 Open Access Article
Open Access ArticleDesigning multi-site charge-bifurcation networks in de novo proteins: a kinetic, statistical, and machine-learning approach
Xiao
Huang
 a,
William F.
DeGrado
a,
William F.
DeGrado
 b,
Michael J.
Therien
b,
Michael J.
Therien
 a and
David N.
Beratan
a and
David N.
Beratan
 *acd
*acd
aDepartment of Chemistry, Duke University, Durham, NC 27708, USA. E-mail: david.beratan@duke.edu
bDepartment of Pharmaceutical Chemistry, University of California, San Francisco, San Francisco, CA 94143, USA
cDepartment of Biochemistry, Duke University, Durham, NC 27710, USA
dDepartment of Physics, Duke University, Durham, NC 27708, USA
First published on 22nd April 2026
Abstract
Electron bifurcation reactions separate electron pairs into high and low potential pools, and these reactions are central to the bioenergetics of living systems. Here, we used kinetic analysis and machine learning to analyze a diverse set of structural and electrochemical landscapes that may guide the design of molecular architectures that could serve as experimental targets that would function to bifurcate holes using light. We find that strong electrostatic repulsion between the holes enhances the quantum yield for bifurcation but reduces the energy efficiency of the process. We find that the quantum yield for hole bifurcation is enhanced by positioning the hot-hole pathway cofactor farther from the hole bifurcation site than its cold-hole pathway counterpart. This integrated design and optimization approach provides design strategies for de novo structures that could realize light-drive hole bifurcation, advancing the aim of employing bioinspired electron bifurcation for energy conversion, photocatalysis, and electrocatalysis. Beyond the specific light-driven hole-bifurcation architecture, our combined kinetic–statistical–machine-learning approach is transferable to other multi-particle, multi-site charge-transport network design challenges, opening paths for designing photochemical and catalytic networks, as well as for designing functional redox networks.
1 Introduction
Electron bifurcation (EB) reactions are poorly understood but central in bioenergetics, splitting electron pairs on a donor cofactor into two spatially separated pools at very different electrochemical potentials.1,2 EB reactions occur in photosynthesis, respiration, and biocatalysis, underpinning nitrogen fixation, carbon dioxide reduction, and hydrogen production. Cytochrome bc1, cytochrome b6f, and certain flavoproteins perform electron bifurcation reactions.1,3,4Realizing electron or hole bifurcation in a de novo protein framework5–8 represents an unrealized challenge, although some basic design principles are emerging.9,10 EB delivers electrons into redox pools at different potentials and, as such, is an open-system reaction. A challenge faced by the EB networks is to avoid electron leakage (or short circuiting) between the low and high potential pathways. Short circuiting avoidance makes the design and synthesis of electron bifurcating networks very challenging.3,4,9,11 In our earlier analysis of light-driven hole bifurcation (HB),10 we described architectures (with specific electrochemical gradients and inter-cofactor distances) that could support light-driven HB. Our earlier photoinduced HB design relied on having two nearby tryptophan residues that are oxidized sequentially by photoactivated (ruthenium) chromophores bound to the de novo protein's surface. Rapid sequential Trp oxidation would produce electrostatic repulsion between two nearby radical cation states, energizing the system. As each hole migrates along a spatially separated pathway, the first and second holes would depart the bifurcation site at two different potentials. The first hole to depart is expected to be more strongly oxidizing, or “hot” due to hole–hole repulsion, while the second hole to leave will be less oxidizing, or “cold”. These holes are directed onto spatially separated transport pathways that lead to terminal sites where they could carry out further redox chemistry. The design is shown schematically in Fig. 1 and 2.
 | ||
| Fig. 1 Schematic representation of a light-driven de novo hole bifurcating system. Two flashes, in rapid succession, oxidize nearby Trp residues using photo-generated Ru(III) species attached to the protein (via a flash-quench scheme).14 The photogenerated holes hop on “hot” and “cold” hole transport pathways. The holes arrive at the termini of the pathways. The redox cycle is completed when the initial electron quencher re-reduces these terminal species.14 This figure is reproduced from ref. 10, available under a CC-BY 4.0 license, and is copyrighted by X. Huang, P. Zhang, J. L. Yuly, W. F. DeGrado, M. J. Therien, and D. N. Beratan. | ||
 | ||
| Fig. 2 An illustrative molecular model of a designed de novo hole bifurcating protein. Two ruthenium flash-quench chromophores (FQC) are covalently bound on the protein surface nearby two Trp (W1 and W2) residues. Following W1 and W2 photooxidation, hole transfer proceeds along two spatially separated pathways: the hot-hole path (W1 → WH1 → WH2 → PMn(CF3)4, shown in the red-arrow pathway) and the cold-hole path (W2 → WL1 → WL2 → Tyr˙-HisH+, in the blue-arrow pathway). The two hole transfer routes terminate on acceptors tuned to different reduction potentials. This design demonstrates how light can initiate bifurcation in a synthetic, protein-based construct. This figure is reproduced from ref. 10, available under a CC-BY 4.0 license, and is copyrighted by X. Huang, P. Zhang, J. L. Yuly, W. F. DeGrado, M. J. Therien, and D. N. Beratan. | ||
Earlier kinetic analysis found that the distances and electrochemical potentials of the cofactors determine the quantum yields for delivering the holes to the chain termini. Even small changes to hopping site distances or electrochemical potentials can dramatically alter the network's bifurcation efficiency. Earlier studies of open system EB in the steady state focused on networks with uniform spacings between the nearest neighbor hopping sites, three (infinite) charge reservoirs, and equal magnitude slopes for the high- and low-electrochemical potential chains.9,12,13 The aim of the present study is to discover molecular designs that could produce high HB quantum yields (while maintaining acceptable energy efficiency) on closed HB networks without enforcing the simplifying constraints used in the earlier models for open system EB. This optimization focuses on the molecular architectures illustrated in Fig. 1 and 2. That is, we aimed to optimize light driven HB in de novo proteins where one high potential and one low potential electron hopping pathway direct charge from the two-hole bifurcation locus.
We combined kinetic network analysis, robust statistical analyses of large hole bifurcation designs, and machine learning models to study hole bifurcation target systems. Using Bayesian optimization of HB network quantum yield and random (unbiased) sampling of the design space, we learned how structural and energetic design parameters influence HB network quantum yields and energy efficiencies. This approach to bifurcation network design produced new insights with respect to earlier EB designs that were focused on near-reversible, open-system biological structures that operate in the steady state in contact with electron reservoirs. Our optimization of closed-system, light-driven hole bifurcation finds that high quantum yield HB can be realized with acceptable energy efficiency across a surprisingly wide range of energy and distance landscapes.
While our immediate focus is on two-hole bifurcation networks in de novo proteins, the challenge that we address, namely how to design functional kinetic networks assisted by machine-learning, is of broader interest. The design strategy that we describe can be mapped to other multi-site, multi-particle charge-transport or bifurcation networks in proteins, molecular materials, or supramolecular assemblies.
2 Methods
2.1 Computational methods and performance assessment for enumerated bifurcation networks
To develop HB design principles for the networks with the structure shown in Fig. 1, we developed a computational protocol to build, evaluate, and analyze a large set of (theoretical) light-driven HB networks with the architecture shown in Fig. 1.The rate constants (kij) for non-adiabatic hole transfer (HT) between pairs of redox active cofactors i and j were calculated using non-adiabatic electron transfer theory, accounting for one high-frequency (quantum) vibrational mode, shown in eqn (1):15,16
 | (1) |
For a given hole transfer reaction, 〈Vij2〉 is the thermally averaged electronic coupling; ΔG(0)ij is the free energy change; λij is the outer-sphere reorganization energy; D is the Huang–Rhys factor, equal to λin/(ħω), where λin is the reorganization energy of the high frequency mode; and ħω is the energy spacing for the (harmonic) high-frequency mode (see Section S1 for the estimated values for the parameters in eqn (1) and (2)).
The mean-squared electronic coupling, 〈Vij2〉, is typically approximated using an exponential decay model reflecting its sensitivity to the inter-cofactor distance Rij,17 shown in eqn (2)
| Vij = V0e−βRij | (2) |
Here, Rij is the edge-to-edge distance between the donor and acceptor heavy atoms.
The occupancy status of each cofactor within the network for any given redox microstate is represented by a vector S. For a system with N cofactors, [C1, C2, …, CN], S is formulated as eqn (3):
| S = [n1, n2, …, nN] | (3) |
The time evolving hole population on the network was simulated using a kinetic master equation based on the system microstates. The probability P(Si,t) of being in microstate Si (defined by the hole occupancy of each cofactor) at time t evolves according to eqn (4):
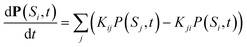 | (4) |
| P(t) = eKtP(0) | (5) |
 | (6) |
The diagonal elements of the rate matrix, kii, are the negative sums of all rates from microstate Si to all other microstates:
 | (7) |
The quantum yield (QY) for HB is defined as the probability for the system to reach the final state where a hole resides on each of the two terminal acceptors for a long time (t = 1 s). This is calculated by summing the probabilities of all microstates S where both terminal cofactors are occupied:
 | (8) |
We define the energy efficiency metric, η, as in eqn (9):
 | (9) |
E
(0)PMn in eqn (5) is the electrochemical potential of the hot-hole acceptor,  is the potential of the second tryptophan (W2) at the bifurcation site (this potential is relevant after the first hole leaves), and Erepulsion is the electrostatic repulsion energy between the two holes at their site formation (W1˙+ and W2˙+). In our earlier study, Erepulsion has been shown to be as much as 1.6 eV according to the DFT calculations, and tunable as a function of edge-to-edge distance between W1˙+ and W2˙+.10
is the potential of the second tryptophan (W2) at the bifurcation site (this potential is relevant after the first hole leaves), and Erepulsion is the electrostatic repulsion energy between the two holes at their site formation (W1˙+ and W2˙+). In our earlier study, Erepulsion has been shown to be as much as 1.6 eV according to the DFT calculations, and tunable as a function of edge-to-edge distance between W1˙+ and W2˙+.10
2.2 Synthetic HB network candidate generation and network parameters space sampling
We generate a large number of candidate bifurcation networks, as described below, and then analyze their performance. Each network candidate has a bifurcation site (with two interacting tryptophan residues, W1 and W2) and two distinct hole transport paths—designated as the low potential (hot-hole) and high potential (cold-hole) branches. Each pathway contains two cofactors (WH1, WH2 and WL1, WL2, respectively). This basic architecture is motivated by the structure of the Nfn1 bifurcating flavoprotein.18 These branches each terminate at a hole acceptor. Our earlier study used a Mn porphyrin cofactor and a Tyr-HisH+ species as terminal hole acceptors.10 The description of a candidate HB landscape requires the enumeration of both redox group positions (edge-to-edge distances between redox species) and the electrochemical potentials of each species. RH and RL are the distances between the edge of the bifurcating cofactor and the nearest high and low potential acceptor. All other nearest-neighbor edge-to-edge distances (i.e., the separations between successive cofactors along both the hot- and cold-hole pathways, excluding RH and RL) were assigned a single, common value R. This global parameter R was then varied during the optimization to explore its impact on network performance. Electrochemical potentials (E(0)i) for each cofactor, reorganization energies (λij), and the electrostatic repulsion between the two oxidized cofactors at the bifurcation site are all included in the landscape models. The midpoint potential for the terminal Tyr-His/Tyr˙-HisH+ redox couple is estimated to be approximately 1.0 V (vs. NHE)19–23 and is not varied. The electrochemical potential of the terminal MnP species is varied, as this potential could be altered by changes to the chemical composition or the protein environment.Table 1 shows the range of electron transfer rate parameters (reorganization energies, electrochemical potentials, distances) that are selected to span biologically and synthetically plausible ranges. The parameter ranges in Table 1 were selected to span the structural and energetic space accessible to de novo proteins. The redox potential ranges for tryptophan and the terminal acceptors were derived from literature reports and previous experimental measurements in various protein environments. Our workflow involved: (1) defining the plausible physiological range for each parameter, (2) performing large-scale sampling of these ranges, and (3) calculating the resulting kinetic performance for each set.
We used two complementary strategies to explore the parameter ranges for HB described in Table 1. The first 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 parameter sets were generated through uniform random sampling across the parameter ranges indicated in Table 1. This random dataset provides unbiased sampling of the parameter space. Then, in order to improve the exposure of high quantum yield design in the dataset, we performed data augmentation by generating 30
000 parameter sets were generated through uniform random sampling across the parameter ranges indicated in Table 1. This random dataset provides unbiased sampling of the parameter space. Then, in order to improve the exposure of high quantum yield design in the dataset, we performed data augmentation by generating 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 more parameter sets, which have high quantum yields (>0.9) identified via Bayesian optimization (BOp), using quantum yield for successful HB as the objective function (see Section S2 for BOp details). We generated a total of 130
000 more parameter sets, which have high quantum yields (>0.9) identified via Bayesian optimization (BOp), using quantum yield for successful HB as the objective function (see Section S2 for BOp details). We generated a total of 130![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 distinct configurations. These datasets allow us to explore crucial factors for the design of high-efficiency HB networks.
000 distinct configurations. These datasets allow us to explore crucial factors for the design of high-efficiency HB networks.
The 130![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 candidate networks represent theoretical “redox landscapes” defined by the parameters in Table 1. Each data point in the training set is a unique combination of distances and potentials, rather than a specific atomistic molecular structure. This approach allows us to discover general design principles that can subsequently be mapped onto specific amino acid sequences and residue positions.
000 candidate networks represent theoretical “redox landscapes” defined by the parameters in Table 1. Each data point in the training set is a unique combination of distances and potentials, rather than a specific atomistic molecular structure. This approach allows us to discover general design principles that can subsequently be mapped onto specific amino acid sequences and residue positions.
2.3 Statistical analysis on generated HB network candidates
We performed statistical analysis on 130![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 enumerated bifurcating networks. These networks were generated as described in Section 2.2. These networks aim to provide sufficient context to establish an understanding of how to control the quantum yield and energy efficiency of HB networks. Correlations between each network's structural and energetic parameters and its performance metrics (quantum yield and energy efficiency) were quantified using Pearson's correlation coefficients and visualized as a heatmap—a standard approach for preliminary feature screening in multivariate studies.24 This linear regression framework enabled us to determine which parameters exert statistically significant effects on quantum yield and energy efficiency, to establish whether each effect is positive or negative (i.e., its sign), and to quantify the strength (effect size) of these relationships under the assumption of linear dependence (as a first-order approximation to capture the primary trends, despite the underlying nonlinearity) between inputs and outputs (see Section S3 for full regression details).
000 enumerated bifurcating networks. These networks were generated as described in Section 2.2. These networks aim to provide sufficient context to establish an understanding of how to control the quantum yield and energy efficiency of HB networks. Correlations between each network's structural and energetic parameters and its performance metrics (quantum yield and energy efficiency) were quantified using Pearson's correlation coefficients and visualized as a heatmap—a standard approach for preliminary feature screening in multivariate studies.24 This linear regression framework enabled us to determine which parameters exert statistically significant effects on quantum yield and energy efficiency, to establish whether each effect is positive or negative (i.e., its sign), and to quantify the strength (effect size) of these relationships under the assumption of linear dependence (as a first-order approximation to capture the primary trends, despite the underlying nonlinearity) between inputs and outputs (see Section S3 for full regression details).
2.4 Machine learning assessment of design parameter influence on quantum yield
As we described in Section 2.2, we generated 130![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 HB network design candidates. We divided them with an 8
000 HB network design candidates. We divided them with an 8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 ratio for the training and testing sets. The training set input information is the set of network edge-to-edge distances, reduction potentials, reorganization energies, electrostatic repulsion energies, and the derived parameter ΔR (ΔR = RH − RL, to show the numerical relationships between RH and RL).
2 ratio for the training and testing sets. The training set input information is the set of network edge-to-edge distances, reduction potentials, reorganization energies, electrostatic repulsion energies, and the derived parameter ΔR (ΔR = RH − RL, to show the numerical relationships between RH and RL).
| Predicted high yield | Predicted low yield | |
|---|---|---|
| True high yield (>0.9) | True positive (TP) | False negative (FN) |
| True low yield (≤0.9) | False positive (FP) | True negative (TN) |
With these outcomes defined, we assessed classifier performance using four standard metrics: accuracy, precision, recall, and the F1-score.26 Accuracy measures the overall fraction of correctly classified parameter sets, shown in eqn (10):
 | (10) |
Precision quantifies the reliability of positive predictions, shown in eqn (11):
 | (11) |
 | (12) |
 | (13) |
Each of these metrics captures a distinct but complementary aspect of model performance. Accuracy provides a global measure of correct classifications, precision ensures that predicted high-yield networks are trustworthy, recall guards against missing genuinely promising designs, and the F1-score balances these competing goals. Considering all four metrics together is particularly important for HB network analysis, since the parameter space is large and highly nonlinear. In this context, false positives (low-yield networks incorrectly predicted as high yield) may waste experimental resources, whereas false negatives (high-yield networks overlooked by the model) risk discarding potentially valuable designs. By jointly evaluating all four metrics, we ensure that the classifier not only identifies high-yield candidates with confidence but also avoids systematically excluding viable configurations that could advance HB experimental realization.
Since this is a theoretical design study aiming to establish fundamental principles, the “true” labels for the ML model are based on the MJL kinetic model. To account for potential discrepancies with future experimental results, we focused on identifying “sweet-spot” ranges rather than single optimal points. These broad ranges suggest that high HB performance is robust to the typical fluctuations and errors inherent in biological redox parameters.
3 Results and discussion
To elucidate critical factors that influence the yield of hole bifurcation (HB) networks, we evaluated the relationships between network parameters (Table 1) and the bifurcation yield. Statistical analysis on the dataset of bifurcation networks, combined with feature importance assessment performed with the machine learning model XGBoost classifier, provided quantitative insight into bifurcation network design (see Section 2.2).3.1 Correlation analysis for the design parameters and the HB performance
To identify the simplest predictive relationships between network parameters and function in our HB network, we first calculated the Pearson correlation coefficients between each input parameter and the bifurcation network performance metrics: quantum yield and energy efficiency. A heatmap shows these correlations in Fig. 3. Notably, the electrostatic repulsion between the two photogenerated holes at the bifurcation site (Erepulsion) has a moderate positive correlation with quantum yield (r = 0.42), indicating that stronger inter-hole repulsion generally promotes more effective bifurcation. In contrast, Erepulsion at the bifurcation site correlates strongly and negatively with η (r = −0.66), indicating a clear trade-off. That is, designs that maximize HB yield tend to incur a larger energetic penalty and thus operate less efficiently. This finding is parallel to recent studies of quantum yield/energy capture tradeoffs in photosynthetic reactions.28We also studied how lowered geometric symmetry between the hot- and cold-hole pathways influences quantum yield (quantified by correlations between quantum yield and the distance difference ΔR = RH − RL, where RH and RL are the edge-to-edge distances from the bifurcation site to the first hot- and cold-pathway cofactors). The value of the parameter ΔR has a robust positive correlation with the quantum yield (r = 0.46), suggesting that a network with a hot-pathway cofactor more distant from the bifurcation site than the cold-pathway cofactor produces higher quantum efficiencies for HB reaction. Taken together, these linear correlations provide two important design insights: (1) sufficiently large electrostatic repulsion is needed to drive high yield hole bifurcation (though this comes with a trade-off, as high repulsion simultaneously lowers the energy efficiency), and (2) maintaining a larger ΔR stabilizes the preferential routing of the hot hole onto the hot-pathway (rather than short-circuiting onto the cold-pathway), thereby enhancing the overall hole bifurcation yield.
3.2 Machine learning-based feature importance for quantum yield
We trained an XGBoost classifier to distinguish between high (quantum yield >0.9) and low (quantum yield ≤0.9) yielding networks to capture the complex, nonlinear effects of design parameters on the performance of the HB networks. Our input features during the training included the full set of geometric and energetic parameters for the 104![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 HB network structures in the training set (e.g., inter-cofactor distances, reduction potentials, electrostatic repulsion, reorganization energy). The classifier achieved 96.2% accuracy (along with similar high score on both precision and recall) on a held-out test set (i.e., the subset of data not used during model training, reserved for independent performance evaluation) (see Table 3 for result details), demonstrating that this model is adequate to predict HB network quantum yields. We then applied SHAP analysis to this model in order to quantify the individual and combined influences of each feature on the predicted quantum yield.
000 HB network structures in the training set (e.g., inter-cofactor distances, reduction potentials, electrostatic repulsion, reorganization energy). The classifier achieved 96.2% accuracy (along with similar high score on both precision and recall) on a held-out test set (i.e., the subset of data not used during model training, reserved for independent performance evaluation) (see Table 3 for result details), demonstrating that this model is adequate to predict HB network quantum yields. We then applied SHAP analysis to this model in order to quantify the individual and combined influences of each feature on the predicted quantum yield.
| Class | Precision | Recall | F1-score | Support |
|---|---|---|---|---|
| High yield (>0.9) | 0.91 | 0.91 | 0.91 | 5520 |
| Low yield (≤0.9) | 0.98 | 0.97 | 0.98 | 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 480 480 |
| Accuracy | 0.96 | 26![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
||
| Macro Avg. | 0.94 | 0.94 | 0.94 | 26![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
| Weighted Avg. | 0.96 | 0.96 | 0.96 | 26![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
SHAP analysis assesses the contribution of each input parameter (i.e., inter-cofactor distances, reduction potentials, electrostatic repulsion, reorganization energy) to the XGBoost classifier's prediction, which distinguishes between networks yielding high (>0.9) versus low (≤0.9) quantum yield (Fig. 4). The summary plot in Fig. 4a shows the SHAP value for each parameter across all 130![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 generated HB network design candidate data samples. On the horizontal axis, a negative SHAP value indicates that a given parameter value shifts the model's output toward predicting a high-yield configuration, while a positive SHAP value favors a low-yield prediction. The color of each point encodes the parameter's magnitude (red = high, blue = low). For example, higher values of Erepulsion, RH, and ΔR consistently produce negative SHAP values, confirming that strong hole–hole repulsion and a larger RH–RL gap are key drivers of high quantum yield. Interestingly, larger total pathway distances R also yield negative SHAP values in this classifier—trained specifically to distinguish configurations with quantum yields above 0.9—suggesting that within the subset of already high-performing designs, shorter individual step distances can be associated with slight variations in yield that are compensated by other optimized parameters.
000 generated HB network design candidate data samples. On the horizontal axis, a negative SHAP value indicates that a given parameter value shifts the model's output toward predicting a high-yield configuration, while a positive SHAP value favors a low-yield prediction. The color of each point encodes the parameter's magnitude (red = high, blue = low). For example, higher values of Erepulsion, RH, and ΔR consistently produce negative SHAP values, confirming that strong hole–hole repulsion and a larger RH–RL gap are key drivers of high quantum yield. Interestingly, larger total pathway distances R also yield negative SHAP values in this classifier—trained specifically to distinguish configurations with quantum yields above 0.9—suggesting that within the subset of already high-performing designs, shorter individual step distances can be associated with slight variations in yield that are compensated by other optimized parameters.
We ranked the network parameters by their overall influence on quantum yield, quantified as the mean absolute SHAP value across all samples, which provides a global measure of feature importance in predicting high quantum yield (Fig. 4b). In line with the Pearson correlation results, R, Erepulsion, and RH emerge as the most critical determinants of high quantum yield. This ranking informs experimental design. For example, de novo protein scaffolds can be engineered systematically to fine-tune these three parameters, focusing first on modulating R and RH by residue positioning. Changing the amino acid composition or geometry near the bifurcation site can be used to tune Erepulsion, either by changing the local dielectric environment, introducing charged or polar residues, or altering the proximity or orientation of the bifurcating tryptophans.
When we just examine cases that produced quantum yields >0.9, we find broad “sweet-spot” ranges for these high quantum yields (i.e., yields in the 20th–80th percentile of the candidates within the high-yield subset): R between 5.4 and 8.8 Å, Erepulsion between 1.25 and 1.71 eV, and RH between 11.2 and 14.5 Å. These parameter windows, which are remarkably wide, suggest that high-performing HB networks are robust to modest changes in structure and energetics, easing the practical design of the targeted de novo proteins.
While the identified “sweet-spot” for Erepulsion (1.25–1.71 eV) is high, such values are physically realizable within the specialized environments of de novo protein cores. The highest value of the sweet-spot has been verified in our previous work.10 By engineering the scaffold to maintain a low local dielectric constant (ε ≈ 2–4) and positioning the bifurcating tryptophans within a hydrophobic pocket shielded from the aqueous interface, the electrostatic repulsion between photogenerated radical cations can be significantly enhanced. Furthermore, our SHAP analysis indicates that the high-yield classification is robust to modest fluctuations in reorganization energy, suggesting that these designs can remain functional despite the dynamic nature of the protein matrix.
To implement these findings, designers can tune Erepulsion by modifying the local dielectric environment—for example, by replacing surface-exposed residues with hydrophobic ones to shield the bifurcation site. Distances like RH and R can be precisely controlled by selecting specific i, i + 7 or i, i + 11 positions on an α-helix to position the cofactors. This provides a concrete roadmap for converting the identified ‘sweet spots’ into experimental protein sequences.
3.3 Visualizing performance trends for the most impactful design parameters
HB network performance depends non-linearly on the key system parameters. We performed kinetic simulations that vary individual parameters while holding others fixed to representative values (see the caption of Fig. 5 for details). Fig. 5 shows the sensitivity of the bifurcation quantum yield and energy efficiency to: (1) the distance between the bifurcation center and the hot-hole pathway (RH) and (2) the electrostatic repulsion energy between the holes at the bifurcation site. | ||
Fig. 5 Simulated quantum yield dependence on key HB network parameters. Both plots utilize representative constant values for several parameters, based on optimized or typical values from our previous work:10RL = 5 Å, inter-cofactor path distance = 5 Å, λ = 1.0 eV,  , E(0)PMn = 1.6 V (and other intermediate potentials as specified in the simulation code). For plot (a), repulsion is fixed at 1.6 eV; for plot (b), RH is fixed at 10 Å. (a) Quantum yield initially increases as RH increases from 5 Å. This improves the condition RH > RL, kinetically favoring the initial transfer of the higher-energy (hot) hole towards the hot pathway (WH1) over the cold pathway (WL1), thereby promoting successful bifurcation. The yield peaks within an optimal range (9–14 Å), consistent with the “sweet spot” identified by SHAP analysis. As RH increases further (>14 Å), the yield drops significantly because the rate of the first crucial hole transfer step , E(0)PMn = 1.6 V (and other intermediate potentials as specified in the simulation code). For plot (a), repulsion is fixed at 1.6 eV; for plot (b), RH is fixed at 10 Å. (a) Quantum yield initially increases as RH increases from 5 Å. This improves the condition RH > RL, kinetically favoring the initial transfer of the higher-energy (hot) hole towards the hot pathway (WH1) over the cold pathway (WL1), thereby promoting successful bifurcation. The yield peaks within an optimal range (9–14 Å), consistent with the “sweet spot” identified by SHAP analysis. As RH increases further (>14 Å), the yield drops significantly because the rate of the first crucial hole transfer step  decreases exponentially with distance due to the decay of the electronic coupling term 〈Vij2〉 in the MJL rate expression (eqn (1)), becoming too slow to compete effectively with potential recombination or undesired pathways within the simulation time. (b) Quantum yield shows a threshold dependence on the electrostatic repulsion energy. Below 0.8 eV, the yield is negligible, indicating insufficient driving force for efficient bifurcation. The yield rises sharply between 0.8 eV and 1.3 eV as repulsion provides the critical energy to drive the hot hole transfer, plateauing near unity for repulsion >1.3 eV. Concurrently, the energy efficiency (orange curve), calculated viaeqn (9), decreases monotonically with increasing repulsion. Plotting both metrics visually demonstrates the crucial design trade-off: maximizing yield requires high repulsion (>1.3 eV), while maximizing energy efficiency favors low repulsion. The optimal range for both high quantum yield and acceptable energy efficiency, therefore, lies in a compromise region, consistent with the SHAP analysis sweet spot (1.25–1.71 eV), where yield is high but the energetic cost of repulsion is not excessive. (a) Quantum yield dependence on RH. Besides RH, which is varied from 5 Å to 20 Å, all other parameters are held constant at the representative values listed in the main caption. (b) Quantum yield and energy efficiency dependence on repulsion energy. Besides the repulsion energy, which is varied from 0.5 eV to 1.6 eV, all other parameters are held constant at the representative values listed in the main caption. decreases exponentially with distance due to the decay of the electronic coupling term 〈Vij2〉 in the MJL rate expression (eqn (1)), becoming too slow to compete effectively with potential recombination or undesired pathways within the simulation time. (b) Quantum yield shows a threshold dependence on the electrostatic repulsion energy. Below 0.8 eV, the yield is negligible, indicating insufficient driving force for efficient bifurcation. The yield rises sharply between 0.8 eV and 1.3 eV as repulsion provides the critical energy to drive the hot hole transfer, plateauing near unity for repulsion >1.3 eV. Concurrently, the energy efficiency (orange curve), calculated viaeqn (9), decreases monotonically with increasing repulsion. Plotting both metrics visually demonstrates the crucial design trade-off: maximizing yield requires high repulsion (>1.3 eV), while maximizing energy efficiency favors low repulsion. The optimal range for both high quantum yield and acceptable energy efficiency, therefore, lies in a compromise region, consistent with the SHAP analysis sweet spot (1.25–1.71 eV), where yield is high but the energetic cost of repulsion is not excessive. (a) Quantum yield dependence on RH. Besides RH, which is varied from 5 Å to 20 Å, all other parameters are held constant at the representative values listed in the main caption. (b) Quantum yield and energy efficiency dependence on repulsion energy. Besides the repulsion energy, which is varied from 0.5 eV to 1.6 eV, all other parameters are held constant at the representative values listed in the main caption. | ||
The quantum yield shows a distinctive optimal range with respect to the distance RH (see Fig. 5a). Increasing RH (while keeping RL short) enhances the yield by directing the first (hot) hole to its high-potential pathway, consistent with the positive correlation found for quantum yield versus ΔR = RH − RL. However, beyond an optimal distance (14 Å in this simulation), the HB yield drops sharply. This decline is attributed to the exponential decay of the hole transfer rate with distance, making the second step hole transfer from W2 to WL1 too slow for productive HB. The peak performance found in this simulation aligns well with the optimal range of RH values identified in the broader SHAP analysis.
The critical role of electrostatic repulsion at the bifurcating site is indicated in Fig. 5b. The quantum yield (blue curve) shows a clear threshold behavior as a function of Erepulsion, rising rapidly when the repulsion between the holes is in the ∼0.8–1.1 eV window. This finding indicates that a sufficient repulsion strength is needed to provide the thermodynamic driving force required to initiate effective HB. Once hole–hole repulsion is sufficiently large (1.3 eV),10 the HB quantum yield approaches unity. The energy efficiency (orange curve) defined by eqn (9), however, shows the opposite trend. That is, the energy efficiency decreases monotonically as the hole–hole repulsion increases (that is, much of the hole–hole repulsion energy is dissipated). Fig. 5b shows key trade-offs in designing HB systems: maximizing quantum yield necessitates a high driving force to deliver the first hole, but growing the driving force for HB comes at the cost of overall energy efficiency. The repulsion energy optimal range for quantum yield identified by SHAP (1.25–1.71 eV) represents a balance among achieving a high quantum yield and an acceptable energy efficiency; near unit quantum yield can only be realized with an energy efficiency of 30–40%. If we require the design to realize very high energy efficiency, the quantum yield for EB would be very small.
The design examples in Fig. 5 underscore the non-linear relationships involved in designing high-performance HB networks, reinforcing the value of comprehensive statistical analysis and machine learning in network design.
3.4 Summary
Our analysis finds three key design principles that are required to realize efficient light-driven hole bifurcation in the design frameworks that we have explored. First, positioning the first hot-hole acceptor further from the bifurcation site than the first cold-hole pathway acceptor (RH > RL) consistently enhances the HB quantum yield. This relationship is supported robustly by both the Pearson correlation and SHAP-based feature importance analysis. Second, while increasing electrostatic repulsion between the two injected holes markedly increases the quantum yield (by providing the driving force needed to accelerate hole transfer), increasing this repulsion energy concomitantly lowers the HB energy efficiency. This finding highlights a central trade-off in the design of closed-system hole or electron transfer.28 Third, our machine learning models pinpoint “sweet-spots” in design space for each critical network parameter, capturing the nonlinear influence of inter-cofactor distances, redox potentials, and electrostatic interactions on quantum yield performance, and thereby offering concrete targets for experimental realizations. Taken together, these findings show that integrating detailed kinetic modeling with statistical and machine learning can direct the design of optimal HB networks by directing us to regions of design space that are not obvious choices based on intuition grounded, largely, in our understanding of single-step redox processes.4 Conclusion
We have explored and characterized the key determinants of efficient hole bifurcation networks. The designs emerged through integrated computational, statistical, and machine learning analysis. Our findings indicate that optimizing HB performance requires thoughtful design of both network geometry and energetics. Positioning the cofactor nearest to the bifurcating site on the hot-hole pathway at longer distances from the bifurcation site compared to that of the cold-hole counterpart (RH > RL) substantially increases the quantum yields, because the larger RH value suppresses short-circuiting of the hot hole onto the cold pathway, enforcing preferential routing of each hole to its intended branch. Installing strong electrostatic repulsion between the two holes on the bifurcation site is crucial for realizing high quantum yields, but negatively impacts the network energy efficiency. We also identified key parameter “islands” that produce high quantum yields and acceptable energy efficiencies. Specific ranges for important parameters, such as RH (11.24–14.5 Å) and electrostatic repulsion (1.25–1.71 eV) balance quantum yield (90% to 100%) and energy efficiency (30% to 40%). Our findings underscore the promise of combining computational modeling with advanced statistical and machine learning methodologies to reveal design principles for electron and hole bifurcation networks.The strategy that we described here (large-scale network enumeration → kinetic simulation → correlation/statistical screening → interpretable ML feature-analysis) constitutes a general design framework for engineered charge-transport networks. While our study focused on light-driven hole bifurcation in de novo proteins, the approach can be extended to design other complex multi-site networks. Networks of this kind might also be used for complex materials assembly or chemical catalysis, enabling the design of functional far-from-equilibrium systems.
Conflicts of interest
There are no conflicts to declare.Data availability
All data that support the findings of this study are available in the manuscript and the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d6cp00473c.Acknowledgements
This work was supported by the W. M. Keck Foundation and the National Science Foundation (CHE-2528383 and CHE-2528384). We also acknowledge support of the Duke University Shared Cluster Resource (DSCR).References
- J. W. Peters, et al., A new era for electron bifurcation, Curr. Opin. Chem. Biol., 2018, 47, 32–38 CrossRef CAS PubMed.
- W. Buckel and R. K. Thauer, Flavin-based electron bifurcation, a new mechanism of biological energy coupling, Chem. Rev., 2018, 118, 3862–3886 CrossRef CAS PubMed.
- J. L. Yuly, C. E. Lubner, P. Zhang, D. N. Beratan and J. W. Peters, Electron bifurcation: progress and grand challenges, Chem. Commun., 2019, 55, 11823–11832 RSC.
- J. L. Yuly, P. Zhang and D. N. Beratan, Energy transduction by reversible electron bifurcation, Curr. Opin. Electrochem., 2021, 29, 100767 CrossRef CAS.
- N. F. Polizzi and W. F. DeGrado, A defined structural unit enables de novo design of small-molecule-binding proteins, Science, 2020, 369, 1227–1233 CrossRef CAS PubMed.
- N. F. Polizzi, Y. Wu, T. Lemmin, A. M. Maxwell, S.-Q. Zhang, J. Rawson, D. N. Beratan, M. J. Therien and W. F. DeGrado, De novo design of a hyperstable non-natural protein-ligand complex with sub-Å accuracy, Nat. Chem., 2017, 9, 1157–1164 CrossRef CAS PubMed.
- S. I. Mann, A. Nayak, G. T. Gassner, M. J. Therien and W. F. DeGrado, De novo design, solution characterization, and crystallographic structure of an abiological Mn-porphyrin-binding protein capable of stabilizing a Mn (V) species, J. Am. Chem. Soc., 2020, 143, 252–259 CrossRef PubMed.
- L. Lu, X. Gou, S. K. Tan, S. I. Mann, H. Yang, X. Zhong, D. Gazgalis, J. Valdiviezo, H. Jo, Y. Wu, M. E. Diolaiti, A. Ashworth, N. F. Polizzi and W. F. DeGrado, De novo design of drug-binding proteins with predictable binding energy and specificity, Science, 2024, 384, 106–112 CrossRef CAS PubMed.
- J. L. Yuly, P. Zhang, C. E. Lubner, J. W. Peters and D. N. Beratan, Universal free-energy landscape produces efficient and reversible electron bifurcation, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 21045–21051 CrossRef CAS PubMed.
- X. Huang, J. Yuly, P. Zhang, W. DeGrado, M. Therien and D. Beratan, Design of light driven hole bifurcating proteins, ACS Cent. Sci., 2025, 11, 1911–1920 CrossRef CAS PubMed.
- C. E. Lubner, et al., Mechanistic insights into energy conservation by flavin-based electron bifurcation, Nat. Chem. Biol., 2017, 13, 655–659 CrossRef CAS PubMed.
- J. L. Yuly, P. Zhang, X. Ru, K. Terai, N. Singh and D. N. Beratan, Efficient and reversible electron bifurcation with either normal or inverted potentials at the bifurcating cofactor, Chem, 2021, 7, 1870–1886 CAS.
- K. Terai, J. L. Yuly, P. Zhang and D. N. Beratan, Correlated particle transport enables biological free energy transduction, Biophys. J., 2023, 122, 1762–1771 CrossRef CAS PubMed.
- H. B. Gray and J. R. Winkler, Electron tunneling through proteins, Q. Rev. Biophys., 2003, 36, 341–372 CrossRef CAS PubMed.
- V. G. Levich and R. R. Dogonadze, Theory of nonradiative electron transitions between ions in solution, Dokl. Akad. Nauk SSSR, 1959, 124, 123–126 CAS.
- J. Jortner, Temperature dependent activation energy for electron transfer between biological molecules, J. Chem. Phys., 1976, 64, 4860–4867 CrossRef CAS.
- J. J. Hopfield, Electron transfer between biological molecules by thermally activated tunneling, Proc. Natl. Acad. Sci. U. S. A., 1974, 71, 3640–3644 CrossRef CAS PubMed.
- C. E. Wise, A. E. Ledinina, D. W. Mulder, K. J. Chou, J. W. Peters, P. W. King and C. E. Lubner, An uncharacteristically low-potential flavin governs the energy landscape of electron bifurcation, Proc. Natl. Acad. Sci. U. S. A., 2022, 119, e2117882119 CrossRef CAS PubMed.
- H. Kless and W. Vermaas, Combinatorial mutagenesis and structural simulations in the environment of the redox-active tyrosine YZ of photosystem II, Biochemistry, 1996, 35, 16458–16464 CrossRef CAS PubMed.
- A.-M. A. Hays, I. R. Vassiliev, J. H. Golbeck and R. J. Debus, Role of D1-His190 in the proton-coupled oxidation of tyrosine YZ in manganese-depleted photosystem II, Biochemistry, 1999, 38, 11851–11865 CrossRef CAS PubMed.
- S. Styring, J. Sjöholm and F. Mamedov, Two tyrosines that changed the world: Interfacing the oxidizing power of photochemistry to water splitting in photosystem II, Biochim. Biophys. Acta, Bioenerg., 2012, 1817, 76–87 CrossRef CAS PubMed.
- T. J. Meyer, M. H. V. Huynh and H. H. Thorp, The possible role of proton-coupled electron transfer (PCET) in water oxidation by photosystem II, Angew. Chem., Int. Ed., 2007, 46, 5284–5304 CrossRef CAS PubMed.
- S. J. Mora, E. Odella, G. F. Moore, D. Gust, T. A. Moore and A. L. Moore, Proton-coupled electron transfer in artificial photosynthetic systems, Acc. Chem. Res., 2018, 51, 445–453 CrossRef CAS PubMed.
- G. James; D. Witten; T. Hastie and R. Tibshirani, An introduction to statistical learning: with applications in R, Springer, 2013, vol. 103 Search PubMed.
- T. Chen and C. Guestrin, XGBoost: A scalable tree boosting system, Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, 2016, pp. 785–794.
- C. Goutte and E. Gaussier, A probabilistic interpretation of precision, recall and F-score, with implication for evaluation, European conference on information retrieval, 2005, pp. 345–359.
- S. M. Lundberg and S.-I. Lee, A unified approach to interpreting model predictions, NIPS'17: Proceedings of the 31st International Conference on Neural Information Processing Systems, 2017, pp. 4768–4777 Search PubMed.
- J. D. Schultz, K. A. Parker, M. J. Therien and D. N. Beratan, Efficiency limits of energy conversion by light-Driven redox chains, J. Am. Chem. Soc., 2024, 146, 32805–32815 CrossRef CAS PubMed.
| This journal is © the Owner Societies 2026 |












