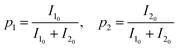Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Moiety-specific mechanism of ATP's hydrotropic action on α-synuclein
Yusuke Shuto a,
Toshifumi Mori†
a,
Toshifumi Mori†
 bc,
Benjamin Kohn‡
bc,
Benjamin Kohn‡
 d,
Erik Walinda
d,
Erik Walinda e,
Daichi Morimoto
e,
Daichi Morimoto f,
Ulrich Scheler
f,
Ulrich Scheler d,
Masatomo So
d,
Masatomo So a,
Ayako Furukawa
a,
Ayako Furukawa a,
Norio Yoshida
a,
Norio Yoshida g and
Kenji Sugase
g and
Kenji Sugase *a
*a
aDivision of Applied Life Sciences, Graduate School of Agriculture, Kyoto University, Kitashirakawa Oiwake-cho, Sakyo-ku, Kyoto 606-8502, Japan. E-mail: sugase.kenji.8c@kyoto-u.ac.jp
bInstitute for Materials Chemistry and Engineering, Kyushu University, Kasuga, Fukuoka 816-8580, Japan
cDepartment of Interdisciplinary Engineering Sciences, Interdisciplinary Graduate School of Engineering Sciences, Kyushu University, Kasuga, Fukuoka 816-8580, Japan
dLeibniz-Institut für Polymerforschung Dresden e.V., Hohe Str. 6, 01069 Dresden, Germany
eDepartment of Molecular and Cellular Physiology, Graduate School of Medicine, Kyoto University, Yoshida Konoe-cho, Sakyo-ku, Kyoto 606-8501, Japan
fDepartment of Molecular Engineering, Graduate School of Engineering, Kyoto University, Nishikyo-ku, Kyoto 615-8510, Japan
gGraduate School of Informatics, Nagoya University, Furo-cho, Chikusa-ku, Nagoya 464-8601, Japan
First published on 24th April 2026
Abstract
Adenosine triphosphate (ATP), the universal energy currency of life, also acts as a biological hydrotrope that maintains protein solubility. However, the molecular mechanism underlying its hydrotropic action, particularly how its distinct chemical moieties contribute to modulating protein conformation and preventing aggregation, remains unclear. Here, we combined NMR spectroscopy and molecular dynamics (MD) simulations to dissect the moiety-specific interactions between ATP and α-synuclein, an intrinsically disordered protein implicated in Parkinson's disease. NMR titration experiments monitoring ATP signals revealed that the adenine ring of ATP formed weak multisite interactions with α-synuclein, whereas the triphosphate group formed fewer but stronger contacts. MD simulations showed that the triphosphate-mediated contacts occurred primarily at N-terminal lysine residues and disrupted long-range intramolecular contacts, resulting in conformational expansion of α-synuclein. Energetic analysis indicated that this expansion incurred a conformational energy cost that was balanced by more favorable solvation. Based on these findings, we propose a “hierarchical binding hydrotrope mechanism”, in which the predominant contribution of each ATP moiety shifts with ATP concentration because the two moieties differ in microscopic affinity and the number of accessible interaction sites. Triphosphate-mediated binding, limited by the number of available binding sites, increases preferentially at lower ATP concentrations, whereas adenine-mediated binding increases progressively at higher concentrations. This mechanism provides a molecular basis for the concentration-dependent hydrotropic effects of ATP and clarifies how this metabolite modulates the conformational properties of aggregation-prone proteins under physiological conditions.
Introduction
The intracellular environment is highly crowded with macromolecules occupying 20–40% of the volume.1 This molecular crowding promotes protein self-association, driving aberrant aggregation such as amyloid fibrillation, and liquid–liquid phase separation (LLPS), which poses a risk of maturing into pathological aggregates. To modulate these conformational states, cells use molecular chaperones that prevent misfolding and aggregation.2 Notably, Patel et al. demonstrated that adenosine triphosphate (ATP), widely known as the universal energy currency, also functions as a hydrotrope,3 a class of small amphiphilic molecules that enhance the solubility of hydrophobic substances.4 They showed that at physiological concentrations (2–8 mM), ATP suppressed the amyloid fibrillation of Aβ1–42 and Mot3, and dissolved liquid droplets of FUS, TAF15, hnRNPA3, and PGL3 in a concentration-dependent manner. The ATP concentrations required for these hydrotropic effects lie in the millimolar range, which is substantially higher than the micromolar concentrations needed for most ATP-dependent enzymatic reactions.3 This distinction suggests that cells maintain high ATP concentrations not only for bioenergetics but also to preserve protein solubility, highlighting that ATP acts as a key factor that prevents aberrant protein aggregation.Subsequent studies have further demonstrated that ATP acts as a hydrotrope within living cells. In Xenopus oocyte nuclei, ATP destabilized nucleolar assemblies via ATP-dependent enzymatic reactions and dispersed phase-separated nucleoli via hydrotropic effects.5 Proteome-wide solubility profiling demonstrated that supplementing human cell lysates with >2 mM ATP solubilized a large fraction of otherwise insoluble proteins, while ATP depletion in cells reduced proteome solubility.6 Complementing these cellular observations, detailed in vitro analyses have revealed that ATP often modulates protein condensation in a biphasic manner. For FUS, ATP promoted droplet formation at submillimolar concentrations but dispersed droplets at millimolar concentrations by disrupting interprotein contacts.7,8 Similarly, for CAPRIN1 and NBDY, ATP promoted LLPS at low concentrations, while higher concentrations inhibited it.9,10 In contrast to these biphasic effects, ATP exhibited distinct behaviors for other proteins. For TDP-43, ATP bound to the N-terminal domain, stabilizing functional oligomers and preventing fibrillation.11,12 For Aβ peptides (Aβ1–42, Aβ1–40, and Aβ16–22), ATP hindered fibrillation by inducing amorphous aggregation or via transient, nonspecific binding.13–15 Collectively, these findings indicate that hydrotropic ATP interacts weakly and nonspecifically with proteins, and that its effects are concentration-dependent.
Given this concentration dependence, α-synuclein (a 140-amino-acid intrinsically disordered protein) is a compelling target for exploring the molecular basis of the hydrotropic effects of ATP in a physiologically relevant context. Its pathological aggregation, a hallmark of Parkinson's disease, is closely linked to the susceptibility of dopaminergic neurons to ATP depletion. These neurons, located in the substantia nigra, are particularly vulnerable because their large axonal arbors and autonomous pacemaking impose exceptionally high ATP demands.16 Age-related mitochondrial decline reduces their ability to maintain high ATP concentrations, resulting in ATP insufficiency. This insufficiency is implicated in the assembly of α-synuclein into toxic oligomers and amyloid fibrils within Lewy bodies.17–19 Consistent with this vulnerability, ATP depletion in mammalian neurons increased axoplasmic viscosity and promoted phase separation and aggregation of α-synuclein, whereas restoring millimolar ATP levels dissolved these condensates.20 In the same system, ATP also modulated the condensation of TDP-43. Together, these findings indicate that the intracellular ATP concentration is a critical determinant of the aggregation propensity of α-synuclein.
Elucidating the mechanism of these effects requires a detailed investigation into how ATP interacts with α-synuclein at the atomic level. Our previous NMR study revealed that ATP interacts weakly and nonspecifically with α-synuclein and slows hydrogen exchange between amide protons and water, and that ATP itself undergoes slight self-aggregation at concentrations of >10 mM.21 However, the molecular mechanism by which these interactions modulate α-synuclein aggregation remains elusive because previous studies on the hydrotropic effects of ATP, including ours, focused primarily on the target proteins, whereas detailed analyses of ATP itself are lacking. In particular, the relative importance of each moiety of ATP, and how these moiety-specific interactions alter the physicochemical properties of proteins underlying aggregation suppression, are unresolved. Therefore, dissecting the specific contributions of each ATP moiety is essential for elucidating the molecular basis of its hydrotropic effect.
In this study, we investigated the moiety-specific interactions between ATP and α-synuclein by combining NMR titration, diffusion NMR, and MD simulations. Our results revealed that the hydrophobic adenine ring binds weakly to many sites on α-synuclein, whereas the negatively charged triphosphate group binds more strongly to fewer sites. Collectively, these interactions induced conformational expansion of α-synuclein. We propose that the interplay between these distinct binding modes of the ATP moieties explains the concentration-dependent nature of its effects and provides new insights into its role as a biological hydrotrope.
Materials and methods
Sample preparation
The recombinant plasmid encoding full-length α-synuclein fused to an N-terminal GST-SUMO tag, constructed in the pET-23a vector, was transformed into Escherichia coli BL21(DE3). To express GST-SUMO-tagged α-synuclein, the transformed colonies were pre-cultured at 37 °C overnight in 200 mL of LB medium supplemented with 50 mg L−1 ampicillin. The pre-culture was transferred to 1.8 L of the same LB medium and incubated with shaking until the OD600 reached approximately 0.6. Protein expression was induced with 1 mM IPTG, followed by overnight incubation at 20 °C. The cells were harvested by centrifugation, washed with 150 mM NaCl, and centrifuged again. The pellet was resuspended in 10 volumes of buffer per gram of wet cell pellet (50 mM Tris–HCl (pH 7.0), 1 mM EDTA, 1 mM DTT, 1 mM PMSF, and one cOmplete protease inhibitor cocktail tablet (Roche)). The suspension was sonicated and centrifuged at 4 °C. The resulting supernatant was filtered through a 0.45 µm syringe filter and loaded onto a 20-mL Glutathione Sepharose 4B column (Cytiva). After washing the column with lysis buffer (50 mM Tris–HCl (pH 7.0), 1 mM EDTA, and 1 mM DTT), the protein was eluted with elution buffer (50 mM Tris–HCl (pH 7.0), 100 mM NaCl, 1 mM EDTA, 1 mM DTT, and 10 mM reduced glutathione), and the pH was adjusted to 7.0 because the addition of reduced glutathione lowered the pH. The eluate was dialyzed overnight against 50 mM Tris–HCl (pH 7.0), 20 mM NaCl, 1 mM EDTA, and 1 mM DTT in the presence of 2 mg GST-SENP2ΔN to cleave the GST-SUMO tag. The tag-cleaved protein was then loaded onto a 5-mL HiTrap Q HP column (Cytiva) using an ÄKTA go system (Cytiva) and eluted with a linear gradient of 150–350 mM NaCl. The eluate was passed through a 20-mL Glutathione Sepharose 4B column, and the flow-through fraction containing α-synuclein was collected. The purity of α-synuclein was confirmed by SDS-PAGE and mass spectrometry. The α-synuclein solution was then buffer-exchanged into ultrapure water, concentrated, and lyophilized.Before NMR experiments, the lyophilized α-synuclein was dissolved in H2O, and the pH was carefully adjusted to 7.0 with 0.1 M NaOH. The solution was lyophilized again and dissolved in an equal volume of D2O. This solvent exchange was performed to suppress the intense water resonance in 1H NMR spectra, which would otherwise reduce sensitivity and obscure signals from α-synuclein and ATP. The concentration of α-synuclein was determined using a DS-11 microvolume spectrophotometer (DeNovix) based on its molar extinction coefficient (ε280 = 5960 M−1 cm−1). NMR samples of ATP were prepared in a similar manner by dissolving Na2-ATP, carefully adjusting the pH to 7.0, lyophilizing the solution, and dissolving it in 99.8% D2O. The ATP concentration was determined using its molar extinction coefficient (ε259 = 15.4 M−1 cm−1).
NMR titration experiments
Samples were individually prepared by dissolving ATP at a final concentration of 100 µM and α-synuclein at 0, 50, 100, 200, 400, or 800 µM in 99.8% D2O containing 150 mM NaCl. Both 1H NMR and 31P NMR experiments were performed on an AVANCE NEO 500 MHz spectrometer equipped with a BBO CryoProbe (Bruker). 1H 1D NMR spectra were acquired using excitation sculpting, while 31P 1D NMR spectra were acquired using the inverse-gated 1H-decoupling sequence, with 256 and 8192 scans, respectively. 1H and 31P NMR spectra were referenced to DSS in 99.8% D2O and 85% phosphoric acid (H3PO4) in 5% D2O, respectively.Diffusion NMR experiments
α-Synuclein was prepared at a final concentration of 94.4 µM, and ATP was used at concentrations ranging from 0 to 12.3 mM. In addition, ATP-only samples were analyzed, and the concentration range was extended to include 16.4 and 24.5 mM to assess self-association at higher concentrations. Diffusion NMR experiments were performed on an Avance III 500 MHz spectrometer equipped with a Diff 30 probehead (Bruker) using a pulsed-field gradient stimulated-echo sequence, with 16 scans.22 Calibration of the gradient strength was achieved by measuring the known diffusion coefficient of H2O. The gradient strength was varied from 80 to 400 G cm−1 for α-synuclein and from 8.5 to 170 G cm−1 for ATP because the larger size of α-synuclein results in slower translational diffusion, which required stronger gradients to achieve comparable signal attenuation. Standard measurements used 16 increments of gradient strength, but 32 increments were used for ATP self-association to determine deviations from a monodisperse decay because the changes in signal intensity relative to the non-aggregated state were small and stronger gradients reduced the overall signal intensity.The diffusion coefficient (D) was calculated using the Stejskal–Tanner equation:
 | (1) |
where I is the signal intensity with the applied gradient and I0 is the reference intensity without the gradient, γ is the gyromagnetic ratio (267.513 × 106 rad−1 T−1), g is the gradient strength, δ is the gradient pulse duration (1 ms), and Δ is the diffusion time (50 ms). Data points with signal intensities at or near the noise level were excluded from the fitting. For high-concentration ATP samples, data points showing statistically significant deviations from linearity in Stejskal–Tanner plots were excluded from the fits to eqn (2).
For ATP self-association, which may involve multiple species, the analysis was simplified by assuming two species, and the data were fitted to a two-component model:
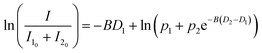 | (2) |
where the subscripts 1 and 2 denote the two molecular species.
The hydrodynamic radius (Rh) was calculated from D using the Stokes–Einstein equation:
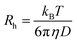 | (3) |
Computational methods
All force field parameters and equilibration steps prior to MD simulations follow our work described previously.23 In short, the Amber20 program was employed with Amber ff99SB-ILDN force field for protein and Meagher's force field for ATP.24–27 The TIP4P-D water model was used to enhance the conformational sampling28 and to prevent the artificial ATP aggregation observed with the standard TIP3P model.23 In MD simulations of α-synuclein, REUS was employed to generate initial structures consistent with the Rh range estimated by diffusion NMR. To this end, a snapshot of α-synuclein, randomly extracted from the previous work,23 was solvated in a water box of ∼13 × 13 × 13 nm3, either with or without 18 ATP molecules (i.e., ∼16 mM ATP). This high concentration was selected to ensure sufficient statistical sampling of weak, transient interactions within the limited simulation timescale, a strategy validated in our previous study.23 The systems were neutralized by adding sodium ions. Each system was then equilibrated as described previously.23 To generate the structures with REUS, a short umbrella sampling of 10 ns per restraint was performed. The restraint was applied on Rg, calculated from the Cα atoms of α-synuclein in the MD trajectories to represent the backbone structure using the following equation:
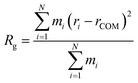 | (4) |
To identify each interaction site, contact probabilities for intramolecular α-synuclein and α-synuclein–ATP interactions were calculated from MD trajectories using pairwise heavy-atom distances. In each frame, a contact was defined when the minimum heavy-atom distance between two amino acid residues, or between an amino-acid residue and an ATP molecule, was ≤7.0 Å. For a residue pair i–j or a pair of the i-th residue and the j-th ATP molecule, the contact probability was computed as:
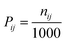 | (5) |
To explore the energetics of the ATP-induced conformational change of α-synuclein, we employed the 3D-RISM29–31 theory for solvation free energy evaluation using the RISMiCal program package.32,33 The AmberTools was used for the structural and intermolecular interaction energies.28 The α-synuclein and ATP structures used for the energy calculations were extracted every 1 ns from the corresponding trajectories of the REUS simulations. The same force field parameters of α-synuclein and ATP as in the REUS simulations were employed. The solvent was assumed to be an aqueous 0.1 M NaCl solution, and the TIP3P model was used for water.34 The temperature of the solvent was set to be the same as that used in the REUS simulations, and its density was set to 0.9728 g cm−3.
Results and discussion
Quantification of ATP-moiety-specific binding to α-synuclein by NMR titration
Although intracellular ATP exists primarily as Mg-ATP, we used Na2-ATP for all experiments to preclude complex ternary interactions mediated by Mg2+, which binds to the acidic C-terminal region of α-synuclein.21 Because Na2-ATP retains the hydrotropic function of ATP,35–37 this approach enabled us to isolate the specific contributions of the adenine and triphosphate moieties.Previous protein-observed NMR titrations estimated the dissociation constants of ATP for α-synuclein to be 4–10 mM,36 whereas complementary ligand-observed experiments had not been reported. We therefore quantitatively evaluated the binding by monitoring chemical shift perturbations of 100 µM ATP upon titration with 0–800 µM α-synuclein. The NMR spectra resolved ATP signals for the adenine (H2, H8), ribose (H1′), and triphosphate (Pα, Pβ) moieties with sufficient sensitivity and resolution (Fig. 1a, b; Fig. S1a, b). Other ATP signals were excluded due to spectral overlap with α-synuclein or attenuation by water suppression (Fig. S1c). The Pγ signal broadened significantly with increasing α-synuclein concentration (Fig. S1d). Even at an ATP concentration of 10 mM, the Pγ signal exhibited slight line broadening (Fig. S1e). This broadening arises in part from chemical exchange between ATP4− and HATP3− species with a pKa of 6.46,38 which is close to the sample pH of 7. Furthermore, additional broadening upon α-synuclein titration was likely caused by chemical exchange between free and bound ATP, which occurred in the intermediate exchange regime on the chemical shift timescale. The Pβ signal also broadened upon titration, indicating that it was likewise in the intermediate exchange regime. In contrast, the Pα and 1H signals showed only slight broadening (Fig. 1a and b), consistent with the fast exchange regime, which reflects weaker binding compared to the intermediate exchange regime. Therefore, these observations suggest that Pβ interacts more strongly with α-synuclein than the Pα, ribose, and adenine moieties.
Apparent dissociation constants 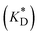 for ATP binding to α-synuclein were estimated by fitting the individual 1H and 31P chemical shift changes (Fig. 1c) to eqn (1) using the GLOVE program:39
for ATP binding to α-synuclein were estimated by fitting the individual 1H and 31P chemical shift changes (Fig. 1c) to eqn (1) using the GLOVE program:39
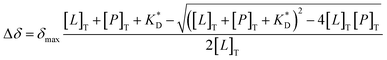 | (6) |
The observed chemical shifts represent population averages arising from multiple binding sites, and individual binding events were not resolved. Therefore, a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 stoichiometry was assumed to simplify the calculation of
1 stoichiometry was assumed to simplify the calculation of  . The estimated
. The estimated  values were 2 µM for H2, 132 µM for H8, 147 µM for H1′, 598 µM for Pα, and 2624 µM for Pβ (Fig. 1c). These values yielded a mean
values were 2 µM for H2, 132 µM for H8, 147 µM for H1′, 598 µM for Pα, and 2624 µM for Pβ (Fig. 1c). These values yielded a mean  of 701 ± 1099 µM, reflecting the heterogeneous and weak nature of the ATP–α-synuclein interactions. Notably, the
of 701 ± 1099 µM, reflecting the heterogeneous and weak nature of the ATP–α-synuclein interactions. Notably, the  for ribose H1′ was comparable to that for adenine H8. Given their spatial proximity, this similarity suggests that ribose H1′ and adenine H8 interact with α-synuclein in a similarly weak manner, possibly through the same or neighboring binding sites. Although H2 exhibited small chemical shift changes, which generally imply weak interactions, the estimate
for ribose H1′ was comparable to that for adenine H8. Given their spatial proximity, this similarity suggests that ribose H1′ and adenine H8 interact with α-synuclein in a similarly weak manner, possibly through the same or neighboring binding sites. Although H2 exhibited small chemical shift changes, which generally imply weak interactions, the estimate  was 2 µM, suggesting unexpectedly strong binding. In contrast, Pβ exhibited the largest chemical shift change but yielded the largest
was 2 µM, suggesting unexpectedly strong binding. In contrast, Pβ exhibited the largest chemical shift change but yielded the largest  of 2624 µM, suggesting weak binding. Importantly, the magnitudes of the chemical shift changes correlated with the exchange regimes: H2 showed only a small chemical shift change in fast exchange, whereas Pβ exhibited a large chemical shift change in intermediate exchange. Notably, although 31P signals inherently exhibit wider dispersion than 1H signals, the observed 31P chemical shift changes were >30-fold larger in ppm and >10-fold larger in Hz, comparable to those observed in specific ATP/ADP–enzyme binding (∼1–2 ppm). The substantial magnitude of the 31P chemical shift changes implies that the phosphate groups interact more strongly with α-synuclein than the adenine ring. However, the
of 2624 µM, suggesting weak binding. Importantly, the magnitudes of the chemical shift changes correlated with the exchange regimes: H2 showed only a small chemical shift change in fast exchange, whereas Pβ exhibited a large chemical shift change in intermediate exchange. Notably, although 31P signals inherently exhibit wider dispersion than 1H signals, the observed 31P chemical shift changes were >30-fold larger in ppm and >10-fold larger in Hz, comparable to those observed in specific ATP/ADP–enzyme binding (∼1–2 ppm). The substantial magnitude of the 31P chemical shift changes implies that the phosphate groups interact more strongly with α-synuclein than the adenine ring. However, the  values suggest the opposite trend, implying stronger binding for adenine H2 than for Pβ. This discrepancy arises because δmax (the magnitude of the chemical shift change) reflects the intrinsic binding affinity (microscopic KD) but serves only as a scaling factor in eqn (1). In contrast, the apparent KD
values suggest the opposite trend, implying stronger binding for adenine H2 than for Pβ. This discrepancy arises because δmax (the magnitude of the chemical shift change) reflects the intrinsic binding affinity (microscopic KD) but serves only as a scaling factor in eqn (1). In contrast, the apparent KD  derived from this equation reflects how readily each ATP moiety approaches saturation at low α-synuclein concentrations due to multiple binding sites.
derived from this equation reflects how readily each ATP moiety approaches saturation at low α-synuclein concentrations due to multiple binding sites.
According to the model proposed by Koshland, Némethy, and Filmer,40 the apparent KD approximates the microscopic KD divided by the number of binding sites, n  . This relationship explains the distinct moiety-specific affinities: the adenine ring yields a smaller
. This relationship explains the distinct moiety-specific affinities: the adenine ring yields a smaller  due to its interactions with a large number of hydrophobic sites, whereas the triphosphate group exhibits a larger
due to its interactions with a large number of hydrophobic sites, whereas the triphosphate group exhibits a larger  , suggesting stronger interactions restricted to fewer sites. This model also rationalizes the discrepancy between the ATP-observed
, suggesting stronger interactions restricted to fewer sites. This model also rationalizes the discrepancy between the ATP-observed  (micromolar) and the protein-observed KD (millimolar). Since we previously reported that ATP interacts with nearly all 140 residues of α-synuclein,21 scaling the protein-observed KD (4–10 mM) by this factor yields values (28–71 µM) comparable to the adenine
(micromolar) and the protein-observed KD (millimolar). Since we previously reported that ATP interacts with nearly all 140 residues of α-synuclein,21 scaling the protein-observed KD (4–10 mM) by this factor yields values (28–71 µM) comparable to the adenine  . The deviation of the phosphate
. The deviation of the phosphate  from this range further reinforces the conclusion that the triphosphate group recognizes a smaller subset of binding sites. This analytical approach is corroborated by a recent study on α-synuclein binders, which similarly demonstrated that high macroscopic affinity arises from the accumulation of weak interactions across the entire sequence.41 As detailed in the MD simulation sections below, this interpretation is consistent with atomic-level contact patterns that resolve the apparent discrepancy in the
from this range further reinforces the conclusion that the triphosphate group recognizes a smaller subset of binding sites. This analytical approach is corroborated by a recent study on α-synuclein binders, which similarly demonstrated that high macroscopic affinity arises from the accumulation of weak interactions across the entire sequence.41 As detailed in the MD simulation sections below, this interpretation is consistent with atomic-level contact patterns that resolve the apparent discrepancy in the  values through the Koshland model.
values through the Koshland model.
ATP oligomerization and ATP-induced conformational changes in α-synuclein characterized by diffusion NMR
Although the titration experiments quantified ATP–α-synuclein interactions, they yielded no structural information. Given the intrinsic disorder of α-synuclein, subtle conformational changes induced by weak ATP binding are difficult to detect by conventional structural methods. Therefore, to probe conformational changes, we determined diffusion coefficients (D) and calculated the hydrodynamic radius (Rh) via the Stokes–Einstein equation (eqn (3)).Since we previously reported that ATP oligomerizes above 10 mM,21 we first examined free ATP by diffusion NMR at concentrations ranging from 1.2 to 24.5 mM (Fig. S3) to assess whether oligomeric species exist at physiological concentrations (2–8 mM) and influence the interaction with α-synuclein. The adenine H2 signal at 8.5 ppm was analyzed because it was well-resolved and the most intense among the observed ATP signals (Fig. 2a; Fig. S2). At ATP concentrations up to 8.6 mM, the Stejskal–Tanner plots were linear (Fig. S3), indicating that ATP is monomeric. As 8.6 mM is close to the physiological range (2–8 mM), these data suggest that ATP oligomers have limited physiological relevance. At concentrations ≥12.3 mM, a second diffusion component corresponding to oligomers became distinguishable (Fig. 2b, left; Fig. S3). The average Rh of the monomeric species across all concentrations was 0.48 ± 0.02 nm (Fig. 2c), which is close to the reported value (0.6 ± 0.1 nm).21 Conversely, the oligomeric component exhibited larger Rh values (0.6–1.2 nm), which became better defined with increasing concentration (Table S1). Taken together, these results indicate that ATP oligomers are observed only at high concentrations exceeding the physiological range.
We next measured the diffusion of ATP in the presence of 94.4 µM α-synuclein, with ATP concentrations ranging up to 12.3 mM (Fig. 2b, middle; Fig. S4). Given the minimal oligomerization of free ATP in this range, we applied only a single-component model to the linear region of the Stejskal–Tanner plots. The diffusion coefficients of ATP and the corresponding Rh values remained largely unchanged relative to those of free ATP (Fig. 2c). This result is expected under conditions of excess ATP, where the observed diffusion coefficient represents a weighted average dominated by free ATP.
In contrast, analysis of the methyl signal of free α-synuclein at 0.8 ppm yielded a diffusion coefficient of (7.89 ± 0.03) × 10−11 m2 s−1 and an Rh of 2.21 ± 0.01 nm (Fig. 2c; Fig. S5). This value represents an apparent Rh because the Stokes–Einstein equation assumes a rigid sphere, whereas α-synuclein is intrinsically disordered. Nevertheless, this value is directly comparable to previously reported values based on the same assumption, including 2.55 ± 0.62 nm determined by dynamic light scattering42 and 2.66 ± 0.05 nm by diffusion NMR.43 Theoretically, protein Rh values can be estimated using the following scaling relationships for globular and denatured states:
| Rh,globular = (4.75 ± 1.11)N0.29±0.02 | (7) |
| Rh,denatured = (2.21 ± 1.07)N0.57±0.02 | (8) |
As the ATP concentration increased from 0 to 12.3 mM, the diffusion coefficient of α-synuclein decreased and the apparent Rh increased, asymptotically reaching (6.34 ± 0.03) × 10−11 m2 s−1 and 2.75 nm, respectively (Fig. S5; Fig. 2c). The increase in apparent Rh from 2.21 to 2.75 nm corresponds to a volume increase of approximately 42 nm3. Assuming that this increase arises solely from ATP binding, and given that the ATP Rh is 0.48 nm (volume 0.46 nm3), the volume expansion is equivalent to 91 bound ATP molecules. Based on a KD of 4–10 mM, the fractional occupancy at 12.3 mM ATP is estimated to be 0.55–0.75. Thus, the number of potential binding sites required to accommodate 91 bound ATP molecules ranges from 121 (KD = 4 mM) to 165 (KD = 10 mM). These results imply that ATP needs to interact with nearly the entire protein sequence, which aligns with the heterogeneous binding behavior indicated by the wide range of moiety-specific  values (2–2624 µM). However, attributing the entire volume increase to binding would require an unrealistic local ATP concentration of 3.6 M, far exceeding the bulk concentration of 12.3 mM. This discrepancy suggests that the observed Rh increase results not only from ATP binding but also from conformational expansion of α-synuclein.
values (2–2624 µM). However, attributing the entire volume increase to binding would require an unrealistic local ATP concentration of 3.6 M, far exceeding the bulk concentration of 12.3 mM. This discrepancy suggests that the observed Rh increase results not only from ATP binding but also from conformational expansion of α-synuclein.
Specifically, the ATP triphosphate group likely screens the electrostatic interactions between the N- and C-termini that normally maintain α-synuclein's compact ensemble.45 This interpretation is consistent with previous paramagnetic relaxation enhancement (PRE) NMR studies demonstrating that ATP disrupts long-range N–C interactions, favoring extended conformations.36
ATP-induced conformational changes of α-synuclein explored by MD simulations
While the titration experiments revealed moiety-specific binding modes and the diffusion NMR suggested an ATP-induced conformational expansion, atomic details of the structural rearrangements and precise binding interfaces remained unclear. To complement the NMR results, we employed all-atom MD simulations using the Amber ff99SB-ILDN force field for protein, Meagher's force field for ATP, and TIP4P-D water model.23 To thoroughly explore the conformational space of the intrinsically disordered α-synuclein, we performed replica exchange umbrella sampling (REUS) using the radius of gyration (Rg) of α-synuclein as the collective variable. The resulting free energy profiles revealed a shift in the global minimum from Rg = 2.49 to 2.71 nm upon ATP binding (Fig. S6). These values are comparable to the Rh range (2.21–2.75 nm) determined by diffusion NMR, indicating that the simulations captured structural ensembles consistent with the experimental data.To characterize the ensembles corresponding to these energy minima, we extracted two independent 500-ns trajectories from the replicas with umbrella restraints at Rg = 2.2 and 2.6 nm in the absence of ATP, and two with restraints at Rg = 2.6 and 3.0 nm in the presence of ATP (Fig. S6a). Since the two trajectories in each set displayed consistent Rg fluctuation patterns within each condition (Fig. 3a), they were combined for subsequent analyses. Analysis of the Rg distributions showed that ATP binding increased the ensemble-averaged Rg from 2.40 to 2.79 nm (Fig. 3b), indicating ATP-induced conformational expansion. Furthermore, ATP binding led to a notably broader Rg distribution compared to that in the absence of ATP, suggesting that α-synuclein exhibits enhanced conformational flexibility in the presence of ATP. Because α-synuclein does not adopt a single rigid structure, the Rg distributions under both conditions naturally show substantial overlap. However, the significant shift of the PMF minimum, the consistent increase in the ensemble-averaged Rg, and the systematic changes in the intramolecular contact maps indicate a clear transition of the entire ensemble rather than a minor fluctuation.
To identify the amino acid residues underlying the conformational expansion, we computed residue–residue contact maps for all structures sampled at 1-ns intervals (Fig. 3c). In the absence of ATP, the contact map revealed diverse intramolecular interactions, including short-range contacts within the N-terminal region (e.g., E13–Y39, Y39–E61), medium-range contacts between the non-amyloid-β component (NAC) and C-terminal regions (Q79–L113), and long-range contacts between the N- and C-terminal regions (K10–Y125) (Fig. 3c, left). The presence of such long-range interactions is consistent with previous studies reporting that α-synuclein adopts conformations associated with transient long-range intramolecular contacts.46 In the presence of ATP, these contacts were markedly weakened, with contact map scores decreasing for E13–Y39 (5 → 0), Y39–E61 (17 → 2), Q79–L113 (10 → 0), and K10–Y125 (8 → 0). Concurrently, new contacts emerged around the NAC region, including N65–T75 (2 → 22), Y39–T75 (0 → 9), and Y39–L113 (0 → 14) (Fig. 3c, right).
To determine the specific contributions of ATP moieties to these changes, we calculated contact maps of ATP–α-synuclein interactions for the entire ATP molecule as well as separately for the adenine and triphosphate moieties (Fig. 3d). These maps visualize the probabilities and spatial distribution of contacts (distinct from the intrinsic binding affinity). The analysis showed that ATP molecules predominantly interact with the lysine-rich N-terminal region and partially with the NAC region, while making minimal contacts with the C-terminal region (Fig. 3d, top). The contact map of the adenine ring alone closely resembled that of the entire ATP molecule (Fig. 3d, middle). In contrast, contacts involving the triphosphate group were fewer and relatively confined to the N-terminal region (Fig. 3d, bottom). These findings indicate that the adenine ring largely determines the overall interaction pattern, whereas the triphosphate groups, some of which do not directly bind, recruit water molecules, thereby increasing the hydrophilicity of the region. This observation supports the interpretation from the titration experiments, which suggested that the adenine ring interacts with multiple sites on α-synuclein, while the triphosphate group binds to fewer sites.
The predominance of ATP contacts with the N-terminal region suggests that electrostatic and/or cation–π interactions between ATP and lysine residues in this region interfere with long-range intramolecular contacts between the positively charged N-terminal and negatively charged C-terminal regions. This disruption likely drives the observed conformational expansion of α-synuclein. Simultaneously, exposure of the NAC region to water may promote new short-range contacts, including interactions with the hydrophobic adenine ring of ATP, possibly as a compensatory mechanism to shield the hydrophobic NAC region from water.
ATP-induced conformational and energetic changes in α-synuclein
To visualize the structural consequences of the observed contact rearrangements, we examined representative MD snapshots and confirmed that α-synuclein maintained compact conformations in the absence of ATP, whereas it adopted expanded and highly dynamic conformations in the presence of ATP (Fig. 4a; Fig. S7). Reflecting this dynamic behavior, the number of bound ATP molecules varied markedly throughout the MD simulations (Fig. 4b), indicating repeated association and dissociation. Quantitative analysis using a 7 Å cutoff revealed that an average of 5.4 ATP molecules were localized near α-synuclein (Fig. 4c). These ATP molecules formed oligomers on the surface of α-synuclein (Fig. 4a; Fig. S7). While ATP oligomerization was not observed in the NMR titration experiments, the formation of these oligomers can be attributed to the higher concentration (∼16 mM) used to enhance the sampling of weak interactions.23 Importantly, our experimental diffusion NMR data (Fig. 2c) show that the hydrodynamic radius of α-synuclein increases and saturates as the ATP concentration rises, confirming that such high concentrations do not reverse the structural expansion trend. Therefore, the qualitative behaviors observed in our MD simulations match the experimental data and are not high-concentration artifacts. The ATP oligomers were internally stabilized by Na+-coordinated triphosphate groups and π–π stacking between adenine rings (Fig. S8a). The oligomers anchored primarily to the N-terminal region, where residues including K6, K10, K43, and H50 formed salt bridges or hydrogen bonds with the triphosphate or ribose moieties (Fig. S8b). Although positively charged lysine residues readily form salt bridges with the negatively charged triphosphate groups of ATP, our MD simulations revealed that sodium ions within the ATP oligomers mediated the interaction with the N-terminal lysine residues by partially neutralizing the triphosphate groups. This neutralization facilitated cation–π interactions between the lysine side chains and the adenine rings, consistent with a previous report.47To elucidate the energetic basis of the ATP-induced effects on α-synuclein, we analyzed the conformational energy (ΔEmm), solvation free energy (ΔGsolv), and total free energy (ΔGtotal) of α-synuclein in the presence and absence of ATP. Both energetic components exhibited considerable fluctuations throughout the trajectories (Fig. 4d, top and middle), reflecting the highly dynamic nature of α-synuclein under both conditions. Upon ATP binding, the conformational energy increased from 1249 ± 252 kJ mol−1 to 1703 ± 258 kJ mol−1, while the solvation free energy decreased from −4330 ± 229 kJ mol−1 to −4762 ± 242 kJ mol−1. Although the absolute values of these energies depend on specific force field parameters and the 3D-RISM model, our analysis focuses on the relative energy changes associated with structural transitions. Because 3D-RISM is an established method for evaluating relative solvation behavior, the qualitative trends remain physically consistent regardless of the absolute baselines. The changes of +454 kJ mol−1 in conformational energy and −432 kJ mol−1 in solvation free energy correspond to the energy scale of tens of hydrogen bonds or more than ten salt bridges, indicating that ATP weakens intramolecular interactions within α-synuclein. The observed energy balance suggests that the penalty of disrupting compact intramolecular contacts is largely compensated by favorable protein–water interactions. However, further analysis revealed that the total free energy increased from −3081 ± 41 kJ mol−1 to −3059 ± 41 kJ mol−1 upon ATP binding, leaving an average net energetic penalty of +22 kJ mol−1 (Fig. 4d, bottom). Although intrinsically disordered proteins are highly fluctuating systems where statistical averaging is difficult, these two independent energy components changed by similar magnitudes in opposite directions, and their net difference consistently converged to the scale of tens of kJ mol−1. This energy scale is physically reasonable because it directly aligns with the weak, millimolar-range binding observed in our NMR experiments.
To identify the driving force that overcomes this energetic penalty to induce the conformational expansion of α-synuclein, we devised a thermodynamic scheme involving a hypothetical intermediate state in which α-synuclein retains its ATP-bound conformation in the absence of ATP (Fig. 4e). This scheme separates the overall process into two steps: the conformational transition (ΔG1 = +22 kJ mol−1), which corresponds to the energetic penalty identified above, and the ATP binding event (ΔG2). We then calculated ΔG2 as the sum of the intermolecular interaction energy between α-synuclein and ATP and the change in solvation free energy upon ATP binding. Since the number of bound ATP molecules varied (Fig. 4b), the calculated free energies were normalized by the number of bound ATP molecules. This analysis yielded an interaction energy of −427 kJ mol−1 and a solvation free energy change of +390 kJ mol−1 per ATP molecule, resulting in a net binding free energy (ΔG2) of −37 kJ mol−1. This binding free energy outweighs the energetic penalty (+22 kJ mol−1), resulting in a negative net free energy change (−15 kJ mol−1). Taken together, these results demonstrate that ATP binding drives conformational expansion of α-synuclein through multiple ion- and π-mediated contacts, where the favorable binding free energy overcomes the energetic penalty, leading to spontaneous expansion.
Hierarchical binding hydrotrope mechanism
In this study, we elucidated the molecular basis of ATP's hydrotropic action on α-synuclein by dissecting the contributions of its individual chemical moieties (Fig. 5). While previous NMR studies relied on protein-observed signals to characterize ATP binding,7,9,11,12,21,36 our ATP-observed NMR approach enabled quantitative differentiation between the binding modes of the adenine and triphosphate moieties. This NMR analysis, combined with MD simulations, revealed that the adenine ring engaged in weak multisite interactions, whereas the triphosphate group formed stronger, more specific contacts anchored to N-terminal lysine residues. These collective interactions disrupted native long-range intramolecular contacts of α-synuclein, leading to a conformational expansion whose thermodynamic cost was compensated by favorable solvation free energy and ATP binding free energy, thereby maintaining the protein in a soluble state without net thermodynamic destabilization.The difference in intrinsic binding affinity between the triphosphate group and the adenine ring provides a mechanistic explanation for the concentration-dependent and often biphasic effects of ATP that have been observed in various protein systems. We propose a model, termed the “hierarchical binding hydrotrope mechanism,” in which the predominant contribution of each ATP moiety shifts with ATP concentration because the two moieties differ in microscopic affinity and the number of accessible interaction sites. In this model, relatively high-affinity interactions of the triphosphate group with cationic residues dominate at lower ATP concentrations. In contrast, adenine-mediated contacts are distributed over a larger ensemble of weakly interacting regions and become prevalent at higher concentrations due to the greater number of accessible sites.
The dominance of triphosphate-mediated interactions at lower ATP concentrations accounts for the promotion of structural or oligomeric transitions by anchoring to specific arginine and lysine residues. FUS LLPS was enhanced at low millimolar ATP concentrations.7 A similar pattern was observed in the prion-like domain of TDP-43, where NMR studies confirmed that specific arginine residues serve as high-affinity binding sites for ATP-induced LLPS.9,10 ATP also promoted CAPRIN1 LLPS through interactions with arginine-rich regions,9 as well as LLPS in basic intrinsically disordered proteins via lysine and arginine residues.48 Furthermore, ATP triggered LLPS followed by amyloid fibril formation in insulin-derived chimeric peptides via electrostatic interactions with lysine-rich regions.49
Conversely, at higher ATP concentrations, adenine-mediated interactions become functionally dominant, typically resulting in condensate dissolution or aggregation inhibition. This high-concentration regime accounts for the dissolution of droplets observed in FUS, TDP-43, and CAPRIN1,7,9,11,12 as well as the inhibition of late-stage fibril formation in α-synuclein.36 Even non-aggregating proteins, such as lysozyme and IgG1, underwent ATP-induced LLPS in this regime.50,51 The role of the adenine ring was also supported by MD simulations showing that ATP formed clusters on hydrophobic patches, such as those on Aβ and FUS.8,13 These adenine-mediated interactions are consistent with reports of nonspecific contacts involving glutamine, serine, and tyrosine residues in the prion-like domain of FUS, and polar and hydrophobic regions in Aβ.8,13,14 Energy decomposition analysis of Aβ peptides (e.g., Aβ16–22 and Aβ1–40) also showed that adenine-driven van der Waals interactions dominated fibrillation suppression.
The hierarchical binding hydrotrope mechanism provides a unified framework that reconciles active debates about ATP's mode of action. Previous proposals include the Hofmeister effect model,52 which emphasizes the kosmotropic properties of the triphosphate group; the cosolute model,47 which categorizes ATP interactions into specific ligand binding, kosmotropic stabilization, and nonspecific binding; and the classical hydrotrope model, which posits that ATP acts as an amphiphilic molecule to solubilize hydrophobic patches. Our findings refine the classical hydrotrope hypothesis by dissecting the distinct roles of the ATP moieties. By characterizing the process as a sequential engagement in which specific, high-affinity electrostatic anchoring is progressively complemented by widespread, nonspecific hydrophobic shielding, this model integrates key aspects of previous theories into a single mechanism that explains the biphasic hydrotropic action of ATP.
Although the biphasic switching of interaction modes is characteristic of in vitro titration experiments, the physiological ATP concentration range (2–8 mM) is sufficient to simultaneously engage both triphosphate-mediated electrostatic interactions and adenine-mediated hydrophobic contacts. Consequently, in healthy cells, ATP functions as a dual-mode modulator of protein interactions. The hierarchical nature of these interactions provides a mechanistic explanation for the link between bioenergetic compromise and protein aggregation pathology. Under physiological conditions, intracellular ATP concentrations are maintained at levels that effectively saturate the triphosphate-mediated electrostatic sites while simultaneously occupying the numerous low-affinity adenine-mediated hydrophobic sites. Our model suggests that this “adenine shield” is essential for maintaining aggregation-prone proteins in soluble states. However, in pathological states associated with mitochondrial dysfunction, where ATP levels decline substantially, the weak, nonspecific adenine-mediated interactions are preferentially lost because of their lower affinity. The resulting loss of hydrophobic shielding promotes transitions toward aggregation-competent states. The proposed mechanism relies on fundamental physicochemical properties, such as electrostatic interactions with cationic residues and hydrophobic interactions. Therefore, this mechanism is likely applicable to not only α-synuclein but also other proteins implicated in neurodegenerative diseases. For instance, Tau and TDP-43, which possess cationic regions together with aggregation-prone hydrophobic segments, may similarly rely on high intracellular ATP concentrations to suppress aberrant transitions.
Conclusions
This study established the hierarchical binding hydrotrope mechanism as a framework in which specific electrostatic anchoring and diffuse hydrophobic shielding cooperatively modulate protein aggregation states. This model reconciles prior debates about ATP's mode of action and clarifies the mechanistic link between cellular energy status and proteostasis. Given its fundamental physicochemical basis, the mechanism is likely applicable to a broad range of proteins and supports therapeutic strategies, including the design of dual-mode small molecules that reinforce the adenine shield to restore proteome integrity in neurodegenerative diseases.Author contributions
Conceptualization: K. S.; writing – original draft and investigation (protein preparation and NMR measurements): Y. S.; methodology and software (molecular dynamics simulations and data collection): T. M.; formal analysis (MD data analysis): N. Y.; investigation (NMR measurements support): B. K., D. M., U. S., and K. S.; supervision: K. S.; writing – review and editing: all authors. All authors have read and agreed to the published version of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The datasets with analysis and discussion supporting this article have been incorporated and presented as part of the comprehensive supplementary information (SI). Supplementary information: overall 1D NMR spectra; fitting results of diffusion NMR recorded at various concentrations; structural ensembles of α-synuclein obtained from replica exchange umbrella sampling; and representative structural snapshots from the trajectories, including Fig. S1–S8 and Table S1. See DOI: https://doi.org/10.1039/d6cp00884d.Acknowledgements
We thank Dr Hajime Sato (Bruker) for his assistance with NMR measurements. This work was supported by JSPS KAKENHI (including a DC2 fellowship; grant numbers 21H02442, 22H05088, 22KK0098, 22H05089, 23K23303, 23KK0254, 24K21756, 24K01434, 24H00282, 25H00913, 25H02464, 25K02235, and 25K02240), the Japan Keirin-Association grant (JKA; grant number 2023M-363), the Kyoto University 125th Anniversary Fund “Kusunoki 125”, JST (ASPIRE; grant number JPMJAP2315), the MEXT Program “Data Creation and Utilization-Type Material Research and Development Project” (Grant No. JPMXP1122714694), and the Applied Quantum Computing Challenge Program by the National Research Council of Canada (Project No. AQC203). This work was performed in part using the NMR spectrometers with the ultra-high magnetic fields under the Collaborative Research Program of the Institute for Protein Research, Osaka University (NMRCR-24-05). MD simulations were partially carried out at the Research Center for Computational Sciences in Okazaki (Grant No. 24-IMS-C105, 25-IMS-C068 and 25-IMS-C105), the GENKAI supercomputer by Research Institute for Information Technology, Kyushu University, the Center for Computational Sciences, University of Tsukuba, the Supercomputer Fugaku under the Data-Driven Research Methods Development and Materials Innovation Led by Computational Materials Science, JPMXP1020230327, (Project ID: hp230212, hp240223, hp250229).References
- M. Grimaldo, F. Roosen-Runge, F. Zhang, F. Schreiber and T. Seydel, Dynamics of proteins in solution, Q. Rev. Biophys., 2019, 52, e7 CrossRef.
- A. C. Mistry, D. Chowdhury, S. Chakraborty and S. Haldar, Elucidating the novel mechanisms of molecular chaperones by single-molecule technologies, Trends Biochem. Sci., 2024, 49, 38–51 CrossRef CAS PubMed.
- A. Patel, L. Malinovska, S. Saha, J. Wang, S. Alberti, Y. Krishnan and A. A. Hyman, ATP as a biological hydrotrope, Science, 2017, 356, 753–756 CrossRef CAS PubMed.
- L. Y. Zakharova, E. A. Vasilieva, A. B. Mirgorodskaya, S. V. Zakharov, R. V. Pavlov, N. E. Kashapova and G. A. Gaynanova, Hydrotropes: Solubilization of nonpolar compounds and modification of surfactant solutions, J. Mol. Liq., 2023, 370, 120923 CrossRef CAS.
- M. H. Hayes, E. H. Peuchen, N. J. Dovichi and D. L. Weeks, Dual roles for ATP in the regulation of phase separated protein aggregates in Xenopus oocyte nucleoli, eLife, 2018, 7, e35224 CrossRef PubMed.
- S. Sridharan, N. Kurzawa, T. Werner, I. Günthner, D. Helm, W. Huber, M. Bantscheff and M. M. Savitski, Proteome-wide solubility and thermal stability profiling reveals distinct regulatory roles for ATP, Nat. Commun., 2019, 10, 1155 CrossRef PubMed.
- J. Kang, L. Lim and J. Song, ATP enhances at low concentrations but dissolves at high concentrations liquid-liquid phase separation (LLPS) of ALS/FTD-causing FUS, Biochem. Biophys. Res. Commun., 2018, 504, 545–551 CrossRef CAS PubMed.
- S. Lin, G. Hu, M. Zhang and J. Li, ATP Binding and Inhibition of Intrinsically Disordered Protein Interactions, Langmuir, 2025, 41, 3315–3324 CrossRef CAS PubMed.
- T. H. Kim, B. J. Payliss, M. L. Nosella, I. T. W. Lee, Y. Toyama, J. D. Forman-Kay and L. E. Kay, Interaction hot spots for phase separation revealed by NMR studies of a CAPRIN1 condensed phase, Proc. Natl. Acad. Sci. U. S. A., 2021, 118, e2104897118 CrossRef CAS PubMed.
- F. Liu and J. Wang, ATP Acts as a Hydrotrope to Regulate the Phase Separation of NBDY Clusters, JACS Au, 2023, 3, 2578–2585 CrossRef CAS PubMed.
- L. Wang, L. Lim, M. Dang and J. Song, A novel mechanism for ATP to enhance the functional oligomerization of TDP-43 by specific binding, Biochem. Biophys. Res. Commun., 2019, 514, 809–814 CrossRef CAS PubMed.
- M. Dang, L. Lim, J. Kang and J. Song, ATP biphasically modulates LLPS of TDP-43 PLD by specifically binding arginine residues, Commun. Biol., 2021, 4, 714 CrossRef CAS PubMed.
- M. Kuramochi, M. Nakamura, H. Takahashi, T. Komoriya, T. Takita, N. T. K. Pham, K. Yasukawa and K. Yoshimune, Adenosine triphosphate induces amorphous aggregation of amyloid β by increasing Aβ dynamics, Sci. Rep., 2024, 14, 8134 CrossRef CAS PubMed.
- S. Sarkar, S. Gupta, C. Mahato, D. Das and J. Mondal, Elucidating ATP's role as solubilizer of biomolecular aggregate, eLife, 2024, 13, RP99150 CrossRef CAS PubMed.
- T. M. Do, D. Horinek and N. Matubayasi, How ATP suppresses the fibrillation of amyloid peptides: analysis of the free-energy contributions, Phys. Chem. Chem. Phys., 2024, 26, 11880–11892 RSC.
- A. Reeve, E. Simcox and D. Turnbull, Ageing and Parkinson's disease: Why is advancing age the biggest risk factor?, Ageing Res. Rev., 2014, 14, 19–30 CrossRef CAS PubMed.
- J. Wang, L. Dai, S. Chen, Z. Zhang, X. Fang and Z. Zhang, Protein–protein interactions regulating α-synuclein pathology, Trends Neurosci., 2024, 47, 209–226 CrossRef CAS PubMed.
- M. Sharma and J. Burré, α-Synuclein in synaptic function and dysfunction, Trends Neurosci., 2023, 46, 153–166 CrossRef CAS PubMed.
- D. J. Surmeier, J. A. Obeso and G. M. Halliday, Selective neuronal vulnerability in Parkinson disease, Nat. Rev. Neurosci., 2017, 18, 101–113 CrossRef CAS PubMed.
- L. Guillaud, A. Garanzini, S. Zakhia, S. De la Fuente, D. Dimitrov, S. Boerner and M. Terenzio, Loss of intracellular ATP affects axoplasmic viscosity and pathological protein aggregation in mammalian neurons, Sci. Adv., 2025, 11, eadq6077 CrossRef CAS PubMed.
- M. Nishizawa, E. Walinda, D. Morimoto, B. Kohn, U. Scheler, M. Shirakawa and K. Sugase, Effects of Weak Nonspecific Interactions with ATP on Proteins, J. Am. Chem. Soc., 2021, 143, 11982–11993 CrossRef CAS PubMed.
- W. S. Price, Pulsed-field gradient nuclear magnetic resonance as a tool for studying translational diffusion: Part 1. Basic theory, Concepts Magn. Reson., 1997, 9, 299–336 CrossRef CAS.
- T. Mori and N. Yoshida, Tuning the ATP–ATP and ATP–disordered protein interactions in high ATP concentration by altering water models, J. Chem. Phys., 2023, 159, 035102 CrossRef CAS PubMed.
- D. A. Case, H. M. Aktulga, K. Belfon, D. S. Cerutti, G. A. Cisneros, V. W. D. Cruzeiro, N. Forouzesh, T. J. Giese, A. W. Götz, H. Gohlke, S. Izadi, K. Kasavajhala, M. C. Kaymak, E. King, T. Kurtzman, T.-S. Lee, P. Li, J. Liu, T. Luchko, R. Luo, M. Manathunga, M. R. Machado, H. M. Nguyen, K. A. O’Hearn, A. V. Onufriev, F. Pan, S. Pantano, R. Qi, A. Rahnamoun, A. Risheh, S. Schott-Verdugo, A. Shajan, J. Swails, J. Wang, H. Wei, X. Wu, Y. Wu, S. Zhang, S. Zhao, Q. Zhu, T. E. I. Cheatham, D. R. Roe, A. Roitberg, C. Simmerling, D. M. York, M. C. Nagan and K. M. Merz, AmberTools, J. Chem. Inf. Model., 2023, 63, 6183–6191 CrossRef CAS PubMed.
- R. Salomon-Ferrer, A. W. Götz, D. Poole, S. Le Grand and R. C. Walker, Routine Microsecond Molecular Dynamics Simulations with AMBER on GPUs. 2. Explicit Solvent Particle Mesh Ewald, J. Chem. Theory Comput., 2013, 9, 3878–3888 CrossRef CAS PubMed.
- K. Lindorff-Larsen, S. Piana, K. Palmo, P. Maragakis, J. L. Klepeis, R. O. Dror and D. E. Shaw, Improved side-chain torsion potentials for the Amber ff99SB protein force field, Proteins:Struct., Funct., Bioinf., 2010, 78, 1950–1958 CrossRef CAS PubMed.
- K. L. Meagher, L. T. Redman and H. A. Carlson, Development of polyphosphate parameters for use with the AMBER force field, J. Comput. Chem., 2003, 24, 1016–1025 CrossRef CAS PubMed.
- D. Song, R. Luo and H.-F. Chen, The IDP-Specific Force Field ff14IDPSFF Improves the Conformer Sampling of Intrinsically Disordered Proteins, J. Chem. Inf. Model., 2017, 57, 1166–1178 CrossRef CAS PubMed.
- D. Beglov and B. Roux, An Integral Equation To Describe the Solvation of Polar Molecules in Liquid Water, J. Phys. Chem. B, 1997, 101, 7821–7826 CrossRef CAS.
- A. Kovalenko and F. Hirata, Three-dimensional density profiles of water in contact with a solute of arbitrary shape: a RISM approach, Chem. Phys. Lett., 1998, 290, 237–244 CrossRef CAS.
- N. Yoshida, S. Phongphanphanee and F. Hirata, Selective Ion Binding by Protein Probed with the Statistical Mechanical Integral Equation Theory, J. Phys. Chem. B, 2007, 111, 4588–4595 CrossRef CAS PubMed.
- N. Yoshida, The Reference Interaction Site Model Integrated Calculator (RISMiCal) program package for nano- and biomaterials design, IOP Conf. Ser.:Mater. Sci. Eng., 2020, 773, 012062 Search PubMed.
- Y. Maruyama and N. Yoshida, RISMiCal: A software package to perform fast RISM/3D-RISM calculations, J. Comput. Chem., 2024, 45, 1470–1482 CrossRef CAS PubMed.
- W. L. Jorgensen, J. Chandrasekhar, J. D. Madura, R. W. Impey and M. L. Klein, Comparison of simple potential functions for simulating liquid water, J. Chem. Phys., 1983, 79, 926–935 CrossRef CAS.
- X. Ou, Y. Lao, J. Xu, Y. Wutthinitikornkit, R. Shi, X. Chen and J. Li, ATP Can Efficiently Stabilize Protein through a Unique Mechanism, JACS Au, 2021, 1, 1766–1777 CrossRef CAS PubMed.
- E. R. Kamski-Hennekam, J. Huang, R. Ahmed and G. Melacini, Toward a molecular mechanism for the interaction of ATP with alpha-synuclein, Chem. Sci., 2023, 14, 9933–9942 RSC.
- Y. Zhu, S. Lin, L. Meng, M. Sun, M. Liu, J. Li, C. Tang and Z. Gong, ATP promotes protein coacervation through conformational compaction, J. Mol. Cell Biol., 2024, 16, mjae038 CrossRef CAS PubMed.
- R. Tribolet and H. Sigel, Influence of the protonation degree on the self-association properties of adenosine 5′-triphosphate (ATP), Eur. J. Biochem., 1988, 170, 617–626 CrossRef CAS PubMed.
- K. Sugase, T. Konuma, J. C. Lansing and P. E. Wright, Fast and accurate fitting of relaxation dispersion data using the flexible software package GLOVE, J. Biomol. NMR, 2013, 56, 275–283 CrossRef CAS PubMed.
- D. E. Koshland, G. Némethy and D. Filmer, Comparison of Experimental Binding Data and Theoretical Models in Proteins Containing Subunits*, Biochemistry, 1966, 5, 365–385 CrossRef CAS PubMed.
- M. Sevenich, I. Gering, B. Kass, M. Vollmer, S. Aghabashlou Saisan, M. Tusche, T. Kupreichyk, T. Pauly, M. Stoldt, W. Hoyer, A. Willuweit, J. Kutzsche, N.-A. Lakomek, L. Nagel-Steger, L. Gremer, G. Tamgüney, J. Mohrlüder and D. Willbold, Direct disassembly of α-syn preformed fibrils into α-syn monomers by an all-D-peptide, npj Parkinson's Dis., 2025, 11, 271 CrossRef CAS PubMed.
- H. Gang, C. Galvagnion, G. Meisl, T. Müller, M. Pfammatter, A. K. Buell, A. Levin, C. M. Dobson, B. Mu and T. P. J. Knowles, Microfluidic Diffusion Platform for Characterizing the Sizes of Lipid Vesicles and the Thermodynamics of Protein–Lipid Interactions, Anal. Chem., 2018, 90, 3284–3290 CrossRef CAS PubMed.
- A. S. Morar, A. Olteanu, G. B. Young and G. J. Pielak, Solvent-induced collapse of α-synuclein and acid-denatured cytochrome c, Protein Sci., 2001, 10, 2195–2199 CrossRef CAS PubMed.
- D. K. Wilkins, S. B. Grimshaw, V. Receveur, C. M. Dobson, J. A. Jones and L. J. Smith, Hydrodynamic Radii of Native and Denatured Proteins Measured by Pulse Field Gradient NMR Techniques, Biochemistry, 1999, 38, 16424–16431 CrossRef CAS PubMed.
- C. W. Bertoncini, Y.-S. Jung, C. O. Fernandez, W. Hoyer, C. Griesinger, T. M. Jovin and M. Zweckstetter, Release of long-range tertiary interactions potentiates aggregation of natively unstructured α-synuclein, Proc. Natl. Acad. Sci. U. S. A., 2005, 102, 1430–1435 CrossRef CAS PubMed.
- C. Zhang, Y. Pei, Z. Zhang, L. Xu, X. Liu, L. Jiang, G. J. Pielak, X. Zhou, M. Liu and C. Li, C-terminal truncation modulates α-Synuclein's cytotoxicity and aggregation by promoting the interactions with membrane and chaperone, Commun. Biol., 2022, 5, 798 CrossRef CAS PubMed.
- A. Hautke and S. Ebbinghaus, The emerging role of ATP as a cosolute for biomolecular processes, Biol. Chem., 2023, 404, 897–908 CrossRef CAS PubMed.
- D. Kota, R. Prasad and H.-X. Zhou, Adenosine Triphosphate Mediates Phase Separation of Disordered Basic Proteins by Bridging Intermolecular Interaction Networks, J. Am. Chem. Soc., 2024, 146, 1326–1336 CrossRef CAS PubMed.
- R. Dec, M. W. Jaworek, W. Dzwolak and R. Winter, Liquid-Droplet-Mediated ATP-Triggered Amyloidogenic Pathway of Insulin-Derived Chimeric Peptides: Unraveling the Microscopic and Molecular Processes, J. Am. Chem. Soc., 2023, 145, 4177–4186 CrossRef CAS PubMed.
- Z. Tian and F. Qian, Adenosine Triphosphate-Induced Rapid Liquid–Liquid Phase Separation of a Model IgG1 mAb, Mol. Pharm., 2021, 18, 267–274 CrossRef CAS PubMed.
- M. Zalar, J. Bye and R. Curtis, Nonspecific Binding of Adenosine Tripolyphosphate and Tripolyphosphate Modulates the Phase Behavior of Lysozyme, J. Am. Chem. Soc., 2023, 145, 929–943 CrossRef CAS PubMed.
- J. Mehringer, T.-M. Do, D. Touraud, M. Hohenschutz, A. Khoshsima, D. Horinek and W. Kunz, Hofmeister versus Neuberg: is ATP really a biological hydrotrope?, Cell Rep. Phys. Sci., 2021, 2, 100343 CrossRef CAS.
Footnotes |
| † Current address: Department of Molecular Engineering, Graduate School of Engineering, Kyoto University, Nishikyo-ku, Kyoto 615-8510, Japan. |
| ‡ Current address: Department of Radiology, Washington University in Saint Louis, St. Louis, MO 63110, USA. |
| This journal is © the Owner Societies 2026 |