 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Graph neural network architectures for predicting the electrophilicity index: insights from 2D and 3D molecular graph representations
Sandeep Kumar *a and
Annika Bande
*a and
Annika Bande ab
ab
aHelmholtz-Zentrum Berlin für Materialien und Energie GmbH, Hahn-Meitner-Platz 1, 14109 Berlin, Germany. E-mail: sandeep.kumar@helmholtz-berlin.de; annika.bande@helmholtz-berlin.de
bLeibniz University Hannover, Institute of Inorganic Chemistry, Callinstr. 9, 30167 Hannover, Germany
First published on 5th May 2026
Abstract
The electrophilicity index (ω) is a fundamental quantum chemical descriptor that quantifies a molecule's ability to accept electrons, playing a critical role in assessing reactivity and stability, and guiding molecular design. In this study, we benchmark a diverse set of graph neural network (GNN) architectures for predicting ω, using data derived from the QM9 dataset of organic molecules. As ω is a global quantum-chemical property derived from frontier molecular orbital energies, typically sensitive to the local chemical structure, we compare 3D molecular geometry models (SchNet, ALIGNN, GemNet), which account for the full atomic structure, with connectivity-based 2D models (attentive FP, GCN, GraphSAGE, GIN, GINE, GATv2) that consider only molecular topology. The results indicate that ω depends not only on molecular topology but also on the complete 3D atomic arrangement, as reflected by the superior predictive accuracy of 3D models—particularly ALIGNN. Nevertheless, some 2D GNNs provide a computationally efficient alternative. Notably, GINE achieves more than an order-of-magnitude reduction in training time compared to ALIGNN, while exhibiting only about a one percent-level decrease in predictive accuracy.
I. Introduction
The concept of electrophilicity plays a fundamental role in understanding chemical reactivity and molecular stability, particularly in the context of frontier molecular orbital theory.1,2 The electrophilicity index (ω), introduced by Parr et al.,3 quantifies the tendency of a molecule to accept electrons, making it an essential descriptor in theoretical chemistry. It is particularly valuable for understanding molecular reactivity in electrophilic interactions and thus for practical applications such as drug design4 and catalysis.5 The electrophilicity index is defined as:3
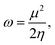 | (1) |
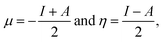 | (2) |
According to Koopmans’ theorem,6 the ionization potential and electron affinity can be approximated from the highest occupied molecular orbital (HOMO) and lowest unoccupied molecular orbital (LUMO) energies, respectively. Using the expressions for μ and η from eqn (2), ω can be written in a unified form as
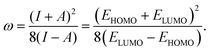 | (3) |
Although ω is defined in terms of the HOMO and LUMO energies, EHOMO and ELUMO, it is a nonlinear function of these quantities, involving both their sum and difference at different exponents in a ratio form. As a result, small errors in the HOMO or LUMO energies can propagate nonlinearly and lead to disproportionately large deviations in ω. Physically, the ω values, being a global reactivity descriptor derived from frontier molecular orbital energies, are particularly sensitive to both local atomic features and long-range interactions. This behavior distinguishes ω from more commonly predicted quantum-chemical properties and renders its direct prediction from the molecular structure a more challenging learning task than reconstructing it from individual orbital energies.
Moreover, the expression in eqn (3) relies on Koopmans' theorem, which assumes that molecular orbitals do not relax upon electron removal or addition. However, a common standard methodology to quantum chemistry calculations for larger molecular structures is density functional theory (DFT).7,8 Among the available functionals, only recent hybrid, range-separated, or double-hybrid functionals provide reliable estimates of frontier orbital energies and related reactivity descriptors.9,10 But such calculations are computationally demanding and are therefore typically limited to relatively small reference datasets. This limitation restricts their applicability in large-scale molecular screening and high-throughput studies.11–18
Consequently, there is growing interest in alternative approaches that can approximate quantum chemical properties at significantly reduced computational cost. Indeed, more recently, ω has gained renewed attention in the application of machine-learning models,19,20 which are likely to deliver predictions faster than DFT.
In this context, data-driven methods such as graph neural networks (GNNs) have emerged as a promising solution, enabling learning directly from existing molecular datasets. By leveraging molecular graph representations, GNNs offer a fast, scalable, and accurate alternative for predicting ω, bypassing the need for expensive quantum chemical calculations. Recent advances in machine learning, particularly the development of GNNs, have enabled the modeling of molecular properties directly from graph-based representations of chemical structures.21–43 Molecules can naturally be encoded as graphs, with atoms as nodes and bonds as edges, which makes GNNs especially well-suited for molecular property prediction. These models learn complex, high-dimensional relationships between structure and properties without the need for handcrafted features or expensive quantum mechanical calculations.
In this work, we investigate the use of GNNs to predict ω using data from the QM9 database,44 a widely used benchmark dataset that contains quantum chemical properties of approximately 134![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 small organic molecules calculated using DFT at the B3LYP/6-31G(2df,p) level of theory. Instead of computing ω directly through DFT for new molecules, we aim to build predictive models that leverage the existing QM9 data to generalize to unseen compounds. Previous studies have applied machine-learning models based on molecular descriptors to estimate electrophilicity-related quantities;19,45 however, these approaches do not fully leverage molecular graph structure. Moreover, a systematic comparison of 2D and 3D GNNs for direct prediction of the conceptual DFT ω remains absent. Here, we conduct a systematic comparative evaluation of 2D and 3D GNN architectures within a unified training and evaluation framework, quantifying the relative contributions of molecular topology and explicit geometric information, as well as the associated accuracy–computational cost trade-off. To the best of our knowledge, this is the first study to directly predict the conceptual DFT ω using GNNs.
000 small organic molecules calculated using DFT at the B3LYP/6-31G(2df,p) level of theory. Instead of computing ω directly through DFT for new molecules, we aim to build predictive models that leverage the existing QM9 data to generalize to unseen compounds. Previous studies have applied machine-learning models based on molecular descriptors to estimate electrophilicity-related quantities;19,45 however, these approaches do not fully leverage molecular graph structure. Moreover, a systematic comparison of 2D and 3D GNNs for direct prediction of the conceptual DFT ω remains absent. Here, we conduct a systematic comparative evaluation of 2D and 3D GNN architectures within a unified training and evaluation framework, quantifying the relative contributions of molecular topology and explicit geometric information, as well as the associated accuracy–computational cost trade-off. To the best of our knowledge, this is the first study to directly predict the conceptual DFT ω using GNNs.
To predict the ω efficiently from molecular graph representations, we employ nine widely used GNN architectures, grouped into 2D and 3D molecular graphs, selected for their proven effectiveness in quantum chemistry and molecular property prediction tasks. All models were trained on ω values derived from the QM9 dataset using eqn (3), with atomic and bond features extracted directly from the corresponding molecular structures. The 2D GNN models considered in this study include graph convolutional network (GCN),46 attentive fingerprints (AFP),47 graph isomorphism network (GIN),48 graph isomorphism network with edge features (GINE),49 graph attention network v2 (GATv2),50 and graph sample and aggregate (GraphSAGE),51 all operating on graphs constructed from SMILES representations.52 To capture geometric effects, we also employ 3D GNN architectures—SchNet,53 atomistic line graph neural network (ALIGNN),26 and geometric message passing network (GemNet)54—which incorporate spatial information such as atomic coordinates and interatomic distances. Together, these models provide a comprehensive framework for evaluating the relative importance of topological and geometric molecular information in predicting ω. Importantly, the models are transferable and can be fine-tuned to higher-quality or experimental datasets via transfer learning.55
It is important to emphasize that the electrophilicity index ω considered in this work is a conceptual density functional theory descriptor derived from frontier molecular orbital energies rather than a directly measurable experimental observable. Consequently, the aim of this study is not to reproduce experimental electrophilicity scales, but rather to investigate whether modern GNN architectures can learn the nonlinear relationship between molecular structure and this theoretical reactivity descriptor. Predicting such descriptors using machine learning can significantly accelerate computational chemistry workflows by providing rapid estimates of quantum-chemical properties without performing explicit electronic-structure calculations. This capability is particularly useful for large-scale molecular screening and high-throughput exploration of chemical space.
The structure of the paper is organized as follows: Section II describes the datasets and exploratory data analysis, the graph-based representation of molecular structures, details of the GNN models and their training procedures, hyperparameter optimization strategies, and the evaluation metrics used for predicting the ω. Section III presents and discusses the results, starting with model performance on the regression tasks. Finally, Section IV summarizes the key findings and main conclusions of the study. Additional results and analyses are provided in the SI, available as a separate file.
II. Methods
A. Dataset overview and preliminary analysis
As introduced earlier, the database of ω employed in this work was constructed from the QM9 dataset. To reveal insights into the dataset composition, the distribution of molecule counts by atom type within the dataset is shown in Fig. S1(a). Carbon is the most prevalent element (133![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 722 molecules), followed by oxygen (113
722 molecules), followed by oxygen (113![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 853), nitrogen (82
853), nitrogen (82![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 699), and fluorine (2120). This composition reflects the dominance of common organic elements, with relatively few fluorinated molecules, consistent with trends typically observed in drug-like and organic chemistry datasets.
699), and fluorine (2120). This composition reflects the dominance of common organic elements, with relatively few fluorinated molecules, consistent with trends typically observed in drug-like and organic chemistry datasets.
Furthermore, to analyze the statistical characteristics of the data, we plotted the distribution of ω values as a histogram in Fig. 1 (left). The resulting distribution is strongly right-skewed, with the majority of values clustered near zero and a long tail extending toward higher ω. Skewness, which quantifies the asymmetry of a distribution, confirms this deviation from normality. While a normal distribution is symmetric with zero skewness, the observed right-skew indicates that the mean of the ω values lies above the median, reflecting the presence of relatively few molecules with exceptionally large ω values. The overlaid red density curve56 further emphasizes the asymmetry and lack of normality in the raw data. This indicates that the majority of molecules have low electrophilic reactivity, consistent with the small, DFT-optimized organic molecules present in the QM9 database. The long right tail represents a minority of highly reactive molecules and makes the distribution less suitable for standard statistical modeling and visualization.
 | ||
| Fig. 1 Distribution of electrophilicity index (ω) values. (left) Raw values showing a right-skewed distribution. (right) Log-transformed (ωlog) values approximating a normal distribution. | ||
To address this skewness, a logarithmic transformation was applied to the ω values,
| ωlog = log(ω + ε). | (4) |
To prevent numerical instabilities caused by log(0) and extremely small ω values, a small constant ε = 1 × 10−6 was added prior to the logarithmic transformation. This standard stabilization technique ensures well-defined gradients and smooth training behavior.57,58 The logarithmic transformation compresses large values while expanding the range of small ones, resulting in a more symmetric, approximately Gaussian distribution. In addition, it also improves visualization by revealing subtle differences among low-reactivity molecules, stabilizes variance, and facilitates regression or classification tasks using GNNs. Fig. 1 (right) presents the distribution of the log-transformed ω values, ωlog. The resulting histogram appears markedly more symmetric.
To check how well the ω agrees with its log-transformed data ωlog, Spearman's rank correlation (ρ) was calculated59 (Fig. S1(b)). The perfect alignment of points along the diagonal (ρ = 1.0) confirms that the relative ordering of molecules by their ω values remains unchanged after logarithmic transformation. These results demonstrate that logarithmic transformation of ω preserves the underlying chemical relationships among molecules while improving the statistical uniformity of the data for reliable modeling.
B. Machine-learning workflow
The workflow for predicting ω using GNNs is illustrated in Fig. 2, and the individual steps are described below.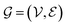 , where the set of nodes
, where the set of nodes 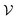 corresponds to the atoms in the molecule and the set of edges
corresponds to the atoms in the molecule and the set of edges 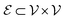 represents the chemical bonds connecting these atoms. In this framework, each atom is treated as a node and each bond as an edge, allowing GNNs to naturally capture both the structural connectivity and chemical properties of molecules.60
represents the chemical bonds connecting these atoms. In this framework, each atom is treated as a node and each bond as an edge, allowing GNNs to naturally capture both the structural connectivity and chemical properties of molecules.60Molecular structures are encoded using SMILES strings52 and parsed with RDKit61 to construct 2D molecular graphs. For 3D GNN models, atomic coordinates provided in QM9 are used directly to build geometry-aware graph representations. By representing molecules as graphs, GNNs can effectively learn intricate structural patterns and chemical interactions, capturing both local atomic neighborhoods and the global connectivity of the molecular graph. In the resulting graph representation, atoms correspond to nodes and chemical bonds correspond to edges. Node features include atomic number, valence, hybridization state, aromaticity, and formal charge, while edge features encode bond type, conjugation, and ring membership, as derived from the molecular structure using RDKit. For geometry-aware architectures, additional spatial information derived from the atomic coordinates, such as interatomic distances and bond-angle relationships, is incorporated to capture three-dimensional structural interactions, as illustrated in Fig. 3.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 725 molecular structures is randomly shuffled and split into 80% for training (106
725 molecular structures is randomly shuffled and split into 80% for training (106![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 980 samples) and 20% for testing (26
980 samples) and 20% for testing (26![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 745 samples). This split ensures a sufficiently large training set while reserving an independent test set for unbiased performance assessment.
745 samples). This split ensures a sufficiently large training set while reserving an independent test set for unbiased performance assessment.Hyperparameter optimization is carried out using the Optuna framework,62 which combines random sampling with adaptive pruning to systematically explore the hyperparameter space and identify optimal training configurations for each GNN architecture. Final models are trained using the Adam optimizer,63 with early stopping applied based on validation loss to prevent overfitting.64 All models are implemented using the PyTorch framework65 and PyTorch-Geometric.66 The hyperparameters for both 3D and 2D GNN models were carefully optimized to ensure stable training and accurate prediction of ω. For 3D models (GemNet, SchNet, ALIGNN), training employed small-to-moderate batch sizes (8–32) and low learning rates (10−4–10−3) to ensure stable convergence. Hidden dimensions were high for GemNet and ALIGNN (256), while SchNet used a lower-dimensional embedding (64) with 256 filters. Model depth ranged from 4 to 5 layers, and regularization was applied via weight decay (10−6–10−5). The number of epochs (105–300) reflects differing convergence rates across architectures.
For 2D models (GINE, GIN, GATv2, GraphSAGE, AFP, GCN), batch sizes (32–128) and learning rates (10−5–10−3) varied more widely, indicating architecture-dependent optimization sensitivity. Deeper networks (up to 6 layers) were used for GIN and GATv2, while others employed 4–5 layers. Hidden dimensions remained consistently large (192–256), and regularization included weight decay (10−6–10−5) and model-specific dropout (0.0–0.2). Overall, the selected configurations balance model complexity, convergence stability, and generalization across diverse GNN architectures. The optimized hyperparameters for all architectures are summarized in Tables S1 and S2
III. Results and discussion
A. Insights from interpreting 2D and 3D molecular graphs
In the following section, we analyze the performance of the 2D and 3D models based on evaluation metrics computed from the log-transformed data, with results converted back to the original ω values.To illustrate the comparative performance of 2D models, Fig. 4 and 5 showcase six GNN models—GINE, attentive FP, GIN, GATv2, GraphSAGE, and GCN—in predicted versus actual ω values from molecular structures. In each figure, the left panels show how well the predicted values match the actual values, while the right panels show the prediction errors (residuals). Overall, all six 2D models perform well, with R2 values above 0.95 for both training (blue points) and test sets (rose points), which means they can capture the relationship between molecular structure and ω values accurately.
 | ||
| Fig. 4 Left panels depict predicted versus actual ω values, while right panels present the residuals plotted against the actual ω values for the GINE, AFP, and GIN models (top to bottom). | ||
 | ||
| Fig. 5 Left panels depict predicted versus actual ω values, while right panels present the residuals plotted against the actual ω values for GATv2, GraphSAGE, and GCN models (top to bottom). | ||
Among the models in Fig. 4, GINE (top) gave the best performance (Rtest2 = 0.9816), followed closely by GIN (Rtest2 = 0.9779, bottom) and attentive FP (Rtest2 = 0.9772, center). These models are especially effective because they account for detailed molecular features during message passing. A minor reduction in R2 is observed when converting predictions from the log-transformed scale back to the original scale, which is expected due to the non-linear nature of the inverse transformation, leading to slightly lower values in the original scale. The performance of all 2D models on the log-transformed data is shown in Fig. S2 and S3 of the SI.
The remaining three 2D models, shown in Fig. 5, also demonstrate strong performance. GATv2 achieved the highest accuracy in this group (Rtest2 = 0.9732), followed by GraphSAGE (Rtest2 = 0.9667) and GCN (Rtest2 = 0.9571). The residual plots on the right side show that most prediction errors are very small and centered around zero, though there is a slightly larger spread of errors for higher ω values. The GCN model, in particular, shows a wider distribution of residuals, which explains its slightly lower performance compared to the other models. In summary, all the GNN architectures tested can successfully predict the ω values, with some models achieving higher accuracy than others. Comparing their performance on the test set, GINE outperformed all other models, GIN and attentive FP performed competitively, GATv2 and GraphSAGE achieved moderate accuracy, while GCN ranked lowest among the six architectures.
To gain further insight, we performed 5-fold cross-validation for all six 2D GNN models. The results, summarized in Table 1, include both a single train/test split and 5-fold CV. The models are ranked from best to worst based on the CV RMSE. For the single train/test split, GINE achieved the lowest test MAE (0.0015) and RMSE (0.0028), along with the highest test R2 (0.9816), indicating superior predictive accuracy. In contrast, GCN showed the poorest performance, with the highest test MAE (0.0024) and RMSE (0.0042) and the lowest R2 (0.9571). The close agreement between training and test metrics across all models suggests minimal overfitting. The 5-fold cross-validation results confirm these trends.
| Metric | GINE | AFP | GIN | GATv2 | GraphSAGE | GCN |
|---|---|---|---|---|---|---|
| Train/test split | ||||||
| Epochs | 214 | 70 | 135 | 215 | 300 | 180 |
| Train MAE | 0.0013 | 0.0017 | 0.0015 | 0.0018 | 0.0019 | 0.0023 |
| Test MAE | 0.0015 | 0.0017 | 0.0017 | 0.0019 | 0.0020 | 0.0024 |
| Train RMSE | 0.0022 | 0.0031 | 0.0026 | 0.0032 | 0.0035 | 0.0041 |
| Test RMSE | 0.0028 | 0.0030 | 0.0030 | 0.0033 | 0.0037 | 0.0042 |
| Train R2 | 0.9875 | 0.9763 | 0.9836 | 0.9741 | 0.9693 | 0.9587 |
| Test R2 | 0.9816 | 0.9772 | 0.9779 | 0.9732 | 0.9667 | 0.9571 |
| 5-Fold cross-validation (mean ± std) | ||||||
| MAE | 0.0017 ± 0.0002 | 0.0018 ± 0.0001 | 0.0021 ± 0.0001 | 0.0023 ± 0.0002 | 0.0024 ± 0.0001 | 0.0026 ± 0.0002 |
| RMSE | 0.0030 ± 0.0002 | 0.0034 ± 0.0002 | 0.0037 ± 0.0002 | 0.0042 ± 0.0003 | 0.0044 ± 0.0003 | 0.0046 ± 0.0002 |
| R2 | 0.9780 ± 0.0034 | 0.9711 ± 0.0025 | 0.9656 ± 0.0044 | 0.9564 ± 0.0069 | 0.9529 ± 0.0066 | 0.9480 ± 0.0038 |
GINE again outperformed the other models, achieving the lowest CV RMSE (0.0030 ± 0.0002) and the highest CV R2 (0.9780 ± 0.0034), followed by AFP and GIN. GATv2, GraphSAGE, and GCN showed progressively higher CV errors and lower R2, indicating reduced generalization performance. MSE values are near zero for all models, reflecting high prediction precision. Overall, GINE provides the most accurate and robust predictions for ω on the original scale, while GCN consistently exhibits the lowest performance among the architectures tested.
In Fig. S4(a) and (b), the per-fold R2 scores are reported, showing both the average predictive performance and consistency of each model across cross-validation splits, providing insight into robustness and generalizability. GINE achieves the highest R2 in every fold, demonstrating superior performance, while other models show lower mean R2 and higher variability, indicating less reliable predictions. The plot visually conveys both central tendency and stability of each model.
For 3D molecular graph representations, Fig. 6 illustrates the performance of three geometry-aware GNNs—GemNet, SchNet, and ALIGNN—in predicting ω values. As with the 2D models, the left panels show predicted versus actual ω values for both training and test sets, while the right panels display the residuals plotted against the actual values.
All three models demonstrate excellent predictive accuracy, with test set R2 values exceeding 0.98, confirming that 3D-aware message-passing architectures are highly effective in capturing the structure–property relationships governing the target ω values. Among the evaluated models, ALIGNN achieves the strongest performance (Rtest2 = 0.9941), highlighting the advantage of incorporating higher-order geometric information through line-graph message passing. SchNet follows closely (Rtest2 = 0.9883), benefiting from continuous-filter convolutions that encode smooth distance-dependent interactions, while GemNet also performs competitively (Rtest2 = 0.9882), though with slightly larger residual variations at higher ω values.
The residual plots show that the majority of prediction errors for all 3D models are tightly clustered around zero, indicating stable and unbiased generalization across the full range of ω values. Compared to standard 2D GNN baselines, the 3D models achieve consistently higher or comparable accuracy, suggesting that explicit incorporation of molecular geometry provides a measurable performance advantage. However, incorporating 3D structural information improves model performance, comparable ω predictions can still be obtained even without explicitly using 3D geometry. Within the family of 3D methods, ALIGNN emerges as the top-performing architecture, followed by SchNet and GemNet, demonstrating the benefit of increasingly expressive geometric message-passing schemes for learning molecular reactivity descriptors. In addition, results for 3D models applied to the log-transformed dataset are presented in Fig. S5.
Besides R2, we also examine additional metrics such as MAE and RMSE to provide a more comprehensive evaluation of model performance, as summarized in Table 2. To obtain a more reliable assessment, we compare all 3D models using five-fold cross-validation (5-CV). Based on the CV results, ALIGNN achieves the lowest prediction errors, with a MAE of 0.0009 ± 0.0002 and an RMSE of 0.0020 ± 0.0003, indicating superior accuracy and robustness across folds. GemNet follows with an MAE of 0.0013 ± 0.0000 and an RMSE of 0.0026 ± 0.0002, while SchNet shows slightly higher errors, with an MAE of 0.0015 ± 0.0002 and an RMSE of 0.0027 ± 0.0002. These results confirm that incorporating directional and geometric information, particularly in ALIGNN, leads to improved performance in ω prediction.
| Metric | ALIGNN | GemNet | SchNet |
|---|---|---|---|
| Train/test split (original scale) | |||
| Epochs | 300 | 163 | 105 |
| Train MAE | 0.0007 | 0.0008 | 0.0010 |
| Test MAE | 0.0009 | 0.0011 | 0.0011 |
| Train RMSE | 0.0011 | 0.0013 | 0.0017 |
| Test RMSE | 0.0016 | 0.0022 | 0.0022 |
| Train R2 | 0.9968 | 0.9957 | 0.9929 |
| Test R2 | 0.9941 | 0.9882 | 0.9883 |
| 5-Fold cross-validation (mean ± std, original scale) | |||
| MAE | 0.0009 ± 0.0002 | 0.0013 ± 0.0000 | 0.0015 ± 0.0002 |
| RMSE | 0.0020 ± 0.0003 | 0.0026 ± 0.0002 | 0.0027 ± 0.0002 |
| R2 | 0.9897 ± 0.0033 | 0.9830 ± 0.0021 | 0.9823 ± 0.0034 |
As with the 2D models, the per-fold R2 values for the 3D GNNs are presented in Fig. S4(c) and (d). ALIGNN consistently attains the highest R2 across all folds with minimal variance, demonstrating both excellent predictive accuracy and stable generalization. GemNet and SchNet also achieve high and relatively consistent R2 values, confirming the robustness of geometry-aware message-passing approaches for molecular property prediction.
The improved performance of geometry-aware models highlights the importance of incorporating spatial information when predicting electronic descriptors. While 2D GNNS capture molecular connectivity, they do not explicitly encode geometric relationships between atoms. In contrast, 3D architectures incorporate interatomic distances and angular interactions that influence the electronic structure of the molecule. The ALIGNN model further extends this representation by introducing a line graph that enables message passing over bond–angle relationships, allowing the network to capture higher-order geometric interactions that are relevant for ω prediction.
B. Functional group influence on electrophilicity index
To gain further chemical insight into the factors influencing ω, we analyzed the distribution of ω values across representative functional group categories identified using SMARTS (SMiles ARbitrary target specification)70–based substructure matching implemented in RDKit. The resulting distributions are shown in Fig. S6. Molecules containing strongly electron-withdrawing substituents, such as carbonyl (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) or nitrile (C
O) or nitrile (C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) N) groups, tend to exhibit larger ω values because these groups stabilize the LUMO energy and thereby enhance the electron-accepting ability of the molecule. These trends can be understood in terms of frontier molecular orbital theory, where electron-withdrawing substituents lower the LUMO energy and thereby increase ω. In contrast, hydrocarbon frameworks composed solely of carbon and hydrogen atoms generally exhibit lower ω values due to the absence of strongly electronegative substituents. Functional groups such as alcohols, ethers, and amines exhibit intermediate behavior because their heteroatoms influence the electronic structure but do not withdraw electron density as strongly as carbonyl-containing groups.
N) groups, tend to exhibit larger ω values because these groups stabilize the LUMO energy and thereby enhance the electron-accepting ability of the molecule. These trends can be understood in terms of frontier molecular orbital theory, where electron-withdrawing substituents lower the LUMO energy and thereby increase ω. In contrast, hydrocarbon frameworks composed solely of carbon and hydrogen atoms generally exhibit lower ω values due to the absence of strongly electronegative substituents. Functional groups such as alcohols, ethers, and amines exhibit intermediate behavior because their heteroatoms influence the electronic structure but do not withdraw electron density as strongly as carbonyl-containing groups.
The relatively broad distributions observed for some functional group categories can be attributed to the diversity of molecular environments in which these groups occur. In many molecules, multiple functional groups coexist and may exert competing electronic effects on the frontier orbital energies. For example, electron-withdrawing substituents may appear together with electron-donating groups, producing a wider range of ω values. This effect is particularly evident in molecules containing mixed heteroatoms, where combinations of oxygen- and nitrogen-containing groups generate a variety of electronic environments.
C. Computational cost of 2D and 3D GNN models
In summary, the choice between 2D and 3D GNN models should be determined by the specific requirements and constraints of the study. When high predictive accuracy and molecular detail are essential—such as in the precise prediction of ω—and reliable 3D structural information is available, 3D GNN models are preferable due to their superior capacity to capture spatial molecular features. However, 3D models demand significantly more computational resources: for example, ALIGNN, GemNet, and SchNet required approximately 3525, 333, and 188 minutes of training, in contrast to the 2D models, GINE, GATv2, GraphSAGE, GCN, GIN, and attentive FP, which were trained in roughly 224, 183, 127, 48, 39, and 21 minutes, respectively. This high computational cost of ALIGNN arises from its coupled message passing on both the atom graph and its line graph, where bonds are treated as nodes and bond pairs encode angular (three-body) interactions, substantially increasing graph size, message-passing operations, and memory usage.26 Note that all model training was conducted on a single NVIDIA RTX 5000 GPU equipped with 32 GB of memory.When computational efficiency, scalability, or the lack of accurate 3D conformations is a concern, 2D GNN models provide a robust and efficient alternative that remains broadly applicable. Therefore, using 3D models when high accuracy is required and 2D models when computational efficiency or scalability is important, provides a practical and balanced approach for predicting molecular properties. All execution times for 2D and 3D GNNs are shown in Fig. 7.
IV. Conclusion
In this study, we performed a comprehensive evaluation of graph neural network (GNN) architectures for predicting the electrophilicity index (ω), a key quantum chemical descriptor related to molecular reactivity. We benchmarked both 2D models—such as GINE, GIN, attentive FP, GraphSAGE, GATv2, and GCN—that utilize molecular graph connectivity, and 3D models—GemNet, SchNet, and ALIGNN—that incorporate spatial features including atomic coordinates, distances, and angles. To the best of our knowledge, this work represents the first study to directly predict the conceptual DFT ω using GNNs.Our results show that while 3D GNNs generally achieve superior accuracy and consistency due to their explicit exploitation of molecular geometry, several 2D models attain competitive performance, highlighting their value for efficient and scalable prediction, especially when 3D structural data are unavailable or costly to obtain. Among the 2D models, GINE outperforms all other architectures, while among the 3D models, ALIGNN achieves the best performance, followed by SchNet and GemNet.
Overall, our study provides a robust foundation for selecting and applying GNN models tailored to molecular property prediction tasks, balancing accuracy and computational feasibility across the chemical space represented by the QM9 dataset. While the present study focuses on small organic molecules from QM9, extending these conclusions to larger and more chemically diverse systems warrants further investigation. Nevertheless, the present results demonstrate that nonlinear conceptual DFT descriptors can be accurately predicted without explicit quantum calculations, enabling scalable, data-driven reactivity modeling.
Author contributions
Sandeep Kumar: conceptualization, methodology, software, data curation, formal analysis, writing – original draft; Annika Bande: conceptualization, supervision, funding acquisition, writing – review & editing.Conflicts of interest
There are no competing interests to declare.Data availability
Supplementary information (SI) includes additional figures, tables, and supporting analyses for this work. See DOI: https://doi.org/10.1039/d6cp00677a.The full code used in this work for creating, training, and analyzing the models is publicly available and can be found at https://github.com/sandeeplu/gnnepi, along with the derived dataset of the electrophilicity index (ω) from QM9.
Acknowledgements
S. K. and A. B. would like to acknowledge our ML subgroup members at Helmholtz-Zentrum Berlin (HZB) and Leibniz University Hannover for their valuable discussions and feedback. The authors also thank Mr Martin Ulrich (HZB) for his technical support in the setup and configuration of the GPU workstation, which was essential for the computational aspects of this work.References
- K. Fukui, T. Yonezawa and H. Shingu, A Molecular Orbital Theory of Reactivity in Aromatic Hydrocarbons, J. Chem. Phys., 1952, 20, 722–725 CrossRef CAS.
- K. Fukui, Recognition of Stereochemical Paths by Orbital Interaction, Acc. Chem. Res., 1971, 4, 57–64 CrossRef CAS.
- R. G. Parr, L. V. Szentpaly and S. Liu, Electrophilicity index, J. Am. Chem. Soc., 1999, 121, 1922–1924 CrossRef CAS.
- L. R. Domingo, M. Ríos-Gutiérrez and P. Pérez, Molecular electron density theory: a modern view of reactivity in organic chemistry, Molecules, 2016, 21, 748 CrossRef PubMed.
- P. Geerlings, F. De Proft and W. Langenaeker, Conceptual density functional theory: chemical hardness, softness, and electrophilicity, Chem. Rev., 2003, 103, 1793–1874 CrossRef CAS PubMed.
- T. Koopmans, Über die Zuordnung von Wellenfunktionen und Eigenwerten zu den einzelnen Elektronen eines Atoms, Physica, 1934, 1, 104–113 CrossRef.
- P. Hohenberg and W. Kohn, Inhomogeneous Electron Gas, Phys. Rev., 1964, 136, B864–B871 CrossRef.
- W. Kohn and L. J. Sham, Self-Consistent Equations Including Exchange and Correlation Effects, Phys. Rev., 1965, 140, A1133–A1138 CrossRef.
- M. Bursch, J.-M. Mewes, A. Hansen and S. Grimme, Best-practice DFT protocols for basic molecular computational chemistry, Angew. Chem., 2022, 134, e202205735 CrossRef.
- G. Santra, R. Calinsky and J. M. Martin, Benefits of range-separated hybrid and doublehybrid functionals for a large and diverse data set of reaction energies and barrier heights, J. Phys. Chem. A, 2022, 126, 5492–5505 CrossRef CAS PubMed.
- L. A. Curtiss, K. Raghavachari, P. C. Redfern and J. A. Pople, Assessment of Gaussian-3 and density functional theories for a larger experimental test set, J. Chem. Phys., 2000, 112, 1125–1132 CrossRef CAS.
- P. K. Chattaraj and S. Giri, Electrophilicity index within a conceptual DFT framework, Annu. Rep. Prog. Chem., Sect. C: Phys. Chem., 2009, 105, 13–39 RSC.
- F. Pereira, D. A. Latino and J. Aires-de Sousa, Estimation of mayr electrophilicity with a quantitative structure–property relationship approach using empirical and DFT descriptors, J. Org. Chem., 2011, 76, 9312–9319 CrossRef CAS PubMed.
- F. Jensen, Introduction to computational chemistry, John Wiley & Sons, 2017 Search PubMed.
- B. Lee, J. Yoo and K. Kang, Predicting the chemical reactivity of organic materials using a machine-learning approach, Chem. Sci., 2020, 11, 7813–7822 RSC.
- M. Tavakoli, A. Mood, D. Van Vranken and P. Baldi, Quantum mechanics and machine learning synergies: graph attention neural networks to predict chemical reactivity, J. Chem. Inf. Model., 2022, 62, 2121–2132 CrossRef CAS PubMed.
- R. Pal and P. K. Chattaraj, Electrophilicity index revisited, J. Comput. Chem., 2023, 44, 278–297 CrossRef CAS PubMed.
- M. Ríos-Gutiérrez, A. Saz Sousa and L. R. Domingo, Electrophilicity and nucleophilicity scales at different DFT computational levels, J. Phys. Org. Chem., 2023, 36, e4503 CrossRef.
- Y. Liu, Q. Yang, J. Cheng, L. Zhang, S. Luo and J.-P. Cheng, Prediction of Nucleophilicity and Electrophilicity Based on a Machine-Learning Approach, ChemPhysChem, 2023, 24, e202300162 CrossRef CAS PubMed.
- S. Barman and U. Sarkar, Prediction of Chemical Reactivity Parameters via Data-Driven Approach, Adv. Theory Simul., 2025, 2401517 CrossRef CAS.
- F. Scarselli, et al., The graph neural network model, IEEE Trans. Neural Networks, 2009, 20, 61–80 Search PubMed.
- D. K. Duvenaud, D. Maclaurin, J. Aguilera-Iparraguirre, R. Gomez-Bombarelli, T. Hirzel, A. Aspuru-Guzik and R. P. Adams, Convolutional networks on graphs for learning molecular fingerprints, Adv. Neural Inf. Process. Syst., 2015, 2224–2232 Search PubMed.
- J. Gilmer, S. S. Schoenholz, P. F. Riley, O. Vinyals and G. E. Dahl, Neural message passing for quantum chemistry, Proceedings of the 34th International Conference on Machine Learning (ICML), 2017, vol. 70, pp. 1263–1272 Search PubMed.
- T. Xie and J. C. Grossman, Crystal graph convolutional neural networks for an accurate and interpretable prediction of material properties, Phys. Rev. Lett., 2018, 120, 145301 CrossRef CAS PubMed.
- Y. Rong, Y. Bian, T. Xu, W. Xie, Y. Wei, W. Huang and J. Huang, Self-supervised graph transformer on large-scale molecular data, Adv. Neural Inf. Process. Syst., 2020, 33, 12559–12571 Search PubMed.
- K. Choudhary and B. DeCost, Atomistic line graph neural network for improved materials property predictions, npj Comput. Mater., 2021, 7, 185 CrossRef.
- K. Singh, et al., Machine learning in computational chemistry, J. Chem. Theory Comput., 2022, 18, 4408–4417 CrossRef CAS PubMed.
- W. Nie, D. Liu, S. Li, H. Yu and Y. Fu, Nucleophilicity prediction using graph neural networks, J. Chem. Inf. Model., 2022, 62, 4319–4328 CrossRef CAS PubMed.
- P. Reiser, M. Neubert, A. Eberhard, L. Torresi, C. Zhou, C. Shao, H. Metni, C. van Hoesel, H. Schopmans and T. Sommer, et al., Graph neural networks for materials science and chemistry, Commun. Mater., 2022, 3, 93 CrossRef CAS PubMed.
- Y. Wang, J. Wang, Z. Cao and A. Barati Farimani, Molecular contrastive learning of representations via graph neural networks, Nat. Mach. Intell., 2022, 4, 279–287 CrossRef.
- X. Fang, L. Liu, J. Lei, D. He, S. Zhang, J. Zhou, F. Wang, H. Wu and H. Wang, Geometry-enhanced molecular representation learning for property prediction, Nat. Mach. Intell., 2022, 4, 127–134 CrossRef.
- A. Kotobi, et al., GNNs for molecular property prediction, J. Am. Chem. Soc., 2023, 145, 22584–22598 CrossRef CAS PubMed.
- Y. Wang, M. Huang, H. Deng, W. Li, Z. Wu, Y. Tang and G. Liu, Identification of vital chemical information via visualization of graph neural networks, Briefings Bioinf., 2023, 24, bbac577 CrossRef PubMed.
- V. P. Dwivedi, C. K. Joshi, A. T. Luu, T. Laurent, Y. Bengio and X. Bresson, Benchmarking graph neural networks, J. Mach. Learn. Res., 2023, 24, 1–48 Search PubMed.
- G. Corso, H. Stark, S. Jegelka, T. Jaakkola and R. Barzilay, Graph neural networks, Nat. Rev. Methods Primers, 2024, 4, 17 CrossRef CAS.
- X. Shi, L. Zhou, Y. Huang, Y. Wu and Z. Hong, A review on the applications of graph neural networks in materials science at the atomic scale, Mater. Genome Eng. Adv., 2024, 2, e50 CrossRef CAS.
- V. H. C. Gil and C. N. Rowley, Graph neural networks for identifying protein-reactive compounds, Digital Discovery, 2024, 3, 1776–1792 RSC.
- R. Satheeskumar, Enhancing drug discovery with AI: Predictive modeling of pharmacokinetics using Graph Neural Networks and ensemble learning, Intell. Pharm., 2025, 3, 127–140 Search PubMed.
- N. Ulrich, K. Voigt, A. Kudria, A. Böhme and R.-U. Ebert, Prediction of the water solubility by a graph convolutional-based neural network on a highly curated dataset, J. Cheminf., 2025, 17, 55 Search PubMed.
- C. Bai, L. Wu, R. Li, Y. Cao, S. He and X. Bo, Machine Learning-Enabled Drug-Induced Toxicity Prediction, Adv. Sci., 2025, 2413405 CrossRef CAS PubMed.
- T. W. Ko, B. Deng, M. Nassar, L. Barroso-Luque, R. Liu, J. Qi, E. Liu, G. Ceder, S. Miret and S. P. Ong, Materials Graph Library (MatGL), an open-source graph deep learning library for materials science and chemistry, arXiv, 2025, preprint, arXiv:2503.03837 DOI:10.48550/arXiv.2503.03837.
- F. Therrien, E. H. Sargent and O. Voznyy, Using GNN property predictors as molecule generators, Nat. Commun., 2025, 16, 1–7 Search PubMed.
- S. Kumar and A. Bande, manuscript under editorial review in ACS Omega, 2025, under review.
- R. Ramakrishnan, P. O. Dral, M. Rupp and O. A. von Lilienfeld, Quantum chemistry structures and properties of 134 kilo molecules, Sci. Data, 2014, 1, 140022 CrossRef CAS PubMed.
- S. A. Cuesta, M. Moreno, R. A. Lopez, J. R. Mora, J. L. Paz and E. A. Marquez, Electro-Predictor: An application to predict Mayr's electrophilicity E through implementation of an ensemble model based on machine learning algorithms, J. Chem. Inf. Model., 2023, 63, 507–521 CrossRef CAS PubMed.
- T. N. Kipf and M. Welling, Semi-Supervised Classification with Graph Convolutional Networks, arXiv, 2017, preprint, arXiv:1609.02907 DOI:10.48550/arXiv.1609.02907.
- Z. Xiong, D. Wang, X. Liu, F. Zhong, X. Wan, X. Li, Z. Li, X. Luo, K. Chen, H. Jiang and M. Zheng, Pushing the boundaries of molecular representation for drug discovery with the graph attention mechanism, J. Med. Chem., 2019, 63, 8749–8760 CrossRef PubMed.
- K. Xu, W. Hu, J. Leskovec and S. Jegelka, How powerful are graph neural networks? International Conference on Learning Representations (ICLR), 2019.
- W. Hu, B. Liu, J. Gomes, M. Zitnik, P. Liang, V. Pande and J. Leskovec, Strategies for pre-training graph neural networks, International Conference on Learning Representations (ICLR), 2020.
- S. Brody, U. Alon and E. Yahav, How Attentive are Graph Attention Networks? International Conference on Learning Representations (ICLR), 2022.
- W. L. Hamilton, R. Ying and J. Leskovec, Inductive representation learning on large graphs, Adv. Neural Inf. Process. Syst., 2017, 1024–1034 Search PubMed.
- D. Weininger, SMILES, a chemical language and information system. 1. Introduction to methodology and encoding rules, J. Chem. Inf. Comput. Sci., 1988, 28, 31–36 CrossRef CAS.
- K. T. Schütt, P.-J. Kindermans, G. Felix, S. Chmiela, A. Tkatchenko and K.-R. Müller, SchNet: A continuous-filter convolutional neural network for modeling quantum interactions, NeurIPS, 2017 Search PubMed.
- J. Gasteiger, J. Groß and S. Günnemann, GemNet: Universal directional graph neural networks for molecules, Adv. Neural Inf. Process. Syst., 2021, 6790–6802 Search PubMed.
- T. Kirschbaum and A. Bande, Transfer learning for molecular property predictions from small datasets, AIP Adv., 2024, 14, 105119 CrossRef CAS.
- B. W. Silverman, Density estimation for statistics and data analysis, Routledge, 2018 Search PubMed.
- F. Changyong, W. Hongyue, L. Naiji, C. Tian, H. Hua, L. Ying and M. T. Xin, Logtransformation and its implications for data analysis, Shanghai Arch. Psychiatry, 2014, 26, 105 Search PubMed.
- R. M. West, Best practice in statistics: The use of log transformation, Ann. Clin. Biochem., 2022, 59, 162–165 CrossRef PubMed.
- M. M. Mukaka, A guide to appropriate use of correlation coefficient in medical research, Malawi Med. J., 2012, 24, 69–71 CAS.
- S. Kearnes, K. McCloskey, M. Berndl, V. Pande and P. Riley, Molecular graph convolutions: moving beyond fingerprints, J. Comput.-Aided Mol. Des., 2016, 30, 595–608 CrossRef CAS PubMed.
- G. Landrum, RDKit: Open-source cheminformatics, 2016, https://www.rdkit.org/.
- T. Akiba, S. Sano, T. Yanase, T. Ohta and M. Koyama, Optuna: A Next-generation Hyperparameter Optimization Framework, Proceedings of the 25th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining, 2019, pp. 2623–2631 Search PubMed.
- D. P. Kingma, A method for stochastic optimization, arXiv, 2014, preprint, arXiv:1412.6980 DOI:10.48550/arXiv.1412.6980.
- L. Prechelt, in Neural Networks: Tricks of the Trade, ed. G. B. Orr and K.-R. Müller, Springer, Berlin, Heidelberg, 1998, pp. 55–69 Search PubMed.
- A. Paszke, S. Gross, F. Massa, A. Lerer, J. Bradbury, G. Chanan, T. Killeen, Z. Lin, N. Gimelshein and L. Antiga, et al., PyTorch: An Imperative Style, High-Performance Deep Learning Library, Adv. Neural Inf. Process. Syst., 2019, 32, 8026–8037 Search PubMed.
- M. Fey and J. E. Lenssen, Fast graph representation learning with PyTorch Geometric, arXiv, 2019, preprint, arXiv:1903.02428 DOI:10.48550/arXiv.1903.02428.
- T. Hastie, R. Tibshirani and J. Friedman, The Elements of Statistical Learning: Data Mining, Inference, and Prediction, Springer, 2nd edn, 2009 Search PubMed.
- T. Chai and R. R. Draxler, Root mean square error (RMSE) or mean absolute error (MAE)? – Arguments against avoiding RMSE in the literature, Geosci. Model Dev., 2014, 7, 1247–1250 CrossRef.
- C. J. Willmott and K. Matsuura, Advantages of the mean absolute error (MAE) over the root mean square error (RMSE) in assessing average model performance, Clim. Res., 2005, 30, 79–82 CrossRef.
- Daylight Chemical Information Systems, Inc. SMARTS – A language for describing molecular patterns. Daylight Chemical Information Systems, Inc., 2008, Daylight Theory Manual.
| This journal is © the Owner Societies 2026 |




