However, the requisite additional reagent-loading steps impede device miniaturisation, rendering them less suitable for minimalist, portable POCT intended for resource-limited settings.
If nanoprobes with high photothermal conversion efficiency (e.g., nanorods or hollow nanostructures) are used, the T-line is irradiated with a near-infrared laser following the initial colourimetric readout. The nanoprobes convert the light energy into thermal energy, causing a localised temperature increase. This photothermal signal can be quantitatively measured by infrared thermal imaging cameras or low-cost thermal sensors. In our group, we first developed Ag@Au triangular nanoplates (Ag@Au TNPs), which exhibit tuneable plasmonic absorption and a high photothermal conversion efficiency of 61.4%, enabling dual-mode LFA detection of SARS-CoV-2 nucleocapsid protein (Fig. 4D).43 To further amplify the signal, we subsequently engineered nanoprobes by densely loading Au nanoshells onto Fe3O4 nanoparticles. This design enabled magnetic enrichment and separation, thereby integrating target preconcentration with enhanced photothermal/colourimetric detection in a single platform.53 Photothermal results remain unaffected by sample colour or turbidity, and are highly resistant to interference, making them suitable for complex matrix samples. Moreover, photothermal signals can be measured by portable infrared thermometers, enabling on-site quantitative analysis.176,177 Nevertheless, they are susceptible to environmental thermal noise (such as ambient temperature fluctuations and air currents), along with the photothermal conversion efficiency of the material and thermal diffusion effects, all of which necessitate reference calibration.
MMNPs also exhibit powerful SERS effects, thereby significantly enhancing the Raman signals of reporter molecules adsorbed on their surfaces. For colourimetric–SERS dual-mode LFAs, apart from their inherently strong colour signals, they can provide near-background-free, highly specific quantitative signals.116,178,179 A prime example of such engineered MMNPs is the AgMBA@Au probe developed in our group.112 We employed a ligand-assisted epitaxial growth strategy, utilising sulfite coordination to lower the redox potential of gold and prevent oxidative etching of the silver core. This allowed for the precise construction of a core–shell nanostructure featuring a silver core, an ultrathin gold shell (∼2 nm), and the Raman reporter molecule (4-mercaptobenzoic acid) embedded within the gap (Fig. 5A). This unique architecture provides both intense electromagnetic enhancement and excellent signal stability, making it an ideal SERS tag for quantitative dual-mode biosensing. The coating of the Au shell not only enhanced stability but also generated a strong electromagnetic field enhancement at the gap, which effectively improved the SERS properties of the nanoprobes. Furthermore, by loading AgMBA@Au NPs onto Fe3O4, we further integrated magnetic functionality into the platform.180 This not only amplified the SERS signal through nanoparticle assembly but also enabled efficient magnetic enrichment, which significantly reduced matrix interference and allowed for highly specific and sensitive differential diagnosis of multiple pathogens in complex biological samples (e.g., whole blood). However, as SERS signals critically depend on the homogeneity and aggregation state of the nanostructure substrate, challenges arise in preparation and batch-to-batch control. Moreover, the requirement for specialised optical systems and spectrometers makes the system costly and operationally complex, currently limiting its practical application in routine clinical or field settings.
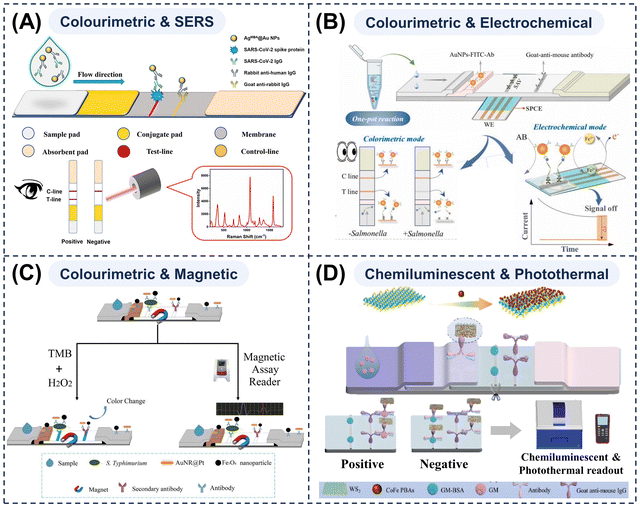 | ||
| Fig. 5 (A) Schematic structure and detection principle of a colourimetric–SERS dual-mode LFA strip. Reprinted with permission from ref. 112. Copyright 2022, American Chemical Society. (B) Schematic illustration of colourimetric and electrochemical LFA strips for accurate quantification of Salmonella. Reprinted with permission from ref. 144. Copyright 2025, Elsevier. (C) Schematic of the catalytic-magnetic dual-mode LFA strips for Salmonella typhimurium detection. Reprinted with permission from ref. 149. Copyright, 2022 Elsevier. (D) Schematic of a chemiluminescent-photothermal dual-mode LFA strip for gentamicin detection. Reprinted with permission from ref. 183. Copyright, 2024 Elsevier. | ||
In addition to the combination of dual optical modes, electrochemical modes may also be incorporated into LFAs. By integrating microelectrodes onto test strips, it enables the simultaneous capture of naked-eye colour signals and the measurement of electrical changes in current, potential, or impedance generated by nanoprobes on T-lines via electrochemical sensors on the same platform. This allows for the synchronous readout of highly sensitive quantitative signals. Chen et al.144 developed a streamlined, integrated RPA-CRISPR/Cas12a electrochemical LFA for the precise detection of Salmonella (Fig. 5B). AuNPs functioned concurrently as colourimetric signal sources and catalytic labels for in situ redox reactions. The redox reaction involved the reduction of ferricyanide to Fe2+ by ammonium borohydride in the presence of AuNPs, thereby significantly increasing the current output. This innovative approach facilitates the rapid, semi-quantitative and quantitative detection of Salmonella in diverse application scenarios. Electrochemical modes are highly sensitive and selective, and the associated readout devices are readily miniaturised and integrated, making them well suited to rapid testing applications requiring high sensitivity.141,146,181 However, owing to the complexity of integrating electrodes onto test strips, and the susceptability of electrodes to contamination and passivation in complex matrices, their stability and reproducibility remain suboptimal. Further optimisation is therefore required before large-scale field applications can be realised.
Similarly, when magnetic MMNPs are employed as nanoprobes, alongside conventional colourimetric results, magnetic signal outputs can be obtained via magnetic sensors. Simultaneously, an external magnetic field can enrich and purify targets, reducing matrix interference and significantly improving signal-to-noise ratios. It can also control the chromatographic process and accelerate flow. Du et al.149 developed a dual-mode LFA that leverages a nanozyme-assisted signal amplification strategy (using AuNR@Pt for colourimetry) combined with a magnetic nanoparticle-based flow control strategy (using Fe3O4 for magnetometry) for the detection of Salmonella typhimurium (Fig. 5C). Zheng et al.182 also developed a novel colourimetric-magnetic dual-mode LFA based on Fe3O4@Pt NPs. However, owing to the specialised and expensive magnetic sensors, the magnetic properties of MMNPs are currently seldom employed directly for quantification. Instead, they primarily serve as a strong sample preparation tool.53,58 After enrichment, colourimetric, fluorescent, photothermal or other signal readouts are readily obtained, thereby facilitating the implementation of highly sensitive and low-background LFAs.
Other combinations of detection modes are also possible. For instance, Wu et al.183 constructed a sensitive, portable dual-mode LFA that combines chemiluminescent and photothermal readouts (Fig. 5D). By anchoring CoFe Prussian blue analogue nanozymes onto high-surface-area WS2 nanosheets, they prepared a multi-component nanocomposite. This material effectively mediated both the enhancement of chemiluminescence in the luminol-H2O2 system and the generation of photothermal signals in the TMB–H2O2 system. Thereby, this platform merges the high sensitivity of laboratory analyses with the portability required for on-site testing. In another work, Sun and colleagues184 immobilised Raman dyes onto Au@Ag NPs, which displayed intense SERS signals and characteristic electrochemical redox peaks. Based on this design, they fabricated an electrochemical-SERS dual-mode LFA. The resulting platform demonstrated high sensitivity, high selectivity, a low detection limit, and long-term stability. The realisation of these novel dual-mode LFA systems hinges critically on the sophisticated design and controlled synthesis of multi-functional nanomaterials. To translate them from proof-of-concept into practical applications, collaborative innovation is equally required in engineering integrated devices that streamline sample handling, detection, and intelligent readout, with the ultimate aims of simplifying operation, enabling intuitive interpretation, and achieving full miniaturisation.
Dual-mode detection ingeniously integrates the composition and functionalities of MMNPs, combining colourimetric, fluorescent, SERS, photothermal, electrochemical, and other modes to confer traditional LFAs with unprecedented analytical capabilities upon traditional LFAs. This integration enriches data through complementary qualitative and quantitative information and ensures robustness via internal cross-validation. Thus, it bridges the gap between rapid, low-cost screening and accurate quantitative analysis.
Hierarchical multi-modal LFAs for performance enhancement. One representative strategy uses MMNPs simultaneously exhibiting catalytic activity and photothermal effects to construct colorimetric–catalytic–photothermal tri-mode LFAs. In such systems, three levels of integration are realised, namely visual observation, catalytic signal amplification, and instrumental photothermal quantification. The intrinsic colorimetric mode offers intuitive qualitative or semi-quantitative readouts for rapid on-site interpretation; catalytically enhanced colorimetric signals intensify colour via nanozyme-mediated substrate oxidation, markedly lowering limits of detection (LODs); and the photothermal mode adds an interference-resistant quantitative signal that can be measured using portable infrared thermometers, greatly improving the reliability and accuracy of field quantification and making these assays particularly suitable for on-site monitoring requiring high-precision quantitative results.
Li et al.185 proposed a tri-mode LFA based on onion flower-like Au–Pd NPs integrating colorimetric, catalytically enhanced colorimetric, and photothermal modes (Fig. 6A). The bimetallic/polydopamine composition, multi-branched morphology, and broad absorption profile endowed these MMNPs with excellent colorimetric performance, catalytic activity, and photothermal properties, enabling ultra-sensitive detection through the cross-validation of all three modes. Xu et al.186,187 developed LFAs for respiratory syncytial virus detection using wheatgrass-like MoSe2@Pt heterojunctions and Fe3O4@MoS2@Pt-based LFAs for the detection of the SARS-CoV-2 nucleocapsid protein and influenza A virus, thereby realising integrated colorimetric–catalytic–photothermal tri-mode detection by combining catalytically active metals with photothermal materials.
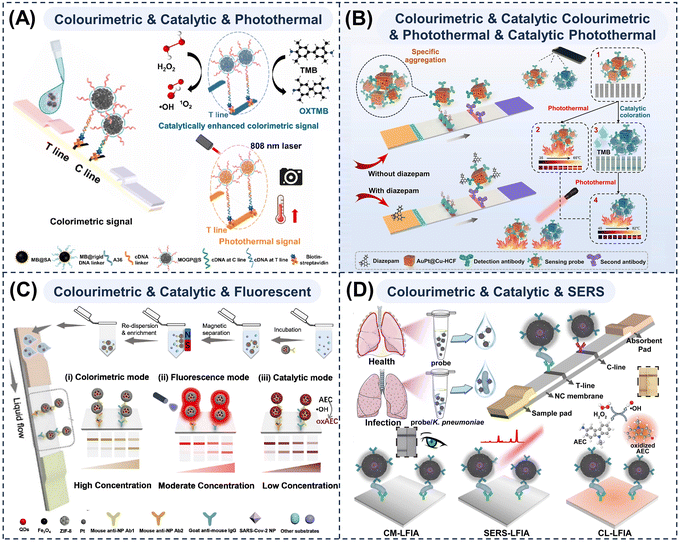 | ||
| Fig. 6 Schematic diagrams of multi-mode LFAs for enabling cross-validation and signal amplification. Schematic illustrations of (A) a complementary colourimetric–catalytic–photothermal tri-mode LFA strip based on Au–Pd nanozymes. Reprinted with permission from ref. 185. Copyright 2025, Elsevier. (B) a quad-mode LFA strip featuring colourimetric–catalytic colourimetric–photothermal–catalytic photothermal readouts, exploiting cascade nanozyme-mediated signal amplification. Reprinted with permission from ref. 189. Copyright 2018, Springer Nature. (C) a colourimetric–catalytic–fluorescent tri-mode LFA strip for the detection of SARS-CoV-2 antigen using SQF@ZIF-8/Pt NPs. Reprinted with permission from ref. 190. Copyright 2022, American Chemical Society. (D) an Au@Au@Ag/Pt probe-integrated colourimetric–catalytic–SERS tri-mode LFA strip for Klebsiella pneumonia detection. Reprinted with permission from ref. 192. Copyright 2025, Elsevier. | ||
Given that oxidised TMB (oxTMB) generated during nanozyme-catalysed TMB oxidation exhibits a strong photothermal conversion capability, laser irradiation applied after the catalytic colorimetric step can yield an additional photothermal signal. Liu et al.188 exploited this principle to design a tri-mode LFA for Staphylococcus aureus detection based on highly catalytically active Pd/Pt NPs, which catalysed the oxidation of TMB for a colorimetric readout and enabled the photothermal detection of oxTMB under excitation at 808 nm. Chen et al.171 synthesised core (Au)–shell (Mn) nanostructures for a colorimetric–catalytic–photothermal tri-mode LFA targeting Escherichia coli O157:H7. Both systems exploit the catalytic generation of photothermally active oxTMB. By integrating these four signals, Zhang et al.189 proposed a ‘colourimetric–catalytic colourimetric–photothermal–catalytic photothermal’ quad-mode LFA based on the copper hexacyanoferrate nanozymes doped with Au and Pt (AuPt@Cu-HCF) (Fig. 6B). This cascade amplification strategy illustrates how a single function (catalysis) can be harnessed to enhance two intrinsic modalities (colourimetric and photothermal responses), yielding a highly sensitive ‘four-in-one’ LFA platform with four tuneable detection ranges and detection limits tailored to diverse application scenarios.
The multi-mode configuration can be re-engineered by replacing the photothermal mode with the more sensitive and flexible fluorescence mode. This strategy requires nanoprobes that combine enzyme-mimicking catalytic activity with intrinsic fluorescence, for which composites of highly catalytic active metals such as Pt with fluorescent quantum dots or metal–organic frameworks (MOFs) are particularly attractive. For instance, Huang et al.190 designed a spatially hierarchical dual-porous nanostructure integrating colourimetric, fluorescent and catalytic functions for the rapid and sensitive detection of SARS-CoV-2 nucleocapsid protein (Fig. 6C). The hierarchical assembly of dendritic mesoporous SiO2 and MOFs enabled controlled loading of red-emissive quantum dots and Fe3O4 within the framework and promoted synergistic catalytic enhancement between Fe3O4 and Pt nanozymes. The spatial distribution and ratio of these three functional units were optimised to yield discrete sensing modes with continuous overlapping dynamic ranges and ensured high sensitivity even at low analyte levels, while overlapping fluorescence signals provided built-in self-verification and reduced external interference. Similarly, Sun et al.191 synthesised ZrFe-MOF@Pt NP nanocomposites via an immersion-reduction route. These nanocomposites exhibited the benefits of broad optical absorption, high peroxidase-like activity and solvent stability, and efficient antibody conjugation and were used to construct colorimetric-catalytic-fluorescence tri-mode LFAs.
The SERS mode targets ultra-high sensitivity and specificity in applications requiring molecular fingerprint recognition. In a colorimetric–catalytic–SERS integrated scheme, the intuitive visual readout and nanozyme-mediated colorimetric amplification are combined with the molecular specificity and ultra-high sensitivity of SERS to achieve the unambiguous identification of low-concentration pathogens in complex matrices. The realisation of this configuration demands the integration of high-quality SERS-active metal substrates with nanozyme components in a bifunctional MMNP architecture and the careful spatial arrangement of different metal elements because certain nanozyme materials are not favourable for SERS enhancement. Zhi et al.192 developed multilayer Au@Au@Ag/Pt NPs featuring a peroxidase-mimicking outer shell and a tuneable plasmonic nanogap and employed them as probes in an LFA for the tri-mode colourimetric–catalytic–SERS detection of Klebsiella pneumoniae in complex biological matrices (Fig. 6D). This system achieved a naked-eye LOD of 104 CFU mL−1, catalytic colourimetric LOD of 103 CFU mL−1, and SERS mode LOD of 38 CFU mL−1, thereby accommodating diverse sensitivity requirements within a single platform.
Multi-modal MMNP platforms for comprehensive optical biosensing. MMNP-driven multi-mode detection strategies enable rapid naked-eye screening (colourimetric), followed, where necessary, by precise quantification using portable instruments (photothermal, fluorescence, and SERS). Given that direct and catalytically enhanced colorimetric readouts are colour-based, earlier configurations essentially exploited only two signal dimensions from the nanotags. By contrast, more advanced tri-mode formats integrate three detection technologies based on distinct physical principles to enrich the dimensionality of analytical information.
A representative example is the combination of colorimetric, fluorescence, and SERS readouts, which jointly utilise optical intensity information (colour and fluorescence) and spectral frequency information (SERS) provided by the nanotags. In this configuration, the colorimetric channel ensures operational simplicity and immediate qualitative results, the fluorescence channel—being mature, highly sensitive, and widely supported by existing instrumentation—provides sensitive quantitative outputs, and the SERS channel offers unmatched specificity and ultra-high sensitivity for trace-level analysis in strongly interfering matrices, all without introducing extra reactions or substrates during measurement. The precise co-assembly of components with vivid colour, strong fluorescence, and robust SERS activity within a single NP allows these three optical modes to complement and reinforce one another, exemplifying an innovative strategy for maximising performance via multi-mode cooperative integration. Li et al.193 developed a tri-mode system for C-reactive protein detection using nano-assemblies of Eu chelate-doped polystyrene particles and Au NPs (Fig. 7A). Chen et al.194 also demonstrated such a system for microRNA detection in bodily fluids using LFAs co-loaded with upconversion NPs (UCNPs) and Au@Ag NPs. Au@Ag NPs acted as chromophores and SERS substrates, while their proximity to UCNPs induced fluorescence resonance energy transfer (FRET)-mediated fluorescence modulation for the third readout.
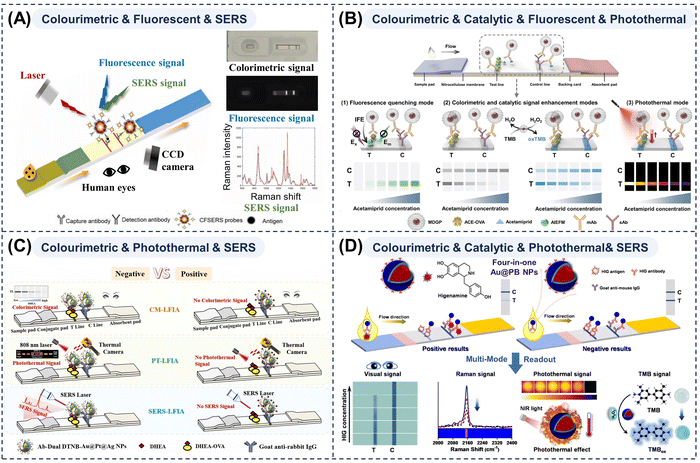 | ||
| Fig. 7 Schematic illustrations of (A) a colourimetric–fluorescent–SERS tri-mode LFA strip for determining C-reactive protein levels. Reprinted with permission from ref. 193. Copyright 2026, Elsevier. (B) a colourimetric–catalytic–fluorescent–photothermal multi-mode LFA strip for higenamin detection. Reprinted with permission from ref. 195. Copyright 2024, Wiley-VCH. (C) a colourimetric–photothermal–SERS tri-mode LFA strip using dual DTNB-Au@Pt@Ag NPs. Reprinted with permission from ref. 197. Copyright 2022, Elsevier. (D) a colourimetric–catalytic–photothermal–SERS quad-mode LFA strip for acetamiprid detection using a four-in-one multi-functional dandelion-like Au@Pt NP probe. Reprinted with permission from ref. 200. Copyright 2025, American Chemical Society. | ||
Wang et al.195 developed an on-site microanalyzer integrating a quad-mode LFA strip capable of colorimetric, catalytic, photothermal, and fluorescence readouts, utilizing multi-functional dandelion-like Au@Pt nanoparticles. This integration offers significant advantages, particularly in scenarios with optical background interference or when equipment portability is crucial (Fig. 7B). Through rational nanostructure design, they maximized the performance of each detection modality. The unique porous architecture provided a large specific surface area to enhance catalytic activity, while intraparticle coupling improved optical absorption efficiency, resulting in a high photothermal conversion efficiency of 65.84%. In addition, the broad absorption spectrum enabled fluorescence quenching based on dual-spectral-overlap, leading to a highly sensitive “signal-on” fluorescence response.
The colorimetric–photothermal–SERS architecture represents another flexible and powerful tri-mode union. Beyond the fundamental colorimetric function, it exploits the photothermal conversion and local electromagnetic field enhancement capacities of MMNPs under laser irradiation to generate photothermal and SERS signals, with all three readouts originating from the same NP and therefore being inherently consistent. These independent cross-validating channels substantially reduce the risk of false positives and negatives compared with single- or dual-mode assays and provide seamless coverage from rapid screening through portable quantitative measurement to fingerprint-level molecular confirmation, supporting an end-to-end field-to-laboratory diagnostic workflow. Core–shell,196,197 multi-branched198,199 or hollow70 NPs are particularly attractive because they often exhibit deeper colours, favourable photothermal behaviour, and strong SERS activity and have been used in tri-mode LFAs. Yang et al.197 synthesised Au@Pt@Ag core–shell–shell particles loaded with dual layers of the Raman reporter 5,5′-dithiobis(2-nitrobenzoic acid) (DTNB), whose broad ultraviolet-visible-NIR absorption and optimised shell thickness afforded a pronounced colour contrast, strong photothermal effects, and intense SERS signals and enabled the colorimetric–photothermal–SERS tri-mode LFA of dehydroepiandrosterone with tuneable LODs and detection ranges (Fig. 7C). Lin et al.169 created a multi-functional heterostructure by selectively growing Au nanostars on CuS nanoplates to generate a probe with plasmon-enhanced catalytic, photothermal, and SERS capabilities and used it in a tri-mode LFA for S. pneumoniae and K. pneumoniae with PCR-comparable accuracy and assay times under 15 min.
Ding et al.200 reported a quad-mode LFA incorporating colorimetric, catalytically enhanced colorimetric, SERS, and photothermal modes for ultra-sensitive higenamine detection (Fig. 7D). The corresponding nanoprobes comprised Au NP cores coated with Prussian blue (PB) shells: the Au NPs provided colour signals and served as SERS substrates, whereas PB contributed intrinsic peroxidase-like activity and NIR-driven photothermal effects. These probes generated Raman signals at 2153 cm−1 in the Raman-silent region, effectively avoiding background interference from complex matrices such as urine and food. This work shows that nanoprobe design must secure not only multiple readouts but also robust reliable performance for each mode under practical conditions.
Magnetically integrated multi-mode LFAs for end-to-end detection. Magnetic responses have also been incorporated into multi-mode LFAs by embedding sample pre-treatment directly within the strip, thus further streaming the overall detection process. The intrinsic magnetic properties of MMNPs enable both efficient magnetic separation and generation of ultra-low-background magnetic signals, providing significant advantages in sample purification, signal amplification, and detection of low-abundance pathogen. This strategy advances LFA systems towards truly “sample-in, answer-out” POCT devices, enhancing its practicality for real-world diagnostic applications. Liu et al.170 developed a multifunctional four-in-one nanotag composed of an Fe3O4 magnetic core, a bimetallic Au/Pt shell, and a Raman reporter, thereby endowing the probe with integrated magnetic, optical, peroxidase-like, and SERS characteristics (Fig. 8A). Following viral capture and magnetic separation, the nanotags provided three complementary readouts—colourimetric, catalytically enhanced colorimetric, and SERS—enabling the tri-mode LFA detection of the monkeypox virus A29L protein and the inactivated virus within 35 min. This system achieved a wide linear detection range and ultra-low quantification LODs of 0.0016 pg mL−1 and 17.724 pfu mL−1, respectively. Yang et al.201 also developed a colorimetric–magnetic–photothermal tri-mode LFA utilizing Fe3O4@Au particles that integrated the strong magnetic and NIR photothermal characteristics of Fe3O4 with the intrinsic colorimetric response of Au (Fig. 8B). These hybrid particles enabled magnetic isolation of biogenic amines from real samples while generating three complementary optical and magnetic signals that collectively enhances assay sensitivity. In practice, however, interpreting magnetic signals typically requires specialised magnetometry instrumentation; therefore, magnetic channels are seldom employed as routine quantitative readouts and more commonly serve to facilitate magnetic separation.
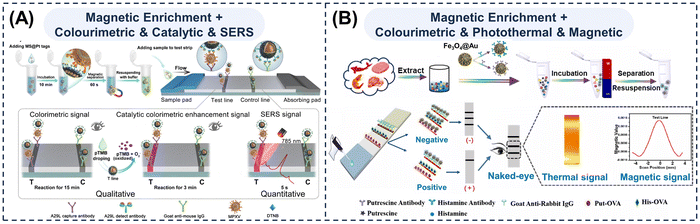 | ||
| Fig. 8 Schematic diagrams of multi-mode LFAs featuring full-process integration with magnetic pre-treatment. Schematic illustrations of (A) an MS@Pt-based tri-mode LFA strip incorporating magnetic enrichment for qualitative and quantitative detection of monkeypox virus A29L protein. Reprinted with permission from ref. 170. Copyright 2024, Elsevier. (B) an Fe3O4@Au-based LFA strip integrating magnetic enrichment with colourimetric, photothermal, and magnetic triple readouts. Reprinted with permission from ref. 201. Copyright 2023, Elsevier. | ||
The remarkable progress of multi-mode LFA systems is from the diversified structural designs and precise syntheses of MMNPs. By meticulously tailoring their composition and morphology to meet functional requirements, and by employing rigorously controlled synthetic methodologies, the distinct physicochemical properties of different metals can be integrated into highly potent multi-functional signal tags and amplifiers. This integrated functionality enables multi-mode LFAs to merge detection technologies based on different physical principles and to evolve from simple functional superposition with internal calibration towards genuine synergistic signal amplification and scenario-adaptive operation. Crucially, this structural and functional evolution empowers MMNP-based LFAs to fulfil the three principal criteria of advanced POCT. Specifically, synergistic signal amplification markedly improves sensitivity and specificity, ensuring far greater accuracy; cross-validation between different detection modes fortifies analytical reliability against complex sample matrices; and the consolidated single-strip format preserves the fundamental operational speed. Consequently, these systems significantly enhance sensitivity, specificity, and accuracy, while introducing new dimensions in instrumentation, quantification, and digitalisation—thus achieving a true 1 + 1 + 1 > 3 synergistic enhancement effect. To quantitatively support these claims, Table 2 summarised the analytical performance of representative MMNP-based multi-mode LFAs. As shown, the incorporation of multiple detection modes within MMNP-based LFAs represents a transition from simple qualitative rapid tests to powerful multi-dimensional information-acquisition platforms, providing unprecedented analytical capability for rapid in vitro diagnostics. By flexibly selecting the most suitable detection modes according to available instrumentation and required sensitivity, multi-mode LFAs can be tailored to demanding applications such as comprehensive pathogen serotyping, multiplex biomarker profiling in complex diseases, or highly reliable testing in resource-limited environments.
| Analysis method | MMNP type | Target biomarker | Detection mode & LOD | Sensitivity improvement | Real sample matrix | Ref. |
|---|---|---|---|---|---|---|
| Dual-mode | PtMnIr | Salmonella typhimurium | Colourimetric: 104 CFU mL−1 | Increased 50 times compared to AuNPs-LFA | Chicken and vegetable | 36 |
| Catalytic: 103 CFU mL−1 | ||||||
| Au@Ag–Pt | Cardiac troponin I | Colourimetric: 2 ng mL−1 | 2 orders of magnitude more sensitive compared with AuNPs-LFA | Serum | 47 | |
| Catalytic: 20.41 pg mL−1 | ||||||
| Ag@SiO2@dye@SiO2 | SARS-CoV-2 antigen | Colourimetric: — | over 30 times lower than commercial colorimetric LFAs | Nasopharyngeal swab | 108 | |
| Fluorescent: 65 pg mL−1 | ||||||
| Litchi-like Au–Ag | Pyrimethanil | Colourimetric: 0.957 ng mL−1 | 2.54- and 3.41-fold lower than AuNPs-LFA | Cucumber and grape | 173 | |
| Fluorescent: 0.713 ng mL−1 | ||||||
| MXene–Au | Dexamethasone | Colourimetric: 0.0018, 0.12, and 0.084 µg kg−1 | 231-fold more sensitive than the reported LFAs | Milk, beef and pork | 174 | |
| Fluorescent: 0.0013, 0.080, and 0.070 µg kg−1 (in milk, beef, and pork) | ||||||
| AuNPs and plasmonic fluors | IL-6 | Colourimetric: 166![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) pg mL−1 pg mL−1 |
More than 1000-fold improvement over conventional LFAs | Plasma and nasopharyngeal swab | 107 | |
| Fluorescent: 93 fg mL−1 | ||||||
| SARS-CoV-2 antibodies | Colourimetric: 1.05 µg mL−1 | |||||
| Fluorescent: 185 pg mL−1 | ||||||
| SARS-CoV-2 antigen | Colourimetric: 76 ng mL−1 | |||||
| Fluorescent: 212 pg mL−1 | ||||||
| AuNPs | α-Fetoprotein | Colourimetric: 5 ng mL−1 | — | Serum | 175 | |
| Chemiluminescent: 0.27 ng mL−1 | ||||||
| Folic acid | Colourimetric: 0.1 ng mL−1 | Milk powder | ||||
| Chemiluminescent: 0.22 ng mL−1 | ||||||
| Co–Fe@hemin | SARS-CoV-2 spike antigen | Colourimetric: 25 ng mL−1 | — | Pseudo-SARS-CoV-2 | 128 | |
| Chemiluminescent: 0.1 ng mL−1 | ||||||
| Ag@Au nanoplates | SARS-CoV-2 nucleocapsid (N) protein | Colourimetric: 1 ng mL−1 | — | Saliva and nasal swab | 43 | |
| Photothermal: 40 pg mL−1 | ||||||
| High-density Aushell–Fe3O4 | SARS-CoV-2 N protein | Colourimetric: 1 ng mL−1 | 1000 times lower than commercial AuNPs-LFA | Artificial saliva | 53 | |
| Photothermal: 43.64 pg mL−1 | ||||||
| Au nanocages | Influenza A | Colourimetric: 1.8 ng mL−1 | 8000-fold more sensitive than traditional AuNPs-LFA | Saliva | 72 | |
| Photothermal: 1.51 pg mL−1 | ||||||
| Star-like Au@Pt | SARS-CoV-2 N protein antibody | Colourimetric: 1 ng mL−1 | 4000 times more sensitive than AuNPs-LFA | Serum | 202 | |
| Photothermal: 24.91 pg mL−1 | ||||||
| ZnFe2O4 | Clenbuterol | Colourimetric: 0.025 ng mL−1 | 162-fold more sensitive than traditional AuNPs-LFA | Pork and milk | 203 | |
| Photothermal: 0.012 ng mL−1 | ||||||
| Ag@Au | SARS-CoV-2 IgG | Colourimetric: 0.1, 1 ng mL−1 | Much lower compared with those using other labels | Serum | 112 | |
| SERS: 0.22, 0.52 pg mL−1 (in PBS and serum) | ||||||
| Fe3O4–AgMBA@Au | SARS-CoV-2 N protein antibody | Colourimetric: 10−8 mg mL−1 | — | Serum | 180 | |
| SERS: 0.08 pg mL−1 | ||||||
| Hollow Au–Ag garland-like | Squamous cell carcinoma antigen | Colourimetric: 0.14 pg mL−1 | — | Serum | 114 | |
| SERS: 0.063 pg mL−1 | ||||||
| Au nanocrown | SARS-CoV-2 spike 1 protein | Colourimetric: 91.24 pg mL−1 | — | Saliva | 116 | |
| SERS: 57.21 fg mL−1 | ||||||
| AuNPs | Salmonella | Colourimetric: 38.4 CFU mL−1 | — | Milk, orange juice, eggs and salmon | 144 | |
| Electrochemical: 1.96 CFU mL−1 | ||||||
| DMSNs/AuNPs@Ag | α-Fetoprotein | Colourimetric: — | — | Serum | 142 | |
| Electrochemical: 0.85 ng mL−1 | ||||||
| AuNR@Pt and Fe3O4 | Salmonella typhimurium | Colourimetric: 50 CFU mL−1 | 1000 times lower than traditional LFAs | Milk | 149 | |
| Magnetic: 75 CFU mL−1 | ||||||
| CoFe PBAs/WS2 | Gentamicin | Chemiluminescent: 0.33 pg mL−1 | — | Milk, urine and serum | 183 | |
| Photothermal: 16.67 pg mL−1 | ||||||
| Au@Ag | Neuron-specific enolase | Electrochemical: 0.04 ng mL−1 | — | Serum | 184 | |
| SERS: 0.6 ng mL−1 | ||||||
| S100-β protein | Electrochemical: 0.01 ng mL−1 | |||||
| SERS: 0.4 ng mL−1 | ||||||
| Tri-mode | Onion flower-like Au-Pd | Tetrodotoxin | Colourimetric: 1 ng mL−1 | — | Pufferfish | 185 |
| Catalytic: 0.01 ng mL−1 | ||||||
| Photothermal: 0.025 ng mL−1 | ||||||
| MoSe2@Pt | Respiratory syncytial virus | Colourimetric: 105 copies mL−1 | Over 10-folds and 50-folds more sensitive than conventional AuNPs-LFA | Nose swab | 186 | |
| Catalytic: 3162 copies mL−1 | ||||||
| Photothermal: 1202 copies mL−1 | ||||||
| Fe3O4@MoS2@Pt | SARS-CoV-2 | Colourimetric: 1 ng mL−1 | About 100 times more sensitive than commercial AuNPs-LFA | Simulated nose swab | 187 | |
| N protein | Catalytic: 80 pg mL−1 | |||||
| Photothermal: 10 pg mL−1 | ||||||
| Influenza A | Colourimetric: 0.1 µg mL−1 | |||||
| Catalytic: 20 ng mL−1 | ||||||
| Photothermal: 8 ng mL−1 | ||||||
| Pd/Pt | Staphylococcus aureus | Colourimetric: 103 CFU mL−1 | — | Urine | 188 | |
| Catalytic: — | ||||||
| Colourimetric photothermal: 4 CFU mL−1 | ||||||
| Multibranched Au@Mn | Escherichia coli O157:H7 | Colourimetric: 2034 CFU mL−1 | 37.21-fold lower than AuNPs-LFA | Milk, apple juice and river water | 171 | |
| Catalytic: 1048 CFU mL−1 | ||||||
| Colourimetric photothermal: 239 CFU mL−1 | ||||||
| SQF@ZIF-8/Pt | SARS-CoV-2 | Colourimetric: 1.56 ng mL−1 | — | Throat swab | 190 | |
| N protein | Fluorescent: — | |||||
| Catalytic: 0.0302 ng mL−1 | ||||||
| ZrFe-MOF@PtNPs | Aflatoxins | Colourimetric: 0.0636 ng mL−1 | two orders of magnitude more sensitive than AuNPs-LFA | Milk and milk powder | 191 | |
| Catalytic: 0.0179 ng mL−1 | ||||||
| Fluorescent: 0.0062 ng mL−1 | ||||||
| Au@Au@Ag/Pt | Klebsiella pneumonia | Colourimetric: 104 CFU mL−1 | — | Bronchoalveolar lavage fluid | 192 | |
| Catalytic: 103 CFU mL−1 | ||||||
| SERS: 38 CFU mL−1 | ||||||
| Au@Ag and UCNPs | microRNA-21 | No LOD mentioned | — | Serum and saliva | 194 | |
| Linear range: 1 fM–2 nM | ||||||
| Europium chelate-doped polystyrene nanoparticles | C-Reactive protein | Colourimetric: 125 ng mL−1 | — | Urine | 193 | |
| Fluorescent: 9.81 ng mL−1 | ||||||
| SERS: 77.15 ng mL−1 | ||||||
| Au–Ag hollow nanoshells | SARS-CoV-2 neutralizing antibody | Colourimetric: 0.2 µg mL−1 | — | Serum | 70 | |
| Photothermal: 20 ng mL−1 | ||||||
| SERS: 20 ng mL−1 | ||||||
| Au@Pt@Ag | Dehydro-epiandrosterone | Colourimetric: 1 ng mL−1 | Over 100-fold, 200-fold and 7000-fold more sensitive than conventional AuNPs-LFA | Milk, orange juice and green tea | 197 | |
| Photothermal: 0.42 ng mL−1 | ||||||
| SERS: 0.013 ng mL−1 | ||||||
| Au@Ag | Influenza A | Colourimetric: — | 16-fold increase | Pharyngeal swab | 196 | |
| Photothermal: 5.63 pg mL−1 | ||||||
| SERS: 31.25 pg mL−1 | ||||||
| Influenza B | Colourimetric: — | 8-fold increase | ||||
| Photothermal: 187.5 pg mL−1 | ||||||
| SERS: 93.75 pg mL−1 | ||||||
| SARS-CoV-2 | Colourimetric: — | 160-fold increase (compared with visual-based LFA) | ||||
| N protein | Photothermal: 15.63 pg mL−1 | |||||
| SERS: 31.25 pg mL−1 | ||||||
| CuS–Au | Streptococcus pneumoniae | Colourimetric: 6.5 × 102 CFU mL−1 | 3–5 orders of magnitude more sensitive compared with AuNPs-LFA | Saliva, urine and river water | 169 | |
| Photothermal: 3.6 × 102 CFU mL−1 | ||||||
| SERS: 2.0 CFU mL−1 | ||||||
| Klebsiella pneumoniae | Colourimetric: 2.9 × 102 CFU mL−1 | |||||
| Photothermal: 1.8 × 102 CFU mL−1 | ||||||
| SERS: 2.0 CFU mL−1 | ||||||
| Fe3O4@Au | Putrescine | Colourimetric: 10 ng mL−1 | — | Fish, prawns, beef and pork | 201 | |
| Photothermal: 2.31 ng mL−1 | ||||||
| Magnetic: 0.17 ng mL−1 | ||||||
| Histamine | Colourimetric: 10 ng mL−1 | |||||
| Photothermal: 4.39 ng mL−1 | ||||||
| Magnetic: 0.31 ng mL−1 | ||||||
| Au/Pt co-decorated Fe3O4 | Monkeypox virus A29L protein | Colourimetric: 0.5 ng mL−1 | 2–3 orders of magnitude more sensitive compared with AuNPs-LFA | Serum and throat swab | 170 | |
| Catalytic: 0.005 ng mL−1 | ||||||
| SERS: 0.0016 ng mL−1 | ||||||
| Quad-mode | Dandelion-like Au@Pt | Acetamiprid | Colourimetric: 0.098 ng mL−1 | 4.6-,9.2-,11.8-,11.2-, and 56.2-fold lower than traditional AuNPs-LFA | Apple sample extracts | 195 |
| Catalytic: 0.049 ng mL−1 | ||||||
| Fluorescent: 0.038 ng mL−1 | ||||||
| Photothermal: 0.04 ng mL−1 | ||||||
| Colourimetric photothermal: 0.008 ng mL−1 | ||||||
| AuPt@Cu-HCF | Diazepam | Colourimetric: 0.82 ng mL−1 | — | Crucian carp and lake water | 189 | |
| Photothermal: 12.82 pg mL−1 | ||||||
| Catalytic: 12.26 pg mL−1 | ||||||
| Colourimetric photothermal: 4.43 pg mL−1 | ||||||
| Au@PB | Higenamine | Colourimetric: 1.07 ng mL−1 | 2 orders of magnitude more sensitive than most chromatography-based methods | Urine and functional beverage | 200 | |
| Catalytic: 0.68 ng mL−1 | ||||||
| SERS: 0.01 ng mL−1 | ||||||
| Photothermal: 0.71 ng mL−1 | ||||||
| Au nanostar@PtOs nanocluster | Breast cancer exosome | Colourimetric: 1.2 × 105 exosomes per µL | 2–4 orders of magnitude more sensitive than AuNPs-LFA | Serum | 198 | |
| Catalytic: 2.6 × 103 exosomes per µL | ||||||
| Photothermal: 4.6 × 102 exosomes per µL | ||||||
| SERS: 41 exosomes per µL |
2.3. Multiplex LFAs
Clinical diagnosis, food safety surveillance, and agricultural quality control demand the rapid, accurate, and simultaneous quantitative detection of multiple analytes such as mycotoxins, pathogenic microorganisms, and pesticide residues.204–210 In these contexts, both the absolute concentrations and relative ratios of biomarkers serve as crucial diagnostic indicators; therefore, assays should encompass the maximum possible number of relevant biomarkers to generate a sufficient information for decision-making. As one of the most extensively adopted POCT platforms, LFAs have achieved notable advancements in multiplex detection through continuous refinement of strip architectures. The current structural formats for multiplex LFAs include multi-signal, single-line multi-path, multi-line, and microarray configurations (Fig. 9), collectively broadening the analytical capacity of LFAs to detect multiple targets within a single assay.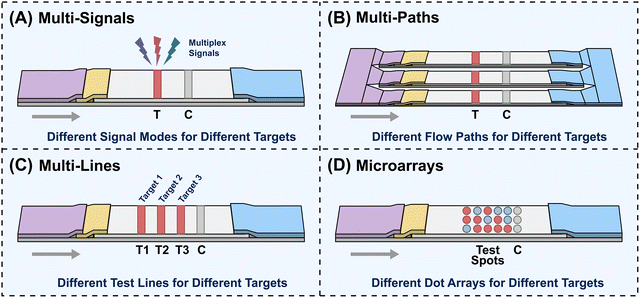 | ||
| Fig. 9 Schematic illustrations of (A) single-line, multi-signal LFA, (B) multi-path LFA, (C) multi-line LFA, and (D) microarray-type LFA. | ||
However, several limitations persist. In complex matrices such as dark-coloured fruit juices or high-fat milk, visual colour judgement can be compromised. To address this limitation, SERS-encoded nanotags offer a robust solution by converting the detection zone into a “fingerprint reader”. By exploiting the high intensity and distinct spectral peaks of Raman reporters, this approach enables the simultaneous quantification of multiple targets, thereby significantly expanding the dynamic range and sensitivity of single-line assays.212–215 Wang et al.213 developed a SERS-encoded platform specifically designed for the single-line simultaneous detection of carbendazim and imidacloprid. The strategy relied on the synthesis of AuNPs-based SERS labels, each encoded with a specific Raman reporter molecule possessing non-overlapping vibrational fingerprints. This design allowed distinct Raman peaks to be resolved from a single detection zone without spectral crosstalk (Fig. 10A). Notably, this “fingerprint readout” offered a quantitative dynamic range significantly superior to visual inspection, thereby validating the potential of SERS-LFAs in high-throughput food-safety monitoring. However, an inherent limitation arises from spatial proximity of labels in single-line multi-signal LFAs: different labels immobilized on the same line may experience steric hindrance or direct competition for binding sites, resulting in signal inaccuracy. Moreover, the accurate discrimination of optical signatures typically requires a specialised Raman reader, which hinders complete detachment from laboratory-based equipment.
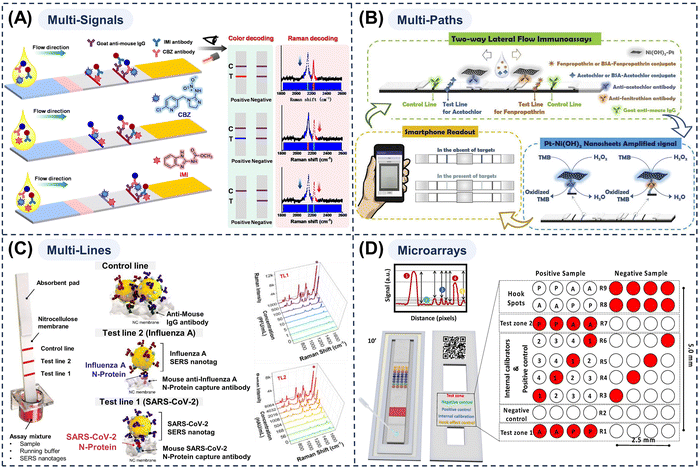 | ||
| Fig. 10 Schematic illustrations of (A) a SERS-encoded LFA strip for single-line multi-signal detection. Reprinted with permission from ref. 213. Copyright 2024, Elsevier. (B) a two-way LFA strip based on Pt–Ni(OH)2 nanosheets with smartphone readout. Reprinted with permission from ref. 217. Copyright 2019, Elsevier. (C) a multi-line SERS-LFA strip for the simultaneous differentiation of SARS-CoV-2 and Influenza A. The design utilizes sequential detection zones to ensure independent immunoreactions. Reprinted with permission from ref. 219. Copyright 2022, Springer Nature. (D) a high-density LFµIA: device featuring a patterned array of 36 discrete spots on a single membrane for simultaneous multiplex allergen detection. Reprinted with permission from ref. 223. Copyright 2022, MDPI. | ||
The main advantages of this architecture are the full independence of each detection channel, which eliminates signal crosstalk between analytes and allows reaction conditions (such as buffer and antibody concentrations) to be individually optimised for each target, thereby ensuring high accuracy. However, several limitations should also be acknowledged. If a shared inlet is not used, multiple sample loading steps are required, thereby increasing operational complexity, while total sample consumption rises because each strip needs an adequate volume. In addition, the integrated cartridge is bulkier than that used in standard single-strip LFAs, slightly compromising portability, and the more complex design and assembly typically lead to higher manufacturing costs.
The principal advantages of the multi-line format are its simple operation, straightforward visual interpretation of T-line signals, and minimal dependence on sophisticated instrumentation, making it highly suitable for rapid on-site screening. However, several intrinsic limitations persist. The typical NC membrane length (approximately 4 cm) constrains the number of T-lines to around 3 to 5, owing to the need for a spacing of at least about 2 mm to prevent signal diffusion and overlap. Upstream T-lines partially deplete the target analytes, thereby lowering the sensitivity of downstream lines, while highly viscous samples such as honey or thick fruit juices may induce non-uniform capillary flow, leading to inconsistent signal development.
The key strengths of this mode include high analytical throughput (often exceeding 10 targets per test), outstanding space efficiency (maximising detection capacity within a confined area), and minimal signal cross-talk between discrete spots, thereby enabling reliable detection in complex samples containing multiple pathogens or toxins. Nevertheless, several challenges remain. The fabrication procedure is inherently complex, requiring precision microarray spotters (e.g. microfluidic spotters) and stringent environmental control—as temperature and humidity directly influence spot uniformity. In addition, the use of advanced instrumentation and specialised labelling reagents drives up production costs, while reliance on high-resolution readers such as fluorescence microarray scanners reduces portability and limits field applications.
2.4. Translational potential of multi-mode LFAs in diverse actual sample matrices
Full realisation of the potential of LFAs in POCT requires rigorous validation of their analytical performance under practical conditions. However, when directly applied to real-world scenarios, LFAs often encounter significant challenges arising from the intrinsic complexity of biological and environmental matrices. Unlike idealised buffer systems, complex samples—such as whole blood, serum, plasma, saliva, urine, nasopharyngeal swabs, or even crude food homogenates (e.g., milk or meat extracts)—contain multiple interfering substances that can markedly impact assay performance.224 For instance, highly pigmented components such as haemoglobin in blood or myoglobin in meat, and endogenous molecules with self-luminescent properties, can absorb and scatter light, leading to interference in colorimetric or fluorescent readouts. Furthermore, viscous mucins in saliva and dense lipid particles or solid debris in food matrices can notably increase sample viscosity, resulting in slow capillary flow, membrane clogging, or delayed detection. Abundant non-target proteins in these matrices may induce non-specific adsorption, elevating background noise, whereas significant variations in pH and ionic strength—particularly in urine—can destabilize nanoprobes by disrupting their double-layer structures, leading to aggregation and loss of signal. Collectively, these matrix effects degrade assay sensitivity and accuracy, often causing false-positive or false-negative outcomes. Consequently, conventional LFAs require laborious and time-consuming sample pre-treatments (e.g., centrifugation, filtration, extraction) prior to analysis, which contradict the intrinsic goals of rapid and user-friendly on-site testing. In contrast, MMNPs offer distinctive physicochemical advantages that help overcome matrix-induced interferences. Their mechanisms for mitigating matrix effects and ensuring detection stability can be broadly categorized into three strategies:In summary, these three complementary strategies collectively mitigate severe matrix effects, enabling MMNPs to bridge the gap between idealized laboratory studies and real-world diagnostic applications. The evaluation of MMNP-based LFAs across diverse real-sample matrices, as summarized in Table 2, further demonstrates their strong translational potential. This robust matrix resilience represents a key prerequisite for advancing toward precision diagnostics, ensuring that LFAs truly fulfil their promise of rapid, reliable, and accurate POCT. As MMNPs effectively capture and amplify multimodal signals within complex biological backgrounds, the next crucial challenge involves the accurate interpretation, integration, and standardization of these diverse outputs—necessitating the incorporation of AI and digital connectivity technologies, which are further discussed in the following section.
3. Intelligent LFA systems: AI-powered analysis and digital connectivity
Accurate and reproducible analysis requires minimising environmental and substrate interferences, recognising weak or overlapping signals, and discerning meaningful correlations among multiple detection channels. At the same time, conventional LFAs function as isolated information islands, providing results that cannot be easily stored, transmitted, or integrated into broader medical information systems. This lack of digital connectivity limits applications in telemedicine, epidemiological surveillance, and real-time clinical decision support. To address these shortcomings, the LFA technology is rapidly undergoing an intelligent transformation through its integration with AI and the IoT, making a shift from subjective standalone diagnostic devices to comprehensive analytic platforms characterised by objective detection, quantitative evaluation, and intelligent data processing.1,231This section outlines the three technological pillars underpinning intelligent LFA systems, as shown in Fig. 11. (1) AI- and ML-based multi-modal data analysis: Advanced algorithms (e.g. convolutional neural networks (CNNs), random forests (RFs), and support vector machines (SVMs)) can process the complex signals generated by multi-modal LFAs, enabling the identification of subtle features, standardisation of result interpretation, and enhanced multi-target diagnostic precision. These models further support adaptive calibration and self-learning, thereby reducing operator bias and dependency on environmental conditions. (2) Smartphone-based objective imaging and quantification: Mobile devices equipped with high-resolution cameras and image-processing applications replace subjective visual assessments with quantitative signal readouts. When paired with simple optical adaptors or dongles, smartphones achieve near-laboratory-level sensitivity while retaining portability and user-friendliness, thereby supporting field deployment and home diagnostics. (3) IoT-enabled connectivity and telemedicine integration: Through Bluetooth, Wi–Fi, or cloud networking, contemporary LFAs can transmit data to healthcare servers or public health databases in real time, which allows automated electronic health record integration, remote clinical consultation, population-level outbreak monitoring, and longitudinal patient management. Synergistic advances across these three components are propelling the evolution of LFAs from simple rapid screening tools to fully intelligent diagnostic ecosystems capable of self-calibration, multi-parameter analysis, and real-time reporting. Essentially, these digital integrations enable modern LFAs to satisfy the key criteria of POCT. Automated, data-driven interpretation through AI and smartphones eliminates subjective visual bias, guaranteeing high accuracy and reliability. Simultaneously, instantaneous on-site data processing and rapid IoT transmission both maintain and enhance the speed necessary for immediate clinical decision-making. Such next-generation smart LFAs are expected to underpin personalised medicine, connected healthcare, and proactive global disease surveillance.
3.1. Application of AI and ML to for multi-modal data interpretation
The rapid evolution of multi-mode LFAs has extended their analytical capabilities beyond conventional single-signal colorimetric formats. Contemporary platforms can simultaneously generate colorimetric, fluorescence, SERS, electrochemical, and other outputs, producing high-dimensional datasets that encode rich molecular information but are difficult to interpret using traditional methods. Each channel exhibits distinct non-linear behaviour, LODs, and noise characteristics that depend on reagent chemistry, illumination, and substrate morphology. Hence, simple thresholding or visual inspection cannot capture inter-mode dependencies or time-resolved signal dynamics.In this context, AI and ML have become essential tools for converting heterogeneous multi-modal LFA readouts into reliable quantitative information. By jointly analysing colourimetric, spectral, and temporal features, supervised and deep learning models (e.g. CNNs, gradient boosting, hybrid time-series architectures) can automatically correct background variations and recognise weak positives that are frequently missed by human readers. For instance, ML-based classifiers for COVID-19 LFAs have been shown to improve interpretation accuracy and reduce the incidence of false positives and negatives by consistently identifying weak T-lines that are frequently misread by users.
AI-based methods offer a powerful solution to the limitations of human perception and the rigidity of classical calibration curve-fitting. Rather than depending on pre-defined analytical or empirical relationships, ML models learn directly from data, autonomously identifying patterns that capture the non-linear coupling among optical density, spectral signatures, and reaction kinetics (Fig. 12). Classical supervised algorithms such as K-nearest neighbours (KNNs) and SVMs are commonly deployed to classify faint or ambiguous T-line signals, effectively separating weak positive outcomes from true negatives even under challenging illumination and imaging conditions.232–234 Ensemble learning methods, including RFs and gradient boosting trees, aggregate multiple decision models to improve robustness against experimental variability and environmental noise, thereby enhancing reliability and generalisation performance. Similarly, Gaussian process regression introduces a probabilistic framework that quantifies prediction uncertainty through Bayesian inference, enabling confidence interval estimation for each predicted concentration.235–238
Deep learning extends these concepts to non-linear high-dimensional data environments. ANNs and CNNs can model intricate correlations across multi-modal features such as spectral profiles, colorimetric gradients, and fluorescence-intensity distributions.239–243 CNNs automatically extract spatial patterns that correlate with analyte concentration by recognising subtle visual cues—such as gradient intensity or nano-colloid aggregation behaviour—often imperceptible to human observers. Hybrid CNN-SVM architectures combine deep feature extraction with conventional classification accuracy, effectively using textural and morphological information to facilitate the discrimination of overlapping signals more effectively.239,240
For instance, classical supervised models such as KNNs and SVMs are widely used to classify faint or borderline T-line signals, thereby distinguishing weak positives from true negatives under variable lighting conditions.232–234 Ensemble methods such as RFs and Gradient Boosting Trees aggregate multiple weak learners to enhance robustness against experimental noise, while Gaussian process regression (GPR) incorporates probabilistic modelling to provide confidence intervals for predictions.235–238 Deep learning approaches, including ANNs and CNNs, extend these principles to nonlinear, high-dimensional data.239–243 CNNs, in particular, are highly effective for automatically recognising subtle colour gradients or fluorescence distributions corresponding to analyte concentration, while hybrid CNN-SVM architectures enable the extraction of texture-level features associated with nanoparticle aggregation.239,240 Representative AI models and their typical applications in LFA data analysis are summarized in Table 3.
| Model type | Algorithm/abbreviation | Core principle | Typical input/feature type | Strengths | Limitations/notes |
|---|---|---|---|---|---|
| Classical ML | K-Nearest neighbours (KNN) | Distance-based classification; assigns label based on nearest feature vectors | Intensity ratios, RGB values, texture features | Simple, interpretable, effective for small datasets | Sensitive to noise and data scaling; not suitable for high-dimensional data |
| Support vector machine (SVM) | Finds optimal hyperplane separating classes with maximal margin | Spectral features, morphological descriptors | High accuracy on small but well-separated datasets | Requires parameter tuning (kernel choice); less scalable | |
| Random forest (RF) | Ensemble of decision trees via bagging | Mixed features (colourimetric + SERS + fluorescence) | Handles nonlinear features; interpretable feature importance | Can overfit noisy data; less effective on extrapolation | |
| Gradient boosting (GBM/XGBoost) | Sequential ensemble learning minimizing residual error | Multi-mode quantitative data | High prediction accuracy; captures nonlinear trends | Computationally heavier; risk of overfitting | |
| Gaussian process regression (GPR) | Probabilistic regression using kernel-based covariance functions | Spectral intensities, calibration curves | Quantifies uncertainty; suitable for small data | Poor scalability with large datasets (O(n3)) | |
| Deep learning | Artificial neural network (ANN) | Multi-layer nonlinear mapping between input and output | Multi-channel numeric inputs, spectral data | Captures nonlinear correlations; flexible architecture | Requires large datasets; black-box nature |
| Convolutional neural network (CNN) | Extracts spatial patterns via convolution filters | LFA strip images, ROI pixel maps | Automatic feature learning; high performance in image analysis | Needs many training samples; sensitive to overfitting | |
| Recurrent neural network (RNN/LSTM) | Sequential modelling of temporal or positional dependencies | Time-resolved signal, flow assay dynamics | Effective for time-sequence data; can model signal drift | Training instability; vanishing gradients | |
| Autoencoder (AE) | Learns compressed latent representation via reconstruction | High-dimensional spectra, noise-rich data | Unsupervised; enhances generalization | Risk of losing subtle signal features | |
| Transfer learning (ResNet/VGG-based) | Reuses pre-trained deep models for similar visual tasks | RGB image datasets (few-shot training) | Reduces data demand; enables cross-device calibration | Requires similar data domain for effective transfer | |
| Hybrid/ensemble | CNN-RF/ANN-SVM | Combines deep feature extraction with robust classifier | Multi-mode image + numerical features | Combines advantages of deep and shallow models; robust | More complex training; requires balanced dataset |
| Probabilistic/statistical | Partial least squares-discriminant analysis (PLS-DA) | Projects features into latent variables maximizing covariance | Raman spectra, multivariate LFA signals | Efficient for small, correlated datasets | Assumes linear relation; limited in nonlinear systems |
| Principal component analysis (PCA) | Orthogonal decomposition for variance-based feature reduction | Spectral and colorimetric data | Reduces dimensionality; highlights main variance | Unsupervised; may lose minor but relevant features | |
| Advanced/recent trends | Attention-based networks | Weighting most informative signal channels dynamically | Multi-mode inputs (colour, SERS, fluorescence) | Interpretable; adaptive channel fusion | Complex architecture; training requires large data |
| Graph neural networks (GNN) | Models relational dependencies among sensor nodes or test zones | Multi-test LFAs, spatial-structural data | Captures inter-line correlation; flexible graph topology | Requires labelled relational data |
Unlike earlier review frameworks that primarily grouped AI applications by measurement modality (e.g., SERS vs. fluorescence) or algorithmic family (e.g., ML vs. DL), this review adopts a stage-wise, function-oriented classification termed SMEDI (signal-model-enhancement-decision-interpretation), as depicted in Fig. 13. This framework maps computational methods onto the natural data flow in LFA systems, establishing a structured hierarchy from raw signal capture to system-level decision-making. At the signal stage, image acquisition and pre-processing operations such as normalisation and denoising are applied to standardise input quality and multi-gate variations arising from illumination, camera hardware, and substrate heterogeneity.
The model stage centres on extracting representative features from these processed inputs, whether via CNN-based visual encoders for image data or principal component analysis (PCA) and related methods for compressing high-dimensional spectral signals. The enhancement stage employs techniques such as transfer learning and domain adaptation to improve model performance to improve cross-device robustness and maintain performance across different sample matrices and operating environments.236 At the decision stage, regression and classification models convert abstract feature representations into quantitative diagnostic outputs and categorical labels. Finally, the interpretation stage integrates automation, explainability, and clinical feedback, closing the loop from sensing to intelligence and enabling human-centred understanding, model auditing, and iterative refinement.244–247
The SMEDI paradigm underscores the fact that AI within LFAs functions not as a single isolated module, but as an end-to-end intelligence pipeline that unifies sensing, data processing, and interpretive reasoning. By structuring AI integration across all stages of the analytical workflow, this framework provides a conceptual foundation for designing next-generation LFA platforms that are self-calibrating, cross-modal, and increasingly autonomous, thereby supporting more reliable and adaptable diagnostic solutions.
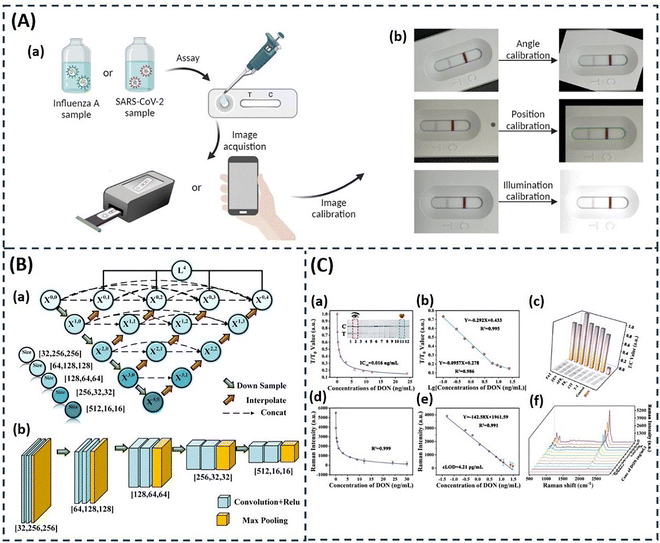 | ||
| Fig. 14 (A) Evaluation of CV-assisted sensing performance using an LFA reader and a smartphone. (a) Schematic overview of the acquisition and processing workflow; (b) post-calibration image showing the correction of geometric distortion, illumination non-uniformity, and colour imbalance via reference-card-based normalisation. Reprinted with permission from ref. 232. Copyright 2023, Royal Society of Chemistry. (B) U-Net++ network architecture and feature extraction pathway. (a) Basic nested U-Net++ structure with dense skip connections; (b) evolution of feature-map size and channel depth along the down-sampling path used for fluorescence LFA line segmentation. Reprinted with permission from ref. 240. Copyright 2024, Royal Society of Chemistry. (C) Performance of dual-mode CR-SERS LFA for deoxynivalenol (DON) detection. (a) Visual CR-LFA responses for DON concentrations 0, 0.1, 0.3, 0.6, 1.2, 3, 6, 9, 12, 24, 30, and 50 ng mL−1 (samples 1–12, respectively) and (d) corresponding SERS-LFA responses. Calibration curves obtained from (b) CR-LFA and (e) SERS-LFA channels; (c) specificity assessment of CR-LFA against potential interferents; (f) representative Raman spectra acquired from RASP-based SERS-LFA strips. Reprinted with permission from ref. 241. Copyright 2025, American Chemical Society. | ||
Once image quality has been standardised, the model stage uses ML algorithms to construct compact feature representations. For relatively simple colourimetric data, classical models such as SVM and KNN can classify intensity or colour profiles of T- and C-lines and have been successfully deployed on smartphone-based LFA platforms to distinguish weak positives from true negatives with a reliability exceeding that provided by human readers.233 In contrast, more complex signals—including fluorescence, SERS patterns, or nanostructured scattering signatures—require non-linear models capable of capturing texture, morphology, and fine spatial gradients. CNNs automatically learn multi-level visual features that map onto chemical or physical changes on the strip. Fairooz et al.239 developed hybrid CNN-SVM models simultaneously exploiting texture descriptors and deep spatial features to enhance thyroid-stimulating hormone assay sensitivity, achieving accuracies above 97% and specificities exceeding 99% under variable lighting conditions.
For fluorescence-based LFAs and line profiles with complex shapes, segmentation-focused architectures such as U-Net, UNet++, and attention-augmented variants offer pixel- or curve-level delineation of T- and C-lines that may be invisible to the naked eye (Fig. 14B).240 These models provide precise masks or integration windows for signal quantification, even when the fluorescence distribution is diffuse or obscured by background noise. In a recent study on fluorescence LFA data, an improved U-Net segmentation model achieved intersection-over-union scores approximately 0.97 for separating C- and T-peak regions, illustrating the potential of deep segmentation networks to standardise quantitative readouts in high-sensitivity assays. In spectral imaging systems such as SERS- and fluorescence-based LFAs, dimensionality reduction methods such as PCA and independent component analysis (ICA) are frequently coupled with classifiers such as SVM or RF (Fig. 14C).241,242 These pipelines compress thousands of wavelength channels into a small set of principal or independent components, improving interpretability and reducing noise while preserving the quantitative information required for accurate classification and regression.246,247
Lightweight ML models derived from these reduced feature sets can be directly deployed on smartphones or embedded readers, enabling on-device inference without reliance on cloud connectivity—an important advantage for decentralised testing, resource-limited settings, and privacy-sensitive applications. Through this integrated workflow, raw, unstructured visual and spectral data are transformed into structured numerical representations well suited for downstream modelling. The synergy between robust CV-based pre-processing and ML-driven feature extraction underpins reproducible quantification even in uncontrolled field environments, forming a core computational pillar of intelligent LFA systems.
At the enhancement stage, transfer learning and domain adaptation are particularly valuable. Pre-trained deep architectures such as ResNet, VGG, and DenseNet can be fine-tuned on small task-specific LFA datasets, sharply reducing the need for extensive manual annotation while retaining strong feature-extraction capabilities. Wang et al.236 demonstrated this strategy in an AI-reinforced UCNP-based LFA, where transfer learning allowed robust quantitative fluorescence detection and maintained high accuracy even when up to 30% artificial noise was introduced, effectively addressing data scarcity and edge-device computational limits (Fig. 15A). In parallel, data augmentation techniques—including rotation, cropping, flipping, Gaussian blurring, and controlled noise injection—expand the diversity of training images, mitigate overfitting, and improve generalisation across different readers, lighting conditions, and strip batches, which is critical for multi-source signal fusion.240
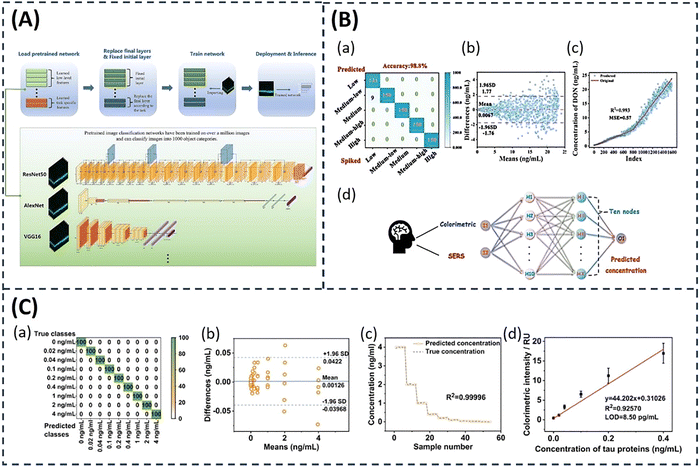 | ||
| Fig. 15 (A) Implementation process of transfer learning. Pre-trained neural network models, which have already consumed substantial computational resources and learned rich visual feature representations from large image datasets, are fine-tuned on LFA images so that their learned “skills” are efficiently adapted to related diagnostic tasks. Compared with training models from randomly initialised weights, this transfer learning strategy notably accelerates convergence and simplifies optimisation while maintaining high performance. Reprinted with permission from ref. 236. Copyright 2023, Elsevier. (B) Application of classical ML and ANN models for the dual-mode LFA of DON. (a) Confusion matrix of the KNN classifier at different DON contamination levels; (b) Bland–Altman plot comparing predicted and reference DON concentrations from ANN outputs; (c) signal response curve showing the relationship between ANN-predicted and original concentrations; (d) schematic of the ANN architecture developed to predict quantitative results from dual-mode LFA signals. Reprinted with permission from ref. 241. Copyright 2025, American Chemical Society. (C) Use of KNN and Gaussian process regression (GPR) models for tau protein analysis in PBS samples. (a) KNN confusion matrix for classification of tau concentrations into discrete classes; (b) Bland–Altman plot comparing GPR-predicted and true tau protein concentrations; (c) signal response curve comparing GPR predictions with reference values; (d) linear correlation between analyte concentration and colourimetric intensity for tau proteins in the 0–0.4 ng mL−1 range (n = 3) demonstrating the feasibility of regression-based quantification. Reprinted with permission from ref. 235. Copyright 2024, Wiley-VCH. | ||
Through this combination of pre-trained models, domain adaptation, and augmentation, multi-modal signals are mapped into a common latent representation that captures the shared structure while preserving modality-specific information. Subsequent decision-layer models (e.g. multi-modal CNNs, ensemble regressors, or attention-based fusion networks) can then exploit cross-channel redundancies and complementarities—such as combining robust but coarse colorimetric trends with highly sensitive but noisy SERS or fluorescence features—to generate more accurate and reliable diagnostic outputs than any single mode alone.
At the decision stage, ensemble and deep regression models integrate heterogeneous features into unified diagnostic outputs. RF and gradient boosting combine numerous weak learners into robust classifiers and regressors, performing particularly well on small or noisy datasets. Artificial neural networks (ANNs) and hybrid CNN-RF architectures can jointly process image, spectral, and numerical inputs to perform simultaneous classification and concentration regression. In a dual-mode colourimetric–SERS LFA, Sun et al.241 embedded both modalities into an ANN framework and achieved an R2 of 0.993 and accuracy of 98.8%, thus realising a sensitivity ∼37-fold higher than that of conventional colourimetric LFAs (Fig. 15B). Wang et al.235 combined ultrasound-assisted enrichment with ML analysis to reach sub-picogram sensitivity for tau proteins in an enhanced colourimetric LFA, illustrating how physical preconcentration and computational learning synergistically amplify detection capabilities (Fig. 15C).
Beyond straightforward data fusion, AI models uncover correlations across physical domains, such as linking subtle spectral shifts to changes in colourimetric contrast or fluorescence decay kinetics, whereas attention-based multimodal networks dynamically weight the most informative channels to improve robustness. These computational strategies constitute the core intelligence layer of multi-mode LFAs, transforming heterogeneous raw sensor outputs into coherent, reliable diagnostic insights suitable for high-performance POCT applications.178,235,236,241,242
Despite these advancements, a formidable challenge in transitioning multimodal LFA systems to real-world clinical environments is the emergence of signal discrepancies across disparate detection channels. Such inconsistencies—for instance, where a sample yields an ambiguous visual absorbance signal despite a robust SERS peak—are often symptomatic of physical perturbations such as localized membrane heterogeneity or stochastic non-specific binding. In these scenarios, a decision-level fusion strategy is prioritized over rudimentary data concatenation; this approach entails performing independent categorical assessments for each sensing modality followed by a high-level reconciliation step.248,249 A representative implementation is the hepatocellular carcinoma diagnostic framework developed by Cheng et al., which utilized a differential weighting scheme (approximately 7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3) for digital and molecular biomarkers, respectively. By assigning these weighted priorities, the model effectively insulated the global diagnostic output from the interference of single-channel noise.248
3) for digital and molecular biomarkers, respectively. By assigning these weighted priorities, the model effectively insulated the global diagnostic output from the interference of single-channel noise.248
Building upon such static models, recent architectures have begun incorporating attention mechanisms to dynamically recalibrate these weights based on real-time signal quality, further enhancing the system's robustness against single-channel noise. Complementing these structural fusion methods, the integration of uncertainty quantification (UQ) protocols significantly enhances diagnostic fidelity by enabling the system to autonomously evaluate its “inferential confidence” when faced with labile or incomplete signal sets.250,251 Tang and Shen demonstrated this through a model based on predictive entropy, which is capable of detecting high-uncertainty states in real-time. Upon detection, the system triggers compensatory algorithms—such as Kalman filtering or majority voting protocols—to refine the output.250 By effectively converging inherent physical variabilities into a mathematically grounded consensus, these logical frameworks facilitate the transition from raw, often conflicting analytical data to clinically rigorous and reliable decision-making in POC settings.
Federated learning enables continuous model optimisation across diverse datasets while preserving patient privacy. Local devices perform on-site training and share model parameters (and not raw data) with a centralised aggregator to yield global updates. This architecture not only accelerates model evolution but also supports geographically adaptive calibration, thereby connecting high-end laboratories with low-resource field settings.247 Automation must operate alongside standardisation protocols that harmonise optical, chemical, and computational variability. Techniques such as grey-world and white-patch normalisation correct illumination inconsistencies,232,246 while Bayesian-regularised neural networks provide uncertainty quantification, improving interpretability and robustness. Cloud integration further facilitates real-time epidemiological mapping, cross-laboratory harmonization, and regulatory traceability.247 These combined advancements ensure analytical consistency across time, geography, and operator expertise.
Although multimodal XAI research tailored for LFAs is in its infancy, the field is increasingly leveraging established techniques validated in medical imaging to surmount the “black-box” opacity inherent in deep learning models. The integration of these XAI frameworks is a quintessential prerequisite for “interpretable diagnostics” within intelligent LFA platforms. This transition from purely predictive to transparent modeling is fundamental to bolstering clinical trust and ensuring the accuracy of POCT.252,253
Among the most prominent architectures, Gradient-weighted Class Activation Mapping (Grad-CAM) and saliency-based techniques—extensively validated in X-ray and MRI—are being adapted to provide spatial transparency. These methods generate diagnostic heatmaps that allow clinicians to verify whether the model's heuristic attention aligns with relevant biochemical features, such as the distinct signal intensities of test and control lines, rather than being confounded by background noise or substrate interference.254 Complementing these spatial insights, SHapley Additive exPlanations (SHAP) and local interpretable model-agnostic explanations (LIME) facilitate a granular understanding of the decision-making process. SHAP, rooted in cooperative game theory, enables the rigorous quantification of each input feature's contribution—from biomarker concentrations to environmental variables—elucidating their global and local influence on the final output. Concurrently, LIME utilizes local surrogate modeling to decipher the rationale behind individual classifications, ensuring that each POCT result is not only numerically accurate but clinically justifiable.255,256
Ultimately, the convergence of automation and standardisation redefines the POC concept. LFAs are no longer static diagnostic tools but autonomous, self-learning biosensing systems capable of continuous adaptation. Coupled with explainable AI and connected infrastructures, this stage completes the SMEDI intelligence loop, transforming LFAs into active, adaptive, and cloud-connected diagnostic ecosystems for large-scale healthcare deployment.178,235,244–247
3.2. Smartphone-based objective and quantitative signal readout
Conventional LFA systems heavily rely on the subjective visual interpretation of T- and C-lines, which renders results sensitive to ambient lighting, individual visual acuity, and operator experience and thus limits analytical accuracy. Smartphone-based readers address these issues by converting line intensities into digital signals using standardised imaging and computational analysis, which improves reproducibility and enables the detection of weak positives difficult to distinguish by the naked eyes. Smartphones are not only ubiquitous communication tools but also integrated analytical platforms that combine high-performance processors, large storage capacity, high-resolution cameras, and connectivity modules such as global positioning system (GPS), Bluetooth, Wi-Fi, near-field communication (NFC), and 3G/4G/5G. In addition, built-in light, proximity, infrared, and motion sensors, together with the support of operating systems such as Android, iOS, and (to a lesser extent) Windows, enable the development of customised, user-friendly applications for image capture, on-device analysis, secure storage, and wireless transmission and thus make anytime, anywhere POCT technically feasible.Beyond colourimetric signals, smartphone platforms can capture and analyse photothermal, fluorescence, SERS, electrochemical, and other transduction modes to enhance sensitivity and multiplexing (Fig. 16).72,141,146,176,181,257,264–266 Atta et al.72 reported a colourimetric–photothermal dual-mode LFA based on Au nanocages for highly sensitive influenza A virus detection, integrating a compact laser, 3D printed housing, an LFA strip, smartphone, and smartphone-compatible thermal imager into a portable photothermal platform that maintained high sensitivity and stability for spiked saliva samples over several months (Fig. 16A). For fluorescent readouts, Wang et al.265 developed a highly sensitive ratiometric fluorescent LFA for detecting heart-type fatty acid-binding protein. To enable portable and accurate quantitative analysis, they designed a compact smart device comprising a 3D-printed attachment integrated with a smartphone (Fig. 16B). The LED excitation light (365 nm) passes through a band-pass filter and is directed at a 45° angle onto the test zone of the LFA strip. The emitted fluorescence is subsequently filtered through a 500 nm long-pass filter to suppress background noise before being captured by the smartphone's CMOS image sensor. The hue and RGB values are then extracted from the recorded images using an application named Colour Picker.
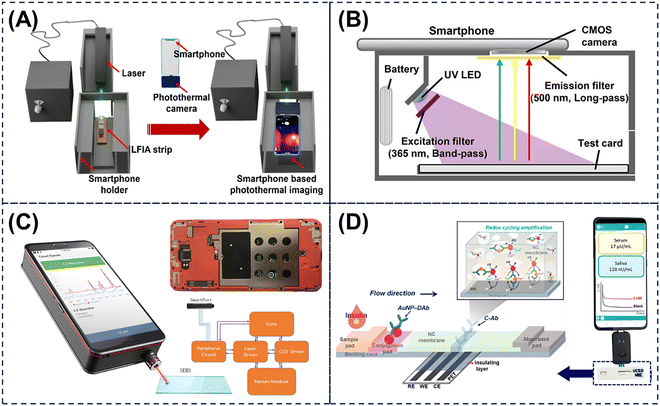 | ||
| Fig. 16 (A) Smartphone-based photothermal LFA reader integrating a portable laser, LFA strip, and smartphone-mounted thermal imager for quantitative temperature mapping of the test zone. Reprinted with permission from ref. 72. Copyright 2025, American Chemical Society. (B) Schematic illustration of a smartphone-based portable fluorescence reader employing dedicated excitation, optical filtering, and collection optics to enhance lateral-flow fluorescence detection sensitivity. Reprinted with permission from ref. 265. Copyright 2025, Wiley-VCH (C) Smartphone-integrated Raman system in which an external miniaturized Raman module and on-phone spectrometer enable SERS-based readout on paper or LFA substrates. Reprinted with permission from ref. 266. Copyright 2019, IEEE. (D) Simplified mobile electrochemical LFA platform combining a screen-printed electrode-based strip with a compact potentiostat interfaced to a smartphone for label-free or labelled electrochemical signal measurement. Reprinted with permission from ref. 146. Copyright 2023, American Chemical Society. | ||
Moreover, as SERS detection typically requires a precision Raman spectrometer, Mu et al.266 developed a miniaturised, high-sensitivity Raman detection system optimised for smartphone integration (Fig. 16C). The design employs fixed-focal-length lenses, bulk phase gratings, and direct slit coupling technology to improve resolution and signal sensitivity while reducing system size. A cloud-based network architecture, established via the smartphone's wireless communication interface, enables rapid on-site identification of substances and big-data analysis, thereby fulfilling field-testing requirements in complex environments. Electrochemical coupling further expands smartphone readout capabilities. Zhang et al.181 developed a smartphone-based e/v-LFA dual-readout POCT strategy for methicillin-resistant S. aureus (MRSA), incorporating a screen-printed carbon electrode beneath an NC membrane to transduce binding events into electrical signals while simultaneously quantifying colour changes via smartphone RGB analysis in a black box. In this dual-mode configuration, a portable electrochemical workstation relayed current signals to the smartphone, enabling accurate, sensitive, and portable MRSA detection in real samples. Nandhakumar et al.146 developed a simplified smartphone-based electrochemical LFA (eLFA) strip for insulin detection by integrating electron-transferring reactions with signal amplification directly onto the LFA strip (Fig. 16D). For portable and user-friendly operation, the eLFA strip was enclosed in a custom-fabricated plastic case designed to ensure precise strip-electrode alignment and mechanical stability during measurement. This streamlined casing design enables consistent electrical contact, minimises user-dependent variability, and demonstrates the feasibility of decentralised, smartphone-integrated electrochemical LFA systems. Collectively, these examples highlights the smartphone's potential to act as a universal hardware interface—standardising signal acquisition across colourimetric, photothermal, and electrochemical modalities—and bridging physical test strips with digital analytical frameworks.
Furthermore, GPS and data connectivity functions enable geo-tagging of tests and immediate sharing of results, adding layers of epidemiological and telemedical utility to the basic diagnostic act. Yentongchai et al.271 employed NFC technology to develop a label-free electrochemical LFAs for S. typhimurium detection. By combining NFC as a potentiometric interface, the proposed sensor enables seamless data transmission to smartphones, and the electrochemical sensitivity was effectively integrated with a user-friendly, field-deployable diagnostic system, facilitating portable wireless signal acquisition and real-time on-site analysis. Gonzalez-Macia et al.272 also used the NFC-enabled potentiostat to achieve the electrochemical detection of maize mosaic virus. In addition, Brangel et al.273 proposed an on-site testing device composed of LFA strips and a smartphone reader. This platform used a specially developed smartphone application to enable rapid and portable testing, data storage and sharing, and the geo-tagging of tested individuals in Uganda. The system held significant potential as an on-site tool for diagnosis, vaccine development and treatment evaluation.
This synergistic integration of standardized hardware acquisition and embedded intelligent software represents a transformative leap, enabling a digital, interconnected POCT platform that delivers laboratory-grade analysis in a ubiquitous, low-cost form factor. This development signifies a substantial advancement for personalised medicine and telemedicine, by enabling professional-grade testing in households worldwide, opening a promising avenue within smart healthcare.244,279,280
3.3. Telemedicine and IoT integration
The deep integration of telemedicine and IoT has accelerated the evolution of LFA technology toward intelligent, interconnected diagnostic platforms. The IoT enables real-time communication and data exchange across distributed device networks,266,281 and its healthcare implementation—often termed the Internet of Medical Things (IoMT)—provides the core infrastructure for smart LFA systems by supporting multidimensional data acquisition, analysis, and remote supervision.282 Modern intelligent LFA platforms exploit cloud-based architectures and mobile communication technologies such as Bluetooth and 4G/5G to offload computationally intensive tasks to remote servers. This approach not only improves analytical accuracy and scalability but also allows users in resource-constrained or geographically isolated regions to access reliable diagnostic support without dependence on centralised laboratories. IoT-enabled LFA systems also support continuous health monitoring and dynamic data integration.283 Patients can track test results and engage in teleconsultations via mobile applications, while public health authorities and healthcare institutions can utilise aggregated anonymised cloud data to build predictive models and design targeted intervention strategies.284 This approachreduces the need for frequent in-person contact between patients and clinicians and enhances the overall responsiveness and systemic efficiency of healthcare services.Bayin et al.153 developed an LFA platform based on superparamagnetic NPs and giant magnetoresistance sensing for the rapid, quantitative, and simultaneous detection of anti-SARS-CoV-2 IgM and IgG, integrating IoMT connectivity to enable result transmission via Bluetooth to a smartphone application and remote sharing with healthcare centres. Guo et al.285 reported a fluorescence LFA sensor for quantitative C-reactive protein detection using mesoporous silica-coated UNCPs (UCNPs@mSiO2), which was further integrated with Bluetooth-enabled smartphones and cloud services to achieve suitability for IoT scenarios (Fig. 17A). This fluorescent sensing platform was subsequently combined with 5G communication to achieve the highly sensitive quantitative detection of SARS-CoV-2 spike and nucleocapsid proteins together with remote medical monitoring (Fig. 17B).286 The compact hardware incorporated Bluetooth and 5G modules to stream data in real time to fog-computing nodes and cloud servers, where embedded fuzzy logic and deep learning algorithms provide intelligent result interpretation and early-warning functions. These IoMT-enabled solutions show great potential for the early diagnosis, proactive alerting, and intelligent prevention and control of COVID-19 and other infectious diseases, while greatly simplifying the remote review of infection history, treatment planning, and epidemiological surveillance by medical staff.
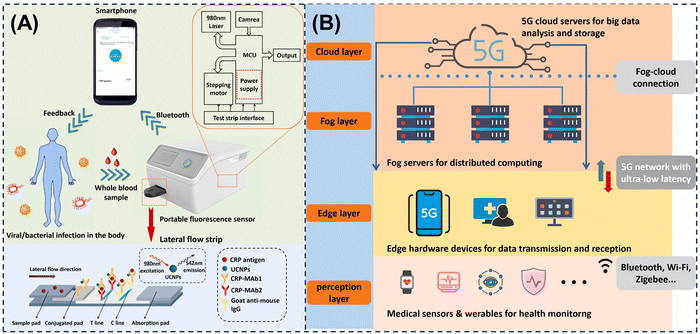 | ||
| Fig. 17 (A) Schematic of the UCNP-LFA-based Internet od Medical Things platform for human C-reactive protein quantification in which a portable fluorescence reader acquires LFA signals and transmits processed results via Bluetooth to a smartphone application, enabling cloud connectivity and remote clinical access. Reprinted with permission from ref. 285. Copyright 2023, American Chemical Society. (B) Conceptual architecture of the 5G-enabled Internet of Medical Things illustrating the layered integration of wearable and POC sensors, edge/fog computing nodes, and 5G cloud infrastructure to support ultra-low-latency high-reliability transmission, large-scale data analytics, and proactive telemedicine services. Reprinted with permission from ref. 286. Copyright 2021, Elsevier. | ||
4. Conclusions and future perspectives: towards next-generation intelligent and synergistic LFAs
This review systematically traces the evolution of LFAs from simple qualitative POCTs to sophisticated, intelligent, and multi-modal diagnostic platforms. Central to this transformation is the advent of MMNPs, whose intrinsic compositional and structural complexities endow them with functionalities that remain unattainable with conventional single-component nanomaterials.Importantly, this review delineates two fundamentally distinct paradigms of multi-modal LFAs: (i) systems in which a single nanostructured probe intrinsically generates multiple, orthogonal signals, and (ii) systems that achieve multi-signal output through the parallel use of different tags or detection modalities. While both approaches strengthen diagnostic reliability, the unique merit of MMNPs lies primarily in the former. By synergistically integrating plasmonic, catalytic, magnetic, photothermal, and electronic properties within a single particle, MMNPs enable the simultaneous production of cross-validating signals—such as colourimetric, fluorescence, SERS, photothermal, and magnetic responses—from a unified sensing interface. This intrinsic multi-signal capability broadens the dynamic range, enhances resilience against environmental and operational fluctuations, and fortifies analytical robustness, thereby fulfilling key criteria of POC diagnostics: speed, reliability, and accuracy.
Building on these material-level innovations, LFA architectures have progressed beyond conventional multi-line formats toward microarray-based and spatially multiplexed configurations, enabling the simultaneous detection of multiple biomarkers from a single specimen. This evolution marks a conceptual transition from single-pathogen or single-analyte testing to comprehensive health assessment. In parallel, the integration of AI and digital connectivity has addressed long-standing challenges in signal interpretation and result standardisation. ML algorithms now enable automated image recognition, multi-signal fusion, and high-throughput data processing, while smartphones serve as ubiquitous computational interfaces that bridge physical assays with cloud-based and IoT-enabled healthcare networks. Consequently, LFAs have been transformed from passive diagnostic strips into active, network-connected analytical nodes that facilitate real-time monitoring, remote diagnostics, and data-driven clinical decision-making.
Despite this rapid progress, several critical challenges must be overcome to fully realise next-generation intelligent LFAs.
1. Design-on-demand synthesis of multi-functional nanomaterials. Despite the enormous potential and multi-modal functionalities of MMNPs in LFAs, their synthesis remains time-consuming, labour-intensive, and technically demanding. These preparations typically involve complex multi-step operations requiring precise control of temperature, reagents, and capping agents. In practical applications, batch-to-batch variations in size, morphology, and composition can undermine analytical consistency. Furthermore, the dependence on precious metals and intricate synthesis routes raises overall production costs, while the long-term stability of MMNPs under various storage conditions requires rigorous evaluation. Bridging the gap between milligram-scale laboratory synthesis and kilogram-scale industrial production remains a major bottleneck. To address this, future MMNPs should be designed with enhanced structural precision, reproducibility, and scalable manufacturability. The development of continuous-flow microfluidic synthesis reactors integrated with real-time quality control modules will be pivotal for ensuring uniformity in particle size, shape, and surface functionality at commercial scale. In parallel, integrating computational modelling and AI-assisted simulation offers a route toward data-driven nanomaterial design, moving beyond traditional trial-and-error synthesis. Closed-loop workflows interlinking AI prediction, targeted synthesis, and performance validation will be essential to tailor MMNPs for specific diagnostic applications.
2. Integration of fully automated and user-centric systems. To achieve genuine sample-in–result-out functionality, LFA systems should incorporate sample preparation, reagent storage, and integrated multi-modal microfluidic readout interfaces within a single, compact device. Low-power, portable readers capable of synchronous multi-signal acquisition will enable intuitive, one-touch operation, delivering laboratory-quality diagnostics in home, primary-care, and resource-limited environments.
3. Clinical translation and multi-mode data integration. While current AI applications primarily focus on image processing and signal extraction, future platforms must exploit multi-dimensional data fusion for clinical decision support. Harmonising heterogeneous signals—optical, electrical, and magnetic—through advanced weighting and conflict-resolution algorithms will be crucial for converting multi-modal measurements into actionable clinical insights. Furthermore, moving beyond 'black-box' algorithms toward explainable AI is imperative to gain clinician trust, ensuring that AI-driven diagnostic scores are transparent, interpretable, and biologically valid.
4. Standardisation and commercialisation frameworks. Successful translation from laboratory research to clinical implementation requires standardised protocols spanning the entire innovation pipeline, incorporating reference color charts and cross-device calibration algorithms to mitigate data distortion from environmental and hardware variability. To bolster engineering reliability, multi-dimensional metrics such as F1-scores and explainable AI must be integrated to ensure diagnostic precision and operational transparency. Establishing these technical standards is indispensable for verifying reproducibility through multi-centre clinical trials, thereby laying the foundation for regulatory approval and large-scale commercialization.
In summary, the synergistic integration of MMNPs and intelligent computation is driving a paradigm shift in LFA technology—from single-mode to intrinsically multi-modal sensing, from subjective interpretation to data-driven decision-making, and from isolated strips to interconnected diagnostic networks. Next-generation LFAs are poised to transcend conventional limitations, emerging as high-performance, globally accessible, and analytically robust platforms that advance precision medicine, democratize advanced healthcare diagnostics, and bolster global public health infrastructure.
Conflicts of interest
There are no conflicts to declare.Data availability
This review does not include any primary research results, software, or code, and no new data were generated or analysed as part of this work.Acknowledgements
This study was supported by the Natural Scientific Foundation of Shandong (grant numbers ZR2022JQ07 and ZR2025ZD25), the National Natural Science Foundation of China (grant number 22476214), the Key Project of Qingdao Natural Scientific Foundation (grant number 24-8-4-zrjj-9-jch), the National Research Foundation of Korea (grant numbers 2020R1A5A1018052, RS-2024-00352256, RS-2024-00339674 and RS-2025-02213941), the Technology Innovation Program (grant number RS-2024-00432382) funded by the Ministry of Trade, Industry, and Energy (MOTIE, Korea), and the Regional Innovation System & Education (RISE) through the Seoul RISE Centre, funded by the Ministry of Education (MOE) and the Seoul Metropolitan Government (grant number 2026-RISE-01-024-05).References
- G.-R. Han, A. Goncharov, M. Eryilmaz, S. Ye, B. Palanisamy, R. Ghosh, F. Lisi, E. Rogers, D. Guzman, D. Yigci, S. Tasoglu, D. Di Carlo, K. Goda, R. A. McKendry and A. Ozcan, Nat. Commun., 2025, 16, 3165 CrossRef CAS PubMed.
- S. Lee, L. Bi, H. Chen, D. Lin, R. Mei, Y. Wu, L. Chen, S.-W. Joo and J. Choo, Chem. Soc. Rev., 2023, 52, 8500–8530 RSC.
- D. C. Christodouleas, B. Kaur and P. Chorti, ACS Cent. Sci., 2018, 4, 1600–1616 CrossRef CAS PubMed.
- H. R. Boehringer and B. J. O’Farrell, Clin. Chem., 2021, 68, 52–58 CrossRef PubMed.
- T. Hall, S. Gulati, R. Sang, Z. Jia, F. McKinnirey, G. Vesey, E. Goldys and F. Deng, TrAC, Trends Anal. Chem., 2025, 189, 118275 CrossRef CAS.
- S. Lee, H. Dang, J.-I. Moon, K. Kim, Y. Joung, S. Park, Q. Yu, J. Chen, M. Lu, L. Chen, S.-W. Joo and J. Choo, Chem. Soc. Rev., 2024, 53, 5394–5427 RSC.
- H. He, B. Liu, S. Wen, J. Liao, G. Lin, J. Zhou and D. Jin, Anal. Chem., 2018, 90, 12356–12360 CrossRef CAS PubMed.
- J. Li, Y. Liu, T. Wu, Z. Xiao, J. Du, H. Liang, C. Zhou and J. Zhou, Nat. Commun., 2024, 15, 5603 CrossRef CAS PubMed.
- E. B. Bahadır and M. K. Sezgintürk, TrAC, Trends Anal. Chem., 2016, 82, 286–306 CrossRef.
- Y. Cheng, Y. Li, J. Tian, L. Zhao, Q. Li, B. Li, J. Wang, D. Zhang and I. A. Darwish, Biosens. Bioelectron., 2026, 296, 118368 CrossRef CAS PubMed.
- Y.-L. Ye, Y. Zhang, J. Cao, J. Zhou, L.-F. Yu, L.-X. Yan, Y.-W. Feng and X.-D. Huang, Biosens. Bioelectron., 2026, 297, 118373 CrossRef CAS PubMed.
- X. Wang, N. Choi, Z. Cheng, J. Ko, L. Chen and J. Choo, Anal. Chem., 2017, 89, 1163–1169 CrossRef CAS PubMed.
- X. Fu, Z. Cheng, J. Yu, P. Choo, L. Chen and J. Choo, Biosens. Bioelectron., 2016, 78, 530–537 Search PubMed.
- E. Sarathkumar and R. S. Jayasree, TrAC, Trends Anal. Chem., 2025, 193, 118478 CrossRef CAS.
- Y. Wang, M. Wang, J. Cui and H. Zhang, Trends Food Sci. Technol., 2025, 163, 105180 CrossRef CAS.
- H. Tong, C. Cao, M. You, S. Han, Z. Liu, Y. Xiao, W. He, C. Liu, P. Peng, Z. Xue, Y. Gong, C. Yao and F. Xu, Biosens. Bioelectron., 2022, 213, 114449 CrossRef CAS PubMed.
- J. Du, C. Cao, Z. Xue, W. Wang, X. Lu, Y. Wei, J. Huang, L. Zhao, L. Wang, F. Xu, C. Yao, T. Wen and M. You, Anal. Chem., 2025, 97, 24196–24208 Search PubMed.
- A. D. Beggs, C. C. S. Caiado, M. Branigan, P. Lewis-Borman, N. Patel, T. Fowler, A. Dijkstra, P. Chudzik, P. Yousefi, A. Javer, B. Van Meurs, L. Tarassenko, B. Irving, C. Whalley, N. Lal, H. Robbins, E. Leung, L. Lee and R. Banathy, Cell Rep. Med., 2022, 3, 100784 CrossRef PubMed.
- H. Ma, L. Hu, F. Ding, J. Liu, J. Su, K. Tu, J. Peng, W. Lan and L. Pan, Biosens. Bioelectron., 2024, 263, 116577 CrossRef CAS PubMed.
- T. Phan-Xuan, S. Schweidler, S. Hirte, M. Schüller, L. Lin, A. Khandelwal, K. Wang, J. Schützke, M. Reischl, C. Kübel, H. Hahn, G. Bello, J. Kirchmair, J. Aghassi-Hagmann, T. Brezesinski, B. Breitung and L. A. Dailey, ACS Nano, 2024, 18, 19024–19037 Search PubMed.
- J. Gargiulo, M. Herran, I. L. Violi, A. Sousa-Castillo, L. P. Martinez, S. Ezendam, M. Barella, H. Giesler, R. Grzeschik, S. Schlücker, S. A. Maier, F. D. Stefani and E. Cortés, Nat. Commun., 2023, 14, 3813 Search PubMed.
- Y.-H. Wang, L. Zhu, E. Mariani, E. Pensa, O. Henrotte, Y. Xia, K. Müller-Caspary, T.-Y. Zhang, M.-R. Gao and E. Cortés, J. Am. Chem. Soc., 2026, 148, 8612–8620 CrossRef CAS PubMed.
- E. B. Aydın, M. Aydın and M. K. Sezgintürk, J. Hazard. Mater., 2026, 505, 141414 CrossRef PubMed.
- S. Sarkar, P. K. Samanta, M. Easton and A. Nag, ACS Catal., 2026, 16, 4550–4560 CrossRef.
- N. Jiang, R. Ahmed, M. Damayantharan, B. Ünal, H. Butt and A. K. Yetisen, Adv. Healthcare Mater., 2019, 8, e1900244 CrossRef PubMed.
- Y. Liu, L. Zhan, Z. Qin, J. Sackrison and J. C. Bischof, ACS Nano, 2021, 15, 3593–3611 Search PubMed.
- K. Omidfar, F. Khorsand and M. Darziani Azizi, Biosens. Bioelectron., 2013, 43, 336–347 Search PubMed.
- V.-T. Nguyen, S. Song, S. Park and C. Joo, Biosens. Bioelectron., 2020, 152, 112015 Search PubMed.
- A. Sena-Torralba, R. Álvarez-Diduk, C. Parolo, A. Piper and A. Merkoçi, Chem. Rev., 2022, 122, 14881–14910 Search PubMed.
- B. Fang, G. Zhang, Y. Li, J. Wu, C. Chang, Z. Gong, X. Liu, Y. Xiong and W. Lai, Coord. Chem. Rev., 2025, 542, 216900 CrossRef CAS.
- M. Ha, J.-H. Kim, M. You, Q. Li, C. Fan and J.-M. Nam, Chem. Rev., 2019, 119, 12208–12278 CrossRef CAS PubMed.
- X. Liu, X. Liang, J. Yu, K. Xu, J.-W. Shen, W. Duan and J. Zeng, TrAC, Trends Anal. Chem., 2023, 169, 117386 CrossRef CAS.
- C. Amiens, D. Ciuculescu-Pradines and K. Philippot, Coord. Chem. Rev., 2016, 308, 409–432 CrossRef CAS.
- L. Tong, D. Li, M. Huang, L. Huang and J. Wang, Anal. Chem., 2023, 95, 17318–17327 CrossRef CAS PubMed.
- M. Park, C. S. H. Hwang and K.-H. Jeong, ACS Appl. Mater. Interfaces, 2018, 10, 290–295 CrossRef CAS PubMed.
- R. Shu, S. Liu, M. Wang, X. Yin, Y. Cheng, J. Wang and D. Zhang, Chem. Eng. J., 2025, 506, 159862 CrossRef CAS.
- W. Huang, Z. Ren, X. Li, R. Chen, D. Fan, J. Da, Y. Zha and Y. Xu, Microchim. Acta, 2025, 192, 408 Search PubMed.
- L. Zhang, Z. Li, W. Bai, L. Dai, C. Wang, L. Zhang, Y. Liu, Q. Jin, S. Zheng and S. Wang, Sens. Actuators, B, 2025, 444, 138508 CrossRef CAS.
- X. Zhao, G. Zhou, Y. Geng, F. Chen, H. Zhu, C. Zhao, H. Suo and L. Ding, Chem. Eng. J., 2025, 512, 162199 CrossRef CAS.
- G. Chen, J. Zhang, W. Chen, R. Lu, C. Ma, Z. Wang and Y. Han, Chem. Sci., 2024, 15, 12550–12558 RSC.
- B. Gosselin, G. Bruylants and I. Jabin, ACS Appl. Nano Mater., 2024, 7, 6169–6177 CrossRef CAS.
- J. Sun, J. Ji, Y. Sun, M. H. Abdalhai, Y. Zhang and X. Sun, Biosens. Bioelectron., 2015, 70, 239–245 CrossRef CAS PubMed.
- K. Wang, X. Liu, X. Liang, Y. Jiang, C.-Y. Wen and J. Zeng, Anal. Chem., 2024, 96, 3208–3216 CAS.
- L. Ji, L. Zhang, H. Yang, S. Liang, J. Pan, Y. Zou, S. Li, Q. Li and S. Zhao, J. Colloid Interface Sci., 2022, 621, 489–498 CrossRef CAS PubMed.
- L. Shi, J. Wang, Y. Li, X. Wang, H. Jia, S. Liu, J. Sun, G. Huang and J. Wang, Biosens. Bioelectron., 2025, 287, 117685 CrossRef CAS PubMed.
- J. Ma, X. Yin, Y. Cheng, C. Wang, Q. Wu, Q. Zhang, L. Zhao, J. Zhou, J. Wang and D. Zhang, Chem. Eng. J., 2024, 498, 155341 CrossRef CAS.
- T. Bai, L. Wang, M. Wang, Y. Zhu, W. Li, Z. Guo and Y. Zhang, Biosens. Bioelectron., 2022, 208, 114218 Search PubMed.
- W. Wang, Q. Cao, J. He, Y. Xie, Y. Zhang, L. Yang, M.-H. Duan, J. Wang and W. Li, Nano Lett., 2024, 24, 8311–8319 CrossRef CAS PubMed.
- X. Meng, W. Zuo, P. Wu, Y. Song, G.-J. Yang, S. Zhang, J. Yang, X. Zou, W. Wei, D. Zhang, J. Dai and Y. Ju, Nano Lett., 2024, 24, 51–60 CrossRef CAS PubMed.
- N. Meng, X. Ma, C. Wang, Y. Wang, R. Yang, J. Shao, Y. Huang, Y. Xu, B. Zhang and Y. Yu, ACS Nano, 2022, 16, 9095–9104 CrossRef CAS PubMed.
- C. Wang, J. Li, Y. Yuan, B. Ouyang, Z. Guo, C. Lin, X. Yang, B. Kang, C. Li, Y. Sun and K. Xu, Angew. Chem., Int. Ed., 2025, 64, e202505616 CrossRef CAS PubMed.
- C. Qin, D. Zhang, P. Wang, F. Guo, D. Ni, Z. Yu and P. Liang, Chem. Eng. J., 2025, 525, 170656 CrossRef CAS.
- X. Li, D. Yu, H. Li, R. Sun, Z. Zhang, T. Zhao, G. Guo, J. Zeng and C. Y. Wen, Biosens. Bioelectron., 2023, 241, 115688 CrossRef CAS PubMed.
- X. Zhao, X. Liu, D. Chen, G. Shi, G. Li, X. Tang, X. Zhu, M. Li, L. Yao, Y. Wei, W. Song, Z. Sun, X. Fan, Z. Zhou, T. Qiu and Q. Hao, Nat. Commun., 2024, 15, 5855 Search PubMed.
- Y. Xu, L. Shi, X. Jing, H. Miao and Y. Zhao, ACS Appl. Mater. Interfaces, 2022, 14, 3293–3301 CrossRef CAS PubMed.
- T. Sun, Y. Wu, H. Ma, C. Zhang, C. Li, B. Man, C. Yang and Z. Li, Nano Lett., 2024, 24, 15324–15330 CrossRef CAS PubMed.
- Y. Huang, Y. Zhang, W. Hao, H. Lu, H. Dong and X. Zhang, Sens. Actuators, B, 2023, 375, 132945 CrossRef CAS.
- J. Ren, P. Wang, Y. Yang, X. Li, X. Pan, Y. Ma, F. Tan, X. Zeng, X. Zhang, H. Yang and J. Zeng, Talanta, 2026, 297, 128670 CrossRef CAS PubMed.
- S. Cheng, L. Luo, M. Bao, T. Bao, Y. Gao, Z. Wu, X. Zhang, S. Wang and W. Wen, Anal. Chim. Acta, 2025, 1338, 343588 CrossRef PubMed.
- K. Wang, D. Yu, X. Liang, W. Lu, X. Jiang, K. Tashpulatov, J. Zeng and C.-Y. Wen, Microchem. J., 2024, 204, 111104 CrossRef CAS.
- Y. Liu, L. Chen, Z. Chen, M. Liu, X. Li, Y. Kou, M. Hou, H. Wang, X. Li, B. Tian and J. Dong, ACS Nano, 2023, 17, 8167–8182 CrossRef CAS PubMed.
- T. Yu, J. Wu, Y. Shen, A. Penkova, W. Qi and R. Su, Chem. Eng. J., 2024, 498, 155144 CrossRef CAS.
- W. Duan, J. Wang, X. Peng, S. Cao, J. Shang, Z. Qiu, X. Lu and J. Zeng, Biosens. Bioelectron., 2023, 223, 115022 Search PubMed.
- L. Zhai, S. T. Gebre, B. Chen, D. Xu, J. Chen, Z. Li, Y. Liu, H. Yang, C. Ling, Y. Ge, W. Zhai, C. Chen, L. Ma, Q. Zhang, X. Li, Y. Yan, X. Huang, L. Li, Z. Guan, C.-L. Tao, Z. Huang, H. Wang, J. Liang, Y. Zhu, C.-S. Lee, P. Wang, C. Zhang, L. Gu, Y. Du, T. Lian, H. Zhang and X.-J. Wu, Nat. Commun., 2023, 14, 2538 CrossRef CAS PubMed.
- Z. Lyu, S. Zhu, L. Xu, Z. Chen, Y. Zhang, M. Xie, T. Li, S. Zhou, J. Liu, M. Chi, M. Shao, M. Mavrikakis and Y. Xia, J. Am. Chem. Soc., 2021, 143, 149–162 CrossRef CAS PubMed.
- M. Ghosh, U. Dasgupta, R.-A. Doong, J. Vidic, P. C. Sadhukhan, E. Y. Park and A. Dutta Chowdhury, ACS Sens., 2026, 11, 299–309 CrossRef CAS PubMed.
- G. Guo, T. Zhao, R. Sun, M. Song, H. Liu, S. Wang, J. Li and J. Zeng, Chin. Chem. Lett., 2024, 35, 109198 CrossRef CAS.
- Y. Dong, C.-Y. Wen, Y. She, Y. Zhang, Y. Chen and J. Zeng, Small, 2021, 17, 2104596 CrossRef CAS PubMed.
- C.-Y. Wen, L.-J. Zhao, Y. Wang, K. Wang, H.-W. Li, X. Li, M. Zi and J.-B. Zeng, Microchim. Acta, 2023, 190, 57 CrossRef CAS PubMed.
- T. Zhao, P. Liang, J. Ren, J. Zhu, X. Yang, H. Bian, J. Li, X. Cui, C. Fu, J. Xing, C. Wen and J. Zeng, Anal. Chim. Acta, 2023, 1255, 341102 CrossRef CAS PubMed.
- Z. Gao, S. Shao, W. Gao, D. Tang, D. Tang, S. Zou, M. J. Kim and X. Xia, ACS Nano, 2021, 15, 2428–2438 CrossRef CAS PubMed.
- S. Atta, Y. Zhao, S. Sanchez, S. V. Yampolsky and T. Vo-Dinh, Anal. Chem., 2025, 97, 6427–6437 CrossRef CAS PubMed.
- L. Zhang, L. Ji, M. Lin, R. Liu, H. Yang, J. Zhao and S. Zhao, Microchim. Acta, 2024, 191, 330 Search PubMed.
- G. Zhang, H. Hu, S. Deng, X. Xiao, Y. Xiong, J. Peng and W. Lai, Biosens. Bioelectron., 2023, 225, 115090 CrossRef CAS PubMed.
- C. Ran, Q. Zhang, Z. Long, Y. Li, X. He, S. Li, C. Chen, C. Cheng, Y. Xu, W. Zhang, W. Guo and Y. Shen, Chem. Eng. J., 2025, 515, 163350 CrossRef CAS.
- M. Liang, X. Hu, D. Wang, X. Tang, Q. Zhang and P. Li, Biosens. Bioelectron., 2025, 277, 117223 CrossRef CAS PubMed.
- H. Park, A. Ashok, M. K. Masud, J. Otte, M. Kim, S. M. Alshehri, T. Ahamad, Y. Bando, M. S. A. Hossain, Y. V. Kaneti and Y. Yamauchi, ACS Nano, 2025, 19, 12777–12786 Search PubMed.
- X.-X. Lin, J.-T. Lin, C.-Y. Tseng, S.-C. Lin, Z.-Y. He, Y. Chen, C.-K. Lin, K.-F. Lee, C.-C. Yang, C.-H. Wang, K.-H. Lin and T.-H. Yang, Adv. Funct. Mater., 2025, e19243 Search PubMed.
- K. M. Mayer, S. Lee, H. Liao, B. C. Rostro, A. Fuentes, P. T. Scully, C. L. Nehl and J. H. Hafner, ACS Nano, 2008, 2, 687–692 CrossRef CAS PubMed.
- R. G. Nuzzo, B. R. Zegarski and L. H. Dubois, J. Am. Chem. Soc., 1987, 109, 733–740 Search PubMed.
- J. C. Love, L. A. Estroff, J. K. Kriebel, R. G. Nuzzo and G. M. Whitesides, Chem. Rev., 2005, 105, 1103–1170 CrossRef CAS PubMed.
- L. Tian, S. Tadepalli, S. Hyun Park, K.-K. Liu, J. J. Morrissey, E. D. Kharasch, R. R. Naik and S. Singamaneni, Biosens. Bioelectron., 2014, 59, 208–215 Search PubMed.
- C. Dallari, C. Capitini, M. Calamai, A. Trabocchi, F. S. Pavone and C. Credi, Nanomaterials, 2021, 11, 665 Search PubMed.
- P. Peluso, D. S. Wilson, D. Do, H. Tran, M. Venkatasubbaiah, D. Quincy, B. Heidecker, K. Poindexter, N. Tolani, M. Phelan, K. Witte, L. S. Jung, P. Wagner and S. Nock, Anal. Biochem., 2003, 312, 113–124 Search PubMed.
- G. Kumari, J. Kandula and C. Narayana, J. Phys. Chem. C, 2015, 119, 20057–20064 Search PubMed.
- Y. Zhang, M. Ma, H. A. Aisa and L. Chen, Chemosensors, 2025, 13, 50 CrossRef CAS.
- S. Yoo, J. Park, D. H. Nam, S. Kim, D. Jeong, M.-K. Lee and S. Lee, Nanoscale Adv., 2025, 7, 2171–2181 Search PubMed.
- J. Gu, L. Wang, C. Liang, Q. Zhuang and J. Kong, J. Alloys Compd., 2018, 745, 430–435 Search PubMed.
- F. Ma and R. B. Lennox, Langmuir, 2000, 16, 6188–6190 Search PubMed.
- F. Frederix, K. Bonroy, W. Laureyn, G. Reekmans, A. Campitelli, W. Dehaen and G. Maes, Langmuir, 2003, 19, 4351–4357 Search PubMed.
- Z. Chen, Langmuir, 2022, 38, 4483–4489 Search PubMed.
- C. Leng, H.-C. Hung, S. Sun, D. Wang, Y. Li, S. Jiang and Z. Chen, ACS Appl. Mater. Interfaces, 2015, 7, 16881–16888 Search PubMed.
- Z. G. Estephan, P. S. Schlenoff and J. B. Schlenoff, Langmuir, 2011, 27, 6794–6800 Search PubMed.
- R. G. Chapman, E. Ostuni, L. Yan and G. M. Whitesides, Langmuir, 2000, 16, 6927–6936 Search PubMed.
- S. Chen, L. Li, C. L. Boozer and S. Jiang, Langmuir, 2000, 16, 9287–9293 Search PubMed.
- C. Singh, Y. Hu, B. P. Khanal, E. R. Zubarev, F. Stellacci and S. C. Glotzer, Nanoscale, 2011, 3, 3244–3250 Search PubMed.
- K. E. Nelson, L. Gamble, L. S. Jung, M. S. Boeckl, E. Naeemi, S. L. Golledge, T. Sasaki, D. G. Castner, C. T. Campbell and P. S. Stayton, Langmuir, 2001, 17, 2807–2816 Search PubMed.
- J. I. Cutler, E. Auyeung and C. A. Mirkin, J. Am. Chem. Soc., 2012, 134, 1376–1391 Search PubMed.
- S. Song, L. Wang, J. Li, C. Fan and J. Zhao, TrAC, Trends Anal. Chem., 2008, 27, 108–117 Search PubMed.
- S. Dalirirad and A. J. Steckl, Sens. Actuators, B, 2019, 283, 79–86 Search PubMed.
- D. Caprara, F. Ripanti, A. Capocefalo, A. Sarra, F. Brasili, C. Petrillo, C. Fasolato and P. Postorino, Colloids Surf., A, 2020, 589, 124399 Search PubMed.
- C. Zhou, Y. Yang, H. Li, F. Gao, C. Song, D. Yang, F. Xu, N. Liu, Y. Ke, S. Su and P. Wang, Nano Lett., 2020, 20, 3155–3159 Search PubMed.
- M. Shin, W. Kim, K. Yoo, H.-S. Cho, S. Jang, H.-J. Bae, J. An, J.-C. Lee, H. Chang, D.-E. Kim, J. Kim, L. P. Lee and B.-H. Jun, Nano Convergence, 2024, 11, 42 Search PubMed.
- L. Rivas, L. Hu, C. Parolo, A. Idili and A. Merkoçi, ACS Appl. Nano Mater., 2023, 6, 4151–4161 Search PubMed.
- Y. Yuan, F. Feng, R. Kong, C. Liu, J. Yang, D. Ji and H. Liu, Microchem. J., 2025, 213, 113721 Search PubMed.
- B. Lee, B. Park, D. Kim, C. Jung, J. H. Park, J.-H. Park, Y. E. Lee, M. G. Shin, M.-G. Kim, N. E. Yu, J. H. Kim and K. Kim, Nat. Commun., 2025, 16, 3377 Search PubMed.
- R. Gupta, P. Gupta, S. Wang, A. Melnykov, Q. Jiang, A. Seth, Z. Wang, J. J. Morrissey, I. George, S. Gandra, P. Sinha, G. A. Storch, B. A. Parikh, G. M. Genin and S. Singamaneni, Nat. Biomed. Eng., 2023, 7, 1556–1570 Search PubMed.
- Y. Hang, A. Wang, W. Tan, K. Bess, A. Eaton and N. Wu, Anal. Chem., 2025, 97, 1221–1228 Search PubMed.
- D. Hong, E.-J. Jo, D. Bang, C. Jung, Y. E. Lee, Y.-S. Noh, M. G. Shin and M.-G. Kim, ACS Nano, 2023, 17, 16607–16619 Search PubMed.
- S. K. Kim, J. U. Lee, M. J. Jeon, S. K. Kim, S. H. Hwang, M. E. Hong and S. J. Sim, RSC Adv., 2023, 13, 27225–27232 Search PubMed.
- W. Wang, S. Srivastava, A. Garg, C. Xiao, S. Hawks, J. Pan, N. Duggal, G. Isaacman-VanWertz, W. Zhou, L. C. Marr and P. J. Vikesland, Environ. Sci. Technol., 2024, 58, 4926–4936 Search PubMed.
- P. Liang, Q. Guo, T. Zhao, C.-Y. Wen, Z. Tian, Y. Shang, J. Xing, Y. Jiang and J. Zeng, Anal. Chem., 2022, 94, 8466–8473 Search PubMed.
- J. Li, M. Liu, J. Zhu, Y. Jiao and J. Zeng, Talanta, 2025, 285, 127351 Search PubMed.
- Y. Fu, J. Zhu, G. Weng, J. Li and J. Zhao, Sens. Actuators, B, 2025, 444, 138402 Search PubMed.
- J. Sun, L. Li, R. Sun, H. Yin and J. Liu, Anal. Chem., 2025, 97, 16515–16524 Search PubMed.
- S. Atta, Y. Zhao, J. Q. Li and T. Vo-Dinh, Anal. Chem., 2024, 96, 4783–4790 Search PubMed.
- T. Dong, X. Zhang, J. Yuan, Z. Lin, P. Yin, H. Yu, M. Wang and A. Liu, Anal. Chem., 2023, 95, 12532–12540 Search PubMed.
- Y. Chen, M. Zhang, X. Wang, X. Wang, Z. Majid, K. Huang, W. Xu, Y. Luo and C. Nan, Sens. Actuators, B, 2023, 383, 133601 Search PubMed.
- C. Wang, W. Shen, Z. Li, X. Xia, J. Li, C. Xu, S. Zheng and B. Gu, ACS Nano, 2024, 18, 25865–25879 Search PubMed.
- B. Wang, J. Li, J. Ai, Z. Xu, Y. Luo, C. He, J. Hu, Z. Li, C. Wang and B. Gu, Sens. Actuators, B, 2026, 446, 138654 Search PubMed.
- J. Cao, T. Li, X. Liao, J. Zhang, X. Yang, H. Yan, B. Sun, Y. Li, J. Huang, Y. Wang, X. Hu and L. Wang, Chem. Eng. J., 2025, 514, 163221 CrossRef CAS.
- Y. Sun, M. Mao, S. Tai, M. Chao, H. Xu, Y. Cai, C. Peng, W. Ma and Z. Wang, Talanta, 2025, 286, 127557 Search PubMed.
- S. E. Son, S. H. Cheon, W. Hur, H. B. Lee, D. H. Kim, C. H. Ha, S. J. Lee, D. K. Han and G. H. Seong, Biosens. Bioelectron., 2024, 243, 115752 Search PubMed.
- V. Panferov, N. Ivanov, W. Zhang, S. Wang and J. Liu, ACS Sens., 2025, 10, 3785–3793 Search PubMed.
- F. Cun, H. Li, H. Wang, B. Yang, J. Kong and H. Chen, Small, 2025, 21, 2408101 Search PubMed.
- Z. Ren and Y. Cui, Anal. Chim. Acta, 2025, 1371, 344405 Search PubMed.
- A. Roda, S. Cavalera, F. Di Nardo, D. Calabria, S. Rosati, P. Simoni, B. Colitti, C. Baggiani, M. Roda and L. Anfossi, Biosens. Bioelectron., 2021, 172, 112765 Search PubMed.
- D. Liu, C. Ju, C. Han, R. Shi, X. Chen, D. Duan, J. Yan and X. Yan, Biosens. Bioelectron., 2021, 173, 112817 CrossRef CAS PubMed.
- J. M. Park, H. W. Jung, Y. W. Chang, H. S. Kim, M. J. Kang and J. C. Pyun, Anal. Chim. Acta, 2015, 853, 360–367 Search PubMed.
- J. Feng, X. Lin, L. Kang, M. Duan, N. Duan, Z. Wang and S. Wu, Biosens. Bioelectron., 2026, 296, 118315 CrossRef CAS PubMed.
- Z. Li, J. Hou, W. Shi, K. Zhai, J. Shen, Z. Cai, B. Zhang and C. Liu, Talanta, 2026, 300, 129234 CrossRef CAS PubMed.
- L. Guo, D.-M. Zhao, S. Chen and Y.-L. Yu, Microchem. J., 2025, 208, 112593 CrossRef CAS.
- Y. Zhao, Y. Huang, X. Zhao, J. F. McClelland and M. Lu, Nanoscale, 2016, 8, 19204–19210 Search PubMed.
- Y. Zhong, X. T. Zheng, Q.-L. Li, X. J. Loh, X. Su and S. Zhao, Biosens. Bioelectron., 2023, 224, 115033 CrossRef CAS PubMed.
- M. Ding, L. Dou, T. Bu, Z. Li, Y. Mao, M. Dang, X. Huang, L. Song, Z. Wang and X. Zhang, Biosens. Bioelectron., 2025, 267, 116779 CrossRef CAS PubMed.
- L. Bian, Q. Fu, Z. Gan, Z. Wu, Y. Song, Y. Xiong, F. Hu and L. Zheng, Adv. Sci., 2024, 11, 2305774 CrossRef CAS PubMed.
- J. Wang, F. Cao, S. He, Y. Xia, X. Liu, W. Jiang, Y. Yu, H. Zhang and W. Chen, Talanta, 2018, 176, 444–449 Search PubMed.
- L. Anfossi, F. Di Nardo, S. Cavalera, C. Giovannoli, G. Spano, E. S. Speranskaya, I. Y. Goryacheva and C. Baggiani, Microchim. Acta, 2018, 185, 94 Search PubMed.
- A. Perju and N. Wongkaew, Anal. Bioanal. Chem., 2021, 413, 5535–5549 CrossRef CAS PubMed.
- T. A. Ivandini, W. P. Wicaksono, E. Saepudin, B. Rismetov and Y. Einaga, Talanta, 2015, 134, 136–143 Search PubMed.
- A. Miglione, F. Di Nardo, S. Cavalera, T. Serra, C. Baggiani, S. Cinti and L. Anfossi, Anal. Chem., 2024, 96, 2297–2302 CrossRef CAS PubMed.
- D. Shi, Y. Yin, X. Li and J. Yuan, ACS Sens., 2024, 9, 5293–5301 CrossRef CAS PubMed.
- Y. Blickenstorfer, V. Jirasko, A. Tanno, S. Dräger, D. Hoven, J. Löhle, S. Leuch, Y. Mamedava, S. L. Müller, K. Leuzinger, M. Osthoff and J. Vörös, Biosens. Bioelectron., 2024, 262, 116524 CrossRef CAS PubMed.
- Q. Chen, H. Wang, H. Xu, Y. Peng, B. Yao, Z. Chen, J. Yang, S. Adeloju and W. Chen, Biosens. Bioelectron., 2025, 285, 117529 CrossRef CAS PubMed.
- P. D. Sinawang, L. Fajs, K. Elouarzaki, J. Nugraha and R. S. Marks, Sens. Actuators, B, 2018, 259, 354–363 Search PubMed.
- P. Nandhakumar, C. Muñoz San Martín, B. Arévalo, S. Ding, M. Lunker, E. Vargas, O. Djassemi, S. Campuzano and J. Wang, ACS Sens., 2023, 8, 3892–3901 CrossRef CAS PubMed.
- A. Sharma, A. I. Y. Tok, C. Lee, R. Ganapathy, P. Alagappan and B. Liedberg, Sens. Actuators, B, 2019, 285, 431–437 CrossRef CAS.
- W. Ren and J. Irudayaraj, ACS Omega, 2022, 7, 29204–29210 Search PubMed.
- Z. Du, Y. Wang, D. He, E. Xu, Q. Chai, Z. Jin, Z. Wu and B. Cui, Food Chem., 2022, 397, 133756 Search PubMed.
- Q. Wu, Y. Xiao, X. Yang, A. Zhu, W. Cao, L. Cai, X. Lin, Z. Zhao, Q. Zhang and X. Zhou, Talanta, 2025, 290, 127834 Search PubMed.
- J. Chen, X. Liu, Z. Liu, J. Ma, J. Han, Y. Sun, J. Liang, H. Han, J. Zhao, B. Wang, R. Xiao and Y. Wang, Sens. Actuators, B, 2024, 409, 135598 CrossRef CAS.
- J. Liang, J. Zeng, X. Huang, T. Zhu, Y. Gong, C. Dong, X. Wang, L. Zhao, L. Xie, K. Liang, Q. Tan, Y. Cui, B. Kong and W. Hui, Nano Res., 2023, 16, 1242–1251 Search PubMed.
- Q. Bayin, L. Huang, C. Ren, Y. Fu, X. Ma and J. Guo, Talanta, 2021, 227, 122207 CrossRef CAS PubMed.
- J. Park, Sens. Actuators, A, 2016, 250, 55–59 Search PubMed.
- X.-H. Mu, H.-F. Liu, Z.-Y. Tong, B. Du, S. Liu, B. Liu, Z.-W. Liu, C. Gao, J. Wang and H. Dong, Sens. Actuators, B, 2019, 284, 638–649 Search PubMed.
- S. Oh, S. Anandakumar, C. Lee, K. W. Kim, B. Lim and C. Kim, Sens. Actuators, B, 2011, 160, 747–752 Search PubMed.
- C. Ren, Q. Bayin, S. Feng, Y. Fu, X. Ma and J. Guo, Biosens. Bioelectron., 2020, 165, 112340 Search PubMed.
- V. A. Bragina, S. L. Znoyko, A. V. Orlov, A. V. Pushkarev, M. P. Nikitin and P. I. Nikitin, Anal. Chem., 2019, 91, 9852–9857 Search PubMed.
- A. V. Orlov, S. L. Znoyko, V. R. Cherkasov, M. P. Nikitin and P. I. Nikitin, Anal. Chem., 2016, 88, 10419–10426 CrossRef CAS PubMed.
- A. V. Orlov, J. A. Malkerov, D. O. Novichikhin, S. L. Znoyko and P. I. Nikitin, Food Chem., 2022, 383, 132427 Search PubMed.
- L. Hong, K. Wang, W. Yan, H. Xu, Q. Chen, Y. Zhang, D. Cui, Q. Jin and J. He, Theranostics, 2018, 8, 6121–6131 CrossRef CAS PubMed.
- Y. Wang, H. Xu, M. Wei, H. Gu, Q. Xu and W. Zhu, Mater. Sci. Eng., C, 2009, 29, 714–718 Search PubMed.
- K. Wang, T. Li, B. Cao, H. Xu, Y. Cheng, C. Zheng, W. Zheng and D. Cui, Sens. Actuators, A, 2022, 333, 113299 CrossRef CAS.
- J. Eveness, J. Kiely, P. Hawkins, P. Wraith and R. Luxton, Sens. Actuators, B, 2009, 139, 538–542 CrossRef CAS.
- B. Wang, T. Peng, Z. Jiang, J. Xu, J. Qu and X. Dai, ACS Sens., 2023, 8, 4512–4520 Search PubMed.
- J. R. Sonawane, R. Jundale and A. A. Kulkarni, Mater. Horiz., 2025, 12, 364–400 Search PubMed.
- J. W. M. Crawley, I. E. Gow, N. Lawes, I. Kowalec, L. Kabalan, C. R. A. Catlow, A. J. Logsdail, S. H. Taylor, N. F. Dummer and G. J. Hutchings, Chem. Rev., 2022, 122, 6795–6849 CrossRef CAS PubMed.
- B. A. Yusuf, W. Yaseen, J. Xie, A. A. Babangida, A. I. Muhammad, M. Xie and Y. Xu, Nano Energy, 2022, 104, 107959 Search PubMed.
- X. Lin, P. Zhou, M. An, C. Zhu, Y. Pang and R. Xiao, Adv. Sci., 2025, 12, e2500134 Search PubMed.
- X. Liu, X. Yang, Z. Zhao, X. Li, J. Liang, Y. Sun, R. Xiao and G. Wang, Chem. Eng. J., 2024, 499, 155995 CrossRef CAS.
- Z. Chen, K. Lai, A. Wang, H. Ji, S. Yu, Z. Fang, D. Liu, J. Peng and W. Lai, ACS Nano, 2025, 19, 9282–9291 CrossRef CAS PubMed.
- B. Tian, J. Zhang, S. Zhou, C. Liu, Q. Wang, Y. Liu, M. Lu, G. Sun, C. Wang and B. Gu, Chem. Eng. J., 2025, 515, 163557 CrossRef CAS.
- X. Shu, P. Guo, G. Zhang, W. Zhang, H. Hu, J. Peng, Y. Xiong, B. Ma and W. Lai, Food Chem., 2024, 450, 139380 Search PubMed.
- Y. Deng, Y. Wang, M. Lin, Y. Chen, Z.-J. Qian, J. Liu and X. Li, Anal. Chem., 2024, 96, 5106–5114 Search PubMed.
- J. Deng, M. Yang, J. Wu, W. Zhang and X. Jiang, Anal. Chem., 2018, 90, 9132–9137 Search PubMed.
- Y. Huang, Y. Ji, M. Zheng, X. Li, Y. Luan, Y. Liu and X. Zhang, ACS Sens., 2024, 9, 2815–2825 Search PubMed.
- L. Shi, Z. Wang, Y. Li, J. Wang, J. Shan, J. Zhuo, X. Yin, J. Sun, D. Zhang and J. Wang, J. Agric. Food Chem., 2024, 72, 4405–4414 Search PubMed.
- W. Shen, J. Li, S. Zheng, S. Wang, C. Wang, J. Yin and C. Wang, Chem. Eng. J., 2025, 514, 163223 CrossRef CAS.
- M. Li, D. Chen, Y. Guo, T. Li, W. Guo, R. Feng, D. Shao, Z. Jiang, Y. Li and J. Zan, Food Chem., 2026, 503, 147785 CrossRef CAS PubMed.
- J. Li, P. Liang, T. Zhao, G. Guo, J. Zhu, C. Wen and J. Zeng, Anal. Bioanal. Chem., 2023, 415, 545–554 CrossRef CAS PubMed.
- S. Zhang, S. Wang, B. Sun, S. Chen, Q. Ma, K. Han, C. Yin, X. Wang and H. Jiang, Sens. Actuators, B, 2025, 425, 137000 Search PubMed.
- C. Zheng, Q. Jiang, K. Wang, T. Li, W. Zheng, Y. Cheng, Q. Ning and D. Cui, Analyst, 2022, 147, 1678–1687 Search PubMed.
- L. Wu, Z. Zhu, J. Xue, L. Zheng, H. Liu, H. Ouyang, Z. Fu and Y. He, Biosens. Bioelectron., 2024, 265, 116711 CrossRef CAS PubMed.
- J. Sun, Y. Zhao, Y. Hou, H. Li, M. Yang, Y. Wang and B. Sun, New J. Chem., 2019, 43, 13381–13387 RSC.
- Y. Li, S. Qi, P. Chen, C. Peng and Z. Wang, Biosens. Bioelectron., 2025, 286, 117619 Search PubMed.
- M. Xu, S. Zhao, W. Zhang, D. Li, Y. Peng, M. Tanemura, Z. Huang, M. Liu and Y. Yang, Sens. Actuators, B, 2025, 426, 137020 CrossRef CAS.
- M. Xu, S. Zhao, C. Lin, Y. Li, W. Zhang, Y. Peng, R. Xiao, Z. Huang and Y. Yang, ACS Appl. Mater. Interfaces, 2024, 16, 11172–11184 Search PubMed.
- S. Liu, Z. Li, C. Li, S. Liu, Y. Lang, X. Zhang, B. Zhang and C. Liu, Sens. Actuators, B, 2024, 413, 135877 CrossRef CAS.
- F. Zhang, J. Wang, W. Liu, Q. Shi, H. Liu, Y. Sang and X. Wang, Talanta, 2025, 285, 127275 CrossRef CAS PubMed.
- L. Huang, H. Chang, H. Han, L. Ao and J. Wang, Anal. Chem., 2025, 97, 17631–17640 Search PubMed.
- B. Sun, V. Panferov, X. Guo, J. Xiong, S. Zhang, L. Qin, C. Yin, X. Wang, C. Liu, K. Han, S. Wang and H. Jiang, Biosens. Bioelectron., 2025, 267, 116797 CrossRef CAS PubMed.
- W. Zhi, X. Guo, Y. Nie, L. Nie, T. He, L. Dai, J. Xu, X. Zong, J. Xu, H. Cai, Y. Tang, Y. Cong, J. Pi, Y. Zhu, P. Sun, J. Guo, H. Chen, X. Huang and H. Zhou, Chem. Eng. J., 2025, 520, 166410 Search PubMed.
- W. Li, X. Wang, M. Zhu, X. Huang, P. H. Umutoni, T.-H. Chen, J. Lu, S.-C. Chen, G. Tan, B. P. Yan and B. L. Khoo, Talanta, 2025, 293, 128100 CrossRef CAS PubMed.
- C. Chen, S. Hu, L. Tian, M. Qi, Z. Chang, L. Li, L. Wang and B. Dong, Biosens. Bioelectron., 2024, 252, 116135 CrossRef CAS PubMed.
- Z. Wang, R. Zou, J. Yi, Y. Wang, H. Hu, C. Qi, W. Lai, Y. Guo and Y. Xianyu, Small, 2024, 20, 2310869 CrossRef CAS PubMed.
- J. Liang, L. Wu, Y. Wang, W. Liang, Y. Hao, M. Tan, G. He, D. Lv, Z. Wang, T. Zeng, X. Zhang, C. Lu, Q. Song, B. Peng, J. Zhao, B. Zhu and Y. Tang, Sens. Actuators, B, 2023, 389, 133875 CrossRef CAS.
- H. Yang, Q. He, M. Lin, L. Ji, L. Zhang, H. Xiao, S. Li, Q. Li, X. Cui and S. Zhao, J. Hazard. Mater., 2022, 435, 129082 CrossRef CAS PubMed.
- X. Lin, P. Zhou, Q. Li and Y. Pang, Anal. Chem., 2024, 96, 10686–10695 CrossRef CAS PubMed.
- P. Wu, W. Zuo, Y. Wang, Q. Yuan, J. Yang, X. Liu, H. Jiang, J. Dai, F. Xue and Y. Ju, Chem. Eng. J., 2023, 451, 139021 CrossRef CAS.
- J. Ding and X. Gao, ACS Appl. Nano Mater., 2025, 8, 16823–16832 CrossRef CAS.
- S. Yang, J. Du, M. Wei, Y. Huang, Y. Zhang, Y. Wang, J. Li, W. Wei, Y. Qiao, H. Dong and X. Zhang, Anal. Chim. Acta, 2023, 1239, 340660 CrossRef CAS PubMed.
- J. Zhu, G. Guo, J. Liu, X. Li, X. Yang, M. Liu, C. Fu, J. Zeng and J. Li, Anal. Chim. Acta, 2024, 1292, 342241 CrossRef CAS PubMed.
- C. Wang, Y. Cheng, X. Yin, Q. Wu, J. Ma, Q. Zhang, L. Zhao, J. Wang and D. Zhang, J. Agric. Food Chem., 2024, 72, 18171–18180 CrossRef CAS PubMed.
- X. Peng, X. Mei, J. Yang, J. Liu and Y. Li, Anal. Chem., 2023, 95, 5807–5814 CrossRef CAS PubMed.
- X. Cai, F. Ma, J. Jiang, X. Yang, Z. Zhang, Z. Jian, M. Liang, P. Li and L. Yu, J. Hazard. Mater., 2023, 441, 129853 CrossRef CAS PubMed.
- Y. Tang, P. Ma, I. M. Khan, W. Cao, Y. Zhang and Z. Wang, Food Chem., 2024, 460, 140398 CrossRef CAS PubMed.
- X. Lai, W. Cao, G. Zhang, E. H. Ang, L. Su, C. Liu, W. He, W. Lai and S. Deng, Chem. Eng. J., 2024, 501, 157565 CrossRef CAS.
- Q. Wu, B. Liu, A. M. Abd El-Aty, X. Zhang, L. Chen, G. Liu, X. Xu, J. Wang, M. Jin, Q. Wang, X. Huang, G. Chen and D. Xu, TrAC, Trends Anal. Chem., 2025, 193, 118455 CrossRef CAS.
- S. Liu, Y. Liao, R. Shu, J. Sun, D. Zhang, W. Zhang and J. Wang, ACS Nano, 2024, 18, 27167–27205 CrossRef CAS PubMed.
- D. M. Kinyua, D. M. Memeu, C. N. Mugo Mwenda, B. D. Ventura and R. Velotta, Sensors, 2025, 25, 5414 CrossRef CAS PubMed.
- Y. Zhang, Y. Yu and J. Y. Ying, Adv. Funct. Mater., 2022, 32, 2109553 CrossRef CAS.
- C. Wang, C. Wang, J. Li, Z. Tu, B. Gu and S. Wang, Biosens. Bioelectron., 2022, 214, 114525 CrossRef CAS PubMed.
- M. Wang, J. Feng, J. Ding, J. Xiao, D. Liu, Y. Lu, Y. Liu and X. Gao, Chem. Eng. J., 2024, 487, 150666 CrossRef CAS.
- M. Sánchez-Purrà, M. Carré-Camps, H. de Puig, I. Bosch, L. Gehrke and K. Hamad-Schifferli, ACS Infect. Dis., 2017, 3, 767–776 Search PubMed.
- D. Zhang, L. Huang, B. Liu, E. Su, H.-Y. Chen, Z. Gu and X. Zhao, Sens. Actuators, B, 2018, 277, 502–509 Search PubMed.
- Y. Zhao, H. Wang, P. Zhang, C. Sun, X. Wang, X. Wang, R. Yang, C. Wang and L. Zhou, Sci. Rep., 2016, 6, 21342 CrossRef CAS PubMed.
- N. Cheng, Q. Shi, C. Zhu, S. Li, Y. Lin and D. Du, Biosens. Bioelectron., 2019, 142, 111498 CrossRef CAS PubMed.
- F. Schenk, P. Weber, J. Vogler, L. Hecht, A. Dietzel and G. Gauglitz, Anal. Bioanal. Chem., 2018, 410, 863–868 CrossRef CAS PubMed.
- M. Lu, Y. Joung, C. S. Jeon, S. Kim, D. Yong, H. Jang, S. H. Pyun, T. Kang and J. Choo, Nano Convergence, 2022, 9, 39 CrossRef CAS PubMed.
- C.-Y. Wen, X. Yang, T.-Y. Zhao, J. Qu, K. Tashpulatov and J. Zeng, Biosens. Bioelectron., 2025, 271, 117030 CrossRef CAS PubMed.
- Y. Wu, Y. Zhou, H. Huang, X. Chen, Y. Leng, W. Lai, X. Huang and Y. Xiong, Sens. Actuators, B, 2020, 316, 128107 CrossRef CAS.
- N. A. Byzova, Y. Y. Vengerov, S. G. Voloshchuk, A. V. Zherdev and A. B. B. Dzantiev, Sensors, 2019, 19, 5494 Search PubMed.
- A. Sena-Torralba, J. Gabaldón-Atienza, A. Cubells-Gómez, P. Casino, Á. Maquieira and S. Morais, Biosensors, 2022, 12, 980 CrossRef CAS PubMed.
- C. Jiang, G. Wang, R. Hein, N. Liu, X. Luo and J. J. Davis, Chem. Rev., 2020, 120, 3852–3889 CrossRef CAS PubMed.
- S. Gao, X. Zheng, J. Zhu, Y. Zhang, R. Zhou, T. Wang, J. Katona, D. Zhang and X. Zou, Coord. Chem. Rev., 2025, 534, 216588 CrossRef CAS.
- D. Cialla-May, A. Bonifacio, T. Bocklitz, A. Markin, N. Markina, S. Fornasaro, A. Dwivedi, T. Dib, E. Farnesi, C. Liu, A. Ghosh and J. Popp, Chem. Soc. Rev., 2024, 53, 8957–8979 RSC.
- M. Wei, H. Rao, Z. Niu, X. Xue, M. Luo, X. Zhang, H. Huang, Z. Xue and X. Lu, Coord. Chem. Rev., 2021, 447, 214149 CrossRef CAS.
- C. Fang, J. Li, B. Lin, Y. Wang, Y. Yao, L. Chen, Y. Zeng, L. Li and L. Guo, Anal. Chem., 2024, 96, 721–729 CrossRef CAS PubMed.
- Y. Pang, Z. Yang, X. Liu, X. Shen, H. Lei and X. Li, Foods, 2025, 14, 3264 Search PubMed.
- J. Jeon, S. H. Lee, Y. Joung, K. Kim, N. Choi and J. Choo, Sens. Actuators, B, 2020, 321, 128521 Search PubMed.
- S. Lee, S. Kim, D. S. Yoon, J. S. Park, H. Woo, D. Lee, S.-Y. Cho, C. Park, Y. K. Yoo, K.-B. Lee and J. H. Lee, Nat. Commun., 2023, 14, 2361 CrossRef CAS PubMed.
- S. Lee, Y. K. Yoo, S. I. Han, D. Lee, S.-Y. Cho, C. Park, D. Lee, D. S. Yoon and J. H. Lee, Analyst, 2023, 148, 6001–6010 RSC.
- T. Fairooz, S. E. McNamee, D. Finlay, K. Y. Ng and J. McLaughlin, Biosens. Bioelectron., 2023, 223, 115016 CrossRef CAS PubMed.
- S. Uddin, I. Haque, H. Lu, M. A. Moni and E. Gide, Sci. Rep., 2022, 12, 6256 CrossRef CAS PubMed.
- S. Wang, Y. Zhu, Z. Zhou, Y. Luo, Y. Huang, Y. Liu and T. Xu, Adv. Sci., 2024, 11, e2406196 CrossRef PubMed.
- W. Wang, K. Chen, X. Ma and J. Guo, Fundam. Res., 2023, 3, 544–556 Search PubMed.
- V. L. Deringer, A. P. Bartók, N. Bernstein, D. M. Wilkins, M. Ceriotti and G. Csányi, Chem. Rev., 2021, 121, 10073–10141 CrossRef CAS PubMed.
- F. Leclercq, Phys. Rev. D, 2018, 98, 063511 CrossRef CAS.
- T. Fairooz, S. E. McNamee, D. Finlay, K. Y. Ng and J. McLaughlin, Biosensors, 2024, 14, 611 CrossRef CAS PubMed.
- S. Zhang, L. Chen, Y. Tan, S. Wu, P. Guo, X. Jiang and H. Pan, Anal. Methods, 2024, 16, 6715–6725 Search PubMed.
- B. Sun, H. Wu, T. Fang, Z. Wang, K. Xu, H. Yan, J. Cao, Y. Wang and L. Wang, Anal. Chem., 2025, 97, 4824–4831 Search PubMed.
- Y. Zha, Y. Li, J. Zhou, X. Liu, K. S. Park and Y. Zhou, Anal. Chem., 2024, 96, 12197–12204 CrossRef CAS PubMed.
- L. Alzubaidi, J. Zhang, A. J. Humaidi, A. Al-Dujaili, Y. Duan, O. Al-Shamma, J. Santamaría, M. A. Fadhel, M. Al-Amidie and L. Farhan, J. Big Data, 2021, 8, 53 CrossRef PubMed.
- D. Bermejo-Peláez, N. Medina, E. Álamo, J. C. Soto-Debran, O. Bonilla, M. Luengo-Oroz, J. L. Rodriguez-Tudela and A. Alastruey-Izquierdo, J. Fungi, 2023, 9, 217 CrossRef PubMed.
- D. Bermejo-Peláez, A. Alastruey-Izquierdo, N. Medina, D. Capellán-Martín, O. Bonilla, M. Luengo-Oroz and J. L. Rodríguez-Tudela, IMA Fungus, 2024, 15, 27 Search PubMed.
- X. Zang, Y. Zhou, S. Li, G. Shi, H. Deng, X. Zang, J. Cao, R. Yang, X. Lin, H. Deng, Y. Huang, C. Yang, N. Wu, C. Song, L. Wu and X. Xue, Talanta, 2025, 284, 127254 CrossRef CAS PubMed.
- S. Martin, M. A. Kohn, J. Bollyky and J. Parsonnet, Diagn. Microbiol. Infect. Dis., 2022, 104, 115763 Search PubMed.
- N. Cheng, Y. Tao, J. Yang, C. Shen, C. Zhang and B. Lou, Biosens. Bioelectron., 2026, 296, 118290 CrossRef CAS PubMed.
- T. U. Rehman, M. Alruwaili, M. H. Siddiqi, Y. Alhwaiti, S. Anwar, Z. Halim and M. Alam, Complex Intell. Syst., 2025, 11, 398 Search PubMed.
- L. Tang and C. Shen, Front. Cardiovasc. Med., 2025, 12, 1606159 Search PubMed.
- S. Chaabene, A. Boudaya, B. Bouaziz and L. Chaari, Int. J. Data Sci. Anal., 2025, 20, 3093–3117 CrossRef.
- E. Nasarian, R. Alizadehsani, U. R. Acharya and K.-L. Tsui, Inf. Fusion, 2024, 108, 102412 CrossRef.
- E. Tjoa and C. Guan, IEEE Trans. Neural Netw. Learn. Syst., 2021, 32, 4793–4813 Search PubMed.
- M. Haupt, M. H. Maurer and R. P. Thomas, Diagnostics, 2025, 15, 1399 Search PubMed.
- M. A. Talukder, A. S. Talaat, M. Kazi and A. Khraisat, Artif. Intell. Rev., 2025, 58, 385 Search PubMed.
- R. Agrawal, T. Gupta, S. Gupta, S. Chauhan, P. Patel and S. Hamdare, Diagn. Pathol., 2025, 20, 105 CrossRef PubMed.
- J. Feng, M. Xiong, Y. Qiao, P. Wang, X. Lin, L. Kang, N. Duan, Z. Wang and S. Wu, Food Chem., 2025, 481, 144058 CrossRef CAS PubMed.
- Y. Huang, Y. Ji, M. Wu, W. Xu, L. Wang, Y. Liu, T. Xu and X. Zhang, Sens. Actuators, B, 2025, 438, 137779 Search PubMed.
- S. Mahari, D. Prakashan and S. Gandhi, Colloids Surf., B, 2023, 226, 113319 CrossRef CAS PubMed.
- P. Phangwipas, B. Thangavel and J. H. Shin, Chemosensors, 2023, 11, 36 CrossRef CAS.
- C. Parolo, A. Sena-Torralba, J. F. Bergua, E. Calucho, C. Fuentes-Chust, L. Hu, L. Rivas, R. Álvarez-Diduk, E. P. Nguyen, S. Cinti, D. Quesada-González and A. Merkoçi, Nat. Protoc., 2020, 15, 3788–3816 CrossRef CAS PubMed.
- N. Cheng, Y. Song, M. M. A. Zeinhom, Y. C. Chang, L. Sheng, H. Li, D. Du, L. Li, M. J. Zhu, Y. Luo, W. Xu and Y. Lin, ACS Appl. Mater. Interfaces, 2017, 9, 40671–40680 CrossRef CAS PubMed.
- N. Cheng, Y. Song, Q. Shi, D. Du, D. Liu, Y. Luo, W. Xu and Y. Lin, Anal. Chem., 2019, 91, 13986–13993 CrossRef CAS PubMed.
- J. Wang, H. Jiang, Y. Chen, X. Zhu, Q. Wu, W. Chen, Q. Zhao, J. Wang and P. Qin, Biosens. Bioelectron., 2024, 249, 116046 CrossRef CAS PubMed.
- J. Wang, C. Jiang, J. Jin, L. Huang, W. Yu, B. Su and J. Hu, Angew. Chem., Int. Ed., 2021, 60, 13042–13049 CrossRef CAS PubMed.
- T. Mu, S. Li, H. Feng, C. Zhang, B. Wang, X. Ma, J. Guo, B. Huang and L. Zhu, IEEE J. Sel. Top. Quantum Electron., 2019, 25, 1–6 Search PubMed.
- S. Dutta, TrAC, Trends Anal. Chem., 2019, 110, 393–400 CrossRef CAS.
- W. Xiao, C. Huang, F. Xu, J. Yan, H. Bian, Q. Fu, K. Xie, L. Wang and Y. Tang, Sens. Actuators, B, 2018, 266, 63–70 CrossRef CAS PubMed.
- H. Xu, A. Xia, J. Luo, M. Gao, R. Liao, F. Li, Q. Zhong, W. Zhang, Y. Wang, J. Cui, W. Fu, K. Chang, M. Gan, W. Jiang and M. Chen, Sens. Actuators, B, 2020, 308, 127750 CrossRef CAS.
- L. Huang, W. Xiao, T. Xu, H. Chen, Z. Jin, Z. Zhang, Q. Song and Y. Tang, Sens. Actuators, B, 2021, 327, 128893 CrossRef CAS.
- M. Yentongchai, K. Chomthong, S. Nuanualsuwan, N. Khongchareonporn, S. Ruantip, M. M. Sain, K. Pungjunun and S. Chaiyo, Sens. Actuators, B, 2025, 441, 138003 CrossRef CAS.
- L. Gonzalez-Macia, Y. Li, K. Zhang, E. Nunez-Bajo, G. Barandun, Y. Cotur, T. Asfour, S. Olenik, P. Coatsworth, J. Herrington and F. Güder, Biosens. Bioelectron., 2024, 251, 116124 CrossRef CAS PubMed.
- P. Brangel, A. Sobarzo, C. Parolo, B. S. Miller, P. D. Howes, S. Gelkop, J. J. Lutwama, J. M. Dye, R. A. McKendry, L. Lobel and M. M. Stevens, ACS Nano, 2018, 12, 63–73 CrossRef CAS PubMed.
- D. A. Mendels, L. Dortet, C. Emeraud, S. Oueslati, D. Girlich, J. B. Ronat, S. Bernabeu, S. Bahi, G. J. H. Atkinson and T. Naas, Proc. Natl. Acad. Sci. U. S. A., 2021, 118, e2019893118 CrossRef CAS PubMed.
- J. L. D. Nelis, Y. Zhao, L. Bura, K. Rafferty, C. T. Elliott and K. Campbell, Anal. Chem., 2020, 92, 7852–7860 CrossRef CAS PubMed.
- J. Sivakumar, J. H. Yang, M. S. Kelly, A. Koh and D. Won, Expert Syst. Appl., 2022, 210, 118471 CrossRef.
- M. Hoque Tania, K. T. Lwin, A. M. Shabut, M. Najlah, J. Chin and M. A. Hossain, Expert Syst. Appl., 2020, 139, 112843 CrossRef.
- M. Colombo, L. Bezinge, A. Rocha Tapia, C. J. Shih, A. J. de Mello and D. A. Richards, Sens. Diagn., 2023, 2, 100–110 RSC.
- D. Bermejo-Peláez, D. Marcos-Mencía, E. Álamo, N. Pérez-Panizo, A. Mousa, E. Dacal, L. Lin, A. Vladimirov, D. Cuadrado, J. Mateos-Nozal, J. C. Galán, B. Romero-Hernandez, R. Cantón, M. Luengo-Oroz and M. Rodriguez-Dominguez, JMIR Public Health Surveill., 2022, 8, e38533 CrossRef PubMed.
- S. Kumar, T. Ko, Y. Chae, Y. Jang, I. Lee, A. Lee, S. Shin, M. H. Nam, B. S. Kim, H. S. Jun and S. Seo, Biosensors, 2023, 13, 623 CrossRef PubMed.
- P. P. Ray, J. King Saud Univ. Comput. Inf. Sci., 2018, 30, 291–319 CrossRef.
- R. M. L. Gulraiz, J. Joyia, A. Farooq and S. Rehman, J. Commun., 2017, 12, 240–247 Search PubMed.
- S. Li, L. D. Xu and S. Zhao, J. Ind. Inf. Integr., 2018, 10, 1–9 Search PubMed.
- S.-m Park, D. D. Won, B. J. Lee, D. Escobedo, A. Esteva, A. Aalipour, T. J. Ge, J. H. Kim, S. Suh, E. H. Choi, A. X. Lozano, C. Yao, S. Bodapati, F. B. Achterberg, J. Kim, H. Park, Y. Choi, W. J. Kim, J. H. Yu, A. M. Bhatt, J. K. Lee, R. Spitler, S. X. Wang and S. S. Gambhir, Nat. Biomed. Eng., 2020, 4, 624–635 CrossRef PubMed.
- J. Guo, S. Chen, S. Tian, K. Liu, X. Ma and J. Guo, Talanta, 2021, 230, 122335 CrossRef CAS PubMed.
- J. Guo, S. Chen, S. Tian, K. Liu, J. Ni, M. Zhao, Y. Kang, X. Ma and J. Guo, Biosens. Bioelectron., 2021, 181, 113160 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |