Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
pH-dependent structural alterations in LHCII revealed by two-dimensional infrared spectroscopy
Thanh Nhut Do ,
Kinga Hajduk
,
Kinga Hajduk ,
Roberta Croce
,
Roberta Croce and
John T. M. Kennis
and
John T. M. Kennis *
*
Department of Physics and Astronomy, Faculty of Science, Vrije Universiteit Amsterdam, De Boelelaan 1100, 1081 HZ Amsterdam, The Netherlands. E-mail: j.t.m.kennis@vu.nl
First published on 23rd March 2026
Abstract
Light-harvesting complex II (LHCII) is the major light-harvesting antenna, which is responsible for harvesting sunlight energy and photoprotection of photosystem II. In this study, we assessed the structural changes in LHCII that are induced by lowering the pH. Absorption, circular dichroism and emission spectra of LHCII indicate that the trimeric LHCII samples were well intact and maintained in the light-harvesting state, exhibiting a single 3-ns exponential fluorescence decay. Fourier-transform and two-dimensional infrared spectroscopies reveal a frequency upshifting of amide I band of some α-helical elements upon increasing acidity, which could be interpreted as a combination of refolding of a 310 helix and a C-terminal coil to form new α-helices. Some sub-stoichiometry formations of β-motifs were also observed upon acidification with the β-sheet's size estimated at around 30–40 amino acids. This β-sheet can be formed with the N-terminal coil or co-folding between the N-terminus and the stromal loop. Our results demonstrate that lowering the pH induces significant structural changes in LHCII, but that they are not sufficient to switch LHCII into a quenched state. In the thylakoid membrane, such pH-dependent structural changes may enable specific protein–protein interactions between LHCII and other membrane proteins, in particular PsbS, that are required for non-photochemical quenching.
Introduction
Light-harvesting complex II (LHCII) is the major light-harvesting antenna of photosystem II (PSII) in vascular plants. It has been well known that PSII is a highly efficient enzyme that can harvest sunlight energy (with the assistance of LHCII) and perform photochemistry to extract protons and electrons from water and drive subsequent processes of photosynthesis. Due to the high efficiency, PSII is under a potential of being over-excited under harsh high-light conditions. Hence, a process called non-photochemical quenching (NPQ) is utilized by plants to dissipate the excess absorbed light energy as heat.1–4 The molecular mechanism(s) of NPQ are still under debate with various proposed models, spanning a large range of timescales from seconds to minutes, and involving many different factors, including LHCII,5,6 PsbS,7,8 xanthophyll cycle,9–12 and even membrane reorganization.13 The fastest phase of NPQ happens within seconds and is triggered by the lumen acidification under high light.5,14 This is called energy-dependent quenching or qE component of NPQ. For a recent review, see ref. 4.The most generally accepted picture of the NPQ's qE phase usually involves both PsbS and LHCII.1,2,4,15 PsbS is known to be essential for NPQ (the npq4 mutant plant lacks PsbS and cannot perform NPQ7), having several pH-sensitive glutamate (Glu) residues on the lumenal side helices.16–18 The pH-sensitivity of Glu residues of PsbS was confirmed to be in the physiological range of the values of the lumen by molecular dynamics (MD) simulation.19,20 Hence, NPQ is proposed to begin with the lumen acidification causing protonation of Glu residues on the lumenal helices of PsbS. Subsequently, PsbS can undergo structural changes,21 and then this could lead to an increase of PsbS's affinity with other proteins in PSII supercomplex. A mutagenesis study using the PsbS mutant plants lacking one or two pH-sensitive Glu residues near the lumenal side (E122 and E226) observed a lack of quenching, indicating the importance of PsbS protonation step to initiate NPQ.8,18
After Glu protonation, PsbS may interact with other light-harvesting antenna proteins to switch them to a quenched state. It has been shown that almost all lhcb proteins can be quenched4,22,23 but the major quenching and hence the best candidate for PsbS interaction is LHCII.6,24–26 In this picture, the association with PsbS will lead to structural changes switching LHCII from the light-harvesting state to the quenched state. The two states of LHCII are characterized by two different fluorescent lifetimes of ∼3–4 ns in the light-harvesting state and a much shorter (below 0.5 ns) in the quenched state.27–30 In the quenched state LHCII dissipates large part of the harvested energy, thus delivering less energy to PSII reaction center as compared to the light-harvesting state, thereby lowering the probability of over-excitation.
The molecular mechanism for PsbS–LHCII interaction was investigated in several studies. In proteoliposome systems, both low pH and PsbS were determined to be essential to induce quenching26 and PsbS–LHCII interaction was proposed to induce Chl–Chl charge-transfer state formation in LHCII,31 or enhance the Chl–carotenoid coupling,24 which are able to quench LHCII. A cryo-electron transmission microscopy (cryo-TEM) study suggested that PsbS can associate with LHCII and compress the protein scaffold of LHCII along the membrane direction, switching LHCII to the quenched state.32 An MD simulation study on pH-dependent structural changes of PsbS predicted that the amphipathic helix H3 is the docking site responsible for interaction with other light-harvesting complex.19
The necessity of PsbS in triggering NPQ was questioned by a biochemical study.33 Using diaminodurene (DAD), the lumenal pH was reduced to around 4, which is well below the physiological value of 5.5. This highly acidic condition was found to enable similar levels of NPQ in both wild-type (WT) and npq4 mutant chloroplasts. Hence, the authors suggested that PsbS is not indispensable for NPQ and acts as an allosteric regulator for LHCII, allowing the switch to a quenched state to occur at a higher pH value.
In a recent cryo-EM study on LHCII, various structural differences were observed in the protein scaffolds between two pH conditions (7.8 and 5.4). However, LHCII trimers were not quenched upon lowering the pH to the physiological value of 5.4 in both detergent-solubilized and nanodisc forms as no significant changes were observed in the sub-ns component of the fluorescent decay kinetics.34
In this work, we used a much more acidic condition (pH 4.4) that in vivo was shown to be sufficient induce quenching in LHCII even in the absence of PsbS.33 We therefore performed a comprehensive spectroscopic analysis to investigate whether these conditions could alter the light-harvesting dynamics and structural organization of LHCII.
We investigated trimeric LHCII solubilized at two pH conditions using different spectroscopic techniques. These include the state-of-the-art two-dimensional infrared (2DIR) as well as Fourier transform infrared (FTIR) spectroscopy to probe the structure of LHCII protein, due to the high sensitivity of the amide I vibrational modes of the protein backbone to the secondary protein structural motifs. To complement the IR results providing structural information of the protein scaffold, we also report the spectroscopic observables in the visible region, probing the electronic spectral properties of chlorophyll (Chl) pigments in LHCII.
Results and discussion
In this section, we present the visible spectroscopic results to validate the integrity of LHCII trimers after buffer exchange procedure. Subsequently, IR spectroscopies focusing on protein structural information will be discussed. We note that the IR spectroscopic observations specifically focus on the amide I band region around 1650 cm−1, strongly overlapping with H2O absorption.35 Therefore, it is a common practice to substitute H2O with D2O, and all proteins and buffers used in this study were deuterium exchanged to the maximal possible extent. All pH values in this study will be henceforth mentioned as pD, and the pD value is determined as pD = pH* + 0.4, where pH* is the reading from the commercial glass electrode pH meter.36 All the visible spectroscopies were performed on the deuterated samples as IR spectroscopies for the sake of consistency.Absorption, circular dichroism and fluorescence spectroscopy
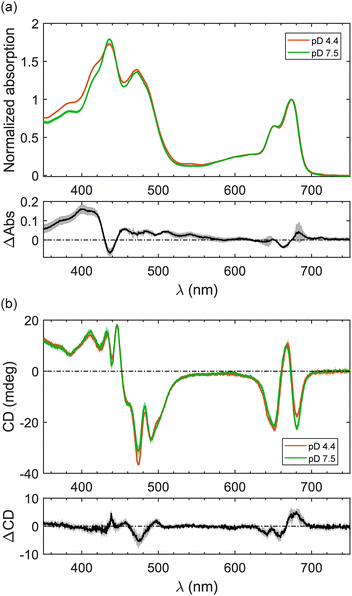
In order to investigate the electronic spectral properties in LHCII, we performed steady-state absorption and circular dichroism (CD) measurements on the solubilized trimeric LHCII samples at pD 4.4 and pD 7.5. Henceforth, the pD-4.4 and pD-7.5 samples will be called acidic and neutral samples, respectively. All the one-dimensional spectra will be colored in orange/green for acidic/neutral samples and the difference spectra will be defined as acidic-minus-neutral spectra and plotted in black, unless otherwise stated.Steady-state absorption spectra of both samples exhibit the typical shape of LHCII's absorption comprising two main bands: the Qy band in the red region and the Soret band in the blue-green region, as observed in Fig. 1(a). The absorption spectrum of the acidic sample shows an additional small absorption at 540 nm and in the 400–420 nm region. This indicates that under highly acidic conditions, a minor fraction of Chls has been converted to pheophytin.
CD spectra are sensitive to excitonic couplings of the Chl pigments. Fig. 1(b) shows that the typical W-like lineshape at the Qy spectral region is well conserved in both samples. This indicates that the excitonic coupling networks of LHCII trimers are largely retained. The trimeric form of LHCII remains intact under highly acidic conditions as monomerization would collapse the negative feature at 475 nm of the trimeric LHCII's CD spectra,37 which is not observed here.
As described before, the aggregation of LHCII can be sensitively detected by low-temperature fluorescence with the characteristic F700 band.38 Hence, to further verify the integrity of trimeric LHCII samples, we performed both steady-state and time-resolved fluorescence measurements and the results are presented in Fig. 2.
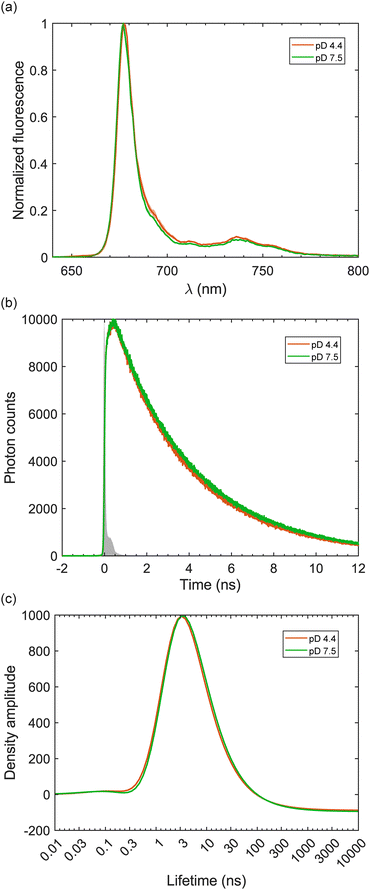 | ||
| Fig. 2 Fluorescence measurements of LHCII at two pD conditions. (a) 77-K steady-state fluorescence spectra excited at Chl a's Soret band 440 nm. (b) Time-correlated single photon counting (TCSPC) traces detected at 680 nm upon excitation at 440 nm. The grey shaded area depicts the instrument response function (IRF) measured with pinacyanol iodide in methanol. (c) Lifetime density of the TCSPC traces obtained by performing Laplace transformation of the traces in (b) taking into account the IRF and using 0.3% intensity regularization with Tikhonov algorithm.39 | ||
Low temperature (77 K) steady-state fluorescence spectra of both LHCII samples are presented in Fig. 2(a). The excitation wavelength 440 nm was chosen to mainly excite Chl a's Soret band. Despite some minimal variations within ±1 standard deviation uncertainty, both fluorescence spectra are virtually identical. The putative F700 band, which is a marker for aggregation,38 is clearly absent in our samples. Two more excitation wavelengths at 475 nm and 500 nm, preferentially exciting Chls b and carotenoids, respectively, were also used to measure 77-K steady-state fluorescence. The same spectra were obtained for these excitation wavelengths (shown in Fig. S1 of the SI), indicating that excitation energy transfer between pigments is properly retained at the two pD conditions. The time-correlated single photon counting (TCSPC) traces in Fig. 2(b) are strongly overlapped and also identical within the noise level. The Laplace transformation of two TCSPC decay traces yields the corresponding lifetime distribution of the decay dynamics shown in Fig. 2(c). The lifetime distributions of both samples show a single peak at ∼3–4 ns, which is the well-established lifetime of the light-harvesting state of LHCII.26,27 No sub-ns lifetime, which is characteristic for the quenched state, can be resolved.
With the above-described analysis and interpretations on UV-vis spectroscopic observables, we can confidently confirm that the trimeric LHCII samples were intact after the buffer exchange procedure and the excitonic couplings between Chl pigments were altered very minimally. However, these small changes do not form any quenched population of LHCII in both pD conditions.
FTIR spectroscopy
We further applied infrared spectroscopy to study the effect of acidification on the LHCII protein structure. As mentioned above, the amide I band is specifically sensitive to the protein secondary structure. The amide I vibrational mode is mainly contributed by the stretching vibration of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O in the amide bond together with a minor contribution of the bending motion of the adjacent N–H bond. Once folded to form secondary structures, almost all C
O in the amide bond together with a minor contribution of the bending motion of the adjacent N–H bond. Once folded to form secondary structures, almost all C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O groups will form hydrogen bonds (H-bonds) with N–H (and/or N–D) of the proximal residues. On top of that, for well-defined secondary structural motifs like α-helices or β-sheets/strands, all C
O groups will form hydrogen bonds (H-bonds) with N–H (and/or N–D) of the proximal residues. On top of that, for well-defined secondary structural motifs like α-helices or β-sheets/strands, all C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bonds are in an almost parallel/anti-parallel alignment. These make the amide I vibrational modes delocalize over the broad spatial extent of the corresponding protein backbones40 and hence, are sensitive to the structures of the protein motifs.
O bonds are in an almost parallel/anti-parallel alignment. These make the amide I vibrational modes delocalize over the broad spatial extent of the corresponding protein backbones40 and hence, are sensitive to the structures of the protein motifs.
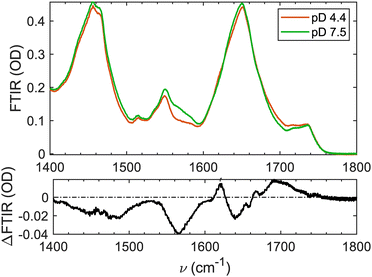
In Fig. 3, the solvent-subtracted FTIR spectra of LHCII at both pDs are presented. Three protein-related spectral regions are observed in the FTIR spectra. The amide II’ region between 1400–1500 cm−1 is dominated by the N–D vibrational motions and quite structurally insensitive, hence will not be discussed further.41,42
The intermediate band between 1500–1600 cm−1 is complex as this region can be contributed by various factors in the protein. This can be the remnant amide II vibration (N–H vibration) due to non-exchangeable H atoms, which are deeply buried inside the hydrophobic core of the protein. Two sharp vibrational modes at around 1515 and 1550 cm−1 can be attributed to the Chls’ and carotenoids’ C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretch motions, respectively.43,44 A broad band between 1565 and 1585 cm−1 can be attributed to the asymmetric stretch of the deprotonated carboxylate group on the sidechain of aspartate (Asp) and Glu residues.45 This is consistent with the fact that the FTIR intensity in this region is lower in acidic conditions. The high-frequency spectral region above 1700 cm−1 exhibits some minor signal. These signal can be assigned as the vibrations of C
C stretch motions, respectively.43,44 A broad band between 1565 and 1585 cm−1 can be attributed to the asymmetric stretch of the deprotonated carboxylate group on the sidechain of aspartate (Asp) and Glu residues.45 This is consistent with the fact that the FTIR intensity in this region is lower in acidic conditions. The high-frequency spectral region above 1700 cm−1 exhibits some minor signal. These signal can be assigned as the vibrations of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O in ester groups of Chls.46 In addition, the protonation/deuteration of Asp and Glu residues shifts the C
O in ester groups of Chls.46 In addition, the protonation/deuteration of Asp and Glu residues shifts the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O vibrational frequencies of the carboxylate sidechains to the 1710–1720 cm−1 region.45 Hence, this can contribute to the increase of pD-4.4 signal in this region as observed in ΔFTIR spectrum.
O vibrational frequencies of the carboxylate sidechains to the 1710–1720 cm−1 region.45 Hence, this can contribute to the increase of pD-4.4 signal in this region as observed in ΔFTIR spectrum.
The amide I region between 1600–1700 cm−1 contains lots of information about the protein secondary structure. The amide I spectra of both sample are dominated by a sharp peak at 1650 cm−1. This can be ascribed to the α-helices’ signals, which are usually resolved as an asymmetric peak at 1650 cm−1. The asymmetry of the α-helix peak is explained by the fact that for the idealized infinitely long α-helix, the amide I band is comprised of two modes called A and E1 having transition frequencies split by around 10 cm−1,41 which is smaller than the linewidth of the IR transition peaks (20–25 cm−1). Hence, in the complicated large-size protein like LHCII, these two modes can be treated as one single asymmetric α-helix signal. The domination of α-helix signal in the amide I band is expected since the protein motif of LHCII contains mostly α-helices, resolved by various structural techniques,47,48 under different pH conditions.34
We note that the area-normalization forces the ΔFTIR spectrum to be conservative, i.e., integrated area = 0 between 1600–1700 cm−1. This can be true with the assumption that the area of amide I is linearly proportional to the number of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O oscillators. In reality, the transition dipole moments of C
O oscillators. In reality, the transition dipole moments of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bonds are not the same in all protein motifs, e.g., the random coils exhibit lower transition dipole moments than α-helices, despite similar transition frequencies.49 In addition, solvent subtraction can also introduce some artificially tilted baseline, which can make the area-normalization less certain. Hence, we will refrain from making extensive quantitative interpretations and only focus on overall features when discussing difference spectra (both ΔFTIR and Δ2DIR) in IR regimes.
O bonds are not the same in all protein motifs, e.g., the random coils exhibit lower transition dipole moments than α-helices, despite similar transition frequencies.49 In addition, solvent subtraction can also introduce some artificially tilted baseline, which can make the area-normalization less certain. Hence, we will refrain from making extensive quantitative interpretations and only focus on overall features when discussing difference spectra (both ΔFTIR and Δ2DIR) in IR regimes.
Lowering the pD from 7.5 to 4.4 leads to the overall broadening of the amide I band. Area-normalization slightly increases the peak intensity of the pD-7.5 spectrum relatively to the pD-4.4 one. This results in the negative feature at 1640 cm−1 in the ΔFTIR spectrum (Fig. 3). Concomitantly, there is an intensity increase at higher frequency region extending from 1660–1740 cm−1. This signal is relatively broad and non-trivial to assign as it is extended towards rather high frequency, which is not the central frequency of normal amide I vibration. The ΔFTIR signal above 1700 cm−1 is due to the Asp/Glu protonation as described previously, and the lower frequency part (<1700 cm−1) is due to amide I band spectral changes. In addition, there is a small increase in intensity of the pD-4.4 spectrum at around 1620 cm−1. This can be assigned to some additional β-motifs that have been formed at lower pD. Another explanation for this positive 1620-cm−1 signal is the downshifting of the low-frequency mode of some β-motifs that increase in size when lowering the pD. These assignments will later be reaffirmed and discussed with 2DIR spectra.
2DIR spectroscopy
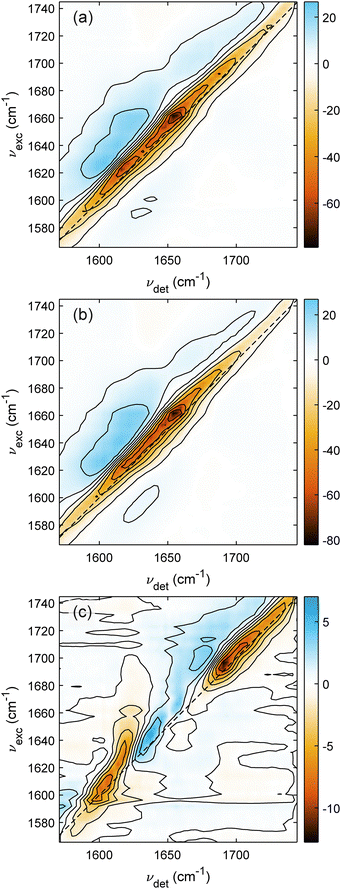
To further disentangle the spectral changes observed in the ΔFTIR spectrum, we performed 2DIR measurements on LHCII at neutral and acidic conditions. With the additional frequency axis, 2D spectroscopy was extensively utilized as a powerful technique to disentangle the congested spectral features.50–52 In this study, the 2DIR spectra will be presented with the axis convention as: the vertical axis depicts the excitation frequency νexc and the horizontal axis shows the detection frequency νdet. All signals on each 2D spectrum will be denoted as a (νexc, νdet) coordinate, except the diagonal signals, which have νexc = νdet. The sign convention for the 2DIR signal is adopted from transient absorption spectroscopy, with the negative diagonal signals indicating the ground-state bleach (GSB) of v = 0 → v = 1 vibrational transitions, and the red-shifted positive signals of excited-state absorption (ESA) corresponding to the v = 1 → v = 2 transitions.Fig. 4 presents the 2DIR spectra of LHCII recorded at a population time of 500 fs using the perpendicular (ZZYY) polarization scheme to enhance contrast for the excitonic coupling cross-peaks.53 First of all, the 2DIR spectra at the two pDs are nearly identical, which is not unexpected, as can be seen in the FTIR spectra in Fig. 3 that the amide I bands of the two samples are very similar. Both of them contain a diagonally elongated negative feature of GSB, accompanied by the red-shifted positive band of ESA. The diagonally elongated 2D peakshape indicates that the amide I band of LHCII is broadened mainly due to the inhomogeneous distribution of the transition frequencies of different oscillators, not due to the homogeneously broadening lineshapes.54 Almost no excitonic coupling cross-peaks are observed in Fig. 4(a) and (b). Hence, we can interpret the diagonally elongated signals in both 2DIR spectra as (mostly) the combination of various signals coming from different parts of the protein.
The acidic-minus-neutral Δ2DIR spectrum in Fig. 4(c) exhibits three main features along the diagonal. As being defined as the acidic-minus-neutral difference spectrum, together with the fact that the GSB signals along the diagonal line are negative, the negative/positive diagonal signals of the Δ2DIR spectrum correspond to an intensity increase/decrease in acidic conditions, relative to the neutral spectrum. The most noticeable feature in the Δ2DIR spectrum is the broad negative feature extending diagonally from around 1680 cm−1 to more than 1720 cm−1. This corresponds to the increased signal in acidic conditions as observed in Fig. 3 as the broad positive signal in the ΔFTIR spectrum. As described before, this increased signal in acidic spectra can be partially explained by the protonation/deuteration of the Asp and Glu sidechains, hence, shifting the signals from 1560–1580 cm−1 (not covered in 2DIR) to higher frequency region of around 1700–1720 cm−1.
The increased signal in acidic conditions at a high frequency regime is not just contributed by the protonation of Asp and Glu residues, but also by some intensity reduction around 1640 cm−1 as observed as the positive diagonal signal in the Δ2DIR spectrum, corresponding to the negative signal in the ΔFTIR spectrum at the same region. In both difference spectra (ΔFTIR and Δ2DIR), this 1640-cm−1 feature has a narrower linewidth, extending between 1620–1660 cm−1. This frequency region is slightly lower than the normal central frequency of the idealized α-helix. Together with the fact that the spectral linewidths of α-helices are usually broad (∼30–40 cm−1),42 we assign this reduced signal in the acidic sample to be due to the frequency upshifting of some α-helices. As the spectral linewidth is broader than the frequency upshifting, this frequency shift will result in the reduction of the low-frequency signal and the increase at the high-frequency edge of the bandwidth. While near the peak center, most of the signals will be canceled out in the difference spectra (compensating effect). This explains the broad linewidth of the increased signal in ΔFTIR and Δ2DIR spectra at the high-frequency region, and the near-zero signal around 1660 cm−1 in the Δ2DIR spectrum.
The assignment of α-helices frequency upshifting is supported by the structural alterations of LHCII protein motifs at low pH observed by Ruan et al. using cryo-EM.34 The overall comparison of the protein manifold of LHCII in detergent at pH 7.8 and pH 5.4 is presented in Fig. 5(a). Three main helices (helix A, B and C) are largely similar between the two structures, evidenced by almost perfect structural overlay in Fig. 5(a). On the other hand, noticeable differences can be observed around helix E, helix D and the C-terminal coil.
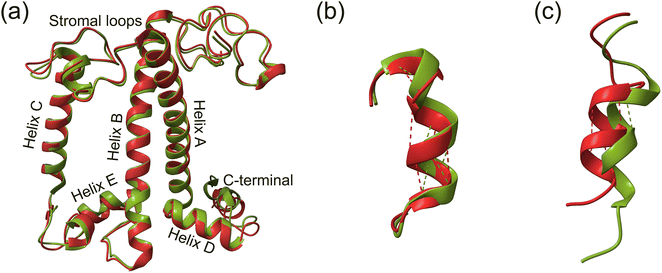 | ||
| Fig. 5 Demonstration for the structural alterations of the LHCII protein motifs. (a) Overlaying the protein scaffolds of one monomeric unit of LHCII in detergent at neutral pH 7.8 (green) and acidic pH 5.4 (red) fetched from the Protein Data Bank cryo-EM structures (PDB IDs: 8IWX and 8IWY).34 (b) and (c) The structural comparison focusing on helix E (b) and the C-terminal coil (c). Dashed lines indicate the H-bonds. The protein figures are prepared with ChimeraX software.55 | ||
At pH 5.4, helix E was observed to refold from a 310 helix to an α-helix. As depicted in Fig. 5(b), the green 310 helix is the pH-7.8 structure exhibiting a slightly smaller radius compared to the red α-helix of the pH-5.4 structure. The H-bond network is also more periodic and regular in the α-helix. As 310 helix is expected to exhibit slightly lower amide I frequency compared to the α-helix (∼5–10 cm−1),56 hence, the refolding of helix E can slightly upshift (part of) the amide I band of LHCII in acidic conditions.
In addition, the C-terminal coil near helix D was also observed to fold to an α-helix when lowering pH from 7.8 to 5.4. As shown in Fig. 5(c), despite the ribbon cartoon, the green C-terminal coil looks more unstructured compared to the red α-helix, with only one detectable H-bond while the red α-helix shows 4 H-bonds with regular periodic spacing pattern. The random coils are usually expected to have a lower transition dipole moment, while exhibiting very similar transition frequency as α-helix. In addition, a coil element is expected to yield a much lower signal in 2DIR than a helical element because of the 4th power scaling of the transition dipole moment in 2D spectroscopy.57 Hence, the folding of the C-terminal coil to an α-helix will increase the signal in acidic IR spectra, and this can make the ΔFTIR and Δ2DIR become non-conservative.
Another possible contribution to the amide I band frequency upshifting is the movement of any helical elements towards the hydrophobic core of the protein. The frequency upshifting of α-helix is explained as the stabilization of the overall helical structure when being buried in the more compact hydrophobic core. This structural stabilization enhances the couplings of the vibrational excitons, together with altered surrounding refractive index,58 and/or the smaller degree of hydration59 causing the amide I frequency upshifting. This effect has been reported in a similar membrane protein PsbS with the amphipathic helix retraction towards the membrane,21 as well as artificial peptide using α-helix stabilization agent.58 In the case of LHCII, a small translational motion of helix D can be observed in the cryo-EM structures.34 The red helix D in Fig. 5(a) is slightly translated towards the core of the protein compared to the green helix D. Ruan et al. explained this motion as the allosteric effect caused by the refolding of both helix E and C-terminal coil forming two new α-helices and pushing helix D towards the protein core upon acidification. We note that, although the translation of helix D in cryo-EM structures is quite small, the acidic pH used in the cryo-EM study was 5.4, while in our study the pD is 4.4, which is more acidic and hence, can enhance the movement of helix D.
The third feature observed in the amide I region of Δ2DIR is the increase of acidic signal at lower frequency of around 1600–1620 cm−1. This frequency is very low compared to the normal central frequency of α-helices. The signal in this frequency range usually indicates β-motifs. In β-sheets/strands, the regular periodic repetitions of coupling pattern between adjacent amino acids create a large delocalization effect of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O vibrations over the whole spatial extension of the β-sheets/strands. This delocalization effect yields the redistribution of transition dipole moments and usually two strong vibrational modes can be observed for β-motifs. The stronger mode is called ν⊥ and exhibits lower central frequency in the range of around 1600–1640 cm−1, depending on the β-sheet's size.60 The weaker mode is called ν‖ having higher frequency of around 1690 cm−1. These two modes are excitonically coupled as they are formed by the same collection of oscillators. Hence, the cross-peaks connecting these two modes are the signature for β-motifs and should be resolved in 2DIR spectra. However, inspecting Fig. 4 does not provide any signature of the cross-peaks coupling two β excitonic modes as expected.
O vibrations over the whole spatial extension of the β-sheets/strands. This delocalization effect yields the redistribution of transition dipole moments and usually two strong vibrational modes can be observed for β-motifs. The stronger mode is called ν⊥ and exhibits lower central frequency in the range of around 1600–1640 cm−1, depending on the β-sheet's size.60 The weaker mode is called ν‖ having higher frequency of around 1690 cm−1. These two modes are excitonically coupled as they are formed by the same collection of oscillators. Hence, the cross-peaks connecting these two modes are the signature for β-motifs and should be resolved in 2DIR spectra. However, inspecting Fig. 4 does not provide any signature of the cross-peaks coupling two β excitonic modes as expected.
To visualize the small features in 2DIR spectra, we reset the color bars and contours of Fig. 4(a) and (b) to 10% of the corresponding maximum. The new plots are presented in Fig. 6(a) and (b). With the new scaling, the small features in 2DIR spectra are enhanced and two cross-peaks corresponding to the β-motifs can be easily recognized. The below-diagonal cross-peak is located at (νexc, νdet) = (1610, 1690) cm−1, while the above diagonal cross-peak at (1720, 1620) cm−1 is a bit off from the expected coordinate, which should be a mirror-image of the other cross-peak via the diagonal line. This non-symmetric cross-peak pair can be explained as the above-diagonal cross-peak is strongly distorted by the large ESA positive signal as the major protein motif of the whole trimeric LHCII is still the α-helix.
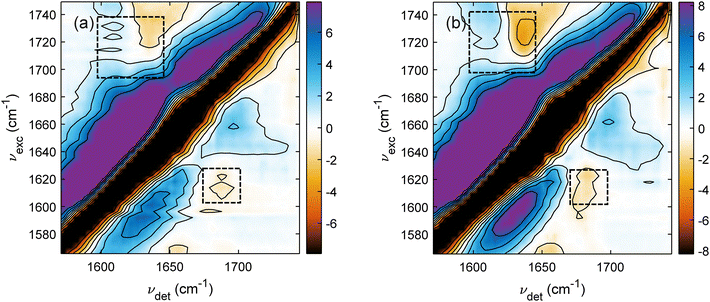 | ||
| Fig. 6 Same as Fig. 4(a) and (b) with the maximal intensities set at 10% of the original maximal intensities to visualize the small features. The dashed rectangles roughly indicate the cross-peaks position corresponding to the β-motifs. | ||
The quasi-transient absorption (quasi-TA) spectrum can be extracted from a 2D spectrum by taking a horizontal slice at a certain excitation frequency νexc (or integrating several horizontal slices within a certain range around νexc). The quasi-TA spectrum, as described by the name, can be considered analogous as the TA spectrum recorded using ultra-narrowband excitation frequency νexc. Any signal along νdet dimension is the spectral signature of molecular factors induced/created by the excitation of νexc. Inspecting the quasi-TA spectra extracted from two 2DIR spectra at two pDs (see SI) reveals that the cross-peaks are somewhat clearer in the pD-4.4 spectrum. Hence, it would be interesting to inspect the quasi-TA spectra extracted from the Δ2DIR spectrum, which can highlight the pD-dependent β-motifs.
The low-frequency quasi-TA spectra (νexc = 1610–1620 cm−1) in Fig. 7(a) exhibit some diagonal features due to the newly formed β-motifs upon acidification, as discussed above. These diagonal peaks are coupled with the cross-peaks (indicated with arrows) having dispersive lineshape with a positive feature at νexc = 1680 cm−1 and a negative peak at νexc = 1690 cm−1. The clearest dispersive lineshape of the cross-peak is observed in the quasi-TA spectra of νexc = 1620–1630 cm−1.
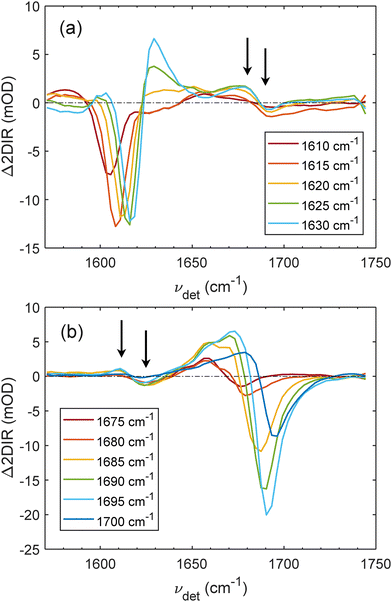 | ||
| Fig. 7 Quasi-TA spectra extracted from Δ2DIR spectrum in Fig. 4(c). The denoted frequencies in the legend are the corresponding excitation frequency νexc. The low-frequency quasi-TA spectra are presented in (a) and the high-frequency ones are presented in (b) highlighting the below and above diagonal cross-peaks, respectively. The arrows indicate the maxima of the positive and negative features of the cross-peaks as discussed below. | ||
The high-frequency quasi-TA spectra are presented in Fig. 7(b). In this region, beside the increased signal from β-motifs, some contribution from α-helices also appears as we observe the frequency upshifting related to the α-helices discussed above. As we can see the dispersive lineshape of the cross-peak is resolved clearest with the 1685–1695-cm−1 spectra, and the diagonal features are also strongest in these spectra. The cross-peak has a negative peak around 1625 cm−1 and a positive peak around 1610 cm−1, as indicated by the arrows in Fig. 7(b). Together with the interpretations obtained from low-frequency quasi-TA spectra, we can conclude that two excitonic modes ν⊥ and ν‖ of β-motifs form upon acidification are centered at around 1625 cm−1 and 1690 cm−1, respectively. For these frequencies, recent theoretical calculations estimate that the β-sheet size should be around 30–40 amino acids,60 depending on how long each β-strand is and whether the β-folding motif is parallel or anti-parallel.
As mentioned before, the increase signal around 1620 cm−1 in acidic spectra observed in Δ2DIR and ΔFTIR spectra can also be explained as the frequency downshifting of the ν⊥ mode. Due to the strong dependence of the ν⊥ mode frequency on the extension degree of the β-motif, if the acidification enlarges the β-sheet size, the ν⊥ mode frequency will be downshifted (probably from 1630–1640 cm−1 to 1620 cm−1), and will result in the decrease/increase of 1640/1620-cm−1 signal as observed in both Δ2DIR and ΔFTIR spectra. However, this possibility can be excluded by fitting the cross-peak region of quasi-TA spectra extracted at νexc = 1690 and 1695 cm−1. The fitting results presented in the SI show that the frequency of ν⊥ mode stays constant around 1622 ± 4 cm−1 in all six quasi-TA spectra inspected. Hence, the β-sheets observed in both 2DIR spectra at two pDs exhibit the same size within the fitting uncertainty and the increase of signal around 1620 cm−1 region is really due to the increase amount of β-motifs formed upon acidification.
pD-dependent structural changes in LHCII: implications for the NPQ mechanism
We confidently determine that lowering the pD from 7.5 to 4.4 causes substantial alterations to the protein manifold of trimeric LHCII. The refolding of the 310 helix E and C-terminal coil to α-helices creates allosteric effects to push helix D deeper into the protein hydrophobic core, causing frequency upshifting of amide I band. Some β-motif formation is well-resolved with two excitonic coupling cross-peaks indicating two exciton bands ν⊥ and ν‖ at 1622 ± 4 cm−1 and 1690 cm−1, respectively. Theoretical calculations give an estimation of around 30–40 amino acids forming this β-sheet, corresponding to around 10–15% of the total number of amino acids in lhcb1 sequence.47 This amount of β-motif is not so small in percentage per sequence but the very low signal in 2DIR (1–2 mOD signal of β-sheet cross-peaks out of 80 mOD maximum) indicates that the β-sheet was formed in sub-stoichiometry upon acidification. We note that this estimation about the amount of newly formed β-sheet upon acidification depends on the accuracy of the theoretical calculation used to estimate the β-sheet size.60 We emphasize that all the structural alterations inferred from the 2DIR spectra aided with cryo-EM structures are related to the membrane-exposed region of the LHCII protein (helices D and E, C-terminal coil and N-terminal coil mentioned later). This is reasonable as LHCII is a membrane protein, with the hydrophobic core embedded in the membrane in vivo or well-protected by the detergent shell in solubilized state. Therefore, no or very minimal effects of the acidification are expected to happen in the hydrophobic core region of the LHCII.In the context of NPQ, we hypothesize that the structural response of LHCII to lowering pH is required to enable specific interactions with PsbS that switch LHCII to a quenching state, which may involve repositioning and refolding of amphipathic helices at the lumenal side.19,21 In additions, even if no β-motifs were resolved in LHCII structural studies,34,47,48 we present clear evidence of the presence of β-sheets at both pD conditions, and that the amount of β-sheet increases upon lowering the pD. The observation of β-motifs is in line with a recent MD study, indicating that monomeric LHCII, upon MD equilibration, forms some β-motifs at the lumenal loop near helix E, the stromal loop, and the N-terminal coil.61 Hence, most of the disordered elements, which are not well-resolved in the crystal and cryo-EM structures, may in fact form β-motifs. With our estimation of around 30–40 amino acids for the pD-dependent β-sheet detected in Δ2DIR, the lumenal loop of LHCII, having 33 amino acids (90th–122th residues), is not long enough to form β-sheet of this size (and helix E occupies the 96th–106th amino acids). On the other hand, the N-terminal coil has 53 amino acids and the stromal loop has 31 residues (137th to 168th). We hence propose that upon acidification of LHCII in vitro, the β-sheet could be formed either by the N-terminal coil, which contains sufficient number of residues required for the proposed β-sheet size of 30–40 amino acids, or by co-folding of the N-terminal coil and the stromal loop. In vivo, only the lumenal side is acidified under high light conditions, and it is hence unclear whether in the thylakoid membrane, a similar increase of β-sheet content in LHCII occurs upon lumen acidification. Nevertheless, our results clearly show that LHCII has a propensity to form β-sheets at the stromal side, and we tentatively propose that through transmembrane allosteric effects, β-sheet formation in the stromal elements may occur. The stromal disordered elements, which are not well resolved in any published structures, were proposed to be the interaction region of LHCII and might play a role in grana stacking.62
Conclusion
In this study, we performed a comprehensive spectroscopic investigation on the major light-harvesting complex, LHCII, in two pD conditions. The steady-state absorption and CD results revealed a very minimal change in the excitonic network of LHCII in highly acidic conditions of pD 4.4. The low pD did not switch LHCII from light-harvesting state to quenched state, evidenced by the persistent 3-ns single exponential decay lifetime. Aggregation and/or precipitation issue was also ruled out by the lack of the typical fluorescence emission band at 700 nm in the 77-K steady-state fluorescence spectra. Our results show that even if LHCII undergoes significant and specific protein structural changes upon lowering the pH, such changes are not sufficient to switch LHCII into a quenched state. We propose that in vivo, lumen acidification may result in an intrinsic structural response of LHCII similar to that reported here (even if the pH does not attain the very low values applied here). However, interactions with other components in the thylakoid membrane, in particular PsbS, are likely required to induce a proper qE response.Infrared spectroscopies shed light on the protein secondary structure of LHCII. Upon acidification, a major frequency upshifting of the amide I band was observed in both FTIR and 2DIR spectra. This frequency upshifting could be explained as the combination of various structural alterations reported previously by cryo-EM, including refolding of a 310 helix and a C-terminal coil to form new α-helices, and possibly an allosteric effect of the newly formed helices pushing helix D towards the protein hydrophobic core. These changes at the lumenal side of the membrane may align with corresponding pH-dependent structural changes in PsbS19,21 and facilitate the interactions that are required for a qE response.
In addition, we demonstrated the presence of a small but significant amount of β-sheet motifs at both pD conditions, and that the amount of β-sheet increases upon acidification. The Δ2DIR spectrum revealed two excitonic cross-peaks, which can be unmistakably assigned to the ν⊥ and ν‖ excitons of the β-sheet. An estimation of around 30–40 amino acids for the pD-dependent β-sheet size was made based on the frequencies of two β-sheet excitons. With this size, the β-sheet observed in our 2DIR results was likely formed in the stromal disordered elements either by the N-terminal coil itself or by the co-folding of stromal loop and N-terminal coil together. Although the stroma is not acidified under NPQ conditions in vivo, β-sheet formation at the stromal side through allosteric effects cannot be ruled out and may play a role in mediating interactions with other components in the thylakoid membrane.
Author contributions
T. N. D.: methodology, software, investigation, validation, formal analysis, visualization, writing – original draft, writing – reviews & editing. K. H.: methodology, investigation, validation, writing – reviews & editing. R. C.: conceptualization, resources, writing – reviews & editing, supervision, project administration, funding acquisition. J. T. M. K.: conceptualization, validation, resources, writing – reviews & editing, supervision, project administration, funding acquisition.Conflicts of interest
There are no conflicts to declare.Data availability
The spectroscopic data presented in this article are available at Zenodo, DOI: https://doi.org/10.5281/zenodo.18457629. The supplementary data about experimental procedures have been included in the supplementary information (SI). Supplementary information includes 77-K steady-state fluorescence spectra with 480-nm and 550-nm excitation, additional quasi-TA 2DIR spectra, cross-peak fitting and experimental procedures. See DOI: https://doi.org/10.1039/d6cp00501b.Acknowledgements
This project was supported by the Dutch organization for scientific research (NWO) via the NWO Groot grant “Nanoscale regulators of photosynthesis” (OCENW.GROOT.2019.086) to R. C. and J. T. M. K.References
- A. V. Ruban, M. P. Johnson and C. D. Duffy, The photoprotective molecular switch in the photosystem II antenna, Biochim. Biophys. Acta, 2012, 1817(1), 167–181, DOI:10.1016/j.bbabio.2011.04.007.
- K. K. Niyogi and T. B. Truong, Evolution of flexible non-photochemical quenching mechanisms that regulate light harvesting in oxygenic photosynthesis, Curr. Opin. Plant Biol., 2013, 16(3), 307–314, DOI:10.1016/j.pbi.2013.03.011.
- A. V. Ruban, Light harvesting control in plants, FEBS Lett., 2018, 592(18), 3030–3039, DOI:10.1002/1873-3468.13111.
- H. van Amerongen and R. Croce, Nonphotochemical quenching in plants: Mechanisms and mysteries, Plant Cell, 2025, 37(11), koaf240, DOI:10.1093/plcell/koaf240.
- P. Horton and A. V. Ruban, Regulation of Photosystem II, Photosynth. Res., 1992, 34(3), 375–385, DOI:10.1007/BF00029812.
- L. Nicol, W. J. Nawrocki and R. Croce, Disentangling the sites of non-photochemical quenching in vascular plants, Nat. Plants, 2019, 5(11), 1177–1183, DOI:10.1038/s41477-019-0526-5.
- X. P. Li, O. Bjorkman, C. Shih, A. R. Grossman, M. Rosenquist, S. Jansson and K. K. Niyogi, A pigment-binding protein essential for regulation of photosynthetic light harvesting, Nature, 2000, 403(6768), 391–395, DOI:10.1038/35000131.
- X. P. Li, A. M. Gilmore, S. Caffarri, R. Bassi, T. Golan, D. Kramer and K. K. Niyogi, Regulation of photosynthetic light harvesting involves intrathylakoid lumen pH sensing by the PsbS protein, J. Biol. Chem., 2004, 279(22), 22866–22874, DOI:10.1074/jbc.M402461200.
- B. Demmig, K. Winter, A. Kruger and F. C. Czygan, Photoinhibition and zeaxanthin formation in intact leaves: a possible role of the xanthophyll cycle in the dissipation of excess light energy, Plant Physiol., 1987, 84(2), 218–224, DOI:10.1104/pp.84.2.218.
- M. Nilkens, E. Kress, P. Lambrev, Y. Miloslavina, M. Muller, A. R. Holzwarth and P. Jahns, Identification of a slowly inducible zeaxanthin-dependent component of non-photochemical quenching of chlorophyll fluorescence generated under steady-state conditions in Arabidopsis, Biochim. Biophys. Acta, 2010, 1797(4), 466–475, DOI:10.1016/j.bbabio.2010.01.001.
- H. Y. Yamamoto, T. O. Nakayama and C. O. Chichester, Studies on the light and dark interconversions of leaf xanthophylls, Arch. Biochem. Biophys., 1962, 97, 168–173, DOI:10.1016/0003-9861(62)90060-7.
- B. Demmig-Adams and W. W. Adams, The carotenoid zeaxanthin and ‘high-energy-state quenching’ of chlorophyll fluorescence, Photosynth. Res., 1990, 25(3), 187–197, DOI:10.1007/BF00033160.
- M. P. Johnson, T. K. Goral, C. D. Duffy, A. P. Brain, C. W. Mullineaux and A. V. Ruban, Photoprotective energy dissipation involves the reorganization of photosystem II light-harvesting complexes in the grana membranes of spinach chloroplasts, Plant Cell, 2011, 23(4), 1468–1479, DOI:10.1105/tpc.110.081646.
- L. A. I. Ramakers, J. Harbinson, E. Wientjes and H. van Amerongen, Unravelling the different components of nonphotochemical quenching using a novel analytical pipeline, New Phytol., 2025, 245(2), 625–636, DOI:10.1111/nph.20271.
- W. Marulanda Valencia and A. Pandit, Photosystem II Subunit S (PsbS): A Nano Regulator of Plant Photosynthesis, J. Mol. Biol., 2024, 436(5), 168407, DOI:10.1016/j.jmb.2023.168407.
- M. Fan, M. Li, Z. Liu, P. Cao, X. Pan, H. Zhang, X. Zhao, J. Zhang and W. Chang, Crystal structures of the PsbS protein essential for photoprotection in plants, Nat. Struct. Mol. Biol., 2015, 22(9), 729–735, DOI:10.1038/nsmb.3068.
- K. K. Niyogi, X. P. Li, V. Rosenberg and H. S. Jung, Is PsbS the site of non-photochemical quenching in photosynthesis?, J. Exp. Bot., 2005, 56(411), 375–382, DOI:10.1093/jxb/eri056.
- X. P. Li, A. Phippard, J. Pasari and K. K. Niyogi, Structure-function analysis of photosystem II subunit S (PsbS) in vivo, Funct. Plant Biol., 2002, 29(10), 1131–1139, DOI:10.1071/FP02065.
- N. Liguori, S. R. R. Campos, A. M. Baptista and R. Croce, Molecular Anatomy of Plant Photoprotective Switches: The Sensitivity of PsbS to the Environment, Residue by Residue, J. Phys. Chem. Lett., 2019, 10(8), 1737–1742, DOI:10.1021/acs.jpclett.9b00437.
- M. G. Chiariello, F. Gru, R. Zarmiento-Garcia and S. J. Marrink, pH-Dependent Conformational Switch Impacts Stability of the PsbS Dimer, J. Phys. Chem. Lett., 2023, 14(4), 905–911, DOI:10.1021/acs.jpclett.2c03760.
- M. Krishnan-Schmieden, P. E. Konold, J. T. M. Kennis and A. Pandit, The molecular pH-response mechanism of the plant light-stress sensor PsbS, Nat. Commun., 2021, 12(1), 2291, DOI:10.1038/s41467-021-22530-4.
- L. Dall'Osto, S. Cazzaniga, M. Bressan, D. Palecek, K. Zidek, K. K. Niyogi, G. R. Fleming, D. Zigmantas and R. Bassi, Two mechanisms for dissipation of excess light in monomeric and trimeric light-harvesting complexes, Nat. Plants, 2017, 3, 17033, DOI:10.1038/nplants.2017.33.
- A. R. Holzwarth, Y. Miloslavina, M. Nilkens and P. Jahns, Identification of two quenching sites active in the regulation of photosynthetic light-harvesting studied by time-resolved fluorescence, Chem. Phys. Lett., 2009, 483(4–6), 262–267, DOI:10.1016/j.cplett.2009.10.085.
- L. Wilk, M. Grunwald, P. N. Liao, P. J. Walla and W. Kühlbrandt, Direct interaction of the major light-harvesting complex II and PsbS in nonphotochemical quenching, Proc. Natl. Acad. Sci. U. S. A., 2013, 110(14), 5452–5456, DOI:10.1073/pnas.1205561110.
- V. Correa-Galvis, G. Poschmann, M. Melzer, K. Stuhler and P. Jahns, PsbS interactions involved in the activation of energy dissipation in Arabidopsis, Nat. Plants, 2016, 2, 15225, DOI:10.1038/nplants.2015.225.
- L. Nicol and R. Croce, The PsbS protein and low pH are necessary and sufficient to induce quenching in the light-harvesting complex of plants LHCII, Sci. Rep., 2021, 11(1), 7415, DOI:10.1038/s41598-021-86975-9.
- C. Ilioaia, M. P. Johnson, P. Horton and A. V. Ruban, Induction of efficient energy dissipation in the isolated light-harvesting complex of Photosystem II in the absence of protein aggregation, J. Biol. Chem., 2008, 283(43), 29505–29512, DOI:10.1074/jbc.M802438200.
- B. van Oort, R. van Grondelle and I. H. van Stokkum, A Hidden State in Light-Harvesting Complex II Revealed By Multipulse Spectroscopy, J. Phys. Chem. B, 2015, 119(16), 5184–5193, DOI:10.1021/acs.jpcb.5b01335.
- S. Wilson, D. H. Li and A. V. Ruban, The Structural and Spectral Features of Light-Harvesting Complex II Proteoliposomes Mimic Those of Native Thylakoid Membranes, J. Phys. Chem. Lett., 2022, 13(24), 5683–5691, DOI:10.1021/acs.jpclett.2c01019.
- A. V. Ruban, R. Berera, C. Ilioaia, I. H. van Stokkum, J. T. Kennis, A. A. Pascal, H. van Amerongen, B. Robert, P. Horton and R. van Grondelle, Identification of a mechanism of photoprotective energy dissipation in higher plants, Nature, 2007, 450(7169), 575–578, DOI:10.1038/nature06262.
- K. Pawlak, S. Paul, C. Liu, M. Reus, C. Yang and A. R. Holzwarth, On the PsbS-induced quenching in the plant major light-harvesting complex LHCII studied in proteoliposomes, Photosynth. Res., 2020, 144(2), 195–208, DOI:10.1007/s11120-020-00740-z.
- S. Wilson, C. D. Clarke, M. A. Carbajal, R. Buccafusca, R. A. Fleck, V. Daskalakis and A. V. Ruban, Hydrophobic Mismatch in the Thylakoid Membrane Regulates Photosynthetic Light Harvesting, J. Am. Chem. Soc., 2024, 146(21), 14905–14914, DOI:10.1021/jacs.4c05220.
- M. P. Johnson and A. V. Ruban, Restoration of rapidly reversible photoprotective energy dissipation in the absence of PsbS protein by enhanced DeltapH, J. Biol. Chem., 2011, 286(22), 19973–19981, DOI:10.1074/jbc.M111.237255.
- M. Ruan, H. Li, Y. Zhang, R. Zhao, J. Zhang, Y. Wang, J. Gao, Z. Wang, Y. Wang and D. Sun, et al., Cryo-EM structures of LHCII in photo-active and photo-protecting states reveal allosteric regulation of light harvesting and excess energy dissipation, Nat. Plants, 2023, 9(9), 1547–1557, DOI:10.1038/s41477-023-01500-2.
- Z. Ganim, H. S. Chung, A. W. Smith, L. P. Deflores, K. C. Jones and A. Tokmakoff, Amide I two-dimensional infrared spectroscopy of proteins, Acc. Chem. Res., 2008, 41(3), 432–441, DOI:10.1021/ar700188n.
- A. Krezel and W. Bal, A formula for correlating pKa values determined in D2O and H2O, J. Inorg. Biochem., 2004, 98(1), 161–166, DOI:10.1016/j.jinorgbio.2003.10.001.
- N. Fehr, C. Dietz, Y. Polyhach, T. von Hagens, G. Jeschke and H. Paulsen, Modeling of the N-terminal Section and the Lumenal Loop of Trimeric Light Harvesting Complex II (LHCII) by Using EPR, J. Biol. Chem., 2015, 290(43), 26007–26020, DOI:10.1074/jbc.M115.669804.
- Y. Miloslavina, A. Wehner, P. H. Lambrev, E. Wientjes, M. Reus, G. Garab, R. Croce and A. R. Holzwarth, Far-red fluorescence: a direct spectroscopic marker for LHCII oligomer formation in non-photochemical quenching, FEBS Lett., 2008, 582(25–26), 3625–3631, DOI:10.1016/j.febslet.2008.09.044.
- G. F. Dorlhiac, C. Fare and J. J. van Thor, PyLDM – An open source package for lifetime density analysis of time-resolved spectroscopic data, PLoS Comput. Biol., 2017, 13(5), e1005528, DOI:10.1371/journal.pcbi.1005528.
- C. Fang, A. Senes, L. Cristian, W. F. DeGrado and R. M. Hochstrasser, Amide vibrations are delocalized across the hydrophobic interface of a transmembrane helix dimer, Proc. Natl. Acad. Sci. U. S. A., 2006, 103(45), 16740–16745, DOI:10.1073/pnas.0608243103.
- N. A. Nevskaya and Y. N. Chirgadze, Infrared spectra and resonance interactions of amide-I and II vibration of alpha-helix, Biopolymers, 1976, 15(4), 637–648, DOI:10.1002/bip.1976.360150404.
- L. P. Deflores, Z. Ganim, R. A. Nicodemus and A. Tokmakoff, Amide I′-II′ 2D IR spectroscopy provides enhanced protein secondary structural sensitivity, J. Am. Chem. Soc., 2009, 131(9), 3385–3391, DOI:10.1021/ja8094922.
- H. C. Yu, D. N. Ding, Y. W. Huang, Y. X. Yuan, J. K. Song and Y. Yin, Characteristic information analysis of Raman spectrum of cucumber chlorophyll content and hardness and detection model construction, J. Food Meas. Charact., 2024, 18(5), 3492–3501, DOI:10.1007/s11694-024-02419-8.
- A. Pascal, U. Wacker, K. D. Irrgang, P. Horton, G. Renger and B. Robert, Pigment binding site properties of two photosystem II antenna proteins. A resonance Raman investigation, J. Biol. Chem., 2000, 275(29), 22031–22036, DOI:10.1074/jbc.M000658200.
- A. Barth, The infrared absorption of amino acid side chains, Prog. Biophys. Mol. Biol., 2000, 74(3–5), 141–173, DOI:10.1016/s0079-6107(00)00021-3.
- J. Breton, Fourier transform infrared spectroscopy of primary electron donors in type I photosynthetic reaction centers, Biochim. Biophys. Acta, 2001, 1507(1–3), 180–193, DOI:10.1016/s0005-2728(01)00206-7.
- Z. Liu, H. Yan, K. Wang, T. Kuang, J. Zhang, L. Gui, X. An and W. Chang, Crystal structure of spinach major light-harvesting complex at 2.72 A resolution, Nature, 2004, 428(6980), 287–292, DOI:10.1038/nature02373.
- X. Su, J. Ma, X. Wei, P. Cao, D. Zhu, W. Chang, Z. Liu, X. Zhang and M. Li, Structure and assembly mechanism of plant C(2)S(2)M(2)-type PSII-LHCII supercomplex, Science, 2017, 357(6353), 815–820, DOI:10.1126/science.aan0327.
- M. Ye, Q. L. Zhang, H. Li, Y. X. Weng, W. C. Wang and X. G. Qiu, Infrared spectroscopic discrimination between the loop and alpha-helices and determination of the loop diffusion kinetics by temperature-jump time-resolved infrared spectroscopy for cytochrome c, Biophys. J., 2007, 93(8), 2756–2766, DOI:10.1529/biophysj.107.106799.
- F. D. Fuller and J. P. Ogilvie, Experimental implementations of two-dimensional Fourier transform electronic spectroscopy, Annu. Rev. Phys. Chem., 2015, 66, 667–690, DOI:10.1146/annurev-physchem-040513-103623.
- A. Gelzinis, R. Augulis, V. Butkus, B. Robert and L. Valkunas, Two-dimensional spectroscopy for non-specialists, Biochim. Biophys. Acta, Bioenerg., 2019, 1860(4), 271–285, DOI:10.1016/j.bbabio.2018.12.006.
- P. H. Lambrev, P. Akhtar and H. S. Tan, Insights into the mechanisms and dynamics of energy transfer in plant light-harvesting complexes from two-dimensional electronic spectroscopy, Biochim. Biophys. Acta, Bioenerg., 2020, 1861(4), 148050, DOI:10.1016/j.bbabio.2019.07.005.
- M. T. Zanni, N. H. Ge, Y. S. Kim and R. M. Hochstrasser, Two-dimensional IR spectroscopy can be designed to eliminate the diagonal peaks and expose only the crosspeaks needed for structure determination, Proc. Natl. Acad. Sci. U. S. A., 2001, 98(20), 11265–11270, DOI:10.1073/pnas.201412998.
- M. Khalil, N. Demirdöven and A. Tokmakoff, Coherent 2D IR spectroscopy: Molecular structure and dynamics in solution, J. Phys. Chem. A, 2003, 107(27), 5258–5279, DOI:10.1021/jp0219247.
- E. F. Pettersen, T. D. Goddard, C. C. Huang, E. C. Meng, G. S. Couch, T. I. Croll, J. H. Morris and T. E. Ferrin, UCSF ChimeraX: Structure visualization for researchers, educators, and developers, Protein Sci., 2021, 30(1), 70–82, DOI:10.1002/pro.3943.
- R. A. Silva, S. C. Yasui, J. Kubelka, F. Formaggio, M. Crisma, C. Toniolo and T. A. Keiderling, Discriminating 3(10)- from alpha-helices: vibrational and electronic CD and IR absorption study of related Aib-containing oligopeptides, Biopolymers, 2002, 65(4), 229–243, DOI:10.1002/bip.10241.
- E. B. Dunkelberger, M. Grechko and M. T. Zanni, Transition Dipoles from 1D and 2D Infrared Spectroscopy Help Reveal the Secondary Structures of Proteins: Application to Amyloids, J. Phys. Chem. B, 2015, 119(44), 14065–14075, DOI:10.1021/acs.jpcb.5b07706.
- K. R. Webb, K. A. Hess, A. Shmidt, K. D. Segner and L. E. Buchanan, Probing local changes to alpha-helical structures with 2D IR spectroscopy and isotope labeling, Biophys. J., 2023, 122(8), 1491–1502, DOI:10.1016/j.bpj.2023.03.014.
- S. Mukherjee, P. Chowdhury and F. Gai, Infrared study of the effect of hydration on the amide I band and aggregation properties of helical peptides, J. Phys. Chem. B, 2007, 111(17), 4596–4602, DOI:10.1021/jp0689060.
- J. P. Lomont, J. S. Ostrander, J. J. Ho, M. K. Petti and M. T. Zanni, Not All beta-Sheets Are the Same: Amyloid Infrared Spectra, Transition Dipole Strengths, and Couplings Investigated by 2D IR Spectroscopy, J. Phys. Chem. B, 2017, 121(38), 8935–8945, DOI:10.1021/acs.jpcb.7b06826.
- E. Elias, N. Liguori and R. Croce, The origin of pigment-binding differences in CP29 and LHCII: the role of protein structure and dynamics, Photochem. Photobiol. Sci., 2023, 22(6), 1279–1297, DOI:10.1007/s43630-023-00368-7.
- P. Albanese, S. Tamara, G. Saracco, R. A. Scheltema and C. Pagliano, How paired PSII-LHCII supercomplexes mediate the stacking of plant thylakoid membranes unveiled by structural mass-spectrometry, Nat. Commun., 2020, 11(1), 1361, DOI:10.1038/s41467-020-15184-1.
| This journal is © the Owner Societies 2026 |