 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
How cysteine oxidation affects protein stability and binding studied by free energy calculations
Tristan Alexander
Mauck
 and
Martin
Zacharias
and
Martin
Zacharias
 *
*
Center of Protein Assemblies, Technical University of Munich, Garching, Germany. E-mail: zacharias@tum.de
First published on 7th April 2026
Abstract
The extracellular environment but also cellular metabolism can generate oxidative stress that can chemically modify and damage protein molecules. The sulfur containing amino acid cysteine (CYS) is particularly vulnerable to oxidation. The molecular details of how CYS oxidation can modulate stability and binding of proteins is still not well understood. Using alchemical free energy simulations, we calculate the change in protein stability and association upon CYS oxidation to different oxidation states for two example proteins. In the case of the URN1 splicing factor FF domain (URN1-FF) the simulations predict a significant decrease in stability upon oxidation in agreement with experiment and the effect also depends on the final oxidation state. In addition, the oxidation leads to conformational changes and partial unfolding at the protein C-terminus. In contrast, for the second system, Parkinson disease protein 7 (DJ-1), CYS oxidation enhances significantly the protein monomer stability again in agreement with the experimental observation and slightly destabilizes homo dimerization. Analysis of the molecular details associated with CYS oxidation in the folded proteins allows us to gain insights into why both stabilizing as well as destabilizing effects can be observed. The CYS oxidation simulation methodology could also serve as a general protocol to analyze single and multiple CYS oxidations in other protein systems and its influence on protein binding and stability.
1 Introduction
Oxidation plays a key role in many metabolic processes and as an inevitable side effect these processes also produce reactive oxygen species that may lead to oxidative damage of proteins and other biomolecules. Due to the free electron pairs of the sulfur atom, cysteine (CYS) and methionine (MET) are particularly vulnerable to being oxidized by reactive oxygen species.1 Cysteine oxidation by both one and two electron oxidants can lead to the formation of cysteine sulfenic acid (CSO). CSO can either be converted back to cysteine with the help of oxidoreductases, form a disulfide bridge with another cysteine, be oxidized irreversibly to cysteine sulfinic acid (CSD) or remain in place stabilized by nearby residues. Cysteine sulfinic acid, in turn, can also be oxidized irreversibly, forming the final oxidation state, cysteine sulfonic acid (OCS). All of these states may convert to their deprotonated form, depending on ambient pH and local environment. The negative charging upon deprotonation is especially relevant for cysteine sulfinic and sulfonic acid with very acidic pKas but also for cysteine sulfenic acid with a pKa slightly below physiological pH (see Fig. 1 panel (A)). The biological function of cysteine oxidation remains not fully understood.2 For example, changes in enzyme activity have been reported in both directions, with tyrosine phosphatases being inactivated by cysteine oxidation while for example in the kinase domain of EGFR an increase in activity has been observed.3,4 Experimental studies have also linked the oxidation of cysteine residues to increased amyloid formation compared to the native (non-oxidized) wild type state under physiological conditions.5 In addition, similar to methionine oxidation, cysteine oxidation, significantly alters the polarity (and possibly total charge) of the residue which has also been linked to changes in thermodynamic stability of proteins. This is presumed to be a widespread effect as a recent large scale computational screening of 969 proteins including solvent exposed and non-disulfide-bonded cysteine residues predicted a significant median destabilization of 2.28 kcal mol−1. It was concluded that potentially numerous physiologically relevant proteins are prone to destabilization.5 However, it is important to note that these predictions are approximate as they were introduced as in silico mutation of cysteine to aspartic acid (ASP) and used the FoldX protein design algorithm to predict ΔΔG.6 | ||
| Fig. 1 (A) CYS and its three oxidation states. Note that only the first oxidation is reversible and can be reverted by oxido-reductases. Oxidation of CYS lowers pKa value compared to native cysteine.2 (B) Cartoon representation of yeast URN1-FF domain protein and the CYS residue at position 57.7 It is located near the C-terminus of the α3-helix and opposite to a loop segment joining α1- and α2-helices. (C) Schematic view of the DJ-1 homodimer.8 In the bound state, the monomers attain mutually inverted orientations (180° rotation). Note the location of the CYS-106, marked in red, positioned at the turn of a loop reaching into a solvent accessible cavity formed by several loops of the same strand. | ||
In the present study we aim to quantify the impact of cysteine oxidation on the thermodynamic stability rigorously by using alchemical free energy methods coupled with molecular dynamics simulations. Due to the computational costs associated with the methodology, we focus on two relevant examples for which experimental data for direct comparison is available.
The yeast URN1 splicing factor FF domain (URN1-FF) is a small globular protein found in yeast whose native fold comprised of three α-helices and one 310-helix almost resembles a loose cubic structure (see Fig. 1). It is reported that exposure to hydrogen peroxide, leading to the formation of cysteine sulfonic acid at position 57, causes a decrease in folding stability of ≈3.5 kcal mol−1.5 Additional investigations using mutations to serine and aspartic acid led to the conclusion that this drop in stability is mainly caused by the insertion of a charge rather than the loss of the thiol group. Forming the last winding of the third α3-helix, CYS-57 can be seen in the crystal structures to interact strongly with several neighboring residues.7
The second system we investigate is the DJ-1 homo dimer protein also known as Parkinson disease protein 7. This dimeric protein complex has been the focus of extensive research since its discovery in 1997 for its role in early onset Parkinson's disease.9 While the protein has been linked to a number of functions, its full spectrum remains unclear. Among the known functions are mitochondrial homeostasis, antioxidant activity and regulating inflammatory responses including protection against oxidative stress.10–12 Based on observations of non-functional proteins in patients, it is believed, that the loss of the protective functions of this protein might be an important factor in the development of Parkinson disease.11 From a structural point of view, DJ-1 forms several helices and beta sheets and contains three cysteine residues at positions 46, 53 and 106. Of these, the residue CYS-106 is most easily oxidized.10 On the molecular scale, it can be motivated by the close interaction with the surrounding residues, most importantly with residue GLU-H-18, lowering the cysteine residue's pKa to about 5.4.12 The protonation of GLU-H-18 has been observed experimentally12 and it stabilizes the deprotonated state of CYS-106 by forming a hydrogen bond. The high tendency to be oxidized is believed to be critical for the protein's functions as CYS-106 can act as a switch or sensor to changes in the concentrations of oxidants. The critical importance of the residue is further supported by the observation that a mutation to alanine lead to a general inactivation and loss of protection with respect to oxidative stress.13 While the first and second oxidation states, CSO- and CSD-, are believed to correspond to functional forms of DJ-1, oxidation to cysteine sulfonic acid (OCS-) has been connected to a loss of function. Due to its functional importance and tendency to get oxidized, our computational study will focus on the oxidation of CYS-106. It is expected that the oxidation of CYS-106 causes changes in the stability of the protein dimer but possibly also of the monomer. To this end, Kiss et al. recently published a combined experimental and computational study.14 The authors found that the oxidation of CYS-106 leads to close interactions with adjacent residues. While the reduced form of the residue only showed weak interactions. A CSO- (106) was found to interact with the backbone atoms of ALA-107 and GLY-75. When adding the second oxygen, increased interactions with GLU-H-18 were observed. Finally, the inactive form, featuring a OCS- at position 106, was found to interact with GLY-157 and HIS-126. Using the MM/GBSA (molecular mechanics generalized Born surface area) method, an approximation for the binding free energy of the dimer was calculated finding that the over-oxidation (to OCS-) seems to weaken the binding of the dimer.
The result is in line with the observation that the over-oxidation leads to a loss of function. Finally, Kiss et al. measured the thermodynamic stability of the dimer experimentally. Interestingly, both the oxidized and over-oxidized CYS-106 resulted in dimers with strongly increased melting temperature of about 13 K compared to the reduced form.14
In the present study, we apply atomistic molecular dynamics simulations and alchemical free energy methods to investigate how oxidation of CYS residues affects the stability and binding of the URN1-FF and DJ-1 protein systems. The results are in good qualitative agreement with experimental observations and provide molecular and structural details to explain the calculated and observed destabilization of URN1-FF but stabilization in case of the DJ-1 system upon CYS oxidation. In addition, the simulations give also insights into the specific effects of the different levels of CYS oxidation which are difficult to distinguish in experimental studies. Hence, our computational study extends and complements existing experimental and theoretical work to better understand the variety of CYS oxidation effects.
2 Methods
New sets of partial charges for the three oxidized states of cysteine were derived according to the AMBER force field methodology. To this end, several geometry optimizations of the capped residues were conducted at B3LYP/6-31G* level using GAUSSIAN09.15 ESP data of the resulting ensemble of low energy structures, generated at HF/6-31G* level, was then used as input for the RESP method to fit partial charges.16 In the procedure, backbone charges were fixed at the values common between charged or neutral amino acids. Bonded parameters were found with the parmchk software using entries of AMBER 14SB and GAFF libraries.17–19 The force field parameters are listed in AMBER format in the SI.The simulations were started from experimental structures available at the protein data bank.20 These are pdb2JUC (URN1-FF domain) and pdb3SF8 (DJ-1 dimer).7,8 Selected oxidized cysteine states were created using AMBER's tleap software.21 Following the methodology established by Pitera and Kollman, the unfolded state was approximated as a capped tripeptide.22 Hence, we assume that this state is fully characterized by interaction with the solvent and possibly by neighboring residues and interactions beyond this are assumed to have negligible contribution to the calculations. Wildtype and oxidized states were merged into a dual topology to be used in AMBER's free energy implementation.17,23–26 All systems were solvated in TIP3P water and neutralized with sodium or chloride ions. The salt concentration was kept low for URN1-FF and set to 0.15 M for the DJ-1 simulations. It was taken care that the box boundaries were placed at least 12 Å from the protein. Probable protonation states of the protein residues were determined using the H++ server and added in the simulations.27–29 In the sensitive case of CYS-106, the likely protonation state was determined using a free energy calculation (see Results section). Each system was first energy-minimized (2500 steps), followed by a gradual heating to 300 K within 100 ps with positional restraints on all protein heavy atoms. The setup was continued by a 100 ps NPT equilibration with the Langevin thermostat with a collision frequency of 5 per ps and the Berendsen barostat. All production runs used a time step of 2 fs and applied the SHAKE algorithm on all bonds involving hydrogen. The challenge of transforming a neutral into a charged residue and thereby altering the system's total charge (yeast URN1-FF domain) was addressed by placing both alchemical paths of the thermodynamic cycle into the same box and running the transformations in opposite direction.30,31 That means that both the transformation in the folded as well as in the unfolded state were placed in one system, however started from the opposite ends. The necessary independence of the two paths was guaranteed by positional restraining one central carbon atom in both molecules, thereby keeping a distance of at least 30 Å at all times (see Fig. 2) but otherwise allowing for full conformational freedom. Each transformation step was sampled in at least three independent production runs of 20 ns to obtain relevant averages from independent simulations. For the transformations, van-der-Waals and Coulomb parameters were altered simultaneously using a soft-core implementation.32–34 Here, the simulations of URN1-FF used standard parameters (α = 0.5, β = 12.0), while the α value was scaled down for the DJ-1 system to 0.2 to prevent Coulomb forces overpowering the van-der-Waals forces.
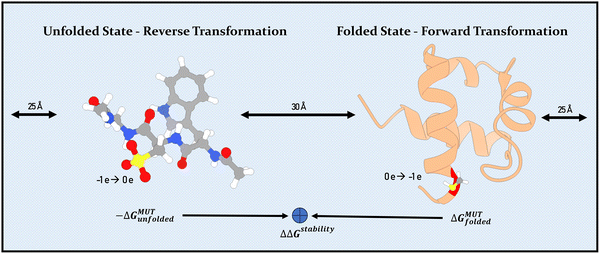 | ||
| Fig. 2 Schematic representation (not to scale) of the simulation for the alchemical transformations involving charge alteration (yeast URN1-FF domain). To avoid artifacts from changes in the system's total charge, we follow the method by Gapsys et al. (ref. 30 and 35), combining both alchemical transformation pathways in the same box thereby keeping the system overall neutral. Note, the large distance maintained between both molecules, ensures independence of the paths. The overall recorded change in free energy is the desired result. The representation of the unfolded state as a capped tri-peptide can be seen on the left. | ||
From previous studies,36 the following distribution of lambda windows was found to be an efficient distribution: {0.0, 0.025, 0.05, 0.075, 0.1, 0.15, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.85, 0.9, 0.925, 0.95, 0.975, 1.0}. For the computation of the free energy difference, 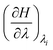 was recorded every 2 ps37,38 to approximate the final free energy difference using the trapezoidal rule. The error and convergence analysis relied on the comparison of the independent runs. The final result ΔGfinal is then computed as the average over all conducted runs. The error analysis was conducted as follows: for each result
was recorded every 2 ps37,38 to approximate the final free energy difference using the trapezoidal rule. The error and convergence analysis relied on the comparison of the independent runs. The final result ΔGfinal is then computed as the average over all conducted runs. The error analysis was conducted as follows: for each result 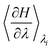 from each run, a block average analysis was conducted, yielding a statistical error for the respective result.39 Using the error propagation formula, the error carried to the average
from each run, a block average analysis was conducted, yielding a statistical error for the respective result.39 Using the error propagation formula, the error carried to the average 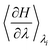 over all runs was determined. As these values would then be used in the trapezoidal rule, again the propagated final error was calculated. The total workflow is illustrated in SI Fig. S1. In addition to the TI production runs, free molecular dynamics simulation of at least 1 µs length were conducted to investigate structural impacts of CYS oxidations.
over all runs was determined. As these values would then be used in the trapezoidal rule, again the propagated final error was calculated. The total workflow is illustrated in SI Fig. S1. In addition to the TI production runs, free molecular dynamics simulation of at least 1 µs length were conducted to investigate structural impacts of CYS oxidations.
3 Results and discussion
3.1 CYS-57 oxidation in the URN1 FF domain protein
The small URN1-FF domain protein is involved in the mRNA splicing process in yeast and sensitive to oxidative damage. In experiments oxidation led to a destabilization of the folded state of about 3.5 kcal mol−1.5 Molecular dynamics simulations were started from the experimental folded structure with CYS-57, GLY-58 and GLU-59 forming the final segment of the α3-helix. During the free molecular dynamics runs we detected that this region undergoes significant dynamics at room temperature. For the wild type, the turn repeatedly opened and closed during the 1 µs simulation. The motion involves the last two residues but not CYS-57 itself. The effect is illustrated in Fig. 3(A), depicting relevant residues in atomic detail. Oxidation of CYS-57 increases the structural flexibility significantly and results in transient unfolding of the last α3-helix turn (fraction folded α3-helix shown in Fig. 3B, see also SI Fig. S2). The changes in thermodynamic stability calculated from the alchemical TI are presented in Table 1. These results are computed from five runs of 19 independent windows of 20 ns production time for each lambda step. Clearly, the oxidation has an unfavorable effect on the folding stability.| Oxidation | ΔΔGstability [kcal mol−1] |
|---|---|
| CYS → CSO | +1.29 ± 0.18 |
| CYS → CSO- | +5.33 ± 0.13 |
| CYS → CSD- | +5.70 ± 0.16 |
| CYS → OCS- | +3.29 ± 0.16 |
Whereas the oxidation to a neutral CSO-57 destabilizes the protein moderately by ≈1.3 kcal mol−1 the oxidation to a charged residue 57 causes a more significant decrease (≈3.3–5.7 kcal mol−1). Surprisingly, the oxidation to the highest oxidation state (OCS-) is predicted to result in less destabilization compared to transformation to CSO- or CSD-. The calculated result for the highest oxidation state (OCS-) is in very good agreement with experiment (3.5 kcal mol−1 (ref. 5)). Additionally, the significant destabilizing caused by the transformation to a negatively charged residue at position 57 also agrees with experimental mutagenesis studies. Here, it was observed that a mutation to serine only caused a small destabilization in comparison to a strong destabilisation due to the mutation to aspartic acid.5
In the following the effects of the oxidation on the local structure are investigated through analysis of the free molecular dynamics simulations.41 It seems, that the cysteine fulfills a specific stabilizing role in the wild type folded protein. The hydrogen of the thiol group engages in 50% of the observed frames in a hydrogen bond with the oxygen of the carbonyl group from the phenylalanine at position 53 (PHE-53) (see Fig. 4 panel C)). Occasionally (<5%), it also interacts with LYS-23, located in the disordered linker between α1- and α2-helices. While the first interaction likely helps stabilizing the α3-helix, the latter interaction could serve as a clamp-like feature, preventing the two parts of the protein from drifting apart. Upon insertion of the additional oxygen atom by the oxidation, the interactions of the oxidized cysteine residue are frequently broken. While still being able to act as a hydrogen bond donor, the increased length of the oxidized side chain hampers the interaction with the backbone of PHE-53. However, clamp-like interactions become more frequent in case of the oxidized residues. To this end, interactions are observed with both LYS-23 and SER-25. Additionally, solvent interactions with the oxidized CYS-57 become increasingly important, being present in more than 50% of the observed frames due to the partial unfolding of the final turn of the α3-helix. In agreement with the increased tendency to interact with the solvent, the graph in Fig. 4D illustrates a decreased average presence of protein residues (both hydrophilic and hydrophobic) in close vicinity (6 Å, around the central sulfur atom of residue 57). Special attention should be paid to the fact that the former hydrogen bond acceptor (backbone of PHE-53) is now only in the vicinity of the oxidized residue in 60% of the observed frames. The removal of a proton, negatively charging the residue, strongly destabilizes the protein with respect to the wild state. This can be explained by a significant change in interaction behavior. Having lost the hydrogen atom, the side chain can no longer function as a hydrogen bond donor and having turned into a strong hydrophilic residue and becomes more solvent exposed. This trend is observed for all oxidation states including OCS- and the average number of protein residues in the vicinity is reduced again (Fig. 4D). Surprisingly, the free energy calculation of CYS-57 to OCS- and hence the insertion of three oxygens, predict a stabilization relative to CSO- or CSD- by roughly 1.8 kcal mol−1. Next to the increased number of hydrogen bonds which can be formed by OCS- and the additional oxygen that enhances also van der Waals interactions an additional explanation might be the symmetry of the side chain. The arrangement of the oxygen atoms of cysteine sulfonic acid is symmetric and therefore less polar compared to the other two oxidation states.
Besides mean structural changes the effect of CYS oxidation on conformational dynamics is of interest. The loss of the interaction of residue 57 upon oxidation with the carbonyl oxygen of PHE53 does not affect the motion and root-mean-square-fluctuation (RMSF) significantly (SI Fig. S5) probably due to the localization in a stable α3-helix. LYS-23, on the other hand, can be seen to show increased fluctuations in all oxidation states compared to the wild type except for CSO- (Fig. 4B). Being located in a disordered part and being positively charged, the increased accumulation of polarity and charge in the vicinity likely causes it to move out of its native position. As expected, the oxidized CYS-57 shows increased fluctuations throughout all oxidized states because of native-like contacts. The plot also captures the significant appearance of the partly unfolded state in the long simulations for all oxidized states. This results in large ΔRMSF values for the terminal residues. The analysis also identifies a few not so obvious consequences of the oxidations. First, CSO- seems to have an impact on ARG-49 dynamics which is located more N-terminal in the -helix likely due to electrostatic interactions between the positively charged arginine and the negative charge of cysteine sulfenic acid. Additionally, all oxidized CYS-57 states influence the fluctuations of residues 15, 16 and 17 from the second helix. As these residues were not part of the neighborhood analysis, this cannot be explained from changes in direct interactions. Possibly, these are secondary effects cause by one of the changes mentioned earlier, hinting at the destabilization of the core of the protein. Overall, we found that the oxidation of CYS-57 significantly alters the local interaction pattern, causing increased fluctuations in the fold and reducing the thermodynamic stability.
3.2 Influence of CYS-106 oxidation on DJ-1 stability and dimerisation
The DJ-1 (Parkinson disease protein 7) homodimer involved in early onset Parkinson's disease has been observed to become more stable upon CYS oxidation associated with a significantly increased unfolding melting temperature compared to the wild type.14 We used alchemical free energy simulations to evaluate the change in three quantities in response to cysteine oxidation including the stability of the folded monomer, the stability of the folded dimer and the binding free energy between the two folded monomers. Since experimentally only structures of the homo dimer have been solved we used one monomer from the experimental dimer structure to represent the monomer start structure, Hence, putative large scale conformational changes upon unbinding are not considered. It was also assumed that both cysteine residues in the dimer are oxidized to the same level.The likely protonation state of CYS-106 was computed using an alchemical transformation using the relation between the change in free energy upon deprotonation in solvent and the protein environment to the change in pKa due to the protein environment.42 The charge conservation was again ensured using the setup illustrated in Fig. 2. Three independent TI runs gave a free energy difference between the two mutation paths of ΔΔG = −5.48 ± 0.45 kcal mol−1, which translates to a decrease to pKa = 4.15 ± 0.33. This result differs slightly from an experimental result by Witt et al. reporting a decrease of the pKa = 5.4, but agrees that the local protein environment supports the deprotonated state12 (termed CYM). The results from the stability and binding calculations are displayed in Tables 2 and 3. Since the DJ-1 system is less flexible than the URN1-FF domain protein three separate TI runs of 19 independent lambda windows (20 ns per window) were sufficient to obtain good convergence. We found that the oxidation affects all of the investigated properties. Especially, the double oxidations to 2× CSO- and 2× CSD- stabilize the folds of the monomer and dimer in a significant way. The oxidation to OCS- is also found to stabilize the fold, but to a lesser extent. The binding, on the other hand, is found to be weakened with each oxidation step. These effects are, however, smaller in scale. To calculated stability values require one more correction to be compared to experimental findings: due to the local surroundings of CYS-106, the deprotonated version of the cysteine states was taken as reference for the calculations. This is appropriate for the oxidized states with their low pKa values, but this is not realistic for the cysteine in the unfolded state, following the native pKa. This means that the stabilizing effect is underestimated in our model and a correction of −1.77 kcal mol−1 has to be applied per monomer (see SI Section S1.2). The alchemical free energy results agree qualitatively with the experimental results observing a fairly large increase in thermodynamic stability for CYS-106 oxidations to CSD- and OCS-.14 Still, these results support the increased activity of DJ-1 in the CSD- state as the strong shift in equilibrium concentrations to the folded monomer will also increase the concentration of bound dimers outbalancing the predicted reduced binding free energy. Interestingly, in the crystal structure of DJ-1 a CSD- was identified at position 10613 which our alchemical simulations also predict as the CYS oxidation state with the most favorable stability change (see Table 3). For the highest oxidation state (OCS-) the predicted modest increase in stability is almost offset by the decreased dimer affinity which may explain the loss of oxidative damage sensing function of this highest oxidation state of CYS-106.13,14
| Oxidation | ΔΔGdimerMUT [kcal mol−1] | ΔΔGmonomerMUT [kcal mol−1] | ΔΔGunfoldedMUT [kcal mol−1] |
|---|---|---|---|
| CYM → CSO- | −6.29 ± 0.10 | −3.41 ± 0.07 | +1.92 ± 0.06 |
| CYM → CSD- | +0.10 ± 0.10 | −0.39 ± 0.06 | +6.07 ± 0.07 |
| CYM → OCS- | +2.39 ± 0.13 | +0.35 ± 0.08 | +3.38 ± 0.06 |
| Oxidation | ΔΔGmonomerstability [kcal mol−1] | ΔΔGdimerstability [kcal mol−1] | ΔΔGdimerbinding [kcal mol−1] |
|---|---|---|---|
| CYM → CSO- | −5.33 ± 0.13 | −10.13 ± 0.22 | +0.53 ± 0.24 |
| CYM → CSD- | −6.46 ± 0.13 | −12.04 ± 0.24 | +0.88 ± 0.22 |
| CYM → OCS- | −3.03 ± 0.14 | −4.37 ± 0.25 | +1.69 ± 0.29 |
The CYS-106 is positioned in a solvent accessible cavity in each monomer, created by protein loops of the monomer and dimerization partner. As a consequence the surrounding residues are mostly hydrophilic. We found that the residue's environment is mostly conserved (see Fig. 6(C)). An interesting feature of this neighbor analysis is the fact that for all states residues of the opposite monomer are also included. These are ARG-216, PRO-372 and VAL-374 from monomer B, which are found in the neighborhood of CYS-106 (in monomer A) and vice versa for the second monomer (see Fig. 6 panel (A)). Hence, the oxidation of CYS-106 can influence the binding affinity between the monomers. All the diagrams for the native state and the three oxidation states are very similar, suggesting only small structural alterations. Slightly less interaction partners are recorded in the third oxidation state, potentially suggesting a motion away from the protein and more into the solvent. In addition, the inter-dimer interaction is seen to be reduced in CSD- and OCS- oxidation states of residue 106. From earlier studies it is known that hydrogen bonds play a critical role in the local interaction of CYS-106.14 To this end, Fig. 6 panel (B) illustrates the respective hydrogen bond frequencies with nearby residues. Next to the solvent, regular interactions are detected with three residues: GLH-18, ALA-107 and GLY-75. Based on their position relative to CYS-106 (see Fig. 6), each of the three interactions supports the native fold. However, interactions with GLU-H-18 and GLY-75 can be expected to have a larger impact as these residues are part of other strands of the fold. Formed hydrogen bonds can act here in a clamp-like fashion.
In its native conformation, CYS-106 (and its respective copy in the second monomer) interacts roughly half the time with solvent and the other half with either GLU-H-18 or to a lesser extend with ALA-107. The next two CYS oxidation states both show a reinforcement of intra-molecular hydrogen bonds overall, probably attributed to the oxygen being a better acceptor than the sulfur atom. In both cases, the oxidized residue clearly favors the interaction with GLU-H-18. A selection of frames demonstrating these interactions is provided in Fig. 5. For cysteine sulfenic acid, no interaction with ALA-107 is observed. Instead, bonds are formed with the backbone of GLY-75, a residue that is located at the start of a helix right above the cysteine. The oxidation to cysteine sulfinic acid modifies this pattern slightly. The interaction with GLU-H-18 remains the most prominent hydrogen bond partner, with GLY-75 and ALA-107 appearing on roughly similar frequency. It is noteworthy, that for cysteine sulfinic acid, a stable orientation seems to be present, with one oxygen (O4) interacting with GLU-H-18 and the other (O2) interacting with either ALA-107 and GLY-75. This pattern seems to be broken up by the introduction of cysteine sulfonic acid. Here, interactions with all previously identified partners are observed. However, no defined order as before seems to exist. Due to the symmetric arrangement of oxygen atoms in the cysteine sulfonic acid, likely several energetically equivalent orientations are possible.
Comparing the results, a possible conclusion seems that the degree of interaction with GLU-H-18 is a critical feature for DJ-1 stability. For the first two oxidation states (CSO- and CSD-) which are predicted to stabilize the folded form more than the OCS- oxidation (Table 3), an increase in interaction relative to wild type is observed. For the OCS- oxidation state still more interactions with GLU-H-18 compared to the wild type is observed in line with the observed order of folding stability. The results from this analysis agree qualitatively with the observations made by Kiss et al., in a previous study.14
To assess the effects on the conformational fluctuations, the RMSF along the chains observed during the continuous MD simulations was analyzed (SI Fig. S8). For the wild type protein dimer fluctuations of most parts of the protein are relatively low (≈1 Å). An exception are the regions of the residues 59 to 69 and 126 to 140, showing fluctuations of up to 3 Å. These regions each feature an exposed helix and a disordered coil positioned in front of the CYS-106 cavity and at the other side of the protein, respectively. The changes in RMSF upon oxidation indicate a reduction of conformational mobility of regions close to the oxidized CYS-106 by about 1 Å (to a lesser degree seen in monomer B). Interestingly, this stabilization seems to go hand in hand with a destabilization of the disordered part following the helix. Next to the changes in this domain, there are changes in the mobility of specific residues in the cavity around the CYS-106 residue (see color-coded depiction of the location of these features in Fig. 6(A)). In most of the graphs, three residues, ASN-76 (yellow), ARG-48 (lime) and ARG-156 (light blue), show reduced motion, suggesting that they have been stabilized indirectly by altered interactions of the oxidized CYS-106. As arginine is a positively charged residue and asparagine shows significant polarity, a stabilizing electrostatic interaction may also contribute. Each of the three residues is part of a different protein strand reaching into the pocket, explaining an improved folding stability.
4 Conclusions
Reactive oxygen species generated during many cellular stress events and also during metabolic processes can cause oxidation of CYS residues which influence protein stability and modulate protein function. Also, different levels of oxidation are possible and the molecular details of how these CYS oxidation states affect folding stability and binding affinity are not well understood. Our alchemical free energy simulations in explicit solvent allow us to accurately calculate changes in stability and binding due to CYS oxidation and to characterize changes in structure and dynamics in atomistic detail. It includes the analysis of the effect of different CYS oxidation states, the independent treatment of the protonated and deprotonated states of cysteine sulfenic acid or the selective investigation of the cysteine oxidation on monomer stability and dimer association in case of DJ-1. The effect of these oxidation processes are difficult to investigate selectively in experiments.Interestingly, our simulations demonstrate for CYS oxidation in the two systems opposite effects on protein folding stability. For the yeast URN1-FF domain CYS-57 oxidation destabilizes and for the DJ-1 CYS-106 oxidation significantly stabilizes the folded state. The results agree with experimental stability measurements on the two protein systems. For the DJ-1 dimer our simulations predict a slight dimer destabilization which is, however, likely outbalanced by the significantly increased folding stability of the monomers due to CYS-106 oxidation. The changes in the interaction pattern due to CYS oxidation seem to be a result of the altered geometry of the side chain and creation of stronger H-bond acceptor and donor features. It was found that for the different CYS oxidation states reorientations of the side chain and surrounding residues are often driven by an increased affinity to polar water. It is likely that the increased affinity to water is a general mechanism of protein destabilization due to CYS oxidation especially for buried CYS residues involved in the hydrophobic core of proteins.
Due to the presence of reactive oxygens numerous proteins in an organism are at risk of being modified by cysteine oxidation. It is likely that such modification results in destabilization as has been indicated in previous studies (e.g. ref. 5). However, this should not mask that cysteine oxidation can also stabilize protein folds and can be of functional nature as in DJ-1 case. Indeed, CYS oxidation can act as a sensor for oxidative stress and due to alteration of fold stability and association can contribute to oxidative stress signaling in cells.5 Future alchemical simulation studies on such systems could be useful to elucidate the molecular mechanism of such CYS oxidation signaling processes.
Author contributions
T. M. performed simulations, wrote programs and scripts, analyzed data, wrote paper draft. M. Z. designed project, supervised computational work, provided computational resources, wrote final manuscript version. T. M. and M. Z. analyzed results, reviewed the manuscript.Conflicts of interest
No competing interest is declared.Data availability
Data for this article including scripts to run and for analysis of the simulations are available at https://doi.org/10.5281/zenodo.18789990. Force field parameters for oxidized cysteine residues and additional tables and figures have been included in the article's supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d6cp00724d.Acknowledgements
The authors thank N. Halbwedl, C. Sustay and Dr M. Kulke and P. Quoika for useful discussions. This work was financially supported by the Deutsche Forschungsgemeinschaft (DFG Za153/29-1) and by local compute cluster (partially funded by DFG INST 95/1610-1 FUGG). We acknowledge additional HPC resources provided by the Erlangen National High Performance Center (NHR@FAU) of the Friedrich-Alexander-Universität Erlangen- Nürnberg, grant b118bb.References
- B. Ezraty, A. Gennaris, F. Barras and J.-F. Collet, Oxidative stress, protein damage and repair in bacteria, Nat. Rev. Microbiol., 2017, 15, 385–396, DOI:10.1038/nrmicro.2017.26.
- D. Garrido Ruiz, A. Sandoval-Perez, A. Vikram Rangarajan, E. L. Gunderson and M. P. Jacobson, Cysteine Oxidation in Proteins: Structure, Biophysics, and Simulation, Biochemistry, 2022, 61, 2165–2176, DOI:10.1021/acs.biochem.2c00349.
- J. M. Held, Redox Systems Biology: Harnessing the Sentinels of the Cysteine Redoxome, Antioxid. Redox Signaling, 2020, 32, 659–676, DOI:10.1089/ars.2019.7725.
- T. H. Truong and K. S. Carroll, Redox regulation of epidermal growth factor receptor signaling through cysteine oxidation, Biochemistry, 2012, 51, 9954–9965, DOI:10.1021/bi301441e.
- P. Marinelli, S. Navarro, R. Graña-Montes, M. Bañó-Polo, M. Rosario Fernández, E. Papaleo and S. Ventura, A single cysteine post-translational oxidation suffices to compromise globular proteins kinetic stability and promote amyloid formation, Redox Biol., 2018, 14, 566–575, DOI:10.1016/j.redox.2017.10.022.
- J. Schymkowitz, J. Borg, F. Stricher, R. Nys, F. Rousseau and L. Serrano, The FoldX web server: an online force field, Nucleic Acids Res., 2005, 33, W382–W388, DOI:10.1093/nar/gki387.
- R. Bonet, X. Ramirez-Espain and M. J. Macias, Solution structure of the yeast URN1 splicing factor FF domain: Comparative analysis of charge distributions in FF domain structures—FFs and SURPs, two domains with a similar fold, Proteins:Struct., Funct., Bioinf., 2008, 73, 1001–1009, DOI:10.1002/prot.22127.
- L. Premkumar, M. K. Dobaczewska and S. J. Riedl, Identification of an artificial peptide motif that binds and stabilizes reduced human DJ-1, J. Struct. Biol., 2011, 176, 414–418, DOI:10.1016/j.jsb.2011.08.011.
- L. D. Skou, S. K. Johansen, J. Okarmus and M. Meyer, Pathogenesis of DJ-1/PARK7-Mediated Parkinson’s Disease, Cells, 2024, 13(4) DOI:10.3390/cells13040296.
- T. Kinumi, J. Kimata, T. Taira, H. Ariga and E. Niki, Cysteine-106 of DJ-1 is the most sensitive cysteine residue to hydrogen peroxide-mediated oxidation in vivo in human umbilical vein endothelial cells, Biochem. Biophys. Res. Commun., 2004, 317, 722–728, DOI:10.1016/j.bbrc.2004.03.110.
- J. Choi, M. Cameron Sullards, J. A. Olzmann, H. D. Rees, S. T. Weintraub, D. E. Bostwick, M. Gearing, A. I. Levey, L.-S. Chin and L. Li, Oxidative Damage of DJ-1 Is Linked to Sporadic Parkinson and Alzheimer Diseases, J. Biol. Chem., 2006, 281, 10816–10824, DOI:10.1074/jbc.M509079200.
- A. C. Witt, M. Lakshminarasimhan, B. C. Remington, S. Hasim, E. Pozharski and M. A. Wilson, Cysteine pKa Depression by a Protonated Glutamic Acid in Human DJ-1, Biochemistry, 2008, 47, 7430–7440, DOI:10.1021/bi800282d.
- R. M. Canet-Avilés, M. A. Wilson, D. W. Miller, R. Ahmad, C. McLendon, S. Bandyopadhyay, M. J. Baptista, D. Ringe, G. A. Petsko and M. R. Cookson, The Parkinson's disease protein DJ-1 is neuroprotective due to cysteine-sulfinic acid-driven mitochondrial localization, Proc. Natl. Acad. Sci. U. S. A., 2004, 101, 9103–9108, DOI:10.1073/pnas.0402959101.
- R. Kiss, M. Zhu, B. Jójárt, A. Czajlik, K. Solti, B. Fórizs, É. Nagy, F. Zsila, T. Beke-Somfai and G. Tóth, Structural features of human DJ-1 in distinct Cys106 oxidative states and their relevance to its loss of function in disease, Biochim. Biophys. Acta, Gen. Subj., 2017, 1861, 2619–2629, DOI:10.1016/j.bbagen.2017.08.017.
- M. J. Frisch, et al.Gaussian09, Gaussian, Inc., Wallingford CT, 2009 Search PubMed.
- C. I. Bayly, P. Cieplak, W. Cornell and P. A. Kollman, A well-behaved electrostatic potential based method using charge restraints for deriving atomic charges: the RESP model, J. Phys. Chem., 1993, 97, 10269–10280, DOI:10.1021/j100142a004.
- D. A. Case, et al., AmberTools, J. Chem. Inf. Model., 2023, 63, 6183–6191, DOI:10.1021/acs.jcim.3c01153.
- J. Wang, R. M. Wolf, J. W. Caldwell, P. A. Kollman and D. A. Case, Development and testing of a general amber force field, J. Comput. Chem., 2004, 25, 1157–1174, DOI:10.1002/jcc.20035.
- J. A. Maier, C. Martinez, K. Kasavajhala, L. Wickstrom, K. E. Hauser and C. Simmerling, ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from ff99SB, J. Chem. Theory Comput., 2015, 11, 3696–3713, DOI:10.1021/acs.jctc.5b00255.
- H. M. Berman, J. Westbrook, Z. Feng, G. Gilliland, T. N. Bhat, H. Weissig, I. N. Shindyalov and P. E. Bourne, The Protein Data Bank, Nucleic Acids Res., 2000, 28, 235–242, DOI:10.1093/nar/28.1.235.
- D. A. Case, T. E. Cheatham III, T. Darden, H. Gohlke, R. Luo, K. M. Merz Jr., A. Onufriev, C. Simmerling, B. Wang and R. J. Woods, The Amber biomolecular simulation programs, J. Comput. Chem., 2005, 26, 1668–1688, DOI:10.1002/jcc.20290.
- J. W. Pitera and P. A. Kollman, Exhaustive mutagenesis in silico: multicoordinate free energy calculations on proteins and peptides, Proteins, 2000, 41, 385–397, DOI:10.1002/1097-0134(20001115)41:3〈385::AID-PROT80〉3.0.CO;2-Q.
- D. A. Case, et al.Amber2023, University of California, San Francisco, 2023 Search PubMed.
- S. Le Grand, A. W. Götz and R. C. Walker, SPFP: Speed without compromise—A mixed precision model for GPU accelerated molecular dynamics simulations, Comput. Phys. Commun., 2013, 184, 374–380, DOI:10.1016/j.cpc.2012.09.022.
- R. Salomon-Ferrer, A. W. Götz, D. Poole, S. Le Grand and R. C. Walker, Routine Microsecond Molecular Dynamics Simulations with AMBER on GPUs. 2. Explicit Solvent Particle Mesh Ewald, J. Chem. Theory Comput., 2013, 9, 3878–3888, DOI:10.1021/ct400314y.
- A. W. Götz, M. J. Williamson, D. Xu, D. Poole, S. Le Grand and R. C. Walker, Routine Microsecond Molecular Dynamics Simulations with AMBER on GPUs. 1. Generalized Born, J. Chem. Theory Comput., 2012, 8, 1542–1555, DOI:10.1021/ct200909j.
- R. Anandakrishnan, B. Aguilar and A. V. Onufriev, H++ 3.0: automating pK prediction and the preparation of biomolecular structures for atomistic molecular modeling and simulations, Nucleic Acids Res., 2012, 40, W537–W541, DOI:10.1093/nar/gks375.
- J. Myers, G. Grothaus, S. Narayanan and A. Onufriev, A simple clustering algorithm can be accurate enough for use in calculations of pKs in macromolecules, Proteins:Struct., Funct., Bioinf., 2006, 63, 928–938, DOI:10.1002/prot.20922.
- J. C. Gordon, J. B. Myers, T. Folta, V. Shoja, L. S. Heath and A. Onufriev, H++: a server for estimating pKa s and adding missing hydrogens to macromolecules, Nucleic Acids Res., 2005, 33, W368–W371, DOI:10.1093/nar/gki464.
- V. Gapsys, S. Michielssens, J. Henning Peters, B. L. de Groot and H. Leonov, Calculation of Binding Free Energies, in Molecular Modeling of Proteins, ed. A. Kukol, Springer New York, New York, NY, 2015, pp. 173–209. isbn: 978-1-4939-1465-4. DOI:10.1007/978-1-4939-1465-49.
- D. Petrov, J. Walther Perthold, C. Oostenbrink, B. L. de Groot and V. Gapsys, Guidelines for Free-Energy Calculations Involving Charge Changes, J. Chem. Theory Comput., 2024, 20, 914–925, DOI:10.1021/acs.jctc.3c00757.
- T. C. Beutler, A. E. Mark, R. C. van Schaik, P. R. Gerber and W. F. van Gunsteren, Avoiding singularities and numerical instabilities in free energy calculations based on molecular simulations, Chem. Phys. Lett., 1994, 222, 529–539, DOI:10.1016/0009-2614(94)00397-1.
- M. Zacharias, T. P. Straatsma and J. A. McCammon, Separation-shifted scaling, a new scaling method for Lennard-Jones interactions in thermodynamic integration, J. Chem. Phys., 1994, 100, 9025–9031, DOI:10.1063/1.466707.
- T. Steinbrecher, I. Joung and D. A. Case, Soft-core potentials in thermodynamic integration: Comparing one- and two-step transformations, J. Comput. Chem., 2011, 32, 3253–3263, DOI:10.1002/jcc.21909.
- J. S. Hub, B. L. de Groot, H. Grubmüller and G. Groenhof, Quantifying Artifacts in Ewald Simulations of Inhomogeneous Systems with a Net Charge, J. Chem. Theory Comput., 2014, 10, 381–390, DOI:10.1021/ct400626b.
- T. Alexander Mauck and M. Zacharias, Influence of methionine oxidation on protein stability and association studied by free energy simulations, J. Mol. Biol., 2025, 169576 Search PubMed.
- D. J. Mermelstein, C. Lin, G. Nelson, R. Kretsch, J. Andrew McCammon and R. C. Walker, Fast and flexible gpu accelerated binding free energy calculations within the amber molecular dynamics package, J. Comput. Chem., 2018, 39, 1354–1358, DOI:10.1002/jcc.25187.
- T.-S. Lee, Y. Hu, B. Sherborne, Z. Guo and D. M. York, Toward Fast and Accurate Binding Affinity Prediction with pmemdGTI: An Efficient Implementation of GPUAccelerated Thermodynamic Integration, J. Chem. Theory Comput., 2017, 13, 3077–3084, DOI:10.1021/acs.jctc.7b00102.
- H. Flyvbjerg and H. G. Petersen, Error estimates on averages of correlated data, J. Chem. Phys., 1989, 91, 461–466, DOI:10.1063/1.457480.
- A. S. J. S. Mey, et al., Best Practices for Alchemical Free Energy Calculations [Article v1.0], Living J. Comput. Mol. Sci., 2020, 2, 18378, DOI:10.33011/livecoms.2.1.18378.
- D. R. Roe and T. E. Cheatham III, PTRAJ and CPPTRAJ: Software for Processing and Analysis of Molecular Dynamics Trajectory Data, J. Chem. Theory Comput., 2013, 9, 3084–3095, DOI:10.1021/ct400341p.
- E. Alexov, et al., Progress in the prediction of pKa values in proteins, Proteins:Struct., Funct., Bioinf., 2011, 79, 3260–3275, DOI:10.1002/prot.23189.
| This journal is © the Owner Societies 2026 |




