Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Δ-Machine learning toward CCSD accuracy for homohalogenated borane–phosphine adducts: screening low-energy structures from DFT and MP2 libraries
Okan Köksal
Center for Molecular Modeling, Ghent University, Technologiepark 903, BE-9052, Zwijnaarde, Belgium. E-mail: okan.koksal@Ugent.be
First published on 29th April 2026
Abstract
Accurate formation energies for weak boron–phosphorus Lewis adducts are challenging because low-order correlation and density-functional methods can misrank low-energy motifs on shallow potential-energy landscapes and are sensitive to basis-set superposition error (BSSE). Herein, we combine large-scale structure-library sampling with Δ-machine learning (Δ-ML) to approach coupled-cluster accuracy for the homohalogenated adducts F3B–PF3, Cl3B–PCl3, and Br3B–PBr3. DFT (B3LYP-D3) and MP2 libraries comprising several thousand geometries per system are generated and used to train CCSD-referenced Δ-ML models that predict ECCSD from low-level inputs. The resulting models reproduce CCSD energies with low errors and enable efficient screening of the full libraries, after which a compact low-energy subset is refined with targeted CCSD(T) calculations. Counterpoise-corrected results show that MP2 substantially overbinds, especially for the chlorinated and brominated adducts. At the highest level, Cl3B–PCl3 is found to lie at the threshold of binding, whereas Br3B–PBr3 remains clearly bound and F3B–PF3 is weakly bound. Distance-resolved scans and Morokuma-type energy decomposition analyses rationalize the distinct binding regimes across F/Cl/Br in terms of the balance between Pauli repulsion, polarization/exchange, and dispersion. The proposed workflow enables reliable coupled-cluster-level screening of weak donor–acceptor adducts at greatly reduced cost.
I. Introduction
Lewis acid–base adducts featuring B–P donor–acceptor bonds remain central to main-group chemistry, with relevance spanning classical synthesis and reactive intermediates to applications in molecular materials and hydrogen-storage concepts.1–4 Although the textbook picture of a phosphine → borane donation into the vacant boron p orbital is conceptually straightforward, the nature of the B–P interaction can vary continuously from weak, predominantly electrostatic association to substantially covalent bonding. This variability depends sensitively on fragment electronic structure, substituent effects, and the deformation required to bring the Lewis acid into a binding-ready (pyramidalized) geometry.5–7 In haloborane adducts, the stability of the B–P contact reflects a delicate interplay between donor → acceptor charge transfer, the pyramidalization penalty of trigonal-planar BX3, and halogen-dependent polarization, back-donation, and dispersion effects.6,7A recurring theme in theoretical work on B–P donor–acceptor complexes is that computed geometries and formation energies can be strongly method dependent, especially for shallow potentials where multiple low-lying minima or near-degenerate motifs may exist.8–19 This motivates a PES-focused perspective and benchmarking against correlated wave-function methods rather than reliance on a single optimized structure. A striking illustration of the underlying complexity was highlighted by Almas and Pearson,20 who demonstrated that the B–P bond-stretching potential energy surface (PES) in phosphine haloboranes can deviate strongly from a conventional Morse-like form and may exhibit anomalous inflection points. In the extreme case of Cl3B–PH3, the PES even shows two distinct minima.20 Such findings emphasize that reliable ranking and mechanistic interpretation of weak-to-intermediate adducts require accurate characterization of the local PES landscape in regions where shallow basins, shoulders, or competing minima may occur. In this context, high-quality experimental benchmarks for closely related B–P adducts remain valuable reference points for assessing computed structures and trends.21 Recently, we presented a systematic ab initio analysis of donor–acceptor bonding in a broad set of X3B⋯PY3 and X3B⋯AsY3 complexes (X = H, F, Cl, Br; Y = CH3, F, Cl, Br), combining MP2 energies with coupled-cluster benchmarking and complementary bonding analyses (EDA, RDG, NBO).22 That work established clear trends from predominantly electrostatic interactions in weakly bound adducts to strongly bound, more covalent systems, and underscored the stabilizing role of halogen back-donation and charge polarization in BCl3 and BBr3 adducts relative to BF3 complexes.22 Importantly, the three symmetric halogenated phosphane–haloborane pairs F3B–PF3, Cl3B–PCl3, and Br3B–PBr3 were already included in that dataset and served as representative examples of weak-to-intermediate B–P triel bonding motifs, and all three were verified as genuine minima at the MP2/aug-cc-pVTZ level,23 exhibiting no imaginary frequencies in harmonic vibrational analyses.22
In the present contribution, we revisit precisely these three prototypical systems: F3B–PF3, Cl3B–PCl3, and Br3B–PBr3, but with a different emphasis and a stricter accuracy target. Rather than focusing on bonding decomposition at a single minimum, we (i) map the local structural landscape in a statistically meaningful manner, (ii) assess how method choice (DFT vs. MP2) affects the identification and ranking of low-energy structures on a potentially shallow, non-Morse-like PES,20 and (iii) connect these structural ensembles to high-accuracy reference energetics at the CCSD(T) level.24,25 To enable this, we generated extensive structure libraries for each complex and evaluated them at both levels of theory, comprising 5497 (MP2) and 5493 (DFT) geometries for F3B–PF3, 5714 (MP2) and 5713 (DFT) geometries for Cl3B–PCl3, and 5557 (MP2) and 5556 (DFT) geometries for Br3B–PBr3. This dense sampling allows identification of competing minima and near-degenerate motifs, quantification of structural variability close to the global minimum, and a controlled comparison of DFT and MP2 predictions for weak B–P donor–acceptor contacts. To connect the observed structural diversity with a physically transparent picture of the B–P interaction, we further analyze selected B–P separations using energy decomposition concepts.26–29
More broadly, machine learned representations of potential-energy surfaces have advanced substantially over the past decade, with widely used approaches including permutationally invariant polynomials, neural networks, and Gaussian-process-based models.30–34 In the present work, however, we do not attempt a full global PES fit. Instead, we employ a descriptor-based Δ-learning strategy for local low-energy screening, using handcrafted geometric/energetic descriptors and cross-checking the conclusions with rotationally and translationally invariant SOAP35 (smooth overlap of atomic positions) representations (see the SI). Related orbital-based machine-learning strategies have also been developed, most notably molecular-orbital-based machine learning (MOB-ML), in which post-Hartree–Fock correlation energies are learned from Hartree–Fock molecular-orbital information rather than primarily from geometric descriptors.36,37 Such approaches aim at improved transferability across chemical systems by exploiting compact orbital-based representations. Extending such strategies to larger molecular systems while retaining high-level electronic-structure accuracy remains challenging. For shallow donor–acceptor potential-energy surfaces, reliable energetic ranking requires an accurate treatment of electron correlation. In this context, coupled-cluster methods provide particularly important reference data, and CCSD(T) is widely regarded as a benchmark or “gold-standard” approach for molecular energetics. However, the steep computational scaling of conventional CC methods, culminating in the formal O(N7) scaling of CCSD(T), makes direct application to large structure libraries impractical.38–40 Indeed, even for systems of only moderate size, high-level PES construction can require very large coupled-cluster datasets, e.g., Bowman and co-workers reported a 10-atom formic-acid-dimer PES based on 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 475 CCSD(T)-F12/triple-ζ energy points.41
475 CCSD(T)-F12/triple-ζ energy points.41
A practical strategy to overcome this bottleneck is Δ-machine learning, in which a model is trained to learn the difference between a low-cost baseline and a higher-level reference, thereby correcting large low-level datasets toward coupled-cluster accuracy.42 In the context of potential-energy surfaces, this idea is commonly expressed as a correction to a low-level PES, VLL→CC = VLL + ΔVCC–LL, where the smoother correction term can often be learned from a comparatively small number of high-level data points.43 Such Δ-learning strategies have been successfully used to accelerate high-level energetics and PES construction for systems ranging from small molecules to medium-sized species.44,45 Motivated by these developments and by the non-trivial PES features reported for phosphine haloboranes,20 we employ a CCSD-trained Δ-learning strategy to screen and rank low-energy regions of the PES and subsequently refine a targeted subset of ML-selected structures at the CCSD(T) level, thereby approaching CCSD(T)-quality energetics without exhaustive CCSD(T) sampling.
The paper is organized as follows. Section II details the electronic-structure methods, the structure-library generation protocol, and the Δ-learning workflow used to approach CCSD accuracy and to guide the subsequent CCSD(T) refinement of selected low-energy candidates. Section III compares DFT and MP2 structural ensembles, identifies the dominant low-energy motifs, analyzes how high-level corrections reshape the energetic ranking, and uses relaxed scans and energy decomposition analysis to rationalize the resulting bonding trends. Finally, Section IV summarizes the principal conclusions and outlines practical guidance for computational studies of shallow donor–acceptor PESs in halogenated phosphane–haloborane complexes.
II. Methods
A. Structure-library generation
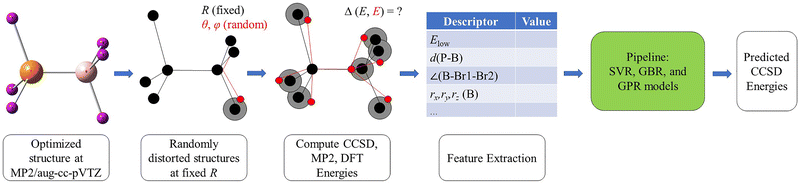
Starting from each equilibrium geometry, we generated randomly distorted structures by applying small, isotropic atomic displacements within a sphere of radius R centered at the original atomic position. For each atom, a displacement length r was sampled uniformly in volume by drawing r′ ∈ [0,1] and setting:
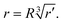 | (1) |
Two angles ϕ ∈ [0,2π) and θ ∈ [0,π] were generated to define the displacement direction in spherical coordinates. The corresponding Cartesian components (dx, dy, dz) were added to the original atomic coordinates to yield the perturbed geometry. Because these displacements were applied directly to the Cartesian coordinates of the atoms, not only the intermolecular B⋯P separation but also the intrafragment B–X and P–X bond lengths and associated angles could vary in the distorted structures. No subsequent geometry relaxation was performed for these library geometries and they were evaluated directly by single-point calculations. In the present work we employed R = 0.001 Å to generate near-equilibrium distortions and R = 0.09 Å to sample more strongly distorted configurations. Consequently, a larger number of structures was generated at larger R to ensure sufficient coverage of the higher-variance regions of the configuration space. As shown in Fig. 1, randomly distorted donor–acceptor geometries were generated by applying small stochastic Cartesian displacements to the atomic coordinates of the reference structure, where the displacement direction for each atom was sampled in spherical coordinates through the angles (θ,ϕ) and the displacement length was sampled within a sphere of radius R. The resulting structures therefore sample variations not only in the intermolecular B⋯P separation, but also in the associated intra- and intermolecular geometric descriptors. Subsequently, we train regression models to learn the correction ΔE = ECCSD − Elow. The trained model is then used to predict CCSD-quality energies for F3B–PF3, Cl3B–PCl3, and Br3B–PBr3 from MP2/DFT inputs across the sampled configurational space.
In this work we focus on the homohalogenated donor–acceptor adducts F3B–PF3, Cl3B–PCl3, and Br3B–PBr3 and combine large-scale structure sampling with correlated reference data and energy-decomposition analysis. All electronic-structure calculations were performed with GAMESS.46 Equilibrium geometries were optimized at the MP2/aug-cc-pVTZ level and, independently, at the DFT (B3LYP-D3)/aug-cc-pVTZ level. The MP2 structure libraries were generated by applying random spherical displacements to the MP2-optimized geometries and evaluating MP2 single-point energies for each sampled structure. The same distorted geometries were then re-evaluated at the DFT (B3LYP-D3)/aug-cc-pVTZ level to construct the corresponding DFT energy libraries. Coupled-cluster reference data were obtained at the CCSD/aug-cc-pVTZ level for a labeled subset of structures and used to train Δ-learning models for the correction ΔE = ECCSD − Elow, where Elow denotes either MP247,48 or DFT (B3LYP-D3).49,50 To assess the impact of perturbative triples in the low-energy region without exhaustive CCSD(T) sampling, targeted CCSD(T)/aug-cc-pVTZ single-point calculations were carried out for a compact ML-selected refinement set for each complex (including the baseline-optimized minimum and the lowest-energy CCSD training-set structure). Hence, these CCSD(T) calculations are best viewed as targeted single-point refinements of an ML-selected low-energy subset, not as an ML-driven search on a full CCSD(T) potential-energy surface. Unless noted otherwise, correlated calculations employed the frozen-core approximation, which reduces computational cost while retaining the dominant valence correlation contribution to bonding.51 Basis-set superposition error (BSSE) was estimated using the Boys–Bernardi counterpoise (CP) procedure.52
III. Results and discussion
A. Geometries and formation energies
Table 1 compares formation energies for the three complexes at MP2, DFT/B3LYP-D3, CCSD (Δ-ML), and CCSD(T) (subset) levels. In triel-bonded systems, the X–B–P angle serves as a useful structural measure of the extent of pyramidalization at the Lewis-acid center, where stronger donor–acceptor interactions are commonly associated with a larger departure from trigonal-planar geometry and α values approaching 90° for the boron fragment.53,54 Furthermore, for F3B–PF3 we report a CCSDopt geometry obtained with numerical gradients in GAMESS, yielding Rab = 3.26410 Å and averaged angles 〈α〉 = 90.641° and 〈β〉 = 119.427° (cf. Table 1). Across the halogen series, MP2 predicts a strongly increasing stabilization (F: −2.05, Cl: −9.30, Br: −16.54 kcal mol−1), whereas CCSD(T) (subset) yields a much more moderate trend (F: −2.19, Cl: −3.82, Br: −9.25 kcal mol−1, cf. Table 1). This indicates that MP2 systematically produces more negative ΔEform values than CCSD(T), with the discrepancy growing substantially from F to Br. In contrast, DFT/B3LYP-D3 gives only slightly negative formation energies for F (−1.57 kcal mol−1) and Br (−3.41 kcal mol−1), but predicts a positive formation energy for the chlorine adduct (+1.71 kcal mol−1, see Table 1). In addition, BSSE corrections reduce the magnitude of the MP2 stabilization across the series (F: −2.05 → −1.17, Cl: −9.30 → −5.42, Br: −16.54 → −11.90 kcal mol−1), which is consistent with increased BSSE for the heavier, more polarizable complexes (cf. Table 1). For DFT/B3LYP-D3, CP corrections are comparatively small for F and Br (F: −1.57 → −1.31, Br: −3.41 → −2.87 kcal mol−1), but further increase the positive formation energy for Cl3B–PCl3 (+1.71 → +2.20 kcal mol−1). At the CCSD (Δ-ML) level, the BSSE correction is decisive for the chlorine adduct: the uncorrected formation energy remains slightly negative (−1.52 kcal mol−1), whereas the CP-corrected value becomes positive (+1.96 kcal mol−1). At the CCSD(T) level, the corresponding CP-corrected formation energy is essentially zero (−0.03 kcal mol−1). Thus, Cl3B–PCl3 emerges as a borderline case whose energetic classification depends sensitively on the level of electron correlation, geometry relaxation, and BSSE correction. This conclusion is sharpened by the ML-assisted optimization, which enabled access to the relevant higher-level minimum and thereby revealed the near-threshold character of the chlorinated adduct much more clearly than would be possible from lower-level structures alone. By contrast, the brominated adduct remains clearly bound even after CP correction at both CCSD and CCSD(T), although the correction substantially reduces the magnitude of the stabilization. At the coupled-cluster level, CP-corrected values could not be obtained for F3B–PF3. The plain supermolecular CCSD and CCSD(T) single-point calculations for this complex converged successfully and displayed a well-behaved single-reference diagnostic, whereas the corresponding BSSE-style coupled-cluster calculations did not converge and yielded a much larger T1 diagnostic. This indicates that the fragment-in-full-basis reference underlying the counterpoise-like decomposition constitutes a more difficult electronic-structure problem than the ordinary supermolecular calculation. For Cl3B–PCl3, the B3LYP-D3 treatment appears to overestimate the destabilization of the adduct, since it predicts a clearly positive formation energy, whereas the correlated CCSD and CCSD(T) results indicate a borderline case with formation energies close to zero once BSSE effects are taken into account. CCSD (Δ-ML) yields negative uncorrected formation energies for all systems and thus recovers the same sign pattern as CCSD(T) before CP correction. Quantitatively, CCSD (Δ-ML) is systematically less negative than CCSD(T), with deviations of ΔE = +0.31 (F), +2.30 (Cl), and +2.75 (Br) kcal mol−1. The increasing gap from F to Br suggests that correlation contributions beyond CCSD become progressively more important for the heavier, more polarizable halogen complexes. For F3B–PF3, the explicitly optimized CCSDopt formation energy (−1.91 kcal mol−1) is close to the CCSD (Δ-ML) value (−1.88 kcal mol−1) and less negative than CCSD(T) (subset) (−2.19 kcal mol−1), indicating that perturbative triples provide additional stabilization even for the weakly bound fluorinated adduct. Importantly, the B–P separations predicted by MP2 and CCSD (Δ-ML) are nearly identical for Cl and Br (Cl: 2.01380 vs. 2.01383 Å, Br: 1.99482 vs. 1.99514 Å, cf. Table 1), indicating that the large differences in ΔEform are not primarily driven by the optimized geometry but instead by the underlying electronic structure treatment. Taken together, Table 1 shows that CCSD (Δ-ML) substantially improves upon MP2 and B3LYP-D3 for these adducts, while the residual shift toward less negative ΔEform relative to CCSD(T) highlights the growing role of perturbative triples (and/or higher-order) effects for Cl and Br. Computational cost and motivation for ML-guided sampling: all CCSD single-point calculations were carried out in parallel on 8 CPU cores (Intel Xeon E5-2683 v4 @ 2.10 GHz). A representative CCSD/aug-cc-pVTZ single-point evaluation required on the order of ∼10 h wall time for the F3B–PF3 complex, which is the smallest system considered here (8 atoms, 74 electrons), with 115 basis-set shells and 450 Cartesian Gaussian basis functions, corresponding to 377 molecular orbitals (37 occupied) in the CCSD calculation (with 12 frozen-core orbitals and no frozen virtual orbitals). The heavier Cl3B–PCl3 and Br3B–PBr3 systems contain 122 and 240 electrons, respectively, and are therefore computationally more demanding.| Complex | Level | Rab (Å) | 〈α〉 (deg) | 〈β〉 (deg) | ΔEform | ΔECPform |
|---|---|---|---|---|---|---|
| a CP-corrected CCSD and CCSD(T) formation energies could not be obtained for this complex because the corresponding BSSE-style coupled-cluster treatment did not permit a reliable determination of the CP-corrected energy. | ||||||
| F3B–PF3 | MP2 | 3.23715 | 90.733 | 119.279 | −2.05 | −1.17 |
| DFT/B3LYP-D3 | 3.39430 | 90.394 | 119.192 | −1.57 | −1.31 | |
| CCSDopt | 3.26410 | 90.641 | 119.427 | −1.91 | —a | |
| CCSD (Δ-ML) | 3.24436 | 90.718 | 119.283 | −1.88 | —a | |
| CCSD(T) (subset) | 3.23715 | 90.733 | 119.279 | −2.19 | —a | |
| Cl3B–PCl3 | MP2 | 2.01380 | 102.880 | 113.441 | −9.30 | −5.42 |
| DFT/B3LYP-D3 | 2.10874 | 102.351 | 113.617 | +1.71 | +2.20 | |
| CCSD (Δ-ML) | 2.01383 | 102.874 | 113.450 | −1.52 | +1.96 | |
| CCSD(T) (subset) | 2.01381 | 102.873 | 113.450 | −3.82 | −0.03 | |
| Br3B–PBr3 | MP2 | 1.99482 | 103.204 | 112.998 | −16.54 | −11.90 |
| DFT/B3LYP-D3 | 2.07699 | 103.123 | 113.015 | −3.41 | −2.87 | |
| CCSD (Δ-ML) | 1.99514 | 103.213 | 113.001 | −6.50 | −2.31 | |
| CCSD(T) (subset) | 1.99514 | 103.213 | 113.001 | −9.25 | −4.63 | |
In total, approximately 1000 CCSD single-point calculations were required in this study, comprising the labeled subsets used for training and for held-out evaluation together with additional low-energy candidates identified by the ML screening and subsequently verified at the CCSD level (see Section III C), corresponding to ∼10.000 h (∼14 months) of serial wall time, or ∼80.000 core-hours. These CCSD calculations comprised both the labeled subset used for Δ-ML training/validation/testing and an additional set of ML-selected low-energy candidates that were recomputed explicitly at the CCSD level after screening of the full low-level library. In practice, these single-point calculations can be executed as large job arrays on HPC resources (e.g., tens of jobs concurrently), substantially reducing the elapsed wall time. In contrast, a fully relaxed CCSD geometry optimization of F3B–PF3 using numerical gradients (NUMGRD=.T. in GAMESS) required nearly three months on 8 CPU cores. Such an optimization is inherently iterative and far less amenable to the same degree of batching because successive optimization steps depend on the preceding geometry. This stark difference in cost between CCSD single-point evaluations and CCSD geometry optimizations underscores the practical importance of the ML-guided workflow, which concentrates high-level calculations on the most informative regions of configuration space while avoiding prohibitively expensive CCSD optimizations for large structure sets.
B. Structure libraries and CCSD-referenced low-energy ordering
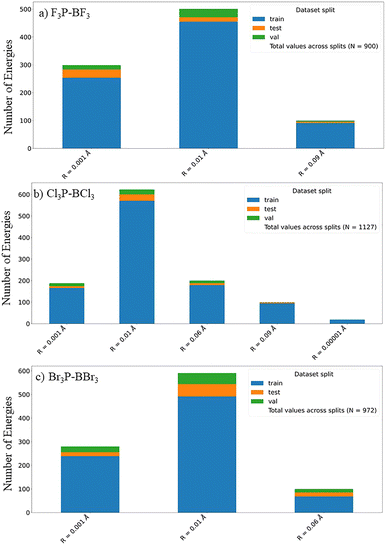
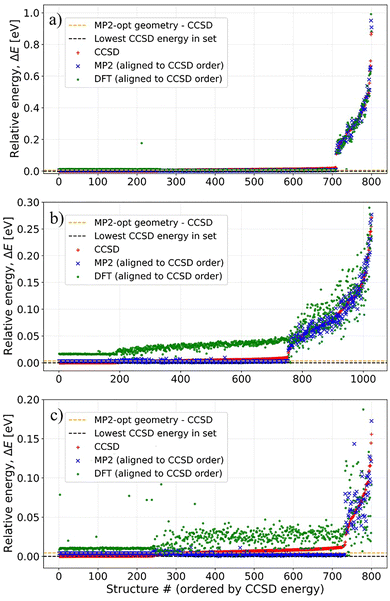
Fig. 2 shows the data for F3B–PF3, Cl3B–PCl3, and Br3B–PBr3 organized into subsets corresponding to different values of the displacement–radius parameter R, which controls the magnitude of the random atomic perturbations applied during structure generation. Each subset is further partitioned into training, test, and validation splits. The stacked-bar representation highlights how the sampling density varies with R for each complex while preserving a consistent split assignment within each R-group. Although CCSD single-point energies at the equilibrium MP2 geometries are included in the training/validation/test datasets, they are intentionally omitted from the plotted categories and only contribute to the reported totals.Fig. 3 contrasts the CCSD relative energies with the associated MP2 and DFT relative energies for the matched CCSD-labeled subset after sorting all structures by increasing CCSD energy. To improve interpretability, the energies are shown relative to the minimum and expressed in eV rather than in absolute atomic units. Plotting MP2 and DFT in the CCSD-imposed order highlights how well these lower-cost methods preserve the CCSD energetic ranking within this evaluated subset, and it also exposes the high-energy tail associated with more strongly distorted configurations. The CCSD relative energy at the MP2-optimized geometry (orange dashed line) and the lowest CCSD energy included in the training set (black dashed line) define reference points for the equilibrium region sampled by the dataset. Complementary histogram analyses of the relative-energy distributions and of the corresponding relative-energy differences for the matched CCSD-labeled subsets are provided in the SI.
C. Δ-ML toward CCSD: descriptors, validation, and screening
To clarify the input representations used in the different machine-learning workflows, Table 2 summarizes the descriptor sets and preprocessing steps for the models considered in this work. For the descriptor-based workflows, the same handcrafted feature set was used across support vector regression (SVR), gradient boosting regression (GBR), and Gaussian process regression (GPR), so that the difference between these models lies in the regression algorithm rather than in the descriptor definition itself. In addition, a SOAP-based representation combined with principal component analysis (PCA) and SVR was evaluated as an alternative structural descriptor in the SI.| Workflow | Input representation | Preprocessing | Role in this work |
|---|---|---|---|
| Descriptor + SVR | Handcrafted geometric descriptors + Elow | Standardization | Final production model |
| Descriptor + GBR | Same as above | Standardization | Benchmarked alternative |
| Descriptor + GPR | Same as above | Standardization | Benchmarked alternative |
| SOAP + SVR | SOAP descriptor + Elow | Standardization + PCA | Robustness check (SI) |
The handcrafted descriptor vector comprised selected Cartesian coordinate components, key intermolecular and intramolecular distances, bond angles, and the corresponding low-level reference energy Elow. Moreover, representative descriptor examples and SOAP hyperparameters are reported (cf. SI).
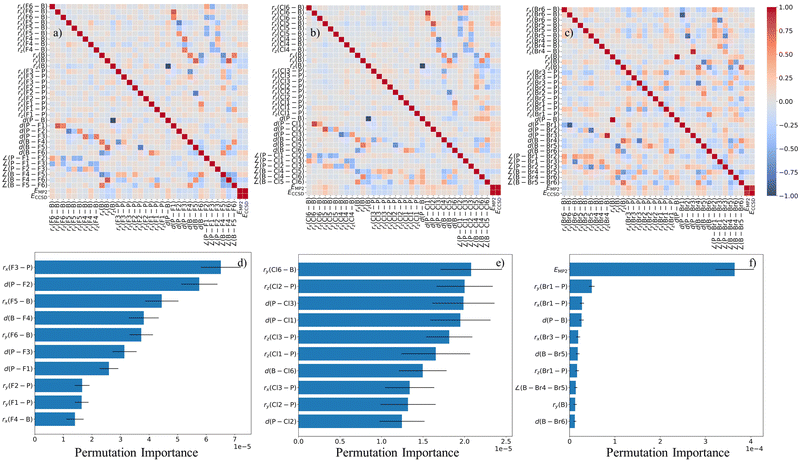
The Δ-ML models were implemented within a pipeline that included SVR, GBR, and GPR. Hyperparameters were optimized with Optuna using 5-fold cross-validation on the training set. For each trial, the model was trained to predict the energy correction ΔE = ECCSD − Elow on four folds and evaluated on the remaining fold, whereas the mean absolute error (MAE) averaged over the five validation folds was used as the optimization objective. Among the tested regressors, SVR yielded the most robust balance between cross-validated MAE and low-energy screening selectivity, and was therefore adopted for the final workflow. Fig. 4 summarizes the descriptor relationships and their predictive relevance for the three donor–acceptor complexes. The correlation heatmaps in panels (a)–(c) quantify linear dependencies among Cartesian coordinate components, selected interatomic distances and angles, and the low-level energy EMP2, together with the CCSD energies available for the labeled subset. Complementarily, the permutation-importance analysis in panels (d)–(f), obtained by shuffling individual features in a trained SVR Δ-learning model, identifies the variables that most strongly affect prediction of ΔE = ECCSD − EMP2 (and thus ECCSD). Notably, EMP2 plays a dominant, non-redundant role for Br3B–PBr3, in agreement with the stronger statistical dependence between ΔE and EMP2 in the brominated system, whereas for F3B–PF3 and Cl3B–PCl3 the correction can be captured largely by geometric descriptors with only minor added benefit from including EMP2 explicitly as an input feature. This behavior can be understood in light of the energetic hierarchy discussed above: F3B–PF3 is weakly bound and Cl3B–PCl3 lies near the threshold of binding once higher-level correlation and BSSE effects are taken into account, whereas Br3B–PBr3 remains clearly bound. Accordingly, geometric descriptors are largely sufficient for the fluorinated and chlorinated systems, while the low-level energy EMP2 becomes more important for the more robustly bound brominated adduct. This system dependence further suggests that orbital-level descriptors, as used in MOB-ML-type frameworks, may provide an interesting alternative route toward a more uniform representation across the halogen series.36,37
In addition to the correlation and permutation-importance analyses shown in Fig. 4, we evaluated the role of including EMP2 as an input feature in the Δ-learning models. The corresponding permutation-importance results (with and without EMP2 appended to the geometric descriptor vector) are reported in the SI. In line with the system-dependent behavior seen in Fig. 4d–f), removing EMP2 from the input features leads to a pronounced degradation in predictive performance for Br3B–PBr3 (e.g., R2 = 0.72 and MAE = 3.09 × 10−4 Eh for the MP2-based SVR model without EMP2, see SI), whereas F3B–PF3 and Cl3B–PCl3 remain largely unaffected (e.g., R2 = 0.997 and MAE = 1.71 × 10−4 Eh for F3B–PF3, and R2 = 0.989 and MAE = 1.05 × 10−4 Eh for Cl3B–PCl3, not shown here). This behavior is in agreement with the stronger dependence of the correction ΔE = ECCSD − EMP2 on EMP2 for the brominated complex, as expressed by the larger magnitude of ρ(ΔE, EMP2) compared with the fluorinated and chlorinated systems. As an additional robustness check on the descriptor representation, we also evaluated an alternative machine learning pipeline based on SOAP descriptors generated with the DScribe package,35 followed by PCA and SVR regression. The resulting SOAP + PCA models and their predictive performance are described in the SI. Importantly, the SOAP-based workflow identifies the same number of low-energy candidate structures for all three complexes as the handcrafted-descriptor models discussed here, supporting the robustness of the ML-guided screening conclusions with respect to the descriptor choice.
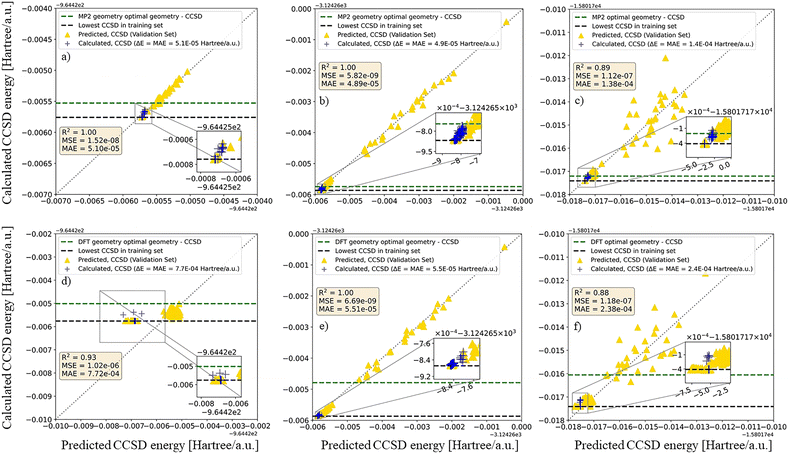
Fig. 5 assesses the accuracy of the Δ-ML models by comparing predicted and calculated CCSD energies on the held-out non-training CCSD-labeled subsets. For F3B–PF3 and Br3B–PBr3, 800 CCSD-labeled structures were used for training, whereas for Cl3B–PCl3 1027 CCSD-labeled structures were used (cf. Fig. 2). The remaining labeled structures were kept outside the training process and partitioned into validation and test subsets during model development. The parity plots are shown for the combined held-out subset in order to provide a compact visual summary of predictive agreement beyond the training data. The MP2-based models (cf. Fig. 5a–c) and DFT-based models (see Fig. 5d–f) are shown separately to highlight the effect of the baseline method on the learned correction. The clustering of points along the parity line demonstrates that CCSD-quality energies are recovered with small errors, particularly in the near-equilibrium region highlighted in the insets. The achieved errors on the held-out non-training subsets are thus appropriate for low-energy screening and energetic ranking, but they are not intended to match the microhartree-level accuracy often required for dynamics- or spectroscopy-grade global PES applications.31 The green and black reference lines mark additional benchmarks corresponding to the optimized equilibrium geometry and the lowest-energy structure included in the training set, respectively. Building on this validated accuracy, we next apply the CCSD-trained models to screen the full structure libraries and to define targeted CCSD(T) refinement sets.
Using the MP2 baseline, this screening step selected 111/156/122 low-energy structures for F3B–PF3, Cl3B–PCl3, and Br3B–PBr3, respectively, with comparable candidate counts obtained from the DFT-baseline models. The screening workflow involves several nested subsets. First, only a labeled subset of the full several-thousand-structure DFT/MP2 libraries was evaluated at the CCSD level for training, validation, and testing of the Δ-ML model. The trained model was then applied to the remaining structures in the low-level libraries to predict CCSD-quality energies and identify a low-energy basin. Candidate structures falling within an MAE-based energy window around the predicted low-energy minimum were subsequently recomputed explicitly at the CCSD level to verify the ML-guided screening, with the MAE obtained on the held-out non-training subset serving as a practical tolerance for candidate selection. Finally, a still smaller subset of the most relevant low-energy candidates was subjected to single-point CCSD(T) calculations. To assess the impact of perturbative triples on the energetic ordering in the low-energy region, we then carried out targeted CCSD(T) single-point calculations for this candidate subset, which includes (i) the baseline-optimized equilibrium structure and (ii) the lowest-energy CCSD structure present in the training data, together with additional ML-selected candidates. This yields CCSD(T) refinement sets of 113/158/124 structures for F3B–PF3, Cl3B–PCl3, and Br3B–PBr3 (MP2 baseline), enabling a focused validation of CCSD-level screening against CCSD(T) energetics without exhaustive CCSD(T) sampling. Accordingly, the present workflow is intended to identify and refine the relevant low-energy basin rather than to guarantee direct recovery of the exact CCSD(T) minimum from the learned model alone.
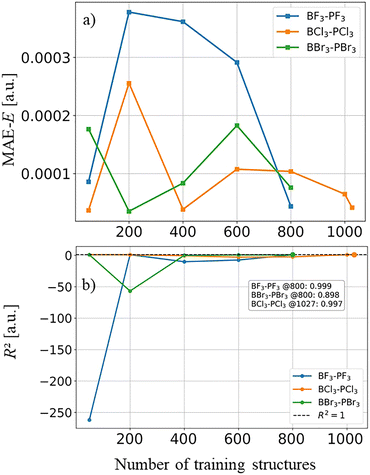
The predictive quality of the trained models as a function of training-set size is illustrated in the learning curves shown in Fig. 6. Panel (a) reports the mean absolute error of the predicted CCSD energies, while panel (b) shows the corresponding R2 values for the SVR Δ-ML models trained on increasing numbers of structures. Across all three donor–acceptor complexes, the MAE decreases rapidly with increasing training-set size and reaches values on the order of 10−4 a.u., indicating that only a few hundred CCSD reference points are sufficient to achieve near chemical accuracy relative to the underlying CCSD reference energies. As the training set grows, the R2 values approach unity, demonstrating that the models capture the overall energetic trends of the sampled configuration space with high fidelity. Among the three systems, F3B–PF3 exhibits the fastest convergence, reaching R2 = 0.999 already at 800 training structures. The Cl3B–PCl3 complex shows similarly strong predictive performance and achieves R2 = 0.997 when trained on 1027 structures. For the heavier Br3B–PBr3 system the convergence is somewhat slower, which can be attributed to its increased configurational flexibility and broader energy distribution of the sampled structures. Nevertheless, even in this case the model reaches R2 = 0.898 with 800 training structures. These learning curves demonstrate that the Δ-ML framework efficiently transfers CCSD accuracy across the configurational landscape and rationalize the high predictive quality observed in the parity plots displayed in Fig. 5.
D. Energy decomposition analysis
Although the relaxed B–P scans and the associated energy-decomposition analysis are not part of the Δ-ML training workflow, they provide a complementary physical interpretation of the low-energy PES region targeted by the ML-guided screening. In particular, this analysis serves two purposes in the present study. First, it allows us to assess whether the homohalogenated systems considered here display unusual bond-stretching behavior of the type previously reported for related phosphine haloboranes by Almas and Pearson.20 Second, it helps rationalize the system-dependent energetic trends and bonding regimes underlying the different screening outcomes for F3B–PF3, Cl3B–PCl3, and Br3B–PBr3.Energy decomposition analysis (EDA) was used to obtain a physically transparent partitioning of the interaction along selected B–P separations. In the spirit of classical partitioning schemes such as the Morokuma approach26 and the related DFT-based framework of Ziegler and Rauk,27 the binding between two fragments is separated into deformation and interaction contributions and the latter is further resolved into physically interpretable terms.28 Here we employ the EDA implementation by Su et al.29 in GAMESS,46 which decomposes the interaction energy as:
| ΔEint = ΔEele + ΔEex + ΔErep + ΔEpol + ΔEdisp. | (2) |
Such decompositions have proven particularly informative for trihaloboranes, where the stability of Lewis adducts reflects a competition between donor → acceptor charge transfer, fragment deformation (pyramidalization), and halogen-dependent polarization/back-donation effects.55 Accordingly, distance-resolved PES scans and EDA evaluations at selected B–P separations are used here to provide a chemically transparent interpretation of the bonding regimes and local PES features relevant to the screened low-energy region.
The Morokuma energy decomposition analysis (EDA) affords a chemically transparent interpretation of the donor–acceptor interaction by separating the total interaction energy into electrostatic (ΔE(ele)), exchange (ΔE(ex)), Pauli repulsion (ΔE(rep)), polarization (ΔE(pol)), and dispersion (ΔE(disp)) contributions (cf. Table 3). For all three complexes, the strongly compressed region (r ≈ 1.5 Å) is dominated by a steep repulsive wall, as reflected by the very large positive ΔE(rep) values (+718, +849, and +868 kcal mol−1 for F3B–PF3, Cl3B–PCl3, and Br3B–PBr3, respectively). This term originates from Pauli exclusion between like-spin electrons when fragment densities overlap strongly. Although ΔE(ele), ΔE(ex), and ΔE(pol) are highly stabilizing at such short separations, their sum cannot compensate the extreme Pauli repulsion, yielding net repulsive interaction energies at r = 1.5 Å in each system.
| Complex | r (Å) | ΔE(ele) | ΔE(ex) | ΔE(rep) | ΔE(pol) | ΔE(disp) | ΔE(int) |
|---|---|---|---|---|---|---|---|
| (1) F3B–PF3 | 1.5 | −160.37 | −304.73 | 718.30 | −191.15 | −8.87 | 53.18 |
| 2.0 | −68.12 | −113.84 | 241.86 | −67.46 | −4.55 | −12.10 | |
| 2.5 | −21.60 | −39.34 | 77.87 | −13.97 | −3.94 | −0.98 | |
| 3.0 | −6.25 | −11.54 | 21.60 | −2.53 | −2.30 | −1.03 | |
| 3.5 | −1.74 | −2.79 | 5.00 | −0.57 | −1.12 | −1.21 | |
| req | −3.41 | −6.00 | 10.98 | −1.21 | −1.65 | −1.29 | |
| (2) Cl3B–PCl3 | 1.5 | −183.43 | −375.04 | 848.83 | −231.41 | −25.00 | 33.96 |
| 2.0 | −82.30 | −160.46 | 326.24 | −94.32 | −16.99 | −27.83 | |
| 2.5 | −27.83 | −63.62 | 120.06 | −22.70 | −13.15 | −7.23 | |
| 3.0 | −8.86 | −23.65 | 41.94 | −4.14 | −8.25 | −2.96 | |
| 3.5 | −2.52 | −7.22 | 12.13 | −0.83 | −4.51 | −2.96 | |
| req | −80.07 | −156.48 | 317.37 | −91.44 | −16.89 | −27.50 | |
| (3) Br3B–PBr3 | 1.5 | −179.12 | −386.07 | 868.29 | −245.00 | −30.12 | 27.98 |
| 2.0 | −82.34 | −172.14 | 346.29 | −101.83 | −21.72 | −31.74 | |
| 2.5 | −29.18 | −71.32 | 132.97 | −26.37 | −17.24 | −11.15 | |
| 3.0 | −9.82 | −28.24 | 49.53 | −5.23 | −11.20 | −4.96 | |
| 3.5 | −2.89 | −9.23 | 15.32 | −1.11 | −6.40 | −4.30 | |
| req | −83.16 | −173.69 | 349.74 | −102.98 | −21.77 | −31.85 | |
Upon increasing the B⋯P separation toward chemically relevant distances, ΔE(rep) decreases rapidly and the balance between attractive and repulsive contributions becomes favorable. In the F3B–PF3 complex, the minimum occurs at a relatively long separation (req = 3.23715 Å), where the interaction is weak (ΔE(int) ≈ −1.29 kcal mol−1). At this distance, the orbital-interaction signatures are modest, with only small exchange and polarization contributions (ΔE(ex) ≈ −6.00 and ΔE(pol) ≈ −1.21 kcal mol−1), which points to limited donor–acceptor mixing at the minimum. Within the conceptual framework employed in our earlier work, such small ΔE(pol) values are characteristic of binding dominated by long-range electrostatics and dispersion, with comparatively weak covalent/dative contributions. Consequently, the F3B–PF3 adduct reaches its most favorable interaction energy in a regime where Pauli repulsion is strongly reduced (ΔE(rep) = +10.98 kcal mol−1) and strong short-range orbital effects remain suppressed.
In contrast, the chlorinated and brominated systems exhibit a qualitatively different energetic signature: their minima occur at short separations (req ≈ 2.0 Å), i.e., in a region of substantial density overlap. For Cl3B–PCl3, the equilibrium interaction energy is strongly stabilizing (ΔE(int) ≈ −27.50 kcal mol−1) and is accompanied by large exchange and polarization terms (ΔE(ex) ≈ −156.48 and ΔE(pol) ≈ −91.44 kcal mol−1). For Br3B–PBr3, the stabilization is even greater (ΔE(int) ≈ −31.85 kcal mol−1) with correspondingly stronger exchange and polarization contributions (ΔE(ex) ≈ −173.69 and ΔE(pol) ≈ −102.98 kcal mol−1). Following the interpretation used in our previous study, pronounced negative ΔE(pol) values indicate substantial reshaping of fragment orbitals upon complex formation, which is characteristic of charge transfer and donor–acceptor orbital mixing, while strongly negative ΔE(ex) values reflect significant overlap-driven stabilization in the bonded region. Together, these features point to a markedly increased covalent/dative component for the BCl3 and BBr3 adducts relative to BF3. A key observation is that the short equilibrium distances in the Cl and Br cases are achieved despite very large Pauli repulsion at the minima: ΔE(rep) = +317.37 kcal mol−1 for Cl3B–PCl3 and ΔE(rep) = +349.74 kcal mol−1 for Br3B–PBr3 as listed in Table 3. Rather than indicating an anomalous result, these values provide a diagnostic fingerprint of a short-range donor–acceptor bond: the system pays a substantial Pauli penalty in order to access geometries where the stabilizing orbital-interaction contributions (exchange and polarization) are even larger in magnitude. In other words, the net attraction at req for the Cl/Br adducts arises from a compensation mechanism in which strong overlap-driven stabilization outweighs strong Pauli repulsion. In contrast, F3B–PF3 avoids this overlap-dominated regime by adopting a larger equilibrium separation, simultaneously reducing ΔE(rep) and suppressing the magnitude of exchange and polarization stabilization. The dispersion contribution introduces an additional systematic stabilization across the halogen series. At equilibrium, ΔE(disp) increases in magnitude from BF3 (≈−1.65 kcal mol−1) to BCl3 (≈−16.89 kcal mol−1) and remains large for BBr3 (≈−21.77 kcal mol−1). This trend follows the increasing polarizability of the heavier halogenated fragments and helps explain the more negative ΔE(int) values for the chlorinated and brominated systems. While dispersion alone does not enforce the short equilibrium distances, it contributes significantly to the overall stabilization in the Cl/Br cases and amplifies the net effect of electrostatics and orbital interactions.
Finally, the distance dependence of the EDA components provides a unified bonding picture for the three adducts. At larger separations (e.g., r = 3.0–3.5 Å), exchange and polarization contributions diminish markedly, as orbital overlap decreases and the interaction becomes increasingly dominated by electrostatics and dispersion. The crucial distinction is that the F3B–PF3 complex reaches its minimum in this weak-interaction regime, whereas Cl3B–PCl3 and Br3B–PBr3 attain their minima at shorter distances where the interaction energy is governed by a balance between very large Pauli repulsion and even larger overlap- and polarization-driven stabilization. Consequently, the Morokuma EDA suggests that F3B–PF3 is best described as a weak donor–acceptor complex with largely electrostatic/dispersion character, whereas the chlorinated and brominated analogues exhibit considerably stronger donor–acceptor bonding with pronounced covalent contributions, as evidenced by their short equilibrium B–P separations and more negative ΔE(int) values.
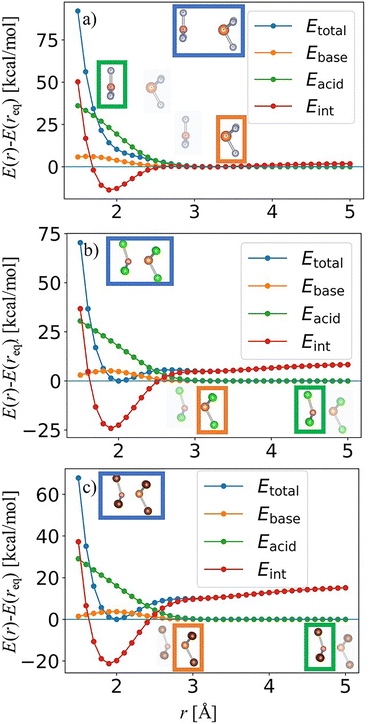
Fig. 7 depicts the relaxed B–P bond-stretching scans for the three homohalogenated trihaloborane–trihalophosphine complexes. For each system, the constrained scan was performed along the B–P distance r using relaxed geometries at each fixed separation, and the resulting energies were decomposed into the total energy Etotal, the fragment energies Eacid (BX3) and Ebase (PX3) evaluated at the complex geometry, and the interaction energy:
| Eint(r) = Etotal(r) − (Eacid(r) + Ebase(r)). | (3) |
All curves are shown as shifted energies, E(r) − E(req), in units of kcal mol−1, to facilitate a direct comparison of the local PES shapes across the three adducts. A key qualitative outcome of Fig. 7 is that none of the three systems exhibits the bimodal (two-minima) dissociation profile reported by Almas and Pearson for Cl3B–PCl3.20 Instead, each complex shows a single-well dissociation curve: following the equilibrium region, Etotal(r) − Etotal(req) increases smoothly toward the separated-fragment limit without developing a second outer minimum. In all three panels, the decomposition reveals the same underlying balance of physical effects. At very short separations, the steep repulsive wall in Etotal arises from (i) substantial fragment deformation, most prominently in the Lewis acid term Eacid (pyramidalization/distortion of BX3), together with (ii) strongly repulsive Eint (Pauli/steric repulsion at overly compressed contacts). Upon increasing r, the deformation penalty decreases rapidly, while Eint transitions from repulsive to attractive (negative), producing a single global minimum where the stabilization gained from interaction is no longer worth additional deformation of the fragments. Beyond the minimum, Eint weakens and the total energy rises monotonically, indicating that any long-range outer-well stabilization is insufficient to generate a second minimum for these homohalogenated pairs within the present scan protocol and level of theory.
The three systems share this common topology but differ in magnitude and range of stabilization. The fluorinated adduct F3B–PF3 (see Fig. 7a) displays the weakest long-range binding signature (a comparatively small energetic rise above the minimum by r = 5 Å), in accordance with a shallow donor–acceptor association at relatively long equilibrium distance. In contrast, the chloro- and bromo-substituted systems (cf. Fig. 7b and c) show markedly stronger short-range stabilization (more negative Eint near the minimum) accompanied by a larger deformation cost at compressed distances, consistent with a more pronounced competition between orbital/charge-transfer stabilization and BX3 pyramidalization in the vicinity of the minimum. Guided by these decompositions and in analogy to the distance-resolved analyses in ref. 20, the subsequent energy-decomposition analysis (EDA) calculations are performed at a small set of representative separations sampling distinct PES regimes: a compressed geometry (r ≈ req − 0.6 Å), the equilibrium geometry (r = req), and one to two post-minimum separations (r ≈ req + 0.6 Å and r ≈ req + 1.2 Å), using the fully relaxed constrained-scan geometries at each chosen r. This selection cleanly separates the repulsive/distortion-dominated region from the near-equilibrium bonding regime and the long-range tail, enabling a physically transparent interpretation of which EDA terms govern the shape of the local PES in each complex.
IV. Summary
This study introduces a CCSD-referenced Δ-machine learning framework for coupled-cluster-quality screening of low-energy structures in homohalogenated borane–phosphine adducts, with F3B–PF3, Cl3B–PCl3, and Br3B–PBr3 as representative cases. Large geometry libraries generated at MP2 and DFT (B3LYP-D3) levels were used to train regression models for the correction ΔE = ECCSD − Elow, enabling CCSD-quality energies across thousands of structures at greatly reduced cost. Analysis of the held-out non-training subsets demonstrates that the resulting Δ-ML models reproduce CCSD energies with low errors and preserve the energetic ordering in the near-equilibrium region, offering a practical route for ranking low-energy structures on shallow potential-energy landscapes. Using the CCSD-trained predictors, we screened the complete MP2/DFT libraries and identified compact low-energy candidate sets for further refinement.Targeted CCSD(T) calculations on the ML-selected subsets show that MP2 markedly overbinds for the chlorinated and brominated adducts, whereas B3LYP-D3 can even predict the wrong sign of the formation energy for Cl3B–PCl3 by placing this borderline system too far on the unfavorable side. In contrast, the CCSD (Δ-ML) screening recovers the correct uncorrected stability sign for all three adducts and yields formation energies that track the CCSD(T) reference trends more closely than the underlying low-level methods. Inclusion of BSSE corrections further sharpens the picture, with Cl3B–PCl3 emerging as a near-threshold adduct and Br3B–PBr3 remaining clearly bound, whereas for F3B–PF3 the supermolecular CCSD and CCSD(T) calculations support weak binding, but CP-corrected coupled-cluster values could not be established because the corresponding BSSE treatment proved numerically unstable.
Distance-resolved relaxed scans and Morokuma-type EDA together establish a coherent physical interpretation of the different binding regimes. F3B–PF3 stabilizes at a longer B⋯P separation in a weak-interaction regime with comparatively small polarization/orbital contributions, whereas the chlorinated and brominated adducts adopt shorter equilibrium distances where large Pauli repulsion is overcompensated by strong exchange and polarization, with dispersion increasing systematically from F to Cl to Br. None of the three homohalogenated systems exhibits a second, outer minimum in the relaxed B–P stretching profile under the present scan protocol.
Overall, the proposed Δ-ML strategy bridges the gap between large-scale structure sampling and correlated wave-function accuracy by enabling CCSD-level screening and selective CCSD(T) validation in the low-energy region. The results also show that for shallow donor–acceptor complexes, BSSE correction, higher-level geometry refinement, and selective high-level validation are all essential for a reliable energetic classification. This workflow is expected to be transferable to other weak donor–acceptor complexes for which correlated reference data are required but exhaustive high-level exploration is computationally prohibitive.
Conflicts of interest
There are no conflicts to declare.Data availability
Supplementary information is available. See DOI: https://doi.org/10.1039/d6cp00985a.The datasets, molecular structures, pretrained models, and Δ-ML workflow implementation supporting this work are available at https://github.com/OkanKoeksal/deltaml4ccsd.
Acknowledgements
I gratefully acknowledge the computational resources (Stevin Supercomputer Infrastructure) and services used in this work that were provided by the VSC (Flemish Supercomputer Center), funded by Ghent University, FWO and the Flemish Government – department EWI.References
- A. Staubitz, A. P. M. Robertson, M. E. Sloan and I. Manners, Chem. Rev., 2010, 110, 4023–4078 CrossRef CAS PubMed.
- D. R. Martin, Chem. Rev., 1944, 34, 461–473 CrossRef CAS.
- H. Booth and D. Martin, Boron Trifluoride and Its Derivatives, John Wiley & Sons, New York, NY, 1949, ch. 4 Search PubMed.
- D. Martin and R. E. Dial, J. Am. Chem. Soc., 1950, 72, 852–856 CrossRef CAS.
- V. Jonas, G. Frenking and M. T. Reetz, J. Am. Chem. Soc., 1994, 116, 8741–8753 CrossRef CAS.
- F. Bessac and G. Frenking, Inorg. Chem., 2003, 42, 7990–7994 CrossRef CAS PubMed.
- F. Bessac and G. Frenking, Inorg. Chem., 2006, 45, 6956–6964 CrossRef CAS PubMed.
- T. A. Holme and T. N. Truong, Chem. Phys. Lett., 1993, 215, 53–57 CrossRef CAS.
- J. O. Jensen, J. Mol. Struct. THEOCHEM, 2004, 686, 165–172 CrossRef CAS.
- T. L. Allen and W. H. Fink, Inorg. Chem., 1993, 32, 4230–4234 CrossRef CAS.
- J. R. Durig and Z. Shen, J. Mol. Struct. THEOCHEM, 1997, 397, 179–190 CrossRef CAS.
- J. A. Plumley and J. D. Evanseck, J. Phys. Chem. A, 2007, 111, 13472–13483 CrossRef CAS PubMed.
- J. A. Plumley and J. D. Evanseck, J. Chem. Theory Comput., 2008, 4, 1249–1253 CrossRef CAS PubMed.
- J. A. Plumley and J. D. Evanseck, J. Phys. Chem. A, 2009, 113, 5985–5992 CrossRef CAS PubMed.
- B. G. Janesko, J. Chem. Theory Comput., 2010, 6, 1825–1833 CrossRef CAS PubMed.
- F. Huang, J. Jiang, M. Wen and Z.-X. Wang, J. Theor. Comput. Chem., 2014, 13, 1350074 CrossRef.
- C. F. Pupim, N. H. Morgon and A. Lopez-Castillo, J. Braz. Chem. Soc., 2015, 26, 1648–1655 CAS.
- T. Leyssens, D. Peeters, A. G. Orpen and J. N. Harvey, New J. Chem., 2005, 29, 1424–1430 RSC.
- H. Anane, S. El Houssame, A. El Guerraze, A. Jarid, A. Boutalib, I. Nebot-Gil and F. Tomas, J. Mol. Struct. THEOCHEM, 2004, 709, 103–107 CrossRef CAS.
- Q. L. Almas and J. K. Pearson, ACS Omega, 2018, 3, 608–614 CrossRef CAS PubMed.
- P. S. Bryan and R. L. Kuczkowski, Inorg. Chem., 1972, 11, 553–559 CrossRef CAS.
- O. Köksal, Comput. Theor. Chem., 2025, 1253, 115398 CrossRef.
- T. H. Dunning Jr., J. Chem. Phys., 1989, 90, 1007 CrossRef.
- J. Čížek, J. Chem. Phys., 1966, 45, 4256 CrossRef.
- O. Christiansen, H. Koch and P. Jørgensen, Chem. Phys. Lett., 1995, 243, 409–418 CrossRef CAS.
- K. Morokuma, J. Chem. Phys., 1971, 55, 1236 CrossRef CAS.
- T. Ziegler and A. Rauk, Theor. Chim. Acta, 1977, 46, 1 CrossRef CAS.
- A. E. Reed, L. A. Curtiss and F. Weinhold, Chem. Rev., 1988, 88, 899 CrossRef CAS.
- P. Su and H. Li, J. Chem. Phys., 2009, 131, 014102 CrossRef PubMed.
- B. J. Braams and J. M. Bowman, Int. Rev. Phys. Chem., 2009, 28, 577–606 Search PubMed.
- C. Qu, Q. Yu and J. M. Bowman, Annu. Rev. Phys. Chem., 2018, 69, 151–175 CrossRef CAS PubMed.
- J. Behler, Phys. Chem. Chem. Phys., 2011, 13, 17930–17955 RSC.
- J. Behler, Int. J. Quantum Chem., 2015, 115, 1032–1050 CrossRef CAS.
- J. Cui and R. V. Krems, J. Phys. B: At., Mol. Opt. Phys., 2016, 49, 224001 CrossRef.
- L. Himanen, M. O. J. Jäger, E. V. Morooka, F. Federici Canova, Y. S. Ranawat, D. Z. Gao, P. Rinke and A. S. Foster, Comput. Phys. Commun., 2020, 247, 106949 CrossRef CAS.
- M. Welborn, L. Cheng and T. F. Miller III, J. Chem. Theory Comput., 2018, 14, 4772–4779 CrossRef CAS PubMed.
- L. Cheng, M. Welborn, A. S. Christensen and T. F. Miller III, J. Chem. Phys., 2019, 150, 131103 CrossRef PubMed.
- T. D. Crawford and H. F. Schaefer III, Rev. Comput. Chem., 1999, 14, 33–136 Search PubMed.
- R. J. Bartlett and M. Musiał, Rev. Mod. Phys., 2007, 79, 291–352 CrossRef CAS.
- K. Raghavachari, G. W. Trucks, J. A. Pople and M. Head-Gordon, Chem. Phys. Lett., 1989, 157, 479–483 CrossRef CAS.
- C. Qu and J. M. Bowman, Phys. Chem. Chem. Phys., 2016, 18, 24835–24840 RSC.
- R. Ramakrishnan, P. O. Dral, M. Rupp and O. A. von Lilienfeld, J. Chem. Theory Comput., 2015, 11, 2087–2096 CrossRef CAS PubMed.
- A. Nandi, C. Qu, P. L. Houston, R. Conte and J. M. Bowman, J. Chem. Phys., 2021, 154, 051102 CrossRef CAS PubMed.
- J. M. Bowman, C. Qu, R. Conte, A. Nandi, P. L. Houston and Q. Yu, J. Chem. Theory Comput., 2023, 19, 1–17 CrossRef CAS PubMed.
- S. S. Khire, N. D. Gurav, A. Nandi and S. R. Gadre, J. Phys. Chem. A, 2022, 126, 1458 CrossRef CAS PubMed.
- G. M. J. Barca, C. Bertoni, L. Carrington, D. Datta, N. De Silva, J. E. Deustua, D. G. Fedorov, J. R. Gour, A. O. Gunina, E. Guidez, T. Harville, S. Irle, J. Ivanic, K. Kowalski, L. Karol, S. Sarom, H. Li, W. Li, J. J. Lutz, I. Magoulas, J. Mato, V. Mironov, H. Nakata, B. Q. Pham, P. Piecuch, D. Poole, S. R. Pruitt, A. P. Rendell, L. B. Roskop, K. Ruedenberg, T. Sattasathuchana, M. W. Schmidt, J. Shen, L. Slipchenko, M. Sosonkina, V. Sundriyal, A. Tiwari, J. L. Galvez Vallejo, B. Westheimer, M. Wloch, P. Xu, F. Zahariev and M. S. Gordon, J. Chem. Phys., 2020, 152, 154102 CrossRef CAS PubMed.
- C. Møller and M. S. Plesset, Phys. Rev., 1934, 46, 618 CrossRef.
- M. Head-Gordon, J. A. Pople and M. J. Frisch, Chem. Phys. Lett., 1988, 153, 503–506 CrossRef CAS.
- A. D. Becke, J. Chem. Phys., 1993, 98, 1372–1377 CrossRef CAS.
- S. Grimme, J. Antony, S. Ehrlich and H. Krieg, J. Chem. Phys., 2010, 132, 154104 CrossRef PubMed.
- W. J. Hehre, L. Radom, P. V. R. Schleyer and J. A. Pople, Ab Initio Molecular Orbital Theory, John Wiley & Sons, New York, 1986 Search PubMed.
- S. F. Boys and F. Bernardi, Mol. Phys., 1970, 19, 553 CrossRef CAS.
- S. J. Grabowski, Molecules, 2020, 25, 4668 CrossRef CAS PubMed.
- S. J. Grabowski, Molecules, 2020, 25, 2703 CrossRef CAS PubMed.
- D. R. Silva, L. A. Santos, M. P. Freitas, C. F. Guerra and T. A. Hamlin, Chem. – Asian J., 2020, 15, 4043 CrossRef PubMed.
| This journal is © the Owner Societies 2026 |