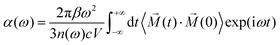Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Ion vibrational spectroscopy of the microhydrated iodate anion: unveiling the onset of heterogeneous ion solvation
Arghya Chakraborty a,
Huiyan Libd,
Zhi-Feng Liu
a,
Huiyan Libd,
Zhi-Feng Liu *bc and
Knut R. Asmis
*bc and
Knut R. Asmis *a
*a
aWilhelm-Ostwald-Institut für Physikalische und Theoretische Chemie, Universität Leipzig, Linnéstrasse 2, D-04103, Leipzig, Germany. E-mail: knut.asmis@uni-leipzig.de
bDepartment of Chemistry and Centre for Scientific Modelling and Computation, Chinese University of Hong Kong, Hong Kong, China. E-mail: zfliu@cuhk.edu.hk
cGanjiang Innovation Academy, Chinese Academy of Sciences, Ganzhou, Jiangxi 341000, China
dCollege of Life Science and Chemistry, Jiangsu Second Normal University, Nanjing, China
First published on 11th May 2026
Abstract
Gas-phase ion vibrational spectroscopy is combined with first principles calculations to provide a bottom-up view of how the iodate anion is microhydrated from one to twenty water molecules. Infrared photodissociation spectra of D2-tagged IO3−(H2O)n (n = 1–8 and 11) are reported in the O–H stretching (2800–3800 cm−1) and the H–O–H bending (1600–1800 cm−1) region. Additionally, infrared multiple photon dissociation spectra of bare IO3−(H2O)n (n = 10–12, 15, and 20) were recorded in the O–H stretching region. Structure assignments are made by comparing experimental spectra with simulated ones derived from MP2 frequency calculations and ab initio molecular dynamics simulations. Two solvation regions around the trigonal pyramidal iodate anion are identified based on their characteristic vibrational signatures. The first water molecules prefer to interact with the negatively polarized O-atoms of iodate by hydrogen bond donation. Notably, interwater hydrogen bonds are first detected at n = 3 as part of a water trimer ring. Starting at n = 4, a water tetramer ring motif is found and it persists in all larger systems. The emergence of a free-OH oscillator feature (∼3713 cm−1) signals the onset of I-atom solvation at n = 11, manifested in the form of Iδ+⋯OH2 interactions as a result of the I atom's electropositive nature in IO3−. The interwater hydrogen bonds associated with I-atom solvation are considerably stronger compared to those associated with O-atom solvation, resulting in characteristically red-shifted (<3000 cm−1) O–H stretching bands. Overall, this study sheds new light on polyoxoanion solvation, complementing insights from condensed-phase experiments.
1. Introduction
The iodate anion (IO3−) is the most abundant iodine species in sea-water, the main source of atmospheric iodine, and plays a critical role in facilitating “new particle formation” required for cloud generation.1–3 However, iodate hydration cannot be accurately described using conventional models that assume a spherically symmetric charge distribution,4,5 as this trigonal pyramidal polyoxyanion exhibits a positively polarized I-atom with a formal charge of +1 separated spatially from three negatively charged O-atoms (−2/3).5 Hence, more realistic solvation models require knowledge of specific ion–water interactions in (at least) the first solvation shell. This motivates the study of microhydrated ions isolated in the gas phase, which are amenable to, both, characterization using highly sensitive and selective gas phase ion spectroscopy techniques as well as to higher level electronic structure calculations.The effect of iodate's asymmetric charge distribution on the structure and dynamics of its solvation shell has been the topic of previous experimental and computational studies.5–9 X-ray absorption fine structure (XAFS) spectroscopy combined with density functional theory-based ab initio molecular dynamics simulations (AIMD) indeed provides evidence for such a bipolar anion with both anionic and cationic regions.7 Interface-specific vibrational spectroscopy reveals that iodate is prevalent at the air–water interface due to the distinct water structure of stronger as well as weaker hydrogen bonding in the iodate's hydration shell.10 Subsequently, the impact of charge anisotropy on the hydration environment in microhydrated iodate anions, IO3−(H2O)n, was investigated using anion photoelectron spectroscopy.9 The most stable structures reported in this publication suggest that the first four water molecules form strong ionic hydrogen bonds to the iodate's O-atoms in either a DD or an ADD fashion, where the letters stand for hydrogen bond acceptor (A) and hydrogen bond donor (D).11 Characteristic anion microhydration moieties have been previously reported and labelled as W1–4 ring motifs according to the size of the hydrogen-bonded network.12,13 A single DD–H2O, donating two hydrogen bonds to IO3−, represents the simplest motif, W1 (see Scheme 1 for an overview of the binding motifs W1 to W4). W2 is composed of an ADD– and a DD–H2O linked by a single interwater hydrogen bond and donates three hydrogen bonds to IO3−. Three ADD–H2O molecules form the W3 motif, a three-membered ring, interconnected by three equatorial interwater hydrogen bonds, and the three axial OHs each donating a hydrogen bond to IO3−. The W4 motif contains a four-membered water ring and is typically part of a larger hydrogen bond network such that each water molecule is at least three-fold coordinated (e.g. four ADD–H2Os). Wang and coworkers found that starting with n = 5 the second solvation shell is populated, i.e., water–water hydrogen bond formation competes with ionic hydrogen bond formation, leading to the generation of particularly stable W4 rings without any dangling O–H bonds.9 They predicted that at n = 12, eight water molecules, all bound to the iodate's three O-atoms, comprise the first solvation-shell.
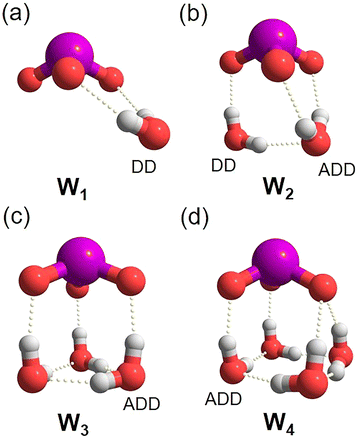 | ||
| Scheme 1 Characteristic binding motifs W1–W4 (labelled a–d) found in microhydrated anions,12,13 in general, and in iodate anions, in particular. Individual water molecules are labeled according to their role as a hydrogen bond acceptor (A) or donor (D). A water molecule can accept up to two and provide up to two hydrogen bonds at maximum coordination (AADD–H2O). W4 typically appears in combination with other motifs in larger clusters. | ||
A comparative study of the influence of four halide anions (F−, Cl−, Br−, and I−) vs. IO3− on the hydrogen bond network in aqueous nanodroplets (n = 30–75) was carried out using gas-phase vibrational action spectroscopy and demonstrated that I− and IO3−, despite having nearly same ionic radii, exhibit differences in hydration motifs that propagate to water molecules beyond the second solvation shell.14 Two distinct solvation regions were also identified in a recent AIMD study on microhydrated iodate anions with 30 or more water molecules.8 This simulation suggests that the iodate's first solvation shell is composed of eleven to twelve water molecules. Of these, around eight to nine H2O molecules engage in hydrogen bonding with one or more of the three O-atoms of the iodate, the so-called oxygen solvation region. The remaining two to three H2O molecules primarily interact with the electropositive iodine center developing the iodine solvation region.
In the present study, we produce microhydrated iodate anions, IO3−(H2O)n, with up to twenty water molecules using a nanospray ion source (see Methodology, Section 4.1). Ion vibrational spectra in the O–H stretching region reveal characteristically red-shifted bands, where the magnitude of the shifts correlates with the strength of the corresponding hydrogen bond.15,16 We assign structures and identify particularly stable microhydration motifs based on a comparison with the results of harmonic frequency calculations as well as AIMD simulations. One of the key experimental findings is the onset of I-atom solvation at n = 11, evidenced by the emergence of a free-OH feature (>3700 cm−1) together with substantially red-shifted bands (<3000 cm−1) associated with hydrogen-bonded O–H oscillators in the iodine solvation region.
2. Results and discussion
2.1 Vibrational spectroscopy
To characterize the structure of microhydrated iodate anions with up to 20 water molecules, we measured vibrational action spectra, using D2 as a messenger tag when possible, in order to obtain IRPD spectra in the linear absorption regime. Since the D2-tagging efficiency decreases with cluster size, it becomes difficult to obtain sufficient tagging yields for n > 8. For n = 11, we made an extra effort, since this is the first spectrum that contains the signature of a free O–H oscillator. For all larger clusters, we report the IRMPD spectra of the untagged microhydrated anion. IRMPD spectra for such clusters are typically less structured, the measured intensities do not necessarily reflect the calculated linear absorption cross sections, in particular, when measuring at longer wavelengths, and therefore we refrain from assigning structures based on a comparison of calculated IR spectra and measured IRMPD spectra.Fig. 1 compares the IRPD spectra (left) of D2-tagged IO3−(H2O)n with n = 1–8 and 11 to the corresponding predicted IR spectra (right) derived from scaled harmonic frequencies of the best fit isomer in the O–H stretching (3800–2800 cm−1) and the H–O–H bending (1800–1600 cm−1) regions. The experimental band positions for n = 1–5 and 11 and their assignments are provided in Table 1. Absorption bands arising from the excitation of O–H stretching modes involved in ionic hydrogen bonds as well as water–water hydrogen bonds are observed between 3680 cm−1 and 2900 cm−1,11 while bands attributed to free OH oscillators are expected above 3680 cm−1.17 The small feature at ∼2960 cm−1 (denoted by an asterisks in Fig. 1) observed up to n = 7 is assigned to excitation of the D–D stretching mode of the messenger molecule. It gains in IR intensity due to polarization and is slightly red-shifted from the frequency of the free D2 molecule (2994 cm−1), suggesting charge transfer into the antibonding σ* orbital of D2.18
| n = 1 | 2 | 3 | 4 | 5 | 11 | Assignment |
|---|---|---|---|---|---|---|
| 3713 (i1) | Free O–H stretch | |||||
| 3524 (a1) | 3550 (b1) | 3335(c2) | 3523 (d1) | 3515 (e1) | O–H stretch (HO–H···OIO2−) | |
| 3504 (a2) | 3526 (b2) | 3428(c3) | 3461 (d2) | |||
| 3411 (d3) | 3425 (e2) | |||||
| 3541(c1) | 3344 (d4) | 3338(e3) | O–H stretch (HO–H···OH2) | |||
| 1687 (a3) | 1676 (b3) | 1679 (c5) | 1690 (d5) | 1683(e4) | HOH bend | |
| 1693 (c4) | ||||||
| 2960 (i4) | O–H stretch (H–O(H)···IO3−) |
Based on the features observed in the O–H stretching region, the experimental IRPD spectra on the left of Fig. 1 can be grouped into four microsolvation regimes. (1) Up to n = 2, the spectra are simple with no significant absorption above 3550 cm−1 or below 3500 cm−1. (2) From n = 3 to n = 5, intense IR features are also observed in the 3200–3550 cm−1 region. (3) The spectra become more congested with size, as expected. Starting at n = 6, substantial absorption is also observed above 3600 cm−1, tailing out to ∼3700 cm−1, however not as intense as the more red-shifted O–H stretching bands. (4) Finally, the IRPD spectrum at n = 11 shows two new features, which are not observed in any of the spectra of the smaller complexes. These are a relatively weak but characteristic free-O–H stretching band at 3713 cm−1 (i1) and substantially red-shifted bands below 3000 cm−1 (i4), both shaded in red in Fig. 1, signaling a characteristic change in the hydration structure at n = 11.
The H2O bending region is less diagnostic, in particular for n > 4, due to overlapping absorption bands between 1640 cm−1 and 1730 cm−1, with band maxima located between 1675 and 1685 cm−1, presumably reflecting the locally slightly different environment of the H2O molecules.
Additional information on the first three of these microsolvation regimes can be obtained by comparing these spectra to those previously reported for the microhydrated perchlorate anion (ClO4−)19 and sulfate dianion (SO4−2).13 For n = 1, a similar IR absorption profile was reported for ClO4−(H2O) and assigned to the W1 motif (see Fig. 1a).19 The substantially more red-shifted band at 3504 cm−1 (a2) for IO3−(H2O) vs. 3575 cm−1 for ClO4−(H2O) suggests stronger ionic hydrogen bonds in the iodate complexes as a result of iodate's larger anion proton affinity. For n = 2 (trace 1b), a similar spectrum is observed with slightly different relative intensities, suggesting a similar assignment. Hence, the second water molecule also likely adopts the W1 motif, leading to a 2W1 arrangement in which iodate's third available O-atom is also engaged, rather than forming a water–water hydrogen bond as part of the W2 motif (see Fig. 1b), which was reported for ClO4−(H2O)2.19 Hence, this first microhydration regime up to n = 2 is characterized by DD–H2O's bound to iodate O-atoms without any interwater hydrogen bonds.
In microhydrated ClO4−, the onset of water–water hydrogen bonding, involving predominantly ADD–H2O's as part of three- and four-membered water rings, is signaled by intense IR absorption features, red-shifted up to 300 cm−1 more than those associated with OH oscillators involved in isolated ionic hydrogen bonds. Indeed, for the iodate anions with n = 3 to n = 5, additional bands are observed from 3500 cm−1 down to 3200 cm−1, favoring a similar assignment. The region 3600–3700 cm−1 is characteristic for the formation of four-fold-coordinated water molecules and we therefore assume that also for iodate the signal above 3600 cm−1 starting with n = 6 signals the formation of AADD–H2Os as the hydrogen-bonded network grows. What causes the additional features starting with n = 11 remains unclear at this point. Further below, we demonstrate that both these features signal the onset of microhydration of the iodine solvation region.
Since messenger-tagging is inefficient for larger microhydrated clusters, we reverted to IRMPD spectroscopy, monitoring the water loss channels, to obtain vibrational information on the larger clusters with up to 20 water molecules. These spectra are shown in Fig. 2. The emergence of the free O–H stretching band at 3713 cm−1 (i1) is evident at n = 11 too, which aligns well with the corresponding IRPD spectrum in Fig. 1i. In the current study, this band is observed up to n = 20. The absorption between 3200 and 3650 cm−1 in Fig. 2 loses any distinctive pattern with increasing number of H2O molecules. A similar absorption profile, including the weak free O–H stretching feature, was reported to persist for larger clusters of up to n = 78.14 The weak absorption signal apparent between 3200 and 3100 cm−1 was attributed to AAD water molecules.14,20 In contrast to the IRPD spectrum of n = 11 (Fig. 1i), the IRMPD spectra in Fig. 2 show no significant signal below 3000 cm−1, which we attribute to IRMPD transparency (see Section 4.1).21 Thus, the IRMPD spectra are less diagnostic when it comes to the more red-shifted O–H stretching bands.
2.2 Microhydration motifs and energetics
In order to identify the structure of the individual iodate–water clusters, we performed a minimum-energy structure search using the MP2/def2-TZVP method. Characteristic low-energy isomers are shown in Fig. 3 [see Fig. S1a–c of the SI for higher energy structures]. Relative energies, without and with zero-point energy (ZPE) correction, are listed in Table 2 (see also Tables S1a and b in the SI). In the text, we refer to the ZPE-corrected energies (ΔE0) if not noted otherwise.| Size n | Isomers | DFT | MP2 | ||
|---|---|---|---|---|---|
| ΔE | ΔE0 | ΔE | ΔE0 | ||
| 1 | 1.2.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| 2 | 2.3.1 | 0.0 | 0.0 | 0.0 | 0.0 |
| 2.4.0 | 3.7 | 1.0 | 3.2 | 0.5 | |
| 3 | 3.3.3 | 0.0 | 0.0 | 0.0 | 0.0 |
| 3.4.2 | 14.8 | 10.5 | 12.9 | 9.0 | |
| 4 | 4.4.4 | 0.0 | 0.0 | 0.0 | 0.0 |
| 4.5.3a | 12.5 | 9.2 | 9.4 | 6.0 | |
| 5 | 5.5.5a | 0.0 | 0.0 | 0.0 | 0.0 |
| 5.5.5b | 7.3 | 5.6 | 7.2 | 6.0 | |
| 6 | 6.5.6a | 0.0 | 0.0 | 0.0 | 0.0 |
| 6.5.6b | 1.6 | 1.0 | 1.1 | 0.7 | |
| 7 | 7.5.9a | 0.0 | 0.0 | 0.0 | 0.0 |
| 7.5.9b | 3.7 | 2.1 | 2.2 | 1.3 | |
| 8 | 8.6.10 | 0.0 | 0.0 | 0.0 | 0.0 |
| 8.7.8 | 13.9 | 10.2 | 11.6 | 8.1 | |
| 11 | 11.7.14 | 0.0 | 0.0 | 0.0 | 0.0 |
| 11.6.14 | 0.6 | 0.4 | 2.2 | 1.5 | |
| 11.7.15 | 7.5 | 7.0 | 7.2 | 6.4 | |
Structures are labelled using the m.i.bw nomenclature scheme, where m refers to the number of water molecules, and i refers to the number of ion–water hydrogen bonds and bw is the number of water–water hydrogen bonds.22 In addition, the letters a–z are used in ascending order according to ΔE0 to differentiate between multiple (m.i.bw) isomers.
| W1 → W2 → W3 → W4 → W4 + W1 → W4 + W2 → W4 + W3 → 2W4 |
For n = 1, 3, 4, 5 and 8, the lowest energy isomer is more than 5 kJ mol−1 lower in energy, while for n = 2, 6, 7 and 11 two low-energy constitutional isomers are predicted only within 2 kJ mol−1.
The minimum-energy structures shown in Fig. 3 predict that up to n = 8 only the oxygen solvation region is populated. The formation of ionic hydrogen bonds competes with the formation of interwater hydrogen bonds already in the smallest possible complex, namely at n = 2, indicating that an increase in the coordination of a water molecule lowers the total energy more than that of one of iodate's O-atoms. It is noted that while 2.3.1 (W2) is predicted to be 0.5 kJ mol−1 more stable than 2.4.0 (2W1), the latter is entropically favored, yielding a lower free energy already at 100 K (Table S2 in the SI). In any case, hydrogen-bonded network structures are formed early on, which also effectively avoids the formation of any energetically unfavorable free O–H oscillators. The first microsolvation shell containing an ADD–H2O is predicted for n = 2 and remains the most abundant species in all the larger complexes. Population of the second solvation shell, i.e. a water molecule without direct contact with the iodate anion, is found for n = 5. The first four-fold-coordinated AADD–H2O is predicted for n = 6, the first complex that exhibits more water–water than ionic hydrogen bonds. Interestingly, the maximum (solvation) coordination number of iodate O-atoms is only two up to n = 8. The structures in Fig. 3 also show that (at least at low temperatures) the W3 and, in particular, the W4 ring motif are particularly stable. Indeed, for n = 6, the labeling as “W4 + W2” above is somewhat misleading. It correctly reflects the stoichiometry of the complex, but its network structure is probably better described as a “W4 + W4” motif, in which two water molecules are shared by each ring.
The first global minimum-energy structure, in which the iodine solvation region is populated, is found for n = 11. 11.7.14 is above 6 kJ mol−1 more stable than the “oxygen solvation region” structure 11.7.15. Besides the presence of an attractive Iδ+⋯OH2 interaction, both iodine solvation region structures in Fig. 3 contain a free OH oscillator, either on an AD–H2O (11.7.14) or an AAD–H2O (11.6.14). The structural analysis of 11.7.14 is discussed in more detail in Section 2.3 (vide infra).
Summarizing, the trends observed in the evolution of the global minimum-energy structures of microhydrated iodate complexes with up to eleven water molecules confirm and consolidate the observations derived from the experimental IRPD spectra. The four identified microsolvation regimes indeed correlate directly with the degree of coordination of the water molecules comprising the solvation shell. In the first regime (n = 1, 2), only DD–H2O molecules are present in the complex. Entropic factors need to be considered in the case of n = 2, in order to bring the experimental observation in agreement with the predictions from the calculations. ADD–H2Os (and DD–H2O) contribute to the second regime (n = 3–5) and finally at least one AADD–H2O is always present in the third regime (n = 6–8). While in the first three regimes microsolvation is limited to the oxygen solvation region, the iodine solvation region is populated at n = 11, which is directly correlated with the presence of (at least) one free OH oscillator.
2.3 Analysis of the vibrational spectra
Microhydrated iodate anions are bound by hydrogen bonds which are considerably weaker than intramolecular bonds and easily rearrange at finite temperature. Their potential energy surfaces are typically flat, with many isomeric structures lying close to each other in energy. It is not always straightforward to assign IRPD spectra based on a comparison to simulated spectra derived from harmonic IR frequencies and intensities of the static minimum-energy structures (at 0 K). Temperature induces free energy and dynamic changes. Therefore, we also performed ab initio molecular dynamic (AIMD) simulations on IO3−(H2O)n clusters to examine such effects (details are provided in Methodology, Section 4).The harmonic IR spectra of the best-fit IO3−(H2O)n isomers (n = 1–8 and 11) are shown in Fig. 1 (right panel). In the following, we provide a more detailed analysis of the experimental IRPD spectra based on a comparison to computed harmonic, and when necessary, to anharmonic IR spectra. This allows us to assign the relevant spectral features and provides the basis for our structure assignment.
The IRPD spectra of IO3−(H2O) and IO3−(H2O)2 in Fig. 4 (trace 4a and 4e) are satisfactorily reproduced by the harmonic IR spectra for 1.2.0 and 2.4.0 (4b and 4f), respectively, which appear similar, since the corresponding structures exclusively exhibit the W1 motif. In contrast, the agreement with the spectrum of 2.3.1 is worse, underscoring our initial finding (see Section 2.1) that the microhydration regime up to n = 2 is characterized by DD–H2O's bound to iodate O-atoms without any interwater hydrogen bonds. For n = 1 (n = 2), IRPD bands a1 (b1), a2 (b2) and a3 (b3) are assigned to the excitation of the H2O antisymmetric stretching, symmetric stretching and bending fundamental, respectively. The weak feature at ∼3350 cm−1 (#) in both IRPD spectra is attributed to the excitation of the H2O bend overtone. It is reproduced when static anharmonic effects are considered (see the SI, trace c of Fig. S2a, for the results of the MP2/VPT2).
 | ||
| Fig. 4 (Left panel) Comparison of the IRPD spectrum (trace a) of D2-tagged IO3−(H2O) with the scaled harmonic MP2/def2TZVP (trace b) and unscaled AIMD simulated IR spectra of the (untagged) lowest energy isomer 1.2.0 at temperatures of 20 K (trace c) and 100 K (trace d). (Right panel) Comparison of the IRPD spectrum (trace e) of D2-tagged IO3−(H2O)2 with the scaled harmonic MP2/def2TZVP (trace f and g) and unscaled AIMD simulated IR spectra (h and i) of (untagged) lowest energy isomers 2.4.0 and 2.3.1, respectively, at a temperature of 100 K. The feature attributed to the wiggling motion of the water molecule is denoted by # and the D2 stretching band is denoted by an asterisk (*). ZPE-corrected relative energies ΔE0 (in kJ mol−1) are reported in parentheses. See Table 1 for band positions and assignments. | ||
The weak signal around 3450 cm−1 is attributed to dynamic effects (see trace 4a vs. 4c), which become more prominent with increasing simulation temperature (trace 4d). It is noted that the AIMD simulation suggests that 2.4.0 efficiently isomerizes to 2.3.1, the global minimum-energy structure, at 20 K, but not anymore at 100 and 200 K. Hence, the free energy barrier separating these two isomers increases at higher simulation temperatures. Indeed, the 100 K spectrum of 2.4.0 (trace 4e vs. 4h, right panel) reproduces the IRPD spectrum quite well, in particular, the characteristic splitting between bands b1 and b2 as well as the weaker features at slightly lower wavenumbers. It is noted that the temperature in an AIMD simulation is scaled to the average kinetic energy, without considering the zero-point energies, and only provides an indication for the extent of the dynamic motion in the cluster rather than the thermodynamic cluster temperature.
The harmonic IR spectra of the two lowest energy isomers for n = 3 and n = 4 are shown in Fig. 5. In both cases, better agreement with the measured IRPD spectrum is found for the lowest energy isomers 3.3.3 and 4.4.4, which both exclusively exhibit ADD–H2O molecules. This confirms the onset of water-ring formation at n = 3, which manifests itself in the calculated IR spectra by additional, more red-shifted bands in the region below 3500 cm−1 (c1). For n = 3, the harmonic spectra allow the band c2 at ∼3425 cm−1 to be assigned to the excitation of the three OH oscillators bound to the iodate core, which are well decoupled from the three OH oscillators comprising the W3-ring contributing to band c1 at ∼3540 cm−1. Hence, the ionic hydrogen bonds in n = 3 are stronger than the water–water hydrogen bonds of the W3 motif. However, the situation is reversed for n = 4 and the most red-shifted band d4 at 3352 cm−1 is indeed assigned to the excitation of OH oscillators in the plane of the W4 motif, reflecting the cooperative stabilization of the water network concomitant with the weakening of the ionic hydrogen bonds to the iodate core. It is noted that multiple peaks in this region (the green-shaded area of Fig. 1) highlight the stronger coupling between ionic hydrogen bond and water–water stretching modes (compared to those in the n = 3 system). Such mode mixing persists up to n = 11, contributing to the spectral complexity in the IRPD spectra of the larger clusters. The spectra in Fig. 6 (vide infra) show that the bands in the HOH bending region are not sufficiently diagnostic and we will therefore not discuss them any further.
The AIMD simulations also emphasize the growing significance of dynamic fluctuations as the cluster size increases. For example, the W3 motif in 3.3.3 comprises three longer equatorial inter-water hydrogen bonds (202 pm) and three shorter axial hydrogen bonds (192 pm) interacting with iodate's three O-atoms (see the SI, Fig. S5). A similar motif was previously identified for NO3−(H2O)3 and shown to be stable during AIMD simulations,12 even at 200 K. However, the present W3 motif in 3.3.3 is less stable and ruptured already at 20 K (see the SI, Fig. S5), likely due to the 30% larger O–O distance in IO3− (290 pm) compared to NO3− (220 pm). Such a deformation strengthens some hydrogen bonds, giving rise to peak C in the AIMD spectrum, which reproduces peak c3 in the IRPD spectrum (Fig. 5d, left panel) that could not be accounted for by comparison to the harmonic spectrum of the two lowest energy isomers (see traces a and b of Fig. 5, left panel).
In contrast, the W4 ring in 4.4.4 remains stable at 100 K, suggesting a better compatibility with the larger size of IO3− (vs. NO3−) and hence a more rigid structure. However, at 200 K, the W4 ring does break, with one H2O molecule moving into the second solvation shell, leading to considerable signal broadening in the OH stretching region (Fig. 5e, right panel). The relative intensities, on the other hand, are less well reproduced, suggesting that capturing the relative contribution of the dynamic solvation motifs is challenging.
Spectral congestion becomes more evident as the number of water molecules (in different coordination environments) increases. For n = 5, the computed IR spectrum of 5.5.5a agrees well with the IRPD spectrum, but does not reproduce the IRPD band e2 (see Fig. 6, left panel), which suggests a contribution from higher energy isomers like 5.5.5b. The AIMD simulations confirm that 5.5.5a remains stable even at 200 K and the corresponding AIMD trace obtained at both 100 K (trace d in Fig. 6) reproduces the IRPD spectrum (trace a) rather well. The OH bonds involved in inter-water hydrogen bond interactions within the W4 ring predominantly contribute to the higher-frequency region (group A in trace 6d, left panel), while the OH bonds associated with axial hydrogen bonds and those from the W1 ring contribute to the lower-frequency region (group B feature).
For n = 6 and n = 7, the harmonic IR spectrum of the lowest energy structures, 6.5.6a and 7.5.9a (see Fig. 3 and Fig. S1) provides the best agreement with the experimental IRPD spectrum, confirming the onset of the formation of structures that contain (four-fold-coordinated) AADD–H2O molecules, but isomers with similar structures are predicted within a few kJ mol−1 (see the SI, Fig. S6 and S7) and probably also contribute. For n = 8, the lowest energy isomer 8.6.10 also provides the best agreement and is likely to be the only isomer present, since this 2W4 motif is 8 kJ mol−1 more stable than other structures.
The above structure assignments confirm that up to eight water molecules exclusively bind in the oxygen solvation region in such a way that the formation of W3 and, in particular, W4 motifs is favored and the formation of free dangling OH moieties is avoided. A characteristic cubic motif involving two O atoms from iodate and six water molecules stacked as W2 on top of W4, first found in 7.5.9a, is predicted to persist in the larger clusters (Fig. 6 and Fig. S8 in the SI) and found quite stable even at 200 K, despite the shuffling of other H2O molecules.
The predicted IR spectra for three low energy isomers of n = 11 are shown in Fig. 7. The spectra are spectrally considerably more congested than those of the smaller clusters and all exhibit multiple IR bands between 3200 and 3600 cm−1, in agreement with the experimental observation. However, only the two lowest energy isomers, 11.7.14 and 11.6.14, contain a free OH group and only their calculated IR spectra reproduce the characteristic IRPD band i1 above 3700 cm−1. The spectra of both isomers also exhibit a characteristic IR band below 3200 cm−1, 11.7.14 a band at 3115 cm−1 and 11.6.14 an even more red-shifted band at 2977 cm−1, which agree well with the IRPD bands i3 and i4, respectively, suggesting that both contribute to the IRPD spectrum. Peaks i3 and i4 correspond to the excitation of an OH oscillator of the water molecule directly interacting with the I-atom. However, the I⋯O interaction comes at the expense of the hydrogen-bonded network among water molecules, resulting in the formation of a free dangling O–H bond. The AIMD simulations starting from 11.6.14 and 11.7.14 (see Fig. 7e and f) confirm the stability of these isomers at 100 K. At 200 K, isomer interconversion is observed, but the free OH group and the I⋯OH2 interaction persist throughout these simulations. In summary, the iodine solvation region is first populated at n = 11, concomitant with the formation of a free OH group, either of the water interacting with the I-atom (11.6.14) or of another (11.7.14).
3. Summary and conclusions
The microhydration of the iodate anion was systematically investigated using IRPD spectroscopy in combination with static electronic structure calculations and AIMD simulations. This study provides a detailed molecular-level understanding of the solvation structure and dynamics of the iodate anion. The findings complement surface-sensitive spectroscopic observations in condensed phases and support recent theoretical predictions. Moreover, the present work offers a benchmark for future comparative studies on the hydration and solvation processes of related pyramidal polyoxyanions, including BrO3−, ClO3−, and SO32−.The primary structural motifs identified in iodate microhydration correspond to four characteristic hydrogen-bonding arrangements (W1–W4), whose stability reflects the competition between ion–water and water–water hydrogen bonds. The cooperative nature of these interactions significantly influences the vibrational features of the water ring motifs. In larger clusters, a particularly stable cubic motif involving two O atoms from iodate and six water molecules is identified. In principle, direct structural assignment based solely on static theoretical spectra becomes increasingly challenging for larger aggregates; however, such comparisons remain highly valuable for identifying systematic trends in solvation behavior, especially when complemented by molecular dynamics simulations. For example, this combined approach enables a reliable quantification of the number of water molecules (n) associated with the completion of oxygen solvation regions followed by the onset of the iodine solvation region of the anion.
In aqueous solution, IO3− resides beneath the topmost interfacial water layer, where its O atoms interact with surface water molecules while the iodine center remains oriented toward the bulk.10 This arrangement results in distinct solvation domains, with the inner iodine solvation region exhibiting stronger hydrogen bonding than the outer oxygen solvation region. The present results obtained for gas-phase model systems mirrors this behavior. At n = 11, the identified low-energy structures display two distinct solvation regions, with hydrogen bonds in the iodine solvation region being over 10% shorter than those in the oxygen region, indicating stronger interactions.
4. Methodology
4.1 Experimental methods
IRPD experiments were performed using the Leipzig 6 K ion trap triple mass spectrometer described previously.22 Briefly, microhydrated iodate anions, IO3−(H2O)n, are generated using a nanospray ion source from a 10 mM solution of KIO3 in the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 mixture of H2O and acetonitrile (CH3CN). The beam of anions is skimmed, thermalized to room temperature in a helium-filled radio-frequency (RF) ion-guide, and then mass-selected using a quadrupole mass filter. The quadruple mass spectrum is presented in Fig. S9 of the SI. Then, the mass-selected ion beam is focused into a RF ring-electrode ion trap held at ∼13 K. Ions are thermalized to the ambient temperature of the trap by many collisions with the buffer gas. At sufficiently low ion-trap temperatures, ion-messenger complexes, in this case D2-tagged anions, are formed via three-body collisions.23
1 mixture of H2O and acetonitrile (CH3CN). The beam of anions is skimmed, thermalized to room temperature in a helium-filled radio-frequency (RF) ion-guide, and then mass-selected using a quadrupole mass filter. The quadruple mass spectrum is presented in Fig. S9 of the SI. Then, the mass-selected ion beam is focused into a RF ring-electrode ion trap held at ∼13 K. Ions are thermalized to the ambient temperature of the trap by many collisions with the buffer gas. At sufficiently low ion-trap temperatures, ion-messenger complexes, in this case D2-tagged anions, are formed via three-body collisions.23
Every 100 ms, all ions are extracted from the ion trap and focused both temporally and spatially into the center of the extraction region of an orthogonally mounted double-focusing reflectron time-of-flight (TOF) tandem mass spectrometer. Background-free action spectra are obtained using the IR1MS2 technique.24,25 The ion packet is accelerated into the reflectron stage; ions spread out in space according to their mass-to-charge ratio (m/z) and are refocused at the initial extraction region. Just before reacceleration towards the MCP detector, ions with a particular m/z value are irradiated by a properly timed widely wavelength tunable IR laser pulse (bandwidth: 3.5 cm−1) from an optical parametric oscillator/amplifier (LaserVision: OPO/OPA/AgGaSe2) laser system.26 An IRPD spectrum is measured by continuously scanning the laser wavelength, which is monitored online using a HighFinesse WS6-600 wavelength meter with a scan speed such that an averaged TOF mass spectrum (over 80 laser shots) is obtained every 2 cm−1. Typically, at least five scans are measured and averaged and the photodissociation cross section (σIRPD) is determined as described previously.22,27 We measured IRPD spectra in two spectral regions: 1600–1800 cm−1 and 2800–3800 cm−1.
For larger clusters (n ≥ 10) the messenger tagging efficiency decreases substantially, making the measurement of IRPD spectra with satisfactory signal/noise ratio increasingly difficult. Therefore, IRMPD measurements without D2-tagging were carried out in the range of 2800–3800 cm−1 by monitoring the H2O dissociation channel. To characterize the differences in the two types of spectra, IRPD and IRMPD measurements for the same n (n = 11) were performed (see Fig. S10 in the SI). The spectra are similar above 3500 cm−1, but the IRMPD spectrum quickly loses relative intensity with decreasing wavenumber and little to no signal is observed below 3200 cm−1. We attribute this to the decrease in efficiency of the multiple photon dissociation process with decreasing photon energy as a result of the substantially higher dissociation energies of the (bare) microhydrated anions compared to the D2-tagged species.21
4.2 Computational methods
A thorough search of the lowest energy structures for n = 1–8 and 10–12 was performed based on previously published structures as well as chemical intuition. Initial geometries for n = 1–8 were taken from ref. 9 considering all low energy structures (ΔE ≤ +10 kJ mol−1). Geometries were then optimized using the B3LYP/def2TZVP and MP2/def2TZVP methods and the energy ordering was determined.28–30 Zero-point-energy (ZPE) correction has been shown to be necessary to obtain a reliable relative energy-ordering for moderately sized microhydrated ions.13,31,32 Therefore, unscaled harmonic frequencies were used to obtain ZPE-corrected relative energies (ΔE0). The MP2/def2TZVP harmonic frequencies in the O–H stretching region (>2800 cm−1) are scaled by a factor 0.96 to correct for anharmonic effects.33 Stick spectra are convoluted with a Gaussian line shape function with a full width at half-maximum of 12 cm−1 for better comparability. Anharmonic MP2/def2TZVP frequency calculations have been carried out using the second-order vibrational perturbation theory (VPT2)34 as implemented by Bloino and Barone. All these electronic structure calculations are carried out using the Gaussian 16 program package.35Ab initio molecular dynamic (AIMD) simulations at finite temperatures are performed using the CP2K package.36 A cluster under simulation is placed at the center of a periodic cubic box with a length of more than twice the largest interatomic distance in the cluster and with the effect of the periodic charge density images eliminated by the decoupling technique developed by Martyna and Tuckerman.37 At every time step, the potential energy and atomic forces are calculated within the framework of density functional theory (DFT). The wave functions are represented by double-zeta Gaussians and the electron density is represented by using a mixed basis set of Gaussians and auxiliary plane waves with a cutoff of 320 Rydberg. The Geodecker–Teter–Hutter (GTH) typed pseudopotentials are employed for the atomic cores.38 For the exchange and correlation energy, the BLYP functional39 is employed with dispersion corrected using Grimme's D3 scheme.40
The atomic motion is treated by Newtonian mechanics, with the temperature controlled using a Nose–Hoover thermostat and with a time step of 0.5 fs. The temperature is first scaled at the desired value for 3 ps, followed by a data collection run in the NVE ensemble for 15 ps. The value of dipole moment is calculated and collected at each time step, so that a vibrational spectrum can be directly simulated by the Fourier transform of the dipole time-correlation function (DTCF),41
![[M with combining right harpoon above (vector)]](https://www.rsc.org/images/entities/i_char_004d_20d1.gif) is the total dipole moment of the system, calculated by the polarization including both ionic and electronic contributions. Dynamic and anharmonic effects are automatically taken into account in such a scheme, although quantum effects at low temperature are not included.
is the total dipole moment of the system, calculated by the polarization including both ionic and electronic contributions. Dynamic and anharmonic effects are automatically taken into account in such a scheme, although quantum effects at low temperature are not included.
Author contributions
AC planned the study and executed the recording of IRPD spectra. AC also performed static computer simulations such as geometry optimization and harmonic and anharmonic spectrum calculations. The analysis and interpretation of the experimental spectra combined with static calculations were performed by AC and KRA. AIMD simulations and their interpretation were made by HL and ZFL. KRA and ZFL coordinated the project and supervised the research. All authors were involved in the preparation of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The authors confirm that the data supporting the findings of this study are available within the article and its supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d6cp01186a.Acknowledgements
This work was supported by the German Research Foundation (DFG) as part of the individual research Grant No. AS133/3-1 “Spectroscopic Characterization of Salt Dissolution in Microhydrated Cluster Ions and at the Water/Vapor Interface.” KRA acknowledges instrumental support from the Fritz Haber Institute of the Max Planck Society.References
- R. C. Tian and E. Nicolas, Mar. Chem., 1995, 48(2), 151 CrossRef CAS.
- C. Moreno, M.-T. Baeza-Romero, M. Sanz, Ó. Gálvez, V. López Arza, J. C. Ianni and E. Espíldora, Phys. Chem. Chem. Phys., 2020, 22(10), 5625 RSC.
- H. Finkenzeller, S. Iyer, X.-C. He, M. Simon, T. K. Koenig, C. F. Lee, R. Valiev, V. Hofbauer, A. Amorim, R. Baalbaki, A. Baccarini, L. Beck, D. M. Bell, L. Caudillo, D. Chen, R. Chiu, B. Chu, L. Dada, J. Duplissy, M. Heinritzi, D. Kemppainen, C. Kim, J. Krechmer, A. Kürten, A. Kvashnin, H. Lamkaddam, C. P. Lee, K. Lehtipalo, Z. Li, V. Makhmutov, H. E. Manninen, G. Marie, R. Marten, R. L. Mauldin, B. Mentler, T. Müller, T. Petäjä, M. Philippov, A. Ranjithkumar, B. Rörup, J. Shen, D. Stolzenburg, C. Tauber, Y. J. Tham, A. Tomé, M. Vazquez-Pufleau, A. C. Wagner, D. S. Wang, M. Wang, Y. Wang, S. K. Weber, W. Nie, Y. Wu, M. Xiao, Q. Ye, M. Zauner-Wieczorek, A. Hansel, U. Baltensperger, J. Brioude, J. Curtius, N. M. Donahue, I. E. Haddad, R. C. Flagan, M. Kulmala, J. Kirkby, M. Sipilä, D. R. Worsnop, T. Kurten, M. Rissanen and R. Volkamer, Nat. Chem., 2023, 15(1), 129 CrossRef CAS PubMed.
- P. Jungwirth and D. J. Tobias, Chem. Rev., 2006, 106(4), 1259 CrossRef CAS PubMed.
- M. D. Baer and C. J. Mundy, Faraday Discuss., 2013, 160, 89–101 RSC ; discussion 103–120.
- W. Kunz, P. Lo Nostro and B. W. Ninham, Curr. Opin. Colloid Interface Sci., 2004, 9(1), 1 CrossRef CAS.
- M. D. Baer, V.-T. Pham, J. L. Fulton, G. K. Schenter, M. Balasubramanian and C. J. Mundy, J. Phys. Chem. Lett., 2011, 2(20), 2650 CrossRef CAS.
- B. Sharma and A. Chandra, J. Comput. Chem., 2018, 39(19), 1226 CrossRef CAS PubMed.
- H. Wen, G.-L. Hou, S. M. Kathmann, M. Valiev and X.-B. Wang, J. Chem. Phys., 2013, 138(3), 31101 CrossRef PubMed.
- S. Saha, S. Roy, P. Mathi and J. A. Mondal, J. Phys. Chem. A, 2019, 123(13), 2924 CrossRef CAS PubMed.
- J.-C. Jiang, Y.-S. Wang, H.-C. Chang, S. H. Lin, Y. T. Lee, G. Niedner-Schatteburg and H.-C. Chang, J. Am. Chem. Soc., 2000, 122(7), 1398 CrossRef CAS.
- H. Li, X. Kong, L. Jiang and Z.-F. Liu, Phys. Chem. Chem. Phys., 2018, 20(42), 26918 RSC.
- H. Knorke, H. Li, J. Warneke, Z.-F. Liu and K. R. Asmis, Phys. Chem. Chem. Phys., 2020, 22(47), 27732 RSC.
- S. Chakrabarty and E. R. Williams, Phys. Chem. Chem. Phys., 2016, 18(36), 25483 RSC.
- M. Rozenberg, A. Loewenschuss and Y. Marcus, Phys. Chem. Chem. Phys., 2000, 2(12), 2699 RSC.
- M. A. Boyer, O. Marsalek, J. P. Heindel, T. E. Markland, A. B. McCoy and S. S. Xantheas, J. Phys. Chem. Lett., 2019, 10(5), 918 CrossRef CAS PubMed.
- W. H. Robertson and M. A. Johnson, Annu. Rev. Phys. Chem., 2003, 54, 173 CrossRef CAS PubMed.
- G. D. Dickenson, M. L. Niu, E. J. Salumbides, J. Komasa, K. S. E. Eikema, K. Pachucki and W. Ubachs, Phys. Rev. Lett., 2013, 110(19), 193601 CrossRef CAS PubMed.
- J. T. Kelly, M. Mayer, A. C. Kennedy, C. Schemel and K. R. Asmis, J. Chem. Phys., 2018, 148(22), 222840 CrossRef PubMed.
- T. Hamashima, K. Mizuse and A. Fujii, J. Phys. Chem. A, 2011, 115(5), 620 CrossRef CAS PubMed.
- N. Heine, T. I. Yacovitch, F. Schubert, C. Brieger, D. M. Neumark and K. R. Asmis, J. Phys. Chem. A, 2014, 118(35), 7613 CrossRef CAS PubMed.
- N. Heine and K. R. Asmis, Int. Rev. Phys. Chem., 2015, 34(1), 1 Search PubMed.
- M. Brümmer, C. Kaposta, G. Santambrogio and K. R. Asmis, J. Chem. Phys., 2003, 119(24), 12700 CrossRef.
- N. Heine, M. R. Fagiani, M. Rossi, T. Wende, G. Berden, V. Blum and K. R. Asmis, J. Am. Chem. Soc., 2013, 135(22), 8266 CrossRef CAS PubMed.
- B. M. Elliott, R. A. Relph, J. R. Roscioli, J. C. Bopp, G. H. Gardenier, T. L. Guasco and M. A. Johnson, J. Chem. Phys., 2008, 129(9), 94303 CrossRef PubMed.
- W. R. Bosenberg and D. R. Guyer, J. Opt. Soc. Am. B, 1993, 10(9), 1716 CrossRef CAS.
- N. Heine and K. R. Asmis, Int. Rev. Phys. Chem., 2016, 35(3), 507 Search PubMed.
- M. J. Frisch, M. Head-Gordon and J. A. Pople, Chem. Phys. Lett., 1990, 166(3), 275 CrossRef CAS.
- F. Weigend, F. Furche and R. Ahlrichs, J. Chem. Phys., 2003, 119(24), 12753 CrossRef CAS.
- A. D. Becke, J. Chem. Phys., 1993, 98(7), 5648 CrossRef CAS.
- A. Chakraborty, T. Brumme, S. Schmahl, H. Weiske, C. Baldauf and K. R. Asmis, Chem. Sci., 2022, 13(44), 13187 RSC.
- A. Chakraborty, S. Schmahl and K. R. Asmis, Chem. Phys. Chem., 2021, 22(11), 1036 CrossRef CAS PubMed.
- R. D. Johnson, NIST Computational Chemistry Comparison and Benchmark Database. NIST Standard Reference Database Number 101, Release 22, May 2022 DOI:10.18434/T47C7Z.
- J. Bloino and V. Barone, J. Chem. Phys., 2012, 136(12), 124108 CrossRef PubMed.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, G. A. Petersson, H. Nakatsuji, X. Li, M. Caricato, A. V. Marenich, J. Bloino, B. G. Janesko, R. Gomperts, B. Mennucci, H. P. Hratchian, J. V. Ortiz, A. F. Izmaylov, J. L. Sonnenberg, D. Williams-Young, F. Ding, F. Lipparini, F. Egidi, J. Goings, B. Peng, A. Petrone, T. Henderson, D. Ranasinghe, V. G. Zakrzewski, J. Gao, N. Rega, G. Zheng, W. Liang, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, K. Throssell, J. A. Montgomery, Jr., J. E. Peralta, F. Ogliaro, M. J. Bearpark, J. J. Heyd, E. N. Brothers, K. N. Kudin, V. N. Staroverov, T. A. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. P. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, J. M. Millam, M. Klene, C. Adamo, R. Cammi, J. W. Ochterski, R. L. Martin, K. Morokuma, O. Farkas, J. B. Foresman and D. J. Fox, 2016.
- J. VandeVondele, M. Krack, F. Mohamed, M. Parrinello, T. Chassaing and J. Hutter, Comput. Phys. Commun., 2005, 167(2), 103 CrossRef CAS.
- G. J. Martyna and M. E. Tuckerman, J. Chem. Phys., 1999, 110(6), 2810 CrossRef CAS.
- M. Krack, Theor. Chem. Acc., 2005, 114(1–3), 145 Search PubMed.
- A. D. Becke, Phys. Rev. A, 1988, 38(6), 3098 CrossRef CAS PubMed.
- S. Grimme, S. Ehrlich and L. Goerigk, J. Comput. Chem., 2011, 32(7), 1456 CrossRef CAS PubMed.
- R. Resta, Rev. Mod. Phys., 1994, 66(3), 899 CrossRef CAS.
| This journal is © the Owner Societies 2026 |